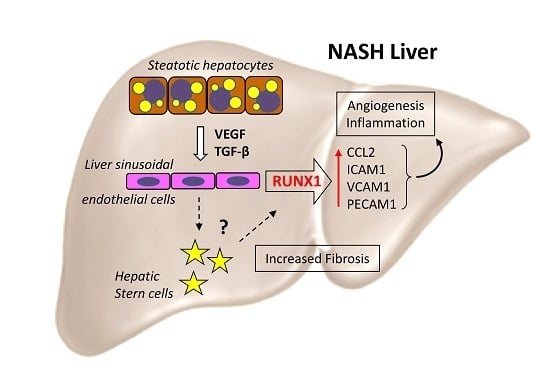
Increased Expression of RUNX1 in Liver Correlates with NASH Activity Score in Patients with Non-Alcoholic Steatohepatitis (NASH)
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects and Collection of Samples
2.2. Differential Gene Expression Studies and qRT-PCRs
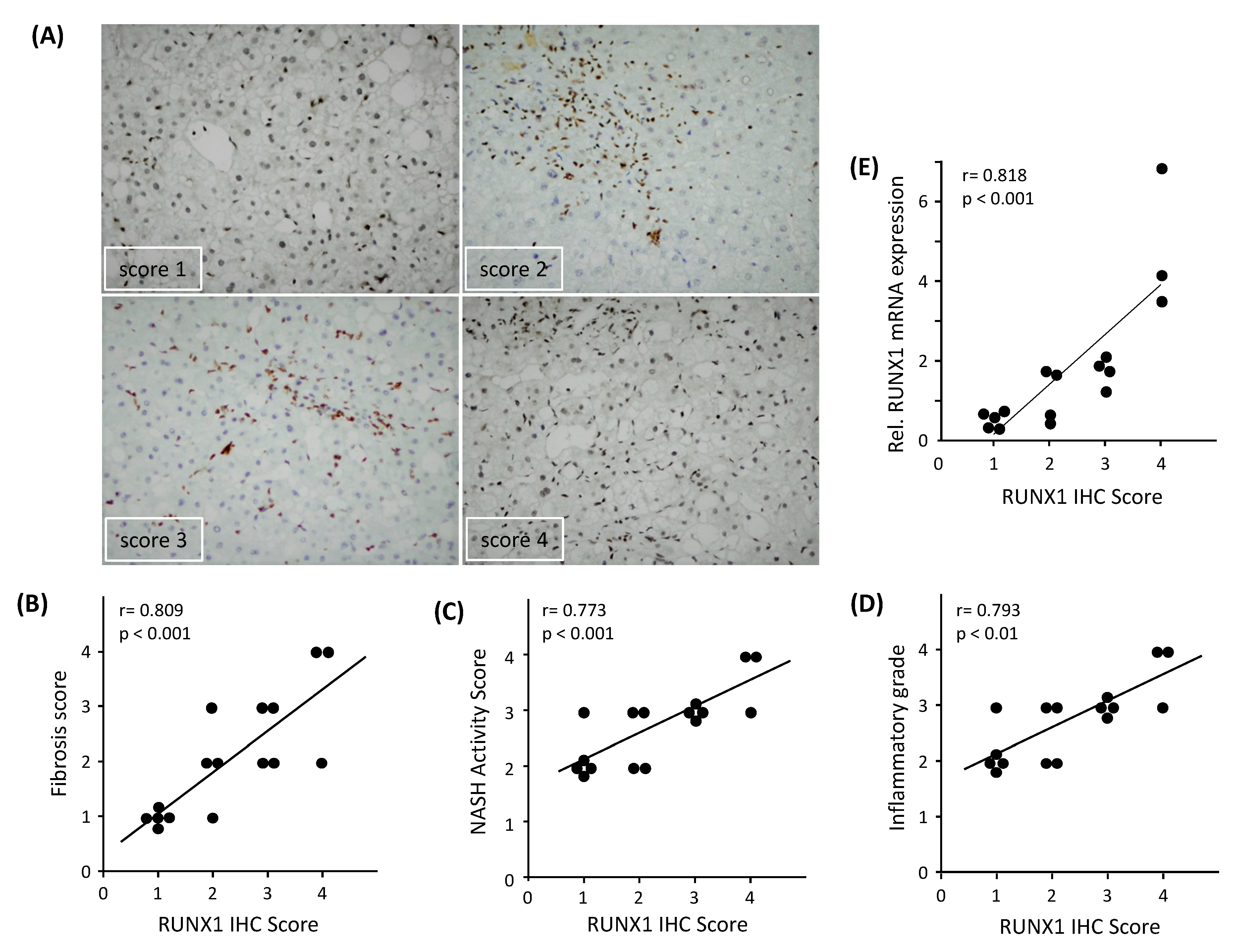
2.3. Immunohistochemistry Analysis
2.4. Culture of Endothelial Cells with Conditioned Medium from Hepatoma Cells Treated with Palmitic Acid
2.5. Induction of RUNX1 Expression in HUVECs
2.6. RUNX1 Inhibition and Overexpression in HUVECs
2.7. Flow Cytometry Analysis
2.8. Statistical Analysis
3. Results
3.1. Expression of Transcription Factor RUNX1 is Increased in NAFLD Controlling Differentially Expressed Genes (Degs) Associated with Angiogenesis
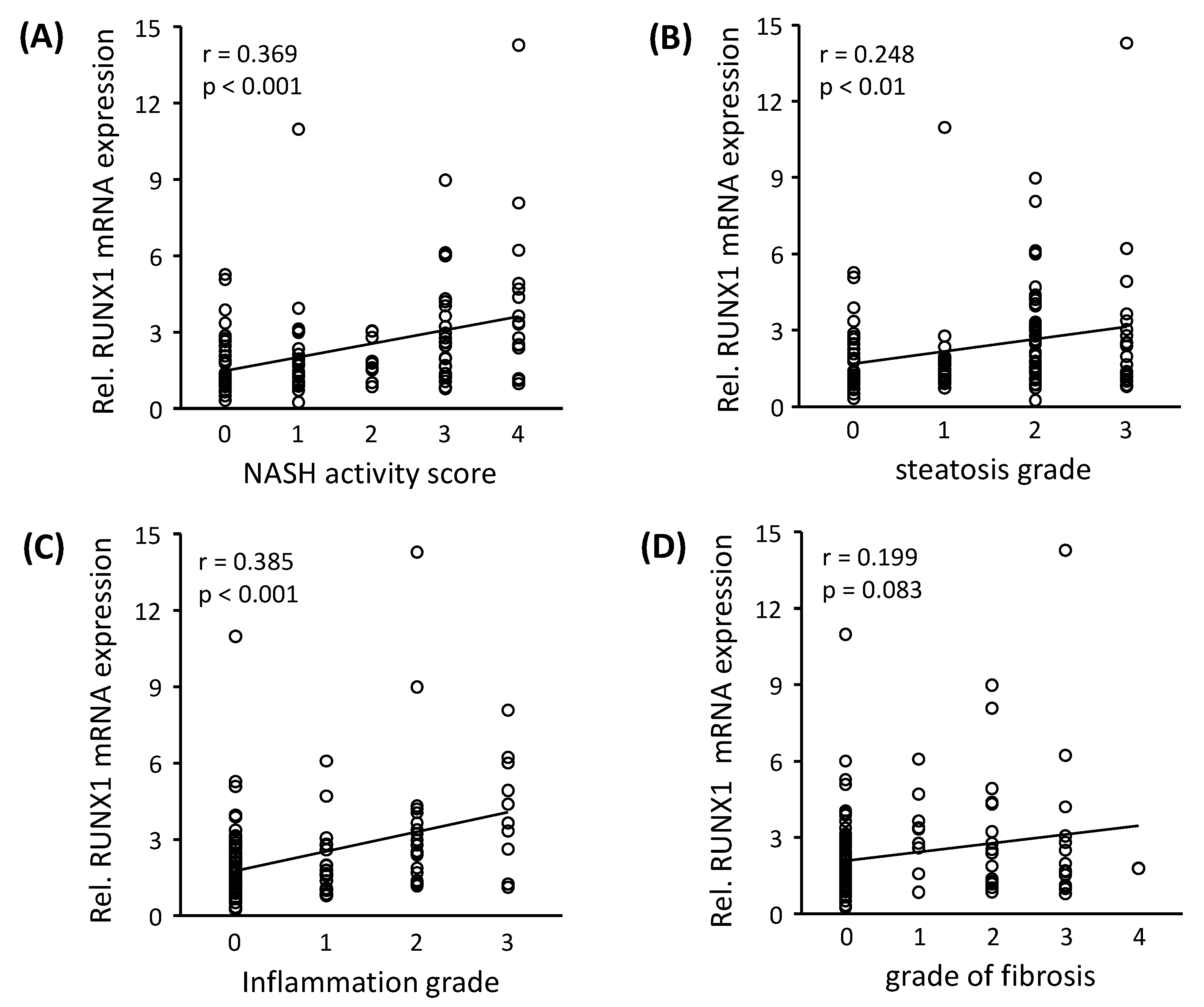
3.2. RUNX1 Expression Correlates with the Severity of NAFLD
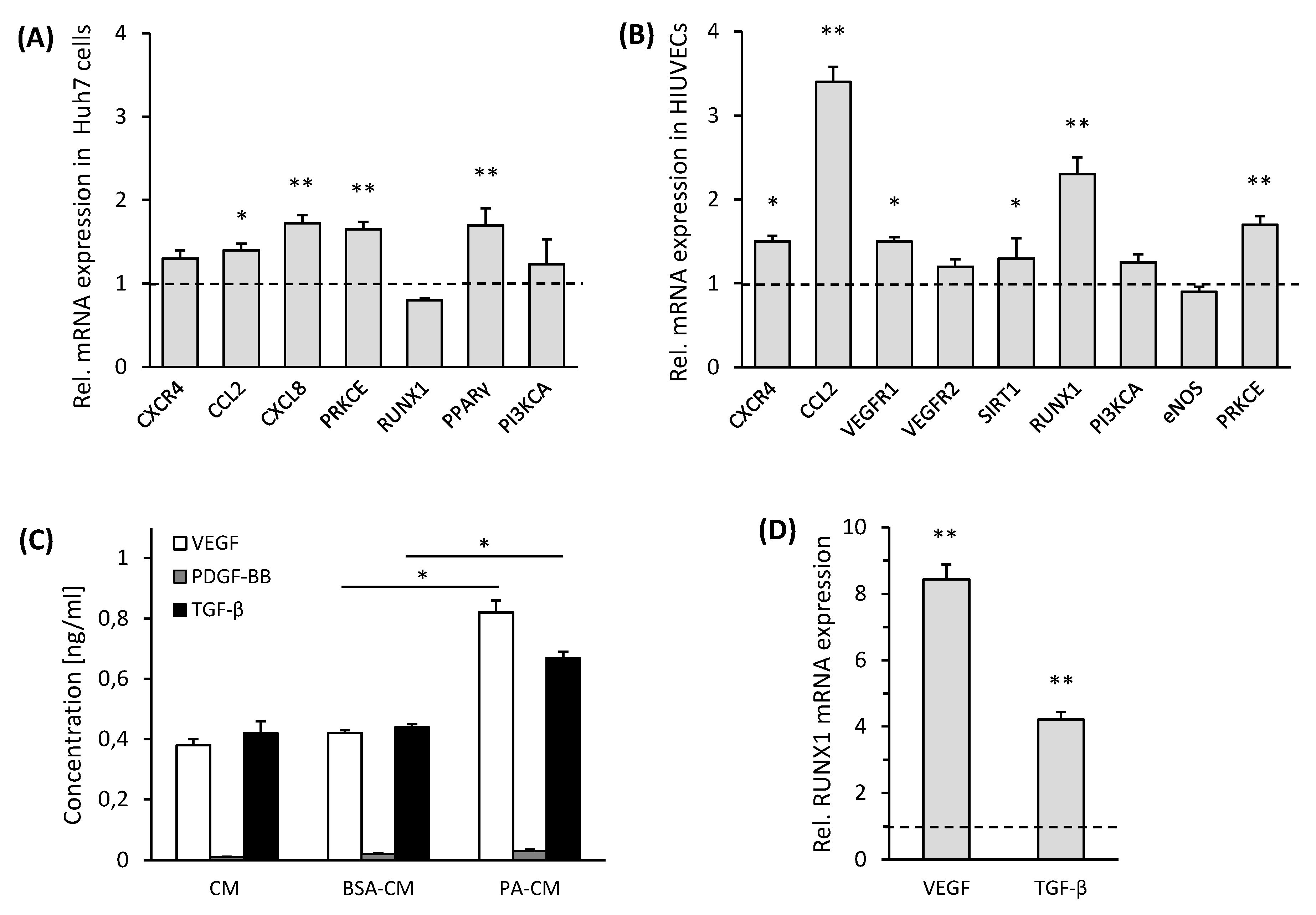
3.3. Palmitic Acid Treated Huh7 Cells Release VEGF and TGF-β-Inducing RUNX1 Gene Expression in ECs
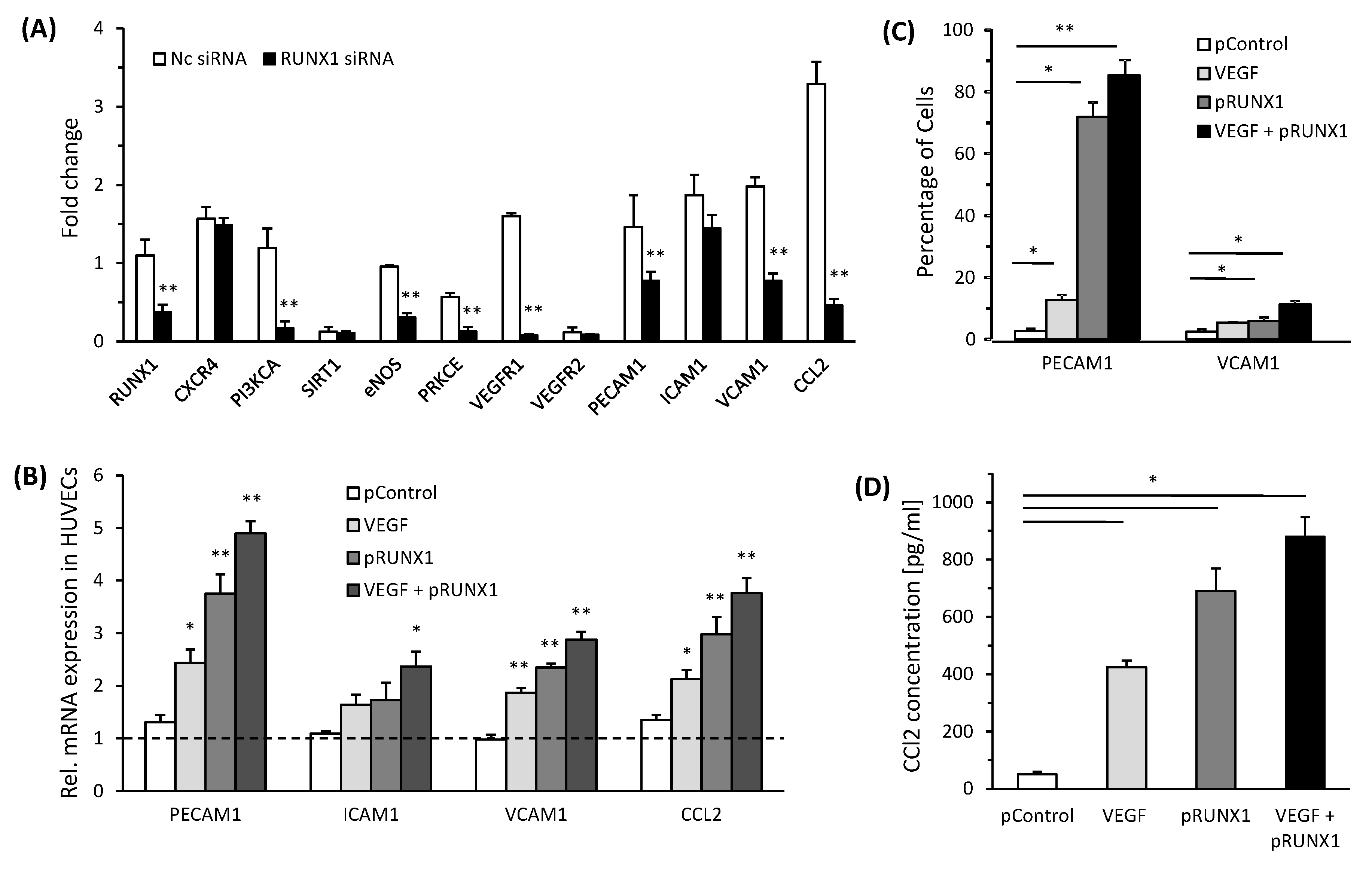
3.4. RUNX1 Enhances Expression of Angiogenic Markers and Adhesion Molecules in HUVECs
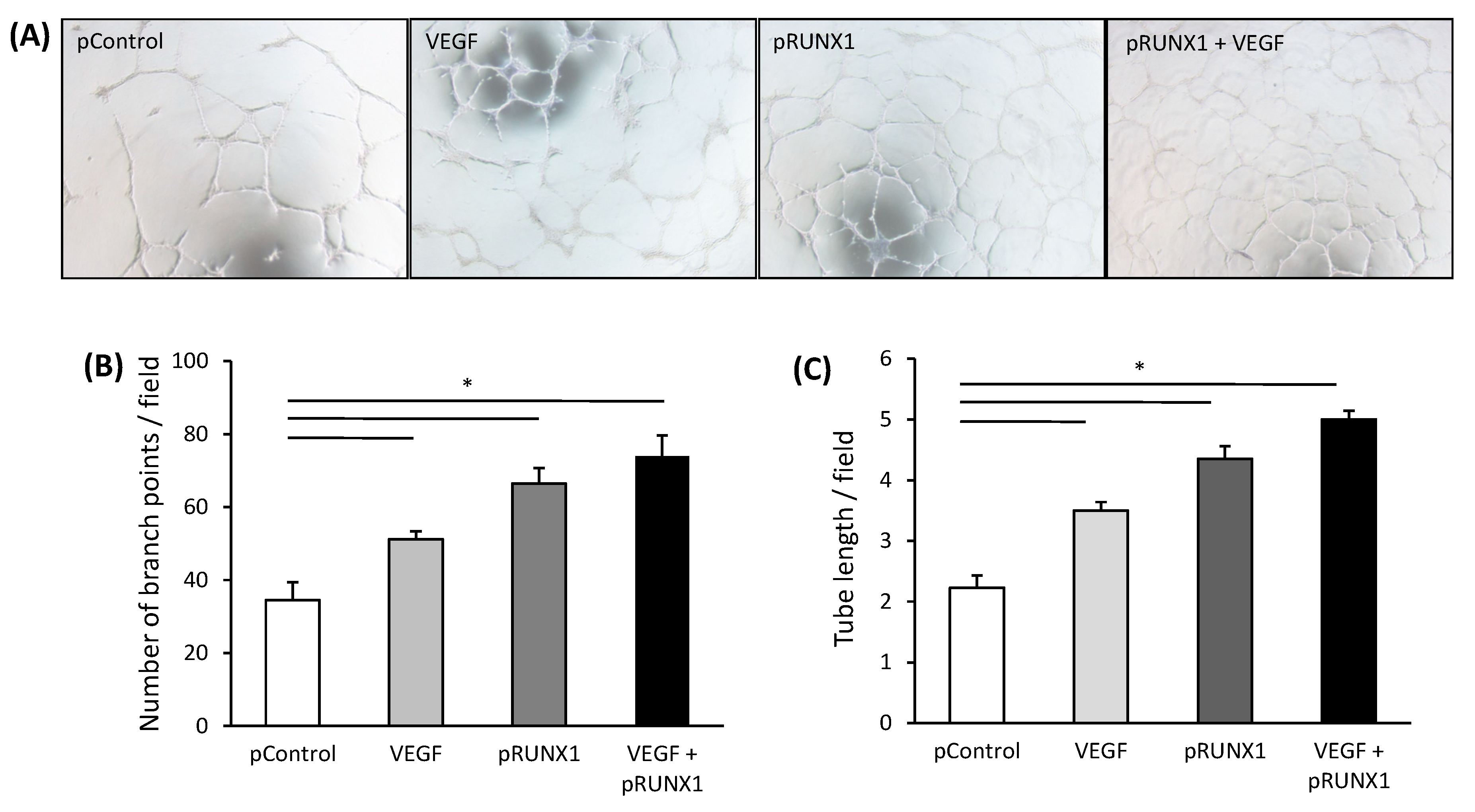
3.5. RUNX1 Increases Angiogenic Activity of HUVECs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Younossi, Z.; Anstee, Q.M.; Marietti, M.; Hardy, T.; Henry, L.; Eslam, M.; George, J.; Bugianesi, E. Global burden of NAFLD and NASH: Trends, predictions, risk factors and prevention. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J.C.; Horton, J.D.; Hobbs, H.H. Human fatty liver disease: old questions and new insights. Science 2011, 332, 1519–1523. [Google Scholar] [CrossRef] [PubMed]
- Fernández, M.; Semela, D.; Bruix, J.; Colle, I.; Pinzani, M.; Bosch, J. Angiogenesis in liver disease. J. Hepatol. 2009, 50, 604–620. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.; Anita, K. Angiogenesis in liver regeneration and fibrosis: “a double-edged sword”. Hepatol. Int. 2013, 7, 959–968. [Google Scholar] [CrossRef]
- Coulon, S.; Francque, S.; Colle, I.; Verrijken, A.; Blomme, B.; Heindryckx, F.; De Munter, S.; Prawitt, J.; Caron, S.; Staels, B.; et al. Evaluation of inflammatory and angiogenic factors in patients with non-alcoholic fatty liver disease. Cytokine 2012, 59, 442–449. [Google Scholar] [CrossRef]
- Coulon, S.; Legry, V.; Heindryckx, F.; Van Steenkiste, C.; Casteleyn, C.; Olievier, K.; Libbrecht, L.; Carmeliet, P.; Jonckx, B.; Stassen, J.-M.; et al. Role of vascular endothelial growth factor in the pathophysiology of nonalcoholic steatohepatitis in two rodent models. Hepatology 2013, 57, 1793–1805. [Google Scholar] [CrossRef]
- Ciupińska-Kajor, M.; Hartleb, M.; Kajor, M.; Kukla, M.; Wyleżoł, M.; Lange, D.; Liszka, L. Hepatic angiogenesis and fibrosis are common features in morbidly obese patients. Hepatol. Int. 2013, 7, 233–240. [Google Scholar] [CrossRef]
- North, T.E.; De Bruijn, M.F.T.R.; Stacy, T.; Talebian, L.; Lind, E.; Robin, C.; Binder, M.; Dzierzak, E.; A Speck, N. Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo. Immunity 2002, 16, 661–672. [Google Scholar] [CrossRef]
- Iwatsuki, K.; Tanaka, K.; Kaneko, T.; Kazama, R.; Okamoto, S.; Nakayama, Y.; Ito, Y.; Satake, M.; Takahashi, S.; et al. Runx1 promotes angiogenesis by downregulation of insulin-like growth factor-binding protein-3. Oncogene 2005, 24, 1129–1137. [Google Scholar] [CrossRef]
- Takakura, N.; Watanabe, T.; Suenobu, S.; Yamada, Y.; Noda, T.; Ito, Y.; Satake, M.; Suda, T. A Role for Hematopoietic Stem Cells in Promoting Angiogenesis. Cell 2000, 102, 199–209. [Google Scholar] [CrossRef]
- Weiss, T.S.; Lupke, M.; Ibrahim, S.; Buechler, C.; Lorenz, J.; Ruemmele, P.; Hofmann, U.; Melter, M.; Dayoub, R. Attenuated lipotoxicity and apoptosis is linked to exogenous and endogenous augmenter of liver regeneration by different pathways. PLOS ONE 2017, 12, e0184282. [Google Scholar] [CrossRef] [PubMed]
- Kleiner, D.E.; Brunt, E.M.; Natta, V.M.; Behling, C.; Contos, M.J.; Cummings, O.W.; Ferrell, L.D.; Liu, Y.C.; Torbenson, M.S.; Unalp-Arida, A.; et al. Nonalcoholic Steatohepatitis Clinical Research Network. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 2005, 41, 1313–1321. [Google Scholar] [CrossRef] [PubMed]
- Rana, P.; Yi, L.; Will, Y. Palmitate Increases the Susceptibility of Cells to Drug-Induced Toxicity: An In Vitro Method to Identify Drugs With Potential Contraindications in Patients With Metabolic Disease. Toxicol. Sci. 2012, 129, 346–362. [Google Scholar]
- Marcher, A.-B.; Bendixen, S.M.; Terkelsen, M.K.; Hohmann, S.S.; Hansen, M.H.; Larsen, B.D.; Mandrup, S.; Dimke, H.; Detlefsen, S.; Ravnskjaer, K. Transcriptional regulation of Hepatic Stellate Cell activation in NASH. Sci. Rep. 2019, 9, 2324. [Google Scholar] [CrossRef]
- Miyoshi, H.; Ohira, M.; Shimizu, K.; Mitani, K.; Hirai, H.; Imai, T.; Yokoyama, K.; Soceda, E.; Ohkl, M. Alternative splicing and genomic structure of theAML1gene involved in acute myeloid leukemia. Nucleic Acids Res. 1995, 23, 2762–2769. [Google Scholar] [CrossRef]
- Bocca, C.; Novo, E.; Miglietta, A.; Parola, M. Angiogenesis and Fibrogenesis in Chronic Liver Diseases. Cell Mol. Gastroenterol. Hepatol. 2015, 1, 477–488. [Google Scholar] [CrossRef]
- Kitade, M.; Yoshiji, H.; Kojima, H.; Ikenaka, Y.; Noguchi, R.; Kaji, K.; Yoshii, J.; Yanase, K.; Namisaki, T.; Yamazaki, M.; et al. Neovascularization and oxidative stress in the progression of non-alcoholic steatohepatitis. Mol. Med. Rep. 2008, 1, 543–548. [Google Scholar] [CrossRef][Green Version]
- Wang, R.; Wang, X.; Zhuang, L. Gene expression profiling reveals key genes and pathways related to the development of non-alcoholic fatty liver disease. Ann. Hepatol. 2016, 15, 190–199. [Google Scholar]
- Tailleux, A.; Wouters, K.; Staels, B. Roles of PPARs in NAFLD: Potential therapeutic targets. Biochim. Et Biophys. Acta (BBA) Mol. Cell Boil. Lipids 2012, 1821, 809–818. [Google Scholar] [CrossRef]
- Dzierzak, E.; Speck, N.A. Of lineage and legacy: The development of mammalian hematopoietic stem cells. Nat. Immunol. 2008, 9, 129–136. [Google Scholar] [CrossRef]
- Lam, K.; Zhang, D.E. RUNX1 and RUNX1-ETO: Roles in hematopoiesis and leukemogenesis. Front. Biosci (Landmark Ed.) 2012, 17, 1120–1139. [Google Scholar] [CrossRef] [PubMed]
- Blyth, K.; Cameron, E.R.; Neil, J.C. The runx genes: Gain or loss of function in cancer. Nat. Rev. Cancer 2005, 5, 376–387. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.-C.; Zhou, S.-Y.; Feng, D.-Y.; Xiao, J.; Li, W.-Y.; Xu, C.-D.; Wang, H.-Y.; Zhou, T. Runt-related Transcription Factor 1 (RUNX1) Binds to p50 in Macrophages and Enhances TLR4-triggered Inflammation and Septic Shock. J. Boil. Chem. 2016, 291, 22011–22020. [Google Scholar] [CrossRef] [PubMed]
- Lam, J.D.; Oh, D.J.; Wong, L.L.; Amarnani, D.; Park-Windhol, C.; Sanchez, A.V.; Cardona-Velez, J.; McGuone, D.; Stemmer-Rachamimov, A.O.; Eliott, D.; et al. Identification of RUNX1 as a Mediator of Aberrant Retinal Angiogenesis. Diabetes 2017, 66, 1950–1956. [Google Scholar] [CrossRef] [PubMed]
- Wildey, G.M.; Howe, P.H. Runx1 is a co-activator with FOXO3 to mediate transforming growth factor beta (TGF β)-induced Bim transcription in hepatic cells. J. Biol. Chem. 2009, 284, 20227–20239. [Google Scholar] [CrossRef] [PubMed]
- Bertrand-Philippe, M.; Ruddell, R.G.; Arthur, M.J.P.; Thomas, J.; Mungalsingh, N.; Mann, D.A. Regulation of Tissue Inhibitor of Metalloproteinase 1 Gene Transcription by RUNX1 and RUNX2. J. Boil. Chem. 2004, 279, 24530–24539. [Google Scholar] [CrossRef] [PubMed]
- Cheah, I.K.; Tang, R.; Ye, P.; Yew, T.S.; Lim, K.H.; Halliwell, B. Liver ergothioneine accumulation in a guinea pig model of non-alcoholic fatty liver disease. A possible mechanism of defence? Free Radic. Res. 2016, 50, 14–25. [Google Scholar] [CrossRef]
- Pimanda, J.E.; Donaldson, I.J.; de Bruijn, M.F.; Kinston, S.; Knezevic, K.; Huckle, L.; Piltz, S.; Landry, J.R.; Green, A.R.; Tannahill, D.; et al. The SCL transcriptional network and BMP signaling pathway interact to regulate RUNX1 activity. Proc. Natl. Acad. Sci. USA 2007, 104, 840–845. [Google Scholar] [CrossRef]
- Zhou, T.; Luo, M.; Cai, W.; Zhou, S.; Feng, D.; Xu, C.; Wang, H. Runt-Related Transcription Factor 1 (RUNX1) Promotes TGF-β-Induced Renal Tubular Epithelial-to-Mesenchymal Transition (EMT) and Renal Fibrosis through the PI3K Subunit p110δ. EBioMedicine 2018, 31, 217–225. [Google Scholar] [CrossRef]
- Karar, J.; Maity, A. PI3K/AKT/mTOR Pathway in Angiogenesis. Front. Mol. Neurosci. 2011, 4, 51. [Google Scholar] [CrossRef]
- Zhang, W.; Liu, Y.; Wu, M.; Zhu, X.; Wang, T.; He, K.; Li, P.; Wu, X. PI3K inhibition protects mice from NAFLD by down-regulating CMKLR1 and NLRP3 in Kupffer cells. J. Physiol. Biochem. 2017, 73, 583–594. [Google Scholar] [CrossRef] [PubMed]
- Jornayvaz, F.R.; Shulman, G.I. Diacylglycerol activation of protein kinase Cε and hepatic insulin resistance. Cell Metab. 2012, 15, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Kumashiro, N.; Erion, D.M.; Zhang, D.; Kahn, M.; Beddow, S.A.; Chu, X.; Still, C.D.; Gerhard, G.S.; Han, X.; Dziura, J.; et al. Cellular mechanism of insulin resistance in nonalcoholic fatty liver disease. Proc. Natl. Acad. Sci. USA 2011, 108, 16381–16385. [Google Scholar] [CrossRef] [PubMed]
- Ter Horst, K.W.; Gilijamse, P.W.; Versteeg, R.I.; Ackermans, M.T.; Nederveen, A.J.; La Fleur, S.E.; Romijn, J.A.; Nieuwdorp, M.; Zhang, D.; Samuel, V.T.; et al. Hepatic Diacylglycerol-Associated Protein Kinase Cε Translocation Links Hepatic Steatosis to Hepatic Insulin Resistance in Humans. Cell Rep. 2017, 19, 1997–2004. [Google Scholar] [CrossRef] [PubMed]
- Ehling, J.; Bartneck, M.; Wei, X.; Gremse, F.; Fech, V.; Möckel, D.; Baeck, C.; Hittatiya, K.; Eulberg, D.; Luedde, T.; et al. CCL2-dependent infiltrating macrophages promote angiogenesis in progressive liver fibrosis. Gut 2014, 63, 1960–1971. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, M. ERK/MAPK-dependent PI3K/Akt phosphorylation through VEGFR-1 after VEGF stimulation in activated hepatic stellate cells. Hepatol. Res. 2003, 26, 232–236. [Google Scholar] [CrossRef]
| Relative mRNA Expression | p-Value | |||||
|---|---|---|---|---|---|---|
| Gene | N (n = 33) | S (n = 46) | NASH (n = 43) | N/S | N/SH | S/SH |
| CCL2 | 1.82 ± 1.47 | 2.30 ± 1.63 | 3.74 ± 2.13 | 0.534 | 0.000 | 0.002 |
| CXCL8 (IL8) | 4.33 ± 4.5 | 7.16 ± 8.99 | 16.97 ± 17.18 | 0.404 | 0.000 | 0.002 |
| CXCR4 | 1.57 ± 1.18 | 2.63 ± 1.70 | 2.87 ± 1.84 | 0.008 | 0.002 | 1.000 |
| EREG | 2.87 ± 2.26 | 5.34 ± 4.86 | 7.48 ± 6.19 | 0.197 | 0.001 | 0.285 |
| FASN | 3.27 ± 2.07 | 6.35 ± 5.75 | 5.67 ± 5.27 | 0.048 | 0.077 | 1.000 |
| HOMX1 (HO1) | 0.71 ± 0.36 | 0.67 ± 0.39 | 0.95 ± 0.66 | 1.000 | 0.126 | 0.024 |
| NOS3 (eNOS) | 0.72 ± 0.32 | 0.82 ± 0.52 | 1.12 ± 0.74 | 1.000 | 0.011 | 0.040 |
| PIK3CA | 0.61 ± 0.21 | 0.98 ± 1.22 | 0.81 ± 0.25 | 0.013 | 0.001 | 1.000 |
| PPARγ | 0.61 ± 0.24 | 0.73 ± 0.34 | 0.97 ± 0.47 | 0.552 | 0.001 | 0.034 |
| PRKCE | 0.71 ± 0.34 | 0.96 ± 0.42 | 1.00 ± 0.60 | 0.004 | 0.014 | 1.000 |
| PROK2 | 2.55 ± 2.17 | 3.66 ± 4.13 | 6.49 ± 7.96 | 1.000 | 0.058 | 0.182 |
| RUNX1 | 1.75 ± 1.21 | 1.90 ± 1.56 | 3.38 ± 2.55 | 1.000 | 0.000 | 0.002 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaur, S.; Rawal, P.; Siddiqui, H.; Rohilla, S.; Sharma, S.; Tripathi, D.M.; Baweja, S.; Hassan, M.; Vlaic, S.; Guthke, R.; et al. Increased Expression of RUNX1 in Liver Correlates with NASH Activity Score in Patients with Non-Alcoholic Steatohepatitis (NASH). Cells 2019, 8, 1277. https://doi.org/10.3390/cells8101277
Kaur S, Rawal P, Siddiqui H, Rohilla S, Sharma S, Tripathi DM, Baweja S, Hassan M, Vlaic S, Guthke R, et al. Increased Expression of RUNX1 in Liver Correlates with NASH Activity Score in Patients with Non-Alcoholic Steatohepatitis (NASH). Cells. 2019; 8(10):1277. https://doi.org/10.3390/cells8101277
Chicago/Turabian StyleKaur, Savneet, Preety Rawal, Hamda Siddiqui, Sumati Rohilla, Shvetank Sharma, Dinesh M Tripathi, Sukriti Baweja, Mohsin Hassan, Sebastian Vlaic, Reinhard Guthke, and et al. 2019. "Increased Expression of RUNX1 in Liver Correlates with NASH Activity Score in Patients with Non-Alcoholic Steatohepatitis (NASH)" Cells 8, no. 10: 1277. https://doi.org/10.3390/cells8101277
APA StyleKaur, S., Rawal, P., Siddiqui, H., Rohilla, S., Sharma, S., Tripathi, D. M., Baweja, S., Hassan, M., Vlaic, S., Guthke, R., Thomas, M., Dayoub, R., Bihari, C., Sarin, S. K., & Weiss, T. S. (2019). Increased Expression of RUNX1 in Liver Correlates with NASH Activity Score in Patients with Non-Alcoholic Steatohepatitis (NASH). Cells, 8(10), 1277. https://doi.org/10.3390/cells8101277