The Putative C2H2 Transcription Factor VadH Governs Development, Osmotic Stress Response, and Sterigmatocystin Production in Aspergillus nidulans
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Culture Conditions
2.2. Nucleic Acid Isolation and Manipulation
2.3. Target Deletion of VadH
2.4. Spore Viability Determination
2.5. Osmotic Stress Assays
2.6. Sterigmatocystin (ST) Determination
2.7. RNA Sequencing (RNA-Seq)
2.8. Statistical Analysis
3. Results
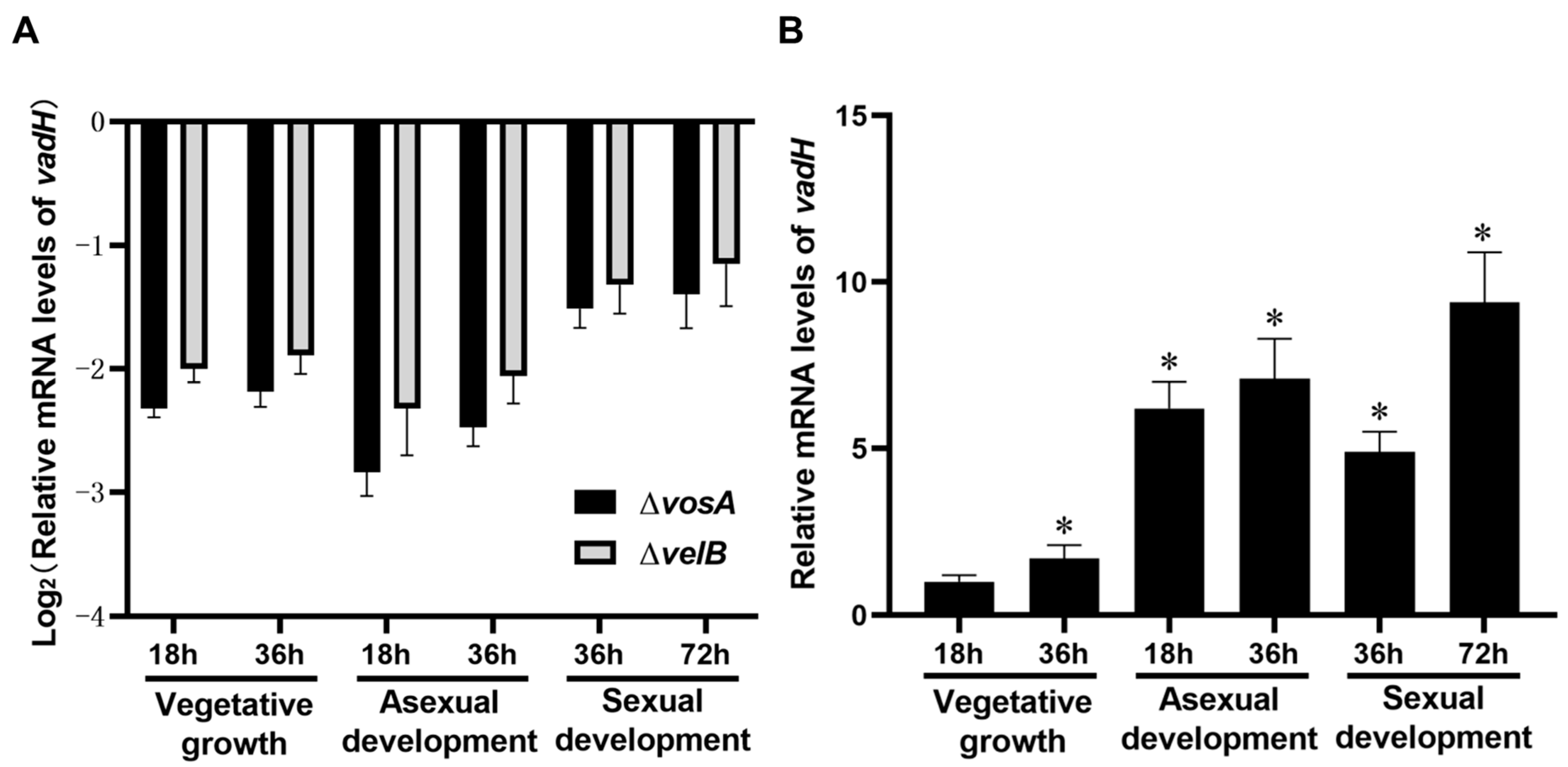
3.1. Characterization of VadH
3.2. VadH Balances Asexual and Sexual Development
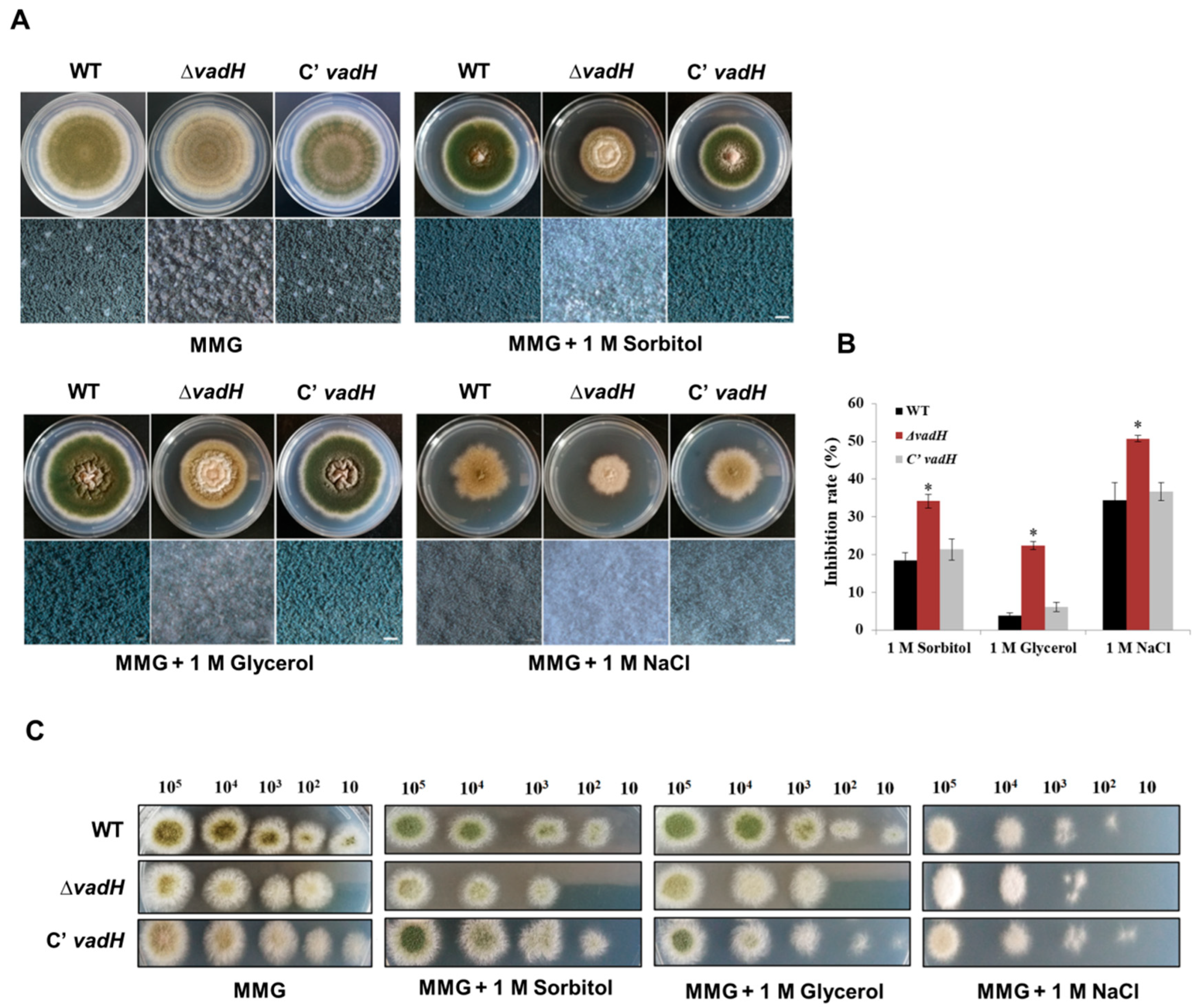
3.3. Disruption of VadH Led to Elevated Sensitivity to Osmotic Stress
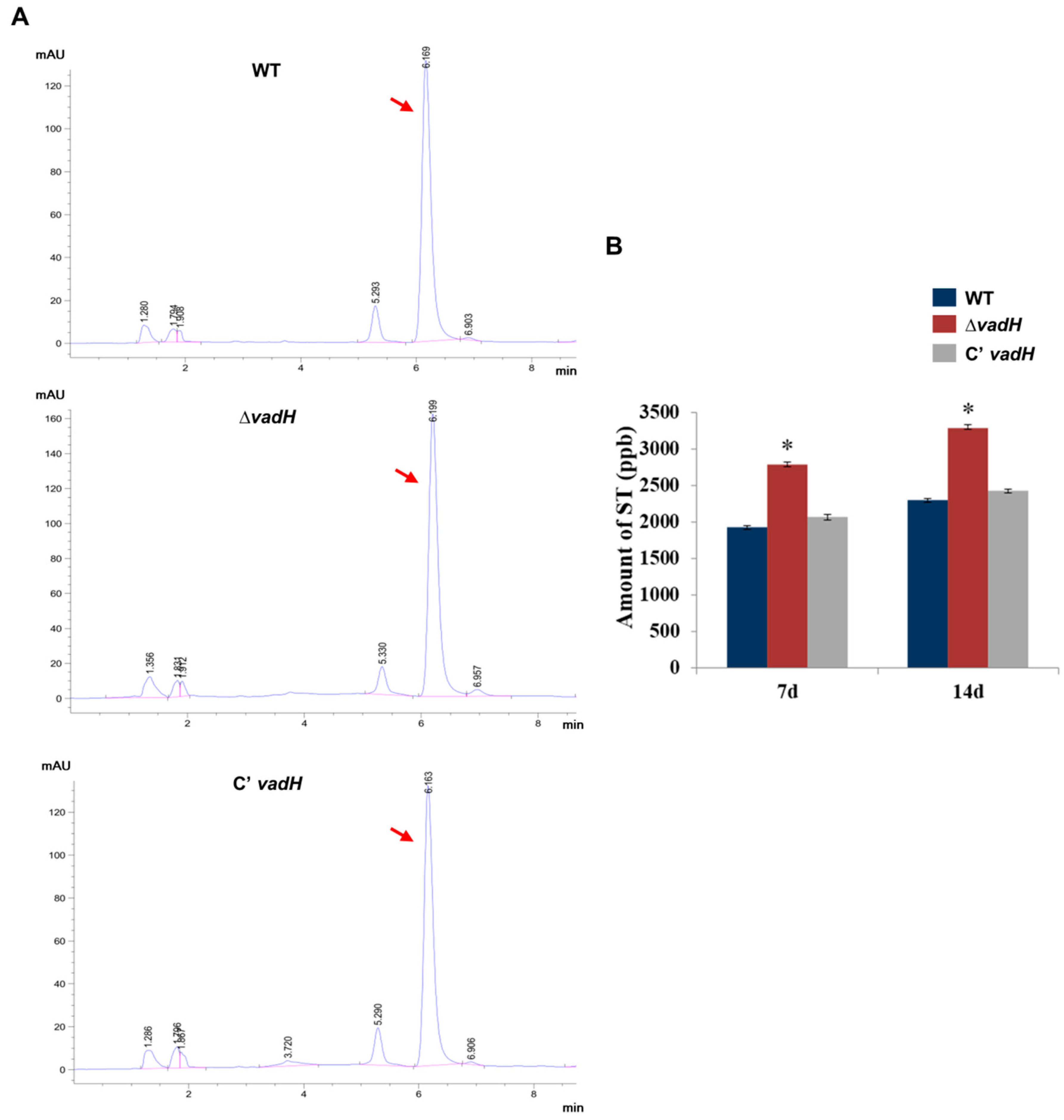
3.4. Deleting VadH Leads to an Increase in ST Production
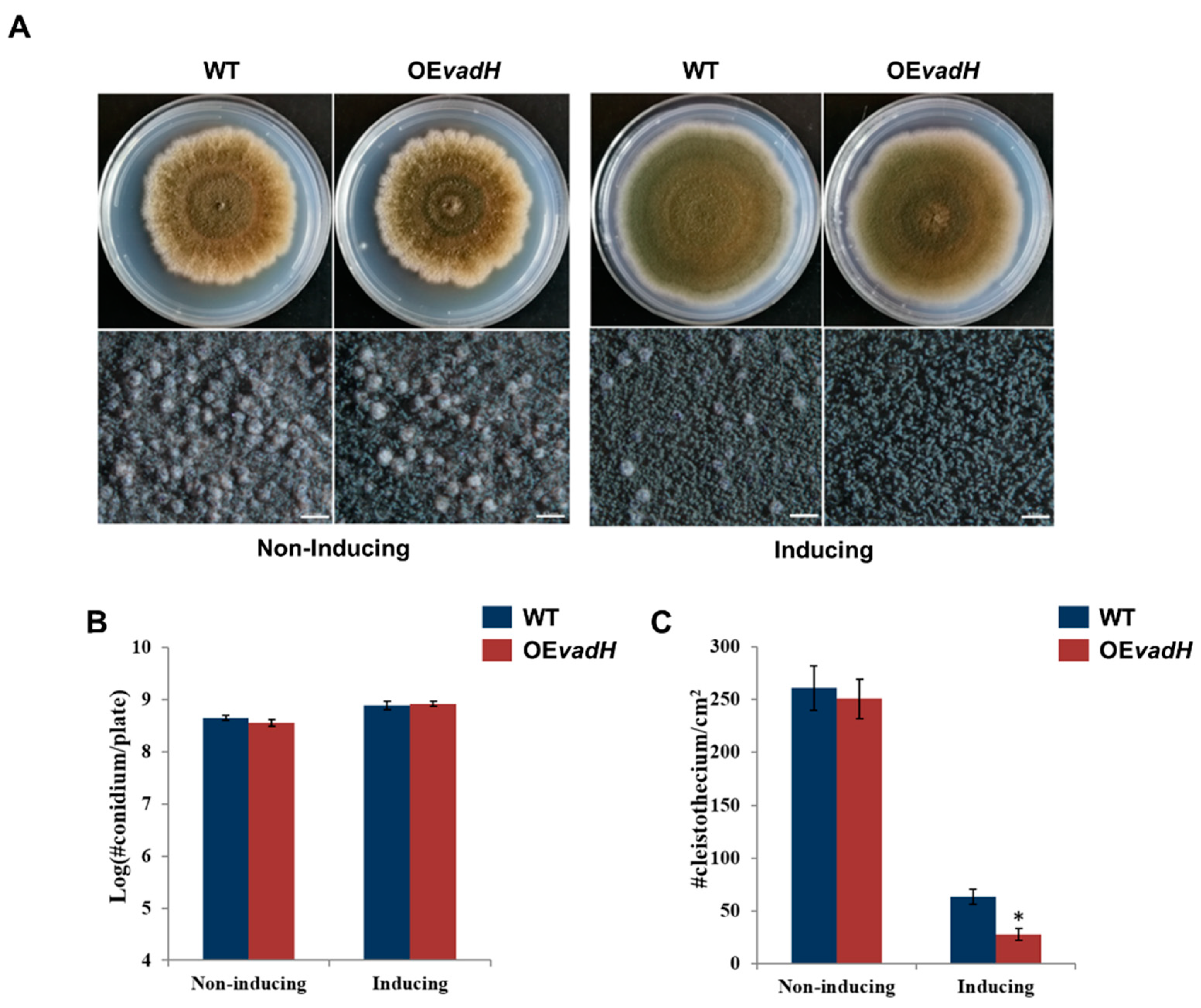
3.5. Overexpression of VadH Suppresses Sexual Development
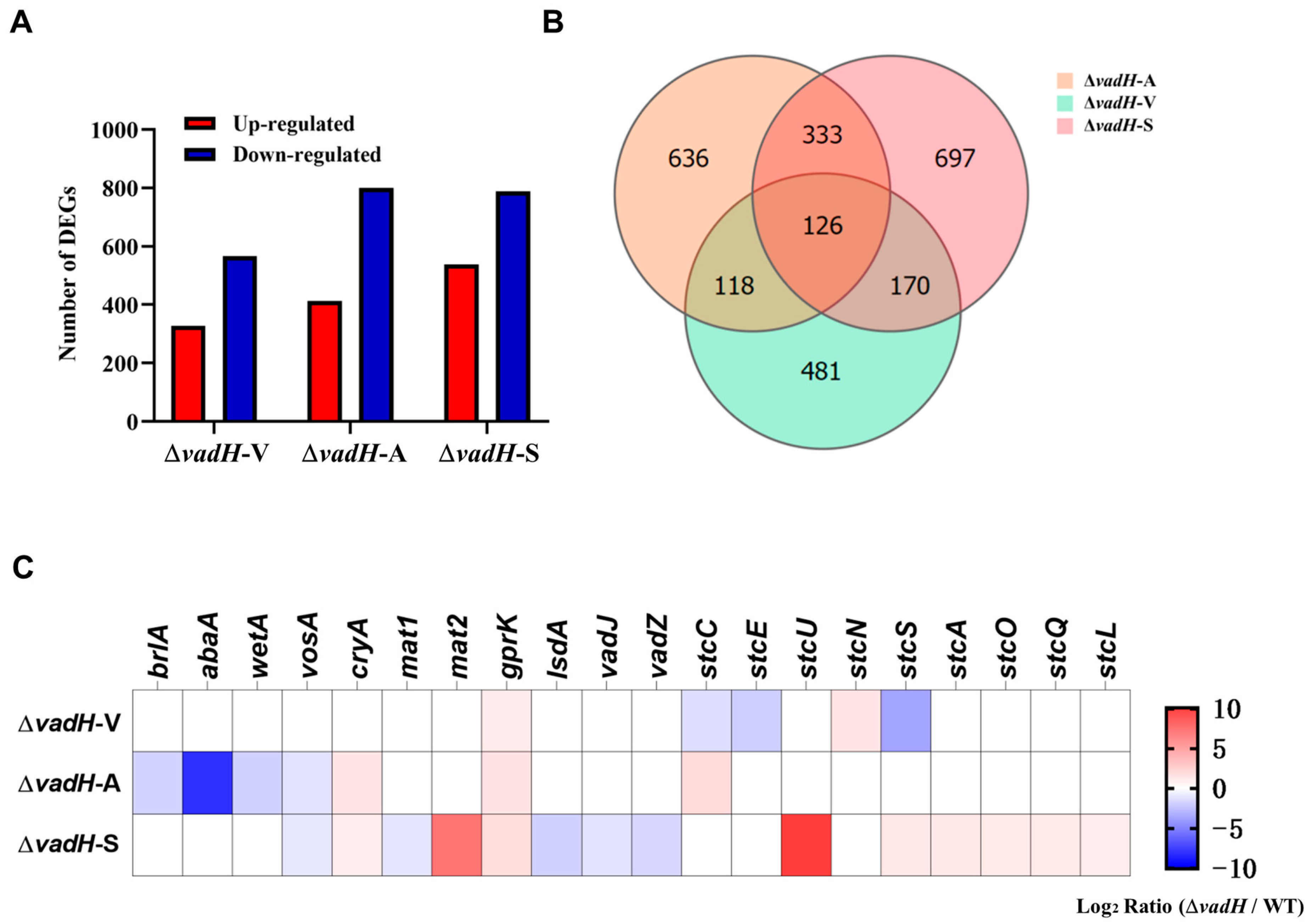
3.6. Transcriptomic Analyses of VadH
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bennett, J.W. An overview of the genus Aspergillus. Aspergillus Mol. Biol. Genom. 2010, 1–17. [Google Scholar]
- Park, H.S.; Yu, J.H. Genetic control of asexual sporulation in filamentous fungi. Curr. Opin. Microbiol. 2012, 15, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Ebbole, D.J. The Conidium. In Cellular and Molecular Biology of Filamentous Fungi; ASM Press: Washington, DC, USA, 2010; pp. 577–590. [Google Scholar]
- Adams, T.H.; Wieser, J.K.; Yu, J.H. Asexual sporulation in Aspergillus nidulans. Microbiol. Mol. Biol. Rev. 1998, 62, 35–54. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H. Regulation of development in Aspergillus nidulans and Aspergillus fumigates. Mycobiology 2010, 38, 229–237. [Google Scholar] [CrossRef]
- Han, S.; Adams, T. Complex control of the developmental regulatory locus brlA in Aspergillus nidulans. Mol. Genet. Genom. 2001, 266, 260–270. [Google Scholar] [CrossRef]
- Andrianopoulos, A.; Timberlake, W.E. The Aspergillus nidulans abaA gene encodes a transcriptional activator that acts as a genetic switch to control development. Mol. Cell. Biol. 1994, 14, 2503–2515. [Google Scholar] [PubMed]
- Marshall, M.A.; Timberlake, W.E. Aspergillus nidulans wetA activates spore-specific gene expression. Mol. Cell. Biol. 1991, 11, 55–62. [Google Scholar]
- Park, H.S.; Nam, T.Y.; Han, K.H.; Kim, S.C.; Yu, J.H. VelC positively controls sexual development in Aspergillus nidulans. PLoS ONE 2014, 9, e89883. [Google Scholar] [CrossRef]
- Park, H.S.; Yu, J.H. Velvet Regulators in Aspergillus spp. Microbiol. Biotechnol. Lett. 2016, 44, 409–419. [Google Scholar] [CrossRef]
- Bayram, O.; Braus, G.H. Coordination of secondary metabolism and development in fungi: The velvet family of regulatory proteins. FEMS Microbiol. Rev. 2012, 36, 1–24. [Google Scholar] [CrossRef]
- Bayram, O.; Krappmann, S.; Ni, M.; Bok, J.W.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef] [PubMed]
- Park, H.S.; Ni, M.; Jeong, K.C.; Kim, Y.H.; Yu, J.H. The role, interaction and regulation of the velvet regulator VelB in Aspergillus nidulans. PLoS ONE 2012, 7, e45935. [Google Scholar] [CrossRef] [PubMed]
- Sarikaya Bayram, O.; Bayram, O.; Valerius, O.; Park, H.S.; Irniger, S.; Gerke, J.; Ni, M.; Han, K.H.; Yu, J.H.; Braus, G.H. LaeA control of velvet family regulatory proteins for light-dependent development and fungal cell-type specificity. PLoS Genet. 2010, 6, e1001226. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, Y.L.; Gerke, J.; Park, H.S.; Bayram, O.; Neumann, P.; Ni, M.; Dickmanns, A.; Kim, S.C.; Yu, J.H.; Braus, G.H.; et al. The velvet family of fungal regulators contains a DNA-binding domain structurally similar to NF-kappaB. PLoS Biol. 2013, 11, e1001750. [Google Scholar] [CrossRef]
- Park, H.S.; Lee, M.K.; Kim, S.C.; Yu, J.H. The role of VosA/VelB-activated developmental gene vadA in Aspergillus nidulans. PLoS ONE 2017, 12, e0177099. [Google Scholar] [CrossRef]
- Park, H.S.; Yu, Y.M.; Lee, M.K.; Maeng, P.J.; Kim, S.C.; Yu, J.H. Velvet-mediated repression of beta-glucan synthesis in Aspergillus nidulans spores. Sci. Rep. 2015, 5, 10199. [Google Scholar] [CrossRef]
- Slattery, M.G.; Liko, D.; Heideman, W. The function and properties of the Azf1 transcriptional regulator change with growth conditions in Saccharomyces cerevisiae. Eukaryot. Cell 2006, 5, 313–320. [Google Scholar] [CrossRef]
- Zhou, Z.Z.; Li, G.H.; Lin, C.H.; He, C.Z. Conidiophore Stalk less1 encodes a putative zinc-finger protein involved in the early stage of conidiation and mycelial infection in Magnaporthe oryzae. Mol. Plant Microbe Interact. 2009, 22, 402–410. [Google Scholar] [CrossRef]
- Li, X.; Ke, Z.; Yu, X.; Liu, Z.; Zhang, C. Transcription factor CgAzf1 regulates melanin production, conidial development and infection in Colletotrichum gloeosporioides. Antonie Van Leeuwenhoek 2019, 112, 1095–1104. [Google Scholar] [CrossRef]
- Shaaban, M.I.; Bok, J.W.; Lauer, C.; Keller, N.P. Suppressor mutagenesis identifies a velvet complex remediator of Aspergillus nidulans secondary metabolism. Eukaryot. Cell 2010, 9, 1816–1824. [Google Scholar] [CrossRef]
- Kwon, N.J.; Shin, K.S.; Yu, J.H. Characterization of the developmental regulator FlbE in Aspergillus fumigatus and Aspergillus nidulans. Fungal Genet. Biol. 2010, 47, 981–993. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Hamari, Z.; Han, K.H.; Seo, J.A.; Reyes-Dominguez, Y.; Scazzocchio, C. Double-joint PCR: A PCR based molecular tool for gene manipulations in filamentous fungi. Fungal Genet. Biol. 2004, 41, 973–981. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Park, H.S.; Yu, J.H. Multi-copy genetic screen in Aspergillus nidulans. Methods Mol. Biol. 2012, 944, 183–190. [Google Scholar] [PubMed]
- Lee, M.K.; Park, H.S.; Han, K.H.; Hong, S.B.; Yu, J.H. High molecular weight genomic DNA mini-prep for filamentous fungi. Fungal Genet. Biol. 2017, 104, 1–5. [Google Scholar] [CrossRef]
- Li, X.; Wu, Y.; Liu, Z.; Zhang, C. The function and transcriptome analysis of a bZIP transcription factor CgAP1 in Colletotrichum gloeosporioides. Microbiol. Res. 2017, 197, 39–48. [Google Scholar] [CrossRef]
- Paoletti, M.; Seymour, F.A.; Alcocer, M.J.C.; Kaur, N.; Calvo, A.M.; Archer, D.B.; Dyer, P.S. Mating type and the genetic basis of self-fertility in the model fungus Aspergillus nidulans. Curr. Biol. 2007, 17, 1384–1389. [Google Scholar] [CrossRef]
- Lee, D.W.; Kim, S.; Kim, S.J.; Han, D.M.; Jahng, K.Y.; Chae, K.S. The lsdA gene is necessary for sexual development inhibition by a salt in Aspergillus nidulans. Curr. Genet. 2001, 39, 237–243. [Google Scholar]
- Zhao, Y.; Lee, M.K.; Lim, J.; Moon, H.; Park, H.S.; Zheng, W.; Yu, J.H. The putative sensor histidine kinase VadJ coordinates development and sterigmatocystin production in Aspergillus nidulans. J. Microbiol. 2021, 59, 746–752. [Google Scholar] [CrossRef]
- Zhao, Y.; Lee, M.K.; Lim, J.; Moon, H.; Park, H.S.; Zheng, W.; Yu, J.H. The velvet-activated putative C6 transcription factor VadZ regulates development and sterigmatocystin production in Aspergillus nidulans. Fungal Biol. 2022, 126, 421–428. [Google Scholar] [CrossRef]
- Kim, M.J.; Jung, W.H.; Son, Y.E.; Yu, J.H.; Lee, M.K.; Park, H.S. The velvet repressed vidA gene plays a key role in governing development in Aspergillus nidulans. J. Microbiol. 2019, 57, 893–899. [Google Scholar] [CrossRef] [PubMed]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Zhao, Y.; Moon, H.; Lim, J.; Park, H.-S.; Liu, Z.; Yu, J.-H. The Putative C2H2 Transcription Factor VadH Governs Development, Osmotic Stress Response, and Sterigmatocystin Production in Aspergillus nidulans. Cells 2022, 11, 3998. https://doi.org/10.3390/cells11243998
Li X, Zhao Y, Moon H, Lim J, Park H-S, Liu Z, Yu J-H. The Putative C2H2 Transcription Factor VadH Governs Development, Osmotic Stress Response, and Sterigmatocystin Production in Aspergillus nidulans. Cells. 2022; 11(24):3998. https://doi.org/10.3390/cells11243998
Chicago/Turabian StyleLi, Xiaoyu, Yanxia Zhao, Heungyun Moon, Jieyin Lim, Hee-Soo Park, Zhiqiang Liu, and Jae-Hyuk Yu. 2022. "The Putative C2H2 Transcription Factor VadH Governs Development, Osmotic Stress Response, and Sterigmatocystin Production in Aspergillus nidulans" Cells 11, no. 24: 3998. https://doi.org/10.3390/cells11243998
APA StyleLi, X., Zhao, Y., Moon, H., Lim, J., Park, H.-S., Liu, Z., & Yu, J.-H. (2022). The Putative C2H2 Transcription Factor VadH Governs Development, Osmotic Stress Response, and Sterigmatocystin Production in Aspergillus nidulans. Cells, 11(24), 3998. https://doi.org/10.3390/cells11243998









