Sex-Biased lncRNA Signature in Fetal Growth Restriction (FGR)
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Clinical Characteristics of Placental Samples
2.3. Library Preparation and High-Throughput Sequencing
2.4. Quality Control, Mapping and Differentially Expressed Analysis
2.5. Identification of Known and Novel lncRNAs
2.6. LncRNA Target Functional Network Analyses
2.7. Prediction of Circular Organization of Transcripts
2.8. Validation of RNA-Seq Results Using External Transcriptomic Datasets
2.9. Validation of RNA-Seq Results Using Quantitative Real-Time PCR (RT-qPCR)
3. Results
3.1. Mapping and Clustering of FGR-Affected RNA-Seq Libraries
3.2. Identification of lncRNAs in FGR-Affected Placentas
3.3. DEGs and lncRNAs in a Female Fetus Affected by FGR (ff-FGR)
3.4. DEGs and lncRNAs in a Male Fetus Affected by FGR (mf-FGR)
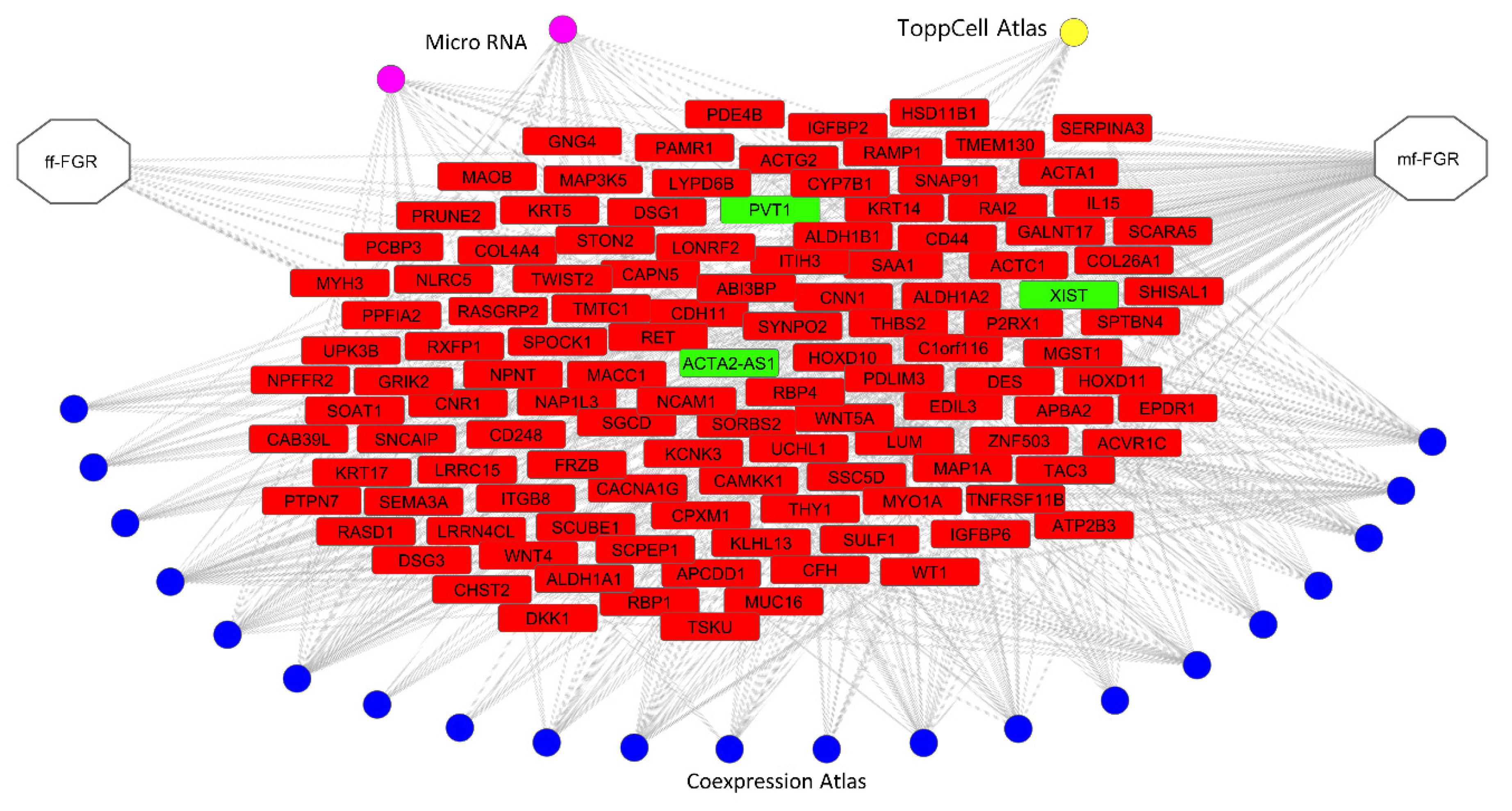
3.5. lncRNA—Target Gene Relationships
3.6. Circular Organization of Transcripts
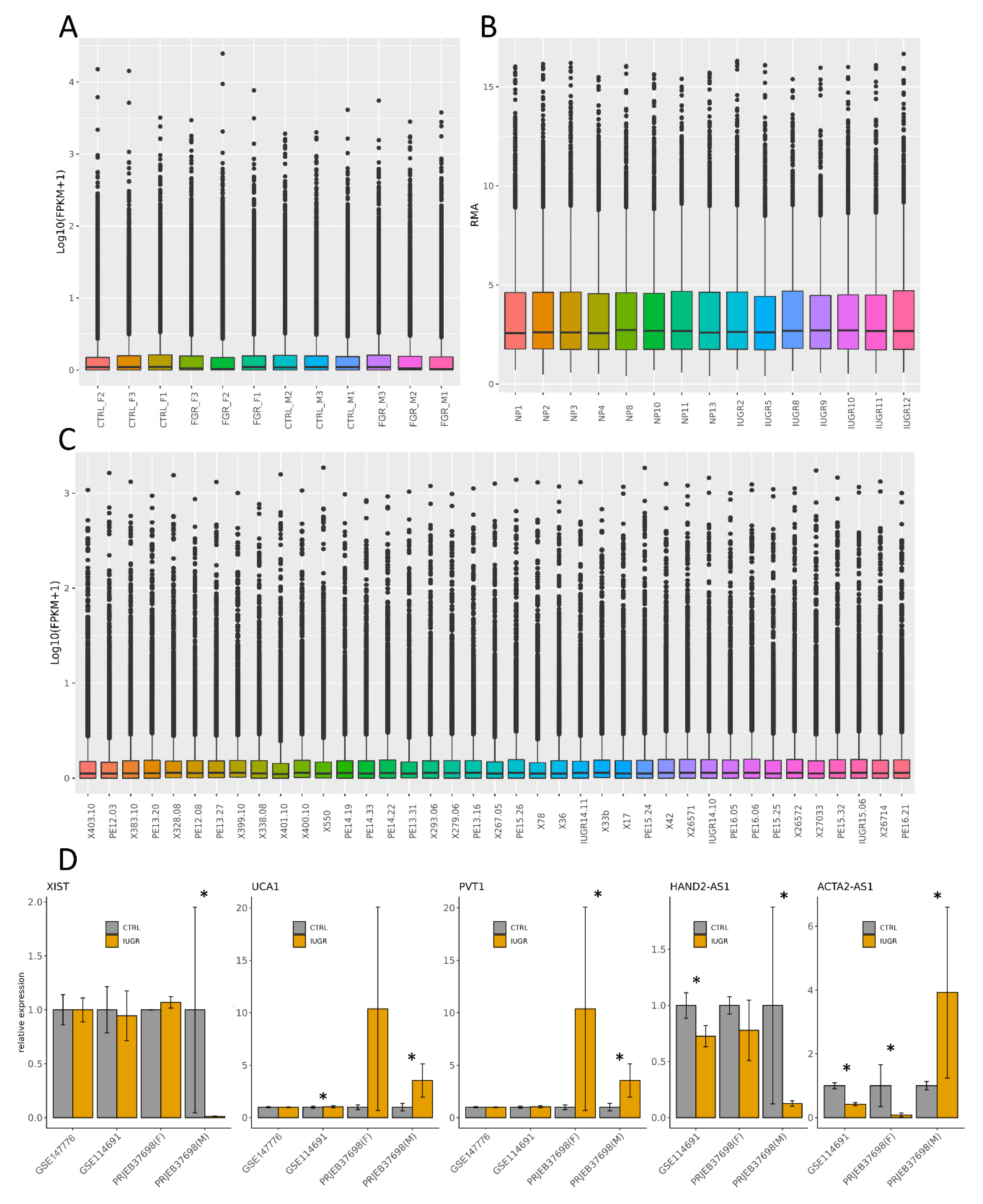
3.7. Validation of RNA-Seq Results Using External Transcriptomic Datasets
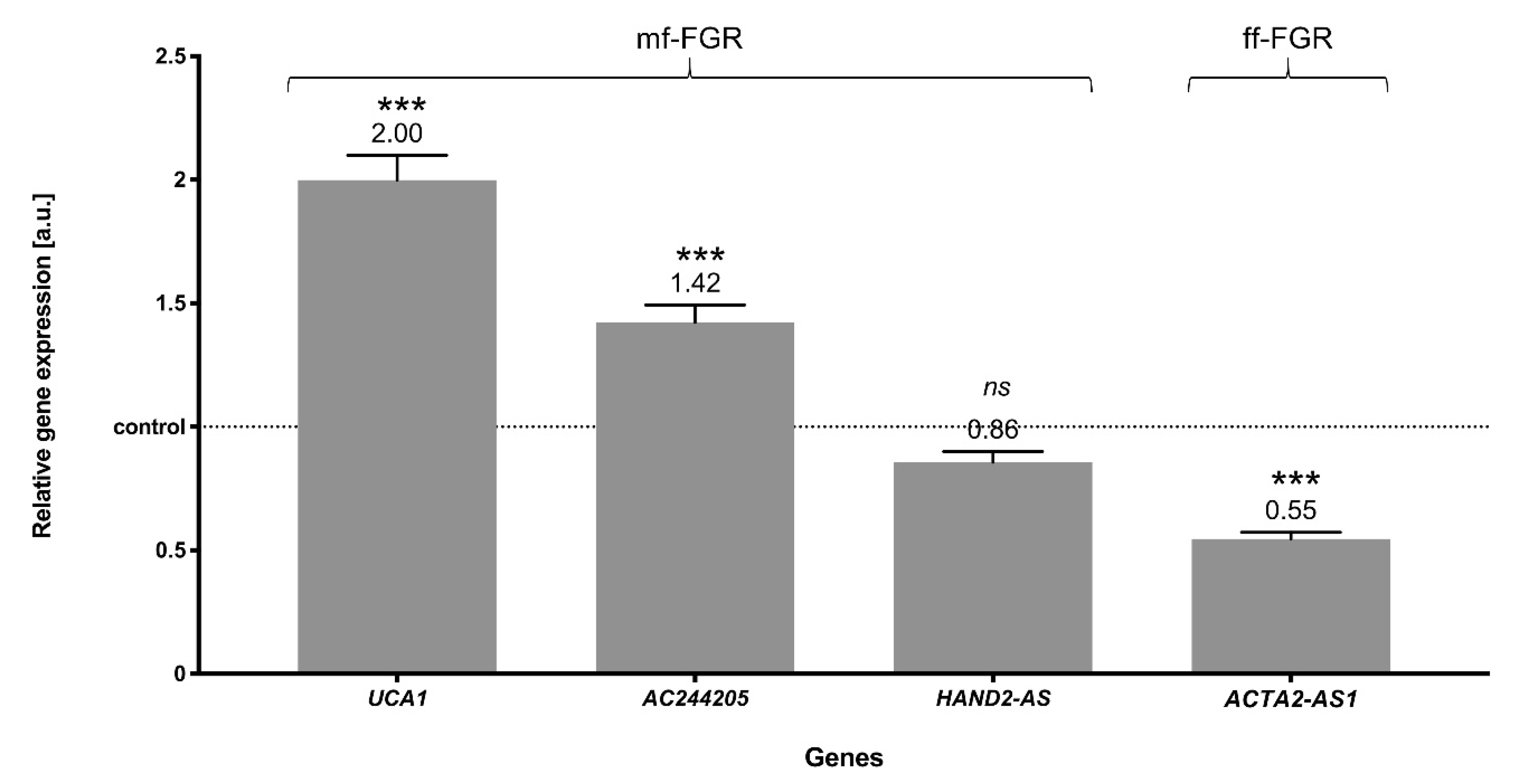
3.8. Validation of RNA-Seq Results Using Quantitative Real-Time PCR (RT-qPCR)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Albu, A.R.; Anca, A.F.; Horhoianu, V.V.; Horhoianu, I.A. Predictive factors for intrauterine growth restriction. J. Med. Life 2014, 7, 165–171. [Google Scholar]
- Zeitlin, J.; El Ayoubi, M.; Jarreau, P.H.; Draper, E.S.; Blondel, B.; Künzel, W.; Cuttini, M.; Kaminski, M.; Gortner, L.; Van Reempts, P.; et al. Impact of fetal growth restriction on mortality and morbidity in a very preterm birth cohort. J. Pediatr. 2010, 157, 733–739.e1. [Google Scholar] [CrossRef] [PubMed]
- Serena, C.; Marchetti, G.; Rambaldi, M.P.; Ottanelli, S.; Di Tommaso, M.; Avagliano, L.; Pieralli, A.; Mello, G.; Mecacci, F. Stillbirth and fetal growth restriction. J. Matern. Neonatal Med. 2013, 26, 16–20. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Shastri, S.; Sharma, P. Intrauterine Growth Restriction: Antenatal and Postnatal Aspects. Clin. Med. Insights Pediatr. 2016, 10, CMPed.S40070. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Sharma, P.; Shastri, S. Genetic, metabolic and endocrine aspect of intrauterine growth restriction: An update. J. Matern. Neonatal Med. 2017, 30, 2263–2275. [Google Scholar] [CrossRef]
- Haroun, H.S.W. Intrauterine Growth Restriction. Anat. Physiol. Biochem. Int. J. 2017, 1, 555572. [Google Scholar] [CrossRef]
- Eydoux, P.; Choiset, A.; le Porrier, N.; Thépot, F.; Szpiro-Tapia, S.; Alliet, J.; Ramond, S.; Viel, J.F.; Gautier, E.; Morichon, N.; et al. Chromosomal prenatal diagnosis: Study of 936 cases of intrauterine abnormalities after ultrasound assessment. Prenat. Diagn. 1989, 9, 255–269. [Google Scholar] [CrossRef]
- Veerbeek, J.H.W.; Nikkels, P.G.J.; Torrance, H.L.; Gravesteijn, J.; Post Uiterweer, E.D.; Derks, J.B.; Koenen, S.V.; Visser, G.H.A.; Van Rijn, B.B.; Franx, A. Placental pathology in early intrauterine growth restriction associated with maternal hypertension. Placenta 2014, 35, 696–701. [Google Scholar] [CrossRef] [PubMed]
- Murthi, P.; Doherty, V.; Said, J.; Donath, S.; Brennecke, S.P.; Kalionis, B. Homeobox Gene HLX1Expression Is Decreased in Idiopathic Human Fetal Growth Restriction. Am. J. Pathol. 2006, 168, 511–518. [Google Scholar] [CrossRef]
- Janzen, C.; Lei, M.Y.Y.; Jeong, I.S.D.; Ganguly, A.; Sullivan, P.; Paharkova, V.; Capodanno, G.; Nakamura, H.; Perry, A.; Shin, B.-C.; et al. Humanin (HN) and glucose transporter 8 (GLUT8) in pregnancies complicated by intrauterine growth restriction. PLoS ONE 2018, 13, e0193583. [Google Scholar] [CrossRef]
- Damodaram, M.; Story, L.; Kulinskaya, E.; Rutherford, M.; Kumar, S. Early adverse perinatal complications in preterm growth-restricted fetuses. Aust. N. Z. J. Obstet. Gynaecol. 2011, 51, 204–209. [Google Scholar] [CrossRef]
- Murray, E.; Fernandes, M.; Fazel, M.; Kennedy, S.; Villar, J.; Stein, A. Differential effect of intrauterine growth restriction on childhood neurodevelopment: A systematic review. BJOG Int. J. Obstet. Gynaecol. 2015, 122, 1062–1072. [Google Scholar] [CrossRef] [PubMed]
- Wixey, J.A.; Chand, K.K.; Colditz, P.B.; Bjorkman, S.T. Review: Neuroinflammation in intrauterine growth restriction. Placenta 2017, 54, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Yzydorczyk, C.; Armengaud, J.B.; Peyter, A.C.; Chehade, H.; Cachat, F.; Juvet, C.; Siddeek, B.; Simoncini, S.; Sabatier, F.; Dignat-George, F.; et al. Endothelial dysfunction in individuals born after fetal growth restriction: Cardiovascular and renal consequences and preventive approaches. J. Dev. Orig. Health Dis. 2017, 8, 448–464. [Google Scholar] [CrossRef]
- Gardosi, J.; Madurasinghe, V.; Williams, M.; Malik, A.; Francis, A. Maternal and fetal risk factors for stillbirth: Population based study. BMJ 2013, 346, f108. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.; Joseph, K.S. Fetal growth: A review of terms, concepts and issues relevant to obstetrics. Ultrasound Obstet. Gynecol. 2013, 41, 136–145. [Google Scholar] [CrossRef]
- Sarvari, S.I.; Rodriguez-Lopez, M.; Nuñez-Garcia, M.; Sitges, M.; Sepulveda-Martinez, A.; Camara, O.; Butakoff, C.; Gratacos, E.; Bijnens, B.; Crispi, F. Persistence of Cardiac Remodeling in Preadolescents With Fetal Growth Restriction. Circ. Cardiovasc. Imaging 2017, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wu, Z.; Fu, X.; Han, W. Long Noncoding RNAs: Insights from Biological Features and Functions to Diseases. Med. Res. Rev. 2013, 33, 517–553. [Google Scholar] [CrossRef] [PubMed]
- Majewska, M.; Lipka, A.; Paukszto, L.; Jastrzebski, J.P.; Myszczynski, K.; Gowkielewicz, M.; Jozwik, M.; Majewski, M.K. Transcriptome profile of the human placenta. Funct. Integr. Genomics 2017, 17, 551–563. [Google Scholar] [CrossRef] [PubMed]
- Mattick, J.S.; Rinn, J.L. Discovery and annotation of long noncoding RNAs. Nat. Struct. Mol. Biol. 2015, 22, 5–7. [Google Scholar] [CrossRef] [PubMed]
- Taylor, D.H.; Chu, E.T.-J.; Spektor, R.; Soloway, P.D. Long non-coding RNA regulation of reproduction and development. Mol. Reprod. Dev. 2015, 82, 932–956. [Google Scholar] [CrossRef]
- Clark, B.S.; Blackshaw, S. Long non-coding RNA-dependent transcriptional regulation in neuronal development and disease. Front. Genet. 2014, 5, 164. [Google Scholar] [CrossRef] [PubMed]
- Khorkova, O.; Myers, A.J.; Hsiao, J.; Wahlestedt, C. Natural antisense transcripts. Hum. Mol. Genet. 2014, 23, R54–R63. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef] [PubMed]
- Barrett, S.P.; Salzman, J. Circular RNAs: Analysis, expression and potential functions. Development 2016, 143, 1838–1847. [Google Scholar] [CrossRef] [PubMed]
- Forsén, T.; Eriksson, J.G.; Tuomilehto, J.; Osmond, C.; Barker, J.P. Growth in utero and during childhood among women who develop coronary heart disease: Longitudinal study. BMJ 1999, 319, 1403–1407. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, J.G.; Kajantie, E.; Osmond, C.; Thornburg, K.; Barker, D.J.P. Boys Live Dangerously in the Womb. Am. J. Hum. Biol. 2010, 22, 330–335. [Google Scholar] [CrossRef]
- Misra, D.P.; Salafia, C.M.; Miller, R.K.; Charles, A.K. Non-linear and gender-specific relationships among placental growth measures and the fetoplacental weight ratio HHS Public Access. Placenta 2009, 30, 1052–1057. [Google Scholar] [CrossRef] [PubMed]
- Buckberry, S.; Bianco-Miotto, T.; Bent, S.J.; Dekker, G.A.; Roberts, C.T. Integrative transcriptome meta-analysis reveals widespread sex-biased gene expression at the human fetal-maternal interface. Mol. Hum. Reprod. 2014, 20, 810–819. [Google Scholar] [CrossRef]
- Majewska, M.; Lipka, A.; Paukszto, L.; Jastrzebski, J.P.; Gowkielewicz, M.; Jozwik, M.; Majewski, M.K. Preliminary RNA-Seq Analysis of Long Non-Coding RNAs Expressed in Human Term Placenta. Int. J. Mol. Sci. 2018, 19, 1894. [Google Scholar] [CrossRef]
- Majewska, M.; Lipka, A.; Paukszto, L.; Jastrzebski, J.P.; Szeszko, K.; Gowkielewicz, M.; Lepiarczyk, E.; Jozwik, M.; Majewski, M.K. Placenta transcriptome profiling in intrauterine growth restriction (IUGR). Int. J. Mol. Sci. 2019, 20, 1510. [Google Scholar] [CrossRef]
- Pouresmaeili, F.; Azari, I.; Arsang-Jang, S.; Taheri, M.; Ghafouri-Fard, S. Association between expression of long noncoding RNAs in placenta and pregnancy features. Per. Med. 2019, 16, 457–466. [Google Scholar] [CrossRef]
- Cui, W.; Qian, Y.; Zhou, X.; Lin, Y.; Jiang, J.; Chen, J.; Zhao, Z.; Shen, B. Discovery and characterization of long intergenic non-coding RNAs (lincRNA) module biomarkers in prostate cancer: An integrative analysis of RNA-Seq data. BMC Genom. 2015, 16, S3. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 2012, 482, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Figueras, F.; Gratacós, E. Update on the diagnosis and classification of fetal growth restriction and proposal of a stage-based management protocol. Fetal Diagn. Ther. 2014, 36, 86–98. [Google Scholar] [CrossRef]
- Hadlock, F.P.; Harrist, R.B.; Sharman, R.S.; Deter, R.L.; Park, S.K. Estimation of fetal weight with the use of head, body, and femur measurements—A prospective study. Am. J. Obstet. Gynecol. 1985, 151, 333–337. [Google Scholar] [CrossRef]
- Burton, G.J.; Sebire, N.J.; Myatt, L.; Tannetta, D.; Wang, Y.-L.; Sadovsky, Y.; Staff, A.C.; Redman, C.W. Optimising sample collection for placental research. Placenta 2014, 35, 9–22. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Babraham Bioinformatics. 2010. Available online: www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 1 January 2021).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Roberts, A.; Pimentel, H.; Trapnell, C.; Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 2011, 27, 2325–2329. [Google Scholar] [CrossRef] [PubMed]
- Warnes, G.R.; Bolker, B.M.; Bonebakker, L.; Gentleman, R.; Liaw, A.; Lumley, T.; Maechler, M.; Magnusson, A.; Moeller, S.; Schwartz, M.; et al. Package ‘ gplots ’. Package, 2013. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Reimand, J.; Arak, T.; Adler, P.; Kolberg, L.; Reisberg, S.; Peterson, H.; Vilo, J. g:Profiler—A web server for functional interpretation of gene lists. Nucleic Acids Res. 2016, 44, W83–W89. [Google Scholar] [CrossRef]
- Kang, Y.J.; Yang, D.C.; Kong, L.; Hou, M.; Meng, Y.Q.; Wei, L.; Gao, G. CPC2: A fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res. 2017, 45, W12–W16. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Zhang, J.; Zhou, Z. PLEK: A tool for predicting long non-coding RNAs and messenger RNAs based on an improved k-mer scheme. BMC Bioinform. 2014, 15, 311. [Google Scholar] [CrossRef] [PubMed]
- Wucher, V.; Legeai, F.; Hédan, B.; Rizk, G.; Lagoutte, L.; Leeb, T.; Jagannathan, V.; Cadieu, E.; David, A.; Lohi, H.; et al. FEELnc: A tool for long non-coding RNA annotation and its application to the dog transcriptome. Nucleic Acids Res. 2017, gkw1306. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Park, H.J.; Dasari, S.; Wang, S.; Kocher, J.-P.; Li, W. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 2013, 41, e74. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Luo, H.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Liu, Y.; Chen, R.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S. Rfam: An RNA family database. Nucleic Acids Res. 2003, 31, 439–441. [Google Scholar] [CrossRef] [PubMed]
- Kalvari, I.; Nawrocki, E.P.; Argasinska, J.; Quinones-Olvera, N.; Finn, R.D.; Bateman, A.; Petrov, A.I. Non-Coding RNA Analysis Using the Rfam Database. Curr. Protoc. Bioinform. 2018, 62, e51. [Google Scholar] [CrossRef] [PubMed]
- Kaimal, V.; Bardes, E.E.; Tabar, S.C.; Jegga, A.G.; Aronow, B.J. ToppCluster: A multiple gene list feature analyzer for comparative enrichment clustering and network-based dissection of biological systems. Nucleic Acids Res. 2010, 38, W96–W102. [Google Scholar] [CrossRef]
- Shannon, P. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Jakobi, T.; Uvarovskii, A.; Dieterich, C. circtools-a one-stop software solution for circular RNA research. Gene Expr. Receiv. 2018. [Google Scholar] [CrossRef]
- Cheng, J.; Metge, F.; Dieterich, C. Specific identification and quantification of circular RNAs from sequencing data. Bioinformatics 2016, 32, 1094–1096. [Google Scholar] [CrossRef] [PubMed]
- Metge, F.; Czaja-Hasse, L.F.; Reinhardt, R.; Dieterich, C. FUCHS-towards full circular RNA characterization using RNAseq. PeerJ 2017, 2017, 1–14. [Google Scholar] [CrossRef]
- Kent, W.J. BLAT—The BLAST-like alignment tool. Genome Res. 2002. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 45e. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.; Acuña, S.; Aoki, J.; Floeter-Winter, L.; Muxel, S. Long Non-Coding RNAs in the Regulation of Gene Expression: Physiology and Disease. Non-Coding RNA 2019, 5, 17. [Google Scholar] [CrossRef] [PubMed]
- McAninch, D.; Roberts, C.T.; Bianco-Miotto, T. Mechanistic insight into long noncoding RNAs and the placenta. Int. J. Mol. Sci. 2017, 18, 1371. [Google Scholar] [CrossRef]
- Zheng, H.; Brennan, K.; Hernaez, M.; Gevaert, O. Benchmark of long non-coding RNA quantification for RNA sequencing of cancer samples. Gigascience 2019, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Meng, T. Long Noncoding RNA in Preeclampsia: Transcriptional Noise or Innovative Indicators? Biomed Res. Int. 2019, 2019, 5437621. [Google Scholar] [CrossRef]
- Basak, T.; Ain, R. Long non-coding RNAs in placental development and disease. Non-Coding RNA Investig. 2019, 3, 14. [Google Scholar] [CrossRef]
- Filardi, T.; Catanzaro, G.; Mardente, S.; Zicari, A.; Santangelo, C.; Lenzi, A.; Morano, S.; Ferretti, E. Non-coding RNA: Role in gestational diabetes pathophysiology and complications. Int. J. Mol. Sci. 2020, 21, 4020. [Google Scholar] [CrossRef]
- Azari, I.; Ghafouri-Fard, S.; Omrani, M.D.; Arsang-Jang, S.; Tamandani, D.M.K.; Rigi, M.S.; Rafiee, S.; Pouresmaeili, F.; Taheri, M. Expression of long non-coding RNAs in placentas of intrauterine growth restriction (IUGR) pregnancies. Rep. Biochem. Mol. Biol. 2019, 8, 25–31. [Google Scholar] [PubMed]
- Lipka, A.; Paukszto, L.; Majewska, M.; Jastrzebski, J.P.; Myszczynski, K.; Panasiewicz, G.; Szafranska, B. Identification of differentially expressed placental transcripts during multiple gestations in the Eurasian beaver (Castor fiber L.). Reprod. Fertil. Dev. 2017, 29, 2073. [Google Scholar] [CrossRef] [PubMed]
- Awamleh, Z.; Gloor, G.B.; Han, V.K.M. Placental microRNAs in pregnancies with early onset intrauterine growth restriction and preeclampsia: Potential impact on gene expression and pathophysiology. BMC Med. Genom. 2019, 12, 91. [Google Scholar] [CrossRef]
- Medina-Bastidas, D.; Guzmán-Huerta, M.; Borboa-Olivares, H.; Ruiz-Cruz, C.; Parra-Hernández, S.; Flores-Pliego, A.; Salido-Guadarrama, I.; Camargo-Marín, L.; Arambula-Meraz, E.; Estrada-Gutierrez, G. Placental Microarray Profiling Reveals Common mRNA and lncRNA Expression Patterns in Preeclampsia and Intrauterine Growth Restriction. Int. J. Mol. Sci. 2020, 21, 3597. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Lee, J.-A.; Nam, Y.K. Molecular characterization of three muscle alpha actin genes in mud loach (Misgurnus mizolepis; Cypriniformes). Fish. Aquat. Sci. 2017, 20, 27. [Google Scholar] [CrossRef]
- Matsson, H.; Eason, J.; Bookwalter, C.S.; Klar, J.; Gustavsson, P.; Sunnegå Rdh, J.; Enell, H.; Jonzon, A.; Vikkula, M.; Gutierrez, I.; et al. Alpha-cardiac actin mutations produce atrial septal defects. Hum. Mol. Genet. 2008. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishna, U.; Albayrak, S.; Zafra, R.; Baraa, A.; Vishweswaraiah, S.; Veerappa, A.M.; Mahishi, D.; Saiyed, N.; Mishra, N.K.; Guda, C.; et al. Placental epigenetics for evaluation of fetal congenital heart defects: Ventricular septal defect (VSD). PLoS ONE 2019, 14, e200229. [Google Scholar] [CrossRef]
- Krcmery, J.; Gupta, R.; Sadleir, R.W.; Ahrens, M.J.; Misener, S. Loss of the Cytoskeletal Protein Pdlim7 Predisposes Mice to Heart Defects and Hemostatic Dysfunction. PLoS ONE 2013, 8, e80809. [Google Scholar] [CrossRef] [PubMed]
- Morales, A.; Painter, T.; Li, R.; Jill, D.; Siegfried, M.S.; Li, D.; Norton, N.; Ray, E.; Hershberger, M. Rare variant mutations in pregnancy-associated or peripartum cardiomyopathy. Circulation, 2010; 121, 2176–2182. [Google Scholar] [CrossRef]
- López-Tello, J.; Pérez-García, V.; Khaira, J.; Kusinski, L.C.; Cooper, W.N.; Andreani, A.; Grant, I.; de Liger, E.F.; Lam, B.Y.H.; Hemberger, M.; et al. Fetal and trophoblast PI3K p110α have distinct roles in regulating resource supply to the growing fetus in mice. Elife 2019, 8, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Ai, X.; Do, A.-T.; Kusche-Gullberg, M.; Lindahl, U.; Lu, K.; Emerson, C.P. Substrate Specificity and Domain Functions of Extracellular Heparan Sulfate 6- O -Endosulfatases, QSulf1 and QSulf2. J. Biol. Chem. 2006, 281, 4969–4976. [Google Scholar] [CrossRef] [PubMed]
- Dhoot, G.K. Regulation of Wnt Signaling and Embryo Patterning by an Extracellular Sulfatase. Science 2001, 293, 1663–1666. [Google Scholar] [CrossRef] [PubMed]
- Moghbeli, M. Genetics of recurrent pregnancy loss among Iranian population. Mol. Genet. Genomic Med. 2019, 7, 1–15. [Google Scholar] [CrossRef]
- Langsdorf, A.; Do, A.-T.; Kusche-Gullberg, M.; Emerson, C.P., Jr.; Ai, X. Sulfs are regulators of growth factor signaling for satellite cell differentiation and muscle regeneration. Dev. Biol. 2007, 311, 464–477. [Google Scholar] [CrossRef]
- Ratzka, A.; Kalus, I.; Moser, M.; Dierks, T.; Mundlos, S.; Vortkamp, A. Redundant function of the heparan sulfate 6-O-endosulfatases Sulf1 and Sulf2 during skeletal development. Dev. Dyn. 2008, 237, 339–353. [Google Scholar] [CrossRef]
- Zahraei, M.; Sheikhha, M.H.; Kalantar, S.M.; Ghasemi, N.; Jahaninejad, T.; Rajabi, S.; Mohammadpour, H. The association of arylendosulfatase 1 (SULF1) gene polymorphism with recurrent miscarriage. J. Assist. Reprod. Genet. 2014, 31, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Fan, X.; Wang, R.; Lu, X.; Dang, Y.L.; Wang, H.; Lin, H.Y.; Zhu, C.; Ge, H.; Cross, J.C.; et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res. 2018, 28, 819–832. [Google Scholar] [CrossRef]
- Altmäe, S.; Segura, M.T.; Esteban, F.J.; Bartel, S.; Brandi, P.; Irmler, M.; Beckers, J.; Demmelmair, H.; López-Sabater, C.; Koletzko, B.; et al. Maternal pre-pregnancy obesity is associated with altered placental transcriptome. PLoS ONE 2017, 12, e169223. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, H.; Ma, Y.; Zhu, X.; Zhang, S.; Li, J. MiR-221-3p is down-regulated in preeclampsia and affects trophoblast growth, invasion and migration partly via targeting thrombospondin 2. Biomed. Pharmacother. 2019, 109, 127–134. [Google Scholar] [CrossRef]
- Wang, Q.; Lu, X.; Li, C.; Zhang, W.; Lv, Y.; Wang, L.; Wu, L.; Meng, L.; Fan, Y.; Ding, H.; et al. Down-regulated long non-coding RNA PVT1 contributes to gestational diabetes mellitus and preeclampsia via regulation of human trophoblast cells. Biomed. Pharmacother. 2019, 120, 109501. [Google Scholar] [CrossRef] [PubMed]
- Lip, S.V.; Van Der Graaf, A.M.; Wiegman, M.J.; Scherjon, S.A.; Boekschoten, M.V.; Plösch, T.; Faas, M.M. Experimental preeclampsia in rats affects vascular gene expression patterns. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]
- Matthews, J.C.; Beveridge, M.J.; Malandro, M.S.; Rothstein, J.D.; Campbell-Thompson, M.; Verlander, J.W.; Kilberg, M.S.; Novak, D.A. Activity and protein localization of multiple glutamate transporters in gestation day 14 vs. day 20 rat placenta. Am. J. Physiol. Cell Physiol. 1998, 274, 603–614. [Google Scholar] [CrossRef]
- Huang, X.; Anderle, P.; Hostettler, L.; Baumann, M.U.; Surbek, D.V.; Ontsouka, E.C.; Albrecht, C. Identification of placental nutrient transporters associated with intrauterine growth restriction and pre-eclampsia. BMC Genom. 2018, 19, 173. [Google Scholar] [CrossRef] [PubMed]
- Bergagnini-Kolev, M.C.; Hebert, M.F.; Easterling, T.R.; Lin, Y.S. Pregnancy increases the renal secretion of N1-methylnicotinamide, an endogenous probe for renal cation transporters, in patients prescribed metformin. Drug Metab. Dispos. 2017, 45, 325–329. [Google Scholar] [CrossRef]
- Bottalico, B.; Larsson, I.; Brodszki, J.; Hernandez-Andrade, E.; Casslén, B.; Marsál, K.; Hansson, S. Norepinephrine Transporter (NET), Serotonin Transporter (SERT), Vesicular Monoamine Transporter (VMAT2) and Organic Cation Transporters (OCT1, 2 and EMT) in Human Placenta from Pre-eclamptic and Normotensive Pregnancies. Placenta 2004, 25, 518–529. [Google Scholar] [CrossRef]
- Jansson, T.; Powell, T.L. Human Placental Transport in Altered Fetal Growth: Does the Placenta Function as a Nutrient Sensor?—A Review. Placenta 2006, 27, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Sferruzzi-Perri, A.; Thakali, K.; Lewis, R.; Dilworth, M.R.; Hayward, C.E.; Lean, S.; Sibley, C.P.; Jones, R.L.; Wareing, M.; Greenwood, S.L. Placental Adaptation: What Can We Learn from Birthweight:Placental Weight Ratio? Front. Physiol. 2016, 7, 28. [Google Scholar] [CrossRef]
- Winterhager, E.; Gellhaus, A. Transplacental nutrient transport mechanisms of intrauterine growth restriction in rodent models and humans. Front. Physiol. 2017, 8, 1–13. [Google Scholar] [CrossRef]
- Westermeier, F.; Sáez, P.J.; Villalobos-Labra, R.; Sobrevia, L.; Farías-Jofré, M. Programming of Fetal Insulin Resistance in Pregnancies with Maternal Obesity by ER Stress and Inflammation. BioMed Res. Int. 2014, 2014, 917672. [Google Scholar] [CrossRef]
- Perichart-Perera, O.; Muñoz-Manrique, C.; Reyes-López, A.; Tolentino-Dolores, M.; Espino, Y.; Sosa, S.; Ramírez-González, M.C. Metabolic markers during pregnancy and their association with maternal and newborn weight status. PLoS ONE 2017, 12, e0180874. [Google Scholar] [CrossRef] [PubMed]
- Renaud, J.M.; Yip, K.; Guimond, J.; Trottier, M.; Pibarot, P.; Turcotte, E.; Maguire, C.; Lalonde, L.; Gulenchyn, K.; Farncombe, T.; et al. Characterization of 3-Dimensional PET Systems for Accurate Quantification of Myocardial Blood Flow. J. Nucl. Med. 2017, 58, 103–109. [Google Scholar] [CrossRef]
- Hamilton, S.A.; Tower, C.L.; Jones, R.L. Identification of Chemokines Associated with the Recruitment of Decidual Leukocytes in Human Labour: Potential Novel Targets for Preterm Labour. PLoS ONE 2013, 8, e56946. [Google Scholar] [CrossRef]
- Dunk, C.; Kwan, M.; Hazan, A.; Walker, S.; Wright, J.K.; Harris, L.K.; Jones, R.L.; Keating, S.; Kingdom, J.C.P.; Whittle, W.; et al. Failure of Decidualization and Maternal Immune Tolerance Underlies Uterovascular Resistance in Intra Uterine Growth Restriction. Front. Endocrinol. 2019, 10, 160. [Google Scholar] [CrossRef] [PubMed]
- Kieckbusch, J.; Gaynor, L.M.; Moffett, A.; Colucci, F. MHC-dependent inhibition of uterine NK cells impedes fetal growth and decidual vascular remodelling. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, Z.; Smith, G.N.; Croy, B.A. Natural killer cell-triggered vascular transformation: Maternal care before birth? Cell. Mol. Immunol. 2011, 8, 1–11. [Google Scholar] [CrossRef]
- Vargas-Leal, V.; Bruno, R.; Derfuss, T.; Krumbholz, M.; Hohlfeld, R.; Meinl, E. Expression and Function of Glial Cell Line-Derived Neurotrophic Factor Family Ligands and Their Receptors on Human Immune Cells. J. Immunol. 2005, 175, 2301–2308. [Google Scholar] [CrossRef] [PubMed]
- Rusmini, M.; Griseri, P.; Lantieri, F.; Matera, I.; Hudspeth, K.L.; Roberto, A.; Mikulak, J.; Avanzini, S.; Rossi, V.; Mattioli, G.; et al. Induction of RET Dependent and Independent Pro-Inflammatory Programs in Human Peripheral Blood Mononuclear Cells from Hirschsprung Patients. PLoS ONE 2013, 8, e59066. [Google Scholar] [CrossRef]
- Meinhardt, G.; Saleh, L.; Otti, G.R.; Haider, S.; Velicky, P.; Fiala, C.; Pollheimer, J.; Knöfler, M. Wingless ligand 5a is a critical regulator of placental growth and survival. Sci. Rep. 2016, 6, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.L.; Mark, P.J.; Keelan, J.A.; Barden, A.; Mas, E.; Mori, T.A.; Waddell, B.J. Maternal dietary omega-3 fatty acid intake increases resolvin and protectin levels in the rat placenta. J. Lipid Res. 2013, 54, 2247–2254. [Google Scholar] [CrossRef]
- Spann, N.J.; Glass, C.K. Sterols and oxysterols in immune cell function. Nat. Immunol. 2013, 14, 893–900. [Google Scholar] [CrossRef] [PubMed]
- Willinger, T. Oxysterols in intestinal immunity and inflammation. J. Intern. Med. 2019, 285, 367–380. [Google Scholar] [CrossRef] [PubMed]
- Ness, R.B.; Sibai, B.M. Shared and disparate components of the pathophysiologies of fetal growth restriction and preeclampsia. Am. J. Obstet. Gynecol. 2006, 195, 40–49. [Google Scholar] [CrossRef]
- Ni, L.; Song, C.; Wu, X.; Zhao, X.; Wang, X.; Li, B.; Gan, Y. RNA-seq transcriptome profiling of porcine lung from two pig breeds in response to Mycoplasma hyopneumoniae infection. PeerJ 2019, 2019, e7900. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Zhang, Q.; Tan, D.; Luo, W.; Zhao, H.; Ma, J.; Liang, H.; Tan, Y. GABA A receptor subunit promotes apoptosis of HTR-8/SVneo trophoblastic cells: Implications in preeclampsia. Int. J. Mol. Med. 2016, 38, 105–112. [Google Scholar] [CrossRef] [PubMed]
- de Barros Mucci, D.; Kusinski, L.C.; Wilsmore, P.; Loche, E.; Pantaleão, L.C.; Ashmore, T.J.; Blackmore, H.L.; Fernandez-Twinn, D.S.; Maria das Graças, T.; Ozanne, S.E. Impact of maternal obesity on placental transcriptome and morphology associated with fetal growth restriction in mice. Int. J. Obes. 2020. [Google Scholar] [CrossRef]
- Rossi, M.N.; Pascarella, A.; Licursi, V.; Caiello, I.; Taranta, A.; Rega, L.R.; Levtchenko, E.; Emma, F.; De Benedetti, F.; Prencipe, G. NLRP2 Regulates Proinflammatory and Antiapoptotic Responses in Proximal Tubular Epithelial Cells. Front. Cell Dev. Biol. 2019, 7, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Mahadevan, S.; Sathappan, V.; Utama, B.; Lorenzo, I.; Kaskar, K.; Van den Veyver, I.B. Maternally expressed NLRP2 links the subcortical maternal complex (SCMC) to fertility, embryogenesis and epigenetic reprogramming. Sci. Rep. 2017, 7, 44667. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Deng, N.; Lei, D.; Huang, J.; Yang, Z.; Fan, C.; Wang, S. Circular RNA expression profiling identifies hsa_circ_0011460 as a novel molecule in severe preeclampsia. Pregnancy Hypertens. 2019, 17, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Ying, W.; Li, F.J.; Wei, S.W.; Li, W.L. Genomic imprinting status of IGF-II and H19 in placentas of fetal growth restriction patients. J. Genet. 2010, 89, 213–216. [Google Scholar] [CrossRef] [PubMed]
- Zuckerwise, L.; Li, J.; Lu, L.; Men, Y.; Geng, T.; Buhimschi, C.S.; Buhimschi, I.A.; Bukowski, R.; Guller, S.; Paidas, M.; et al. H19 Long Noncoding RNA Alters Trophoblast Cell Migration and Invasion by Regulating TβR3 in Placentae with Fetal Growth Restriction. Oncotarget 2016, 7, 38398–38407. [Google Scholar] [CrossRef]
- Buhimschi, I.A.; Nayeri, U.A.; Zhao, G.; Shook, L.L.; Pensalfini, A.; Funai, E.F.; Bernstein, I.M.; Glabe, C.G.; Buhimschi, C.S. Defective Amyloid Processing in Preeclampsia. Sci. Transl. Med. 2014, 92. [Google Scholar] [CrossRef]
- Galbiati, S.; Inversetti, A.; Causarano, V.; Stenirri, S.; Soriani, N.; Ambrosi, A.; Valsecchi, L.; Candiani, M.; Cremonesi, L.; Ferrari, M.; et al. HIF1A and MIF as potential predictive mRNA biomarkers of pre-eclampsia: A longitudinal prospective study in high risk population. Clin. Chem. Lab. Med. 2015, 53, 1339–1347. [Google Scholar] [CrossRef] [PubMed]
- Dunk, C.E.; Roggensack, A.M.; Cox, B.; Perkins, J.E.; Senius, F.; Keating, S.; Weksberg, R.; Kingdom, J.C.P.; Adamson, S.L. A distinct microvascular endothelial gene expression profile in severe IUGR placentas. Placenta 2012, 33, 285–293. [Google Scholar] [CrossRef]
- McKay, J.; Xie, L.; Adriaens, M.; Evelo, C.; Ford, D.; Mathers, J. Organ-Specific Gene Expression Changes in the Fetal Liver and Placenta in Response to Maternal Folate Depletion. Nutrients 2016, 8, 661. [Google Scholar] [CrossRef]
| Biosample Number | Fetus Sex | Fetus Weight [g] | Week of Gestation | Apgar Score | Maternal Age | FGR Stage * |
|---|---|---|---|---|---|---|
| ERX4055529 | Male | 750 | 26 | 2 | 31 | 4 |
| ERX4055530 | Male | 1680 | 30 | 2 | 37 | 3 |
| ERX4055531 | Male | 2060 | 39 | 10 | 28 | 1 |
| ERX4055532 | Female | 2360 | 36 | 10 | 28 | 1 |
| ERX4055533 | Female | 2260 | 37 | 10 | 24 | 1 |
| ERX4055534 | Female | 2000 | 37 | 10 | 27 | 1 |
| ERX4055535 | Male | 3230 | 37 | 10 | 35 | - |
| ERX4055536 | Male | 3420 | 39 | 10 | 37 | - |
| ERX4055537 | Male | 3500 | 39 | 10 | 31 | - |
| ERX4055538 | Female | 2960 | 39 | 10 | 23 | - |
| ERX4055539 | Female | 3300 | 38 | 10 | 29 | - |
| ERX4055540 | Female | 3100 | 38 | 10 | 30 | - |
| Transcript Biotypes | Transcripts |
|---|---|
| antisense_RNA | 8295 |
| lincRNA | 7849 |
| processed_transcript | 484 |
| retained_intron | 441 |
| sense_intronic | 657 |
| sense_overlapping | 250 |
| TEC | 722 |
| others | 176 |
| Gene ID | Gene Name | Log2 Fold Change | ENSEMBL ID | lncRNA |
|---|---|---|---|---|
| XLOC_000619 | - | Inf | NA | - |
| XLOC_006184 | ACTA2-AS1 | −3.55 | ENSG00000180139 | known |
| XLOC_006962 | - | -Inf | NA | - |
| XLOC_013293 | MYO1A,TAC3 | −4.60 | ENSG00000166866, ENSG00000166863 | - |
| XLOC_013509 | PPFIA2 | −4.98 | ENSG00000139220 | - |
| XLOC_019012 | ACTC1 | −6.02 | ENSG00000159251 | - |
| XLOC_019742 | - | -Inf | NA | - |
| XLOC_023789 | CAMKK1,P2RX1 | −2.88 | ENSG00000004660, ENSG00000108405 | - |
| XLOC_023922 | MYH3 | −3.55 | ENSG00000109063 | - |
| XLOC_040198 | - | Inf | NA | - |
| XLOC_042821 | PDLIM3 | −2.30 | ENSG00000154553 | - |
| XLOC_044217 | SGCD | −3.63 | ENSG00000170624 | - |
| XLOC_048901 | THBS2 | −3.24 | ENSG00000186340 | - |
| XLOC_051439 | TMEM130 | −4.53 | ENSG00000166448 | - |
| XLOC_052796 | SULF1 | −4.39 | ENSG00000137573 | - |
| XLOC_053187 | MIR1204,PVT1, PVT1_1,PVT1_3 | 3.38 | ENSG00000283710, ENSG00000249859, ENSG00000276443, ENSG00000278324 | -, novel, known |
| XLOC_054578 | - | -Inf | NA | - |
| XLOC_054962 | ALDH1B1 | −3.37 | ENSG00000137124 | - |
| XLOC_056099 | ANKRD18A,FAM95C | −2.75 | ENSG00000180071, ENSG00000225345, ENSG00000250989, ENSG00000273036, ENSG00000272934, ENSG00000272904, ENSG00000283486 | -, novel, known |
| Term Name | Term ID | Source | p-Value * | Gene Names |
|---|---|---|---|---|
| muscle contraction | GO:0006936 | BP | 3.3074 × 10−3 | P2RX1,SULF1,ACTC1,SGCD,MYH3 |
| muscle system process | GO:0003012 | BP | 1.2096 × 10−2 | P2RX1,SULF1,ACTC1,SGCD,MYH3 |
| actin filament | GO:0005884 | CC | 8.5988 × 10−3 | PDLIM3,ACTC1,MYO1A |
| filamentous actin | GO:0031941 | CC | 3.7185 × 10−2 | PDLIM3,MYO1A |
| actin cytoskeleton | GO:0015629 | CC | 4.0095 × 10−2 | PDLIM3,ACTC1,MYO1A,MYH3 |
| sarcomere | GO:0030017 | CC | 4.2036 × 10−2 | PDLIM3,ACTC1,MYH3 |
| carbohydrate derivative binding | GO:0097367 | MF | 4.4875 × 10−2 | CAMKK1,P2RX1,SULF1,ACTC1,MYO1A,THBS2,MYH3 |
| lncRNA | DEG | lncRNA Name | DEG Name | Correlation | p-Value |
|---|---|---|---|---|---|
| XLOC_006184 | XLOC_019012 | ACTA2-AS1 | ACTC1 | 0.960 | 2.3481 × 10−3 |
| XLOC_006184 | XLOC_042821 | ACTA2-AS1 | PDLIM3 | 0.948 | 4.0051 × 10−3 |
| XLOC_006184 | XLOC_044217 | ACTA2-AS1 | SGCD | 0.943 | 4.8411 × 10−3 |
| XLOC_006184 | XLOC_052796 | ACTA2-AS1 | SULF1 | 0.933 | 6.6462 × 10−3 |
| XLOC_006184 | XLOC_054962 | ACTA2-AS1 | ALDH1B1 | 0.976 | 8.6087 × 10−4 |
| Sample Set | Chr | Start | End | Host Gene | Strand | Region |
|---|---|---|---|---|---|---|
| ff-FGR | 1 | 629,675 | 629,725 | AL669831.3 | - | intron |
| ff-FGR | 1 | 207,336 713 | 207,336,763 | CD55 | + | exon 7 |
| ff-FGR | 11 | 1,997,400 | 1,997,475 | H19 | - | exon 1 |
| ff-FGR | 11 | 1,997,424 | 1,997,475 | H19 | - | exon 1 |
| ff-FGR | 11 | 1,997,697 | 1,997,767 | H19 | - | exon 1 |
| ff-FGR | KI270721.1 | 52,582 | 52,657 | not annotated | - | intergenic |
| ff-FGR | KI270721.1 | 52,606 | 52,657 | not annotated | + | intergenic |
| mf-FGR | 1 | 629,675 | 629,725 | AL669831.3 | - | intron |
| mf-FGR | 11 | 1,997,424 | 1,997,475 | H19 | - | exon 1 |
| mf-FGR | 21 | 41,178,852 | 41,178,960 | BACE2 | + | intron |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lipka, A.; Jastrzebski, J.P.; Paukszto, L.; Makowczenko, K.G.; Lopienska-Biernat, E.; Gowkielewicz, M.; Lepiarczyk, E.; Wiszpolska, M.; Majewski, M.K.; Majewska, M. Sex-Biased lncRNA Signature in Fetal Growth Restriction (FGR). Cells 2021, 10, 921. https://doi.org/10.3390/cells10040921
Lipka A, Jastrzebski JP, Paukszto L, Makowczenko KG, Lopienska-Biernat E, Gowkielewicz M, Lepiarczyk E, Wiszpolska M, Majewski MK, Majewska M. Sex-Biased lncRNA Signature in Fetal Growth Restriction (FGR). Cells. 2021; 10(4):921. https://doi.org/10.3390/cells10040921
Chicago/Turabian StyleLipka, Aleksandra, Jan Pawel Jastrzebski, Lukasz Paukszto, Karol Gustaw Makowczenko, Elzbieta Lopienska-Biernat, Marek Gowkielewicz, Ewa Lepiarczyk, Marta Wiszpolska, Mariusz Krzysztof Majewski, and Marta Majewska. 2021. "Sex-Biased lncRNA Signature in Fetal Growth Restriction (FGR)" Cells 10, no. 4: 921. https://doi.org/10.3390/cells10040921
APA StyleLipka, A., Jastrzebski, J. P., Paukszto, L., Makowczenko, K. G., Lopienska-Biernat, E., Gowkielewicz, M., Lepiarczyk, E., Wiszpolska, M., Majewski, M. K., & Majewska, M. (2021). Sex-Biased lncRNA Signature in Fetal Growth Restriction (FGR). Cells, 10(4), 921. https://doi.org/10.3390/cells10040921







