Somatic Mutations and Autoimmunity
Abstract
1. Introduction
2. The Pathogenesis of Autoimmune Diseases
3. The Somatic Mutation Hypothesis
3.1. The History of the Somatic Mutation Hypothesis
3.2. Somatic Mutations
3.3. Somatic Mutations in Non-Malignant Diseases
3.4. Frequency of Somatic Mutations
3.5. Epidemiological Evidence for the Somatic Mutation Hypothesis
3.6. Somatic Mutation in Autoimmune Diseases
4. Practical Implications and Potential Benefits
5. Future Research Prospects
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cooper, G.S.; Bynum, M.L.; Somers, E.C. Recent insights in the epidemiology of autoimmune diseases: Improved prevalence estimates and understanding of clustering of diseases. J. Autoimmun. 2009, 33, 197–207. [Google Scholar] [CrossRef]
- Wang, L.; Wang, F.S.; Gershwin, M.E. Human autoimmune diseases: A comprehensive update. J. Intern. Med. 2015, 278, 369–395. [Google Scholar] [CrossRef]
- Goodnow, C.C. Multistep pathogenesis of autoimmune disease. Cell 2007, 130, 25–35. [Google Scholar] [CrossRef]
- Chaplin, D.D. Overview of the immune response. J. Allergy Clin. Immunol. 2010, 125, S3–S23. [Google Scholar] [CrossRef]
- Vafiadis, P.; Bennett, S.T.; Todd, J.A.; Nadeau, J.; Grabs, R.; Goodyer, C.G.; Wickramasinghe, S.; Colle, E.; Polychronakos, C. Insulin expression in human thymus is modulated by INS VNTR alleles at the IDDM2 locus. Nat. Genet. 1997, 15, 289–292. [Google Scholar] [CrossRef]
- Goodnow, C.C.; Sprent, J.; Fazekas de St Groth, B.; Vinuesa, C.G. Cellular and genetic mechanisms of self tolerance and autoimmunity. Nature 2005, 435, 590–597. [Google Scholar] [CrossRef]
- Vyse, T.J.; Todd, J.A. Genetic analysis of autoimmune disease. Cell 1996, 85, 311–318. [Google Scholar] [CrossRef]
- Derbinski, J.; Schulte, A.; Kyewski, B.; Klein, L. Promiscuous gene expression in medullary thymic epithelial cells mirrors the peripheral self. Nat. Immunol. 2001, 2, 1032–1039. [Google Scholar] [CrossRef] [PubMed]
- Betterle, C.; Greggio, N.A.; Volpato, M. Clinical review 93: Autoimmune polyglandular syndrome type 1. J. Clin. Endocrinol. Metab. 1998, 83, 1049–1055. [Google Scholar] [CrossRef] [PubMed]
- T., J.H. Horror Autotoxicus and other concepts of Paul Ehrlich. JAMA 1961, 176, 50–51. [Google Scholar] [CrossRef]
- Silverstein, A.M. Autoimmunity versus horror autotoxicus: The struggle for recognition. Nat. Immunol. 2001, 2, 279–281. [Google Scholar] [CrossRef]
- Metalnikoff, S.A. Études sur la spermotoxine. Ann Inst Pasteur 1900, 14, 577–589. [Google Scholar]
- Ehrlich, P.; Morgenroth, J. Zytotoxine als Antikörper. Berl klin Wochenschr 1901, 38, 251–255. [Google Scholar]
- Burnet, F.M. The Clonal Selection Theory of Acquired Immunity; Cambridge University Press: Cambridge, UK, 1959. [Google Scholar]
- Burnet, F.M. Autoimmunity and Autoimmune Disease; Medical and Technical Publishing Co Ltd.: Lancaster, UK, 1972. [Google Scholar]
- Burnet, F.M. A reassessment of the forbidden clone hypothesis of autoimmune disease. Aust. J. Exp. Biol. Med. Sci. 1972, 50, 1–9. [Google Scholar] [CrossRef]
- Buckley, P.G.; Mantripragada, K.K.; Piotrowski, A.; Diaz de Stahl, T.; Dumanski, J.P. Copy-number polymorphisms: Mining the tip of an iceberg. Trends Genet. 2005, 21, 315–317. [Google Scholar] [CrossRef]
- Behjati, S.; Huch, M.; van Boxtel, R.; Karthaus, W.; Wedge, D.C.; Tamuri, A.U.; Martincorena, I.; Petljak, M.; Alexandrov, L.B.; Gundem, G.; et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 2014, 513, 422–425. [Google Scholar] [CrossRef]
- Bae, T.; Tomasini, L.; Mariani, J.; Zhou, B.; Roychowdhury, T.; Franjic, D.; Pletikos, M.; Pattni, R.; Chen, B.J.; Venturini, E.; et al. Different mutational rates and mechanisms in human cells at pregastrulation and neurogenesis. Science 2018, 359, 550–555. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, A.J.; Miller, J.H.; Suzuki, D.T.; Lewontin, R.C.; Gelbart, W.M. Somatic versus germinal mutation. In An Introduction to Genetic Analysis; W. H. Freeman: New York, NY, USA, 2000. [Google Scholar]
- Watson, I.R.; Takahashi, K.; Futreal, P.A.; Chin, L. Emerging patterns of somatic mutations in cancer. Nat. Rev. Genet. 2013, 14, 703–718. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hernandez-Boluda, J.C.; Cervantes, F. Imatinib mesylate (Gleevec, Glivec): A new therapy for chronic myeloid leukemia and other malignancies. Drugs Today 2002, 38, 601–613. [Google Scholar] [CrossRef]
- Laurie, C.C.; Laurie, C.A.; Rice, K.; Doheny, K.F.; Zelnick, L.R.; McHugh, C.P.; Ling, H.; Hetrick, K.N.; Pugh, E.W.; Amos, C.; et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat. Genet. 2012, 44, 642–650. [Google Scholar] [CrossRef]
- Ignatowicz, L.; Kappler, J.; Marrack, P. The repertoire of T cells shaped by a single MHC/peptide ligand. Cell 1996, 84, 521–529. [Google Scholar] [CrossRef][Green Version]
- Laufer, T.M.; DeKoning, J.; Markowitz, J.S.; Lo, D.; Glimcher, L.H. Unopposed positive selection and autoreactivity in mice expressing class II MHC only on thymic cortex. Nature 1996, 383, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Wardemann, H.; Yurasov, S.; Schaefer, A.; Young, J.W.; Meffre, E.; Nussenzweig, M.C. Predominant autoantibody production by early human B cell precursors. Science 2003, 301, 1374–1377. [Google Scholar] [CrossRef]
- Zerrahn, J.; Held, W.; Raulet, D.H. The MHC reactivity of the T cell repertoire prior to positive and negative selection. Cell 1997, 88, 627–636. [Google Scholar] [CrossRef]
- Liston, A.; Lesage, S.; Gray, D.H.; O’Reilly, L.A.; Strasser, A.; Fahrer, A.M.; Boyd, R.L.; Wilson, J.; Baxter, A.G.; Gallo, E.M.; et al. Generalized resistance to thymic deletion in the NOD mouse; a polygenic trait characterized by defective induction of Bim. Immunity 2004, 21, 817–830. [Google Scholar] [CrossRef]
- Krammer, P.H. CD95(APO-1/Fas)-mediated apoptosis: Live and let die. Adv. Immunol. 1999, 71, 163–210. [Google Scholar] [CrossRef]
- Nagata, S.; Golstein, P. The Fas death factor. Science 1995, 267, 1449–1456. [Google Scholar] [CrossRef]
- Frank, S.A. Multistage progression. In Dynamics of Cancer: Incidence, Inheritance, and Evolution; Princeton University Press: Princeton, NJ, USA, 2007. [Google Scholar]
- Egle, A.; Harris, A.W.; Bouillet, P.; Cory, S. Bim is a suppressor of Myc-induced mouse B cell leukemia. Proc. Natl. Acad. Sci. USA 2004, 101, 6164–6169. [Google Scholar] [CrossRef] [PubMed]
- Holzelova, E.; Vonarbourg, C.; Stolzenberg, M.C.; Arkwright, P.D.; Selz, F.; Prieur, A.M.; Blanche, S.; Bartunkova, J.; Vilmer, E.; Fischer, A.; et al. Autoimmune lymphoproliferative syndrome with somatic Fas mutations. N. Engl. J. Med. 2004, 351, 1409–1418. [Google Scholar] [CrossRef]
- Gronbaek, K.; Straten, P.T.; Ralfkiaer, E.; Ahrenkiel, V.; Andersen, M.K.; Hansen, N.E.; Zeuthen, J.; Hou-Jensen, K.; Guldberg, P. Somatic Fas mutations in non-Hodgkin’s lymphoma: Association with extranodal disease and autoimmunity. Blood 1998, 92, 3018–3024. [Google Scholar] [CrossRef] [PubMed]
- Alriyami, M.; Marchand, L.; Li, Q.; Du, X.; Olivier, M.; Polychronakos, C. Clonal copy-number mosaicism in autoreactive T lymphocytes in diabetic NOD mice. Genome Res. 2019, 29, 1951–1961. [Google Scholar] [CrossRef] [PubMed]
- Araten, D.J.; Golde, D.W.; Zhang, R.H.; Thaler, H.T.; Gargiulo, L.; Notaro, R.; Luzzatto, L. A quantitative measurement of the human somatic mutation rate. Cancer Res. 2005, 65, 8111–8117. [Google Scholar] [CrossRef] [PubMed]
- Lichtenauer-Kaligis, E.G.; Thijssen, J.; den Dulk, H.; van de Putte, P.; Tasseron-de Jong, J.G.; Giphart-Gassler, M. Comparison of spontaneous hprt mutation spectra at the nucleotide sequence level in the endogenous hprt gene and five other genomic positions. Mutat. Res. 1996, 351, 147–155. [Google Scholar] [CrossRef]
- Werner, B.; Sottoriva, A. Variation of mutational burden in healthy human tissues suggests non-random strand segregation and allows measuring somatic mutation rates. PLoS Comput. Biol. 2018, 14, e1006233. [Google Scholar] [CrossRef]
- Abyzov, A.; Mariani, J.; Palejev, D.; Zhang, Y.; Haney, M.S.; Tomasini, L.; Ferrandino, A.F.; Rosenberg Belmaker, L.A.; Szekely, A.; Wilson, M.; et al. Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature 2012, 492, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Blokzijl, F.; de Ligt, J.; Jager, M.; Sasselli, V.; Roerink, S.; Sasaki, N.; Huch, M.; Boymans, S.; Kuijk, E.; Prins, P.; et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 2016, 538, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Lodato, M.A.; Rodin, R.E.; Bohrson, C.L.; Coulter, M.E.; Barton, A.R.; Kwon, M.; Sherman, M.A.; Vitzthum, C.M.; Luquette, L.J.; Yandava, C.N.; et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science 2018, 359, 555–559. [Google Scholar] [CrossRef]
- McConnell, M.J.; Lindberg, M.R.; Brennand, K.J.; Piper, J.C.; Voet, T.; Cowing-Zitron, C.; Shumilina, S.; Lasken, R.S.; Vermeesch, J.R.; Hall, I.M.; et al. Mosaic copy number variation in human neurons. Science 2013, 342, 632–637. [Google Scholar] [CrossRef]
- Hoang, M.L.; Kinde, I.; Tomasetti, C.; McMahon, K.W.; Rosenquist, T.A.; Grollman, A.P.; Kinzler, K.W.; Vogelstein, B.; Papadopoulos, N. Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing. Proc. Natl. Acad. Sci. USA 2016, 113, 9846–9851. [Google Scholar] [CrossRef]
- Martincorena, I.; Roshan, A.; Gerstung, M.; Ellis, P.; Van Loo, P.; McLaren, S.; Wedge, D.C.; Fullam, A.; Alexandrov, L.B.; Tubio, J.M.; et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 2015, 348, 880–886. [Google Scholar] [CrossRef] [PubMed]
- Busque, L.; Patel, J.P.; Figueroa, M.E.; Vasanthakumar, A.; Provost, S.; Hamilou, Z.; Mollica, L.; Li, J.; Viale, A.; Heguy, A.; et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat. Genet. 2012, 44, 1179–1181. [Google Scholar] [CrossRef]
- Genovese, G.; Kahler, A.K.; Handsaker, R.E.; Lindberg, J.; Rose, S.A.; Bakhoum, S.F.; Chambert, K.; Mick, E.; Neale, B.M.; Fromer, M.; et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 2014, 371, 2477–2487. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Lu, C.; Wang, J.; McLellan, M.D.; Johnson, K.J.; Wendl, M.C.; McMichael, J.F.; Schmidt, H.K.; Yellapantula, V.; Miller, C.A.; et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat. Med. 2014, 20, 1472–1478. [Google Scholar] [CrossRef]
- Young, A.L.; Challen, G.A.; Birmann, B.M.; Druley, T.E. Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults. Nat. Commun. 2016, 7, 12484. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, L.A.; Rasi, C.; Razzaghian, H.R.; Pakalapati, G.; Waite, L.; Thilbeault, K.S.; Ronowicz, A.; Wineinger, N.E.; Tiwari, H.K.; Boomsma, D.; et al. Age-related somatic structural changes in the nuclear genome of human blood cells. Am. J. Hum. Genet. 2012, 90, 217–228. [Google Scholar] [CrossRef]
- Jacobs, K.B.; Yeager, M.; Zhou, W.; Wacholder, S.; Wang, Z.; Rodriguez-Santiago, B.; Hutchinson, A.; Deng, X.; Liu, C.; Horner, M.J.; et al. Detectable clonal mosaicism and its relationship to aging and cancer. Nat. Genet. 2012, 44, 651–658. [Google Scholar] [CrossRef]
- Jaiswal, S.; Fontanillas, P.; Flannick, J.; Manning, A.; Grauman, P.V.; Mar, B.G.; Lindsley, R.C.; Mermel, C.H.; Burtt, N.; Chavez, A.; et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 2014, 371, 2488–2498. [Google Scholar] [CrossRef]
- Araten, D.J.; Krejci, O.; Ditata, K.; Wunderlich, M.; Sanders, K.J.; Zamechek, L.; Mulloy, J.C. The rate of spontaneous mutations in human myeloid cells. Mutat. Res. 2013, 749, 49–57. [Google Scholar] [CrossRef]
- Muschen, M.; Re, D.; Jungnickel, B.; Diehl, V.; Rajewsky, K.; Kuppers, R. Somatic mutation of the CD95 gene in human B cells as a side-effect of the germinal center reaction. J. Exp. Med. 2000, 192, 1833–1840. [Google Scholar] [CrossRef]
- Li, R.; Montpetit, A.; Rousseau, M.; Wu, S.Y.; Greenwood, C.M.; Spector, T.D.; Pollak, M.; Polychronakos, C.; Richards, J.B. Somatic point mutations occurring early in development: A monozygotic twin study. J. Med. Genet. 2014, 51, 28–34. [Google Scholar] [CrossRef]
- Budowle, B. Molecular genetic investigative leads to differentiate monozygotic twins. Investig. Genet. 2014, 5, 11. [Google Scholar] [CrossRef]
- Araten, D.J.; Nafa, K.; Pakdeesuwan, K.; Luzzatto, L. Clonal populations of hematopoietic cells with paroxysmal nocturnal hemoglobinuria genotype and phenotype are present in normal individuals. Proc. Natl. Acad. Sci. USA 1999, 96, 5209–5214. [Google Scholar] [CrossRef] [PubMed]
- Bernatsky, S.; Boivin, J.F.; Joseph, L.; Rajan, R.; Zoma, A.; Manzi, S.; Ginzler, E.; Urowitz, M.; Gladman, D.; Fortin, P.R.; et al. An international cohort study of cancer in systemic lupus erythematosus. Arthritis Rheum. 2005, 52, 1481–1490. [Google Scholar] [CrossRef] [PubMed]
- Cellier, C.; Delabesse, E.; Helmer, C.; Patey, N.; Matuchansky, C.; Jabri, B.; Macintyre, E.; Cerf-Bensussan, N.; Brousse, N. Refractory sprue, coeliac disease, and enteropathy-associated T-cell lymphoma. French Coeliac Disease Study Group. Lancet 2000, 356, 203–208. [Google Scholar] [CrossRef]
- Zintzaras, E.; Voulgarelis, M.; Moutsopoulos, H.M. The risk of lymphoma development in autoimmune diseases: A meta-analysis. Arch. Intern. Med. 2005, 165, 2337–2344. [Google Scholar] [CrossRef]
- Smedby, K.E.; Hjalgrim, H.; Askling, J.; Chang, E.T.; Gregersen, H.; Porwit-MacDonald, A.; Sundstrom, C.; Akerman, M.; Melbye, M.; Glimelius, B.; et al. Autoimmune and chronic inflammatory disorders and risk of non-Hodgkin lymphoma by subtype. J. Natl. Cancer Inst. 2006, 98, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Ekstrom Smedby, K.; Vajdic, C.M.; Falster, M.; Engels, E.A.; Martinez-Maza, O.; Turner, J.; Hjalgrim, H.; Vineis, P.; Seniori Costantini, A.; Bracci, P.M.; et al. Autoimmune disorders and risk of non-Hodgkin lymphoma subtypes: A pooled analysis within the InterLymph Consortium. Blood 2008, 111, 4029–4038. [Google Scholar] [CrossRef]
- Anderson, L.A.; Gadalla, S.; Morton, L.M.; Landgren, O.; Pfeiffer, R.; Warren, J.L.; Berndt, S.I.; Ricker, W.; Parsons, R.; Engels, E.A. Population-based study of autoimmune conditions and the risk of specific lymphoid malignancies. Int. J. Cancer 2009, 125, 398–405. [Google Scholar] [CrossRef] [PubMed]
- Franklin, J.; Lunt, M.; Bunn, D.; Symmons, D.; Silman, A. Incidence of lymphoma in a large primary care derived cohort of cases of inflammatory polyarthritis. Ann. Rheum. Dis. 2006, 65, 617–622. [Google Scholar] [CrossRef]
- Mellemkjaer, L.; Pfeiffer, R.M.; Engels, E.A.; Gridley, G.; Wheeler, W.; Hemminki, K.; Olsen, J.H.; Dreyer, L.; Linet, M.S.; Goldin, L.R.; et al. Autoimmune disease in individuals and close family members and susceptibility to non-Hodgkin’s lymphoma. Arthritis Rheum. 2008, 58, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Landgren, O.; Engels, E.A.; Pfeiffer, R.M.; Gridley, G.; Mellemkjaer, L.; Olsen, J.H.; Kerstann, K.F.; Wheeler, W.; Hemminki, K.; Linet, M.S.; et al. Autoimmunity and susceptibility to Hodgkin lymphoma: A population-based case-control study in Scandinavia. J. Natl. Cancer Inst. 2006, 98, 1321–1330. [Google Scholar] [CrossRef]
- Bernatsky, S.; Ramsey-Goldman, R.; Isenberg, D.; Rahman, A.; Dooley, M.A.; Sibley, J.; Boivin, J.F.; Joseph, L.; Armitage, J.; Zoma, A.; et al. Hodgkin’s lymphoma in systemic lupus erythematosus. Rheumatology 2007, 46, 830–832. [Google Scholar] [CrossRef]
- Kristinsson, S.Y.; Landgren, O.; Sjoberg, J.; Turesson, I.; Bjorkholm, M.; Goldin, L.R. Autoimmunity and risk for Hodgkin’s lymphoma by subtype. Haematologica 2009, 94, 1468–1469. [Google Scholar] [CrossRef][Green Version]
- Hansen, A.; Lipsky, P.E.; Dorner, T. B-cell lymphoproliferation in chronic inflammatory rheumatic diseases. Nat. Clin. Pract. Rheumatol. 2007, 3, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Varoczy, L.; Gergely, L.; Zeher, M.; Szegedi, G.; Illes, A. Malignant lymphoma-associated autoimmune diseases—A descriptive epidemiological study. Rheumatol. Int. 2002, 22, 233–237. [Google Scholar] [CrossRef] [PubMed]
- Dameshek, W.; Schwartz, R.S. Leukemia and auto-immunization- some possible relationships. Blood 1959, 14, 1151–1158. [Google Scholar] [CrossRef]
- Straus, S.E.; Jaffe, E.S.; Puck, J.M.; Dale, J.K.; Elkon, K.B.; Rosen-Wolff, A.; Peters, A.M.; Sneller, M.C.; Hallahan, C.W.; Wang, J.; et al. The development of lymphomas in families with autoimmune lymphoproliferative syndrome with germline Fas mutations and defective lymphocyte apoptosis. Blood 2001, 98, 194–200. [Google Scholar] [CrossRef]
- Allegretta, M.; Nicklas, J.A.; Sriram, S.; Albertini, R.J. T cells responsive to myelin basic protein in patients with multiple sclerosis. Science 1990, 247, 718–721. [Google Scholar] [CrossRef] [PubMed]
- Sriram, S. Longitudinal study of frequency of HPRT mutant T cells in patients with multiple sclerosis. Neurology 1994, 44, 311–315. [Google Scholar] [CrossRef]
- Albertini, R.J.; Castle, K.L.; Borcherding, W.R. T-cell cloning to detect the mutant 6-thioguanine-resistant lymphocytes present in human peripheral blood. Proc. Natl. Acad. Sci. USA 1982, 79, 6617–6621. [Google Scholar] [CrossRef] [PubMed]
- Kemppinen, A.K.; Baker, A.; Liao, W.; Fiddes, B.; Jones, J.; Compston, A.; Ban, M.; Sawcer, S. Exome sequencing in single cells from the cerebrospinal fluid in multiple sclerosis. Mult. Scler. 2014, 20, 1564–1568. [Google Scholar] [CrossRef] [PubMed]
- Valori, M.; Jansson, L.; Kiviharju, A.; Ellonen, P.; Rajala, H.; Awad, S.A.; Mustjoki, S.; Tienari, P.J. A novel class of somatic mutations in blood detected preferentially in CD8+ cells. Clin. Immunol. 2017, 175, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Savola, P.; Kelkka, T.; Rajala, H.L.; Kuuliala, A.; Kuuliala, K.; Eldfors, S.; Ellonen, P.; Lagstrom, S.; Lepisto, M.; Hannunen, T.; et al. Somatic mutations in clonally expanded cytotoxic T lymphocytes in patients with newly diagnosed rheumatoid arthritis. Nat. Commun. 2017, 8, 15869. [Google Scholar] [CrossRef]
- Polychronakos, C.; Li, Q. Understanding type 1 diabetes through genetics: Advances and prospects. Nat. Rev. Genet. 2011, 12, 781–792. [Google Scholar] [CrossRef]
- Lempainen, J.; Ilonen, J. Influence of type 1 diabetes genes on disease progression: Similarities and differences between countries. Curr. Diab. Rep. 2012, 12, 447–455. [Google Scholar] [CrossRef]
- Surcel, H.M.; Ilonen, J.; Kaar, M.L.; Hyoty, H.; Leinikki, P. Infection by multiple viruses and lymphocyte abnormalities at the diagnosis of diabetes. Acta Paediatr. Scand. 1988, 77, 471–474. [Google Scholar] [CrossRef] [PubMed]
- Redondo, M.J.; Jeffrey, J.; Fain, P.R.; Eisenbarth, G.S.; Orban, T. Concordance for islet autoimmunity among monozygotic twins. N. Engl. J. Med. 2008, 359, 2849–2850. [Google Scholar] [CrossRef]
- Makino, S.; Kunimoto, K.; Muraoka, Y.; Mizushima, Y.; Katagiri, K.; Tochino, Y. Breeding of a non-obese, diabetic strain of mice. Jikken Dobutsu 1980, 29, 1–13. [Google Scholar] [CrossRef]
- Cohen, S.B.; Emery, P.; Greenwald, M.W.; Dougados, M.; Furie, R.A.; Genovese, M.C.; Keystone, E.C.; Loveless, J.E.; Burmester, G.R.; Cravets, M.W.; et al. Rituximab for rheumatoid arthritis refractory to anti-tumor necrosis factor therapy: Results of a multicenter, randomized, double-blind, placebo-controlled, phase III trial evaluating primary efficacy and safety at twenty-four weeks. Arthritis Rheum. 2006, 54, 2793–2806. [Google Scholar] [CrossRef]
- Emery, P.; Fleischmann, R.; Filipowicz-Sosnowska, A.; Schechtman, J.; Szczepanski, L.; Kavanaugh, A.; Racewicz, A.J.; van Vollenhoven, R.F.; Li, N.F.; Agarwal, S.; et al. The efficacy and safety of rituximab in patients with active rheumatoid arthritis despite methotrexate treatment: Results of a phase IIB randomized, double-blind, placebo-controlled, dose-ranging trial. Arthritis Rheum. 2006, 54, 1390–1400. [Google Scholar] [CrossRef] [PubMed]
- Peterfy, C.; Emery, P.; Tak, P.P.; Ostergaard, M.; DiCarlo, J.; Otsa, K.; Navarro Sarabia, F.; Pavelka, K.; Bagnard, M.A.; Gylvin, L.H.; et al. MRI assessment of suppression of structural damage in patients with rheumatoid arthritis receiving rituximab: Results from the randomised, placebo-controlled, double-blind RA-SCORE study. Ann. Rheum. Dis. 2016, 75, 170–177. [Google Scholar] [CrossRef] [PubMed]

| Technical Method | Data | Autoimmune Disease | Description | References |
|---|---|---|---|---|
| Sequencing/candidate gene approach | Single-nucleotide variant | Autoimmune lymphoproliferative syndrome | DNA extracted from phytohemagglutinin-activated lymphocytes or purified double-negative T cells was amplified with oligonucleotides spanning the nine FAS exons and sequenced | [34] |
| Hypoxanthine guanine phosphoribosyltransferase (HPRT) assay | Mutant frequency values | Multiple sclerosis | Identifies the resistance of cultured T cells to 6-thioguanine, caused by somatic mutations inhibition of HPRT gene or other related pathways | [73,74,75] |
| Exome-sequencing | Single-nucleotide variant and their frequency values | Multiple sclerosis | Exome-sequence DNA of CD4+ lymphocytes isolated from patients’ cerebrospinal fluid along with sequencing DNA from peripheral blood to serve as germline reference | [76] |
| Next-generation HiSeq-DNA sequencing using a custom designed gene panel consisting of 986 genes related to immunity and cancer | Single-nucleotide variant | Multiple sclerosis, Myasthenia gravis, and Narcolepsy | Somatic mutations were called using next-generation HiSeq-DNA sequencing targeting 986 immune-related genes and by comparing the sequence data of immune cell subpopulations to other subpopulations of the same patient | [77] |
| Custom deep-sequencing panel of immune-related genes and exome and deep amplicon sequencing | Single-nucleotide variant | Rheumatoid arthritis | Custom deep-sequencing panel of 986 immune-related genes of CD8+ T cells of untreated newly diagnosed patients with RA | [78] |
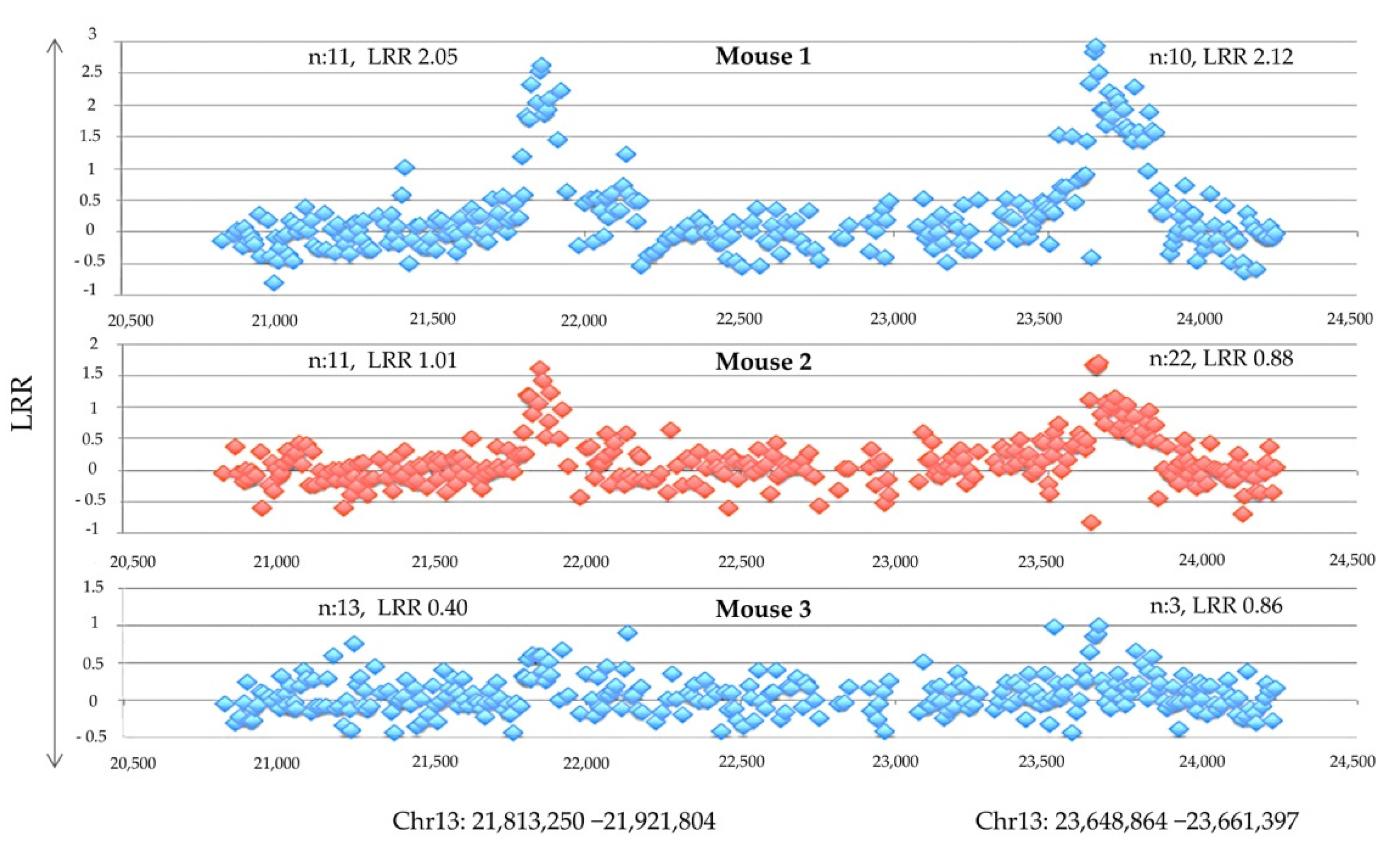
| Comparative genomic hybridization (CGH) assay and validation via multiplex ligation-dependent probe amplification (MLPA) | Copy-number aberrations | Type 1 diabetes | CGH on DNA from memory CD4+ T cells obtained from the pancreatic lymph nodes of diabetic NOD mice and using DNA from a tail-clip sample to serve as germline reference | [36] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alriyami, M.; Polychronakos, C. Somatic Mutations and Autoimmunity. Cells 2021, 10, 2056. https://doi.org/10.3390/cells10082056
Alriyami M, Polychronakos C. Somatic Mutations and Autoimmunity. Cells. 2021; 10(8):2056. https://doi.org/10.3390/cells10082056
Chicago/Turabian StyleAlriyami, Maha, and Constantin Polychronakos. 2021. "Somatic Mutations and Autoimmunity" Cells 10, no. 8: 2056. https://doi.org/10.3390/cells10082056
APA StyleAlriyami, M., & Polychronakos, C. (2021). Somatic Mutations and Autoimmunity. Cells, 10(8), 2056. https://doi.org/10.3390/cells10082056






