The Memory T Cell “Communication Web” in Context with Gastrointestinal Disorders—How Memory T Cells Affect Their Surroundings and How They Are Influenced by It
Abstract
:1. Introduction
2. Characterisation of Memory T Cells
3. Central and Effector Memory T Cells
4. Tissue Resident Memory T Cells
5. Memory T Cells and Their Role in Gut-Related Diseases
6. The Memory T Cell Networking System
6.1. Interaction with Microbiota—Regulatory Proteins and Metabolites
6.2. Interaction with Other Cell Types and Compartments
7. Therapeutical Approaches
8. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alatab, S.; Sepanlou, S.G.; Ikuta, K.; Vahedi, H.; Bisignano, C.; Safiri, S.; Sadeghi, A.; Nixon, M.R.; Abdoli, A.; Abolhassani, H. The global, regional, and national burden of inflammatory bowel disease in 195 countries and territories, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 2020, 5, 17–30. [Google Scholar] [CrossRef]
- Barberio, B.; Zamani, M.; Black, C.J.; Savarino, E.V.; Ford, A.C. Prevalence of symptoms of anxiety and depression in patients with inflammatory bowel disease: A systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 2021, 6, 359–370. [Google Scholar] [CrossRef]
- Trindade, I.A.; Ferreira, C.; Moura-Ramos, M.; Pinto-Gouveia, J. An 18-month study of the effects of IBD symptomatology and emotion regulation on depressed mood. Int. J. Colorectal Dis. 2017, 32, 651–660. [Google Scholar] [CrossRef]
- Fantini, M.C.; Guadagni, I. From inflammation to colitis-associated colorectal cancer in inflammatory bowel disease: Pathogenesis and impact of current therapies. Dig. Liver Dis. 2021, 53, 558–565. [Google Scholar] [CrossRef]
- Miller, K.D.; Nogueira, L.; Mariotto, A.B.; Rowland, J.H.; Yabroff, K.R.; Alfano, C.M.; Jemal, A.; Kramer, J.L.; Siegel, R.L. Cancer treatment and survivorship statistics, 2019. CA A Cancer J. Clin. 2019, 69, 363–385. [Google Scholar] [CrossRef]
- Schmitt, M.; Greten, F.R. The inflammatory pathogenesis of colorectal cancer. Nat. Rev. Immunol. 2021, 21, 653–667. [Google Scholar] [CrossRef] [PubMed]
- Altajar, S.; Moss, A. Inflammatory Bowel Disease Environmental Risk Factors: Diet and Gut Microbiota. Curr. Gastroenterol. Rep. 2020, 22, 57. [Google Scholar] [CrossRef] [PubMed]
- De Souza, H.S.; Fiocchi, C. Immunopathogenesis of IBD: Current state of the art. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 13–27. [Google Scholar] [CrossRef]
- Schenkel, J.M.; Masopust, D. Tissue-resident memory T cells. Immunity 2014, 41, 886–897. [Google Scholar] [CrossRef]
- Rosenblum, M.D.; Gratz, I.K.; Paw, J.S.; Lee, K.; Marshak-Rothstein, A.; Abbas, A.K. Response to self antigen imprints regulatory memory in tissues. Nature 2011, 480, 538–542. [Google Scholar] [CrossRef] [Green Version]
- Gattinoni, L.; Zhong, X.-S.; Palmer, D.C.; Ji, Y.; Hinrichs, C.S.; Yu, Z.; Wrzesinski, C.; Boni, A.; Cassard, L.; Garvin, L.M.; et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 2009, 15, 808–813. [Google Scholar] [CrossRef]
- Raeber, M.E.; Zurbuchen, Y.; Impellizzieri, D.; Boyman, O. The role of cytokines in T-cell memory in health and disease. Immunol. Rev. 2018, 283, 176–193. [Google Scholar] [CrossRef]
- Gray, J.I.; Westerhof, L.M.; MacLeod, M.K. The roles of resident, central and effector memory CD 4 T-cells in protective immunity following infection or vaccination. Immunology 2018, 154, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Gebhardt, T.; Palendira, U.; Tscharke, D.C.; Bedoui, S. Tissue-resident memory T cells in tissue homeostasis, persistent infection, and cancer surveillance. Immunol. Rev. 2018, 283, 54–76. [Google Scholar] [CrossRef]
- Alabed, M.; Shaik, A.S.; Sharif-Askari, N.S.; Sharif-Askari, F.S.; Hafezi, S.; Mdkhana, B.; Ratemi, E.; Al-Muhsen, S.; Hamid, Q.; Halwani, R. Enhanced Infiltration of Central Memory T Cells to the Lung Tissue during Allergic Lung Inflammation. Int. Arch. Allergy Immunol. 2021, 183, 127–141. [Google Scholar] [CrossRef] [PubMed]
- Bošnjak, B.; Kazemi, S.; Altenburger, L.M.; Mokrović, G.; Epstein, M.M. Th2-TRMs Maintain Life-Long Allergic Memory in Experimental Asthma in Mice. Front. Immunol. 2019, 10, 840. [Google Scholar] [CrossRef]
- Zundler, S.; Becker, E.; Spocinska, M.; Slawik, M.; Parga-Vidal, L.; Stark, R.; Wiendl, M.; Atreya, R.; Rath, T.; Leppkes, M. Hobit-and Blimp-1-driven CD4+ tissue-resident memory T cells control chronic intestinal inflammation. Nat. Immunol. 2019, 20, 288–300. [Google Scholar] [CrossRef]
- Bishu, S.; El Zaatari, M.; Hayashi, A.; Hou, G.; Bowers, N.; Kinnucan, J.; Manoogian, B.; Muza-Moons, M.; Zhang, M.; Grasberger, H. CD4+ tissue-resident memory T cells expand and are a major source of mucosal tumour necrosis factor α in active Crohn’s disease. J. Crohn’s Colitis 2019, 13, 905–915. [Google Scholar] [CrossRef] [PubMed]
- Lanzavecchia, A.; Sallusto, F. Understanding the generation and function of memory T cell subsets. Curr. Opin. Immunol. 2005, 17, 326–332. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.N.; Mackay, L.K. Tissue-resident memory T cells: Local specialists in immune defence. Nat. Rev. Immunol. 2016, 16, 79–89. [Google Scholar] [CrossRef]
- Choi, H.; Song, H.; Jung, Y.W. The Roles of CCR7 for the Homing of Memory CD8+ T Cells into Their Survival Niches. Immune Netw 2020, 20, e20. [Google Scholar] [CrossRef]
- Sallusto, F.; Lenig, D.; Förster, R.; Lipp, M.; Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 1999, 401, 708–712. [Google Scholar] [CrossRef]
- Hipkin, W. CCR7 Chemokine Receptor. In xPharm: The Comprehensive Pharmacology Reference; Enna, S.J., Bylund, D.B., Eds.; Elsevier: New York, NY, USA, 2007; pp. 1–10. [Google Scholar] [CrossRef]
- Tian, Y.; Babor, M.; Lane, J.; Schulten, V.; Patil, V.S.; Seumois, G.; Rosales, S.L.; Fu, Z.; Picarda, G.; Burel, J.; et al. Unique phenotypes and clonal expansions of human CD4 effector memory T cells re-expressing CD45RA. Nat. Commun. 2017, 8, 1473. [Google Scholar] [CrossRef] [PubMed]
- Chao, C.C.; Jensen, R.; Dailey, M.O. Mechanisms of L-selectin regulation by activated T cells. J. Immunol. 1997, 159, 1686–1694. [Google Scholar] [PubMed]
- Sallusto, F.; Geginat, J.; Lanzavecchia, A. Central memory and effector memory T cell subsets: Function, generation, and maintenance. Annu. Rev. Immunol. 2004, 22, 745–763. [Google Scholar] [CrossRef]
- Gerlach, C.; Moseman, E.A.; Loughhead, S.M.; Alvarez, D.; Zwijnenburg, A.J.; Waanders, L.; Garg, R.; de la Torre, J.C.; von Andrian, U.H. The Chemokine Receptor CX3CR1 Defines Three Antigen-Experienced CD8 T Cell Subsets with Distinct Roles in Immune Surveillance and Homeostasis. Immunity 2016, 45, 1270–1284. [Google Scholar] [CrossRef] [PubMed]
- Kohlmeier, J.E.; Reiley, W.W.; Perona-Wright, G.; Freeman, M.L.; Yager, E.J.; Connor, L.M.; Brincks, E.L.; Cookenham, T.; Roberts, A.D.; Burkum, C.E.; et al. Inflammatory chemokine receptors regulate CD8(+) T cell contraction and memory generation following infection. J. Exp. Med. 2011, 208, 1621–1634. [Google Scholar] [CrossRef] [PubMed]
- Cui, W.; Kaech, S.M. Generation of effector CD8+ T cells and their conversion to memory T cells. Immunol. Rev. 2010, 236, 151–166. [Google Scholar] [CrossRef]
- Joshi, N.S.; Cui, W.; Chandele, A.; Lee, H.K.; Urso, D.R.; Hagman, J.; Gapin, L.; Kaech, S.M. Inflammation directs memory precursor and short-lived effector CD8(+) T cell fates via the graded expression of T-bet transcription factor. Immunity 2007, 27, 281–295. [Google Scholar] [CrossRef] [Green Version]
- Liu, Q.; Sun, Z.; Chen, L. Memory T cells: Strategies for optimizing tumor immunotherapy. Protein Cell 2020, 11, 549–564. [Google Scholar] [CrossRef]
- Akondy, R.S.; Fitch, M.; Edupuganti, S.; Yang, S.; Kissick, H.T.; Li, K.W.; Youngblood, B.A.; Abdelsamed, H.A.; McGuire, D.J.; Cohen, K.W.; et al. Origin and differentiation of human memory CD8 T cells after vaccination. Nature 2017, 552, 362–367. [Google Scholar] [CrossRef] [PubMed]
- Youngblood, B.; Hale, J.S.; Kissick, H.T.; Ahn, E.; Xu, X.; Wieland, A.; Araki, K.; West, E.E.; Ghoneim, H.E.; Fan, Y.; et al. Effector CD8 T cells dedifferentiate into long-lived memory cells. Nature 2017, 552, 404–409. [Google Scholar] [CrossRef]
- Arenas-Ramirez, N.; Woytschak, J.; Boyman, O. Interleukin-2: Biology, Design and Application. Trends Immunol. 2015, 36, 763–777. [Google Scholar] [CrossRef]
- Damoiseaux, J. The IL-2—IL-2 receptor pathway in health and disease: The role of the soluble IL-2 receptor. Clin. Immunol. 2020, 218, 108515. [Google Scholar] [CrossRef]
- Wang, X.; Rickert, M.; Garcia, K.C. Structure of the Quaternary Complex of Interleukin-2 with Its α, ß, and γc Receptors. Science 2005, 310, 1159–1163. [Google Scholar] [CrossRef]
- Zhang, B.; Zhang, Y.; Xiong, L.; Li, Y.; Zhang, Y.; Zhao, J.; Jiang, H.; Li, C.; Liu, Y.; Liu, X.; et al. CD127 imprints functional heterogeneity to diversify monocyte responses in inflammatory diseases. J. Exp. Med. 2022, 219, e20211191. [Google Scholar] [CrossRef] [PubMed]
- Knörck, A.; Schwär, G.; Alansary, D.; Thurner, L.; Hoth, M.; Schwarz, E.C. Cytotoxic efficiency of human CD8+ T cell memory subtypes. Front. Immunol. 2021, 13, 838484. [Google Scholar] [CrossRef] [PubMed]
- Westerhof, L.M.; McGuire, K.; MacLellan, L.; Flynn, A.; Gray, J.I.; Thomas, M.; Goodyear, C.S.; MacLeod, M.K. Multifunctional cytokine production reveals functional superiority of memory CD4 T cells. Eur. J. Immunol. 2019, 49, 2019–2029. [Google Scholar] [CrossRef] [PubMed]
- Masopust, D.; Jiang, J.; Shen, H.; Lefrançois, L. Direct Analysis of the Dynamics of the Intestinal Mucosa CD8 T Cell Response to Systemic Virus Infection. J. Immunol. 2001, 166, 2348–2356. [Google Scholar] [CrossRef] [Green Version]
- Masopust, D.; Vezys, V.; Marzo, A.L.; Lefrançois, L. Preferential Localization of Effector Memory Cells in Nonlymphoid Tissue. Science 2001, 291, 2413–2417. [Google Scholar] [CrossRef]
- Gebhardt, T.; Wakim, L.M.; Eidsmo, L.; Reading, P.C.; Heath, W.R.; Carbone, F.R. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 2009, 10, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Kumar, B.V.; Ma, W.; Miron, M.; Granot, T.; Guyer, R.S.; Carpenter, D.J.; Senda, T.; Sun, X.; Ho, S.-H.; Lerner, H.; et al. Human Tissue-Resident Memory T Cells Are Defined by Core Transcriptional and Functional Signatures in Lymphoid and Mucosal Sites. Cell Rep. 2017, 20, 2921–2934. [Google Scholar] [CrossRef] [PubMed]
- Mackay, L.K.; Rahimpour, A.; Ma, J.Z.; Collins, N.; Stock, A.T.; Hafon, M.-L.; Vega-Ramos, J.; Lauzurica, P.; Mueller, S.N.; Stefanovic, T.; et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 2013, 14, 1294–1301. [Google Scholar] [CrossRef]
- Simms, P.E.; Ellis, T.M. Utility of flow cytometric detection of CD69 expression as a rapid method for determining poly- and oligoclonal lymphocyte activation. Clin. Diagn. Lab. Immunol. 1996, 3, 301–304. [Google Scholar] [CrossRef] [PubMed]
- Sathaliyawala, T.; Kubota, M.; Yudanin, N.; Turner, D.; Camp, P.; Thome Joseph, J.C.; Bickham Kara, L.; Lerner, H.; Goldstein, M.; Sykes, M.; et al. Distribution and Compartmentalization of Human Circulating and Tissue-Resident Memory T Cell Subsets. Immunity 2013, 38, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Steinert Elizabeth, M.; Schenke Jason, M.; Fraser Kathryn, A.; Beura Lalit, K.; Manlove Luke, S.; Igyártó Botond, Z.; Southern Peter, J.; Masopust, D. Quantifying Memory CD8 T Cells Reveals Regionalization of Immunosurveillance. Cell 2015, 161, 737–749. [Google Scholar] [CrossRef]
- Behr, F.M.; Chuwonpad, A.; Stark, R.; van Gisbergen, K.P.J.M. Armed and Ready: Transcriptional Regulation of Tissue-Resident Memory CD8 T Cells. Front. Immunol. 2018, 9, 1770. [Google Scholar] [CrossRef]
- Topham, D.J.; Reilly, E.C. Tissue-Resident Memory CD8+ T Cells: From Phenotype to Function. Front. Immunol. 2018, 9, 515. [Google Scholar] [CrossRef]
- Bankovich, A.J.; Shiow, L.R.; Cyster, J.G. CD69 suppresses sphingosine 1-phosophate receptor-1 (S1P1) function through interaction with membrane helix 4. J. Biol. Chem. 2010, 285, 22328–22337. [Google Scholar] [CrossRef] [Green Version]
- Danese, S.; Furfaro, F.; Vetrano, S. Targeting S1P in Inflammatory Bowel Disease: New Avenues for Modulating Intestinal Leukocyte Migration. J. Crohns Colitis 2018, 12, S678–S686. [Google Scholar] [CrossRef]
- Baeyens, A.; Fang, V.; Chen, C.; Schwab, S.R. Exit Strategies: S1P Signaling and T Cell Migration. Trends Immunol. 2015, 36, 778–787. [Google Scholar] [CrossRef] [PubMed]
- Cepek, K.L.; Shaw, S.K.; Parker, C.M.; Russell, G.J.; Morrow, J.S.; Rimm, D.L.; Brenner, M.B. Adhesion between epithelial cells and T lymphocytes mediated by E-cadherin and the alpha E beta 7 integrin. Nature 1994, 372, 190–193. [Google Scholar] [CrossRef]
- Farber, D.L.; Yudanin, N.A.; Restifo, N.P. Human memory T cells: Generation, compartmentalization and homeostasis. Nat. Rev. Immunol. 2014, 14, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Behr, F.M.; Kragten, N.A.M.; Wesselink, T.H.; Nota, B.; van Lier, R.A.W.; Amsen, D.; Stark, R.; Hombrink, P.; van Gisbergen, K.P.J.M. Blimp-1 Rather Than Hobit Drives the Formation of Tissue-Resident Memory CD8+ T Cells in the Lungs. Front. Immunol. 2019, 10, 400. [Google Scholar] [CrossRef] [PubMed]
- McMaster, S.R.; Wein, A.N.; Dunbar, P.R.; Hayward, S.L.; Cartwright, E.K.; Denning, T.L.; Kohlmeier, J.E. Pulmonary antigen encounter regulates the establishment of tissue-resident CD8 memory T cells in the lung airways and parenchyma. Mucosal Immunol. 2018, 11, 1071–1078. [Google Scholar] [CrossRef]
- Bromley, S.K.; Akbaba, H.; Mani, V.; Mora-Buch, R.; Chasse, A.Y.; Sama, A.; Luster, A.D. CD49a Regulates Cutaneous Resident Memory CD8+ T Cell Persistence and Response. Cell Rep. 2020, 32, 108085. [Google Scholar] [CrossRef]
- McNamara, H.; Cai, Y.; Wagle, M.; Sontani, Y.; Roots, C.; Miosge, L.; O’Connor, J.; Sutton, H.; Ganusov, V.; Heath, W.R. Up-regulation of LFA-1 allows liver-resident memory T cells to patrol and remain in the hepatic sinusoids. Sci. Immunol. 2017, 2, eaaj1996. [Google Scholar] [CrossRef]
- Sheridan Brian, S.; Pham, Q.-M.; Lee, Y.-T.; Cauley Linda, S.; Puddington, L.; Lefrançois, L. Oral Infection Drives a Distinct Population of Intestinal Resident Memory CD8+ T Cells with Enhanced Protective Function. Immunity 2014, 40, 747–757. [Google Scholar] [CrossRef]
- Herndler-Brandstetter, D.; Ishigame, H.; Shinnakasu, R.; Plajer, V.; Stecher, C.; Zhao, J.; Lietzenmayer, M.; Kroehling, L.; Takumi, A.; Kometani, K.; et al. KLRG1+ Effector CD8+ T Cells Lose KLRG1, Differentiate into All Memory T Cell Lineages, and Convey Enhanced Protective Immunity. Immunity 2018, 48, 716–729.e718. [Google Scholar] [CrossRef] [Green Version]
- El-Asady, R.; Yuan, R.; Liu, K.; Wang, D.; Gress, R.E.; Lucas, P.J.; Drachenberg, C.B.; Hadley, G.A. TGF-β–dependent CD103 expression by CD8+ T cells promotes selective destruction of the host intestinal epithelium during graft-versus-host disease. J. Exp. Med. 2005, 201, 1647–1657. [Google Scholar] [CrossRef]
- Mackay Laura, K.; Wynne-Jones, E.; Freestone, D.; Pellicci Daniel, G.; Mielke Lisa, A.; Newman Dane, M.; Braun, A.; Masson, F.; Kallies, A.; Belz Gabrielle, T.; et al. T-box Transcription Factors Combine with the Cytokines TGF-β and IL-15 to Control Tissue-Resident Memory T Cell Fate. Immunity 2015, 43, 1101–1111. [Google Scholar] [CrossRef]
- Skon, C.N.; Lee, J.-Y.; Anderson, K.G.; Masopust, D.; Hogquist, K.A.; Jameson, S.C. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 2013, 14, 1285–1293. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-Y.; Skon, C.N.; Lee, Y.J.; Oh, S.; Taylor, J.J.; Malhotra, D.; Jenkins, M.K.; Rosenfeld, M.G.; Hogquist, K.A.; Jameson, S.C. The Transcription Factor KLF2 Restrains CD4+ T Follicular Helper Cell Differentiation. Immunity 2015, 42, 252–264. [Google Scholar] [CrossRef] [PubMed]
- Milner, J.J.; Toma, C.; Yu, B.; Zhang, K.; Omilusik, K.; Phan, A.T.; Wang, D.; Getzler, A.J.; Nguyen, T.; Crotty, S.; et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature 2017, 552, 253–257. [Google Scholar] [CrossRef] [PubMed]
- Bartolomé-Casado, R.; Landsverk, O.J.; Chauhan, S.K.; Richter, L.; Phung, D.; Greiff, V.; Risnes, L.F.; Yao, Y.; Neumann, R.S.; Yaqub, S. Resident memory CD8 T cells persist for years in human small intestine. J. Exp. Med. 2019, 216, 2412–2426. [Google Scholar] [CrossRef]
- Bartolomé-Casado, R.; Landsverk, O.J.B.; Chauhan, S.K.; Sætre, F.; Hagen, K.T.; Yaqub, S.; Øyen, O.; Horneland, R.; Aandahl, E.M.; Aabakken, L.; et al. CD4+ T cells persist for years in the human small intestine and display a TH1 cytokine profile. Mucosal Immunol. 2021, 14, 402–410. [Google Scholar] [CrossRef]
- Bergsbaken, T.; Bevan, M.J. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8⁺ T cells responding to infection. Nat. Immunol. 2015, 16, 406–414. [Google Scholar] [CrossRef]
- Bergsbaken, T.; Bevan, M.J.; Fink, P.J. Local inflammatory cues regulate differentiation and persistence of CD8+ tissue-resident memory T cells. Cell Rep. 2017, 19, 114–124. [Google Scholar] [CrossRef]
- Zhang, N.; Bevan, M.J. Transforming growth factor-β signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 2013, 39, 687–696. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Senda, T.; Dogra, P.; Granot, T.; Furuhashi, K.; Snyder, M.E.; Carpenter, D.J.; Szabo, P.A.; Thapa, P.; Miron, M.; Farber, D.L. Microanatomical dissection of human intestinal T-cell immunity reveals site-specific changes in gut-associated lymphoid tissues over life. Mucosal Immunol. 2019, 12, 378–389. [Google Scholar] [CrossRef]
- Paap, E.-M.; Müller, T.M.; Sommer, K.; Neurath, M.F.; Zundler, S. Total Recall: Intestinal TRM Cells in Health and Disease. Front. Immunol. 2021, 11, 623072. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, R.; Beura, L.K.; Quarnstrom, C.F.; Ghoneim, H.E.; Fan, Y.; Zebley, C.C.; Scott, M.C.; Fares-Frederickson, N.J.; Wijeyesinghe, S.; Thompson, E.A.; et al. Developmental plasticity allows outside-in immune responses by resident memory T cells. Nat. Immunol. 2020, 21, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Künzli, M.; King, C.G. Resident Memory T Cells Escape ‘Home Quarantine’. Trends Immunol. 2020, 41, 454–456. [Google Scholar] [CrossRef] [PubMed]
- Lamb, C.A.; Mansfield, J.C.; Tew, G.W.; Gibbons, D.; Long, A.K.; Irving, P.; Diehl, L.; Eastham-Anderson, J.; Price, M.B.; O’Boyle, G.; et al. αEβ7 Integrin Identifies Subsets of Pro-Inflammatory Colonic CD4+ T Lymphocytes in Ulcerative Colitis. J. Crohn’s Colitis 2016, 11, 610–620. [Google Scholar] [CrossRef] [PubMed]
- Sasson, S.C.; Slevin, S.M.; Cheung, V.T.; Nassiri, I.; Olsson-Brown, A.; Fryer, E.; Ferreira, R.C.; Trzupek, D.; Gupta, T.; Al-Hillawi, L. Interferon-Gamma–Producing CD8+ Tissue Resident Memory T Cells Are a Targetable Hallmark of Immune Checkpoint Inhibitor–Colitis. Gastroenterology 2021, 161, 1229–1244.e1229. [Google Scholar] [CrossRef]
- Roosenboom, B.; Wahab, P.J.; Smids, C.; Groenen, M.J.M.; van Koolwijk, E.; van Lochem, E.G.; Horjus Talabur Horje, C.S. Intestinal CD103+CD4+ and CD103+CD8+ T-Cell Subsets in the Gut of Inflammatory Bowel Disease Patients at Diagnosis and During Follow-up. Inflamm. Bowel Dis. 2019, 25, 1497–1509. [Google Scholar] [CrossRef]
- Noble, A.; Durant, L.; Hoyles, L.; Mccartney, A.L.; Man, R.; Segal, J.; Costello, S.P.; Hendy, P.; Reddi, D.; Bouri, S.; et al. Deficient Resident Memory T Cell and CD8 T Cell Response to Commensals in Inflammatory Bowel Disease. J. Crohn’s Colitis 2019, 14, 525–537. [Google Scholar] [CrossRef]
- Noble, A.; Pring, E.T.; Durant, L.; Man, R.; Dilke, S.M.; Hoyles, L.; James, S.A.; Carding, S.R.; Jenkins, J.T.; Knight, S.C. Altered immunity to microbiota, B cell activation and depleted γδ/resident memory T cells in colorectal cancer. Cancer Immunol. Immunother. 2022, 1–11, Online ahead of printing. [Google Scholar] [CrossRef]
- Lauret Marie Joseph, E.; Kirilovsky, A.; Lecoester, B.; El Sissy, C.; Boullerot, L.; Rangan, L.; Marguier, A.; Tochet, F.; Dosset, M.; Boustani, J.; et al. Chemoradiation triggers antitumor Th1 and tissue resident memory-polarized immune responses to improve immune checkpoint inhibitors therapy. J. Immunother. Cancer 2021, 9, e002256. [Google Scholar] [CrossRef]
- Stringhini, M.; Probst, P.; Neri, D. Immunotherapy of CT26 murine tumors is characterized by an oligoclonal response of tissue-resident memory T cells against the AH1 rejection antigen. Eur. J. Immunol. 2020, 50, 1591–1597. [Google Scholar] [CrossRef]
- Overacre-Delgoffe, A.E.; Hand, T.W. Regulation of tissue-resident memory T cells by the Microbiota. Mucosal Immunol. 2022, 15, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, A.N.; West, N.R.; Stubbington, M.J.T.; Wendt, E.; Suijker, K.I.M.; Datsi, A.; This, S.; Danne, C.; Campion, S.; Duncan, S.H.; et al. Circulating and Tissue-Resident CD4+ T Cells with Reactivity to Intestinal Microbiota Are Abundant in Healthy Individuals and Function Is Altered During Inflammation. Gastroenterology 2017, 153, 1320–1337.e1316. [Google Scholar] [CrossRef]
- Hohenberger, M.; Cardwell, L.A.; Oussedik, E.; Feldman, S.R. Interleukin-17 inhibition: Role in psoriasis and inflammatory bowel disease. J. Dermatol. Treat. 2018, 29, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Fan, M.Y.; Low, J.S.; Tanimine, N.; Finn, K.K.; Priyadharshini, B.; Germana, S.K.; Kaech, S.M.; Turka, L.A. Differential Roles of IL-2 Signaling in Developing versus Mature Tregs. Cell Rep. 2018, 25, 1204–1213.e1204. [Google Scholar] [CrossRef] [PubMed]
- Laurence, A.; Tato, C.M.; Davidson, T.S.; Kanno, Y.; Chen, Z.; Yao, Z.; Blank, R.B.; Meylan, F.; Siegel, R.; Hennighausen, L. Interleukin-2 signaling via STAT5 constrains T helper 17 cell generation. Immunity 2007, 26, 371–381. [Google Scholar] [CrossRef]
- Bachem, A.; Makhlouf, C.; Binger, K.J.; de Souza, D.P.; Tull, D.; Hochheiser, K.; Whitney, P.G.; Fernandez-Ruiz, D.; Dähling, S.; Kastenmüller, W.; et al. Microbiota-Derived Short-Chain Fatty Acids Promote the Memory Potential of Antigen-Activated CD8+ T Cells. Immunity 2019, 51, 285–297.e285. [Google Scholar] [CrossRef]
- Balmer, M.L.; Ma, E.H.; Bantug, G.R.; Grählert, J.; Pfister, S.; Glatter, T.; Jauch, A.; Dimeloe, S.; Slack, E.; Dehio, P.; et al. Memory CD8(+) T Cells Require Increased Concentrations of Acetate Induced by Stress for Optimal Function. Immunity 2016, 44, 1312–1324. [Google Scholar] [CrossRef]
- Idzko, M.; Hammad, H.; van Nimwegen, M.; Kool, M.; Willart, M.A.; Muskens, F.; Hoogsteden, H.C.; Luttmann, W.; Ferrari, D.; Di Virgilio, F.; et al. Extracellular ATP triggers and maintains asthmatic airway inflammation by activating dendritic cells. Nat. Med. 2007, 13, 913–919. [Google Scholar] [CrossRef]
- Antonioli, L.; Pacher, P.; Vizi, E.S.; Haskó, G. CD39 and CD73 in immunity and inflammation. Trends Mol. Med. 2013, 19, 355–367. [Google Scholar] [CrossRef] [Green Version]
- Stagg, A.J. Intestinal dendritic cells in health and gut inflammation. Front. Immunol. 2018, 9, 2883. [Google Scholar] [CrossRef]
- Bottois, H.; Ngollo, M.; Hammoudi, N.; Courau, T.; Bonnereau, J.; Chardiny, V.; Grand, C.; Gergaud, B.; Allez, M.; Le Bourhis, L. KLRG1 and CD103 Expressions Define Distinct Intestinal Tissue-Resident Memory CD8 T Cell Subsets Modulated in Crohn’s Disease. Front. Immunol. 2020, 11, 896. [Google Scholar] [CrossRef] [PubMed]
- Brand, S.; Beigel, F.; Olszak, T.; Zitzmann, K.; Eichhorst, S.T.; Otte, J.-M.; Diepolder, H.; Marquardt, A.; Jagla, W.; Popp, A.; et al. IL-22 is increased in active Crohn’s disease and promotes proinflammatory gene expression and intestinal epithelial cell migration. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 290, G827–G838. [Google Scholar] [CrossRef] [PubMed]
- Kaser, A.; Ludwiczek, O.; Holzmann, S.; Moschen, A.R.; Weiss, G.; Enrich, B.; Graziadei, I.; Dunzendorfer, S.; Wiedermann, C.J.; Mürzl, E.; et al. Increased Expression of CCL20 in Human Inflammatory Bowel Disease. J. Clin. Immunol. 2004, 24, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Corridoni, D.; Antanaviciute, A.; Gupta, T.; Fawkner-Corbett, D.; Aulicino, A.; Jagielowicz, M.; Parikh, K.; Repapi, E.; Taylor, S.; Ishikawa, D.; et al. Single-cell atlas of colonic CD8+ T cells in ulcerative colitis. Nat. Med. 2020, 26, 1480–1490. [Google Scholar] [CrossRef]
- Ferreira, C.; Barros, L.; Baptista, M.; Blankenhaus, B.; Barros, A.; Figueiredo-Campos, P.; Konjar, Š.; Lainé, A.; Kamenjarin, N.; Stojanovic, A.; et al. Type 1 Treg cells promote the generation of CD8+ tissue-resident memory T cells. Nat. Immunol. 2020, 21, 766–776. [Google Scholar] [CrossRef]
- Kotlarz, D.; Marquardt, B.; Barøy, T.; Lee, W.S.; Konnikova, L.; Hollizeck, S.; Magg, T.; Lehle, A.S.; Walz, C.; Borggraefe, I.; et al. Human TGF-β1 deficiency causes severe inflammatory bowel disease and encephalopathy. Nat. Genet. 2018, 50, 344–348. [Google Scholar] [CrossRef]
- Velikova, T.V.; Miteva, L.; Stanilov, N.; Spassova, Z.; Stanilova, S.A. Interleukin-6 compared to the other Th17/Treg related cytokines in inflammatory bowel disease and colorectal cancer. World J. Gastroenterol. 2020, 26, 1912–1925. [Google Scholar] [CrossRef] [PubMed]
- McDaniel, M.M.; Chawla, A.S.; Jain, A.; Meibers, H.E.; Saha, I.; Gao, Y.; Jain, V.; Roskin, K.; Way, S.S.; Pasare, C. Effector memory CD4+ T cells induce damaging innate inflammation and autoimmune pathology by engaging CD40 and TNFR on myeloid cells. Sci. Immunol. 2022, 7, eabk0182. [Google Scholar] [PubMed]
- Thompson, E.A.; Darrah, P.A.; Foulds, K.E.; Hoffer, E.; Caffrey-Carr, A.; Norenstedt, S.; Perbeck, L.; Seder, R.A.; Kedl, R.M.; Loré, K. Monocytes Acquire the Ability to Prime Tissue-Resident T Cells via IL-10-Mediated TGF-β Release. Cell Rep. 2019, 28, 1127–1135.e1124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desai, P.; Tahiliani, V.; Stanfield, J.; Abboud, G.; Salek-Ardakani, S. Inflammatory monocytes contribute to the persistence of CXCR3hiCX3CR1lo circulating and lung-resident memory CD8+ T cells following respiratory virus infection. Immunol. Cell Biol. 2018, 96, 370–378. [Google Scholar] [CrossRef]
- Vermeire, S.; Loftus, E.V., Jr.; Colombel, J.-F.; Feagan, B.G.; Sandborn, W.J.; Sands, B.E.; Danese, S.; D’Haens, G.R.; Kaser, A.; Panaccione, R.; et al. Long-term Efficacy of Vedolizumab for Crohn’s Disease. J. Crohn’s Colitis 2016, 11, 412–424. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.C.; Hilmi, I.N.; Blake, A.; Bhayat, F.; Adsul, S.; Khan, Q.R.; Wu, D.-C. Low Frequency of Opportunistic Infections in Patients Receiving Vedolizumab in Clinical Trials and Post-Marketing Setting. Inflamm. Bowel Dis. 2018, 24, 2431–2441. [Google Scholar] [CrossRef] [PubMed]
- Noman, M.; Ferrante, M.; Bisschops, R.; De Hertogh, G.; Van den Broeck, K.; Rans, K.; Rutgeerts, P.; Vermeire, S.; Van Assche, G. Vedolizumab Induces Long-term Mucosal Healing in Patients with Crohn’s Disease and Ulcerative Colitis. J. Crohn’s Colitis 2017, 11, 1085–1089. [Google Scholar] [CrossRef] [PubMed]
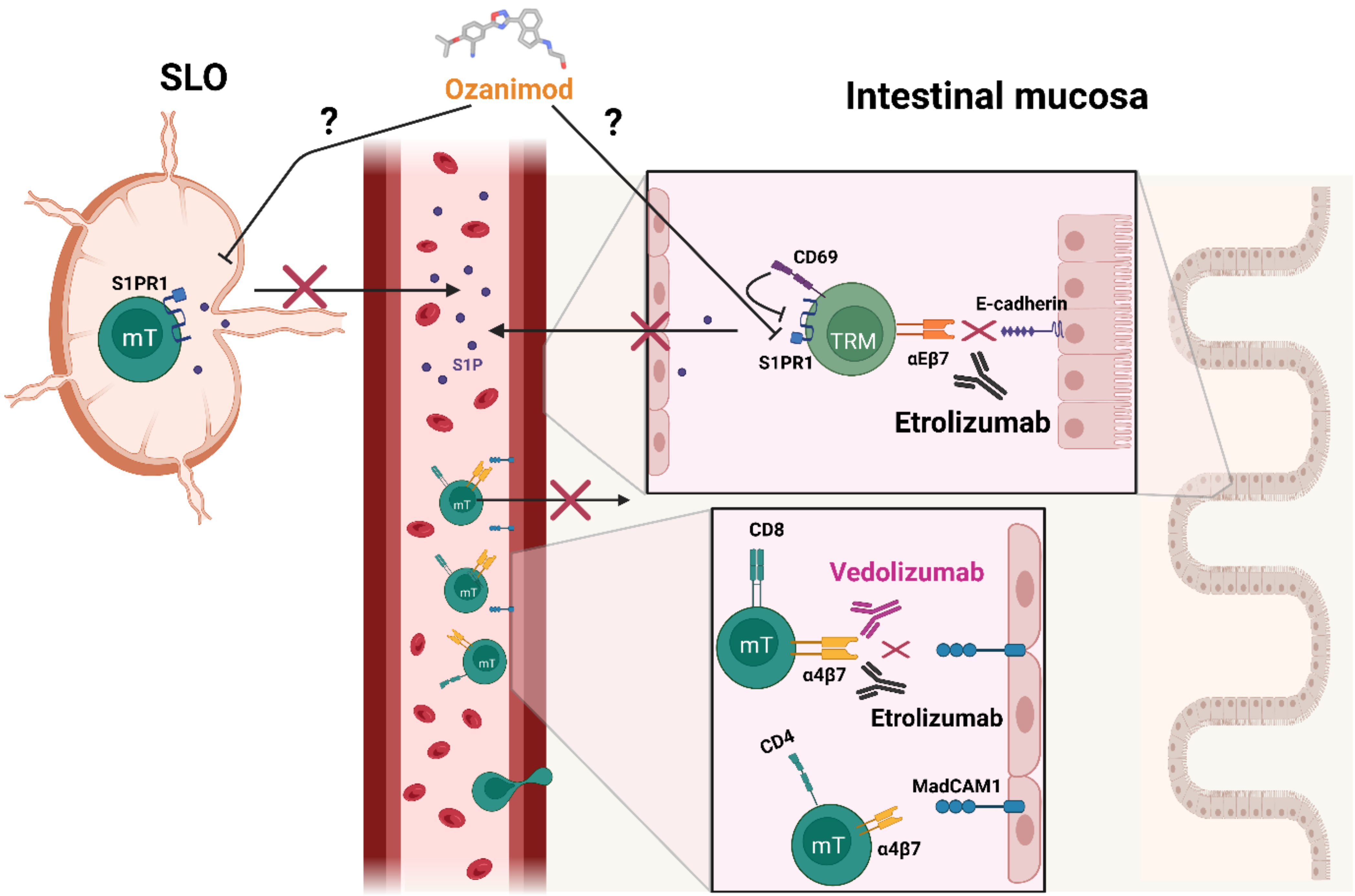
- Soler, D.; Chapman, T.; Yang, L.-L.; Wyant, T.; Egan, R.; Fedyk, E.R. The binding specificity and selective antagonism of vedolizumab, an anti-α4β7 integrin therapeutic antibody in development for inflammatory bowel diseases. J. Pharmacol. Exp. Ther. 2009, 330, 864–875. [Google Scholar] [CrossRef]
- Wyant, T.; Fedyk, E.; Abhyankar, B. An Overview of the Mechanism of Action of the Monoclonal Antibody Vedolizumab. J. Crohn’s Colitis 2016, 10, 1437–1444. [Google Scholar] [CrossRef] [PubMed]
- Sandborn, W.J.; Vermeire, S.; Tyrrell, H.; Hassanali, A.; Lacey, S.; Tole, S.; Tatro, A.R.; The Etrolizumab Global Steering, C. Etrolizumab for the Treatment of Ulcerative Colitis and Crohn’s Disease: An Overview of the Phase 3 Clinical Program. Adv. Ther. 2020, 37, 3417–3431. [Google Scholar] [CrossRef] [PubMed]
- Tew, G.W.; Hackney, J.A.; Gibbons, D.; Lamb, C.A.; Luca, D.; Egen, J.G.; Diehl, L.; Eastham Anderson, J.; Vermeire, S.; Mansfield, J.C.; et al. Association Between Response to Etrolizumab and Expression of Integrin αE and Granzyme A in Colon Biopsies of Patients with Ulcerative Colitis. Gastroenterology 2016, 150, 477–487.e479. [Google Scholar] [CrossRef]
- Zundler, S.; Schillinger, D.; Fischer, A.; Atreya, R.; López-Posadas, R.; Watson, A.; Neufert, C.; Atreya, I.; Neurath, M.F. Blockade of αEβ7 integrin suppresses accumulation of CD8+ and Th9 lymphocytes from patients with IBD in the inflamed gut in vivo. Gut 2017, 66, 1936–1948. [Google Scholar] [CrossRef]
- Comi, G.; Kappos, L.; Selmaj, K.W.; Bar-Or, A.; Arnold, D.L.; Steinman, L.; Hartung, H.-P.; Montalban, X.; Kubala Havrdová, E.; Cree, B.A.C.; et al. Safety and efficacy of ozanimod versus interferon beta-1a in relapsing multiple sclerosis (SUNBEAM): A multicentre, randomised, minimum 12-month, phase 3 trial. Lancet Neurol. 2019, 18, 1009–1020. [Google Scholar] [CrossRef]
- Sandborn, W.J.; Feagan, B.G.; Wolf, D.C.; D’Haens, G.; Vermeire, S.; Hanauer, S.B.; Ghosh, S.; Smith, H.; Cravets, M.; Frohna, P.A.; et al. Ozanimod Induction and Maintenance Treatment for Ulcerative Colitis. N. Engl. J. Med. 2016, 374, 1754–1762. [Google Scholar] [CrossRef]
- Sandborn, W.J.; Feagan, B.G.; D’Haens, G.; Wolf, D.C.; Jovanovic, I.; Hanauer, S.B.; Ghosh, S.; Petersen, A.; Hua, S.Y.; Lee, J.H.; et al. Ozanimod as Induction and Maintenance Therapy for Ulcerative Colitis. N. Engl. J. Med. 2021, 385, 1280–1291. [Google Scholar] [CrossRef] [PubMed]
- Feagan, B.G.; Sandborn, W.J.; Danese, S.; Wolf, D.C.; Liu, W.J.; Hua, S.Y.; Minton, N.; Olson, A.; D’Haens, G. Ozanimod induction therapy for patients with moderate to severe Crohn’s disease: A single-arm, phase 2, prospective observer-blinded endpoint study. Lancet Gastroenterol. Hepatol. 2020, 5, 819–828. [Google Scholar] [CrossRef]
| mT Cell Subset | Interacting with | Cytokines/ Mediators Involved | Type of Interaction | Impact on Inflammation | Reference |
|---|---|---|---|---|---|
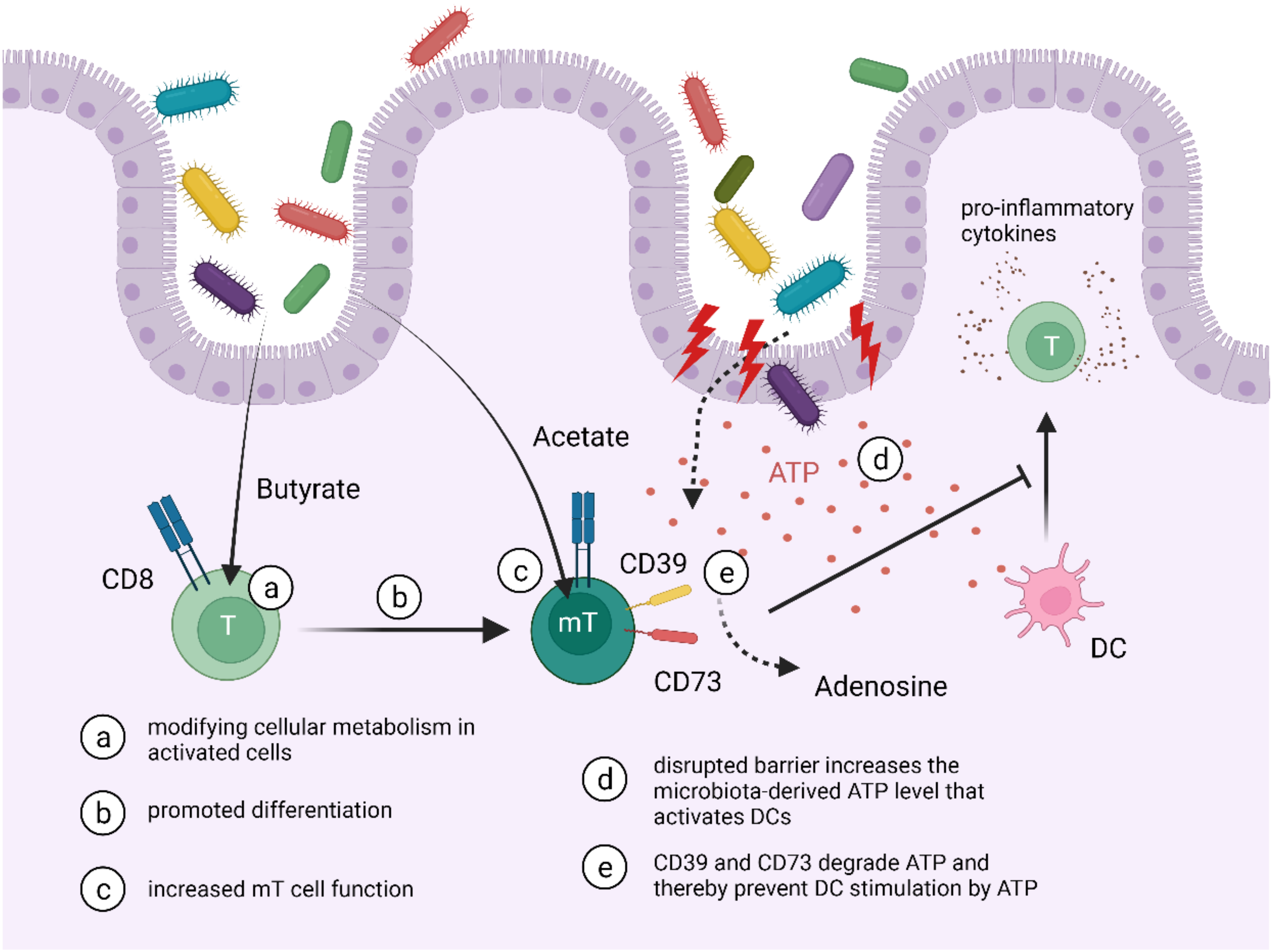
| CD39+ CD73+ TRMs | DCs | ATP | TRMs can degrade ATP released by bacteria preventing DC activation | anti-inflammatory | [78,89,90] |
| CD103+ TRMs | Paneth cells | IL22 | TRMs release IL22 leading to Paneth cell stimulation and release of antimicrobial peptides | anti-inflammatory | [92,93] |
| CD103+ TRMs | CCR6+ cells | CCL20 | CCL20 expressed by TRMs can attract CCR6+ cells | dichotomic | [92] |
| TRMs | Tregs | TGF-β1 | Recruited Tregs lead to production of bioavailable TGF-β1 production and development of TRMs | under pro-inflammatory conditions | [96,97] |
| TEMs | DCs and macrophages | CD40L, TNFα | TEMs expressing CD40L and TNFα activate myeloid cells | pro-inflammatory | [99] |
| TRMs | Monocytes | IL10, TGF-β | IL10 stimulates monocytes to secrete TGF-β leading to upregulation of CD103 | — | [100] |
| CD4+ TRMs | Microbiota (bacterial species associated with IBD) | TNFα, IL2, IL17A | Upon microbiota sensing, TRMs secrete pro-inflammatory cytokines | pro-inflammatory | [83] |
| CD8+ TRMs | Commensals/pathogenic microorganisms | IFNγ | Microorganisms directly evoke TRMs leading to IFNγ signalling | pro-inflammatory | [76] |
| CD8+ TRMs | Microbiota | SCFAs (butyrate, acetate) | Microbiota can induce the CD8+ T cell to CD8+ TRM transition and increased memory properties | — | [87,88] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Knauss, A.; Gabel, M.; Neurath, M.F.; Weigmann, B. The Memory T Cell “Communication Web” in Context with Gastrointestinal Disorders—How Memory T Cells Affect Their Surroundings and How They Are Influenced by It. Cells 2022, 11, 2780. https://doi.org/10.3390/cells11182780
Knauss A, Gabel M, Neurath MF, Weigmann B. The Memory T Cell “Communication Web” in Context with Gastrointestinal Disorders—How Memory T Cells Affect Their Surroundings and How They Are Influenced by It. Cells. 2022; 11(18):2780. https://doi.org/10.3390/cells11182780
Chicago/Turabian StyleKnauss, Annkathrin, Michael Gabel, Markus F. Neurath, and Benno Weigmann. 2022. "The Memory T Cell “Communication Web” in Context with Gastrointestinal Disorders—How Memory T Cells Affect Their Surroundings and How They Are Influenced by It" Cells 11, no. 18: 2780. https://doi.org/10.3390/cells11182780
APA StyleKnauss, A., Gabel, M., Neurath, M. F., & Weigmann, B. (2022). The Memory T Cell “Communication Web” in Context with Gastrointestinal Disorders—How Memory T Cells Affect Their Surroundings and How They Are Influenced by It. Cells, 11(18), 2780. https://doi.org/10.3390/cells11182780








