Exosome-Based Molecular Transfer Activity of Macrophage-Like Cells Involves Viability of Oral Carcinoma Cells: Size Exclusion Chromatography and Concentration Filter Method
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells
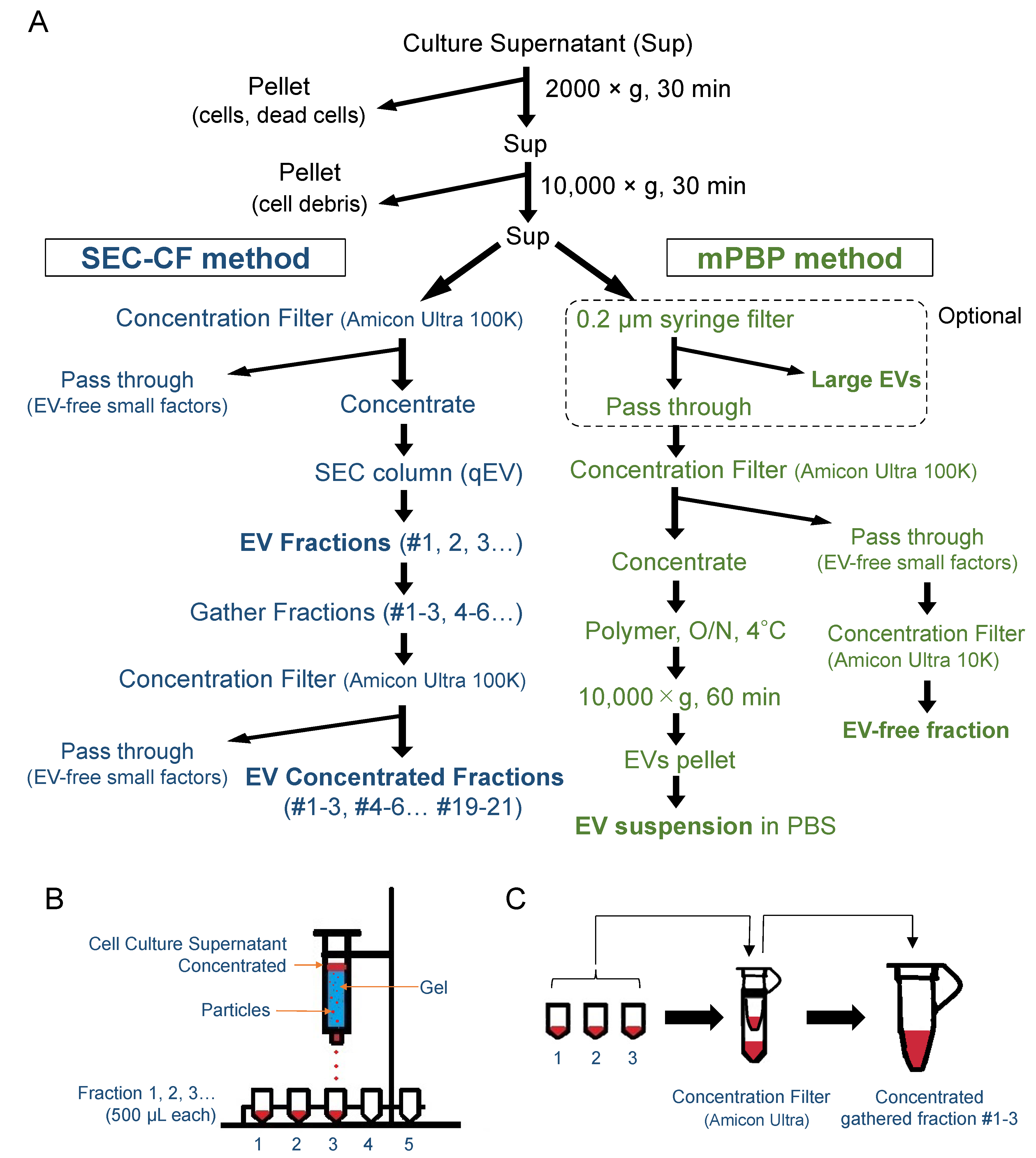
2.2. EV Fractionation (SEC-CF and PBP)
2.3. Particle Size Distribution
2.4. Transmission Electron Microscopy
2.5. Western Blotting
2.6. Fluorescence-Labeling of Cells and EVs
2.7. EV Molecular Transfer Assay
2.8. Cell Viability
2.9. Flow Cytometry
2.10. Statistical Analysis
3. Results
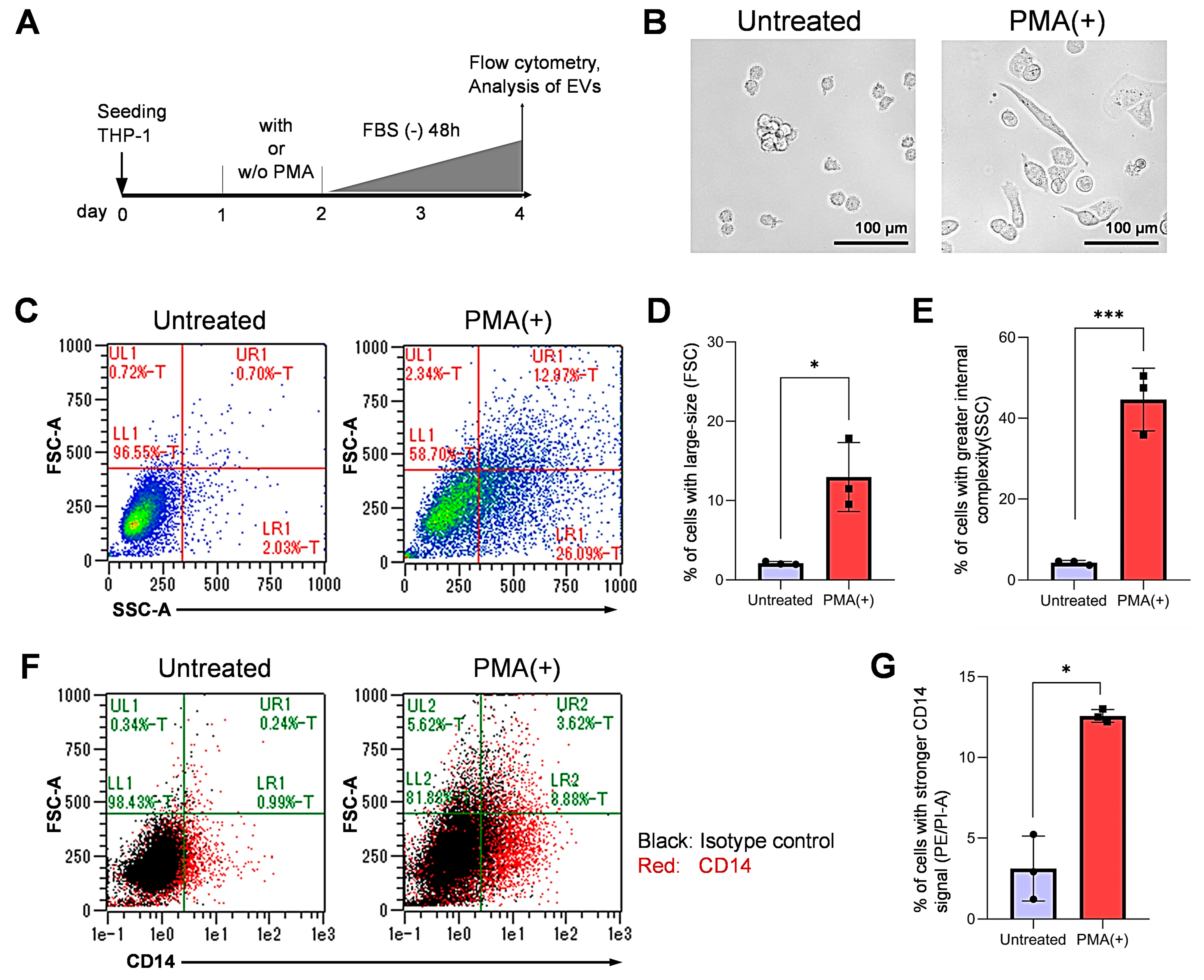
3.1. Differentiation of Monocytic Cells into Macrophage-Like Cells
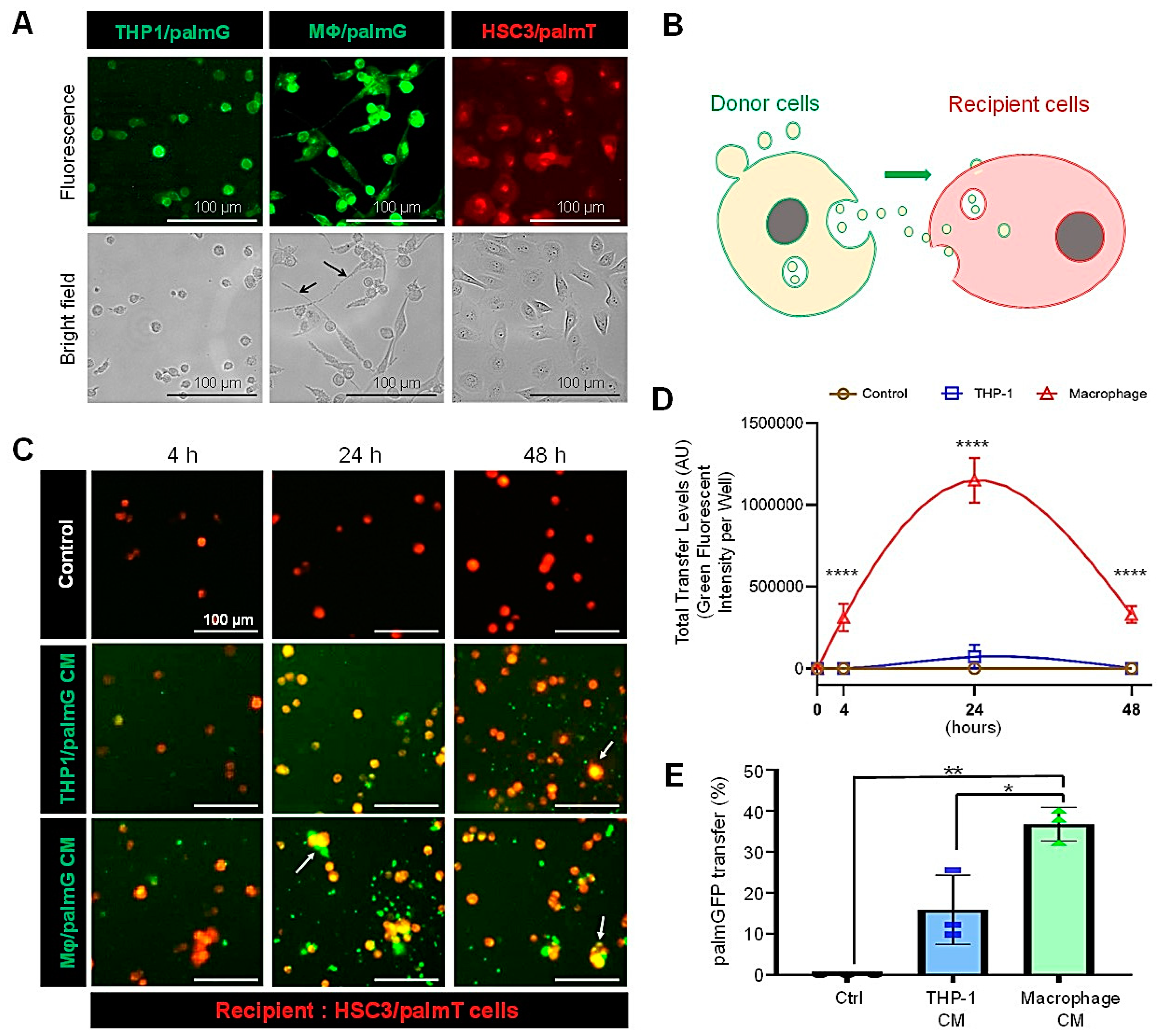
3.2. High Transmission Efficiency of Macrophage-Derived EVs to Oral Carcinoma Cells
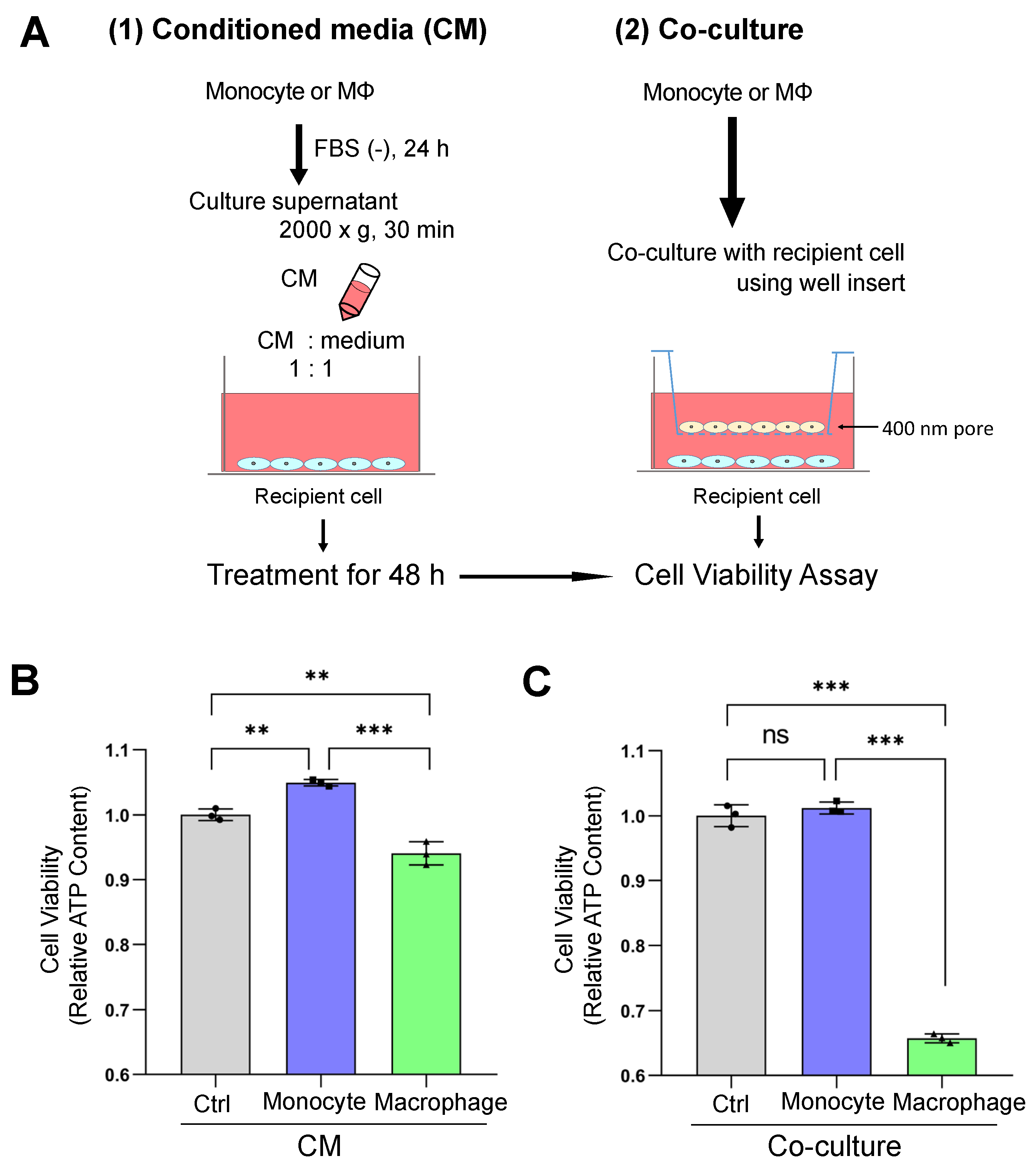
3.3. Macrophage-Secreted Factors Reduced Cell Viability in Receiver Oral Carcinoma Cells
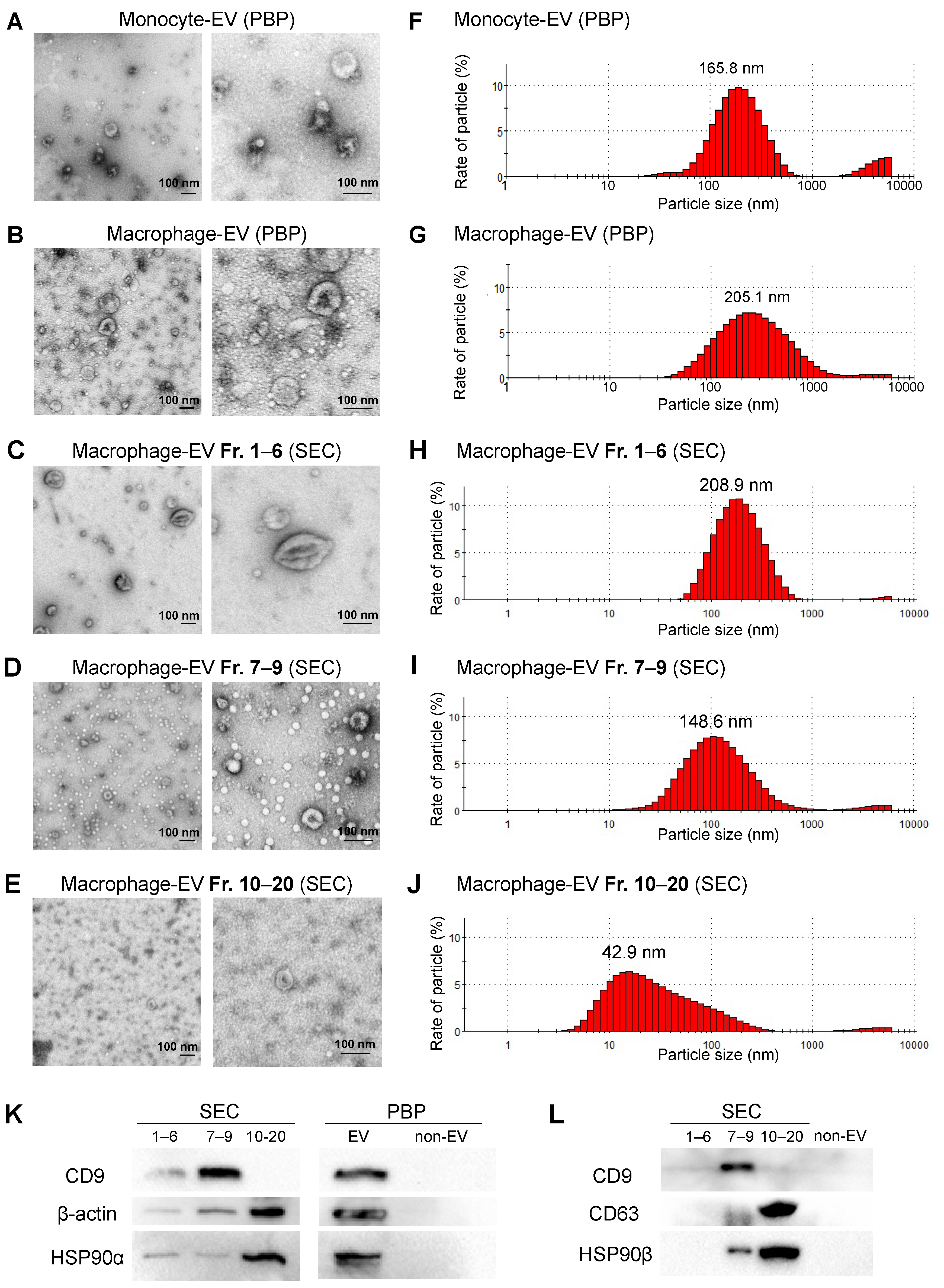
3.4. Separation of Large, Medium, and Small EVs Using the SEC-CF Method
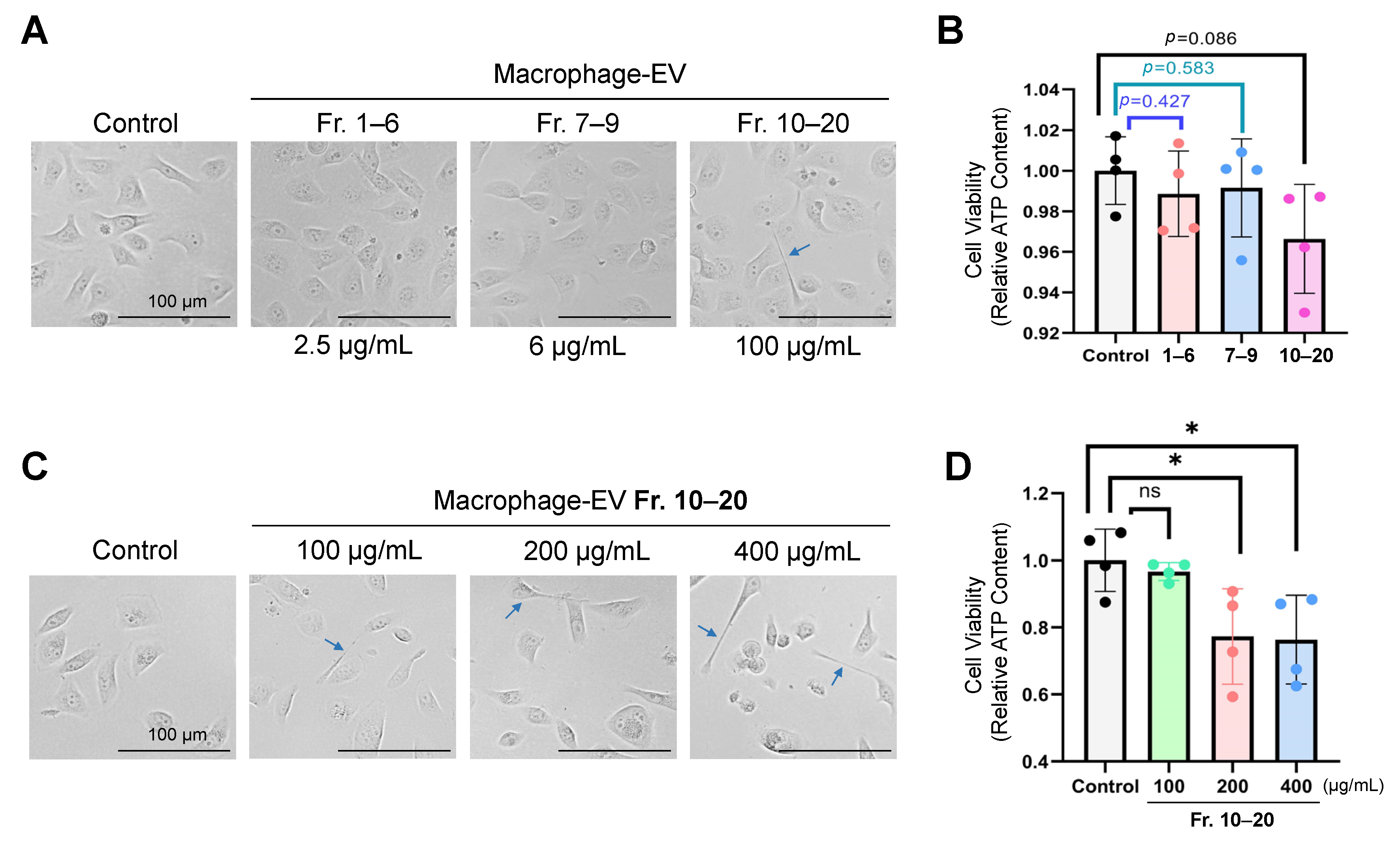
3.5. Macrophage-Secreted Factors (Fraction 10–20 Containing Small Exosomes) Decreased Viability of Oral Carcinoma Cells
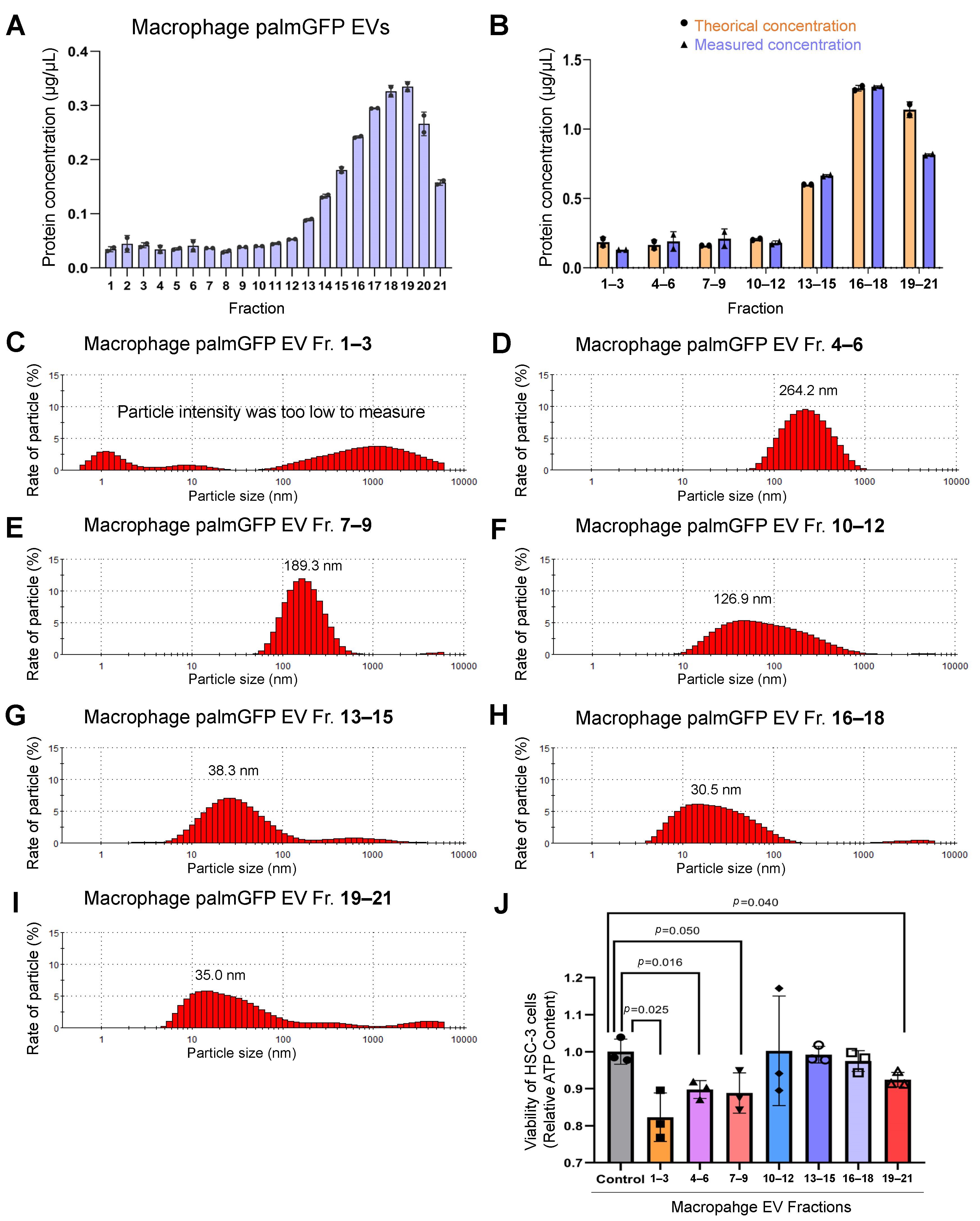
3.6. Size-Based Fractionation of Macrophage/palmGFP-Secreted EVs
3.7. Macrophage-Derived (Large and Small) EVs Reduced Viability of Oral Carcinoma Cells
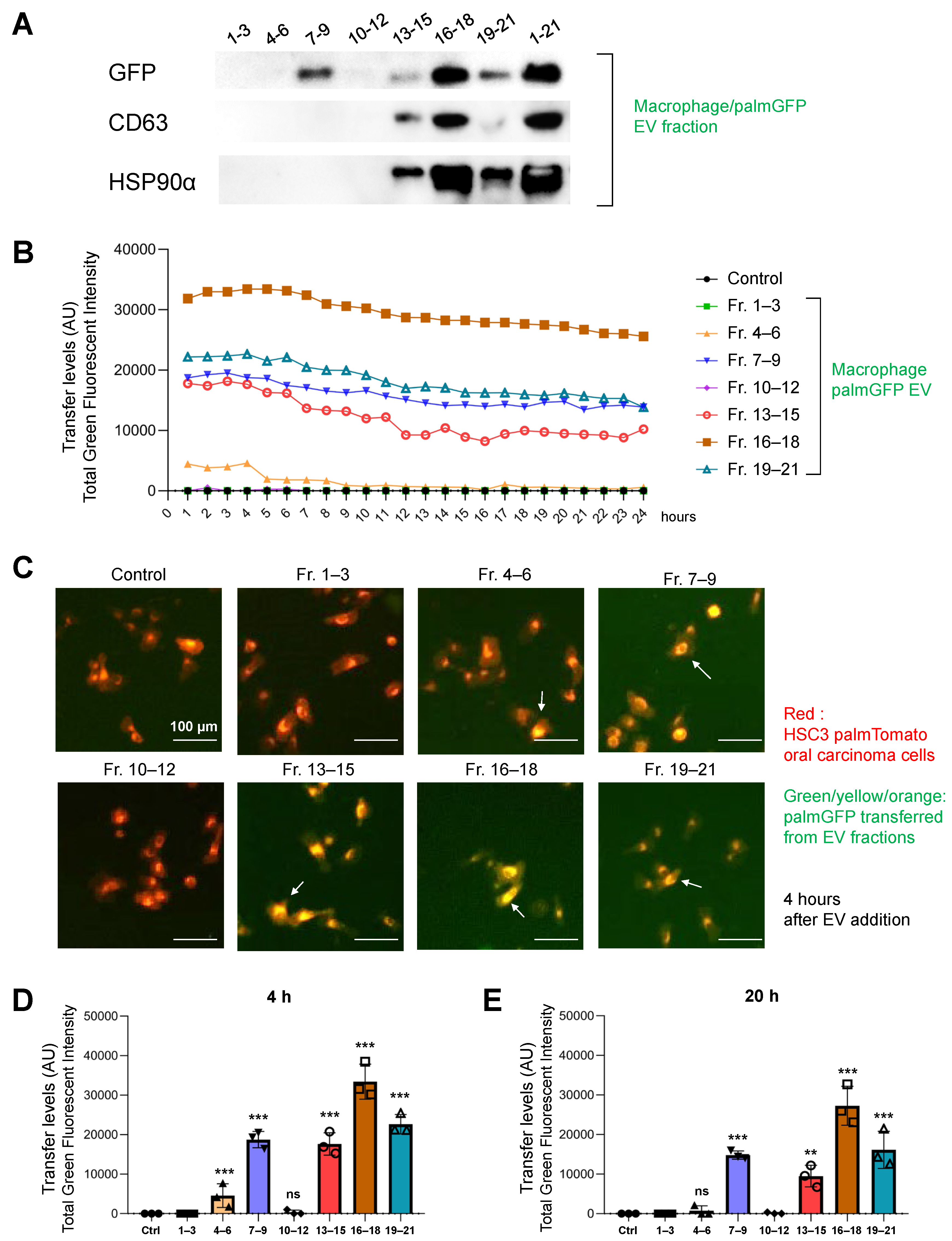
3.8. Molecular Transfer of palmGFP from Macrophage-EVs to Oral Carcinoma Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Colombo, M.; Raposo, G.; Théry, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 2014, 30, 255–289. [Google Scholar] [CrossRef]
- Yáñez-Mó, M.; Siljander, P.R.-M.; Andreu, Z.; Bedina Zavec, A.; Borràs, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef]
- Taha, E.A.; Sogawa, C.; Okusha, Y.; Kawai, H.; Oo, M.W.; Elseoudi, A.; Lu, Y.; Nagatsuka, H.; Kubota, S.; Satoh, A.; et al. Knockout of MMP3 weakens solid tumor organoids and cancer extracellular vesicles. Cancers 2020, 12, 1260. [Google Scholar] [CrossRef]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef]
- Lawson, C.; Vicencio, J.M.; Yellon, D.M.; Davidson, S.M. Microvesicles and exosomes: New players in metabolic and cardiovascular disease. J. Endocrinol. 2016, 228, R57–R71. [Google Scholar] [CrossRef]
- Battistelli, M.; Falcieri, E. Apoptotic bodies: Particular extracellular vesicles involved in intercellular communication. Biology 2020, 9, 21. [Google Scholar] [CrossRef]
- Meehan, B.; Rak, J.; Di Vizio, D. Oncosomes—Large and small: What are they, where they came from? J. Extracell. Vesicles 2016, 5, 33109. [Google Scholar] [CrossRef]
- Vagner, T.; Spinelli, C.; Minciacchi, V.R.; Balaj, L.; Zandian, M.; Conley, A.; Zijlstra, A.; Freeman, M.R.; Demichelis, F.; De, S.; et al. Large extracellular vesicles carry most of the tumour DNA circulating in prostate cancer patient plasma. J. Extracell. Vesicles 2018, 7, 1505403. [Google Scholar] [CrossRef]
- Chen, Q.; Bei, J.J.; Liu, C.; Feng, S.B.; Zhao, W.B.; Zhou, Z.; Yu, Z.P.; Du, X.J.; Hu, H.Y. HMGB1 induces secretion of matrix vesicles by macrophages to enhance ectopic mineralization. PLoS ONE 2016, 11, e0156686. [Google Scholar] [CrossRef]
- Mebarek, S.; Abousalham, A.; Magne, D.; Do, L.D.; Bandorowicz-Pikula, J.; Pikula, S.; Buchet, R. Phospholipases of mineralization competent cells and matrix vesicles: Roles in physiological and pathological mineralizations. Int. J. Mol. Sci. 2013, 14, 5036–5129. [Google Scholar] [CrossRef]
- Huang, Y.; Zucker, B.; Zhang, S.; Elias, S.; Zhu, Y.; Chen, H.; Ding, T.; Li, Y.; Sun, Y.; Lou, J.; et al. Migrasome formation is mediated by assembly of micron-scale tetraspanin macrodomains. Nat. Cell Biol. 2019, 21, 991–1002. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef]
- Seyama, M.; Yoshida, K.; Yoshida, K.; Fujiwara, N.; Ono, K.; Eguchi, T.; Kawai, H.; Guo, J.; Weng, Y.; Haoze, Y.; et al. Outer membrane vesicles of Porphyromonas gingivalis attenuate insulin sensitivity by delivering gingipains to the liver. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165731. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.Y.; Park, H.T.; Dinh, N.T.H.; Choi, S.J.; Lee, J.; Kim, J.H.; Lee, S.-W.; Gho, Y.S. Bacterial outer membrane vesicles suppress tumor by interferon-γ-mediated anti-tumor response. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Eguchi, T.; Sogawa, C.; Okusha, Y.; Uchibe, K.; Iinuma, R.; Ono, K.; Nakano, K.; Murakami, J.; Itoh, M.; Arai, K.; et al. Organoids with cancer stem cell-like properties secrete exosomes and HSP90 in a 3D nanoenvironment. PLoS ONE 2018, 13, e0191109. [Google Scholar] [CrossRef]
- Taha, E.A.; Ono, K.; Eguchi, T. Roles of extracellular HSPs as biomarkers in immune surveillance and immune evasion. Int. J. Mol. Sci. 2019, 20, 4588. [Google Scholar] [CrossRef]
- Eguchi, T.; Sogawa, C.; Ono, K.; Matsumoto, M.; Tran, M.T.; Okusha, Y.; Lang, B.J.; Okamoto, K.; Calderwood, S.K. Cell Stress Induced Stressome Release Including Damaged Membrane Vesicles and Extracellular HSP90 by Prostate Cancer Cells. Cells 2020, 9, 755. [Google Scholar] [CrossRef]
- Ono, K.; Sogawa, C.; Kawai, H.; Tran, M.T.; Taha, E.A.; Lu, Y.; Oo, M.W.; Okusha, Y.; Okamura, H.; Ibaragi, S. Triple knockdown of CDC37, HSP90-alpha and HSP90-beta diminishes extracellular vesicles-driven malignancy events and macrophage M2 polarization in oral cancer. J. Extracell. Vesicles 2020, 9, 1769373. [Google Scholar] [CrossRef]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-’t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2013, 2, 20360. [Google Scholar] [CrossRef]
- Zaborowski, M.P.; Cheah, P.S.; Zhang, X.; Bushko, I.; Lee, K.; Sammarco, A.; Zappulli, V.; Maas, S.L.N.; Allen, R.M.; Rumde, P.; et al. Membrane-bound Gaussia luciferase as a tool to track shedding of membrane proteins from the surface of extracellular vesicles. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef]
- Böing, A.N.; van der Pol, E.; Grootemaat, A.E.; Coumans, F.A.W.; Sturk, A.; Nieuwland, R. Single-step isolation of extracellular vesicles by size-exclusion chromatography. J. Extracell. Vesicles 2014, 3, 23430. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.P.; Kim, E.Y.; Badr, C.E.; Weissleder, R.; Mempel, T.R.; Tannous, B.A.; Breakefield, X.O. Visualization and tracking of tumour extracellular vesicle delivery and RNA translation using multiplexed reporters. Nat. Commun. 2015, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, T.; Eguchi, T.; Sogawa, C.; Ono, K.; Murakami, J.; Ibaragi, S.; ichi Asaumi, J.; Calderwood, S.K.; Okamoto, K.; Kozaki, K.-I. Carcinogenic epithelial-mesenchymal transition initiated by oral cancer exosomes is inhibited by anti-EGFR antibody cetuximab. Oral Oncol. 2018, 86, 251–257. [Google Scholar] [CrossRef]
- Grubisic, Z.; Rempp, P.; Benoit, H. A universal calibration for gel permeation chromatography. J. Polym. Sci. Part. B Polym. Lett. 1967, 5, 753–759. [Google Scholar] [CrossRef]
- Monguió-Tortajada, M.; Gálvez-Montón, C.; Bayes-Genis, A.; Roura, S.; Borràs, F.E. Extracellular vesicle isolation methods: Rising impact of size-exclusion chromatography. Cell. Mol. Life Sci. 2019, 76, 2369–2382. [Google Scholar] [CrossRef]
- Gámez-Valero, A.; Monguió-Tortajada, M.; Carreras-Planella, L.; Franquesa, M.; Beyer, K.; Borràs, F.E. Size-Exclusion Chromatography-based isolation minimally alters Extracellular Vesicles’ characteristics compared to precipitating agents. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Furuta, K.; Eguchi, T. Roles of HSP on Antigen Presentation. In Heat Shock Proteins in Human Diseases; Asea, A.A.A., Kaur, P., Eds.; Springer International Publishing: Cham, Germany, 2021; pp. 275–280. ISBN 978-3-030-62289-3. [Google Scholar]
- Ono, K.; Eguchi, T.; Sogawa, C.; Calderwood, S.K.; Futagawa, J.; Kasai, T.; Seno, M.; Okamoto, K.; Sasaki, A.; Kozaki, K.-i. HSP-enriched properties of extracellular vesicles involve survival of metastatic oral cancer cells. J. Cell. Biochem. 2018, 119, 7350–7362. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 2006, 30, 3.22.1–3.22.29. [Google Scholar] [CrossRef]
- Okusha, Y.; Eguchi, T.; Tran, M.T.; Sogawa, C.; Yoshida, K.; Itagaki, M.; Taha, E.A.; Ono, K.; Aoyama, E.; Okamura, H.; et al. Extracellular vesicles enriched with moonlighting metalloproteinase are highly transmissive, pro-tumorigenic, and trans-activates cellular communication network factor (Ccn2/ctgf): CRISPR against cancer. Cancers 2020, 12, 881. [Google Scholar] [CrossRef]
- Namba, Y.; Sogawa, C.; Okusha, Y.; Kawai, H.; Itagaki, M.; Ono, K.; Murakami, J.; Aoyama, E.; Ohyama, K.; Asaumi, J.I.; et al. Depletion of lipid efflux pump ABCG1 triggers the intracellular accumulation of extracellular vesicles and reduces aggregation and tumorigenesis of metastatic cancer cells. Front. Oncol. 2018, 8, 376. [Google Scholar] [CrossRef]
- Sogawa, C.; Eguchi, T.; Tran, M.T.; Ishige, M.; Trin, K.; Okusha, Y.; Taha, E.A.; Lu, Y.; Kawai, H.; Sogawa, N.; et al. Antiparkinson drug benztropine suppresses tumor growth, circulating tumor cells, and metastasis by acting on SLC6A3/dat and reducing STAT3. Cancers 2020, 12, 523. [Google Scholar] [CrossRef]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367, eaau6977. [Google Scholar] [CrossRef]
- Kowal, J.; Arras, G.; Colombo, M.; Jouve, M.; Morath, J.P.; Primdal-Bengtson, B.; Dingli, F.; Loew, D.; Tkach, M.; Théry, C. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl. Acad. Sci. USA 2016, 113, E968–E977. [Google Scholar] [CrossRef]
- Lu, Y.; Eguchi, T. HSP Stimulation on Macrophages and Dendritic Cells Activates Innate Immune System. In Heat Shock Proteins; Asea, A.A.A., Punit, K., Eds.; Springer: Dordrecht, The Netherland, 2020; pp. 1–15. [Google Scholar]
- Eguchi, T.; Ono, K.; Kawata, K.; Okamoto, K.; Calderwood, S.K. Regulatory Roles of HSP90-Rich Extracellular Vesicles. In Heat Shock Protein 90 in Human Diseases and Disorders; Asea, A.A.A., Kaur, P., Eds.; Springer Nature: Cham, Germany, 2019; pp. 3–17. ISBN 978-3-030-23158-3. [Google Scholar]
- Eustace, B.K.; Sakurai, T.; Stewart, J.K.; Yimlamai, D.; Unger, C.; Zehetmeier, C.; Lain, B.; Torella, C.; Henning, S.W.; Beste, G.; et al. Functional proteomic screens reveal an essential extracellular role for hsp90 alpha in cancer cell invasiveness. Nat. Cell Biol. 2004, 6, 507–514. [Google Scholar] [CrossRef]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Simonsen, J.B. What Are We Looking At? Extracellular Vesicles, Lipoproteins, or Both? Circ. Res. 2017, 121, 920–922. [Google Scholar] [CrossRef]
- Karimi, N.; Cvjetkovic, A.; Jang, S.C.; Crescitelli, R.; Hosseinpour Feizi, M.A.; Nieuwland, R.; Lötvall, J.; Lässer, C. Detailed analysis of the plasma extracellular vesicle proteome after separation from lipoproteins. Cell. Mol. Life Sci. 2018, 75, 2873–2886. [Google Scholar] [CrossRef]
- Schroder, K.; Hertzog, P.J.; Ravasi, T.; Hume, D.A. Interferon-gamma: An overview of signals, mechanisms and functions. J. Leukoc. Biol. 2004, 75, 163–189. [Google Scholar] [CrossRef]
- Murshid, A.; Eguchi, T.; Calderwood, S.K. Stress proteins in aging and life span. Int. J. Hyperth. 2013, 29, 442–447. [Google Scholar] [CrossRef] [PubMed]
- Eguchi, T.; Lang, B.; Murshid, A.; Prince, T.; Gong, J.; Calderwood, S. Regulatory Roles for Hsp70 in Cancer Incidence and Tumor Progression. In Frontiers in Structural Biology; Galigniana, M.D., Ed.; Bentham Science: Dordrecht, The Netherland, 2018; pp. 1–22. ISBN 9781681086156. [Google Scholar]
- Goughnour, P.; Park, M.; Kim, S.; Jun, S.; Yang, W.S.; Chae, S.; Seongmin, C.; Song, C.; Lee, J.-H.; Hyun, J.; et al. Extracellular vesicles derived from macrophage display glycyl-tRNA synthetase 1 exhibits anti-cancer activity. J. Extracell. Vesicles 2020, 10, e12029. [Google Scholar] [CrossRef]
- Park, M.C.; Kang, T.; Jin, D.; Han, J.M.; Kim, S.B.; Park, Y.J.; Cho, K.; Park, Y.W.; Guo, M.; He, W.; et al. Secreted human glycyl-tRNA synthetase implicated in defense against ERK-activated tumorigenesis. Proc. Natl. Acad. Sci. USA 2012, 109, E640–E647. [Google Scholar] [CrossRef] [PubMed]
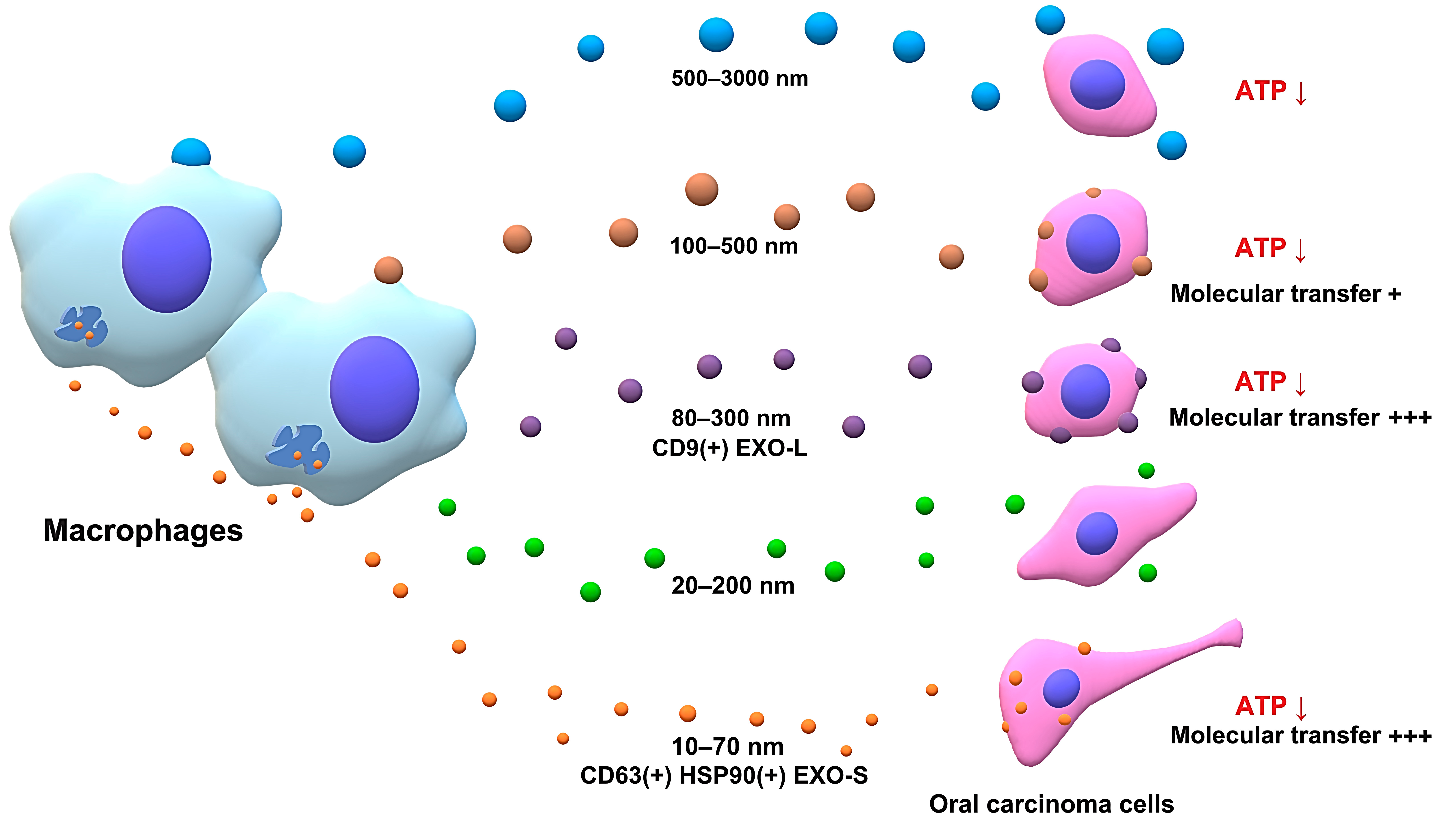
| EV/Particle Fraction | Approx. Size | palmGFP Expression | Molecular Transfer 1 | Approx. Protein Conc. | Marker | ATP Content in Receiver Cells |
|---|---|---|---|---|---|---|
| Fr. 1–3 | 500–3000 nm | nd. | nd. | 0.2 µg/µL | ni. | Decreased |
| Fr. 4–6 | 100–500 nm | nd. | Rapid | 0.2 µg/µL | ni. | Decreased |
| Fr. 7–9 | 80–300 nm | Positive | High | 0.2 µg/µL | CD9 | Decreased |
| Fr. 10–12 | 20–200 nm | nd. | nd. | 0.2 µg/µL | ni. | na. |
| Fr. 13–21 | 10–70 nm | Positive | High | 0.5–2.0 µg/µL | CD63 HSP90α/β | Decreased |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, Y.; Eguchi, T.; Sogawa, C.; Taha, E.A.; Tran, M.T.; Nara, T.; Wei, P.; Fukuoka, S.; Miyawaki, T.; Okamoto, K. Exosome-Based Molecular Transfer Activity of Macrophage-Like Cells Involves Viability of Oral Carcinoma Cells: Size Exclusion Chromatography and Concentration Filter Method. Cells 2021, 10, 1328. https://doi.org/10.3390/cells10061328
Lu Y, Eguchi T, Sogawa C, Taha EA, Tran MT, Nara T, Wei P, Fukuoka S, Miyawaki T, Okamoto K. Exosome-Based Molecular Transfer Activity of Macrophage-Like Cells Involves Viability of Oral Carcinoma Cells: Size Exclusion Chromatography and Concentration Filter Method. Cells. 2021; 10(6):1328. https://doi.org/10.3390/cells10061328
Chicago/Turabian StyleLu, Yanyin, Takanori Eguchi, Chiharu Sogawa, Eman A. Taha, Manh Tien Tran, Toshiki Nara, Penggong Wei, Shiro Fukuoka, Takuya Miyawaki, and Kuniaki Okamoto. 2021. "Exosome-Based Molecular Transfer Activity of Macrophage-Like Cells Involves Viability of Oral Carcinoma Cells: Size Exclusion Chromatography and Concentration Filter Method" Cells 10, no. 6: 1328. https://doi.org/10.3390/cells10061328
APA StyleLu, Y., Eguchi, T., Sogawa, C., Taha, E. A., Tran, M. T., Nara, T., Wei, P., Fukuoka, S., Miyawaki, T., & Okamoto, K. (2021). Exosome-Based Molecular Transfer Activity of Macrophage-Like Cells Involves Viability of Oral Carcinoma Cells: Size Exclusion Chromatography and Concentration Filter Method. Cells, 10(6), 1328. https://doi.org/10.3390/cells10061328







