Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes
Abstract
1. Introduction
2. Materials and Methods
2.1. Anterior Cruciate Ligament (ACL) Injury Model
2.2. Histological Assessment of Disease Severity
2.3. Immunohistochemistry (IHC)
2.4. Single-Cell RNA Sequencing (scRNA-seq)
2.5. scRNA-seq Data Analysis of Chondrocytes from Uninjured Joints
2.6. Analysis of Human Chondrocyte scRNA-seq Data
2.7. Comparison of Chondrocytes from Uninjured and Injured Joints
2.8. Pseudotime Trajectory Finding
2.9. Ontology Enrichment Analysis
3. Results
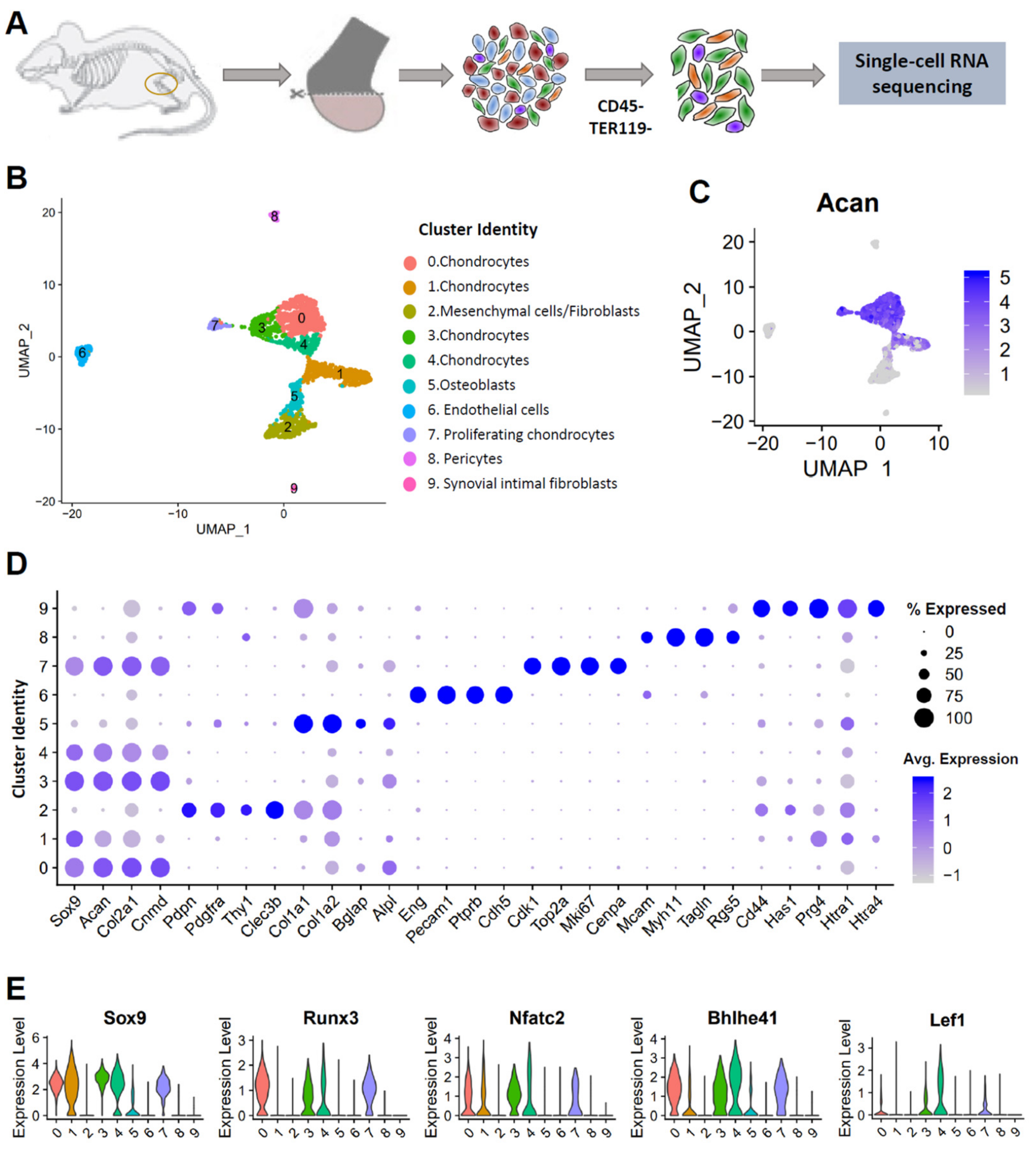
3.1. Single-Cell Profiling Reveals Cellular Heterogeneity in Healthy Murine Knee Joints
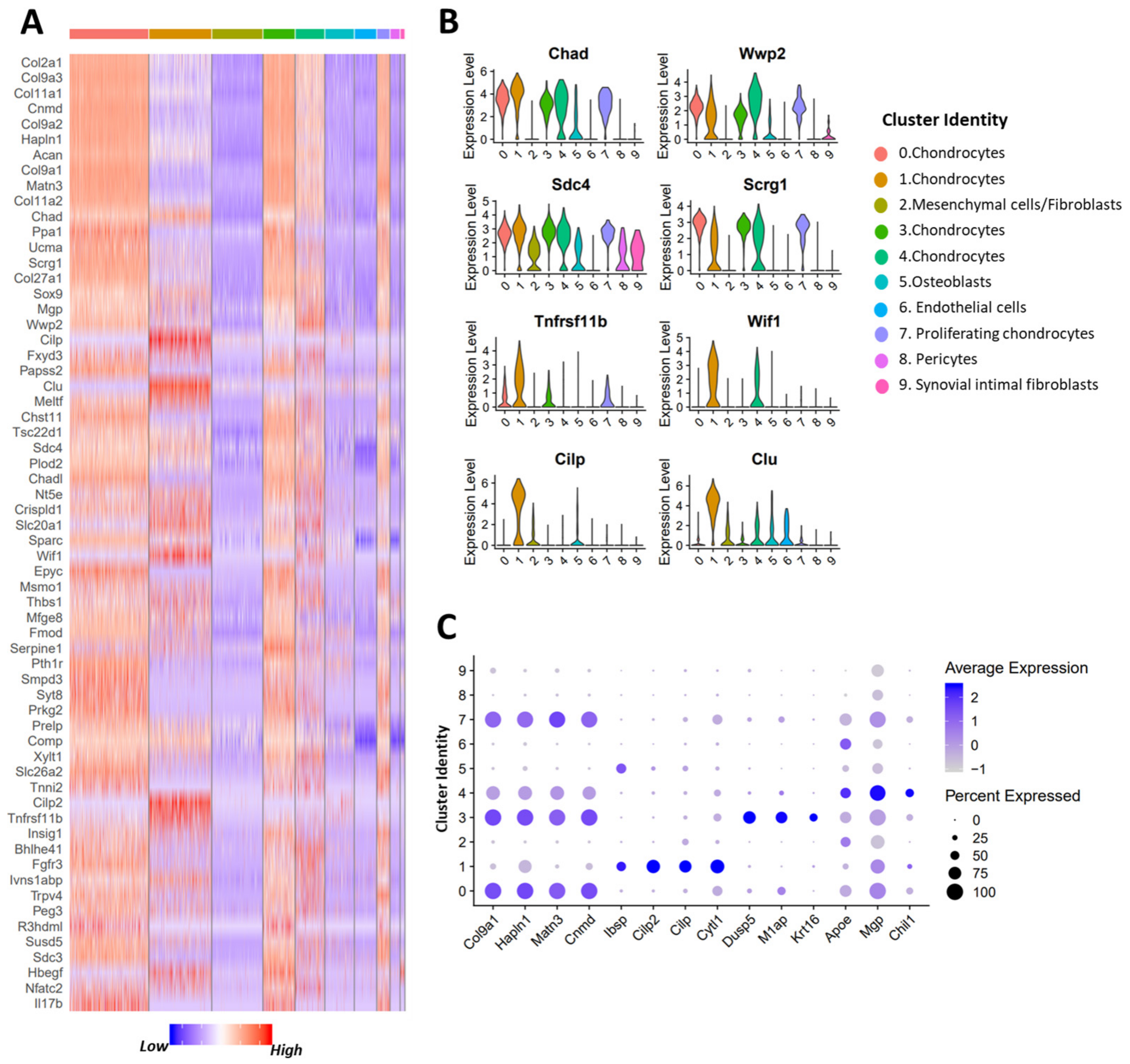
3.2. Identification of Potential OA Targets Enriched in Chondrocytes
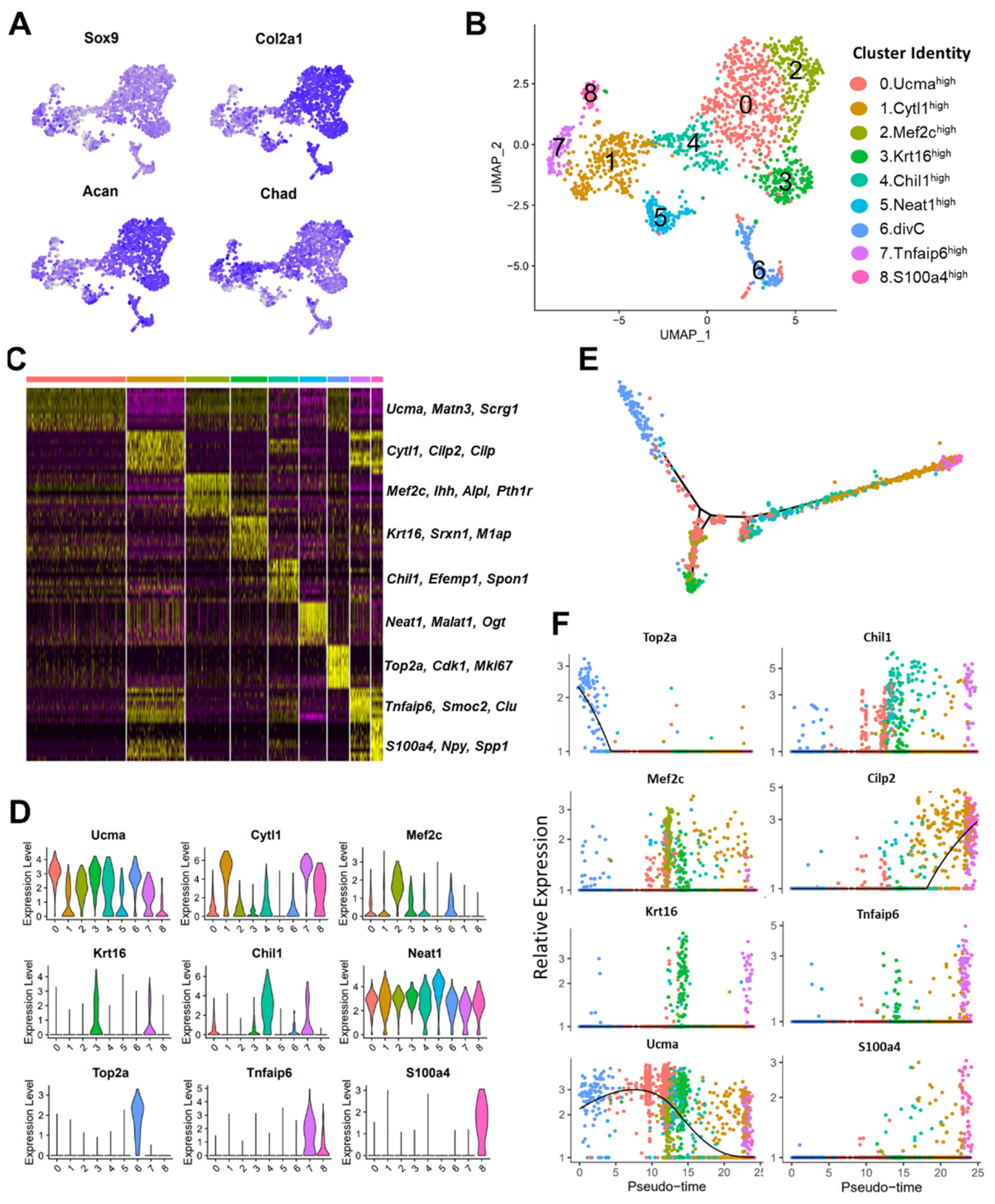
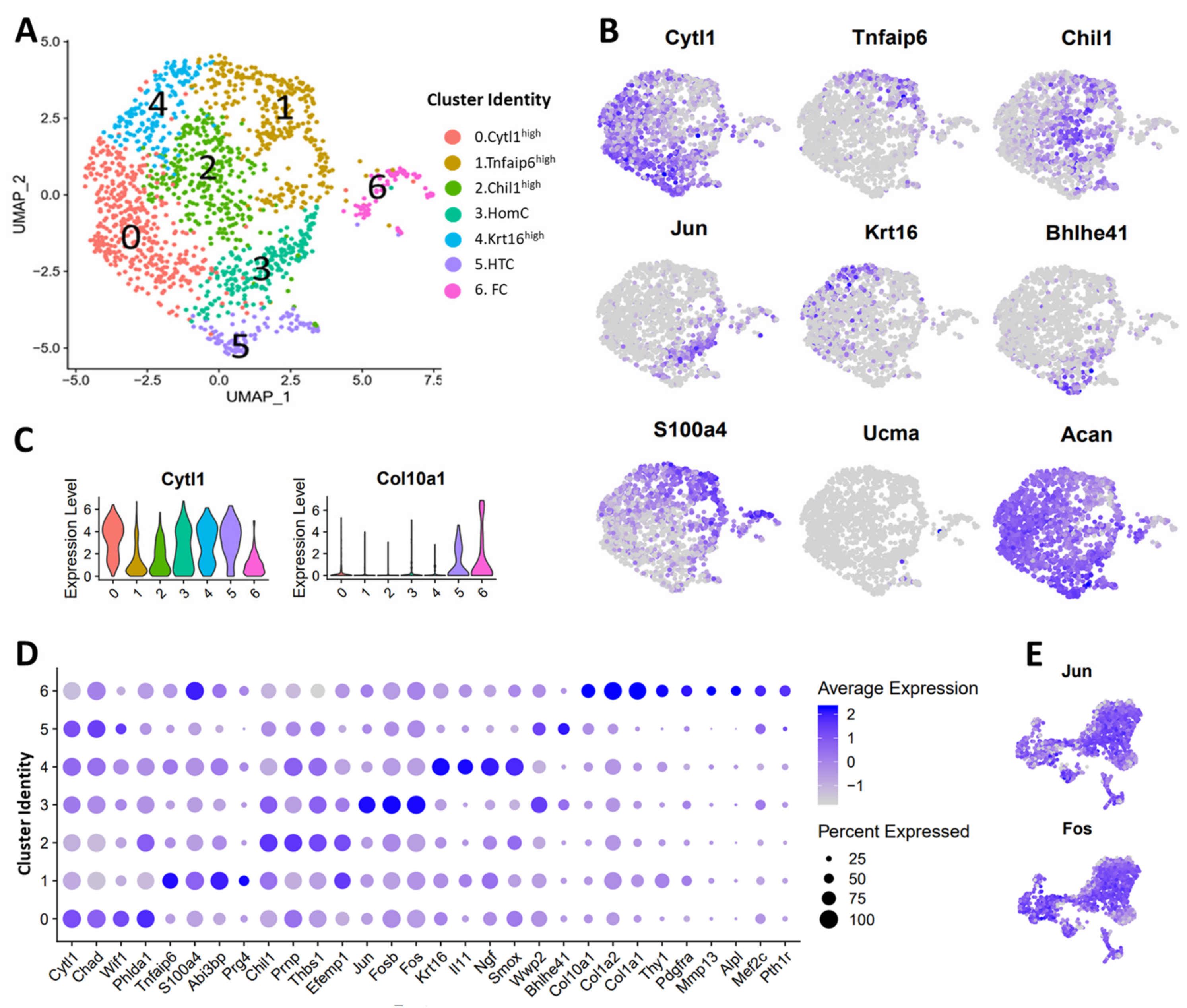
3.3. scRNA-seq Analysis Identified Nine Chondrocyte Subtypes in Mouse Knee Joints
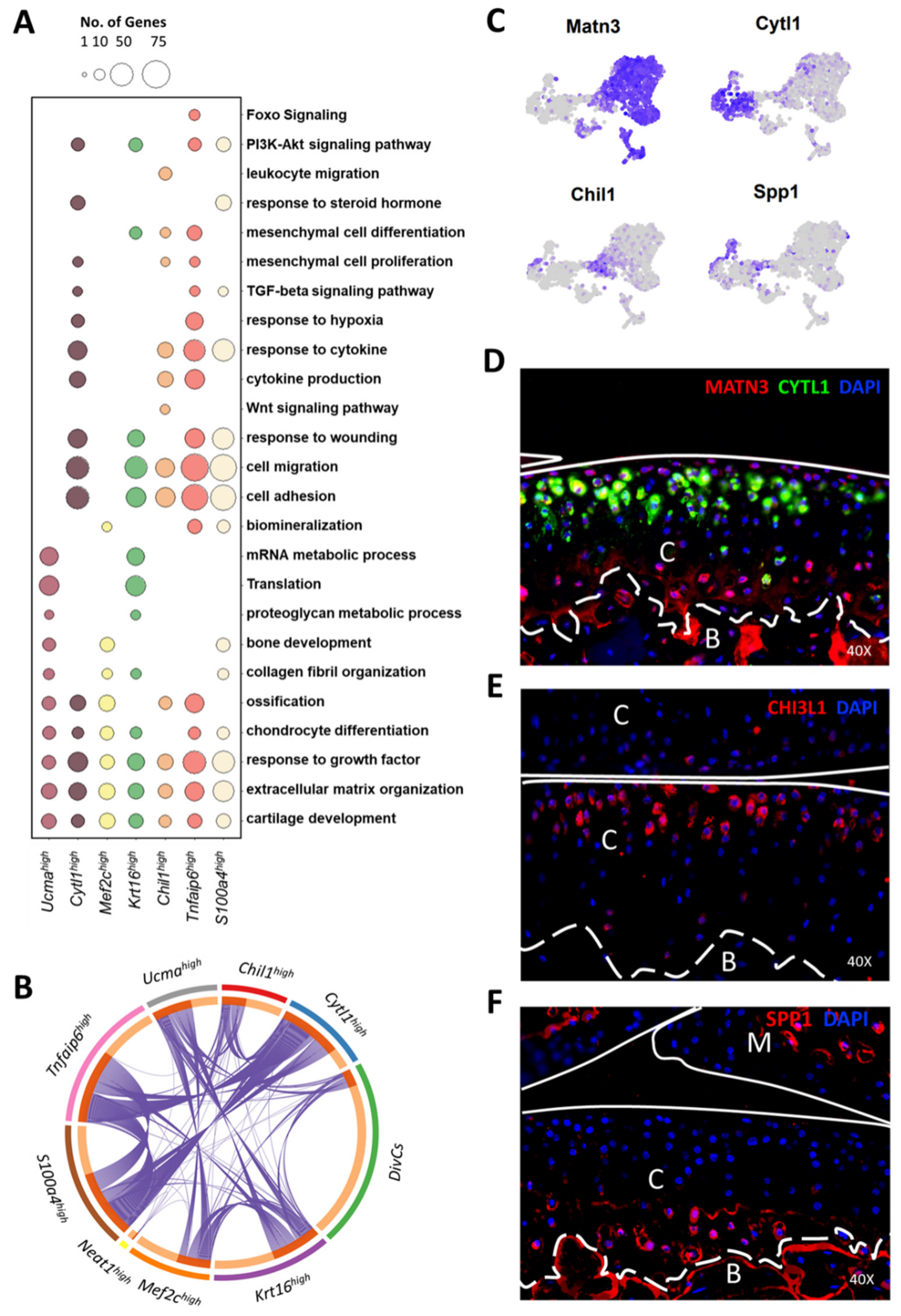
3.4. Molecular and Functional-Level Characterization of Chondrocyte Subpopulations
3.5. Comparative Transcriptomic Analysis Identified Similarities and Differences between Mouse and Human Chondrocyte Subtypes
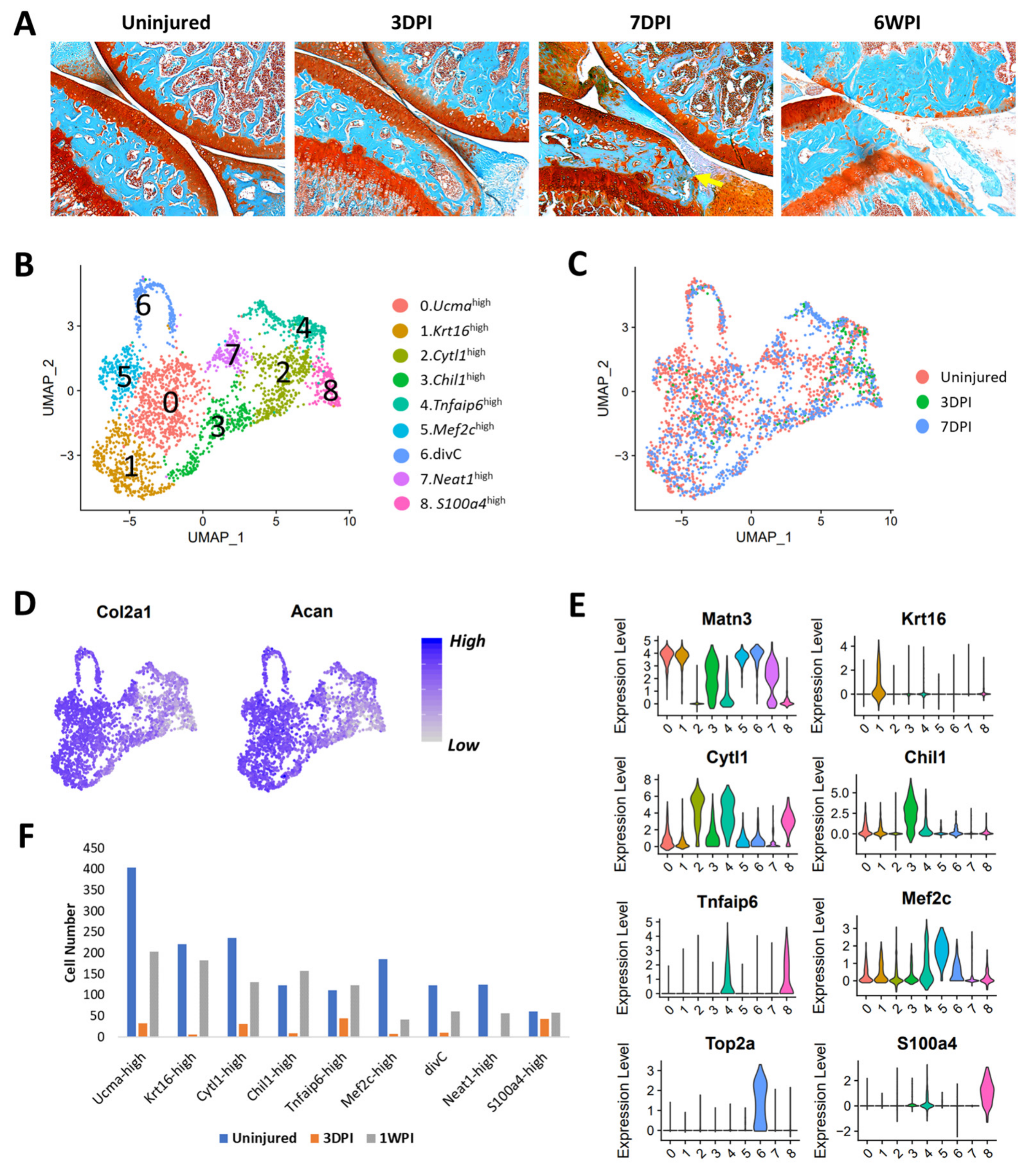
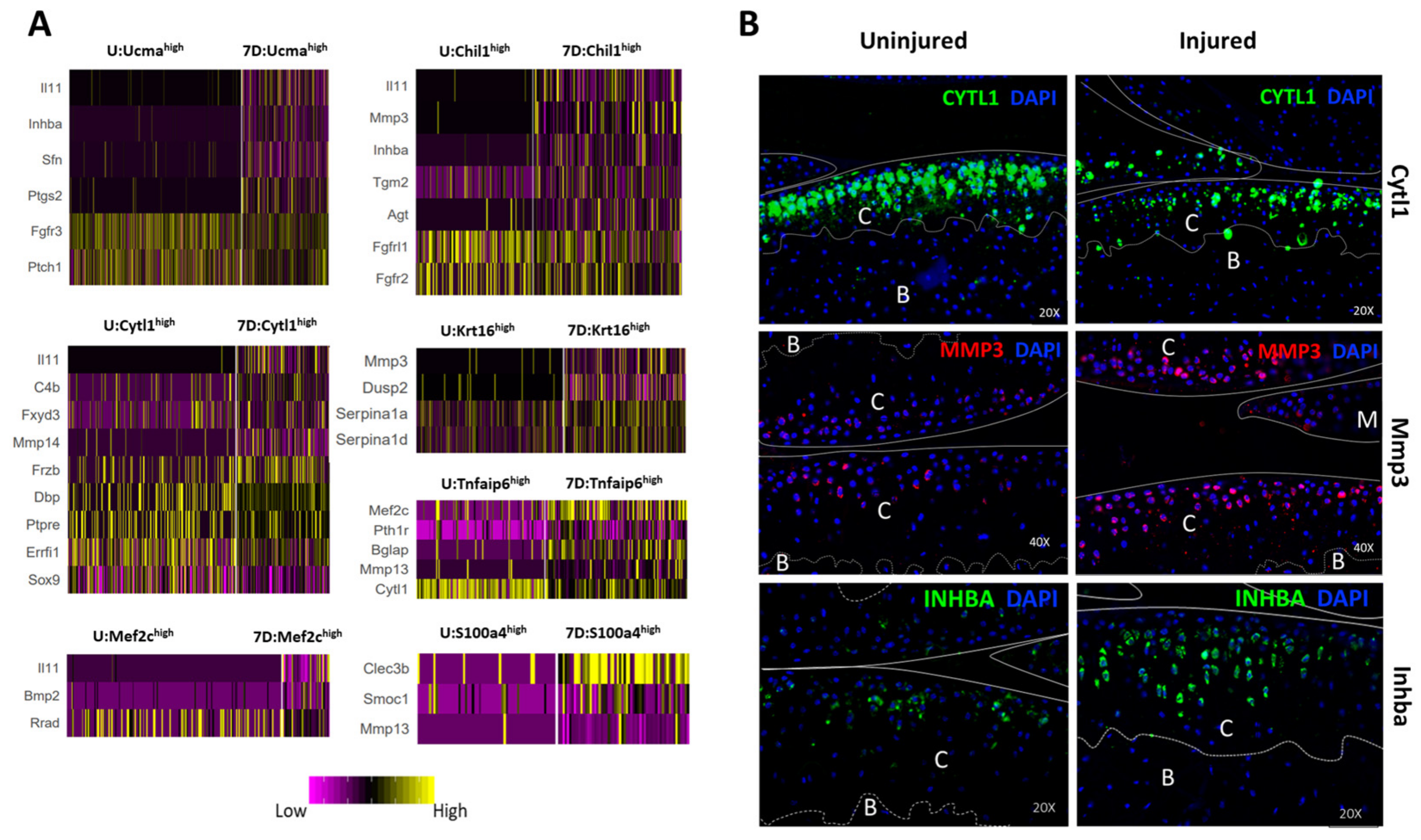
3.6. Identification of Injury-Induced Early Molecular Changes in the Articular Chondrocytes
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, D.; Shen, J.; Zhao, W.; Wang, T.; Han, L.; Hamilton, J.L.; Im, H.J. Osteoarthritis: Toward a comprehensive understanding of pathological mechanism. Bone Res. 2017, 5, 16044. [Google Scholar] [CrossRef] [PubMed]
- Blixen, C.E.; Kippes, C. Depression, social support, and quality of life in older adults with osteoarthritis. Image J. Nurs. Sch. 1999, 31, 221–226. [Google Scholar] [CrossRef] [PubMed]
- Blalock, D.; Miller, A.; Tilley, M.; Wang, J. Joint instability and osteoarthritis. Clin. Med. Insights Arthritis Musculoskelet. Disords 2015, 8, 15–23. [Google Scholar] [CrossRef]
- Steinert, A.F.; Ghivizzani, S.C.; Rethwilm, A.; Tuan, R.S.; Evans, C.H.; Noth, U. Major biological obstacles for persistent cell-based regeneration of articular cartilage. Arthritis Res. Ther. 2007, 9, 213. [Google Scholar] [CrossRef] [PubMed]
- Sophia Fox, A.J.; Bedi, A.; Rodeo, S.A. The basic science of articular cartilage: Structure, composition, and function. Sports Health 2009, 1, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Ji, Q.; Zheng, Y.; Zhang, G.; Hu, Y.; Fan, X.; Hou, Y.; Wen, L.; Li, L.; Xu, Y.; Wang, Y.; et al. Single-cell RNA-seq analysis reveals the progression of human osteoarthritis. Ann. Rheum. Dis. 2019, 78, 100–110. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.H.; Jain, V.; Gibson, J.; Attarian, D.E.; Haraden, C.A.; Yohn, C.B.; Laberge, R.M.; Gregory, S.; Kraus, V.B. Synovial cell cross-talk with cartilage plays a major role in the pathogenesis of osteoarthritis. Sci. Rep. 2020, 10, 10868. [Google Scholar] [CrossRef] [PubMed]
- Ramos, Y.F.; den Hollander, W.; Bovee, J.V.; Bomer, N.; van der Breggen, R.; Lakenberg, N.; Keurentjes, J.C.; Goeman, J.J.; Slagboom, P.E.; Nelissen, R.G.; et al. Genes involved in the osteoarthritis process identified through genome wide expression analysis in articular cartilage; the RAAK study. PLoS ONE 2014, 9, e103056. [Google Scholar] [CrossRef]
- He, A.; Ning, Y.; Wen, Y.; Cai, Y.; Xu, K.; Cai, Y.; Han, J.; Liu, L.; Du, Y.; Liang, X.; et al. Use of integrative epigenetic and mRNA expression analyses to identify significantly changed genes and functional pathways in osteoarthritic cartilage. Bone Joint Res. 2018, 7, 343–350. [Google Scholar] [CrossRef]
- Little, C.B.; Hunter, D.J. Post-traumatic osteoarthritis: From mouse models to clinical trials. Nat. Rev. Rheumatol. 2013, 9, 485–497. [Google Scholar] [CrossRef]
- Loeser, R.F.; Olex, A.L.; McNulty, M.A.; Carlson, C.S.; Callahan, M.; Ferguson, C.; Fetrow, J.S. Disease progression and phasic changes in gene expression in a mouse model of osteoarthritis. PLoS ONE 2013, 8, e54633. [Google Scholar] [CrossRef]
- JChang, C.; Sebastian, A.; Murugesh, D.K.; Hatsell, S.; Economides, A.N.; Christiansen, B.A.; Loots, G.G. Global molecular changes in a tibial compression induced ACL rupture model of post-traumatic osteoarthritis. J. Orthop. Res. 2017, 35, 474–485. [Google Scholar] [CrossRef]
- Sebastian, A.; Chang, J.C.; Mendez, M.E.; Murugesh, D.K.; Hatsell, S.; Economides, A.N.; Christiansen, B.A.; Loots, G.G. Comparative Transcriptomics Identifies Novel Genes and Pathways Involved in Post-Traumatic Osteoarthritis Development and Progression. Int. J. Mol. Sci. 2018, 19, 2657. [Google Scholar] [CrossRef]
- Kung, L.H.W.; Ravi, V.; Rowley, L.; Bell, K.M.; Little, C.B.; Bateman, J.F. Comprehensive Expression Analysis of microRNAs and mRNAs in Synovial Tissue from a Mouse Model of Early Post-Traumatic Osteoarthritis. Sci. Rep. 2017, 7, 17701. [Google Scholar] [CrossRef]
- Zhang, R.; Fang, H.; Chen, Y.; Shen, J.; Lu, H.; Zeng, C.; Ren, J.; Zeng, H.; Li, Z.; Chen, S.; et al. Gene expression analyses of subchondral bone in early experimental osteoarthritis by microarray. PLoS ONE 2012, 7, e32356. [Google Scholar] [CrossRef]
- Tang, F.; Lao, K.; Surani, M.A. Development and applications of single-cell transcriptome analysis. Nat. Methods 2011, 8, S6–S11. [Google Scholar] [CrossRef]
- Jonason, J.H.; Hoak, D.; O′Keefe, R.J. Primary murine growth plate and articular chondrocyte isolation and cell culture. Methods Mol. Biol. 2015, 1226, 11–18. [Google Scholar]
- Christiansen, B.A.; Anderson, M.J.; Lee, C.A.; Williams, J.C.; Yik, J.H.; Haudenschild, D.R. Musculoskeletal changes following non-invasive knee injury using a novel mouse model of post-traumatic osteoarthritis. Osteoarthr. Cartil. 2012, 20, 773–782. [Google Scholar] [CrossRef]
- Chang, J.C.; Christiansen, B.A.; Murugesh, D.K.; Sebastian, A.; Hum, N.R.; Collette, N.M.; Hatsell, S.; Economides, A.N.; Blanchette, C.D.; Loots, G.G. SOST/Sclerostin Improves Posttraumatic Osteoarthritis and Inhibits MMP2/3 Expression After Injury. J. Bone Miner Res. 2018, 33, 1105–1113. [Google Scholar] [CrossRef]
- Butler, A.; Hoffman, P.; Smibert, P.; Papalexi, E.; Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018, 36, 411–420. [Google Scholar] [CrossRef]
- Sebastian, A.; Hum, N.R.; Martin, K.A.; Gilmore, S.F.; Peran, I.; Byers, S.W.; Wheeler, E.K.; Coleman, M.A.; Loots, G.G. Single-Cell Transcriptomic Analysis of Tumor-Derived Fibroblasts and Normal Tissue-Resident Fibroblasts Reveals Fibroblast Heterogeneity in Breast Cancer. Cancers 2020, 12, 1307. [Google Scholar] [CrossRef]
- Trapnell, C.; Cacchiarelli, D.; Grimsby, J.; Pokharel, P.; Li, S.; Morse, M.; Lennon, N.J.; Livak, K.J.; Mikkelsen, T.S.; Rinn, J.L. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 2014, 32, 381–386. [Google Scholar] [CrossRef]
- Chen, J.; Bardes, E.E.; Aronow, B.J.; Jegga, A.G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 2009, 37, W305–W311. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Haseeb, A.; Kc, R.; Angelozzi, M.; de Charleroy, C.; Rux, D.; Tower, R.J.; Yao, L.; da Silva, R.P.; Pacifici, M.; Qin, L.; et al. SOX9 keeps growth plates and articular cartilage healthy by inhibiting chondrocyte dedifferentiation/osteoblastic redifferentiation. Proc. Natl. Acad. Sci. USA 2021, 118, e2019152118. [Google Scholar] [CrossRef]
- Scott, R.E.; Ghule, P.N.; Stein, J.L.; Stein, G.S. Cell cycle gene expression networks discovered using systems biology: Significance in carcinogenesis. J. Cell Physiol. 2015, 230, 2533–2542. [Google Scholar] [CrossRef]
- Zhong, L.; Yao, L.; Tower, R.J.; Wei, Y.; Miao, Z.; Park, J.; Shrestha, R.; Wang, L.; Yu, W.; Holdreith, N.; et al. Single cell transcriptomics identifies a unique adipose lineage cell population that regulates bone marrow environment. Elife 2020, 9, e54695. [Google Scholar] [CrossRef]
- Muhl, L.; Genove, G.; Leptidis, S.; Liu, J.; He, L.; Mocci, G.; Sun, Y.; Gustafsson, S.; Buyandelger, B.; Chivukula, I.V.; et al. Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination. Nat. Commun. 2020, 11, 3953. [Google Scholar] [CrossRef] [PubMed]
- Tikhonova, A.N.; Dolgalev, I.; Hu, H.; Sivaraj, K.K.; Hoxha, E.; Cuesta-Dominguez, A.; Pinho, S.; Akhmetzyanova, I.; Gao, J.; Witkowski, M.; et al. The bone marrow microenvironment at single-cell resolution. Nature 2019, 569, 222–228. [Google Scholar] [CrossRef]
- Thomson, B.R.; Carota, I.A.; Souma, T.; Soman, S.; Vestweber, D.; Quaggin, S.E. Targeting the vascular-specific phosphatase PTPRB protects against retinal ganglion cell loss in a pre-clinical model of glaucoma. Elife 2019, 8, e48474. [Google Scholar] [CrossRef]
- Esteves, C.L.; Donadeu, F.X. Pericytes and their potential in regenerative medicine across species. Cytom. A 2018, 93, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Shrestha, S.; Yen, Y.H.; Scott, M.A.; Soo, C.; Ting, K.; Peault, B.; Dry, S.M.; James, A.W. The pericyte antigen RGS5 in perivascular soft tissue tumors. Hum. Pathol. 2016, 47, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Hess, D.L.; Kelly-Goss, M.R.; Cherepanova, O.A.; Nguyen, A.T.; Baylis, R.A.; Tkachenko, S.; Annex, B.H.; Peirce, S.M.; Owens, G.K. Perivascular cell-specific knockout of the stem cell pluripotency gene Oct4 inhibits angiogenesis. Nat. Commun. 2019, 10, 967. [Google Scholar] [CrossRef] [PubMed]
- Gault, C.R.; Obeid, L.M.; Hannun, Y.A. An overview of sphingolipid metabolism: From synthesis to breakdown. Adv. Exp. Med. Biol. 2010, 688, 1–23. [Google Scholar]
- Luo, M.; Chen, J.; Li, S.; Sun, H.; Zhang, Z.; Fu, Q.; Li, J.; Wang, J.; Hughes, C.E.; Caterson, B.; et al. Changes in the metabolism of chondroitin sulfate glycosaminoglycans in articular cartilage from patients with Kashin-Beck disease. Osteoarthr. Cartil. 2014, 22, 986–995. [Google Scholar] [CrossRef]
- Tasdelen, I.; Berger, R.; Kalkhoven, E. PPARgamma regulates expression of carbohydrate sulfotransferase 11 (CHST11/C4ST1), a regulator of LPL cell surface binding. PLoS ONE 2013, 8, e64284. [Google Scholar] [CrossRef]
- Soul, J.; Barter, M.J.; Little, C.B.; Young, D.A. OATargets: A knowledge base of genes associated with osteoarthritis joint damage in animals. Ann. Rheum. Dis. 2020. [Google Scholar] [CrossRef]
- Li, J.; Dong, S. The Signaling Pathways Involved in Chondrocyte Differentiation and Hypertrophic Differentiation. Stem. Cells Int. 2016, 2016, 2470351. [Google Scholar] [CrossRef]
- Bian, Q.; Cheng, Y.H.; Wilson, J.P.; Su, E.Y.; Kim, D.W.; Wang, H.; Yoo, S.; Blackshaw, S.; Cahan, P. A single cell transcriptional atlas of early synovial joint development. Development 2020, 147, dev185777. [Google Scholar] [CrossRef]
- Kozhemyakina, E.; Lassar, A.B.; Zelzer, E. A pathway to bone: Signaling molecules and transcription factors involved in chondrocyte development and maturation. Development 2015, 142, 817–831. [Google Scholar] [CrossRef]
- Myllarniemi, M.; Tikkanen, J.; Hulmi, J.J.; Pasternack, A.; Sutinen, E.; Ronty, M.; Lepparanta, O.; Ma, H.; Ritvos, O.; Koli, K. Upregulation of activin-B and follistatin in pulmonary fibrosis—A translational study using human biopsies and a specific inhibitor in mouse fibrosis models. BMC Pulm. Med. 2014, 14, 170. [Google Scholar] [CrossRef]
- Song, Z.; Chen, W.; Athavale, D.; Ge, X.; Desert, R.; Das, S.; Han, H.; Nieto, N. Osteopontin takes center stage in chronic liver disease. Hepatology 2020. [Google Scholar] [CrossRef]
- Kishimoto, H.; Akagi, M.; Zushi, S.; Teramura, T.; Onodera, Y.; Sawamura, T.; Hamanishi, C. Induction of hypertrophic chondrocyte-like phenotypes by oxidized LDL in cultured bovine articular chondrocytes through increase in oxidative stress. Osteoarthr. Cartil. 2010, 18, 1284–1290. [Google Scholar] [CrossRef]
- Mokuda, S.; Nakamichi, R.; Matsuzaki, T.; Ito, Y.; Sato, T.; Miyata, K.; Inui, M.; Olmer, M.; Sugiyama, E.; Lotz, M.; et al. Wwp2 maintains cartilage homeostasis through regulation of Adamts5. Nat. Commun. 2019, 10, 2429. [Google Scholar] [CrossRef]
- Mendez, M.E.; Murugesh, D.K.; Sebastian, A.; Hum, N.R.; McCloy, S.A.; Kuhn, E.A.; Christiansen, B.A.; Loots, G.G. Antibiotic Treatment Prior to Injury Improves Post-Traumatic Osteoarthritis Outcomes in Mice. Int. J. Mol. Sci. 2020, 21, 6424. [Google Scholar] [CrossRef]
- Mendez, M.E.; Sebastian, A.; Murugesh, D.K.; Hum, N.R.; McCool, J.L.; Hsia, A.W.; Christiansen, B.A.; Loots, G.G. LPS-Induced Inflammation Prior to Injury Exacerbates the Development of Post-Traumatic Osteoarthritis in Mice. J. Bone Miner Res. 2020. [Google Scholar] [CrossRef]
- Sebastian, A.; Murugesh, D.K.; Mendez, M.E.; Hum, N.R.; Rios-Arce, N.D.; McCool, J.L.; Christiansen, B.A.; Loots, G.G. Global Gene Expression Analysis Identifies Age-Related Differences in Knee Joint Transcriptome during the Development of Post-Traumatic Osteoarthritis in Mice. Int. J. Mol. Sci. 2020, 21, 364. [Google Scholar] [CrossRef]
- Lefebvre, V.; Behringer, R.R.; de Crombrugghe, B. L-Sox5, Sox6 and Sox9 control essential steps of the chondrocyte differentiation pathway. Osteoarthr. Cartil. 2001, 9 (Suppl. SA), S69–S75. [Google Scholar] [CrossRef]
- Greenblatt, M.B.; Ritter, S.Y.; Wright, J.; Tsang, K.; Hu, D.; Glimcher, L.H.; Aliprantis, A.O. NFATc1 and NFATc2 repress spontaneous osteoarthritis. Proc. Natl. Acad. Sci. USA 2013, 110, 19914–19919. [Google Scholar] [CrossRef]
- Yoshida, C.A.; Yamamoto, H.; Fujita, T.; Furuichi, T.; Ito, K.; Inoue, K.; Yamana, K.; Zanma, A.; Takada, K.; Ito, Y.; et al. Runx2 and Runx3 are essential for chondrocyte maturation, and Runx2 regulates limb growth through induction of Indian hedgehog. Genes Dev. 2004, 18, 952–963. [Google Scholar] [CrossRef]
- Spater, D.; Hill, T.P.; O’sullivan, R.J.; Gruber, M.; Conner, D.A.; Hartmann, C. Wnt9a signaling is required for joint integrity and regulation of Ihh during chondrogenesis. Development 2006, 133, 3039–3049. [Google Scholar] [CrossRef]
- Elayyan, J.; Lee, E.J.; Gabay, O.; Smith, C.A.; Qiq, O.; Reich, E.; Mobasheri, A.; Henrotin, Y.; Kimber, S.J.; Dvir-Ginzberg, M. LEF1-mediated MMP13 gene expression is repressed by SIRT1 in human chondrocytes. FASEB J. 2017, 31, 3116–3125. [Google Scholar] [CrossRef]
- Seuffert, F.; Weidner, D.; Baum, W.; Schett, G.; Stock, M. Upper zone of growth plate and cartilage matrix associated protein protects cartilage during inflammatory arthritis. Arthritis Res. Ther. 2018, 20, 88. [Google Scholar] [CrossRef]
- Wu, C.L.; Dicks, A.; Steward, N.; Tang, R.; Katz, D.B.; Choi, Y.R.; Guilak, F. Single cell transcriptomic analysis of human pluripotent stem cell chondrogenesis. Nat. Commun. 2021, 12, 362. [Google Scholar] [CrossRef]
- Davidson, E.N.B.; Vitters, E.L.; van Lent, P.L.; van de Loo, F.A.; van den Berg, W.B.; van der Kraan, P.M. Elevated extracellular matrix production and degradation upon bone morphogenetic protein-2 (BMP-2) stimulation point toward a role for BMP-2 in cartilage repair and remodeling. Arthritis Res. Ther. 2007, 9, R102. [Google Scholar] [CrossRef]
- Shu, B.; Zhang, M.; Xie, R.; Wang, M.; Jin, H.; Hou, W.; Tang, D.; Harris, S.E.; Mishina, Y.; O’Keefe, R.J.; et al. BMP2, but not BMP4, is crucial for chondrocyte proliferation and maturation during endochondral bone development. J. Cell Sci. 2011, 124, 3428–3440. [Google Scholar] [CrossRef] [PubMed]
- Pengas, I.; Eldridge, S.; Assiotis, A.; McNicholas, M.; Mendes, J.E.; Laver, L. MMP-3 in the peripheral serum as a biomarker of knee osteoarthritis, 40 years after open total knee meniscectomy. J. Exp. Orthop. 2018, 5, 21. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Sampson, E.R.; Jin, H.; Li, J.; Ke, Q.H.; Im, H.J.; Chen, D. MMP13 is a critical target gene during the progression of osteoarthritis. Arthritis Res. Ther. 2013, 15, R5. [Google Scholar] [CrossRef]
- Chou, C.H.; Lee, M.T.; Song, I.W.; Lu, L.S.; Shen, H.C.; Lee, C.H.; Wu, J.Y.; Chen, Y.T.; Kraus, V.B.; Wu, C.C. Insights into osteoarthritis progression revealed by analyses of both knee tibiofemoral compartments. Osteoarthr. Cartil. 2015, 23, 571–580. [Google Scholar] [CrossRef]
- Tong, Z.; Liu, Y.; Chen, B.; Yan, L.; Hao, D. Association between MMP3 and TIMP3 polymorphisms and risk of osteoarthritis. Oncotarget 2017, 8, 83563–83569. [Google Scholar] [CrossRef]
- Valdes, A.M.; Loughlin, J.; Timms, K.M.; van Meurs, J.J.; Southam, L.; Wilson, S.G.; Doherty, S.; Lories, R.J.; Luyten, F.P.; Gutin, A.; et al. Genome-wide association scan identifies a prostaglandin-endoperoxide synthase 2 variant involved in risk of knee osteoarthritis. Am. J. Hum. Genet. 2008, 82, 1231–1240. [Google Scholar] [CrossRef] [PubMed]
- Styrkarsdottir, U.; Lund, S.H.; Thorleifsson, G.; Zink, F.; Stefansson, O.A.; Sigurdsson, J.K.; Juliusson, K.; Bjarnadottir, K.; Sigurbjornsdottir, S.; Jonsson, S.; et al. Meta-analysis of Icelandic and UK data sets identifies missense variants in SMO, IL11, COL11A1 and 13 more new loci associated with osteoarthritis. Nat. Genet. 2018, 50, 1681–1687. [Google Scholar] [CrossRef] [PubMed]
- Mastbergen, S.C.; Jansen, N.W.; Bijlsma, J.W.; Lafeber, F.P. Differential direct effects of cyclo-oxygenase-1/2 inhibition on proteoglycan turnover of human osteoarthritic cartilage: An in vitro study. Arthritis Res. Ther. 2006, 8, R2. [Google Scholar] [CrossRef]
- De Boer, T.N.; Huisman, A.M.; Polak, A.A.; Niehoff, A.G.; van Rinsum, A.C.; Saris, D.; Bijlsma, J.W.; Lafeber, F.J.; Mastbergen, S.C. The chondroprotective effect of selective COX-2 inhibition in osteoarthritis: Ex vivo evaluation of human cartilage tissue after in vivo treatment. Osteoarthr. Cartil. 2009, 17, 482–488. [Google Scholar] [CrossRef][Green Version]
- Davidson, R.K.; Jupp, O.; de Ferrars, R.; Kay, C.D.; Culley, K.L.; Norton, R.; Driscoll, C.; Vincent, T.L.; Donell, S.T.; Bao, Y.; et al. Sulforaphane represses matrix-degrading proteases and protects cartilage from destruction in vitro and in vivo. Arthritis Rheum. 2013, 65, 3130–3140. [Google Scholar] [CrossRef]
- Bie, Q.; Jin, C.; Zhang, B.; Dong, H. IL-17B: A new area of study in the IL-17 family. Mol. Immunol. 2017, 90, 50–56. [Google Scholar] [CrossRef]
- Kurowska-Stolarska, M.; Alivernini, S. Synovial tissue macrophages: Friend or foe? RMD Open 2017, 3, e000527. [Google Scholar] [CrossRef]
- Brandt, K.D.; Radin, E.L.; Dieppe, P.A.; van de Putte, L. Yet more evidence that osteoarthritis is not a cartilage disease. Ann. Rheum. Dis. 2006, 65, 1261–1264. [Google Scholar] [CrossRef]
- Loeser, R.F.; Goldring, S.R.; Scanzello, C.R.; Goldring, M.B. Osteoarthritis: A disease of the joint as an organ. Arthritis Rheum. 2012, 64, 1697–1707. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sebastian, A.; McCool, J.L.; Hum, N.R.; Murugesh, D.K.; Wilson, S.P.; Christiansen, B.A.; Loots, G.G. Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes. Cells 2021, 10, 1462. https://doi.org/10.3390/cells10061462
Sebastian A, McCool JL, Hum NR, Murugesh DK, Wilson SP, Christiansen BA, Loots GG. Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes. Cells. 2021; 10(6):1462. https://doi.org/10.3390/cells10061462
Chicago/Turabian StyleSebastian, Aimy, Jillian L. McCool, Nicholas R. Hum, Deepa K. Murugesh, Stephen P. Wilson, Blaine A. Christiansen, and Gabriela G. Loots. 2021. "Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes" Cells 10, no. 6: 1462. https://doi.org/10.3390/cells10061462
APA StyleSebastian, A., McCool, J. L., Hum, N. R., Murugesh, D. K., Wilson, S. P., Christiansen, B. A., & Loots, G. G. (2021). Single-Cell RNA-Seq Reveals Transcriptomic Heterogeneity and Post-Traumatic Osteoarthritis-Associated Early Molecular Changes in Mouse Articular Chondrocytes. Cells, 10(6), 1462. https://doi.org/10.3390/cells10061462






