Preimplantation Genetic Testing for Aneuploidy Improves Live Birth Rates with In Vitro Produced Bovine Embryos: A Blind Retrospective Study
Abstract
1. Introduction
2. Methods
2.1. Study Design
2.2. Ovarian Stimulation and In Vitro Embryo Production
2.3. Embryo Biopsy, WGA, SNP Genotyping, and ET
2.4. Computing and PGT-A
2.5. Limitations
2.6. Statistical Analysis
2.7. Ethical Statement
2.8. Data Availability Statement
3. Results
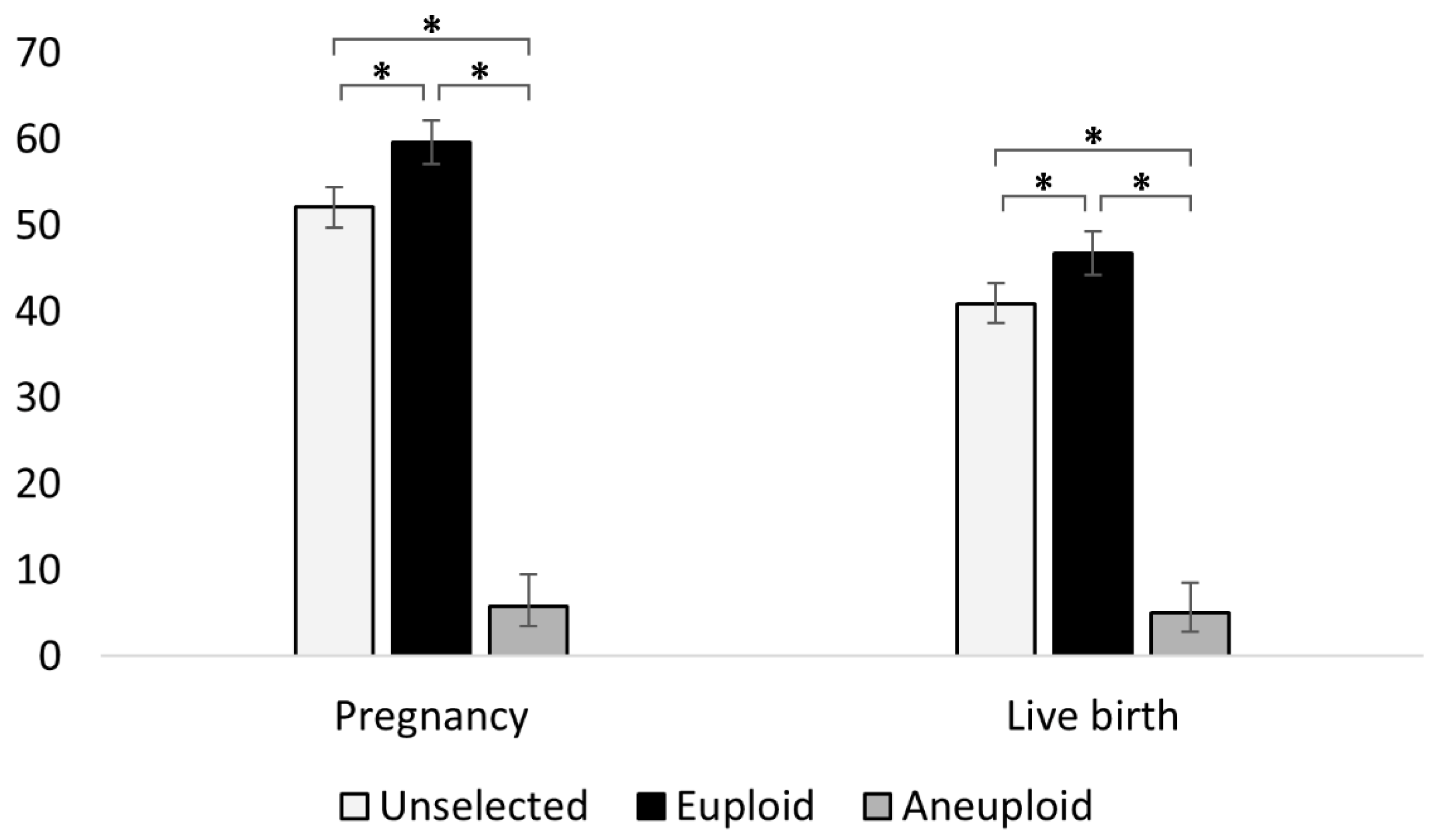
3.1. PGT-A Improves Pregnancy and Live Birth Outcomes
3.2. Chromosomal Abnormality Classes: Maternal Meiotic Errors Are by Far the Most Frequent
3.3. Mosaicism Incidence, Pregnancy, and Live Birth Outcomes in Chromosomally Abnormal Embryos
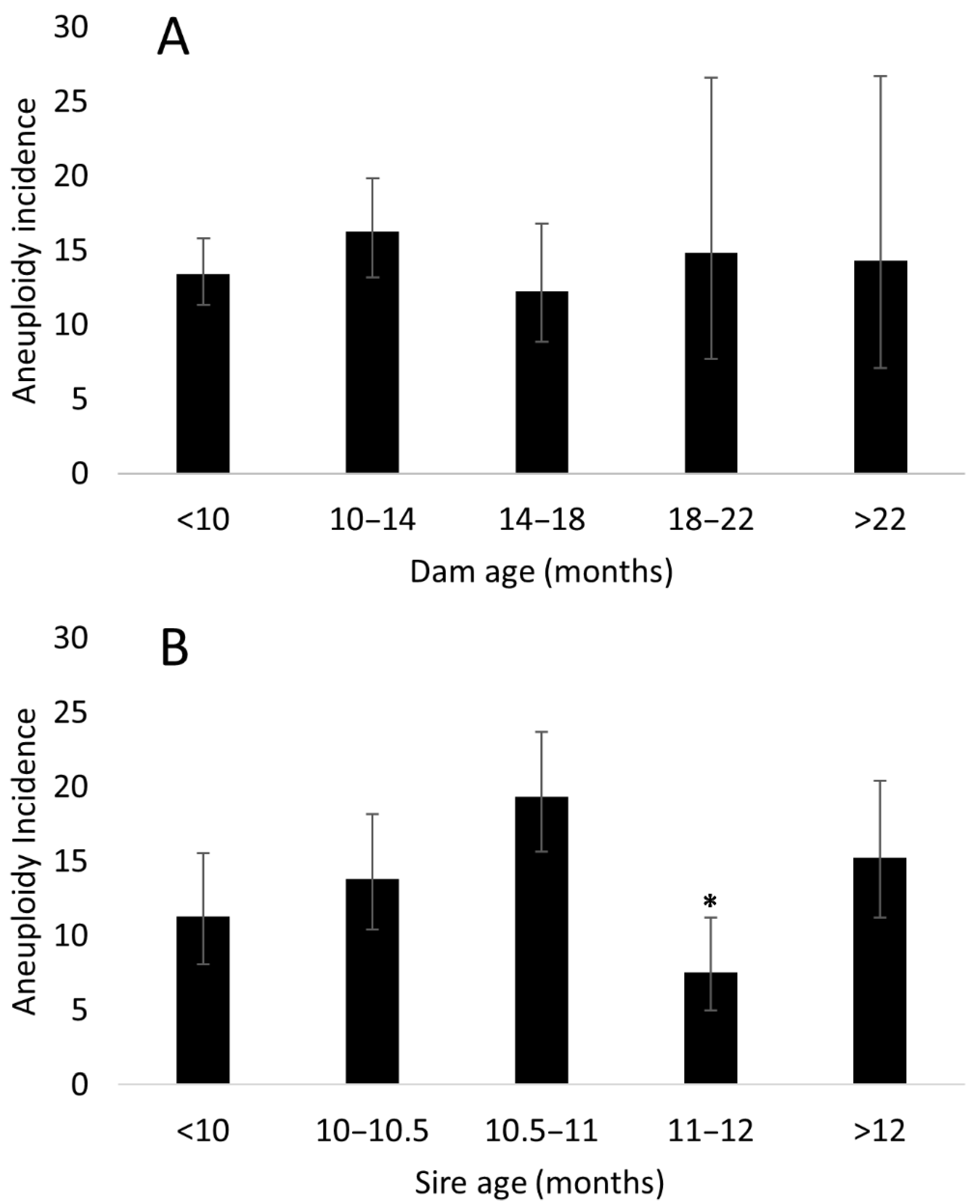
3.4. Parental Effects on Aneuploidy Incidence
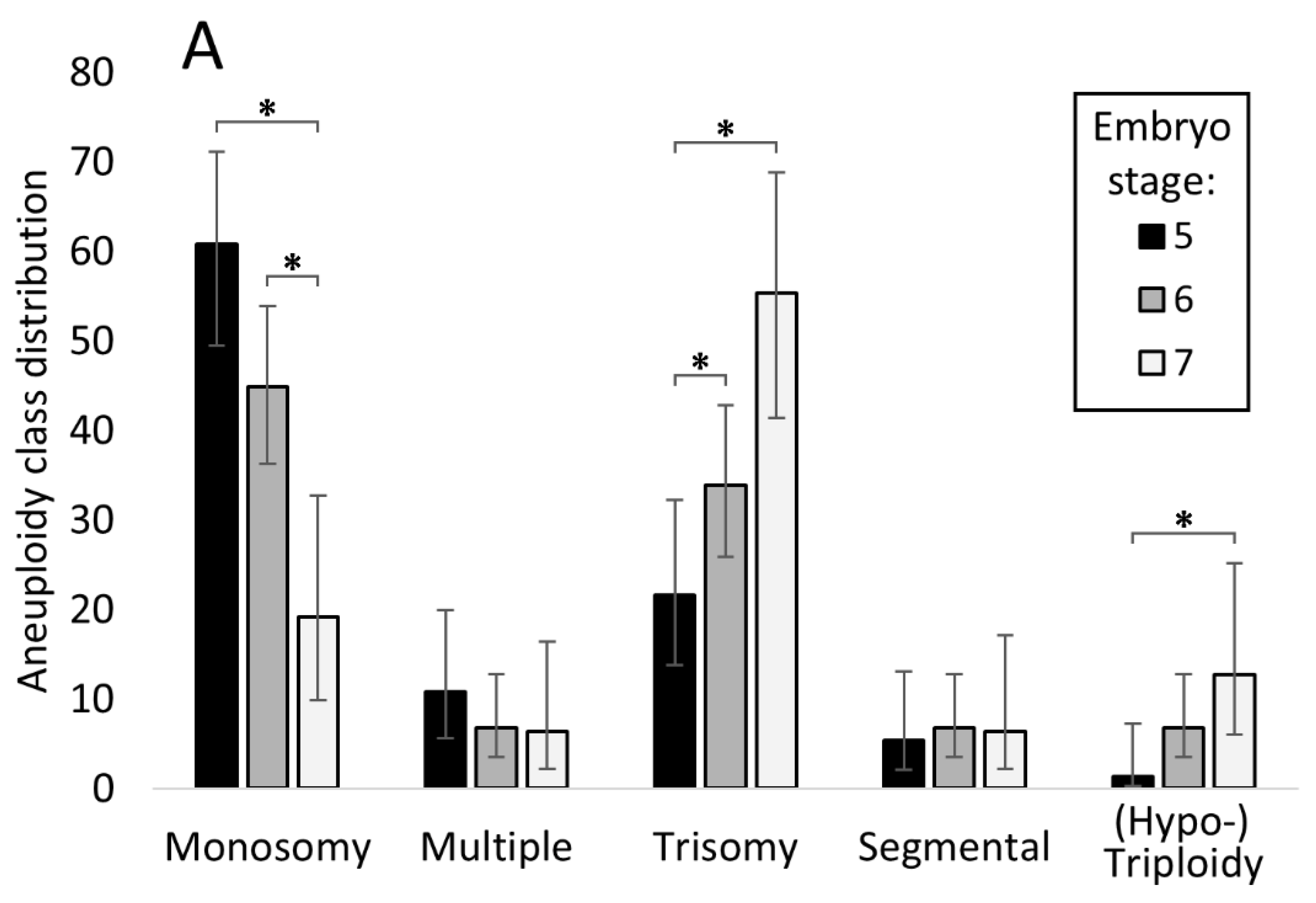
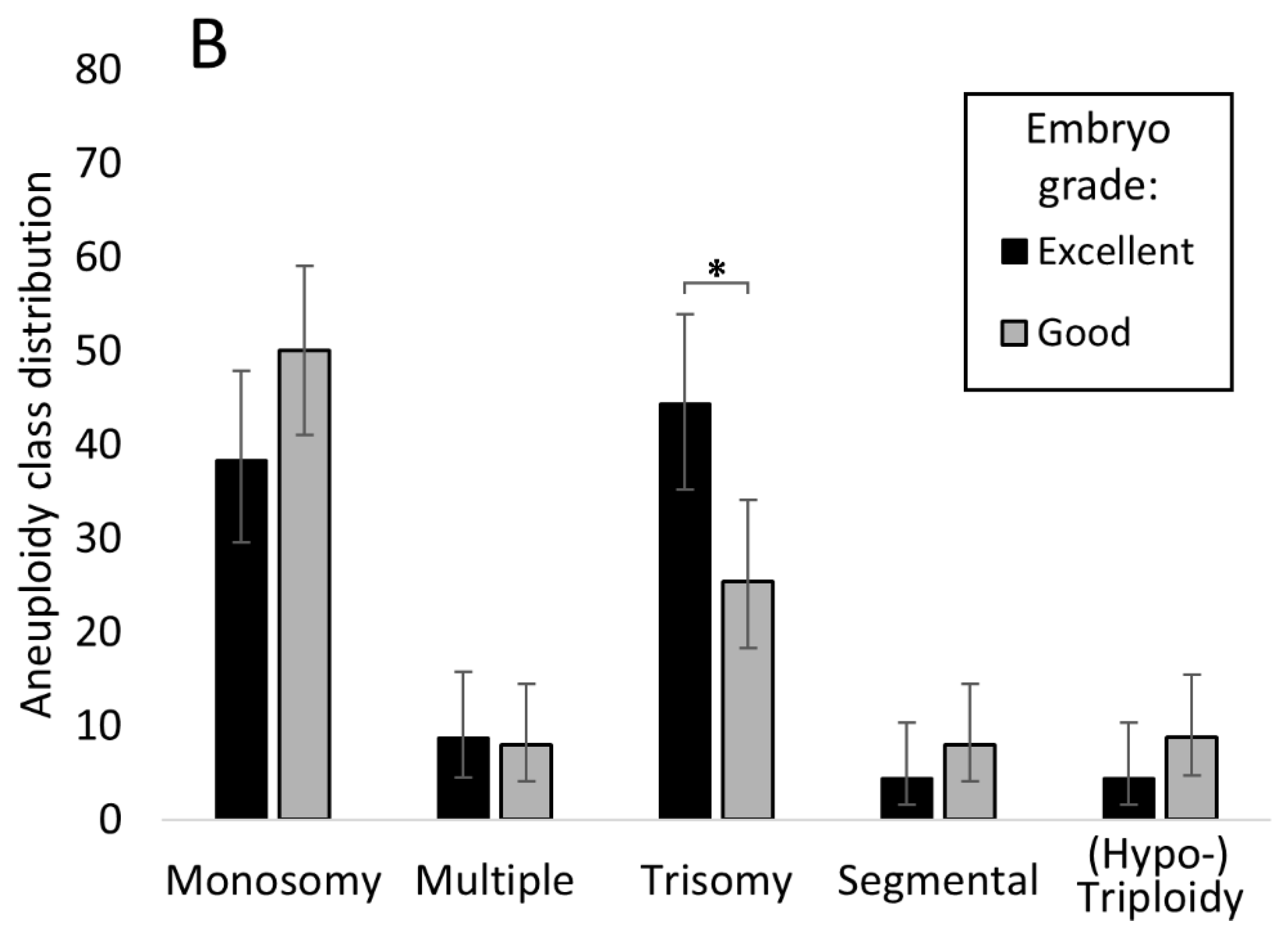
3.5. Relationship between Embryo Stage and Grade with Aneuploidy and Pregnancy/Live Birth Rates
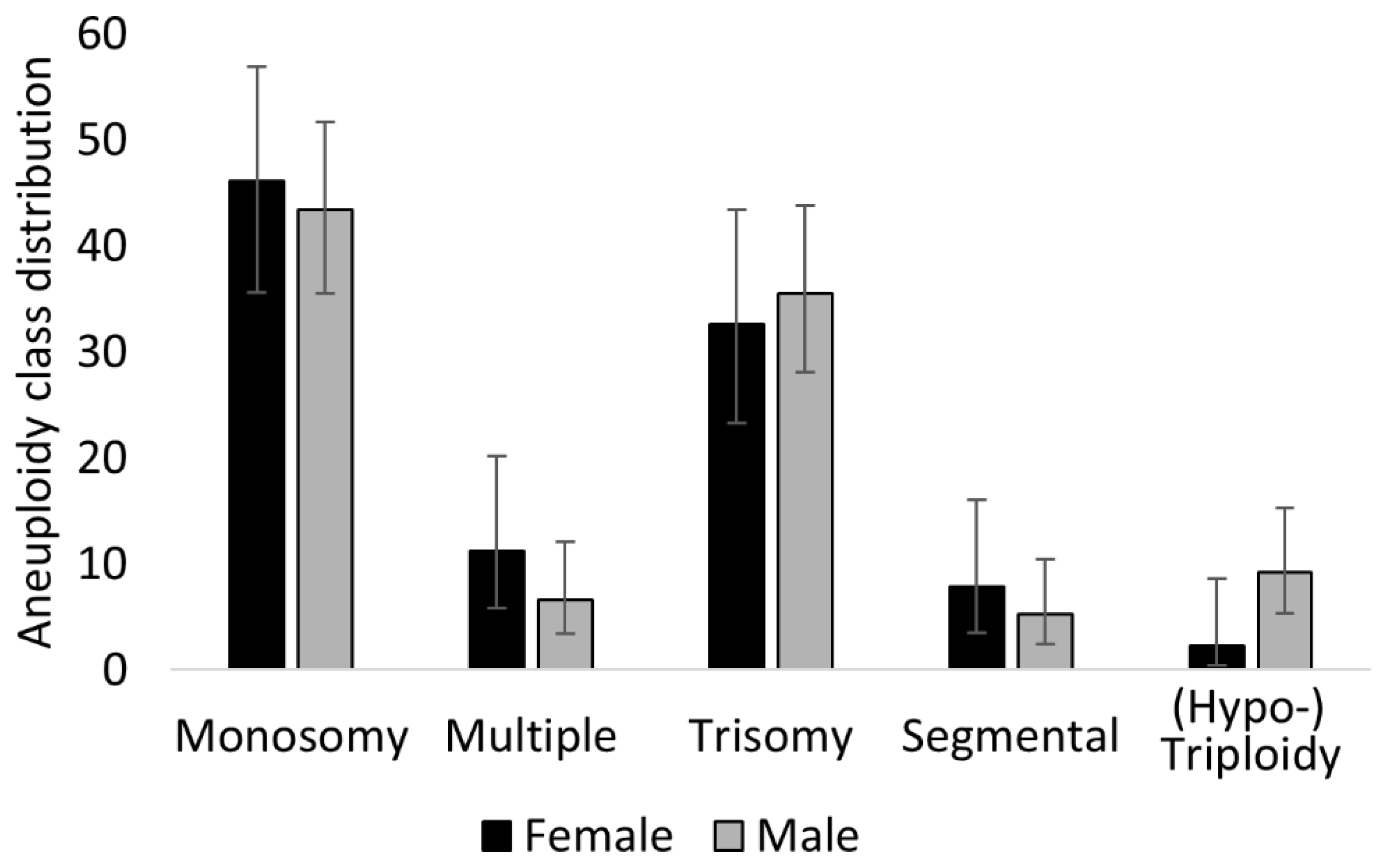
3.6. Male Embryos (XY) Achieve Higher Morphology Scores But Are Disproportionately Affected by Aneuploidy
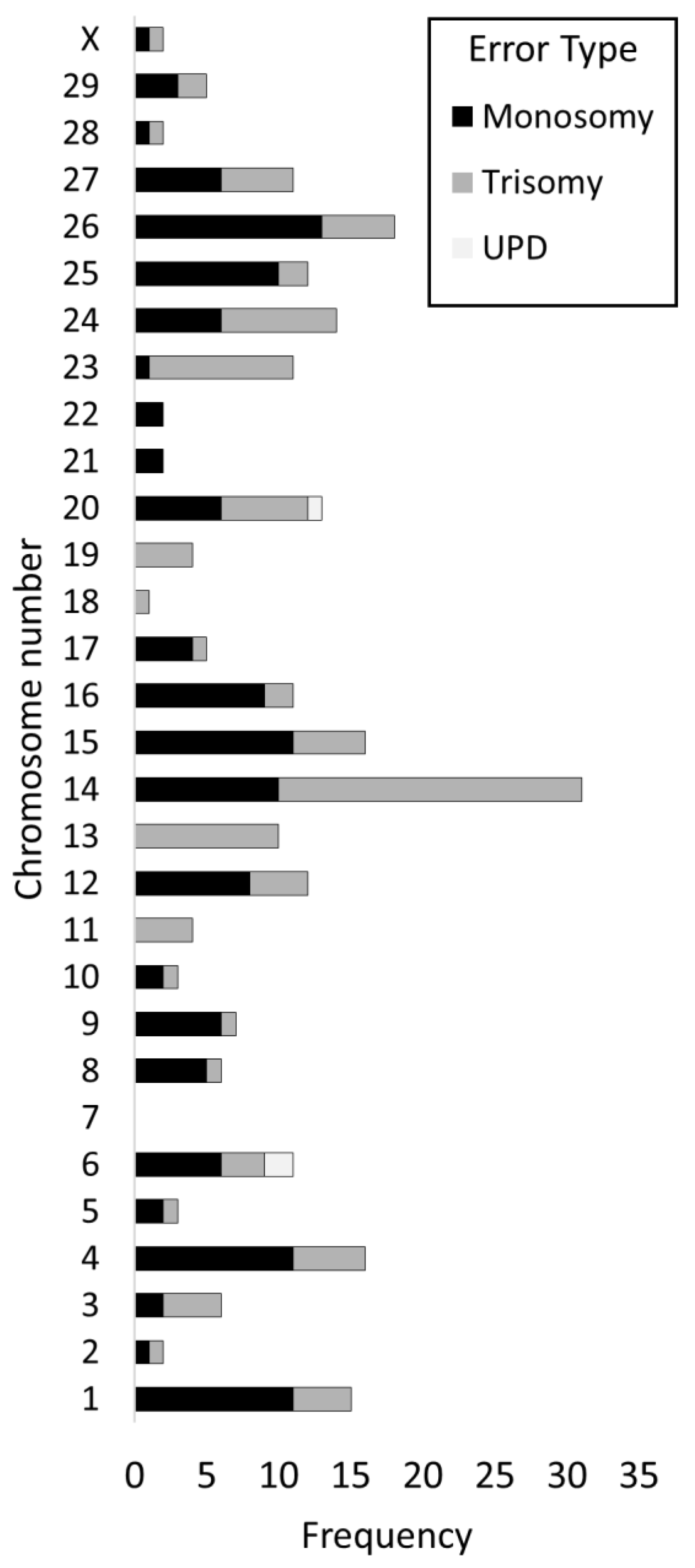
3.7. Incidence of Errors by Chromosome
3.8. Chromosomal Loss in Hypotriploidy
4. Discussion
4.1. Relationship between Chromosomal Abnormality and Embryo Morphology
4.2. Certain Chromosomes Are Preferentially Affected by Aneuploidy
4.3. Trisomic Embryos Survive Longer than Embryos Carrying a Monosomy
4.4. Segmental and Mosaic Errors
4.5. The Origin of Aneuploidy in Cattle IVP
4.6. Differences between XX and XY Embryos
4.7. The Wider Context
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ferré, L.; Kjelland, M.; Strøbech, L.; Hyttel, P.; Mermillod, P.; Ross, P. Recent Advances in Bovine in Vitro Embryo Production: Reproductive Biotechnology History and Methods. Animal 2020, 14, 991–1004. [Google Scholar] [CrossRef]
- Mullaart, E.; Wells, D. Embryo Biopsies for Genomic Selection. In Animal Biotechnology; Niemann, H., Wrenzycki, C., Eds.; Springer Nature: Cham, Switzerland, 2018; Volume 2, pp. 81–94. [Google Scholar]
- Fujii, T.; Naito, A.; Hirayama, H.; Kashima, M.; Yoshino, H.; Hanamure, T.; Domon, Y.; Hayakawa, H.; Watanabe, T.; Moriyasu, S. Potential of Preimplantation Genomic Selection for Carcass Traits in Japanese Black Cattle. J. Reprod. Dev. 2019, 65, 251–258. [Google Scholar] [CrossRef]
- Meuwissen, T.H.; Hayes, B.J.; Goddard, M.E. Prediction of Total Genetic Value using Genome-Wide Dense Marker Maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [CrossRef]
- Hansen, P.J. The Incompletely Fulfilled Promise of Embryo Transfer in cattle—why aren’t Pregnancy Rates Greater and what can we do about it? J. Anim. Sci. 2020, 98, skaa288. [Google Scholar] [CrossRef]
- Munné, S.; Kaplan, B.; Frattarelli, J.L.; Child, T.; Nakhuda, G.; Shamma, F.N.; Silverberg, K.; Kalista, T.; Handyside, A.H.; Katz-Jaffe, M. Preimplantation Genetic Testing for Aneuploidy Versus Morphology as Selection Criteria for Single Frozen-Thawed Embryo Transfer in Good-Prognosis Patients: A Multicenter Randomized Clinical Trial. Fertil. Steril. 2019, 112, 1071–1079. [Google Scholar] [CrossRef]
- Pagliardini, L.; Viganò, P.; Alteri, A.; Corti, L.; Somigliana, E.; Papaleo, E. Shooting STAR: Reinterpreting the Data from the ‘Single Embryo TrAnsfeR of Euploid Embryo’randomized Clinical Trial. Reprod. Biomed. Online 2020, 40, 475–478. [Google Scholar] [CrossRef]
- Griffin, D.K.; Ogur, C. Chromosomal Analysis in IVF: Just how Useful is it? Reproduction 2018, 156, F29–F50. [Google Scholar] [CrossRef] [PubMed]
- Hornak, M.; Kubicek, D.; Broz, P.; Hulinska, P.; Hanzalova, K.; Griffin, D.; Machatkova, M.; Rubes, J. Aneuploidy Detection and mtDNA Quantification in Bovine Embryos with Different Cleavage Onset using a Next-Generation Sequencing-Based Protocol. Cytogenet. Genome Res. 2016, 150, 60–67. [Google Scholar] [CrossRef]
- Turner, K.J.; Silvestri, G.; Black, D.H.; Dobson, G.; Smith, C.; Handyside, A.H.; Sinclair, K.D.; Griffin, D.K. Karyomapping for Simultaneous Genomic Evaluation and Aneuploidy Screening of Preimplantation Bovine Embryos: The First Live-Born Calves. Theriogenology 2019, 125, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Tutt, D.; Silvestri, G.; Serrano-Albal, M.; Simmons, R.; Kwong, W.; Guven-Ates, G.; Canedo-Ribeiro, C.; Labrecque, R.; Blondin, P.; Handyside, A. Analysis of Bovine Blastocysts Indicates Ovarian Stimulation does Not Induce Chromosome Errors, nor Discordance between Inner-Cell Mass and Trophectoderm Lineages. Theriogenology 2021, 161, 108–119. [Google Scholar] [CrossRef]
- Attiyeh, E.F.; Diskin, S.J.; Attiyeh, M.A.; Mosse, Y.P.; Hou, C.; Jackson, E.M.; Kim, C.; Glessner, J.; Hakonarson, H.; Biegel, J.A.; et al. Genomic Copy Number Determination in Cancer Cells from Single Nucleotide Polymorphism Microarrays Based on Quantitative Genotyping Corrected for Aneuploidy. Genome Res. 2009, 19, 276–283. [Google Scholar] [CrossRef][Green Version]
- Handyside, A.H.; Harton, G.L.; Mariani, B.; Thornhill, A.R.; Affara, N.; Shaw, M.A.; Griffin, D.K. Karyomapping: A Universal Method for Genome Wide Analysis of Genetic Disease Based on Mapping Crossovers between Parental Haplotypes. J. Med. Genet. 2010, 47, 651–658. [Google Scholar] [CrossRef]
- Gabriel, A.S.; Hassold, T.J.; Thornhill, A.R.; Affara, N.A.; Handyside, A.H.; Griffin, D.K. An Algorithm for Determining the Origin of Trisomy and the Positions of Chiasmata from SNP Genotype Data. Chromosome Res. 2011, 19, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Landry, D.A.; Bellefleur, A.; Labrecque, R.; Grand, F.; Vigneault, C.; Blondin, P.; Sirard, M. Effect of Cow Age on the in Vitro Developmental Competence of Oocytes obtained After FSH Stimulation and Coasting Treatments. Theriogenology 2016, 86, 1240–1246. [Google Scholar] [CrossRef]
- Miller, G.; Gliedt, D.; Rakes, J.; Rorie, R. Addition of Penicillamine, Hypotaurine and Epinephrine (PHE) Or Bovine Oviductal Epithelial Cells (BOEC) Alone Or in Combination to Bovine in Vitro Fertilization Medium Increases the Subsequent Embryo Cleavage Rate. Theriogenology 1994, 41, 689–696. [Google Scholar] [CrossRef]
- Bó, G.; Mapletoft, R. Evaluation and Classification of Bovine Embryos. Anim. Reprod. (AR) 2018, 10, 344–348. [Google Scholar]
- Blondin, P.; Farin, P.W.; Crosier, A.E.; Alexander, J.E.; Farin, C.E. In Vitro Production of Embryos Alters Levels of Insulin-Like Growth Factor-II Messenger Ribonucleic Acid in Bovine Fetuses 63 Days After Transfer. Biol. Reprod. 2000, 62, 384–389. [Google Scholar] [CrossRef]
- Kubicek, D.; Hornak, M.; Horak, J.; Navratil, R.; Tauwinklova, G.; Rubes, J.; Vesela, K. Incidence and Origin of Meiotic Whole and Segmental Chromosomal Aneuploidies Detected by Karyomapping. Reprod. Biomed. Online 2019, 38, 330–339. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing, 4.0.3; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Gel, B.; Serra, E. KaryoploteR: An R/Bioconductor Package to Plot Customizable Genomes Displaying Arbitrary Data. Bioinformatics 2017, 33, 3088–3090. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Wright, F.A.; Tang, Z.; Nordgard, S.H.; Loo, P.V.; Yu, T.; Kristensen, V.N.; Perou, C.M. Integrated Study of Copy Number States and Genotype Calls using High-Density SNP Arrays. Nucleic Acids Res. 2009, 37, 5365–5377. [Google Scholar] [CrossRef]
- Viana, J. 2017 Statistics of Embryo Production and Transfer in Domestic Farm Animals. Embryo Tech. News. 2018, 36, 8–25. [Google Scholar]
- Alfarawati, S.; Fragouli, E.; Colls, P.; Stevens, J.; Gutiérrez-Mateo, C.; Schoolcraft, W.B.; Katz-Jaffe, M.G.; Wells, D. The Relationship between Blastocyst Morphology, Chromosomal Abnormality, and Embryo Gender. Fertil. Steril. 2011, 95, 520–524. [Google Scholar] [CrossRef]
- Baryshnikova, A.; VanderSluis, B.; Costanzo, M.; Myers, C.L.; Cha, R.S.; Andrews, B.; Boone, C. Global Linkage Map Connects Meiotic Centromere Function to Chromosome Size in Budding Yeast. G3: Genes Genomes Genet. 2013, 3, 1741–1751. [Google Scholar] [CrossRef] [PubMed][Green Version]
- McCoy, R.C. Mosaicism in Preimplantation Human Embryos: When Chromosomal Abnormalities are the Norm. Trends Genet. 2017, 33, 448–463. [Google Scholar] [CrossRef] [PubMed]
- Kahraman, S.; Cetinkaya, M.; Yuksel, B.; Yesil, M.; Pirkevi Cetinkaya, C. The Birth of a Baby with Mosaicism Resulting from a Known Mosaic Embryo Transfer: A Case Report. Hum. Reprod. 2020, 35, 727–733. [Google Scholar] [CrossRef]
- Popovic, M.; Dhaenens, L.; Taelman, J.; Dheedene, A.; Bialecka, M.; De Sutter, P.; Chuva de Sousa Lopes, S.M.; Menten, B.; Heindryckx, B. Extended in Vitro Culture of Human Embryos Demonstrates the Complex Nature of Diagnosing Chromosomal Mosaicism from a Single Trophectoderm Biopsy. Hum. Reprod. 2019, 34, 758–769. [Google Scholar] [CrossRef] [PubMed]
- Hassold, T.; Hunt, P. To Err (Meiotically) is Human: The Genesis of Human Aneuploidy. Nat. Rev. Genet. 2001, 2, 280–291. [Google Scholar] [CrossRef]
- Tšuiko, O.; Catteeuw, M.; Zamani Esteki, M.; Destouni, A.; Bogado Pascottini, O.; Besenfelder, U.; Havlicek, V.; Smits, K.; Kurg, A.; Salumets, A.; et al. Genome stability of bovine in vivo-conceived cleavage-stage embryos is higher compared to in vitro-produced embryos. Hum. Reprod. 2017, 32, 2348–2357. [Google Scholar] [CrossRef] [PubMed]
- Eaton, J.L.; Hacker, M.R.; Barrett, C.B.; Thornton, K.L.; Penzias, A.S. Influence of Embryo Sex on Development to the Blastocyst Stage and Euploidy. Fertil. Steril. 2011, 95, 936–939. [Google Scholar] [CrossRef]
- Wang, A.; Kort, J.; Behr, B.; Westphal, L.M. Euploidy in Relation to Blastocyst Sex and Morphology. J. Assist. Reprod. Genet. 2018, 35, 1565–1572. [Google Scholar] [CrossRef]

| SNP Chip Name | Number of SNPs | Embryos Tested (n) | Parents Tested (n) |
|---|---|---|---|
| GGP Bovine HD 150k v01 | 138,892 | 379 | - |
| GGP Bovine HD 150k v03 | 139,376 | 1241 | - |
| GGP Bovine HD 150k v04 | 140,668 | 112 | - |
| GGP Bovine LD v04 | 30,105 | 5 | - |
| GGP Bovine SNP50 | 45,187 | - | 241 |
| (A) | Origin | |||
| Aneuploidy class | Overall | Dam | Sire | Embryo |
| Segmental errors | 21 | 2 | 14 | 5 |
| Triploidy and hypotriploidy | 16 | 6 | 10 | - |
| Whole chromosome | 255 | 212 | 16 | 27 |
| Total errors | 292 | 220 | 40 | 32 |
| (B) | Origin | |||
| Whole chromosome | Overall | Dam | Sire | Embryo |
| Trisomy | 113 | 90 | 1 | 22 |
| MI | 84 | 83 | 1 | - |
| MII | 7 | 7 | - | - |
| Mitotic | 22 | - | - | 22 |
| Monosomy | 139 | 120 | 15 | 4 |
| UPD | 3 | 2 | - | 1 |
| Sample | Sex | Embryo Stage | Embryo Grade | Diagnosis | Chr | Mosaic | % Mosaicism |
|---|---|---|---|---|---|---|---|
| 1536 | M | 6 | 1 | Maternal Monosomy | 4 | - | - |
| 816 | F | 5 | 2 | Maternal Monosomy | 22 | - | - |
| 1173 | M | 7 | 1 | Maternal Trisomy | 4 | - | - |
| 37 | M | 7 | 1 | Maternal Trisomy | 26 | - | - |
| 1437 | M | 6 | 1 | Maternal Trisomy | 26 | - | - |
| 1127 | M | 6 | 1 | Maternal Trisomy | 27 | - | - |
| 1524 | M | 5 | 2 | Mitotic Trisomy | 6 | Yes | 76.0% |
| 257 | M | 6 | 2 | Segmental Deletion | 17 | Yes | 47.3% |
| 482 | F | 6 | 2 | UPD | 6 | - | - |
| 1021 | M | 8 | 1 | Hypotriploidy (polyspermy) | n/a | Yes | 21.7 to 41.4% |
| 353 | M | 6 | 2 | Hypotriploidy (polyspermy) | n/a | - | - |
| 1407 | M | 6 | 2 | Hypotriploidy (polyspermy) | n/a | Yes | 21.0 to 47.0% |
| Embryo Stage | n | Aneuploidy Incidence (%) | Pregnancy Rate D60 (%) | Live Birth Rate (%) |
|---|---|---|---|---|
| 5 | 308 | 24.0 (19.6–29.1) | 47.7 (42.2–53.3) | 35.4 (30.3–40.9) |
| 6 | 786 | 15.0 (12.7–17.7) | 50.5 (47.0–54.0) | 41.0 (37.6–44.4) |
| 7 | 515 | 9.1 (6.9–11.9) | 54.9 (50.6–59.2) | 42.1 (37.9–46.5) |
| 8–9 | 103 | 1.9 (0.5–6.8) | 62.1 (52.5–70.9) | 49.5 (40.0–59.0) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silvestri, G.; Canedo-Ribeiro, C.; Serrano-Albal, M.; Labrecque, R.; Blondin, P.; Larmer, S.G.; Marras, G.; Tutt, D.A.R.; Handyside, A.H.; Farré, M.; et al. Preimplantation Genetic Testing for Aneuploidy Improves Live Birth Rates with In Vitro Produced Bovine Embryos: A Blind Retrospective Study. Cells 2021, 10, 2284. https://doi.org/10.3390/cells10092284
Silvestri G, Canedo-Ribeiro C, Serrano-Albal M, Labrecque R, Blondin P, Larmer SG, Marras G, Tutt DAR, Handyside AH, Farré M, et al. Preimplantation Genetic Testing for Aneuploidy Improves Live Birth Rates with In Vitro Produced Bovine Embryos: A Blind Retrospective Study. Cells. 2021; 10(9):2284. https://doi.org/10.3390/cells10092284
Chicago/Turabian StyleSilvestri, Giuseppe, Carla Canedo-Ribeiro, María Serrano-Albal, Remi Labrecque, Patrick Blondin, Steven G. Larmer, Gabriele Marras, Desmond A.R. Tutt, Alan H. Handyside, Marta Farré, and et al. 2021. "Preimplantation Genetic Testing for Aneuploidy Improves Live Birth Rates with In Vitro Produced Bovine Embryos: A Blind Retrospective Study" Cells 10, no. 9: 2284. https://doi.org/10.3390/cells10092284
APA StyleSilvestri, G., Canedo-Ribeiro, C., Serrano-Albal, M., Labrecque, R., Blondin, P., Larmer, S. G., Marras, G., Tutt, D. A. R., Handyside, A. H., Farré, M., Sinclair, K. D., & Griffin, D. K. (2021). Preimplantation Genetic Testing for Aneuploidy Improves Live Birth Rates with In Vitro Produced Bovine Embryos: A Blind Retrospective Study. Cells, 10(9), 2284. https://doi.org/10.3390/cells10092284








