DNA Methylation: A Potential Mediator of the Memory Regulatory Effects of taVNS
Abstract
1. Introduction
2. Memory Enhancement Effects of taVNS
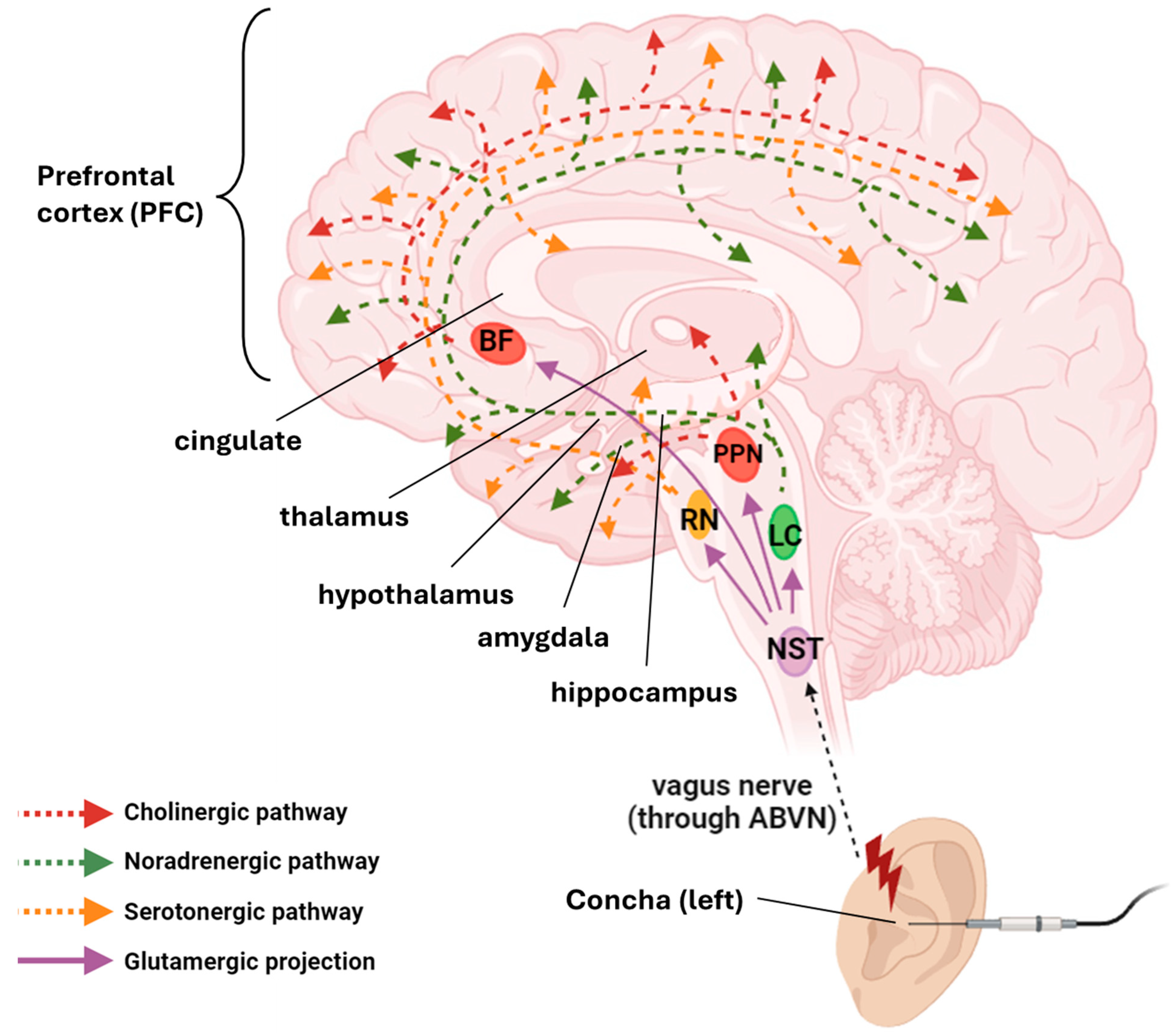
2.1. Anatomical and Physiological Basis of taVNS
2.2. Therapeutic Effects in Humans and Animals
2.3. Selective Memory Modulation by taVNS
2.4. Electrophysiological and Biochemical Effects of taVNS
3. DNA Methylation and Memory: A Functional Perspective
3.1. DNA Methylation Regulators and Readers
3.1.1. DNA Methyltransferases
3.1.2. Demethylases
3.1.3. Methyl-CpG-Binding Protein 2
3.2. Regulation of DNA Methylation by Neuronal Activity
3.2.1. Dynamic Modification of the Spatiotemporal Methylation Pattern
3.2.2. Differential Binding by MeCP2
4. Perspectives for Future Mechanistic Studies on taVNS
5. Summary
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ABVN | auricular branch of the vagus nerve |
| ACC | anterior cingulate cortex |
| ACS | alternating current stimulation |
| AD | Alzheimer’s disease |
| AP | area postrema |
| ASD | autism spectrum disorder |
| BDNF | brain-derived neurotrophic factor |
| BF | basal forebrain |
| CREB | cAMP response element-binding protein |
| DBS | deep brain stimulation |
| DCS | direct current stimulation |
| DLB | dementia with Lewy bodies |
| DMN | dorsal motor nucleus of the VN |
| DNMT | DNA methyltransferase |
| ECT | electroconvulsive therapy |
| 5-caC | 5-carboxylcytosine |
| 5-mC | 5-methyl cytosine |
| 5-fC | 5-formylcytosine |
| 5-hmC | 5-hydroxymethylcytosine |
| GABA | γ-aminobutyric acid |
| Gadd45 | growth arrest and DNA damage 45 |
| GluR1 | glutamatergic receptor 1 |
| LC | locus coeruleus |
| LTP | Long- term potentiation |
| L-VGCC | L-type voltage-gated calcium channel |
| MCI | mild cognitive impairment |
| MeCP2 | methyl-CpG-binding protein 2 |
| MWM | Morris water maze |
| NA | nucleus ambiguus |
| NCoR | nuclear co-repressor |
| NEDD8 | neural precursor cell-expressed developmentally down-regulated gene 8 |
| NMDAR | N-methyl-D-aspartate receptor |
| NORT | novel object recognition test |
| Nrp1 | neuropilin-1 |
| NST | nucleus of the solitary tract |
| NT | neurotrophin |
| PD | Parkinson’s disease |
| PFC | prefrontal cortex |
| PKA | protein kinase A |
| PPN | pedunculopontine nucleus |
| PrL | prelimbic cortex |
| PSD95 | postsynaptic protein 95 |
| RN | raphe nuclei |
| SAMP8 | senescence-accelerated mouse prone 8 |
| SNT | spinal nucleus of the trigeminal nerve |
| TaVNS | transcutaneous auricular vagus nerve stimulation |
| TBRS | Tatton-Brown Rahman syndrome |
| Tet | ten-eleven translocation |
| TMS | transcranial magnetic stimulation |
References
- Alberini, C.M. Mechanisms of memory stabilization: Are consolidation and reconsolidation similar or distinct processes? Trends Neurosci. 2005, 28, 51–56. [Google Scholar] [CrossRef]
- Tan, S.Z.K.; Sheng, V.; Chan, Y.S.; Lim, L.W. Eternal sunshine of the neuromodulated mind: Altering fear memories through neuromodulation. Exp. Neurol. 2019, 314, 9–19. [Google Scholar] [CrossRef] [PubMed]
- D’Ardenne, K.; Eshel, N.; Luka, J.; Lenartowicz, A.; Nystrom, L.E.; Cohen, J.D. Role of prefrontal cortex and the midbrain dopamine system in working memory updating. Proc. Natl. Acad. Sci. USA 2012, 109, 19900–19909. [Google Scholar] [CrossRef] [PubMed]
- Josselyn, S.A.; Tonegawa, S. Memory engrams: Recalling the past and imagining the future. Science 2020, 367, eaaw4325. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.Z.K.; Du, R.; Perucho, J.A.U.; Chopra, S.S.; Vardhanabhuti, V.; Lim, L.W. Dropout in Neural Networks Simulates the Paradoxical Effects of Deep Brain Stimulation on Memory. Front. Aging Neurosci. 2020, 12, 273. [Google Scholar] [CrossRef]
- Barry, D.N.; Maguire, E.A. Remote Memory and the Hippocampus: A Constructive Critique. Trends Cogn. Sci. 2019, 23, 128–142. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, Y.; Wang, J.; Xia, Y.; Zhang, J.; Chen, L. Recent advances in Alzheimer’s disease: Mechanisms, clinical trials and new drug development strategies. Signal Transduct. Target. Ther. 2024, 9, 211. [Google Scholar] [CrossRef]
- Huang, L.-K.; Kuan, Y.-C.; Lin, H.-W.; Hu, C.-J. Clinical trials of new drugs for Alzheimer disease: A 2020–2023 update. J. Biomed. Sci. 2023, 30, 83. [Google Scholar] [CrossRef]
- Lockhart, B.P.; Lestage, P.J. Cognition enhancing or neuroprotective compounds for the treatment of cognitive disorders: Why? when? which? Exp. Gerontol. 2003, 38, 119–128. [Google Scholar] [CrossRef]
- Yu, W.S.; Tse, A.C.; Guan, L.; Chiu, J.L.Y.; Tan, S.Z.K.; Khairuddin, S.; Agadagba, S.K.; Lo, A.C.Y.; Fung, M.L.; Chan, Y.S.; et al. Antidepressant-like effects of transcorneal electrical stimulation in rat models. Brain Stimul. 2022, 15, 843–856. [Google Scholar] [CrossRef]
- Yu, W.S.; Aquili, L.; Wong, K.H.; Lo, A.C.Y.; Chan, L.L.H.; Chan, Y.S.; Lim, L.W. Transcorneal electrical stimulation enhances cognitive functions in aged and 5XFAD mouse models. Ann. Acad. Sci. 2022, 1515, 249–265. [Google Scholar] [CrossRef] [PubMed]
- Goldthorpe, R.A.; Rapley, J.M.; Violante, I.R. A Systematic Review of Non-invasive Brain Stimulation Applications to Memory in Healthy Aging. Front. Neurol. 2020, 11, 575075. [Google Scholar] [CrossRef]
- Jacobs, H.I.L.; Riphagen, J.M.; Razat, C.M.; Wiese, S.; Sack, A.T. Transcutaneous vagus nerve stimulation boosts associative memory in older individuals. Neurobiol. Aging 2015, 36, 1860–1867. [Google Scholar] [CrossRef] [PubMed]
- Burger, A.M.; D’Agostini, M.; Verkuil, B.; Van Diest, I. Moving beyond belief: A narrative review of potential biomarkers for transcutaneous vagus nerve stimulation. Psychophysiology 2020, 57, e13571. [Google Scholar] [CrossRef] [PubMed]
- Sawchenko, P.E. Central connections of the sensory and motor nuclei of the vagus nerve. J. Auton. Nerv. Syst. 1983, 9, 13–26. [Google Scholar] [CrossRef]
- Ruggiero, D.A.; Underwood, M.D.; Mann, J.J.; Anwar, M.; Arango, V. The human nucleus of the solitary tract: Visceral pathways revealed with an “in vitro” postmortem tracing method. J. Auton. Nerv. Syst. 2000, 79, 181–190. [Google Scholar] [CrossRef]
- Knowles, C.H.; Aziz, Q. Basic and clinical aspects of gastrointestinal pain. PAIN 2009, 141, 191–209. [Google Scholar] [CrossRef]
- Choi, S.; Jang, D.C.; Chung, G.; Kim, S.K. Transcutaneous Auricular Vagus Nerve Stimulation Enhances Cerebrospinal Fluid Circulation and Restores Cognitive Function in the Rodent Model of Vascular Cognitive Impairment. Cells 2022, 11, 3019. [Google Scholar] [CrossRef]
- Yu, Y.; Jiang, X.; Fang, X.; Wang, Y.; Liu, P.; Ling, J.; Yu, L.; Jiang, M.; Tang, C. Transauricular Vagal Nerve Stimulation at 40 Hz Inhibits Hippocampal P2X7R/NLRP3/Caspase-1 Signaling and Improves Spatial Learning and Memory in 6-Month-Old APP/PS1 Mice. Neuromodulation Technol. Neural Interface 2023, 26, 589–600. [Google Scholar] [CrossRef]
- Sun, J.-B.; Cheng, C.; Tian, Q.-Q.; Yuan, H.; Yang, X.-J.; Deng, H.; Guo, X.-Y.; Cui, Y.-P.; Zhang, M.-K.; Yin, Z.-X.; et al. Transcutaneous Auricular Vagus Nerve Stimulation Improves Spatial Working Memory in Healthy Young Adults. Front. Neurosci. 2021, 15, 790793. [Google Scholar] [CrossRef]
- Sanders, T.H.; Weiss, J.; Hogewood, L.; Chen, L.; Paton, C.; McMahan, R.L.; Sweatt, J.D. Cognition-Enhancing Vagus Nerve Stimulation Alters the Epigenetic Landscape. J. Neurosci. 2019, 39, 3454–3469. [Google Scholar] [CrossRef]
- Poon, C.H.; Liu, Y.; Pak, S.; Zhao, R.C.; Aquili, L.; Tipoe, G.L.; Leung, G.K.; Chan, Y.S.; Yang, S.; Fung, M.L.; et al. Prelimbic Cortical Stimulation with L-methionine Enhances Cognition through Hippocampal DNA Methylation and Neuroplasticity Mechanisms. Aging Dis. 2023, 14, 112–135. [Google Scholar] [CrossRef]
- Liu, Y.; Poon, C.H.; Tse, L.S.R.; Roy, J.; So, C.T.; Tong, B.C.-K.T.; Yao, K.-M.; Tipoe, G.L.S.; Harry, S.; Aquili, L.; et al. Homeostatic DNMT3a activity is required to restore hippocampal DNA methylation and cognition in a mouse model of Alzheimer’s disease. Aging Dis. 2025, 17. [Google Scholar] [CrossRef]
- Poon, C.H.; Chan, Y.S.; Fung, M.L.; Lim, L.W. Memory and neuromodulation: A perspective of DNA methylation. Neurosci. Biobehav. Rev. 2020, 111, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Poon, C.H.; Tse, L.S.R.; Lim, L.W. DNA methylation in the pathology of Alzheimer’s disease: From gene to cognition. Ann. Acad. Sci. 2020, 1475, 15–33. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, L.; Li, M. Mini-review of DNA Methylation Detection Techniques and Their Potential Applications in Disease Diagnosis, Prognosis, and Treatment. ACS Sens. 2024, 9, 1089–1103. [Google Scholar] [CrossRef]
- Fraga, M.F.; Ballestar, E.; Paz, M.F.; Ropero, S.; Setien, F.; Ballestar, M.L.; Heine-Suñer, D.; Cigudosa, J.C.; Urioste, M.; Benitez, J.; et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proc. Natl. Acad. Sci. USA 2005, 102, 10604–10609. [Google Scholar] [CrossRef]
- Martin, G.M. Epigenetic drift in aging identical twins. Proc. Natl. Acad. Sci. USA 2005, 102, 10413–10414. [Google Scholar] [CrossRef]
- Wilson, V.L.; Smith, R.A.; Ma, S.; Cutler, R.G. Genomic 5-methyldeoxycytidine decreases with age. J. Biol. Chem. 1987, 262, 9948–9951. [Google Scholar] [CrossRef]
- Oliveira, A.M.M.; Hemstedt, T.J.; Bading, H. Rescue of aging-associated decline in Dnmt3a2 expression restores cognitive abilities. Nat. Neurosci. 2012, 15, 1111–1113. [Google Scholar] [CrossRef]
- Huang, J.L.; Zhang, F.; Su, M.; Li, J.; Yi, W.; Hou, L.X.; Yang, S.M.; Liu, J.Y.; Zhang, H.A.; Ma, T.; et al. MeCP2 prevents age-associated cognitive decline via restoring synaptic plasticity in a senescence-accelerated mouse model. Aging Cell 2021, 20, e13451. [Google Scholar] [CrossRef]
- Brito, D.V.C.; Kupke, J.; Karaca, K.G.; Zeuch, B.; Oliveira, A.M.M. Mimicking Age-Associated Gadd45γ Dysregulation Results in Memory Impairments in Young Adult Mice. J. Neurosci. 2020, 40, 1197–1210. [Google Scholar] [CrossRef]
- Guo, J.U.; Ma, D.K.; Mo, H.; Ball, M.P.; Jang, M.-H.; Bonaguidi, M.A.; Balazer, J.A.; Eaves, H.L.; Xie, B.; Ford, E.; et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat. Neurosci. 2011, 14, 1345–1351. [Google Scholar] [CrossRef] [PubMed]
- Miller, C.A.; Sweatt, J.D. Covalent Modification of DNA Regulates Memory Formation. Neuron 2008, 59, 1051. [Google Scholar] [CrossRef]
- Levenson, J.M.; Roth, T.L.; Lubin, F.D.; Miller, C.A.; Huang, I.C.; Desai, P.; Malone, L.M.; Sweatt, J.D. Evidence That DNA (Cytosine-5) Methyltransferase Regulates Synaptic Plasticity in the Hippocampus. J. Biol. Chem. 2006, 281, 15763–15773. [Google Scholar] [CrossRef] [PubMed]
- Berthoud, H.-R.; Neuhuber, W.L. Functional and chemical anatomy of the afferent vagal system. Auton. Neurosci. 2000, 85, 1–17. [Google Scholar] [CrossRef]
- Dockray, G.J. Enteroendocrine cell signalling via the vagus nerve. Curr. Opin. Pharmacol. 2013, 13, 954–958. [Google Scholar] [CrossRef]
- Ruffoli, R.; Giorgi, F.S.; Pizzanelli, C.; Murri, L.; Paparelli, A.; Fornai, F. The chemical neuroanatomy of vagus nerve stimulation. J. Chem. Neuroanat. 2011, 42, 288–296. [Google Scholar] [CrossRef]
- Magdaleno-Madrigal, V.M.; Valdés-Cruz, A.; Martínez-Vargas, D.; Martínez, A.; Almazán, S.; Fernández-Mas, R.; Fernández-Guardiola, A. Effect of Electrical Stimulation of the Nucleus of the Solitary Tract on the Development of Electrical Amygdaloid Kindling in the Cat. Epilepsia 2002, 43, 964–969. [Google Scholar] [CrossRef]
- Lulic, D.; Ahmadian, A.; Baaj, A.A.; Benbadis, S.R.; Vale, F.L. Vagus nerve stimulation. Neurosurg. Focus 2009, 27, E5. [Google Scholar] [CrossRef]
- Yuan, H.; Silberstein, S.D. Vagus Nerve and Vagus Nerve Stimulation, a Comprehensive Review: Part I. Headache J. Head Face Pain 2016, 56, 71–78. [Google Scholar] [CrossRef]
- Mello-Carpes, P.B.; Izquierdo, I. The Nucleus of the Solitary Tract → Nucleus Paragigantocellularis → Locus Coeruleus → CA1 region of dorsal hippocampus pathway is important for consolidation of object recognition memory. Neurobiol. Learn. Mem. 2013, 100, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Fornai, F.; Ruffoli, R.; Giorgi, F.S.; Paparelli, A. The role of locus coeruleus in the antiepileptic activity induced by vagus nerve stimulation. Eur. J. Neurosci. 2011, 33, 2169–2178. [Google Scholar] [CrossRef] [PubMed]
- Yakunina, N.; Kim, S.S.; Nam, E.-C. Optimization of Transcutaneous Vagus Nerve Stimulation Using Functional MRI. Neuromodulation Technol. Neural Interface 2017, 20, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Horinouchi, T.; Nezu, T.; Saita, K.; Date, S.; Kurumadani, H.; Maruyama, H.; Kirimoto, H. Transcutaneous auricular vagus nerve stimulation enhances short-latency afferent inhibition via central cholinergic system activation. Sci. Rep. 2024, 14, 11224. [Google Scholar] [CrossRef]
- Gedankien, T.; Tan, R.J.; Qasim, S.E.; Moore, H.; McDonagh, D.; Jacobs, J.; Lega, B. Acetylcholine modulates the temporal dynamics of human theta oscillations during memory. Nat. Commun. 2023, 14, 5283. [Google Scholar] [CrossRef]
- Luchetti, A.; Bota, A.; Weitemier, A.; Mizuta, K.; Sato, M.; Islam, T.; McHugh, T.J.; Tashiro, A.; Hayashi, Y. Two Functionally Distinct Serotonergic Projections into Hippocampus. J. Neurosci. 2020, 40, 4936–4944. [Google Scholar] [CrossRef]
- Liu, Y.; Aquili, L.; Wong, K.H.; Lu, Z.; Lim, L.W. Past, present, and future of serotonin-targeting therapeutics for Alzheimer’s disease: Perspectives from DNA methylation. Aging Res. Rev. 2025, 108, 102755. [Google Scholar] [CrossRef]
- Muppidi, S.; Gupta, P.K.; Vernino, S. Reversible right vagal neuropathy. Neurology 2011, 77, 1577–1579. [Google Scholar] [CrossRef]
- Ardell, J.L.; Randall, W.C. Selective vagal innervation of sinoatrial and atrioventricular nodes in canine heart. Am. J. Physiol. Heart Circ. Physiol. 1986, 251, H764–H773. [Google Scholar] [CrossRef]
- Cohn, A.E. On the differences in the effects of stimulation of the two vagus nerves on rate and conduction of the dog’s heart. J. Exp. Med. 1912, 16, 732–757. [Google Scholar] [CrossRef]
- Randall, W.; Armour, J. Regional vagosympathetic control of the heart. Am. J. Physiol. Leg. Content 1974, 227, 444–452. [Google Scholar] [CrossRef]
- Bermejo, P.; López, M.; Larraya, I.; Chamorro, J.; Cobo, J.L.; Ordóñez, S.; Vega, J.A. Innervation of the Human Cavum Conchae and Auditory Canal: Anatomical Basis for Transcutaneous Auricular Nerve Stimulation. BioMed Res. Int. 2017, 2017, 7830919. [Google Scholar] [CrossRef] [PubMed]
- Redgrave, J.; Day, D.; Leung, H.; Laud, P.J.; Ali, A.; Lindert, R.; Majid, A. Safety and tolerability of Transcutaneous Vagus Nerve stimulation in humans; a systematic review. Brain Stimul. 2018, 11, 1225–1238. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Silberstein, S.D. Vagus Nerve and Vagus Nerve Stimulation, a Comprehensive Review: Part II. Headache J. Head Face Pain 2016, 56, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Kreuzer, P.M.; Landgrebe, M.; Husser, O.; Resch, M.; Schecklmann, M.; Geisreiter, F.; Poeppl, T.B.; Prasser, S.J.; Hajak, G.; Langguth, B. Transcutaneous Vagus Nerve Stimulation: Retrospective Assessment of Cardiac Safety in a Pilot Study. Front. Psychiatry 2012, 3, 70. [Google Scholar] [CrossRef]
- Mertens, A.; Naert, L.; Miatton, M.; Poppa, T.; Carrette, E.; Gadeyne, S.; Raedt, R.; Boon, P.; Vonck, K. Transcutaneous Vagus Nerve Stimulation Does Not Affect Verbal Memory Performance in Healthy Volunteers. Front. Psychol. 2020, 11, 551. [Google Scholar] [CrossRef]
- Giraudier, M.; Ventura-Bort, C.; Weymar, M. Transcutaneous Vagus Nerve Stimulation (tVNS) Improves High-Confidence Recognition Memory but Not Emotional Word Processing. Front. Psychol. 2020, 11, 1276. [Google Scholar] [CrossRef]
- Ventura-Bort, C.; Giraudier, M.; Weymar, M. Transcutaneous Auricular Vagus Nerve Stimulation Enhances Emotional Processing and Long-Term Recognition Memory: Electrophysiological Evidence Across Two Studies. Psychophysiology 2025, 62, e70034. [Google Scholar] [CrossRef]
- Zhao, R.; Chang, M.-Y.; Cheng, C.; Tian, Q.-Q.; Yang, X.-J.; Du, M.-Y.; Cui, Y.-P.; He, Z.-Y.; Wang, F.-M.; Kong, Y.; et al. Transcutaneous auricular vagus stimulation (taVNS) improves human working memory performance under sleep deprivation stress. Behav. Brain Res. 2023, 439, 114247. [Google Scholar] [CrossRef]
- Shin, J.; Noh, S.; Park, J.; Jun, S.B.; Sung, J.E. Transcutaneous auricular vagus nerve stimulation enhanced working memory in older adults with age-related hearing loss. Sci. Rep. 2025, 15, 26629. [Google Scholar] [CrossRef]
- Thakkar, V.J.; Richardson, Z.A.; Dang, A.; Centanni, T.M. The effect of non-invasive vagus nerve stimulation on memory recall in reading: A pilot study. Behav. Brain Res. 2023, 438, 114164. [Google Scholar] [CrossRef]
- Cibulcova, V.; Koenig, J.; Jackowska, M.; Jandackova, V.K. Influence of a 2-week transcutaneous auricular vagus nerve stimulation on memory: Findings from a randomized placebo controlled trial in non-clinical adults. Clin. Auton. Res. 2024, 34, 447–462. [Google Scholar] [CrossRef]
- Fisicaro, F.; Cortese, K.; Bella, R.; Pennisi, M.; Lanza, G.; Yuasa, K.; Ugawa, Y.; Terao, Y. Effects of off-line auricular transcutaneous vagus nerve stimulation (taVNS) on a short-term memory task: A pilot study. Front. Aging Neurosci. 2025, 17, 1549167. [Google Scholar] [CrossRef]
- Kaan, E.; De Aguiar, I.; Clarke, C.; Lamb, D.G.; Williamson, J.B.; Porges, E.C. A transcutaneous vagus nerve stimulation study on verbal order memory. J. Neurolinguistics 2021, 59, 100990. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, J.; Guo, C.; He, J.; Zhang, S.; Wang, Y.; Zhao, Y.; Li, L.; Wang, J.; Hou, L.; et al. The efficacy and safety of transcutaneous auricular vagus nerve stimulation in patients with mild cognitive impairment: A double blinded randomized clinical trial. Brain Stimul. 2022, 15, 1405–1414. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Oliver, A.; Brambilla-Pisoni, C.; Domingo-Gainza, M.; Maldonado, R.; Ivorra, A.; Ozaita, A. Auricular transcutaneous vagus nerve stimulation improves memory persistence in naïve mice and in an intellectual disability mouse model. Brain Stimul. 2020, 13, 494–498. [Google Scholar] [CrossRef] [PubMed]
- Badran, B.W.; Dowdle, L.T.; Mithoefer, O.J.; LaBate, N.T.; Coatsworth, J.; Brown, J.C.; DeVries, W.H.; Austelle, C.W.; McTeague, L.M.; George, M.S. Neurophysiologic effects of transcutaneous auricular vagus nerve stimulation (taVNS) via electrical stimulation of the tragus: A concurrent taVNS/fMRI study and review. Brain Stimul. Basic Transl. Clin. Res. Neuromodulation 2018, 11, 492–500. [Google Scholar] [CrossRef]
- Frangos, E.; Ellrich, J.; Komisaruk, B.R. Non-invasive Access to the Vagus Nerve Central Projections via Electrical Stimulation of the External Ear: fMRI Evidence in Humans. Brain Stimul. Basic Transl. Clin. Res. Neuromodulation 2015, 8, 624–636. [Google Scholar] [CrossRef]
- Sclocco, R.; Garcia, R.G.; Kettner, N.W.; Fisher, H.P.; Isenburg, K.; Makarovsky, M.; Stowell, J.A.; Goldstein, J.; Barbieri, R.; Napadow, V. Stimulus frequency modulates brainstem response to respiratory-gated transcutaneous auricular vagus nerve stimulation. Brain Stimul. 2020, 13, 970–978. [Google Scholar] [CrossRef]
- van Midden, V.M.; Demšar, J.; Pirtošek, Z.; Kojović, M. The effects of transcutaneous auricular vagal nerve stimulation on cortical GABAergic and cholinergic circuits: A transcranial magnetic stimulation study. Eur. J. Neurosci. 2023, 57, 2160–2173. [Google Scholar] [CrossRef]
- Gerges, A.N.H.; Graetz, L.; Hillier, S.; Uy, J.; Hamilton, T.; Opie, G.; Vallence, A.-M.; Braithwaite, F.A.; Chamberlain, S.; Hordacre, B. Transcutaneous auricular vagus nerve stimulation modifies cortical excitability in middle-aged and older adults. Psychophysiology 2024, 62, e14584. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, J.; Li, H.; Yan, Z.; Liu, X.; Cao, J.; Park, J.; Wilson, G.; Liu, B.; Kong, J. Transcutaneous auricular vagus nerve stimulation at 1 Hz modulates locus coeruleus activity and resting state functional connectivity in patients with migraine: An fMRI study. NeuroImage Clin. 2019, 24, 101971. [Google Scholar] [CrossRef]
- Hansen, N. Memory Reinforcement and Attenuation by Activating the Human Locus Coeruleus via Transcutaneous Vagus Nerve Stimulation. Front. Neurosci. 2019, 12, 955. [Google Scholar] [CrossRef]
- Murphy, A.J.; O’Neal, A.G.; Cohen, R.A.; Lamb, D.G.; Porges, E.C.; Bottari, S.A.; Ho, B.; Trifilio, E.; DeKosky, S.T.; Heilman, K.M.; et al. The Effects of Transcutaneous Vagus Nerve Stimulation on Functional Connectivity Within Semantic and Hippocampal Networks in Mild Cognitive Impairment. Neurotherapeutics 2023, 20, 419–430. [Google Scholar] [CrossRef] [PubMed]
- Gu, Q. Neuromodulatory transmitter systems in the cortex and their role in cortical plasticity. Neuroscience 2002, 111, 815–835. [Google Scholar] [CrossRef] [PubMed]
- de Melo, P.S.; Parente, J.; Rebello-Sanchez, I.; Marduy, A.; Gianlorenco, A.C.; Kyung Kim, C.; Choi, H.; Song, J.-J.; Fregni, F. Understanding the Neuroplastic Effects of Auricular Vagus Nerve Stimulation in Animal Models of Stroke: A Systematic Review and Meta-Analysis. Neurorehabilit. Neural Repair 2023, 37, 564–576. [Google Scholar] [CrossRef] [PubMed]
- Olsen, L.; Moore, R.; Bechmann, N.; Cunningham, S.; Ethridge, V.; Rohan, J.; Hatcher-Solis, C. Vagus nerve stimulation induced cognitive enhancement: Hippocampal neuroplasticity in healthy male rats. Brain Stimul. Basic Transl. Clin. Res. Neuromodulation 2021, 14, 1703–1704. [Google Scholar] [CrossRef]
- Follesa, P.; Biggio, F.; Gorini, G.; Caria, S.; Talani, G.; Dazzi, L.; Puligheddu, M.; Marrosu, F.; Biggio, G. Vagus nerve stimulation increases norepinephrine concentration and the gene expression of BDNF and bFGF in the rat brain. Brain Res. 2007, 1179, 28–34. [Google Scholar] [CrossRef]
- Biggio, F.; Gorini, G.; Utzeri, C.; Olla, P.; Marrosu, F.; Mocchetti, I.; Follesa, P. Chronic vagus nerve stimulation induces neuronal plasticity in the rat hippocampus. Int. J. Neuropsychopharmacol. 2009, 12, 1209–1221. [Google Scholar] [CrossRef]
- Revesz, D.; Tjernstrom, M.; Ben-Menachem, E.; Thorlin, T. Effects of vagus nerve stimulation on rat hippocampal progenitor proliferation. Exp. Neurol. 2008, 214, 259–265. [Google Scholar] [CrossRef]
- Poon, C.H.; Heng, B.C.; Lim, L.W. New insights on brain-derived neurotrophic factor epigenetics: From depression to memory extinction. Ann. Acad. Sci. 2021, 1484, 9–31. [Google Scholar] [CrossRef]
- Griffith, J.S.; Mahler, H.R. DNA ticketing theory of memory. Nature 1969, 223, 580–582. [Google Scholar] [CrossRef]
- Holliday, R. Is there an epigenetic component in long-term memory? J. Theor. Biol. 1999, 200, 339–341. [Google Scholar] [CrossRef]
- Tillotson, R.; Cholewa-Waclaw, J.; Chhatbar, K.; Connelly, J.C.; Kirschner, S.A.; Webb, S.; Koerner, M.V.; Selfridge, J.; Kelly, D.A.; De Sousa, D.; et al. Neuronal non-CG methylation is an essential target for MeCP2 function. Mol. Cell 2021, 81, 1260–1275.e1212. [Google Scholar] [CrossRef] [PubMed]
- McKinney, B.C.; Lin, C.-W.; Rahman, T.; Oh, H.; Lewis, D.A.; Tseng, G.; Sibille, E. DNA methylation in the human frontal cortex reveals a putative mechanism for age-by-disease interactions. Transl. Psychiatry 2019, 9, 39. [Google Scholar] [CrossRef] [PubMed]
- Shireby, G.; Dempster, E.L.; Policicchio, S.; Smith, R.G.; Pishva, E.; Chioza, B.; Davies, J.P.; Burrage, J.; Lunnon, K.; Seiler Vellame, D.; et al. DNA methylation signatures of Alzheimer’s disease neuropathology in the cortex are primarily driven by variation in non-neuronal cell-types. Nat. Commun. 2022, 13, 5620. [Google Scholar] [CrossRef] [PubMed]
- Desplats, P.; Spencer, B.; Coffee, E.; Patel, P.; Michael, S.; Patrick, C.; Adame, A.; Rockenstein, E.; Masliah, E. Alpha-synuclein sequesters Dnmt1 from the nucleus: A novel mechanism for epigenetic alterations in Lewy body diseases. J. Biol. Chem. 2011, 286, 9031–9037. [Google Scholar] [CrossRef]
- Satterstrom, F.K.; Kosmicki, J.A.; Wang, J.; Breen, M.S.; De Rubeis, S.; An, J.-Y.; Peng, M.; Collins, R.; Grove, J.; Klei, L.; et al. Large-Scale Exome Sequencing Study Implicates Both Developmental and Functional Changes in the Neurobiology of Autism. Cell 2020, 180, 568–584.e523. [Google Scholar] [CrossRef]
- Sanders, S.J.; He, X.; Willsey, A.J.; Ercan-Sencicek, A.G.; Samocha, K.E.; Cicek, A.E.; Murtha, M.T.; Bal, V.H.; Bishop, S.L.; Dong, S.; et al. Insights into Autism Spectrum Disorder Genomic Architecture and Biology from 71 Risk Loci. Neuron 2015, 87, 1215–1233. [Google Scholar] [CrossRef]
- Tatton-Brown, K.; Seal, S.; Ruark, E.; Harmer, J.; Ramsay, E.; del Vecchio Duarte, S.; Zachariou, A.; Hanks, S.; O’Brien, E.; Aksglaede, L.; et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat. Genet. 2014, 46, 385–388. [Google Scholar] [CrossRef]
- Armstrong, M.J.; Jin, Y.; Vattathil, S.M.; Huang, Y.; Schroeder, J.P.; Bennet, D.A.; Qin, Z.S.; Wingo, T.S.; Jin, P. Role of TET1-mediated epigenetic modulation in Alzheimer’s disease. Neurobiol. Dis. 2023, 185, 106257. [Google Scholar] [CrossRef]
- Cochran, J.N.; Geier, E.G.; Bonham, L.W.; Newberry, J.S.; Amaral, M.D.; Thompson, M.L.; Lasseigne, B.N.; Karydas, A.M.; Roberson, E.D.; Cooper, G.M.; et al. Non-coding and Loss-of-Function Coding Variants in TET2 are Associated with Multiple Neurodegenerative Diseases. Am. J. Hum. Genet. 2020, 106, 632–645. [Google Scholar] [CrossRef] [PubMed]
- Beck, D.B.; Petracovici, A.; He, C.; Moore, H.W.; Louie, R.J.; Ansar, M.; Douzgou, S.; Sithambaram, S.; Cottrell, T.; Santos-Cortez, R.L.P.; et al. Delineation of a Human Mendelian Disorder of the DNA Demethylation Machinery: TET3 Deficiency. Am. J. Hum. Genet. 2020, 106, 234–245. [Google Scholar] [CrossRef] [PubMed]
- Rose, S.A.; Djukic, A.; Jankowski, J.J.; Feldman, J.F.; Fishman, I.; Valicenti-Mcdermott, M. Rett syndrome: An eye-tracking study of attention and recognition memory. Dev. Med. Child Neurol. 2013, 55, 364–371. [Google Scholar] [CrossRef]
- Brima, T.; Molholm, S.; Molloy, C.J.; Sysoeva, O.V.; Nicholas, E.; Djukic, A.; Freedman, E.G.; Foxe, J.J. Auditory sensory memory span for duration is severely curtailed in females with Rett syndrome. Transl. Psychiatry 2019, 9, 130. [Google Scholar] [CrossRef]
- Fabio, R.A.; Antonietti, A.; Castelli, I.; Marchetti, A. Attention and communication in Rett Syndrome. Res. Autism Spectr. Disord. 2009, 3, 329–335. [Google Scholar] [CrossRef]
- Elefant, C.; Wigram, T. Learning ability in children with Rett syndrome. Brain Dev. 2005, 27, S97–S101. [Google Scholar] [CrossRef]
- Lee, S.; Kim, T.K.; Choi, J.E.; Choi, Y.; You, M.; Ryu, J.; Chun, Y.L.; Ham, S.; Hyeon, S.J.; Ryu, H.; et al. Dysfunction of striatal MeCP2 is associated with cognitive decline in a mouse model of Alzheimer’s disease. Theranostics 2022, 12, 1404–1418. [Google Scholar] [CrossRef] [PubMed]
- Van Esch, H.; Bauters, M.; Ignatius, J.; Jansen, M.; Raynaud, M.; Hollanders, K.; Lugtenberg, D.; Bienvenu, T.; Jensen, L.R.; Gécz, J.; et al. Duplication of the MECP2 Region Is a Frequent Cause of Severe Mental Retardation and Progressive Neurological Symptoms in Males. Am. J. Hum. Genet. 2005, 77, 442–453. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Zhou, Y.; Campbell, S.L.; Le, T.; Li, E.; Sweatt, J.D.; Silva, A.J.; Fan, G. Dnmt1 and Dnmt3a maintain DNA methylation and regulate synaptic function in adult forebrain neurons. Nat. Neurosci. 2010, 13, 423–430. [Google Scholar] [CrossRef] [PubMed]
- Morris, M.J.; Adachi, M.; Na, E.S.; Monteggia, L.M. Selective role for DNMT3a in learning and memory. Neurobiol. Learn. Mem. 2014, 115, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Kupke, J.; Klimmt, J.; Mudlaff, F.; Schwab, M.; Lutsik, P.; Plass, C.; Sticht, C.; Oliveira, A.M.M. Dnmt3a1 regulates hippocampus-dependent memory via the downstream target Nrp1. Neuropsychopharmacology 2024, 49, 1528–1539. [Google Scholar] [CrossRef]
- Zhang, R.-R.; Cui, Q.-Y.; Murai, K.; Lim, Y.C.; Smith, Z.D.; Jin, S.; Ye, P.; Rosa, L.; Lee, Y.K.; Wu, H.-P.; et al. Tet1 Regulates Adult Hippocampal Neurogenesis and Cognition. Cell Stem Cell 2013, 13, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Gontier, G.; Iyer, M.; Shea, J.M.; Bieri, G.; Wheatley, E.G.; Ramalho-Santos, M.; Villeda, S.A. Tet2 Rescues Age-Related Regenerative Decline and Enhances Cognitive Function in the Adult Mouse Brain. Cell Rep. 2018, 22, 1974–1981. [Google Scholar] [CrossRef]
- Antunes, C.; Da Silva, J.D.; Guerra-Gomes, S.; Alves, N.D.; Ferreira, F.; Loureiro-Campos, E.; Branco, M.R.; Sousa, N.; Reik, W.; Pinto, L.; et al. Tet3 ablation in adult brain neurons increases anxiety-like behavior and regulates cognitive function in mice. Mol. Psychiatry 2021, 26, 1445–1457. [Google Scholar] [CrossRef]
- Leach, P.T.; Poplawski, S.G.; Kenney, J.W.; Hoffman, B.; Liebermann, D.A.; Abel, T.; Gould, T.J. Gadd45b knockout mice exhibit selective deficits in hippocampus-dependent long-term memory. Learn. Mem. 2012, 19, 319–324. [Google Scholar] [CrossRef]
- Sultan, F.A.; Wang, J.; Tront, J.; Liebermann, D.A.; Sweatt, J.D. Genetic deletion of Gadd45b, a regulator of active DNA demethylation, enhances long-term memory and synaptic plasticity. J. Neurosci. 2012, 32, 17059–17066. [Google Scholar] [CrossRef]
- Moretti, P.; Levenson, J.M.; Battaglia, F.; Atkinson, R.; Teague, R.; Antalffy, B.; Armstrong, D.; Arancio, O.; Sweatt, J.D.; Zoghbi, H.Y. Learning and Memory and Synaptic Plasticity Are Impaired in a Mouse Model of Rett Syndrome. J. Neurosci. 2006, 26, 319–327. [Google Scholar] [CrossRef]
- He, L.; Caudill, M.S.; Jing, J.; Wang, W.; Sun, Y.; Tang, J.; Jiang, X.; Zoghbi, H.Y. A weakened recurrent circuit in the hippocampus of Rett syndrome mice disrupts long-term memory representations. Neuron 2022, 110, 1689–1699.e1686. [Google Scholar] [CrossRef]
- Samaco, R.C.; McGraw, C.M.; Ward, C.S.; Sun, Y.; Neul, J.L.; Zoghbi, H.Y. Female Mecp2+/− mice display robust behavioral deficits on two different genetic backgrounds providing a framework for pre-clinical studies. Hum. Mol. Genet. 2012, 22, 96–109. [Google Scholar] [CrossRef]
- Asaka, Y.; Jugloff, D.G.M.; Zhang, L.; Eubanks, J.H.; Fitzsimonds, R.M. Hippocampal synaptic plasticity is impaired in the Mecp2-null mouse model of Rett syndrome. Neurobiol. Dis. 2006, 21, 217–227. [Google Scholar] [CrossRef]
- Na, E.S.; Nelson, E.D.; Adachi, M.; Autry, A.E.; Mahgoub, M.A.; Kavalali, E.T.; Monteggia, L.M. A Mouse Model for MeCP2 Duplication Syndrome: MeCP2 Overexpression Impairs Learning and Memory and Synaptic Transmission. J. Neurosci. 2012, 32, 3109–3117. [Google Scholar] [CrossRef]
- Simmons, R.K.; Stringfellow, S.A.; Glover, M.E.; Wagle, A.A.; Clinton, S.M. DNA methylation markers in the postnatal developing rat brain. Brain Res. 2013, 1533, 26–36. [Google Scholar] [CrossRef]
- Feng, J.; Chang, H.; Li, E.; Fan, G. Dynamic expression of de novo DNA methyltransferases Dnmt3a and Dnmt3b in the central nervous system. J. Neurosci. Res. 2005, 79, 734–746. [Google Scholar] [CrossRef] [PubMed]
- Mitchnick, K.A.; Creighton, S.; O’Hara, M.; Kalisch, B.E.; Winters, B.D. Differential contributions of de novo and maintenance DNA methyltransferases to object memory processing in the rat hippocampus and perirhinal cortex—A double dissociation. Eur. J. Neurosci. 2015, 41, 773–786. [Google Scholar] [CrossRef]
- Chen, T.; Ueda, Y.; Xie, S.; Li, E. A novel Dnmt3a isoform produced from an alternative promoter localizes to euchromatin and its expression correlates with active de novo methylation. J. Biol. Chem. 2002, 277, 38746–38754. [Google Scholar] [CrossRef]
- Wu, H.; Coskun, V.; Tao, J.; Xie, W.; Ge, W.; Yoshikawa, K.; Li, E.; Zhang, Y.; Sun, Y.E. Dnmt3a-Dependent Nonpromoter DNA Methylation Facilitates Transcription of Neurogenic Genes. Science 2010, 329, 444–448. [Google Scholar] [CrossRef]
- Zocher, S.; Overall, R.W.; Berdugo-Vega, G.; Rund, N.; Karasinsky, A.; Adusumilli, V.S.; Steinhauer, C.; Scheibenstock, S.; Händler, K.; Schultze, J.L.; et al. De novo DNA methylation controls neuronal maturation during adult hippocampal neurogenesis. Eur. Mol. Biol. Organ. 2021, 40, e107100. [Google Scholar] [CrossRef]
- Lubin, F.D.; Roth, T.L.; Sweatt, J.D. Epigenetic regulation of BDNF gene transcription in the consolidation of fear memory. J. Neurosci. 2008, 28, 10576–10586. [Google Scholar] [CrossRef]
- Kaas, G.A.; Zhong, C.; Eason, D.E.; Ross, D.L.; Vachhani, R.V.; Ming, G.L.; King, J.R.; Song, H.; Sweatt, J.D. TET1 controls CNS 5-methylcytosine hydroxylation, active DNA demethylation, gene transcription, and memory formation. Neuron 2013, 79, 1086–1093. [Google Scholar] [CrossRef]
- Bayraktar, G.; Kreutz, M.R. The Role of Activity-Dependent DNA Demethylation in the Adult Brain and in Neurological Disorders. Front. Mol. Neurosci. 2018, 11, 169. [Google Scholar] [CrossRef] [PubMed]
- Mellén, M.; Ayata, P.; Heintz, N. 5-hydroxymethylcytosine accumulation in postmitotic neurons results in functional demethylation of expressed genes. Proc. Natl. Acad. Sci. USA 2017, 114, e7812–e7821. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yao, B.; Chen, L.; Kang, Y.; Li, Y.; Cheng, Y.; Li, L.; Lin, L.; Wang, Z.; Wang, M.; et al. Ten-eleven translocation 2 interacts with forkhead box O3 and regulates adult neurogenesis. Nat. Commun. 2017, 8, 15903. [Google Scholar] [CrossRef]
- Montalbán-Loro, R.; Lozano-Ureña, A.; Ito, M.; Krueger, C.; Reik, W.; Ferguson-Smith, A.C.; Ferrón, S.R. TET3 prevents terminal differentiation of adult NSCs by a non-catalytic action at Snrpn. Nat. Commun. 2019, 10, 1726. [Google Scholar] [CrossRef] [PubMed]
- Rai, K.; Huggins, I.J.; James, S.R.; Karpf, A.R.; Jones, D.A.; Cairns, B.R. DNA Demethylation in Zebrafish Involves the Coupling of a Deaminase, a Glycosylase, and Gadd45. Cell 2008, 135, 1201–1212. [Google Scholar] [CrossRef]
- Wu, X.; Zhang, Y. TET-mediated active DNA demethylation: Mechanism, function and beyond. Nat. Rev. Genet. 2017, 18, 517–534. [Google Scholar] [CrossRef] [PubMed]
- Kriaucionis, S.; Bird, A. The major form of MeCP2 has a novel N-terminus generated by alternative splicing. Nucleic Acids Res. 2004, 32, 1818–1823. [Google Scholar] [CrossRef]
- Mnatzakanian, G.N.; Lohi, H.; Munteanu, I.; Alfred, S.E.; Yamada, T.; MacLeod, P.J.; Jones, J.R.; Scherer, S.W.; Schanen, N.C.; Friez, M.J.; et al. A previously unidentified MECP2 open reading frame defines a new protein isoform relevant to Rett syndrome. Nat. Genet. 2004, 36, 339–341. [Google Scholar] [CrossRef]
- Chahrour, M.; Jung, S.Y.; Shaw, C.; Zhou, X.; Wong, S.T.C.; Qin, J.; Zoghbi, H.Y. MeCP2, a Key Contributor to Neurological Disease, Activates and Represses Transcription. Science 2008, 320, 1224–1229. [Google Scholar] [CrossRef]
- Szulwach, K.E.; Li, X.; Smrt, R.D.; Li, Y.; Luo, Y.; Lin, L.; Santistevan, N.J.; Li, W.; Zhao, X.; Jin, P. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell Biol. 2010, 189, 127–141. [Google Scholar] [CrossRef]
- Lyst, M.J.; Ekiert, R.; Ebert, D.H.; Merusi, C.; Nowak, J.; Selfridge, J.; Guy, J.; Kastan, N.R.; Robinson, N.D.; de Lima Alves, F.; et al. Rett syndrome mutations abolish the interaction of MeCP2 with the NCoR/SMRT co-repressor. Nat. Neurosci. 2013, 16, 898–902. [Google Scholar] [CrossRef] [PubMed]
- Subbanna, S.; Nagre, N.N.; Shivakumar, M.; Umapathy, N.S.; Psychoyos, D.; Basavarajappa, B.S. Ethanol induced acetylation of histone at G9a exon1 and G9a-mediated histone H3 dimethylation leads to neurodegeneration in neonatal mice. Neuroscience 2014, 258, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Shahbazian, M.D.; Antalffy, B.; Armstrong, D.L.; Zoghbi, H.Y. Insight into Rett syndrome: MeCP2 levels display tissue- and cell-specific differences and correlate with neuronal maturation. Hum. Mol. Genet. 2002, 11, 115–124. [Google Scholar] [CrossRef]
- Skene, P.J.; Illingworth, R.S.; Webb, S.; Kerr, A.R.W.; James, K.D.; Turner, D.J.; Andrews, R.; Bird, A.P. Neuronal MeCP2 Is Expressed at Near Histone-Octamer Levels and Globally Alters the Chromatin State. Mol. Cell 2010, 37, 457–468. [Google Scholar] [CrossRef]
- Olson, C.O.; Zachariah, R.M.; Ezeonwuka, C.D.; Liyanage, V.R.; Rastegar, M. Brain region-specific expression of MeCP2 isoforms correlates with DNA methylation within Mecp2 regulatory elements. PLoS ONE 2014, 9, e90645. [Google Scholar] [CrossRef]
- Li, H.; Zhong, X.; Chau, K.F.; Santistevan, N.J.; Guo, W.; Kong, G.; Li, X.; Kadakia, M.; Masliah, J.; Chi, J.; et al. Cell cycle-linked MeCP2 phosphorylation modulates adult neurogenesis involving the Notch signalling pathway. Nat. Commun. 2014, 5, 5601. [Google Scholar] [CrossRef]
- Smrt, R.D.; Eaves-Egenes, J.; Barkho, B.Z.; Santistevan, N.J.; Zhao, C.; Aimone, J.B.; Gage, F.H.; Zhao, X. Mecp2 deficiency leads to delayed maturation and altered gene expression in hippocampal neurons. Neurobiol. Dis. 2007, 27, 77–89. [Google Scholar] [CrossRef]
- Sun, Y.; Gao, Y.; Tidei, J.J.; Shen, M.; Hoang, J.T.; Wagner, D.F.; Zhao, X. Loss of MeCP2 in immature neurons leads to impaired network integration. Hum. Mol. Genet. 2018, 28, 245–257. [Google Scholar] [CrossRef]
- Khoshnan, A.; Patterson, P.H. Elevated IKKα Accelerates the Differentiation of Human Neuronal Progenitor Cells and Induces MeCP2-Dependent BDNF Expression. PLoS ONE 2012, 7, e41794. [Google Scholar] [CrossRef][Green Version]
- Tsujimura, K.; Abematsu, M.; Kohyama, J.; Namihira, M.; Nakashima, K. Neuronal differentiation of neural precursor cells is promoted by the methyl-CpG-binding protein MeCP2. Exp. Neurol. 2009, 219, 104–111. [Google Scholar] [CrossRef]
- Chen, L.; Chen, K.; Lavery, L.A.; Baker, S.A.; Shaw, C.A.; Li, W.; Zoghbi, H.Y. MeCP2 binds to non-CG methylated DNA as neurons mature, influencing transcription and the timing of onset for Rett syndrome. Proc. Natl. Acad. Sci. USA 2015, 112, 5509–5514. [Google Scholar] [CrossRef]
- Bruel-Jungerman, E.; Davis, S.; Rampon, C.; Laroche, S. Long-Term Potentiation Enhances Neurogenesis in the Adult Dentate Gyrus. J. Neurosci. 2006, 26, 5888–5893. [Google Scholar] [CrossRef] [PubMed]
- Barker, G.R.I.; Warburton, E.C.; Koder, T.; Dolman, N.P.; More, J.C.A.; Aggleton, J.P.; Bashir, Z.I.; Auberson, Y.P.; Jane, D.E.; Brown, M.W. The Different Effects on Recognition Memory of Perirhinal Kainate and NMDA Glutamate Receptor Antagonism: Implications for Underlying Plasticity Mechanisms. J. Neurosci. 2006, 26, 3561–3566. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Miyashita, T.; Fu, T.F.; Lin, W.Y.; Wu, C.L.; Pyzocha, L.; Lin, I.R.; Saitoe, M.; Tully, T.; Chiang, A.S. NMDA receptors mediate olfactory learning and memory in Drosophila. Curr. Biol. 2005, 15, 603–615. [Google Scholar] [CrossRef]
- Bayraktar, G.; Yuanxiang, P.; Confettura, A.D.; Gomes, G.M.; Raza, S.A.; Stork, O.; Tajima, S.; Suetake, I.; Karpova, A.; Yildirim, F.; et al. Synaptic control of DNA methylation involves activity-dependent degradation of DNMT3A1 in the nucleus. Neuropsychopharmacology 2020, 45, 2120–2130. [Google Scholar] [CrossRef]
- Pusalkar, M.; Ghosh, S.; Jaggar, M.; Husain, B.F.A.; Galande, S.; Vaidya, V.A. Acute and Chronic Electroconvulsive Seizures (ECS) Differentially Regulate the Expression of Epigenetic Machinery in the Adult Rat Hippocampus. Int. J. Neuropsychopharmacol. 2016, 19, pyw040. [Google Scholar] [CrossRef]
- Li, X.; Wei, W.; Zhao, Q.-Y.; Widagdo, J.; Baker-Andresen, D.; Flavell, C.R.; D’Alessio, A.; Zhang, Y.; Bredy, T.W. Neocortical Tet3-mediated accumulation of 5-hydroxymethylcytosine promotes rapid behavioral adaptation. Proc. Natl. Acad. Sci. USA 2014, 111, 7120–7125. [Google Scholar] [CrossRef]
- Ma, D.K.; Jang, M.-H.; Guo, J.U.; Kitabatake, Y.; Chang, M.-l.; Pow-anpongkul, N.; Flavell, R.A.; Lu, B.; Ming, G.-l.; Song, H. Neuronal Activity–Induced Gadd45b Promotes Epigenetic DNA Demethylation and Adult Neurogenesis. Science 2009, 323, 1074–1077. [Google Scholar] [CrossRef]
- Pohodich, A.E.; Yalamanchili, H.; Raman, A.T.; Wan, Y.-W.; Gundry, M.; Hao, S.; Jin, H.; Tang, J.; Liu, Z.; Zoghbi, H.Y. Forniceal deep brain stimulation induces gene expression and splicing changes that promote neurogenesis and plasticity. eLife 2018, 7, e34031. [Google Scholar] [CrossRef]
- Yu, H.; Su, Y.; Shin, J.; Zhong, C.; Guo, J.U.; Weng, Y.-L.; Gao, F.; Geschwind, D.H.; Coppola, G.; Ming, G.-l.; et al. Tet3 regulates synaptic transmission and homeostatic plasticity via DNA oxidation and repair. Nat. Neurosci. 2015, 18, 836–843. [Google Scholar] [CrossRef]
- Rudenko, A.; Dawlaty, M.M.; Seo, J.; Cheng, A.W.; Meng, J.; Le, T.; Faull, K.F.; Jaenisch, R.; Tsai, L.-H. Tet1 Is Critical for Neuronal Activity-Regulated Gene Expression and Memory Extinction. Neuron 2013, 79, 1109–1122. [Google Scholar] [CrossRef]
- Tao, J.; Hu, K.; Chang, Q.; Wu, H.; Sherman, N.E.; Martinowich, K.; Klose, R.J.; Schanen, C.; Jaenisch, R.; Wang, W.; et al. Phosphorylation of MeCP2 at Serine 80 regulates its chromatin association and neurological function. Proc. Natl. Acad. Sci. USA 2009, 106, 4882–4887. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Hong, E.J.; Cohen, S.; Zhao, W.N.; Ho, H.Y.; Schmidt, L.; Chen, W.G.; Lin, Y.; Savner, E.; Griffith, E.C.; et al. Brain-specific phosphorylation of MeCP2 regulates activity-dependent Bdnf transcription, dendritic growth, and spine maturation. Neuron 2006, 52, 255–269. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.G.; Chang, Q.; Lin, Y.; Meissner, A.; West, A.E.; Griffith, E.C.; Jaenisch, R.; Greenberg, M.E. Derepression of BDNF Transcription Involves Calcium-Dependent Phosphorylation of MeCP2. Science 2003, 302, 885–889. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.; Gabel, H.W.; Hemberg, M.; Hutchinson, A.N.; Sadacca, L.A.; Ebert, D.H.; Harmin, D.A.; Greenberg, R.S.; Verdine, V.K.; Zhou, Z.; et al. Genome-Wide Activity-Dependent MeCP2 Phosphorylation Regulates Nervous System Development and Function. Neuron 2011, 72, 72–85. [Google Scholar] [CrossRef]
- Engeroff, K.; Warm, D.; Bittner, S.; Blanquie, O. Different activity patterns control various stages of Reelin synthesis in the developing neocortex. Cereb. Cortex 2023, 33, 9376–9386. [Google Scholar] [CrossRef]
- Kundakovic, M.; Chen, Y.; Costa, E.; Grayson, D.R. DNA Methyltransferase Inhibitors Coordinately Induce Expression of the Human Reelin and Glutamic Acid Decarboxylase 67 Genes. Mol. Pharmacol. 2007, 71, 644–653. [Google Scholar] [CrossRef]
- Ebert, D.H.; Gabel, H.W.; Robinson, N.D.; Kastan, N.R.; Hu, L.S.; Cohen, S.; Navarro, A.J.; Lyst, M.J.; Ekiert, R.; Bird, A.P.; et al. Activity-dependent phosphorylation of MeCP2 threonine 308 regulates interaction with NCoR. Nature 2013, 499, 341–345. [Google Scholar] [CrossRef]
- Tai, D.J.C.; Liu, Y.C.; Hsu, W.L.; Ma, Y.L.; Cheng, S.J.; Liu, S.Y.; Lee, E.H.Y. MeCP2 SUMOylation rescues Mecp2-mutant-induced behavioural deficits in a mouse model of Rett syndrome. Nat. Commun. 2016, 7, 10552. [Google Scholar] [CrossRef]
- Zhong, H.; Zhu, J.; Liu, S.; Zhou, D.; Long, Q.; Wu, C.; Zhao, B.; Cheng, C.; Yang, Y.; Wu, Q.; et al. Linking DNA methylation in brain regions to Alzheimer’s disease risk: A Mendelian randomization study. Hum. Mol. Genet. 2025, 34, 1026–1033. [Google Scholar] [CrossRef]
| Study | Memory Task | taVNS Delivery | Stimulation Site | Subjects | Subject Condition | Effect of taVNS |
|---|---|---|---|---|---|---|
| Human studies | ||||||
| [13] | Associative memory | During/after learning (online) | Inner side of the tragus | 50M/50F (Mean age: 60.57 ± 2.54) | Older adults | Higher correct hits in face-name task (p < 0.05) |
| [57] | Recognition memory | After learning | Left cymba conchae | Young: 49M/51F (Mean age: 22.20 ± 1.97) Old: 29M/70F (Mean age: 55.13 ± 6.59) | Healthy adults | No effect on recall or recognition (p > 0.05) |
| [58] | Recognition memory | During learning (online) | Left cymba conchae | 14M/46F (Mean age: 23.39 ± 4.67) | Healthy adults | Increased hits for words remembered with higher subjective confidence (p < 0.05) |
| [65] | Item order memory | During memory task | Left tragus | Control: 10M/23F (Mean age: 19.8) Sham first: 6M/9F (Mean age: 20.4) taVNS first: 4M/10F (Mean age: 20.4) | Healthy adults | Higher accuracy on the order memory task in taVNS groups (p < 0.01) |
| [20] | Spatial working memory | Before/during testing (online/offline) | Left cymba conchae | 36M/24F (Mean age: 19.90 ± 1.49) | Healthy adults | Offline taVNS significantly increased hits in spatial 3-back task (p < 0.05) |
| [66] | Auditory verbal learning test-HuaShan version (AVLT-H) | 24-week intervention | Auricular acupoints: heart (concha, CO15) and kidney (CO10) | taVNS: 5M/20F (Mean age: 66.9 ± 3.66) sham: 4M/23F (Mean age: 67 ± 4.36) | Patients with mild cognitive impairment | Significantly increased N5 (immediate recall) and N7 (delayed recall) for taVNS group (p < 0.001) after intervention |
| [62] | Verbal short-term working memory | During learning | Left posterior tragus | taVNS: 3M/9F (Mean age: 19.17 ± 0.88) sham: 4M/8F (Mean age: 20.50 ± 2.48) | Healthy adults | Improved performance in memory questions (p = 0.007) |
| [60] | Working memory | Before testing | Left cymba conchae | 19M/16F (Mean age: 21.26 ± 1.90) | Healthy adults with 24-hour sleep deprivation | Improved accuracy rate in spatial 3-back tasks (p < 0.05) |
| [63] | Rey Auditory Verbal Learning Test (RAVLT) | 2-week intervention | Left tragus | 30M/46F (Mean age: 48.32) | Healthy adults | Significantly improved immediate recall and short-term memory score in taVNS group after intervention (p < 0.05) |
| [64] | Short-term memory | Before testing | Left tragus | 9M/9F (Mean age: 73.5 ± 4.71) | Older adults | Significantly reduced total error in 7, 8, 9-digit span tasks after 5-min taVNS (p < 0.001) |
| [59] | Recognition memory | During learning | Left cymba conchae | 13M/49F/3NS (Mean age: 24.24 ± 5.25) | Healthy adults | Improved recollection-based memory performance in taVNS group compared to sham 1 week after learning (p = 0.001) |
| [61] | Working memory | During memory task | Cymba conchae (left/right counterbalance) | Typical hearing: 23M/13F (Mean age: 64.03 ± 3.72) Hearing impaired: 14M/6F (Mean age: 65.55 ± 3.66) | Older adults with age-related hearing loss | Improved memory performance in 2-back test for subjects with hearing impairment (p < 0.05) |
| Animal studies | ||||||
| [67] | Novel object-recognition memory (NORT) for long-term memory | After familiarization phase | Left concha | Naive young-adult male CD-1 mice (10–12 weeks old) | Improved object-recognition memory performance at 48 h (p = 0.01) | |
| Fmr1KO young-adult male CD-1 mice (fragile X syndrome model) | Improved object-recognition memory performance at 48 h (p = 0.0003) | |||||
| [18] | NORT, Y-maze test for spatial memory | Before behavioral tests | Left cymba concha | Vascular Cognitive Impairment model: 8-week-old male C57BL/6 mice with common carotid arteries (CCA) dissected | Higher discrimination index scores in the NOR test (p < 0.001) and rates of spontaneous alternations in the Y-maze test (p = 0.008) | |
| [19] | Spatial learning and memory (MWM), NORT | Before and during days of behavioral tests | Bilateral auricular concha | 6-month-old APP/PS1 mice (Alzheimer’s model) | Longer target quadrant time and more platform crossing in MWM (p < 0.05); No change in recognition memory | |
| 6-month-old C57BL/6 mice | No change in long-term spatial memory and recognition memory | |||||
| DNAm-Related Protein | Function | Disease/Condition | Neurocognitive Implications | Ref |
|---|---|---|---|---|
| Human studies | ||||
| DNMT1 | Methylation maintenance | Parkinson’s disease (PD) | Decreased nuclear expression in postmortem human brain samples from PD and DLB patients | [88] |
| Dementia with Lewy Bodies (DLB) | ||||
| DNMT3a | De novo methylation (converting C to 5-mC) | Autism spectrum disorder (ASD) | Heterozygous mutation of DNMT3A gene is linked to ASD | [89,90] |
| Tatton-Brown–Rahman syndrome (TBRS) | Heterozygous DNMT3A variant identified in TBRS patients | [91] | ||
| Tet1 | Converting 5-mC to 5-hmC | Early-onset Alzheimer’s disease (EOAD) | Loss-of-function mutation of TET1 gene is linked to EOAD | [92] |
| Tet2 | Early-onset Alzheimer’s disease (EOAD) | Loss-of-function TET2 variant is linked to EOAD and FTD | [93] | |
| Frontotemporal dementia (FTD) | ||||
| Tet3 | Intellectual disability (ID) | Haploinsufficiency of TET3 due to heterozygous mutation links to ID | [94] | |
| MeCP2 | 5-mC and 5-hmC binder and reader (can also bind 5-fC and 5-acC with lower affinity) | Rett syndrome | Loss-of-function mutation in MeCP2 links to Rett syndrome, characterized by impaired recognition memory, sensory memory, selective attention, and skill acquisition | [95,96,97,98] |
| Alzheimer’s disease (AD) | Downregulated MeCP2 mRNA expression in AD patient hippocampus | [31] | ||
| Upregulated MeCP2 protein expression in AD patient forebrain (putamen, cortex) | [99] | |||
| MeCP2 duplication syndrome | Gain-of-function in MECP2 gene copy causes intellectual disability, developmental delays, and speech difficulties | [100] | ||
| Animal studies | ||||
| DNMT1 and/or DNMT3a | DNMT1: methylation maintenance DNMT3a: De novo methylation (C → 5-mC) | Mice with double knockout of DNMT1 and DNMT3a in neurons | Only the double knockout mice have impaired spatial learning and contextual fear memory | [101] |
| Mice with single knockout of DNMT1 or DNMT3a in neurons | ||||
| Mice with single knockout of DNMT1 or DNMT3a in neurons | Only DNMT3a knockout mice have impaired contextual fear memory and spatial–object recognition memory | [102] | ||
| Aged mice with/without hippocampal DNMT3a2 overexpression | Age-related decline of DNMT1 and DNMT3a2 mRNA expression in hippocampus and cortex is linked to impaired object-location long-term memory and contextual fear memory. Overexpressing DNMT3a2 in hippocampus rescued age-related memory decline | [30] | ||
| Young mice with hippocampal DNMT3a2 knockdown | Depletion of DNMT3a2 in hippocampus impaired object-location long-term memory and contextual fear memory | |||
| Young mice with hippocampal DNMT3a1 knockdown | Depletion of DNMT3a2 in hippocampus impaired object-location long-term memory and contextual fear memory The memory deficit can be rescued by overexpressing its downstream mediator, neuropilin-1 (Nrp1) | [103] | ||
| Tet1 | Converting 5-mC to 5-hmC | Young mice with Tet1 mutation | Impairment in spatial learning and short-term memory Altered expression and methylation of genes involved in neural progenitor proliferation, neuroprotection, and mitochondria function, e.g., Galanin, Ng2, Ngb, Kctd14, and Atp5h | [104] |
| 5xFAD mice (AD model) with/without Tet1 heterozygous mutation | Slightly worse contextual fear memory in mice with Tet1 mutation | [92] | ||
| Tet2 | Adult mature mice | Age-associated hippocampal Tet2 mRNA decline with loss in differentially 5-hydroxymethylated regions (DhMRs) for neurogenic genes. Overexpressing Tet2 in hippocampus improved associative fear memory acquisition | [105] | |
| Young mice with hippocampal Tet2 knockdown | Impaired spatial learning and long-term contextual fear memory | |||
| Tet3 | Young mice with conditional knockout of Tet3 in neuronal cells | Impaired spatial learning | [106] | |
| Gadd45b |
| Gadd45b knockout mice | Impaired long-term contextual fear conditioning | [107] |
| Enhanced contextual fear and long-term spatial memory, with better long-term potentiation in hippocampus | [108] | |||
| Gadd45γ | Aged mice | Reduced Gadd45γ mRNA level in dorsal hippocampus (while not Gadd45α and Gadd45β) | [32] | |
| Young mice with hippocampal Gadd45γ knockdown | Impaired object–space memory and contextual fear memory | |||
| MeCP2 | 5-mC and 5-hmC binder and reader (can also bind 5-fC and 5-acC with lower affinity) | Mice with Mecp2 loss-of-function mutation (Rett syndrome model) | Impaired spatial memory, contextual fear memory, and social memory related to impaired electrophysiology and excitatory neuroplasticity in the hippocampus | [109,110,111,112] |
| Mice with MeCP2 overexpression in neurons (MeCP2 duplication syndrome model) | Impaired contextual fear memory and novel object recognition, associated with deficits in short-term synaptic plasticity and LTP | [113] | ||
| Senescence-accelerated mouse prone 8 (SAMP8) (AD model) | Reduced hippocampal MeCP2 expression (mRNA and protein level) Overexpressing hippocampal MeCP2 rescued deficits in spatial learning and retention memory | [31] | ||
| Amyloid precursor protein (APP)/presenilin1 (PS1) transgenic mice (AD model) | Increased striatal MeCP2 expression (mRNA and protein level). Knockdown of striatal MeCP2 rescued deficits in social memory and spatial memory in 10-month-old mice | [99] | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yee, P.O.P.; Tsui, K.C.; Fung, M.L.; Heng, B.C.; Kocabicak, E.; Jahanshahi, A.; Temel, Y.; Blokland, A.; Aquili, L.; Kalueff, A.; et al. DNA Methylation: A Potential Mediator of the Memory Regulatory Effects of taVNS. Cells 2025, 14, 1327. https://doi.org/10.3390/cells14171327
Yee POP, Tsui KC, Fung ML, Heng BC, Kocabicak E, Jahanshahi A, Temel Y, Blokland A, Aquili L, Kalueff A, et al. DNA Methylation: A Potential Mediator of the Memory Regulatory Effects of taVNS. Cells. 2025; 14(17):1327. https://doi.org/10.3390/cells14171327
Chicago/Turabian StyleYee, Pak On Patrick, Ka Chun Tsui, Man Lung Fung, Boon Chin Heng, Ersoy Kocabicak, Ali Jahanshahi, Yasin Temel, Arjan Blokland, Luca Aquili, Allan Kalueff, and et al. 2025. "DNA Methylation: A Potential Mediator of the Memory Regulatory Effects of taVNS" Cells 14, no. 17: 1327. https://doi.org/10.3390/cells14171327
APA StyleYee, P. O. P., Tsui, K. C., Fung, M. L., Heng, B. C., Kocabicak, E., Jahanshahi, A., Temel, Y., Blokland, A., Aquili, L., Kalueff, A., Wong, K. H., & Lim, L. W. (2025). DNA Methylation: A Potential Mediator of the Memory Regulatory Effects of taVNS. Cells, 14(17), 1327. https://doi.org/10.3390/cells14171327











