MicroRNAs in Cardiac Autophagy: Small Molecules and Big Role
Abstract
1. Overview of Autophagy and MicroRNAs
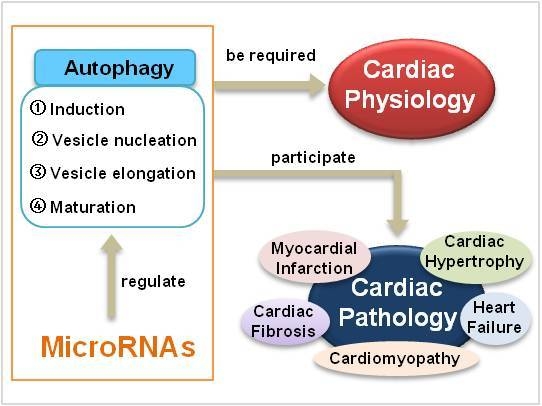
1.1. Autophagy
1.2. MicroRNAs
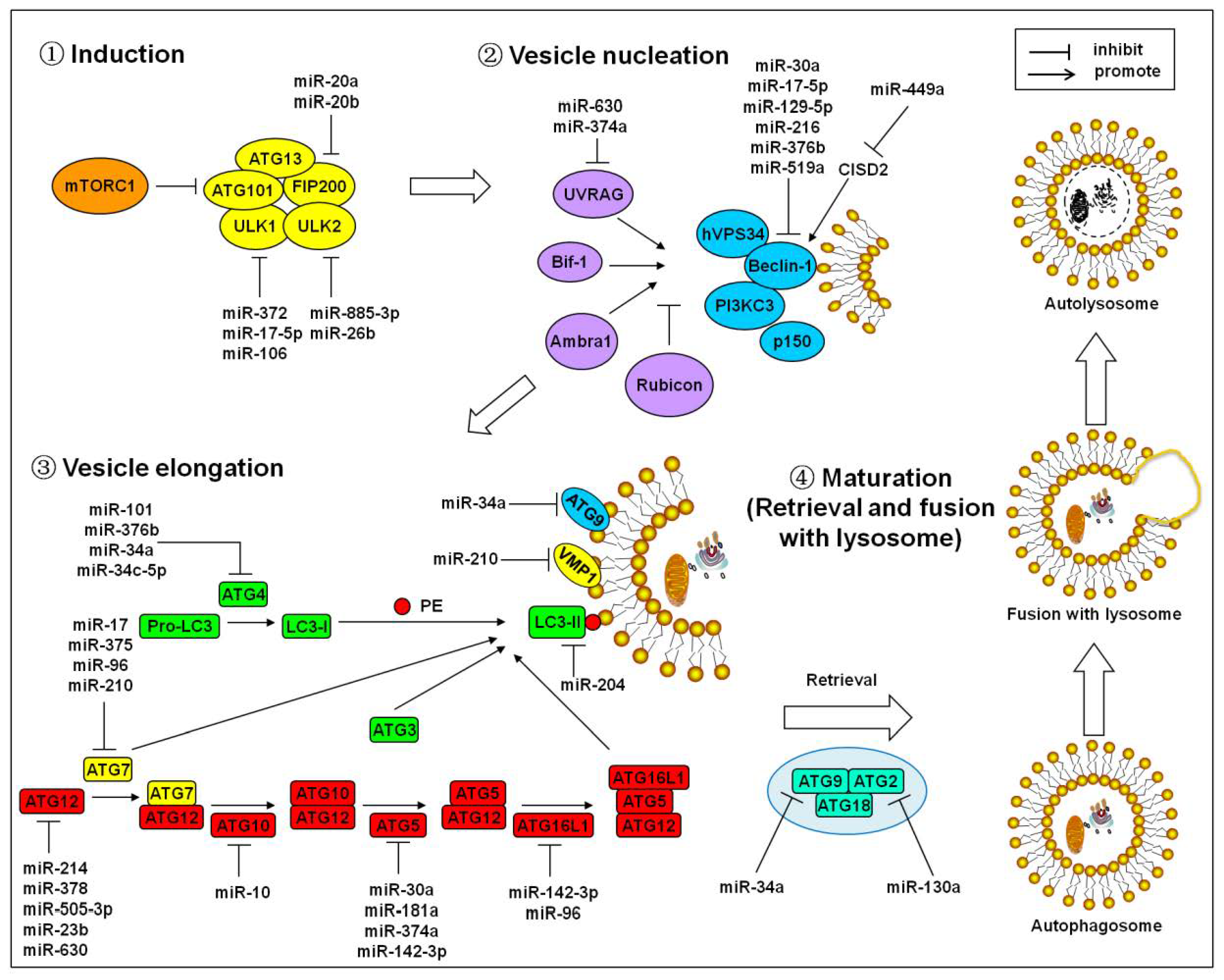
2. MicroRNAs Regulate the Core Autophagy Signaling Cascades
3. The Role of MicroRNAs in Cardiac Autophagy
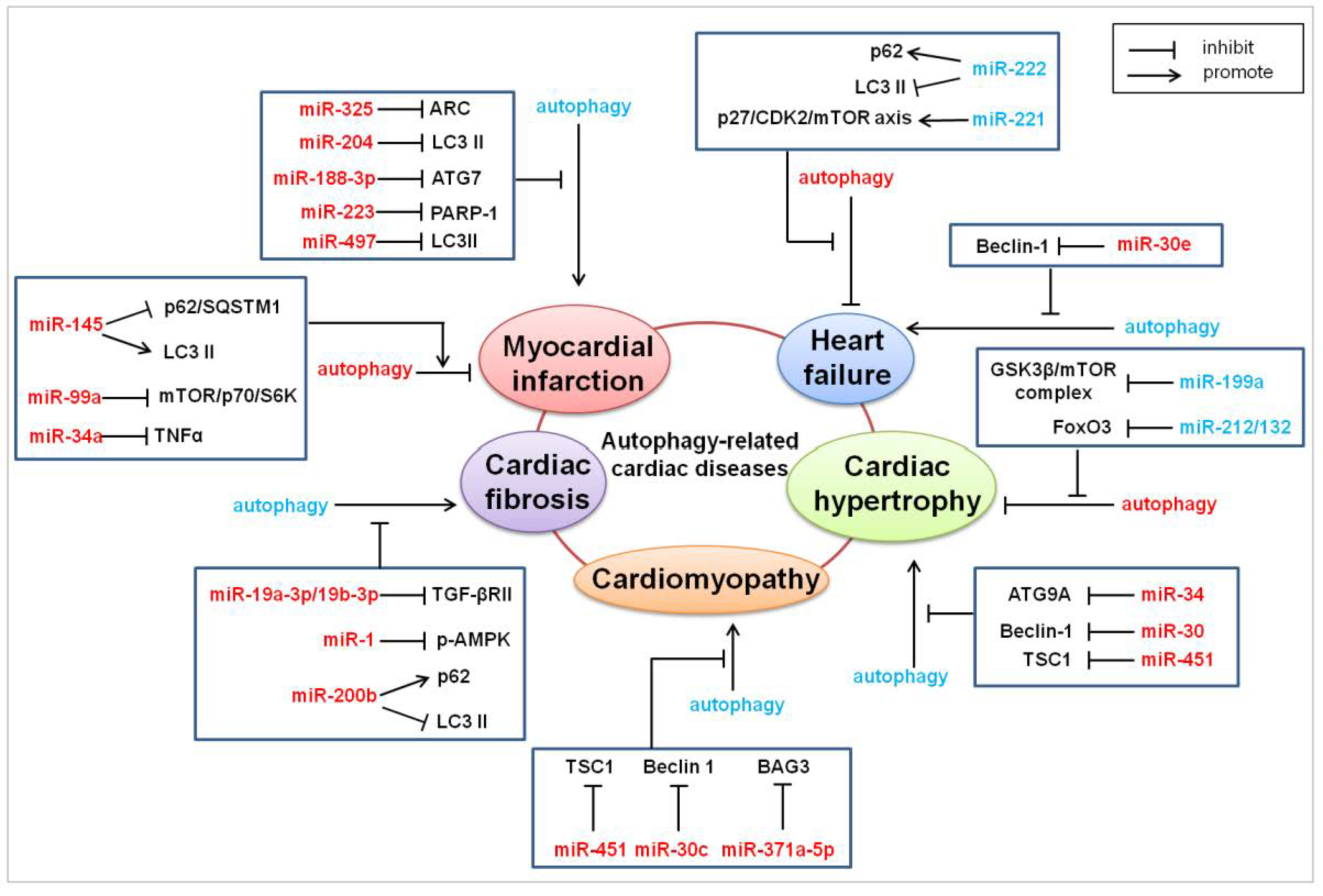
4. MicroRNAs in Autophagy-Related Heart Diseases
4.1. Myocardial Infarction
4.2. Cardiac Hypertrophy
4.3. Cardiac Fibrosis
4.4. Cardiomyopathy
4.5. Heart Failure
5. Conclusions and Perspectives
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| ACE2 | angiotensin-converting enzyme 2 |
| AGOs | argonaute proteins |
| AMPK | AMP- activated protein kinase |
| ARC | apoptosis repressor with caspase recruit domain |
| ATG | autophagy-related gene |
| BAG3 | Bcl-2-associated athanogene 3 |
| Bcl2-L-13 | Bcl2-like protein 13 |
| BNIP3 | Bcl-2 interacting protein 3 |
| BNIP3L | Bcl2 interacting protein 3 like |
| CDS | coding sequence |
| CISD2 | CDGSH iron sulfur domain 2 |
| FIP200 | focal adhesion kinase family interacting protein of 200 kDa |
| FOXO3 | forkhead box O3 |
| FUNDC1 | FUN14 domain containing protein 13 |
| GSK3β | glycogen synthase kinase 3β |
| HCM | hypertrophic cardiomyopathy |
| MFN2 | mitofusion2 |
| miRNAs | microRNAs |
| mTOR | mammalian target of rapamycin |
| mTORC1 | mTOR complex 1 |
| PARP-1 | poly (ADP-ribose) polymerase 1 |
| PE | phosphatidylethanolamine |
| PIK3C3 | phosphatidylinositol 3-kinase catalytic subunit type 3 |
| PINK1 | putative kinase 1 |
| PTEN | phosphatase and tensin homolog deleted on chromosome ten |
| TSC1 | tuberous sclerosis complex 1 |
| TTC | Takatsubo cardiomyopathy |
| ULK | Unc-51-like kinase |
| UTR | untranslated region |
| UVRAG | UV irradiation resistance-associated gene |
References
- He, C.; Klionsky, D.J. Regulation mechanisms and signaling pathways of autophagy. Annu. Rev. Genet. 2009, 43, 67–93. [Google Scholar] [CrossRef] [PubMed]
- Legakis, J.E.; Yen, W.L.; Klionsky, D.J. A cycling protein complex required for selective autophagy. Autophagy 2007, 3, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Kiriyama, Y.; Nochi, H. Intra- and intercellular quality control mechanisms of mitochondria. Cells 2017, 7, 1. [Google Scholar] [CrossRef] [PubMed]
- Harper, J.W.; Ordureau, A.; Heo, J.M. Building and decoding ubiquitin chains for mitophagy. Nat. Rev. Mol. Cell Biol. 2018, 19, 93–108. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, R.; Pattison, J.S. Macroautophagy and chaperone-mediated autophagy in heart failure: The known and the unknown. Oxid. Med. Cell. Longev. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Dai, C.; Fan, Y.; Guo, B.; Ren, K.; Sun, T.; Wang, W. From autophagy to mitophagy: The roles of p62 in neurodegenerative diseases. J. Bioenerg. Biomembr. 2017, 49, 413–422. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Li, T.S. Dual role of mitophagy in cancer drug resistance. Anticancer Res. 2018, 38, 617–621. [Google Scholar] [PubMed]
- Herst, P.M.; Rowe, M.R.; Carson, G.M.; Berridge, M.V. Functional mitochondria in health and disease. Front. Endocrinol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Cummins, N.; Gotz, J. Shedding light on mitophagy in neurons: What is the evidence for pink1/parkin mitophagy in vivo? Cell. Mol. Life Sci. 2018, 75, 1151–1162. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liu, L.; Xue, Y.; Ma, Y.; Liu, X.; Li, Z.; Li, Z.; Liu, Y. Endothelial monocyte-activating polypeptide-ii induces bnip3-mediated mitophagy to enhance temozolomide cytotoxicity of glioma stem cells via down-regulating mir-24-3p. Front. Mol. Neurosci. 2018, 11, 92. [Google Scholar] [CrossRef] [PubMed]
- Murakawa, T.; Yamaguchi, O.; Hashimoto, A.; Hikoso, S.; Takeda, T.; Oka, T.; Yasui, H.; Ueda, H.; Akazawa, Y.; Nakayama, H.; et al. Bcl-2-like protein 13 is a mammalian atg32 homologue that mediates mitophagy and mitochondrial fragmentation. Nat. Commun. 2015, 6, 7527. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Zhu, P.; Wang, J.; Zhu, H.; Ren, J.; Chen, Y. Pathogenesis of cardiac ischemia reperfusion injury is associated with ck2alpha-disturbed mitochondrial homeostasis via suppression of fundc1-related mitophagy. Cell Death Differ. 2018, 25, 1080. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.Y.; Zhang, Y.H.; Li, R.B.; Zhou, L.Y.; An, T.; Zhang, R.C.; Zhai, M.; Huang, Y.; Yan, K.W.; Dong, Y.H.; et al. Lncrna caif inhibits autophagy and attenuates myocardial infarction by blocking p53-mediated myocardin transcription. Nat. Commun. 2018, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Song, Y.; Liu, L.; Hou, N.; An, X.; Zhan, D.; Li, Y. Mir-199a impairs autophagy and induces cardiac hypertrophy through mtor activation. Cell Death Differ. 2017, 24, 1205–1213. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, C.; Turdi, S.; Richmond, K.L.; Zhang, Y.; Ren, J. ALDH2 protects against high fat diet-induced obesity cardiomyopathy and defective autophagy: Role of CaM kinase II, histone H3K9 methyltransferase SUV39H, Sirt1, and PGC-1α deacetylation. Int. J. Obes. 2018, 42, 1073–1087. [Google Scholar] [CrossRef] [PubMed]
- Zou, M.; Wang, F.; Gao, R.; Wu, J.; Ou, Y.; Chen, X.; Wang, T.; Zhou, X.; Zhu, W.; Li, P.; et al. Autophagy inhibition of hsa-miR-19a-3p/19b-3p by targeting TGF-β R II during TGF-β1-induced fibrogenesis in human cardiac fibroblasts. Sci. Rep. 2016, 6, 24747. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Wang, J.; Wang, C.; Wang, X.; Dong, W.; Qiu, W.; Wang, Y.; Zhao, X.; Zou, Y.; Song, L.; et al. MicroRNA-221 inhibits autophagy and promotes heart failure by modulating the p27/CDK2/mTOR axis. Cell Death Differ. 2015, 22, 986–999. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Liu, C.Y.; Zhou, L.Y.; Wang, J.X.; Wang, M.; Zhao, B.; Zhao, W.K.; Xu, S.J.; Fan, L.H.; Zhang, X.J.; et al. APF lncRNA regulates autophagy and myocardial infarction by targeting miR-188-3p. Nat. Commun. 2015, 6, 6779. [Google Scholar] [CrossRef] [PubMed]
- Higashi, K.; Yamada, Y.; Minatoguchi, S.; Baba, S.; Iwasa, M.; Kanamori, H.; Kawasaki, M.; Nishigaki, K.; Takemura, G.; Kumazaki, M.; et al. MicroRNA-145 repairs infarcted myocardium by accelerating cardiomyocyte autophagy. Am. J. Physiology-Heart Circ. Physiol. 2015, 309, H1813–H1826. [Google Scholar] [CrossRef] [PubMed]
- Ucar, A.; Gupta, S.K.; Fiedler, J.; Erikci, E.; Kardasinski, M.; Batkai, S.; Dangwal, S.; Kumarswamy, R.; Bang, C.; Holzmann, A.; et al. The miRNA-212/132 family regulates both cardiac hypertrophy and cardiomyocyte autophagy. Nat. Commun. 2012, 3, 1078. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, Y.; Wang, Y.; Qian, J.; Ma, H.; Wang, X.; Jiang, G.; Liu, M.; An, Y.; Ma, L.; et al. Cardiomyocyte-restricted low density lipoprotein receptor-related protein 6 (LRP6) deletion leads to lethal dilated cardiomyopathy partly through Drp1 signaling. Theranostics 2018, 8, 627–643. [Google Scholar] [CrossRef] [PubMed]
- Carresi, C.; Musolino, V.; Gliozzi, M.; Maiuolo, J.; Mollace, R.; Nucera, S.; Maretta, A.; Sergi, D.; Muscoli, S.; Gratteri, S.; et al. Anti-oxidant effect of bergamot polyphenolic fraction counteracts doxorubicin-induced cardiomyopathy: Role of autophagy and c-kitposCD45negCD31neg cardiac stem cell activation. J. Mol. Cell. Cardiol. 2018, 119, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. Micrornas: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Radmark, O.; Kim, S.; et al. The nuclear RNase III drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lee, Y.; Yeom, K.H.; Nam, J.W.; Heo, I.; Rhee, J.K.; Sohn, S.Y.; Cho, Y.; Zhang, B.T.; Kim, V.N. Molecular basis for the recognition of primary micrornas by the Drosha-DGCR8 complex. Cell 2006, 125, 887–901. [Google Scholar] [CrossRef] [PubMed]
- Lund, E.; Guttinger, S.; Calado, A.; Dahlberg, J.E.; Kutay, U. Nuclear export of microRNA precursors. Science 2004, 303, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; McLachlan, J.; Pasquinelli, A.E.; Balint, E.; Tuschl, T.; Zamore, P.D. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 2001, 293, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: Are the answers in sight? Nat. Rev. Genet. 2008, 9, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Eulalio, A.; Huntzinger, E.; Izaurralde, E. Getting to the root of miRNA-mediated gene silencing. Cell 2008, 132, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Su, Z.; Yang, Z.; Xu, Y.; Chen, Y.; Yu, Q. Micrornas in apoptosis, autophagy and necroptosis. Oncotarget 2015, 6, 8474–8490. [Google Scholar] [CrossRef] [PubMed]
- Llorens, F.; Thune, K.; Marti, E. Regional and subtype-dependent miRNA signatures in sporadic Creutzfeldt-Jakob disease are accompanied by alterations in miRNA silencing machinery and biogenesis. PloS Pathog. 2018, 14, e1006802. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Ma, B.; Han, X. The role of autophagy in angiotensin II induced pathological cardiac hypertrophy. J. Mol. Endocrinol. 2016, 57, R143–R152. [Google Scholar] [CrossRef] [PubMed]
- Frankel, L.B.; Lund, A.H. Microrna regulation of autophagy. Carcinogenesis 2012, 33, 2018–2025. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhang, Z.; Lu, Y.; Song, K.; Liu, X.; Xia, F.; Sun, W. Downregulation of UKL1 by microRNA-372 inhibits the survival of human pancreatic adenocarcinoma cells. Cancer Sci. 2017, 108, 1811–1819. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Zhang, T.; Ding, S.; Wei, J.; Su, C.; Liu, H.; Xu, G. MicroRNA-17-5p modulates bacille calmette-guerin growth in RAW264.7 cells by targeting ULK1. PloS ONE 2015, 10, e0138011. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, S.I.; Gautschi, O.; Batliner, J.; Gugger, M.; Fey, M.F.; Tschan, M.P. MicroRNA-106a targets autophagy and enhances sensitivity of lung cancer cells to Src inhibitors. Lung Cancer 2017, 107, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Chuang, A.Y.; Ratovitski, E.A. Phospho-ΔNp63α/miR-885–3p axis in tumor cell life and cell death upon cisplatin exposure. Cell Cycle 2011, 10, 3938–3947. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, J.; Tang, N. Long noncoding RNA Malat1 is a potent autophagy inducer protecting brain microvascular endothelial cells against oxygen-glucose deprivation/reoxygenation-induced injury by sponging miR-26b and upregulating ULK2 expression. Neuroscience 2017, 354, 1–10. [Google Scholar] [CrossRef] [PubMed]
- John Clotaire, D.Z.; Zhang, B.; Wei, N.; Gao, R.; Zhao, F.; Wang, Y.; Lei, M.; Huang, W. Mir-26b inhibits autophagy by targeting ULK2 in prostate cancer cells. Biochem. Biophys. Res. Commun. 2016, 472, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Qiang, Q.; Shan, H.; Shi, M.; Gan, G.; Ma, F.; Chen, B. MiR-20a and miR-20b negatively regulate autophagy by targeting RB1CC1/FIP200 in breast cancer cells. Life Sci. 2016, 147, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Ren, X.; Hait, W.N.; Yang, J.M. Therapeutic targeting of autophagy in disease: Biology and pharmacology. Pharmacol. Rev. 2013, 65, 1162–1197. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Guerrero-Preston, R.; Ratovitski, E.A. Phospho-ΔNp63α-dependent regulation of autophagic signaling through transcription and micro-RNA modulation. Cell Cycle 2012, 11, 1247–1259. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yu, Y.; Li, S.; Liu, Y.; Zhou, S.; Cao, S.; Yin, J.; Li, G. Micro RNA-30a ameliorates hepatic fibrosis by inhibiting Beclin1-mediated autophagy. J. Cell. Mol. Med. 2017, 21, 3679–3692. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Song, L.; Zhao, Y.; Liu, Q.; Zhang, S. Inhibition of Beclin-1-mediated autophagy by microRNA-17-5p enhanced the radiosensitivity of glioma cells. Oncol. Res. Featur. Preclin. Clin. Cancer Ther. 2017, 25, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Zhang, Y.; Kang, L.; Song, Y.; Wang, K.; Li, S.; Wu, X.; Hua, W.; Shao, Z.; Yang, S.; et al. Methylation of microRNA-129-5p modulates nucleus pulposus cell autophagy by targeting Beclin-1 in intervertebral disc degeneration. Oncotarget 2017, 8, 86264–86276. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Shi, H.; Lin, S.; Ba, M.; Cui, S. MicroRNA-216a enhances the radiosensitivity of pancreatic cancer cells by inhibiting beclin-1-mediated autophagy. Oncol. Rep. 2015, 34, 1557–1564. [Google Scholar] [CrossRef] [PubMed]
- Korkmaz, G.; le Sage, C.; Tekirdag, K.A.; Agami, R.; Gozuacik, D. miR-376b controls starvation and mTOR inhibition-related autophagy by targeting ATG4C and BECN1. Autophagy 2012, 8, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Sun, A.G.; Meng, F.G.; Wang, M.G. Cisd2 promotes the proliferation of glioma cells via suppressing beclin1mediated autophagy and is targeted by microRNA449a. Mol. Med. Rep. 2017, 16, 7939–7948. [Google Scholar] [CrossRef] [PubMed]
- Han, R.; Ji, X.; Rong, R.; Li, Y.; Yao, W.; Yuan, J.; Wu, Q.; Yang, J.; Yan, W.; Han, L.; et al. MiR-449a regulates autophagy to inhibit silica-induced pulmonary fibrosis through targeting Bcl2. J. Mol. Med. 2016, 94, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Gao, X.; Wang, J.; Yang, C.; Wang, Y.; Liu, Y.; Zou, W.; Liu, T. Hypoxia-induced microRNA-146a represses Bcl-2 through Traf6/IRAK1 but not Smad4 to promote chondrocyte autophagy. Biol. Chem. 2017, 398, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.L.; He, G.Y.; Lan, X.L.; Zeng, Z.C.; Guan, J.; Ding, Y.; Qian, X.L.; Liao, W.T.; Ding, Y.Q.; Liang, L. Inhibition of ATG12-mediated autophagy by miR-214 enhances radiosensitivity in colorectal cancer. Oncogenesis 2018, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.; Zhou, C.; Han, S.; Hou, X.; Kang, S.; Zhang, Y. MicroRNA-378 enhances migration and invasion in cervical cancer by directly targeting autophagy-related protein 12. Mol. Med. Rep. 2018, 17, 6319–6326. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Yu, B.; Cheng, C.; Cheng, T.; Yuan, B.; Li, K.; Xiao, J.; Qiu, Z. Mir505-3p regulates axonal development via inhibiting the autophagy pathway by targeting Atg12. Autophagy 2017, 13, 1679–1696. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Liu, A.; Zhang, J.; Ji, W.; Li, Y.; Yang, X.; Wu, Z.; Guo, J. miR-23b improves cognitive impairments in traumatic brain injury by targeting ATG12-mediated neuronal autophagy. Behav. Brain Res. 2018, 340, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; He, Y.; Zhai, N.; Ding, S.; Li, J.; Peng, Z. MicroRNA-181a inhibits autophagy by targeting Atg5 in hepatocellular carcinoma. Front. Biosci. (Landmark edition) 2018, 23, 388–396. [Google Scholar]
- Fu, X.T.; Shi, Y.H.; Zhou, J.; Peng, Y.F.; Liu, W.R.; Shi, G.M.; Gao, Q.; Wang, X.Y.; Song, K.; Fan, J.; et al. MicroRNA-30a suppresses autophagy-mediated anoikis resistance and metastasis in hepatocellular carcinoma. Cancer Lett. 2018, 412, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Chen, J.; Zhou, H.; Chen, Y.; Zhi, Y.; Zhang, B.; Chen, L.; Chu, X.; Wang, R.; Zhang, C. Pu.1/microRNA-142-3p targets ATG5/ATG16L1 to inactivate autophagy and sensitize hepatocellular carcinoma cells to sorafenib. Cell Death Dis. 2018, 9, 312. [Google Scholar] [CrossRef] [PubMed]
- Gan, J.; Cai, Q.; Qu, Y.; Zhao, F.; Wan, C.; Luo, R.; Mu, D. MiR-96 attenuates status epilepticus-induced brain injury by directly targeting Atg7 and Atg16l1. Sci. Rep. 2017, 7, 10270. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Zhang, Z.Z.; Yang, W.; Ouyang, Z.H.; Xue, J.B.; Li, X.L.; Zhang, J.; Chen, W.K.; Yan, Y.G.; Wang, W.J. MiR-210 facilitates ECM degradation by suppressing autophagy via silencing of ATG7 in human degenerated NP cells. Biomed. Pharmacother. 2017, 93, 470–479. [Google Scholar] [CrossRef] [PubMed]
- Comincini, S.; Allavena, G.; Palumbo, S.; Morini, M.; Durando, F.; Angeletti, F.; Pirtoli, L.; Miracco, C. MicroRNA-17 regulates the expression of ATG7 and modulates the autophagy process, improving the sensitivity to temozolomide and low-dose ionizing radiation treatments in human glioblastoma cells. Cancer Biol. Ther. 2013, 14, 574–586. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Shen, W.; Zhu, Z.; Lin, J.; Fang, Q.; Ruan, Y.; Zhao, H. Combined inhibition of EGFR and c-ABL suppresses the growth of fulvestrant-resistant breast cancer cells through miR-375-autophagy axis. Biochem. Bioph. Res. Commun. 2018, 498, 559–565. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Cheng, Y. Inhibition of miR-20 promotes proliferation and autophagy in articular chondrocytes by PI3K/AKT/mTOR signaling pathway. Biomed. Pharmacother. 2018, 97, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Dai, X.; Ni, Z.; Yan, X.; He, F.; Lian, J. The downregulation of ATG4B mediated by microRNA-34a/34c-5p suppresses rapamycin-induced autophagy. Irani. J. Basic Med. Sci. 2017, 20, 1125–1130. [Google Scholar]
- Frankel, L.B.; Wen, J.; Lees, M.; Hoyer-Hansen, M.; Farkas, T.; Krogh, A.; Jaattela, M.; Lund, A.H. MicroRNA-101 is a potent inhibitor of autophagy. EMBO J. 2011, 30, 4628–4641. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Zhu, X.; He, B.; Zhang, Y.; Kang, B.; Wang, Z.; Ni, X. MiR-204 regulates cardiomyocyte autophagy induced by ischemia-reperfusion through LC3-II. J. Biomed. Sci. 2011, 18, 35. [Google Scholar] [CrossRef] [PubMed]
- Ying, Q.; Liang, L.; Guo, W.; Zha, R.; Tian, Q.; Huang, S.; Yao, J.; Ding, J.; Bao, M.; Ge, C.; et al. Hypoxia-inducible microRNA-210 augments the metastatic potential of tumor cells by targeting vacuole membrane protein 1 in hepatocellular carcinoma. Hepatology 2011, 54, 2064–2075. [Google Scholar] [CrossRef] [PubMed]
- Rotter, D.; Rothermel, B.A. Targets, trafficking, and timing of cardiac autophagy. Pharmacol. Res. 2012, 66, 494–504. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.V.; Rothermel, B.A.; Hill, J.A. Autophagy in hypertensive heart disease. J. Biol. Chem. 2010, 285, 8509–8514. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chen, D.; He, Y.; Melendez, A.; Feng, Z.; Hong, Q.; Bai, X.; Li, Q.; Cai, G.; Wang, J.; et al. MiR-34 modulates caenorhabditis elegans lifespan via repressing the autophagy gene atg9. Age 2013, 35, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Kovaleva, V.; Mora, R.; Park, Y.J.; Plass, C.; Chiramel, A.I.; Bartenschlager, R.; Dohner, H.; Stilgenbauer, S.; Pscherer, A.; Lichter, P.; et al. MircoRNA-130a targets ATG2b and DICER1 to inhibit autophagy and trigger killing of chronic lymphocytic leukemia cells. Cancer Res. 2012, 72, 1763–1772. [Google Scholar] [CrossRef] [PubMed]
- Bravo-San Pedro, J.M.; Kroemer, G.; Galluzzi, L. Autophagy and mitophagy in cardiovascular disease. Circ. Res. 2017, 120, 1812–1824. [Google Scholar] [CrossRef] [PubMed]
- Nakai, A.; Yamaguchi, O.; Takeda, T.; Higuchi, Y.; Hikoso, S.; Taniike, M.; Omiya, S.; Mizote, I.; Matsumura, Y.; Asahi, M.; et al. The role of autophagy in cardiomyocytes in the basal state and in response to hemodynamic stress. Nat. Med. 2007, 13, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Taneike, M.; Yamaguchi, O.; Nakai, A.; Hikoso, S.; Takeda, T.; Mizote, I.; Oka, T.; Tamai, T.; Oyabu, J.; Murakawa, T.; et al. Inhibition of autophagy in the heart induces age-related cardiomyopathy. Autophagy 2010, 6, 600–606. [Google Scholar] [CrossRef] [PubMed]
- Ouimet, M.; Ediriweera, H.; Afonso, M.S.; Ramkhelawon, B.; Singaravelu, R.; Liao, X.; Bandler, R.C.; Rahman, K.; Fisher, E.A.; Rayner, K.J.; et al. MicroRNA-33 regulates macrophage autophagy in atherosclerosis. Arterioscleros. Thromb. Vasc. Biol. 2017, 37, 1058–1067. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, W.N.; Xu, S.B.; Wu, H.; Dai, B.; Jian, D.D.; Yang, M.; Wu, Y.T.; Feng, Q.; Zhu, J.H.; et al. MicroRNA-214-3p: A link between autophagy and endothelial cell dysfunction in atherosclerosis. Acta Physiol. 2018, 222, e12973. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Wang, Z.; Hu, X.; Wan, T.; Wu, H.; Jiang, W.; Hu, R. Human aortic smooth muscle cell-derived exosomal miR-221/222 inhibits autophagy via a PTEN/Akt signaling pathway in human umbilical vein endothelial cells. Biochem. Biophys. Res. Commun. 2016, 479, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Sun, Y.T.; Xu, T.H.; Sun, W.; Tian, B.Y.; Sheng, Z.T.; Sun, L.; Liu, L.L.; Ma, J.F.; Wang, L.N.; et al. MicroRNA-30b regulates high phosphorus level-induced autophagy in vascular smooth muscle cells by targeting BECN1. Cell. Physiol. Biochem. 2017, 42, 530–536. [Google Scholar] [CrossRef] [PubMed]
- Pietrocola, F.; Izzo, V.; Niso-Santano, M.; Vacchelli, E.; Galluzzi, L.; Maiuri, M.C.; Kroemer, G. Regulation of autophagy by stress-responsive transcription factors. Semin. Cancer Biol. 2013, 23, 310–322. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Ding, F.; Wang, X.; Huang, Q.; Zhang, L.; Bi, C.; Hua, B.; Yuan, Y.; Han, Z.; Jin, M.; et al. By targeting Atg7 microRNA-143 mediates oxidative stress-induced autophagy of c-kit+ mouse cardiac progenitor cells. EBioMedicine 2018, 32, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Dorn, G.W., 2nd. Mitochondrial pruning by Nix and BNip3: An essential function for cardiac-expressed death factors. J. Cardiovasc. Transl. Res. 2010, 3, 374–383. [Google Scholar] [CrossRef] [PubMed]
- Gong, G.; Song, M.; Csordas, G.; Kelly, D.P.; Matkovich, S.J.; Dorn, G.W., 2nd. Parkin-mediated mitophagy directs perinatal cardiac metabolic maturation in mice. Science 2015, 350, aad2459. [Google Scholar] [CrossRef] [PubMed]
- Andres, A.M.; Hernandez, G.; Lee, P.; Huang, C.; Ratliff, E.P.; Sin, J.; Thornton, C.A.; Damasco, M.V.; Gottlieb, R.A. Mitophagy is required for acute cardioprotection by simvastatin. Antioxid. Redox Signal. 2014, 21, 1960–1973. [Google Scholar] [CrossRef] [PubMed]
- Billia, F.; Hauck, L.; Konecny, F.; Rao, V.; Shen, J.; Mak, T.W. PTEN-inducible kinase 1 (PINK1)/Park6 is indispensable for normal heart function. Proc. Natl. Acad. Sci. USA 2011, 108, 9572–9577. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhang, X.; Zhuang, H.; Chen, H.G.; Chen, Y.; Tian, W.; Wu, W.; Li, Y.; Wang, S.; Zhang, L.; et al. MicroRNA-137 is a novel hypoxia-responsive microRNA that inhibits mitophagy via regulation of two mitophagy receptors FUNDC1 and NIX. J. Biol. Chem. 2014, 289, 10691–10701. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Fiesel, F.C.; Belmonte, K.C.; Hudec, R.; Wang, W.X.; Kim, C.; Nelson, P.T.; Springer, W.; Kim, J. MiR-27a and miR-27b regulate autophagic clearance of damaged mitochondria by targeting PTEN-induced putative kinase 1 (PINK1). Mol. Neurodegener. 2016, 11, 55. [Google Scholar] [CrossRef] [PubMed]
- Cheng, M.; Liu, L.; Lao, Y.; Liao, W.; Liao, M.; Luo, X.; Wu, J.; Xie, W.; Zhang, Y.; Xu, N. MicroRNA-181a suppresses parkin-mediated mitophagy and sensitizes neuroblastoma cells to mitochondrial uncoupler-induced apoptosis. Oncotarget 2016, 7, 42274–42287. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Li, T. MicroRNA-410 is involved in mitophagy after cardiac ischemia/reperfusion injury by targeting high-mobility group box 1 protein. J. Cell. Biochem. 2018, 119, 2427–2439. [Google Scholar] [CrossRef] [PubMed]
- Verjans, R.; van Bilsen, M.; Schroen, B. MiRNA deregulation in cardiac aging and associated disorders. Int. Rev. Cell Mol. Biol. 2017, 334, 207–263. [Google Scholar] [PubMed]
- Yan, L.; Gao, S.; Ho, D.; Park, M.; Ge, H.; Wang, C.; Tian, Y.; Lai, L.; De Lorenzo, M.S.; Vatner, D.E.; et al. Calorie restriction can reverse, as well as prevent, aging cardiomyopathy. Age 2013, 35, 2177–2182. [Google Scholar] [CrossRef] [PubMed]
- Egan, D.F.; Shackelford, D.B.; Mihaylova, M.M.; Gelino, S.; Kohnz, R.A.; Mair, W.; Vasquez, D.S.; Joshi, A.; Gwinn, D.M.; Taylor, R.; et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 2011, 331, 456–461. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kim, Y.C.; Fang, C.; Russell, R.C.; Kim, J.H.; Fan, W.; Liu, R.; Zhong, Q.; Guan, K.L. Differential regulation of distinct Vps34 complexes by AMPK in nutrient stress and autophagy. Cell 2013, 152, 290–303. [Google Scholar] [CrossRef] [PubMed]
- Russell, R.C.; Tian, Y.; Yuan, H.; Park, H.W.; Chang, Y.Y.; Kim, J.; Kim, H.; Neufeld, T.P.; Dillin, A.; Guan, K.L. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat. Cell Biol. 2013, 15, 741–750. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Wang, F.; Hu, S.; Yin, C.; Li, X.; Zhao, S.; Wang, J.; Yan, X. MiR-20a and miR-106b negatively regulate autophagy induced by leucine deprivation via suppression of ULK1 expression in C2C12 myoblasts. Cell. signal. 2012, 24, 2179–2186. [Google Scholar] [CrossRef] [PubMed]
- Wan, G.; Xie, W.; Liu, Z.; Xu, W.; Lao, Y.; Huang, N.; Cui, K.; Liao, M.; He, J.; Jiang, Y.; et al. Hypoxia-induced miR155 is a potent autophagy inducer by targeting multiple players in the mtor pathway. Autophagy 2014, 10, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Martins-Marques, T.; Ribeiro-Rodrigues, T.; Pereira, P.; Codogno, P.; Girao, H. Autophagy and ubiquitination in cardiovascular diseases. DNA Cell Biol. 2015, 34, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Zhong, Y.; Cheng, C.; Liu, B.; Wang, L.; Li, A.; Xiong, L.; Liu, S. MiR-30-regulated autophagy mediates angiotensin II-induced myocardial hypertrophy. PloS ONE 2013, 8, e53950. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Yang, S.; Li, H.; Yin, Z.; Fan, J.; Zhao, Y.; Gong, W.; Yan, M.; Wang, D.W. MiR30c is involved in diabetic cardiomyopathy through regulation of cardiac autophagy via BECN1. Mol. Therapy-Nucleic Acids 2017, 7, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Sun, W.; Huang, H.; Ye, J.; Pan, W.; Zhong, Y.; Cheng, C.; You, X.; Liu, B.; Xiong, L.; et al. MiR-34a modulates angiotensin ii-induced myocardial hypertrophy by direct inhibition of atg9a expression and autophagic activity. PloS ONE 2014, 9, e94382. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Chen, Z.; Wang, C.; Song, L.; Zou, Y.; Zhang, L.; Hui, R.; Wang, J. Cardiac-specific overexpression of miR-222 induces heart failure and inhibits autophagy in mice. Cell. Physiol. Biochem. 2016, 39, 1503–1511. [Google Scholar] [CrossRef] [PubMed]
- Lai, L.; Chen, J.; Wang, N.; Zhu, G.; Duan, X.; Ling, F. MiRNA-30e mediated cardioprotection of ACE2 in rats with doxorubicin-induced heart failure through inhibiting cardiomyocytes autophagy. Life Sci. 2017, 169, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Wang, A.; Xu, Y.; Qiao, S.; An, J.; Li, H.; Wang, C. Role of microRNA-1-mediated AMP-activated protein kinase pathway in cardiac fibroblasts induced by high glucose in rats. Zhonghua wei zhong bing ji jiu yi xue 2018, 30, 145–150. [Google Scholar] [PubMed]
- D’Avenia, M.; Citro, R.; De Marco, M.; Veronese, A.; Rosati, A.; Visone, R.; Leptidis, S.; Philippen, L.; Vitale, G.; Cavallo, A.; et al. A novel miR-371a-5p-mediated pathway, leading to BAG3 upregulation in cardiomyocytes in response to epinephrine, is lost in takotsubo cardiomyopathy. Cell Death Dis. 2015, 6, e1948. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Yang, L.; Wang, L.; Tang, B.; Wang, J.; Li, Q. MicroRNA-34a protects myocardial cells against ischemia-reperfusion injury through inhibiting autophagy via regulating TNAα expression. Biochem. Cell Biol. 2018, 96, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Deng, Y.; Xu, Y.; Jin, W.; Li, H. MicroRNA-223 protects neonatal rat cardiomyocytes and h9c2 cells from hypoxia-induced apoptosis and excessive autophagy via the akt/mtor pathway by targeting parp-1. J. Mol. Cell. Cardiol. 2018, 118, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Dong, Y.H.; Du, W.; Shi, C.Y.; Wang, K.; Tariq, M.A.; Wang, J.X.; Li, P.F. The role of microRNAs in myocardial infarction: From molecular mechanism to clinical application. Int. J. Mol. Sci. 2017, 18, 745. [Google Scholar] [CrossRef] [PubMed]
- Bo, L.; Su-Ling, D.; Fang, L.; Lu-Yu, Z.; Tao, A.; Stefan, D.; Kun, W.; Pei-Feng, L. Autophagic program is regulated by miR-325. Cell Death Differ. 2014, 21, 967–977. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Li, J.; Kou, B.; Yi, Q.; Shi, T. MicroRNA-30e protects the heart against ischemia and reperfusion injury through autophagy and the notch1/Hes1/Akt signaling pathway. Int. J. Mol. Med. 2018, 41, 3221–3230. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Xie, J.; Li, R.; Shi, J.; Sun, J.; Gu, R.; Ding, L.; Wang, L.; Xu, B. Overexpression of microRNA-99a attenuates heart remodelling and improves cardiac performance after myocardial infarction. J. Cell. Mol. Med. 2014, 18, 919–928. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zeng, Z.; Li, Q.; Xu, Q.; Xie, J.; Hao, H.; Luo, G.; Liao, W.; Bin, J.; Huang, X.; et al. Inhibition of microRNA-497 ameliorates anoxia/reoxygenation injury in cardiomyocytes by suppressing cell apoptosis and enhancing autophagy. Oncotarget 2015, 6, 18829–18844. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Cui, T. Autophagy modulation: A potential therapeutic approach in cardiac hypertrophy. Am. J. Physiol. Heart Circ. Physiol. 2017, 313, H304–H319. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Su, M.; Wang, S.; Zou, Y.; Wang, X.; Wang, Y.; Cui, H.; Zhao, P.; Hui, R.; Wang, J. MiR-451 is decreased in hypertrophic cardiomyopathy and regulates autophagy by targeting tsc1. J. Cell. Mol. Med. 2014, 18, 2266–2274. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.D.; Qin, R.H.; Yang, J.J.; Xu, S.S.; Tao, H.; Ding, X.S.; Shi, K.H. DNMT3A controls miR-200b in cardiac fibroblast autophagy and cardiac fibrosis. Inflamm. Res. 2018, 67, 681–690. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, T.; Li, M.-Y.; Li, P.-F.; Cao, J.-M. MicroRNAs in Cardiac Autophagy: Small Molecules and Big Role. Cells 2018, 7, 104. https://doi.org/10.3390/cells7080104
Sun T, Li M-Y, Li P-F, Cao J-M. MicroRNAs in Cardiac Autophagy: Small Molecules and Big Role. Cells. 2018; 7(8):104. https://doi.org/10.3390/cells7080104
Chicago/Turabian StyleSun, Teng, Meng-Yang Li, Pei-Feng Li, and Ji-Min Cao. 2018. "MicroRNAs in Cardiac Autophagy: Small Molecules and Big Role" Cells 7, no. 8: 104. https://doi.org/10.3390/cells7080104
APA StyleSun, T., Li, M.-Y., Li, P.-F., & Cao, J.-M. (2018). MicroRNAs in Cardiac Autophagy: Small Molecules and Big Role. Cells, 7(8), 104. https://doi.org/10.3390/cells7080104