Stem Cells in Cardiovascular Medicine: Historical Overview and Future Prospects
Abstract
1. Introduction
2. Adult Stem Cells
2.1. Skeletal Myoblasts
2.2. Bone-Marrow-Derived SCs
2.3. Cardiac Progenitor Cells and Stem Cell Niches
3. Pluripotent Stem Cells
3.1. Embryonic Stem Cells
3.2. Induced Pluripotent Stem Cells
3.3. Embryonic Stem Cells Versus Induced Pluripotent Stem Cells
4. Cardiac Stem-Ness
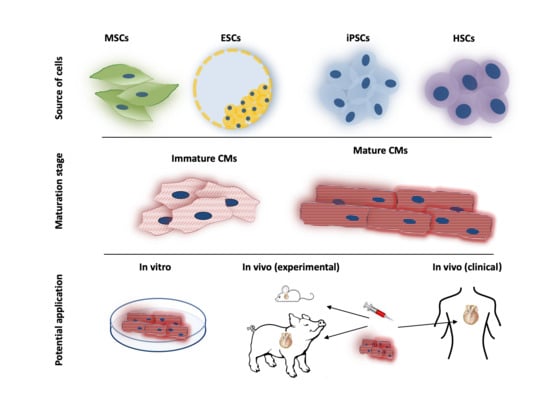
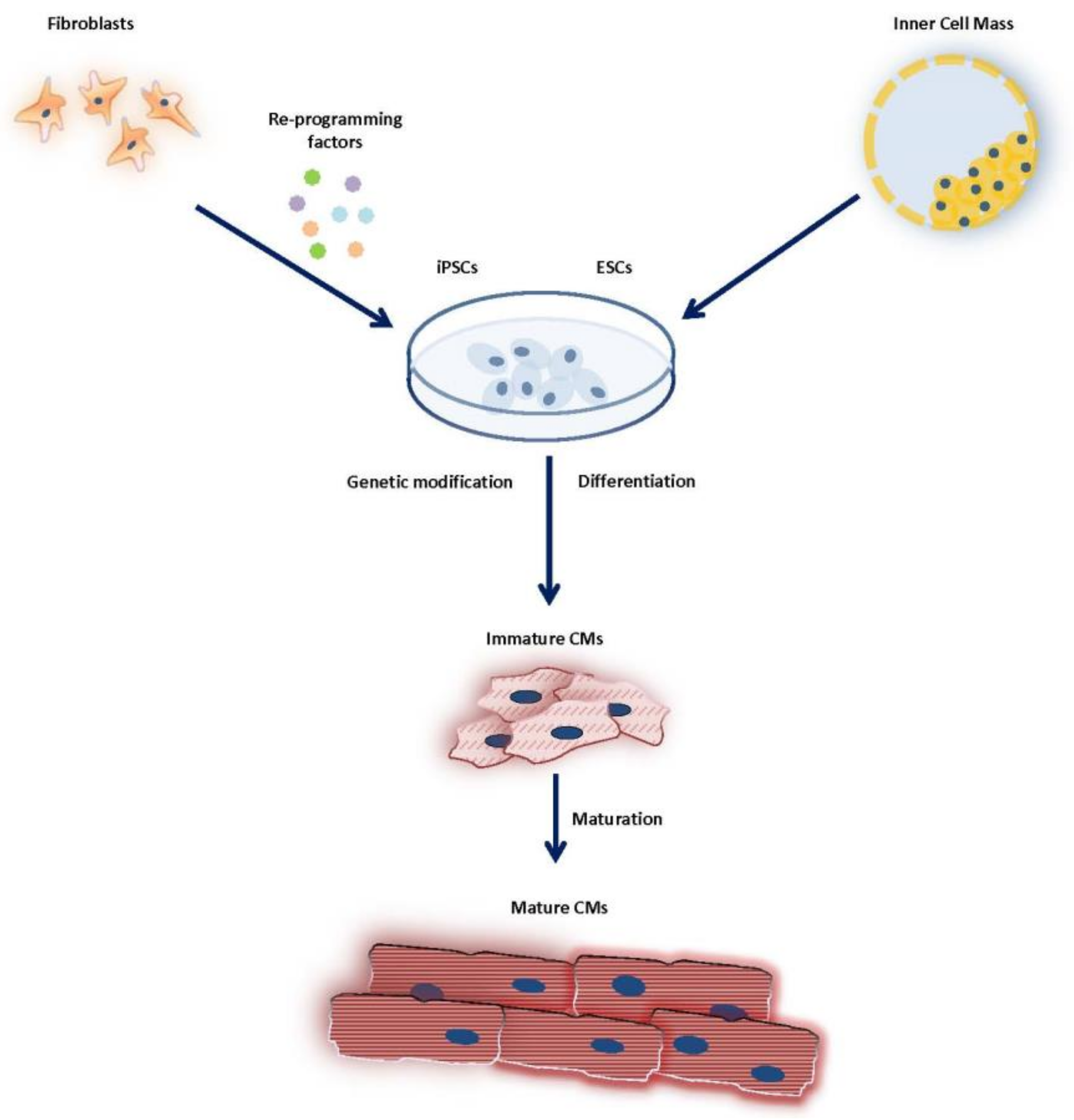
4.1. Generation/Differentiation of Pluripotent Stem Cells
4.2. Maturation of Pluripotent Stem Cells
4.3. Engineered Heart Tissue
5. Applications of PSCs in Cardiovascular Research
5.1. Pluripotent Stem Cells in Cardiovascular Disease Modeling
5.2. Pluripotent Stem Cells in Pharmaceutical Screenings
5.3. Genetic Modification of Pluripotent Stem Cells
6. Translational Potential of PSCs in Cardiovascular Regenerative Therapy
6.1. Pluripotent Stem Cells in Rodent Models
6.2. Pluripotent Stem Cells in Large-Animal Models
6.2.1. Porcine Models
6.2.2. Non-Human Primate Models
7. Pluripotent Stem Cells in First Human Trials
8. Conclusions and Remarks
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CM | cardiomyocytes |
| CPC | cardiac progenitor cells |
| CVD | cardiovascular disease |
| ESC | embryonic stem cells |
| iPSC | induced pluripotent stem cells |
| LVEF | left ventricular ejection fraction |
| MSC | mesenchymal stem cells |
| PSC | pluripotent stem cells |
| SM | skeletal myoblast |
References
- WHO. Cardiovascular Diseases (CVDs). Available online: http://www.who.int/en/news-room/fact-sheets/detail/cardiovascular-diseases- (accessed on 1 June 2019).
- Benjamin, E.J.; Blaha, M.J.; Chiuve, S.E.; Cushman, M.; Das, S.R.; Deo, R.; de Ferranti, S.D.; Floyd, J.; Fornage, M.; Gillespie, C.; et al. Heart Disease and Stroke Statistics-2017 Update: A Report from the American Heart Association. Circulation 2017, 135, e146–e603. [Google Scholar] [CrossRef] [PubMed]
- Samak, M.; Fatullayev, J.; Sabashnikov, A.; Zeriouh, M.; Schmack, B.; Ruhparwar, A.; Karck, M.; Popov, A.F.; Dohmen, P.M.; Weymann, A. Total Arterial Revascularization: Bypassing Antiquated Notions to Better Alternatives for Coronary Artery Disease. Med. Sci. Monit. Basic Res. 2016, 22, 107–114. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fatullayev, J.; Samak, M.; Sabashnikov, A.; Zeriouh, M.; Rahmanian, P.B.; Choi, Y.H.; Schmack, B.; Kallenbach, K.; Ruhparwar, A.; Eghbalzadeh, K.; et al. Continuous-Flow Left Ventricular Assist Device Thrombosis: A Danger Foreseen is a Danger Avoided. Med. Sci. Monit. Basic Res. 2015, 21, 141–144. [Google Scholar] [CrossRef] [PubMed]
- Roger, V.L. Epidemiology of heart failure. Circ. Res. 2013, 113, 646–659. [Google Scholar] [CrossRef]
- Gass, A.L.; Emaminia, A.; Lanier, G.; Aggarwal, C.; Brown, K.A.; Raffa, M.; Kai, M.; Spielvogel, D.; Malekan, R.; Tang, G.; et al. Cardiac Transplantation in the New Era. Cardiol. Rev. 2015, 23, 182–188. [Google Scholar] [CrossRef]
- Gouadon, E.; Moore-Morris, T.; Smit, N.W.; Chatenoud, L.; Coronel, R.; Harding, S.E.; Jourdon, P.; Lambert, V.; Rucker-Martin, C.; Puceat, M. Concise Review: Pluripotent Stem Cell-Derived Cardiac Cells, A Promising Cell Source for Therapy of Heart Failure: Where Do We Stand? Stem Cells 2016, 34, 34–43. [Google Scholar] [CrossRef]
- Sampogna, G.; Guraya, S.Y.; Forgione, A. Regenerative medicine: Historical roots and potential strategies in modern medicine. J. Microsc. Ultrastruct. 2015, 3, 101–107. [Google Scholar] [CrossRef]
- Muller, P.; Lemcke, H.; David, R. Stem Cell Therapy in Heart Diseases-Cell Types, Mechanisms and Improvement Strategies. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2018, 48, 2607–2655. [Google Scholar] [CrossRef]
- Viola, J.L.B.; Grad, O. The Emergence of Tissue Engineering as a Research Field; National Science Foundation: Alexandria, VA, USA, 14 October 2003. [Google Scholar]
- Tajbakhsh, S. Stem cells to tissue: Molecular, cellular and anatomical heterogeneity in skeletal muscle. Curr. Opin. Genet. Dev. 2003, 13, 413–422. [Google Scholar] [CrossRef]
- Al Attar, N.; Carrion, C.; Ghostine, S.; Garcin, I.; Vilquin, J.T.; Hagege, A.A.; Menasche, P. Long-term (1 year) functional and histological results of autologous skeletal muscle cells transplantation in rat. Cardiovasc. Res. 2003, 58, 142–148. [Google Scholar] [CrossRef]
- Suzuki, K.; Murtuza, B.; Suzuki, N.; Smolenski, R.T.; Yacoub, M.H. Intracoronary infusion of skeletal myoblasts improves cardiac function in doxorubicin-induced heart failure. Circulation 2001, 104 (Suppl. 1), I213–I217. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pouly, J.; Hagege, A.A.; Vilquin, J.T.; Bissery, A.; Rouche, A.; Bruneval, P.; Duboc, D.; Desnos, M.; Fiszman, M.; Fromes, Y.; et al. Does the functional efficacy of skeletal myoblast transplantation extend to nonischemic cardiomyopathy? Circulation 2004, 110, 1626–1631. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gavira, J.J.; Perez-Ilzarbe, M.; Abizanda, G.; Garcia-Rodriguez, A.; Orbe, J.; Paramo, J.A.; Belzunce, M.; Rabago, G.; Barba, J.; Herreros, J.; et al. A comparison between percutaneous and surgical transplantation of autologous skeletal myoblasts in a swine model of chronic myocardial infarction. Cardiovasc. Res. 2006, 71, 744–753. [Google Scholar] [CrossRef] [PubMed][Green Version]
- He, K.L.; Yi, G.H.; Sherman, W.; Zhou, H.; Zhang, G.P.; Gu, A.; Kao, R.; Haimes, H.B.; Harvey, J.; Roos, E.; et al. Autologous skeletal myoblast transplantation improved hemodynamics and left ventricular function in chronic heart failure dogs. J. Heart Lung Transplant. 2005, 24, 1940–1949. [Google Scholar] [CrossRef] [PubMed]
- Gavira, J.J.; Nasarre, E.; Abizanda, G.; Perez-Ilzarbe, M.; de Martino-Rodriguez, A.; Garcia de Jalon, J.A.; Mazo, M.; Macias, A.; Garcia-Bolao, I.; Pelacho, B.; et al. Repeated implantation of skeletal myoblast in a swine model of chronic myocardial infarction. Eur. Heart J. 2010, 31, 1013–1021. [Google Scholar] [CrossRef]
- Menasche, P.; Hagege, A.A.; Vilquin, J.T.; Desnos, M.; Abergel, E.; Pouzet, B.; Bel, A.; Sarateanu, S.; Scorsin, M.; Schwartz, K.; et al. Autologous skeletal myoblast transplantation for severe postinfarction left ventricular dysfunction. J. Am. Coll. Cardiol. 2003, 41, 1078–1083. [Google Scholar] [CrossRef]
- Herreros, J.; Prosper, F.; Perez, A.; Gavira, J.J.; Garcia-Velloso, M.J.; Barba, J.; Sanchez, P.L.; Canizo, C.; Rabago, G.; Marti-Climent, J.M.; et al. Autologous intramyocardial injection of cultured skeletal muscle-derived stem cells in patients with non-acute myocardial infarction. Eur. Heart J. 2003, 24, 2012–2020. [Google Scholar] [CrossRef]
- Siminiak, T.; Fiszer, D.; Jerzykowska, O.; Grygielska, B.; Rozwadowska, N.; Kalmucki, P.; Kurpisz, M. Percutaneous trans-coronary-venous transplantation of autologous skeletal myoblasts in the treatment of post-infarction myocardial contractility impairment: The POZNAN trial. Eur. Heart J. 2005, 26, 1188–1195. [Google Scholar] [CrossRef]
- Smits, P.C.; van Geuns, R.J.; Poldermans, D.; Bountioukos, M.; Onderwater, E.E.; Lee, C.H.; Maat, A.P.; Serruys, P.W. Catheter-based intramyocardial injection of autologous skeletal myoblasts as a primary treatment of ischemic heart failure: Clinical experience with six-month follow-up. J. Am. Coll. Cardiol. 2003, 42, 2063–2069. [Google Scholar] [CrossRef]
- Hagege, A.A.; Marolleau, J.P.; Vilquin, J.T.; Alheritiere, A.; Peyrard, S.; Duboc, D.; Abergel, E.; Messas, E.; Mousseaux, E.; Schwartz, K.; et al. Skeletal myoblast transplantation in ischemic heart failure: Long-term follow-up of the first phase I cohort of patients. Circulation 2006, 114, I108–I113. [Google Scholar] [CrossRef]
- Ince, H.; Petzsch, M.; Rehders, T.C.; Chatterjee, T.; Nienaber, C.A. Transcatheter transplantation of autologous skeletal myoblasts in postinfarction patients with severe left ventricular dysfunction. J. Endovasc. Ther. 2004, 11, 695–704. [Google Scholar] [CrossRef]
- Veltman, C.E.; Soliman, O.I.; Geleijnse, M.L.; Vletter, W.B.; Smits, P.C.; ten Cate, F.J.; Jordaens, L.J.; Balk, A.H.; Serruys, P.W.; Boersma, E.; et al. Four-year follow-up of treatment with intramyocardial skeletal myoblasts injection in patients with ischaemic cardiomyopathy. Eur. Heart J. 2008, 29, 1386–1396. [Google Scholar] [CrossRef] [PubMed]
- Reinecke, H.; Poppa, V.; Murry, C.E. Skeletal muscle stem cells do not transdifferentiate into cardiomyocytes after cardiac grafting. J. Mol. Cell. Cardiol. 2002, 34, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Abraham, M.R.; Henrikson, C.A.; Tung, L.; Chang, M.G.; Aon, M.; Xue, T.; Li, R.A.; B, O.R.; Marban, E. Antiarrhythmic engineering of skeletal myoblasts for cardiac transplantation. Circ. Res. 2005, 97, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Brickwedel, J.; Gulbins, H.; Reichenspurner, H. Long-term follow-up after autologous skeletal myoblast transplantation in ischaemic heart disease. Interact. Cardiovasc. Thorac. Surg. 2014, 18, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Dib, N.; Dinsmore, J.; Lababidi, Z.; White, B.; Moravec, S.; Campbell, A.; Rosenbaum, A.; Seyedmadani, K.; Jaber, W.A.; Rizenhour, C.S.; et al. One-year follow-up of feasibility and safety of the first U.S., randomized, controlled study using 3-dimensional guided catheter-based delivery of autologous skeletal myoblasts for ischemic cardiomyopathy (CAuSMIC study). JACC Cardiovasc. Interv. 2009, 2, 9–16. [Google Scholar] [CrossRef]
- Duckers, H.J.; Houtgraaf, J.; Hehrlein, C.; Schofer, J.; Waltenberger, J.; Gershlick, A.; Bartunek, J.; Nienaber, C.; Macaya, C.; Peters, N.; et al. Final results of a phase IIa, randomised, open-label trial to evaluate the percutaneous intramyocardial transplantation of autologous skeletal myoblasts in congestive heart failure patients: The SEISMIC trial. Eurointerv. J. Eur. Collab. Work. Group Interv. Cardiol. Eur. Soc. Cardiol. 2011, 6, 805–812. [Google Scholar] [CrossRef]
- Menasche, P.; Alfieri, O.; Janssens, S.; McKenna, W.; Reichenspurner, H.; Trinquart, L.; Vilquin, J.T.; Marolleau, J.P.; Seymour, B.; Larghero, J.; et al. The Myoblast Autologous Grafting in Ischemic Cardiomyopathy (MAGIC) trial: First randomized placebo-controlled study of myoblast transplantation. Circulation 2008, 117, 1189–1200. [Google Scholar] [CrossRef]
- Povsic, T.J.; O’Connor, C.M.; Henry, T.; Taussig, A.; Kereiakes, D.J.; Fortuin, F.D.; Niederman, A.; Schatz, R.; Spencer, R.; Owens, D.; et al. A double-blind, randomized, controlled, multicenter study to assess the safety and cardiovascular effects of skeletal myoblast implantation by catheter delivery in patients with chronic heart failure after myocardial infarction. Am. Heart J. 2011, 162, 654–662. [Google Scholar] [CrossRef]
- Ferrari, G.; Cusella-De Angelis, G.; Coletta, M.; Paolucci, E.; Stornaiuolo, A.; Cossu, G.; Mavilio, F. Muscle regeneration by bone marrow-derived myogenic progenitors. Science 1998, 279, 1528–1530. [Google Scholar] [CrossRef]
- Orlic, D.; Kajstura, J.; Chimenti, S.; Jakoniuk, I.; Anderson, S.M.; Li, B.; Pickel, J.; McKay, R.; Nadal-Ginard, B.; Bodine, D.M.; et al. Bone marrow cells regenerate infarcted myocardium. Nature 2001, 410, 701–705. [Google Scholar] [CrossRef] [PubMed]
- Strauer, B.E.; Brehm, M.; Zeus, T.; Gattermann, N.; Hernandez, A.; Sorg, R.V.; Kogler, G.; Wernet, P. Intracoronary, human autologous stem cell transplantation for myocardial regeneration following myocardial infarction. Deutsche Medizinische Wochenschrift 1946 2001, 126, 932–938. [Google Scholar] [CrossRef]
- Wen, Y.; Chen, B.; Wang, C.; Ma, X.; Gao, Q. Bone marrow-derived mononuclear cell therapy for patients with ischemic heart disease and ischemic heart failure. Expert Opin. Biol. Ther. 2012, 12, 1563–1573. [Google Scholar] [CrossRef] [PubMed]
- De Jong, R.; Houtgraaf, J.H.; Samiei, S.; Boersma, E.; Duckers, H.J. Intracoronary stem cell infusion after acute myocardial infarction: A meta-analysis and update on clinical trials. Circ. Cardiovasc. Interv. 2014, 7, 156–167. [Google Scholar] [CrossRef] [PubMed]
- Malliaras, K.; Marban, E. Cardiac cell therapy: Where we’ve been, where we are, and where we should be headed. Br. Med. Bull. 2011, 98, 161–185. [Google Scholar] [CrossRef]
- Miao, C.; Lei, M.; Hu, W.; Han, S.; Wang, Q. A brief review: The therapeutic potential of bone marrow mesenchymal stem cells in myocardial infarction. Stem Cell Res. Ther. 2017, 8, 242. [Google Scholar] [CrossRef]
- Erbs, S.; Linke, A.; Schachinger, V.; Assmus, B.; Thiele, H.; Diederich, K.W.; Hoffmann, C.; Dimmeler, S.; Tonn, T.; Hambrecht, R.; et al. Restoration of microvascular function in the infarct-related artery by intracoronary transplantation of bone marrow progenitor cells in patients with acute myocardial infarction: The Doppler Substudy of the Reinfusion of Enriched Progenitor Cells and Infarct Remodeling in Acute Myocardial Infarction (REPAIR-AMI) trial. Circulation 2007, 116, 366–374. [Google Scholar] [CrossRef]
- Asahara, T.; Kawamoto, A.; Masuda, H. Concise review: Circulating endothelial progenitor cells for vascular medicine. Stem Cells 2011, 29, 1650–1655. [Google Scholar] [CrossRef]
- Steinhoff, G.; Nesteruk, J.; Wolfien, M.; Kundt, G.; Borgermann, J.; David, R.; Garbade, J.; Grosse, J.; Haverich, A.; Hennig, H.; et al. Cardiac Function Improvement and Bone Marrow Response−: Outcome Analysis of the Randomized PERFECT Phase III Clinical Trial of Intramyocardial CD133(+) Application After Myocardial Infarction. EBioMedicine 2017, 22, 208–224. [Google Scholar] [CrossRef]
- Karantalis, V.; Schulman, I.H.; Balkan, W.; Hare, J.M. Allogeneic cell therapy: A new paradigm in therapeutics. Circ. Res. 2015, 116, 12–15. [Google Scholar] [CrossRef]
- Dimmeler, S.; Zeiher, A.M. Cell therapy of acute myocardial infarction: Open questions. Cardiology 2009, 113, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Kanelidis, A.J.; Premer, C.; Lopez, J.; Balkan, W.; Hare, J.M. Route of Delivery Modulates the Efficacy of Mesenchymal Stem Cell Therapy for Myocardial Infarction: A Meta-Analysis of Preclinical Studies and Clinical Trials. Circ. Res. 2017, 120, 1139–1150. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.L.; Fang, W.W.; Ye, F.; Liu, Y.H.; Qian, J.; Shan, S.J.; Zhang, J.J.; Chunhua, R.Z.; Liao, L.M.; Lin, S.; et al. Effect on left ventricular function of intracoronary transplantation of autologous bone marrow mesenchymal stem cell in patients with acute myocardial infarction. Am. J. Cardiol. 2004, 94, 92–95. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Lee, S.H.; Youn, Y.J.; Ahn, M.S.; Kim, J.Y.; Yoo, B.S.; Yoon, J.; Kwon, W.; Hong, I.S.; Lee, K.; et al. A randomized, open-label, multicenter trial for the safety and efficacy of adult mesenchymal stem cells after acute myocardial infarction. J. Korean Med Sci. 2014, 29, 23–31. [Google Scholar] [CrossRef]
- Chullikana, A.; Majumdar, A.S.; Gottipamula, S.; Krishnamurthy, S.; Kumar, A.S.; Prakash, V.S.; Gupta, P.K. Randomized, double-blind, phase I/II study of intravenous allogeneic mesenchymal stromal cells in acute myocardial infarction. Cytotherapy 2015, 17, 250–261. [Google Scholar] [CrossRef]
- Gao, L.R.; Pei, X.T.; Ding, Q.A.; Chen, Y.; Zhang, N.K.; Chen, H.Y.; Wang, Z.G.; Wang, Y.F.; Zhu, Z.M.; Li, T.C.; et al. A critical challenge: Dosage-related efficacy and acute complication intracoronary injection of autologous bone marrow mesenchymal stem cells in acute myocardial infarction. Int. J. Cardiol. 2013, 168, 3191–3199. [Google Scholar] [CrossRef]
- Hare, J.M.; Fishman, J.E.; Gerstenblith, G.; DiFede Velazquez, D.L.; Zambrano, J.P.; Suncion, V.Y.; Tracy, M.; Ghersin, E.; Johnston, P.V.; Brinker, J.A.; et al. Comparison of allogeneic vs. autologous bone marrow-derived mesenchymal stem cells delivered by transendocardial injection in patients with ischemic cardiomyopathy: The POSEIDON randomized trial. JAMA 2012, 308, 2369–2379. [Google Scholar] [CrossRef]
- Hare, J.M.; DiFede, D.L.; Rieger, A.C.; Florea, V.; Landin, A.M.; El-Khorazaty, J.; Khan, A.; Mushtaq, M.; Lowery, M.H.; Byrnes, J.J.; et al. Randomized Comparison of Allogeneic Versus Autologous Mesenchymal Stem Cells for Nonischemic Dilated Cardiomyopathy: POSEIDON-DCM Trial. J. Am. Coll. Cardiol. 2017, 69, 526–537. [Google Scholar] [CrossRef]
- Bergmann, O.; Jovinge, S. Cardiac regeneration in vivo: Mending the heart from within? Stem Cell Res. 2014, 13, 523–531. [Google Scholar] [CrossRef]
- Bergmann, O.; Zdunek, S.; Alkass, K.; Druid, H.; Bernard, S.; Frisen, J. Identification of cardiomyocyte nuclei and assessment of ploidy for the analysis of cell turnover. Exp. Cell Res. 2011, 317, 188–194. [Google Scholar] [CrossRef]
- Laflamme, M.A.; Murry, C.E. Heart regeneration. Nature 2011, 473, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Ang, K.L.; Shenje, L.T.; Reuter, S.; Soonpaa, M.H.; Rubart, M.; Field, L.J.; Galinanes, M. Limitations of conventional approaches to identify myocyte nuclei in histologic sections of the heart. Am. J. Physiol. Cell Physiol. 2010, 298, C1603–C1609. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, O.; Bhardwaj, R.D.; Bernard, S.; Zdunek, S.; Barnabe-Heider, F.; Walsh, S.; Zupicich, J.; Alkass, K.; Buchholz, B.A.; Druid, H.; et al. Evidence for cardiomyocyte renewal in humans. Science 2009, 324, 98–102. [Google Scholar] [CrossRef] [PubMed]
- Mollova, M.; Bersell, K.; Walsh, S.; Savla, J.; Das, L.T.; Park, S.Y.; Silberstein, L.E.; Dos Remedios, C.G.; Graham, D.; Colan, S.; et al. Cardiomyocyte proliferation contributes to heart growth in young humans. Proc. Natl. Acad. Sci. USA 2013, 110, 1446–1451. [Google Scholar] [CrossRef] [PubMed]
- Naqvi, N.; Li, M.; Calvert, J.W.; Tejada, T.; Lambert, J.P.; Wu, J.; Kesteven, S.H.; Holman, S.R.; Matsuda, T.; Lovelock, J.D.; et al. A proliferative burst during preadolescence establishes the final cardiomyocyte number. Cell 2014, 157, 795–807. [Google Scholar] [CrossRef] [PubMed]
- Beltrami, A.P.; Barlucchi, L.; Torella, D.; Baker, M.; Limana, F.; Chimenti, S.; Kasahara, H.; Rota, M.; Musso, E.; Urbanek, K.; et al. Adult cardiac stem cells are multipotent and support myocardial regeneration. Cell 2003, 114, 763–776. [Google Scholar] [CrossRef]
- Aguilar-Sanchez, C.; Michael, M.; Pennings, S. Cardiac Stem Cells in the Postnatal Heart: Lessons from Development. Stem Cells Int. 2018, 2018, 1247857. [Google Scholar] [CrossRef]
- Hesse, M.; Fleischmann, B.K.; Kotlikoff, M.I. Concise review: The role of C-kit expressing cells in heart repair at the neonatal and adult stage. Stem Cells 2014, 32, 1701–1712. [Google Scholar] [CrossRef]
- Nigro, P.; Perrucci, G.L.; Gowran, A.; Zanobini, M.; Capogrossi, M.C.; Pompilio, G. c-kit(+) cells: The tell-tale heart of cardiac regeneration? Cell. Mol. Life Sci. 2015, 72, 1725–1740. [Google Scholar] [CrossRef]
- Jesty, S.A.; Steffey, M.A.; Lee, F.K.; Breitbach, M.; Hesse, M.; Reining, S.; Lee, J.C.; Doran, R.M.; Nikitin, A.Y.; Fleischmann, B.K.; et al. c-kit+ precursors support postinfarction myogenesis in the neonatal, but not adult, heart. Proc. Natl. Acad. Sci. USA 2012, 109, 13380–13385. [Google Scholar] [CrossRef]
- Nadal-Ginard, B.; Ellison, G.M.; Torella, D. The cardiac stem cell compartment is indispensable for myocardial cell homeostasis, repair and regeneration in the adult. Stem Cell Res. 2014, 13, 615–630. [Google Scholar] [CrossRef] [PubMed]
- Gude, N.A.; Firouzi, F.; Broughton, K.M.; Ilves, K.; Nguyen, K.P.; Payne, C.R.; Sacchi, V.; Monsanto, M.M.; Casillas, A.R.; Khalafalla, F.G.; et al. Cardiac c-Kit Biology Revealed by Inducible Transgenesis. Circ. Res. 2018, 123, 57–72. [Google Scholar] [CrossRef] [PubMed]
- Hodgkinson, C.P.; Gomez, J.A.; Baksh, S.S.; Payne, A.; Schmeckpeper, J.; Pratt, R.E.; Dzau, V.J. Insights from molecular signature of in vivo cardiac c-Kit(+) cells following cardiac injury and beta-catenin inhibition. J. Mol. Cell. Cardiol. 2018, 123, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; He, L.; Huang, X.; Bhaloo, S.I.; Zhao, H.; Zhang, S.; Pu, W.; Tian, X.; Li, Y.; Liu, Q.; et al. Genetic Lineage Tracing of Nonmyocyte Population by Dual Recombinases. Circulation 2018, 138, 793–805. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lv, Z.; He, L.; Huang, X.; Zhang, S.; Zhao, H.; Pu, W.; Li, Y.; Yu, W.; Zhang, L.; et al. Genetic Tracing Identifies Early Segregation of the Cardiomyocyte and Nonmyocyte Lineages. Circ. Res. 2019, 125, 343–355. [Google Scholar] [CrossRef] [PubMed]
- Elhelaly, W.M.; Cardoso, A.C.; Pereira, A.H.M.; Elnawasany, A.; Ebrahimi, S.; Nakada, Y.; Sadek, H.A. C-Kit Cells Do Not Significantly Contribute to Cardiomyogenesis During Neonatal Heart Regeneration. Circulation 2019, 139, 559–561. [Google Scholar] [CrossRef] [PubMed]
- Tzahor, E.; Poss, K.D. Cardiac regeneration strategies: Staying young at heart. Science 2017, 356, 1035–1039. [Google Scholar] [CrossRef]
- Van Berlo, J.H.; Kanisicak, O.; Maillet, M.; Vagnozzi, R.J.; Karch, J.; Lin, S.C.; Middleton, R.C.; Marban, E.; Molkentin, J.D. c-kit+ cells minimally contribute cardiomyocytes to the heart. Nature 2014, 509, 337–341. [Google Scholar] [CrossRef]
- Sussman, M.A.; Murry, C.E. Bones of contention: Marrow-derived cells in myocardial regeneration. J. Mol. Cell. Cardiol. 2008, 44, 950–953. [Google Scholar] [CrossRef]
- Evans, M.J.; Kaufman, M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature 1981, 292, 154–156. [Google Scholar] [CrossRef]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic stem cell lines derived from human blastocysts. Science 1998, 282, 1145–1147. [Google Scholar] [CrossRef] [PubMed]
- De Trizio, E.; Brennan, C.S. The business of human embryonic stem cell research and an international analysis of relevant laws. J. Biolaw Bus. 2004, 7, 14–22. [Google Scholar] [PubMed]
- Jain, K.K. Ethical and regulatory aspects of embryonic stem cell research. Expert Opin. Biol. Ther. 2005, 5, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Murry, C.E.; Keller, G. Differentiation of embryonic stem cells to clinically relevant populations: Lessons from embryonic development. Cell 2008, 132, 661–680. [Google Scholar] [CrossRef] [PubMed]
- Davis, R.L.; Weintraub, H.; Lassar, A.B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 1987, 51, 987–1000. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- Okita, K.; Ichisaka, T.; Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 2007, 448, 313–317. [Google Scholar] [CrossRef]
- Masip, M.; Veiga, A.; Izpisua Belmonte, J.C.; Simon, C. Reprogramming with defined factors: From induced pluripotency to induced transdifferentiation. Mol. Hum. Reprod. 2010, 16, 856–868. [Google Scholar] [CrossRef]
- Zhao, Y.; Yin, X.; Qin, H.; Zhu, F.; Liu, H.; Yang, W.; Zhang, Q.; Xiang, C.; Hou, P.; Song, Z.; et al. Two supporting factors greatly improve the efficiency of human iPSC generation. Cell Stem Cell 2008, 3, 475–479. [Google Scholar] [CrossRef]
- Han, J.; Yuan, P.; Yang, H.; Zhang, J.; Soh, B.S.; Li, P.; Lim, S.L.; Cao, S.; Tay, J.; Orlov, Y.L.; et al. Tbx3 improves the germ-line competency of induced pluripotent stem cells. Nature 2010, 463, 1096–1100. [Google Scholar] [CrossRef]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007, 131, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Vodyanik, M.A.; Smuga-Otto, K.; Antosiewicz-Bourget, J.; Frane, J.L.; Tian, S.; Nie, J.; Jonsdottir, G.A.; Ruotti, V.; Stewart, R.; et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007, 318, 1917–1920. [Google Scholar] [CrossRef] [PubMed]
- Wernig, M.; Meissner, A.; Foreman, R.; Brambrink, T.; Ku, M.; Hochedlinger, K.; Bernstein, B.E.; Jaenisch, R. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 2007, 448, 318–324. [Google Scholar] [CrossRef] [PubMed]
- Maherali, N.; Sridharan, R.; Xie, W.; Utikal, J.; Eminli, S.; Arnold, K.; Stadtfeld, M.; Yachechko, R.; Tchieu, J.; Jaenisch, R.; et al. Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 2007, 1, 55–70. [Google Scholar] [CrossRef] [PubMed]
- Chin, M.H.; Mason, M.J.; Xie, W.; Volinia, S.; Singer, M.; Peterson, C.; Ambartsumyan, G.; Aimiuwu, O.; Richter, L.; Zhang, J.; et al. Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell 2009, 5, 111–123. [Google Scholar] [CrossRef]
- Doi, A.; Park, I.H.; Wen, B.; Murakami, P.; Aryee, M.J.; Irizarry, R.; Herb, B.; Ladd-Acosta, C.; Rho, J.; Loewer, S.; et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat. Genet. 2009, 41, 1350–1353. [Google Scholar] [CrossRef]
- Marchetto, M.C.; Yeo, G.W.; Kainohana, O.; Marsala, M.; Gage, F.H.; Muotri, A.R. Transcriptional signature and memory retention of human-induced pluripotent stem cells. PLoS ONE 2009, 4, e7076. [Google Scholar] [CrossRef]
- Kim, K.; Doi, A.; Wen, B.; Ng, K.; Zhao, R.; Cahan, P.; Kim, J.; Aryee, M.J.; Ji, H.; Ehrlich, L.I.; et al. Epigenetic memory in induced pluripotent stem cells. Nature 2010, 467, 285–290. [Google Scholar] [CrossRef]
- Hewitt, K.J.; Garlick, J.A. Cellular reprogramming to reset epigenetic signatures. Mol. Asp. Med. 2013, 34, 841–848. [Google Scholar] [CrossRef]
- Gherghiceanu, M.; Barad, L.; Novak, A.; Reiter, I.; Itskovitz-Eldor, J.; Binah, O.; Popescu, L.M. Cardiomyocytes derived from human embryonic and induced pluripotent stem cells: Comparative ultrastructure. J. Cell. Mol. Med. 2011, 15, 2539–2551. [Google Scholar] [CrossRef]
- Gupta, M.K.; Illich, D.J.; Gaarz, A.; Matzkies, M.; Nguemo, F.; Pfannkuche, K.; Liang, H.; Classen, S.; Reppel, M.; Schultze, J.L.; et al. Global transcriptional profiles of beating clusters derived from human induced pluripotent stem cells and embryonic stem cells are highly similar. BMC Dev. Biol. 2010, 10, 98. [Google Scholar] [CrossRef] [PubMed]
- Volarevic, V.; Markovic, B.S.; Gazdic, M.; Volarevic, A.; Jovicic, N.; Arsenijevic, N.; Armstrong, L.; Djonov, V.; Lako, M.; Stojkovic, M. Ethical and Safety Issues of Stem Cell-Based Therapy. Int. J. Med. Sci. 2018, 15, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Kiskinis, E.; Eggan, K. Progress toward the clinical application of patient-specific pluripotent stem cells. J. Clin. Investig. 2010, 120, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, A.; Stacey, G.; Kidane, L.; Seriola, A.; Stachelscheid, H.; Veiga, A. Regulatory insight into the European human pluripotent stem cell registry. Stem Cells Dev. 2014, 23 (Suppl. 1), 51–55. [Google Scholar] [CrossRef]
- Lee, J.Y.; Lee, D.Y.; Choi, Y.S.; Lee, K.J.; Kim, Y.O. Registration of human embryonic stem cell lines: Korea, 2010. Osong Public Health Res. Perspect. 2011, 2, 141–147. [Google Scholar] [CrossRef][Green Version]
- Obama, B. Executive Order 13505 of March 9, 2009: Removing Barriers to Responsible Scientific Research Involving Human Stem Cells. Government Publishing Office. 2009. Available online: https://www.gpo.gov/fdsys/pkg/FR-2009-03-11/pdf/E9-5441.pdf (accessed on 9 March 2009).
- Sanganalmath, S.K.; Bolli, R. Cell therapy for heart failure: A comprehensive overview of experimental and clinical studies, current challenges, and future directions. Circ. Res. 2013, 113, 810–834. [Google Scholar] [CrossRef]
- Olson, E.N. Gene regulatory networks in the evolution and development of the heart. Science 2006, 313, 1922–1927. [Google Scholar] [CrossRef]
- Costello, I.; Pimeisl, I.M.; Drager, S.; Bikoff, E.K.; Robertson, E.J.; Arnold, S.J. The T-box transcription factor Eomesodermin acts upstream of Mesp1 to specify cardiac mesoderm during mouse gastrulation. Nat. Cell Biol. 2011, 13, 1084–1091. [Google Scholar] [CrossRef]
- David, R.; Jarsch, V.B.; Schwarz, F.; Nathan, P.; Gegg, M.; Lickert, H.; Franz, W.M. Induction of MesP1 by Brachyury(T) generates the common multipotent cardiovascular stem cell. Cardiovasc. Res. 2011, 92, 115–122. [Google Scholar] [CrossRef]
- Wu, S.M.; Chien, K.R.; Mummery, C. Origins and fates of cardiovascular progenitor cells. Cell 2008, 132, 537–543. [Google Scholar] [CrossRef]
- Sylva, M.; van den Hoff, M.J.; Moorman, A.F. Development of the human heart. Am. J. Med. Genet. Part A 2014, 164A, 1347–1371. [Google Scholar] [CrossRef] [PubMed]
- De Bakker, B.S.; de Jong, K.H.; Hagoort, J.; de Bree, K.; Besselink, C.T.; de Kanter, F.E.; Veldhuis, T.; Bais, B.; Schildmeijer, R.; Ruijter, J.M.; et al. An interactive three-dimensional digital atlas and quantitative database of human development. Science 2016, 354. [Google Scholar] [CrossRef] [PubMed]
- Gadue, P.; Huber, T.L.; Nostro, M.C.; Kattman, S.; Keller, G.M. Germ layer induction from embryonic stem cells. Exp. Hematol. 2005, 33, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Moretti, A.; Laugwitz, K.L.; Dorn, T.; Sinnecker, D.; Mummery, C. Pluripotent stem cell models of human heart disease. Cold Spring Harb. Perspect. Med. 2013, 3. [Google Scholar] [CrossRef] [PubMed]
- Laflamme, M.A.; Chen, K.Y.; Naumova, A.V.; Muskheli, V.; Fugate, J.A.; Dupras, S.K.; Reinecke, H.; Xu, C.; Hassanipour, M.; Police, S.; et al. Cardiomyocytes derived from human embryonic stem cells in pro-survival factors enhance function of infarcted rat hearts. Nat. Biotechnol. 2007, 25, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Lee, M.Y.; Schliffke, S.; Paavola, J.; Amos, P.J.; Ge, X.; Ye, M.; Zhu, S.; Senyei, G.; Lum, L.; et al. Small molecule Wnt inhibitors enhance the efficiency of BMP-4-directed cardiac differentiation of human pluripotent stem cells. J. Mol. Cell. Cardiol. 2011, 51, 280–287. [Google Scholar] [CrossRef]
- Zeineddine, D.; Papadimou, E.; Mery, A.; Menard, C.; Puceat, M. Cardiac commitment of embryonic stem cells for myocardial repair. Methods Mol. Med. 2005, 112, 175–182. [Google Scholar]
- Kehat, I.; Kenyagin-Karsenti, D.; Snir, M.; Segev, H.; Amit, M.; Gepstein, A.; Livne, E.; Binah, O.; Itskovitz-Eldor, J.; Gepstein, L. Human embryonic stem cells can differentiate into myocytes with structural and functional properties of cardiomyocytes. J. Clin. Investig. 2001, 108, 407–414. [Google Scholar] [CrossRef]
- Mauritz, C.; Schwanke, K.; Reppel, M.; Neef, S.; Katsirntaki, K.; Maier, L.S.; Nguemo, F.; Menke, S.; Haustein, M.; Hescheler, J.; et al. Generation of functional murine cardiac myocytes from induced pluripotent stem cells. Circulation 2008, 118, 507–517. [Google Scholar] [CrossRef]
- Narazaki, G.; Uosaki, H.; Teranishi, M.; Okita, K.; Kim, B.; Matsuoka, S.; Yamanaka, S.; Yamashita, J.K. Directed and systematic differentiation of cardiovascular cells from mouse induced pluripotent stem cells. Circulation 2008, 118, 498–506. [Google Scholar] [CrossRef]
- Mummery, C.; Ward-van Oostwaard, D.; Doevendans, P.; Spijker, R.; van den Brink, S.; Hassink, R.; van der Heyden, M.; Opthof, T.; Pera, M.; de la Riviere, A.B.; et al. Differentiation of human embryonic stem cells to cardiomyocytes: Role of coculture with visceral endoderm-like cells. Circulation 2003, 107, 2733–2740. [Google Scholar] [CrossRef] [PubMed]
- Burridge, P.W.; Matsa, E.; Shukla, P.; Lin, Z.C.; Churko, J.M.; Ebert, A.D.; Lan, F.; Diecke, S.; Huber, B.; Mordwinkin, N.M.; et al. Chemically defined generation of human cardiomyocytes. Nat. Methods 2014, 11, 855–860. [Google Scholar] [CrossRef] [PubMed]
- Lian, X.; Zhang, J.; Azarin, S.M.; Zhu, K.; Hazeltine, L.B.; Bao, X.; Hsiao, C.; Kamp, T.J.; Palecek, S.P. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/beta-catenin signaling under fully defined conditions. Nat. Protoc. 2013, 8, 162–175. [Google Scholar] [CrossRef] [PubMed]
- Lian, X.; Bao, X.; Zilberter, M.; Westman, M.; Fisahn, A.; Hsiao, C.; Hazeltine, L.B.; Dunn, K.K.; Kamp, T.J.; Palecek, S.P. Chemically defined, albumin-free human cardiomyocyte generation. Nat. Methods 2015, 12, 595–596. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Soonpaa, M.H.; Adler, E.D.; Roepke, T.K.; Kattman, S.J.; Kennedy, M.; Henckaerts, E.; Bonham, K.; Abbott, G.W.; Linden, R.M.; et al. Human cardiovascular progenitor cells develop from a KDR+ embryonic-stem-cell-derived population. Nature 2008, 453, 524–528. [Google Scholar] [CrossRef] [PubMed]
- Moretti, A.; Caron, L.; Nakano, A.; Lam, J.T.; Bernshausen, A.; Chen, Y.; Qyang, Y.; Bu, L.; Sasaki, M.; Martin-Puig, S.; et al. Multipotent embryonic isl1+ progenitor cells lead to cardiac, smooth muscle, and endothelial cell diversification. Cell 2006, 127, 1151–1165. [Google Scholar] [CrossRef]
- Elliott, D.A.; Braam, S.R.; Koutsis, K.; Ng, E.S.; Jenny, R.; Lagerqvist, E.L.; Biben, C.; Hatzistavrou, T.; Hirst, C.E.; Yu, Q.C.; et al. NKX2-5(eGFP/w) hESCs for isolation of human cardiac progenitors and cardiomyocytes. Nat. Methods 2011, 8, 1037–1040. [Google Scholar] [CrossRef]
- Dubois, N.C.; Craft, A.M.; Sharma, P.; Elliott, D.A.; Stanley, E.G.; Elefanty, A.G.; Gramolini, A.; Keller, G. SIRPA is a specific cell-surface marker for isolating cardiomyocytes derived from human pluripotent stem cells. Nat. Biotechnol. 2011, 29, 1011–1018. [Google Scholar] [CrossRef]
- Uosaki, H.; Fukushima, H.; Takeuchi, A.; Matsuoka, S.; Nakatsuji, N.; Yamanaka, S.; Yamashita, J.K. Efficient and scalable purification of cardiomyocytes from human embryonic and induced pluripotent stem cells by VCAM1 surface expression. PLoS ONE 2011, 6, e23657. [Google Scholar] [CrossRef]
- Gallo, P.; Grimaldi, S.; Latronico, M.V.; Bonci, D.; Pagliuca, A.; Gallo, P.; Ausoni, S.; Peschle, C.; Condorelli, G. A lentiviral vector with a short troponin-I promoter for tracking cardiomyocyte differentiation of human embryonic stem cells. Gene Ther. 2008, 15, 161–170. [Google Scholar] [CrossRef]
- Kita-Matsuo, H.; Barcova, M.; Prigozhina, N.; Salomonis, N.; Wei, K.; Jacot, J.G.; Nelson, B.; Spiering, S.; Haverslag, R.; Kim, C.; et al. Lentiviral vectors and protocols for creation of stable hESC lines for fluorescent tracking and drug resistance selection of cardiomyocytes. PLoS ONE 2009, 4, e5046. [Google Scholar] [CrossRef] [PubMed]
- Tohyama, S.; Hattori, F.; Sano, M.; Hishiki, T.; Nagahata, Y.; Matsuura, T.; Hashimoto, H.; Suzuki, T.; Yamashita, H.; Satoh, Y.; et al. Distinct metabolic flow enables large-scale purification of mouse and human pluripotent stem cell-derived cardiomyocytes. Cell Stem Cell 2013, 12, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Ransom, J.F.; Li, A.; Vedantham, V.; von Drehle, M.; Muth, A.N.; Tsuchihashi, T.; McManus, M.T.; Schwartz, R.J.; Srivastava, D. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 2007, 129, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.D.; Rushing, S.N.; Lieu, D.K.; Chan, C.W.; Kong, C.W.; Geng, L.; Wilson, K.D.; Chiamvimonvat, N.; Boheler, K.R.; Wu, J.C.; et al. Distinct roles of microRNA-1 and -499 in ventricular specification and functional maturation of human embryonic stem cell-derived cardiomyocytes. PLoS ONE 2011, 6, e27417. [Google Scholar] [CrossRef] [PubMed]
- Hodgkinson, C.P.; Kang, M.H.; Dal-Pra, S.; Mirotsou, M.; Dzau, V.J. MicroRNAs and Cardiac Regeneration. Circ. Res. 2015, 116, 1700–1711. [Google Scholar] [CrossRef]
- Yang, X.; Pabon, L.; Murry, C.E. Engineering adolescence: Maturation of human pluripotent stem cell-derived cardiomyocytes. Circ. Res. 2014, 114, 511–523. [Google Scholar] [CrossRef]
- Friedman, C.E.; Nguyen, Q.; Lukowski, S.W.; Helfer, A.; Chiu, H.S.; Miklas, J.; Levy, S.; Suo, S.; Han, J.J.; Osteil, P.; et al. Single-Cell Transcriptomic Analysis of Cardiac Differentiation from Human PSCs Reveals HOPX-Dependent Cardiomyocyte Maturation. Cell Stem Cell 2018, 23, 586–598. [Google Scholar] [CrossRef]
- Churko, J.M.; Garg, P.; Treutlein, B.; Venkatasubramanian, M.; Wu, H.; Lee, J.; Wessells, Q.N.; Chen, S.Y.; Chen, W.Y.; Chetal, K.; et al. Defining human cardiac transcription factor hierarchies using integrated single-cell heterogeneity analysis. Nat. Commun. 2018, 9, 4906. [Google Scholar] [CrossRef]
- Lundy, S.D.; Zhu, W.Z.; Regnier, M.; Laflamme, M.A. Structural and functional maturation of cardiomyocytes derived from human pluripotent stem cells. Stem Cells Dev. 2013, 22, 1991–2002. [Google Scholar] [CrossRef]
- Bhavsar, P.K.; Dhoot, G.K.; Cumming, D.V.; Butler-Browne, G.S.; Yacoub, M.H.; Barton, P.J. Developmental expression of troponin I isoforms in fetal human heart. FEBS Lett. 1991, 292, 5–8. [Google Scholar]
- Hunkeler, N.M.; Kullman, J.; Murphy, A.M. Troponin I isoform expression in human heart. Circ. Res. 1991, 69, 1409–1414. [Google Scholar] [CrossRef] [PubMed]
- Bedada, F.B.; Chan, S.S.; Metzger, S.K.; Zhang, L.; Zhang, J.; Garry, D.J.; Kamp, T.J.; Kyba, M.; Metzger, J.M. Acquisition of a quantitative, stoichiometrically conserved ratiometric marker of maturation status in stem cell-derived cardiac myocytes. Stem Cell Rep. 2014, 3, 594–605. [Google Scholar] [CrossRef] [PubMed]
- Taegtmeyer, H.; Sen, S.; Vela, D. Return to the fetal gene program: A suggested metabolic link to gene expression in the heart. Ann. N. Y. Acad. Sci. 2010, 1188, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Everett, A.W. Isomyosin expression in human heart in early pre- and post-natal life. J. Mol. Cell. Cardiol. 1986, 18, 607–615. [Google Scholar] [CrossRef]
- Weber, N.; Schwanke, K.; Greten, S.; Wendland, M.; Iorga, B.; Fischer, M.; Geers-Knorr, C.; Hegermann, J.; Wrede, C.; Fiedler, J.; et al. Stiff matrix induces switch to pure beta-cardiac myosin heavy chain expression in human ESC-derived cardiomyocytes. Basic Res. Cardiol. 2016, 111, 68. [Google Scholar] [CrossRef]
- Lahmers, S.; Wu, Y.; Call, D.R.; Labeit, S.; Granzier, H. Developmental control of titin isoform expression and passive stiffness in fetal and neonatal myocardium. Circ. Res. 2004, 94, 505–513. [Google Scholar] [CrossRef]
- Xu, X.Q.; Soo, S.Y.; Sun, W.; Zweigerdt, R. Global expression profile of highly enriched cardiomyocytes derived from human embryonic stem cells. Stem Cells 2009, 27, 2163–2174. [Google Scholar] [CrossRef]
- Blazeski, A.; Zhu, R.; Hunter, D.W.; Weinberg, S.H.; Boheler, K.R.; Zambidis, E.T.; Tung, L. Electrophysiological and contractile function of cardiomyocytes derived from human embryonic stem cells. Prog. Biophys. Mol. Biol. 2012, 110, 178–195. [Google Scholar] [CrossRef]
- Funakoshi, S.; Miki, K.; Takaki, T.; Okubo, C.; Hatani, T.; Chonabayashi, K.; Nishikawa, M.; Takei, I.; Oishi, A.; Narita, M.; et al. Enhanced engraftment, proliferation, and therapeutic potential in heart using optimized human iPSC-derived cardiomyocytes. Sci. Rep. 2016, 6, 19111. [Google Scholar] [CrossRef]
- Synnergren, J.; Ameen, C.; Jansson, A.; Sartipy, P. Global transcriptional profiling reveals similarities and differences between human stem cell-derived cardiomyocyte clusters and heart tissue. Physiol. Genom. 2012, 44, 245–258. [Google Scholar] [CrossRef]
- Lieu, D.K.; Liu, J.; Siu, C.W.; McNerney, G.P.; Tse, H.F.; Abu-Khalil, A.; Huser, T.; Li, R.A. Absence of transverse tubules contributes to non-uniform Ca(2+) wavefronts in mouse and human embryonic stem cell-derived cardiomyocytes. Stem Cells Dev. 2009, 18, 1493–1500. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Bergeron, D.L.; Runge, A.; Dahl, K.D.; Fehling, H.J.; Keller, G.; Simon, M.C. Hypoxia affects mesoderm and enhances hemangioblast specification during early development. Development 2004, 131, 4623–4634. [Google Scholar] [CrossRef] [PubMed]
- Lopaschuk, G.D.; Jaswal, J.S. Energy metabolic phenotype of the cardiomyocyte during development, differentiation, and postnatal maturation. J. Cardiovasc. Pharmacol. 2010, 56, 130–140. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.; Wong, J.; Wen, J.; Wang, S.; Wang, C.; Spiering, S.; Kan, N.G.; Forcales, S.; Puri, P.L.; Leone, T.C.; et al. Studying arrhythmogenic right ventricular dysplasia with patient-specific iPSCs. Nature 2013, 494, 105–110. [Google Scholar] [CrossRef]
- Mills, R.J.; Titmarsh, D.M.; Koenig, X.; Parker, B.L.; Ryall, J.G.; Quaife-Ryan, G.A.; Voges, H.K.; Hodson, M.P.; Ferguson, C.; Drowley, L.; et al. Functional screening in human cardiac organoids reveals a metabolic mechanism for cardiomyocyte cell cycle arrest. Proc. Natl. Acad. Sci. USA 2017, 114, E8372–E8381. [Google Scholar] [CrossRef] [PubMed]
- Hazeltine, L.B.; Simmons, C.S.; Salick, M.R.; Lian, X.; Badur, M.G.; Han, W.; Delgado, S.M.; Wakatsuki, T.; Crone, W.C.; Pruitt, B.L.; et al. Effects of substrate mechanics on contractility of cardiomyocytes generated from human pluripotent stem cells. Int. J. Cell Biol. 2012, 2012, 508294. [Google Scholar] [CrossRef]
- Kim, D.H.; Lipke, E.A.; Kim, P.; Cheong, R.; Thompson, S.; Delannoy, M.; Suh, K.Y.; Tung, L.; Levchenko, A. Nanoscale cues regulate the structure and function of macroscopic cardiac tissue constructs. Proc. Natl. Acad. Sci. USA 2010, 107, 565–570. [Google Scholar] [CrossRef]
- Lee, Y.K.; Ng, K.M.; Chan, Y.C.; Lai, W.H.; Au, K.W.; Ho, C.Y.; Wong, L.Y.; Lau, C.P.; Tse, H.F.; Siu, C.W. Triiodothyronine promotes cardiac differentiation and maturation of embryonic stem cells via the classical genomic pathway. Mol. Endocrinol. 2010, 24, 1728–1736. [Google Scholar] [CrossRef]
- McDevitt, T.C.; Laflamme, M.A.; Murry, C.E. Proliferation of cardiomyocytes derived from human embryonic stem cells is mediated via the IGF/PI 3-kinase/Akt signaling pathway. J. Mol. Cell. Cardiol. 2005, 39, 865–873. [Google Scholar] [CrossRef]
- Zimmermann, W.H.; Schneiderbanger, K.; Schubert, P.; Didie, M.; Munzel, F.; Heubach, J.F.; Kostin, S.; Neuhuber, W.L.; Eschenhagen, T. Tissue engineering of a differentiated cardiac muscle construct. Circ. Res. 2002, 90, 223–230. [Google Scholar] [CrossRef]
- Zimmermann, W.H.; Melnychenko, I.; Wasmeier, G.; Didie, M.; Naito, H.; Nixdorff, U.; Hess, A.; Budinsky, L.; Brune, K.; Michaelis, B.; et al. Engineered heart tissue grafts improve systolic and diastolic function in infarcted rat hearts. Nat. Med. 2006, 12, 452–458. [Google Scholar] [CrossRef] [PubMed]
- Eschenhagen, T.; Eder, A.; Vollert, I.; Hansen, A. Physiological aspects of cardiac tissue engineering. Am. J. Physiol. Heart Circ. Physiol. 2012, 303, H133–H143. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Miao, S.; Esworthy, T.; Zhou, X.; Lee, S.J.; Liu, C.; Yu, Z.X.; Fisher, J.P.; Mohiuddin, M.; Zhang, L.G. 3D bioprinting for cardiovascular regeneration and pharmacology. Adv. Drug Deliv. Rev. 2018, 132, 252–269. [Google Scholar] [CrossRef] [PubMed]
- Sheehy, S.P.; Grosberg, A.; Qin, P.; Behm, D.J.; Ferrier, J.P.; Eagleson, M.A.; Nesmith, A.P.; Krull, D.; Falls, J.G.; Campbell, P.H.; et al. Toward improved myocardial maturity in an organ-on-chip platform with immature cardiac myocytes. Exp. Biol. Med. 2017, 242, 1643–1656. [Google Scholar] [CrossRef] [PubMed]
- Schwan, J.; Kwaczala, A.T.; Ryan, T.J.; Bartulos, O.; Ren, Y.; Sewanan, L.R.; Morris, A.H.; Jacoby, D.L.; Qyang, Y.; Campbell, S.G. Anisotropic engineered heart tissue made from laser-cut decellularized myocardium. Sci. Rep. 2016, 6, 32068. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, F.; Boue, S.; Izpisua Belmonte, J.C. Methods for making induced pluripotent stem cells: Reprogramming a la carte. Nat. Rev. Genet. 2011, 12, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Yamanaka, S. Induced Pluripotent Stem Cells 10 Years Later: For Cardiac Applications. Circ. Res. 2017, 120, 1958–1968. [Google Scholar] [CrossRef]
- Bellin, M.; Mummery, C.L. Inherited heart disease—What can we expect from the second decade of human iPS cell research? FEBS Lett. 2016, 590, 2482–2493. [Google Scholar] [CrossRef]
- Carvajal-Vergara, X.; Sevilla, A.; D’Souza, S.L.; Ang, Y.S.; Schaniel, C.; Lee, D.F.; Yang, L.; Kaplan, A.D.; Adler, E.D.; Rozov, R.; et al. Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature 2010, 465, 808–812. [Google Scholar] [CrossRef]
- Dudek, J.; Cheng, I.F.; Balleininger, M.; Vaz, F.M.; Streckfuss-Bomeke, K.; Hubscher, D.; Vukotic, M.; Wanders, R.J.; Rehling, P.; Guan, K. Cardiolipin deficiency affects respiratory chain function and organization in an induced pluripotent stem cell model of Barth syndrome. Stem Cell Res. 2013, 11, 806–819. [Google Scholar] [CrossRef]
- Fatima, A.; Xu, G.; Shao, K.; Papadopoulos, S.; Lehmann, M.; Arnaiz-Cot, J.J.; Rosa, A.O.; Nguemo, F.; Matzkies, M.; Dittmann, S.; et al. In vitro modeling of ryanodine receptor 2 dysfunction using human induced pluripotent stem cells. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 2011, 28, 579–592. [Google Scholar] [CrossRef] [PubMed]
- Hinson, J.T.; Chopra, A.; Nafissi, N.; Polacheck, W.J.; Benson, C.C.; Swist, S.; Gorham, J.; Yang, L.; Schafer, S.; Sheng, C.C.; et al. HEART DISEASE. Titin mutations in iPS cells define sarcomere insufficiency as a cause of dilated cardiomyopathy. Science 2015, 349, 982–986. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.P.; Chen, P.H.; Hwu, W.L.; Chuang, C.Y.; Chien, Y.H.; Stone, L.; Chien, C.L.; Li, L.T.; Chiang, S.C.; Chen, H.F.; et al. Human Pompe disease-induced pluripotent stem cells for pathogenesis modeling, drug testing and disease marker identification. Hum. Mol. Genet. 2011, 20, 4851–4864. [Google Scholar] [CrossRef] [PubMed]
- Itzhaki, I.; Maizels, L.; Huber, I.; Zwi-Dantsis, L.; Caspi, O.; Winterstern, A.; Feldman, O.; Gepstein, A.; Arbel, G.; Hammerman, H.; et al. Modelling the long QT syndrome with induced pluripotent stem cells. Nature 2011, 471, 225–229. [Google Scholar] [CrossRef]
- Kujala, K.; Paavola, J.; Lahti, A.; Larsson, K.; Pekkanen-Mattila, M.; Viitasalo, M.; Lahtinen, A.M.; Toivonen, L.; Kontula, K.; Swan, H.; et al. Cell model of catecholaminergic polymorphic ventricular tachycardia reveals early and delayed afterdepolarizations. PLoS ONE 2012, 7, e44660. [Google Scholar] [CrossRef]
- Lahti, A.L.; Kujala, V.J.; Chapman, H.; Koivisto, A.P.; Pekkanen-Mattila, M.; Kerkela, E.; Hyttinen, J.; Kontula, K.; Swan, H.; Conklin, B.R.; et al. Model for long QT syndrome type 2 using human iPS cells demonstrates arrhythmogenic characteristics in cell culture. Dis. Models Mech. 2012, 5, 220–230. [Google Scholar] [CrossRef]
- Ma, D.; Wei, H.; Lu, J.; Ho, S.; Zhang, G.; Sun, X.; Oh, Y.; Tan, S.H.; Ng, M.L.; Shim, W.; et al. Generation of patient-specific induced pluripotent stem cell-derived cardiomyocytes as a cellular model of arrhythmogenic right ventricular cardiomyopathy. Eur. Heart J. 2013, 34, 1122–1133. [Google Scholar] [CrossRef]
- Ma, D.; Wei, H.; Zhao, Y.; Lu, J.; Li, G.; Sahib, N.B.; Tan, T.H.; Wong, K.Y.; Shim, W.; Wong, P.; et al. Modeling type 3 long QT syndrome with cardiomyocytes derived from patient-specific induced pluripotent stem cells. Int. J. Cardiol. 2013, 168, 5277–5286. [Google Scholar] [CrossRef]
- Moretti, A.; Bellin, M.; Welling, A.; Jung, C.B.; Lam, J.T.; Bott-Flugel, L.; Dorn, T.; Goedel, A.; Hohnke, C.; Hofmann, F.; et al. Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N. Engl. J. Med. 2010, 363, 1397–1409. [Google Scholar] [CrossRef]
- Yazawa, M.; Hsueh, B.; Jia, X.; Pasca, A.M.; Bernstein, J.A.; Hallmayer, J.; Dolmetsch, R.E. Using induced pluripotent stem cells to investigate cardiac phenotypes in Timothy syndrome. Nature 2011, 471, 230–234. [Google Scholar] [CrossRef]
- Ojala, M.; Prajapati, C.; Polonen, R.P.; Rajala, K.; Pekkanen-Mattila, M.; Rasku, J.; Larsson, K.; Aalto-Setala, K. Mutation-Specific Phenotypes in hiPSC-Derived Cardiomyocytes Carrying Either Myosin-Binding Protein C Or alpha-Tropomyosin Mutation for Hypertrophic Cardiomyopathy. Stem Cells Int. 2016, 2016, 1684792. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; McCain, M.L.; Yang, L.; He, A.; Pasqualini, F.S.; Agarwal, A.; Yuan, H.; Jiang, D.; Zhang, D.; Zangi, L.; et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat. Med. 2014, 20, 616–623. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Yazawa, M.; Liu, J.; Han, L.; Sanchez-Freire, V.; Abilez, O.J.; Navarrete, E.G.; Hu, S.; Wang, L.; Lee, A.; et al. Patient-specific induced pluripotent stem cells as a model for familial dilated cardiomyopathy. Sci. Transl. Med. 2012, 4, 130ra47. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Marceau, C.; Hamaguchi, R.; Burridge, P.W.; Rajarajan, K.; Churko, J.M.; Wu, H.; Sallam, K.I.; Matsa, E.; Sturzu, A.C.; et al. Human induced pluripotent stem cell-derived cardiomyocytes as an in vitro model for coxsackievirus B3-induced myocarditis and antiviral drug screening platform. Circ. Res. 2014, 115, 556–566. [Google Scholar] [CrossRef]
- Gu, M.; Shao, N.Y.; Sa, S.; Li, D.; Termglinchan, V.; Ameen, M.; Karakikes, I.; Sosa, G.; Grubert, F.; Lee, J.; et al. Patient-Specific iPSC-Derived Endothelial Cells Uncover Pathways that Protect against Pulmonary Hypertension in BMPR2 Mutation Carriers. Cell Stem Cell 2017, 20, 490–504. [Google Scholar] [CrossRef]
- Fatima, A.; Kaifeng, S.; Dittmann, S.; Xu, G.; Gupta, M.K.; Linke, M.; Zechner, U.; Nguemo, F.; Milting, H.; Farr, M.; et al. The disease-specific phenotype in cardiomyocytes derived from induced pluripotent stem cells of two long QT syndrome type 3 patients. PLoS ONE 2013, 8, e83005. [Google Scholar] [CrossRef]
- Paci, M.; Hyttinen, J.; Aalto-Setala, K.; Severi, S. Computational models of ventricular- and atrial-like human induced pluripotent stem cell derived cardiomyocytes. Ann. Biomed. Eng. 2013, 41, 2334–2348. [Google Scholar] [CrossRef]
- Paci, M.; Passini, E.; Severi, S.; Hyttinen, J.; Rodriguez, B. Phenotypic variability in LQT3 human induced pluripotent stem cell-derived cardiomyocytes and their response to antiarrhythmic pharmacologic therapy: An in silico approach. Heart Rhythm 2017, 14, 1704–1712. [Google Scholar] [CrossRef]
- Paci, M.; Polonen, R.P.; Cori, D.; Penttinen, K.; Aalto-Setala, K.; Severi, S.; Hyttinen, J. Automatic Optimization of an in Silico Model of Human iPSC Derived Cardiomyocytes Recapitulating Calcium Handling Abnormalities. Front. Physiol. 2018, 9, 709. [Google Scholar] [CrossRef]
- Bellin, M.; Casini, S.; Davis, R.P.; D’Aniello, C.; Haas, J.; Ward-van Oostwaard, D.; Tertoolen, L.G.; Jung, C.B.; Elliott, D.A.; Welling, A.; et al. Isogenic human pluripotent stem cell pairs reveal the role of a KCNH2 mutation in long-QT syndrome. EMBO J. 2013, 32, 3161–3175. [Google Scholar] [CrossRef]
- Rampe, D.; Brown, A.M. A history of the role of the hERG channel in cardiac risk assessment. J. Pharmacol. Toxicol. Methods 2013, 68, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Ahola, A.; Polonen, R.P.; Aalto-Setala, K.; Hyttinen, J. Simultaneous Measurement of Contraction and Calcium Transients in Stem Cell Derived Cardiomyocytes. Ann. Biomed. Eng. 2018, 46, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Bjork, S.; Ojala, E.A.; Nordstrom, T.; Ahola, A.; Liljestrom, M.; Hyttinen, J.; Kankuri, E.; Mervaala, E. Evaluation of Optogenetic Electrophysiology Tools in Human Stem Cell-Derived Cardiomyocytes. Front. Physiol. 2017, 8, 884. [Google Scholar] [CrossRef] [PubMed]
- Klimas, A.; Ambrosi, C.M.; Yu, J.; Williams, J.C.; Bien, H.; Entcheva, E. OptoDyCE as an automated system for high-throughput all-optical dynamic cardiac electrophysiology. Nat. Commun. 2016, 7, 11542. [Google Scholar] [CrossRef]
- Rajamohan, D.; Kalra, S.; Duc Hoang, M.; George, V.; Staniforth, A.; Russell, H.; Yang, X.; Denning, C. Automated Electrophysiological and Pharmacological Evaluation of Human Pluripotent Stem Cell-Derived Cardiomyocytes. Stem Cells Dev. 2016, 25, 439–452. [Google Scholar] [CrossRef]
- Chi, K.R. Revolution dawning in cardiotoxicity testing. Nat. Rev. Drug Discov. 2013, 12, 565–567. [Google Scholar] [CrossRef]
- Miller, J.C.; Tan, S.; Qiao, G.; Barlow, K.A.; Wang, J.; Xia, D.F.; Meng, X.; Paschon, D.E.; Leung, E.; Hinkley, S.J.; et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 2011, 29, 143–148. [Google Scholar] [CrossRef]
- Urnov, F.D.; Rebar, E.J.; Holmes, M.C.; Zhang, H.S.; Gregory, P.D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 2010, 11, 636–646. [Google Scholar] [CrossRef]
- Kim, H.; Kim, J.S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 2014, 15, 321–334. [Google Scholar] [CrossRef]
- Collin, J.; Lako, M. Concise review: Putting a finger on stem cell biology: Zinc finger nuclease-driven targeted genetic editing in human pluripotent stem cells. Stem Cells 2011, 29, 1021–1033. [Google Scholar] [CrossRef]
- Zou, J.; Maeder, M.L.; Mali, P.; Pruett-Miller, S.M.; Thibodeau-Beganny, S.; Chou, B.K.; Chen, G.; Ye, Z.; Park, I.H.; Daley, G.Q.; et al. Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell 2009, 5, 97–110. [Google Scholar] [CrossRef] [PubMed]
- Hockemeyer, D.; Soldner, F.; Beard, C.; Gao, Q.; Mitalipova, M.; DeKelver, R.C.; Katibah, G.E.; Amora, R.; Boydston, E.A.; Zeitler, B.; et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat. Biotechnol. 2009, 27, 851–857. [Google Scholar] [CrossRef] [PubMed]
- Hockemeyer, D.; Wang, H.; Kiani, S.; Lai, C.S.; Gao, Q.; Cassady, J.P.; Cost, G.J.; Zhang, L.; Santiago, Y.; Miller, J.C.; et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 2011, 29, 731–734. [Google Scholar] [CrossRef]
- Yusa, K.; Rashid, S.T.; Strick-Marchand, H.; Varela, I.; Liu, P.Q.; Paschon, D.E.; Miranda, E.; Ordonez, A.; Hannan, N.R.; Rouhani, F.J.; et al. Targeted gene correction of alpha1-antitrypsin deficiency in induced pluripotent stem cells. Nature 2011, 478, 391–394. [Google Scholar] [CrossRef] [PubMed]
- Karakikes, I.; Stillitano, F.; Nonnenmacher, M.; Tzimas, C.; Sanoudou, D.; Termglinchan, V.; Kong, C.W.; Rushing, S.; Hansen, J.; Ceholski, D.; et al. Correction of human phospholamban R14del mutation associated with cardiomyopathy using targeted nucleases and combination therapy. Nat. Commun. 2015, 6, 6955. [Google Scholar] [CrossRef]
- Haghighi, K.; Kolokathis, F.; Gramolini, A.O.; Waggoner, J.R.; Pater, L.; Lynch, R.A.; Fan, G.C.; Tsiapras, D.; Parekh, R.R.; Dorn, G.W., 2nd; et al. A mutation in the human phospholamban gene, deleting arginine 14, results in lethal, hereditary cardiomyopathy. Proc. Natl. Acad. Sci. USA 2006, 103, 1388–1393. [Google Scholar] [CrossRef]
- Van der Zwaag, P.A.; van Rijsingen, I.A.; Asimaki, A.; Jongbloed, J.D.; van Veldhuisen, D.J.; Wiesfeld, A.C.; Cox, M.G.; van Lochem, L.T.; de Boer, R.A.; Hofstra, R.M.; et al. Phospholamban R14del mutation in patients diagnosed with dilated cardiomyopathy or arrhythmogenic right ventricular cardiomyopathy: Evidence supporting the concept of arrhythmogenic cardiomyopathy. Eur. J. Heart Fail. 2012, 14, 1199–1207. [Google Scholar] [CrossRef]
- Ovando-Roche, P.; Georgiadis, A.; Smith, A.J.; Pearson, R.A.; Ali, R.R. Harnessing the Potential of Human Pluripotent Stem Cells and Gene Editing for the Treatment of Retinal Degeneration. Curr. Stem Cell Rep. 2017, 3, 112–123. [Google Scholar] [CrossRef]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef]
- Chaterji, S.; Ahn, E.H.; Kim, D.H. CRISPR Genome Engineering for Human Pluripotent Stem Cell Research. Theranostics 2017, 7, 4445–4469. [Google Scholar] [CrossRef]
- Limpitikul, W.B.; Dick, I.E.; Tester, D.J.; Boczek, N.J.; Limphong, P.; Yang, W.; Choi, M.H.; Babich, J.; DiSilvestre, D.; Kanter, R.J.; et al. A Precision Medicine Approach to the Rescue of Function on Malignant Calmodulinopathic Long-QT Syndrome. Circ. Res. 2017, 120, 39–48. [Google Scholar] [CrossRef]
- Yamamoto, Y.; Makiyama, T.; Harita, T.; Sasaki, K.; Wuriyanghai, Y.; Hayano, M.; Nishiuchi, S.; Kohjitani, H.; Hirose, S.; Chen, J.; et al. Allele-specific ablation rescues electrophysiological abnormalities in a human iPS cell model of long-QT syndrome with a CALM2 mutation. Hum. Mol. Genet. 2017, 26, 1670–1677. [Google Scholar] [CrossRef] [PubMed]
- Brodehl, A.; Ebbinghaus, H.; Deutsch, M.A.; Gummert, J.; Gartner, A.; Ratnavadivel, S.; Milting, H. Human Induced Pluripotent Stem-Cell-Derived Cardiomyocytes as Models for Genetic Cardiomyopathies. Int. J. Mol. Sci. 2019, 20, 4381. [Google Scholar] [CrossRef] [PubMed]
- Te Riele, A.S.; Agullo-Pascual, E.; James, C.A.; Leo-Macias, A.; Cerrone, M.; Zhang, M.; Lin, X.; Lin, B.; Sobreira, N.L.; Amat-Alarcon, N.; et al. Multilevel analyses of SCN5A mutations in arrhythmogenic right ventricular dysplasia/cardiomyopathy suggest non-canonical mechanisms for disease pathogenesis. Cardiovasc. Res. 2017, 113, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Seeger, T.; Shrestha, R.; Lam, C.K.; Chen, C.; McKeithan, W.L.; Lau, E.; Wnorowski, A.; McMullen, G.; Greenhaw, M.; Lee, J.; et al. A Premature Termination Codon Mutation in MYBPC3 Causes Hypertrophic Cardiomyopathy via Chronic Activation of Nonsense-Mediated Decay. Circulation 2019, 139, 799–811. [Google Scholar] [CrossRef] [PubMed]
- Van Laake, L.W.; Passier, R.; Monshouwer-Kloots, J.; Verkleij, A.J.; Lips, D.J.; Freund, C.; den Ouden, K.; Ward-van Oostwaard, D.; Korving, J.; Tertoolen, L.G.; et al. Human embryonic stem cell-derived cardiomyocytes survive and mature in the mouse heart and transiently improve function after myocardial infarction. Stem Cell Res. 2007, 1, 9–24. [Google Scholar] [CrossRef]
- Caspi, O.; Huber, I.; Kehat, I.; Habib, M.; Arbel, G.; Gepstein, A.; Yankelson, L.; Aronson, D.; Beyar, R.; Gepstein, L. Transplantation of human embryonic stem cell-derived cardiomyocytes improves myocardial performance in infarcted rat hearts. J. Am. Coll. Cardiol. 2007, 50, 1884–1893. [Google Scholar] [CrossRef]
- Masumoto, H.; Matsuo, T.; Yamamizu, K.; Uosaki, H.; Narazaki, G.; Katayama, S.; Marui, A.; Shimizu, T.; Ikeda, T.; Okano, T.; et al. Pluripotent stem cell-engineered cell sheets reassembled with defined cardiovascular populations ameliorate reduction in infarct heart function through cardiomyocyte-mediated neovascularization. Stem Cells 2012, 30, 1196–1205. [Google Scholar] [CrossRef]
- Matsuo, T.; Masumoto, H.; Tajima, S.; Ikuno, T.; Katayama, S.; Minakata, K.; Ikeda, T.; Yamamizu, K.; Tabata, Y.; Sakata, R.; et al. Efficient long-term survival of cell grafts after myocardial infarction with thick viable cardiac tissue entirely from pluripotent stem cells. Sci. Rep. 2015, 5, 16842. [Google Scholar] [CrossRef]
- Shiba, Y.; Fernandes, S.; Zhu, W.Z.; Filice, D.; Muskheli, V.; Kim, J.; Palpant, N.J.; Gantz, J.; Moyes, K.W.; Reinecke, H.; et al. Human ES-cell-derived cardiomyocytes electrically couple and suppress arrhythmias in injured hearts. Nature 2012, 489, 322–325. [Google Scholar] [CrossRef]
- Lelovas, P.P.; Kostomitsopoulos, N.G.; Xanthos, T.T. A comparative anatomic and physiologic overview of the porcine heart. J. Am. Assoc. Lab. Anim. Sci. 2014, 53, 432–438. [Google Scholar] [PubMed]
- Kehat, I.; Khimovich, L.; Caspi, O.; Gepstein, A.; Shofti, R.; Arbel, G.; Huber, I.; Satin, J.; Itskovitz-Eldor, J.; Gepstein, L. Electromechanical integration of cardiomyocytes derived from human embryonic stem cells. Nat. Biotechnol. 2004, 22, 1282–1289. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Chang, Y.H.; Xiong, Q.; Zhang, P.; Zhang, L.; Somasundaram, P.; Lepley, M.; Swingen, C.; Su, L.; Wendel, J.S.; et al. Cardiac repair in a porcine model of acute myocardial infarction with human induced pluripotent stem cell-derived cardiovascular cells. Cell Stem Cell 2014, 15, 750–761. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, M.; Miyagawa, S.; Fukushima, S.; Saito, A.; Miki, K.; Ito, E.; Sougawa, N.; Kawamura, T.; Daimon, T.; Shimizu, T.; et al. Enhanced survival of transplanted human induced pluripotent stem cell-derived cardiomyocytes by the combination of cell sheets with the pedicled omental flap technique in a porcine heart. Circulation 2013, 128, S87–S94. [Google Scholar] [CrossRef]
- Kawamura, M.; Miyagawa, S.; Fukushima, S.; Saito, A.; Miki, K.; Funakoshi, S.; Yoshida, Y.; Yamanaka, S.; Shimizu, T.; Okano, T.; et al. Enhanced Therapeutic Effects of Human iPS Cell Derived-Cardiomyocyte by Combined Cell-Sheets with Omental Flap Technique in Porcine Ischemic Cardiomyopathy Model. Sci. Rep. 2017, 7, 8824. [Google Scholar] [CrossRef]
- Ishigami, M.; Masumoto, H.; Ikuno, T.; Aoki, T.; Kawatou, M.; Minakata, K.; Ikeda, T.; Sakata, R.; Yamashita, J.K.; Minatoya, K. Human iPS cell-derived cardiac tissue sheets for functional restoration of infarcted porcine hearts. PLoS ONE 2018, 13, e0201650. [Google Scholar] [CrossRef]
- Gao, L.; Gregorich, Z.R.; Zhu, W.; Mattapally, S.; Oduk, Y.; Lou, X.; Kannappan, R.; Borovjagin, A.V.; Walcott, G.P.; Pollard, A.E.; et al. Large Cardiac Muscle Patches Engineered from Human Induced-Pluripotent Stem Cell-Derived Cardiac Cells Improve Recovery from Myocardial Infarction in Swine. Circulation 2018, 137, 1712–1730. [Google Scholar] [CrossRef]
- Bontrop, R.E. Non-human primates: Essential partners in biomedical research. Immunol. Rev. 2001, 183, 5–9. [Google Scholar] [CrossRef]
- Ishigaki, H.; Shiina, T.; Ogasawara, K. MHC-identical and transgenic cynomolgus macaques for preclinical studies. Inflamm. Regen. 2018, 38, 30. [Google Scholar] [CrossRef]
- Deleidi, M.; Hargus, G.; Hallett, P.; Osborn, T.; Isacson, O. Development of histocompatible primate-induced pluripotent stem cells for neural transplantation. Stem Cells 2011, 29, 1052–1063. [Google Scholar] [CrossRef]
- Shiba, Y.; Gomibuchi, T.; Seto, T.; Wada, Y.; Ichimura, H.; Tanaka, Y.; Ogasawara, T.; Okada, K.; Shiba, N.; Sakamoto, K.; et al. Allogeneic transplantation of iPS cell-derived cardiomyocytes regenerates primate hearts. Nature 2016, 538, 388–391. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, T.; Miyagawa, S.; Fukushima, S.; Maeda, A.; Kashiyama, N.; Kawamura, A.; Miki, K.; Okita, K.; Yoshida, Y.; Shiina, T.; et al. Cardiomyocytes Derived from MHC-Homozygous Induced Pluripotent Stem Cells Exhibit Reduced Allogeneic Immunogenicity in MHC-Matched Non-human Primates. Stem Cell Rep. 2016, 6, 312–320. [Google Scholar] [CrossRef] [PubMed]
- Chong, J.J.; Yang, X.; Don, C.W.; Minami, E.; Liu, Y.W.; Weyers, J.J.; Mahoney, W.M.; Van Biber, B.; Cook, S.M.; Palpant, N.J.; et al. Human embryonic-stem-cell-derived cardiomyocytes regenerate non-human primate hearts. Nature 2014, 510, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Menasche, P.; Vanneaux, V.; Hagege, A.; Bel, A.; Cholley, B.; Cacciapuoti, I.; Parouchev, A.; Benhamouda, N.; Tachdjian, G.; Tosca, L.; et al. Human embryonic stem cell-derived cardiac progenitors for severe heart failure treatment: First clinical case report. Eur. Heart J. 2015, 36, 2011–2017. [Google Scholar] [CrossRef] [PubMed]
- Menasche, P.; Vanneaux, V.; Hagege, A.; Bel, A.; Cholley, B.; Parouchev, A.; Cacciapuoti, I.; Al-Daccak, R.; Benhamouda, N.; Blons, H.; et al. Transplantation of Human Embryonic Stem Cell-Derived Cardiovascular Progenitors for Severe Ischemic Left Ventricular Dysfunction. J. Am. Coll. Cardiol. 2018, 71, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Faculty of Medicine, Osaka University. iPS Cell-Based Therapy for Heart Disease: Clinical Application iPS Cell-Derived Cardiomyocytes. Available online: http://www.med.osaka-u.ac.jp/eng/archives/2777 (accessed on 1 June 2019).
| PSC-Derived CM | Mature/Adult CM |
|---|---|
| Smaller in size, roundish in shape | Larger in size, elongated in shape |
| Disorganized sarcomeres | Organized sarcomeres |
| Slow/skeletal troponin I (TnIs) | Adult cardiac troponin I (TnIc) |
| Titin N2BA isoform | Titin N2B isoform |
| Higher αMHC:βMHC | Lower αMHC:βMHC |
| Poor expression of ion-transport components genes (e.g., KCNJ2, RYR2) | High expression of ion-transport components genes |
| Less efficient Calcium handling | Improved Calcium handling |
| Less negative resting membrane potential | More negative resting membrane potential |
| No or few T-tubules | Abundant T-tubules |
| Few, underdeveloped mitochondria Glucose as major energy source | Dense, well-distributed and developed mitochondria Fatty acids as major energy source |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samak, M.; Hinkel, R. Stem Cells in Cardiovascular Medicine: Historical Overview and Future Prospects. Cells 2019, 8, 1530. https://doi.org/10.3390/cells8121530
Samak M, Hinkel R. Stem Cells in Cardiovascular Medicine: Historical Overview and Future Prospects. Cells. 2019; 8(12):1530. https://doi.org/10.3390/cells8121530
Chicago/Turabian StyleSamak, Mostafa, and Rabea Hinkel. 2019. "Stem Cells in Cardiovascular Medicine: Historical Overview and Future Prospects" Cells 8, no. 12: 1530. https://doi.org/10.3390/cells8121530
APA StyleSamak, M., & Hinkel, R. (2019). Stem Cells in Cardiovascular Medicine: Historical Overview and Future Prospects. Cells, 8(12), 1530. https://doi.org/10.3390/cells8121530