Piezo1 Channels Contribute to the Regulation of Human Atrial Fibroblast Mechanical Properties and Matrix Stiffness Sensing
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.1.1. Cell Types and Maintenance
2.1.2. Transfection
2.1.3. Compounds and Treatments
2.2. Cell Culture Matrices with Different Stiffness
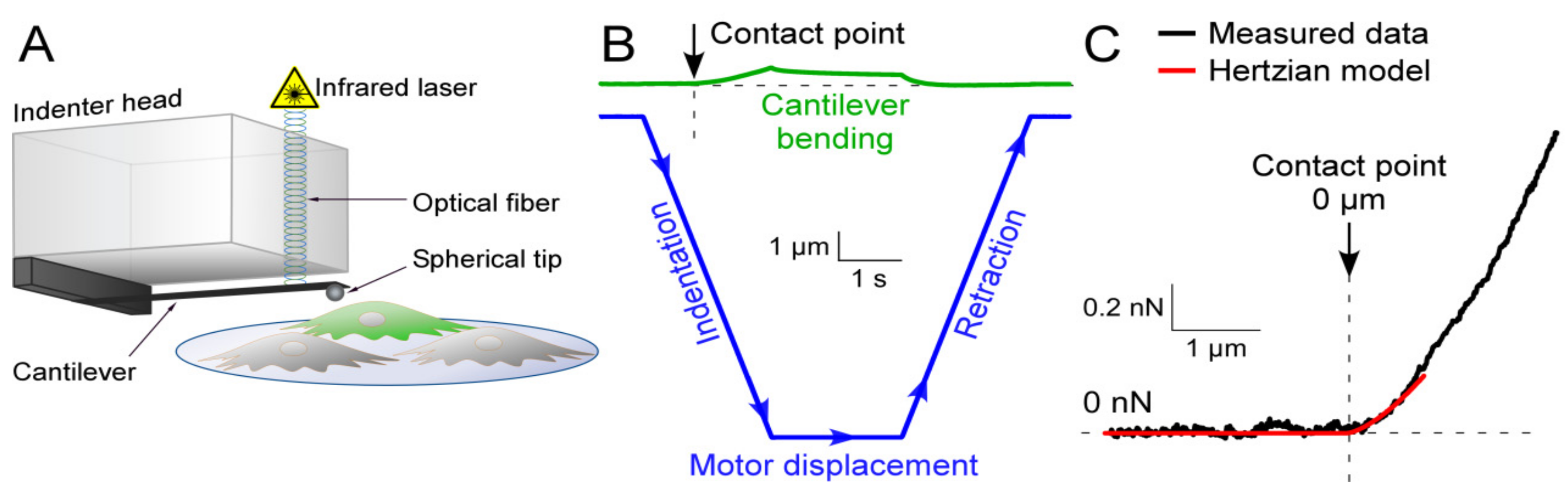
2.3. Nanoindentation
2.4. Cytoskeleton: Staining, Image Acquisition, and Data Analysis
2.5. Gene Expression Analysis
2.6. IL-6 Measurements by Enzyme-Linked Immunosorbent Assay
2.7. Piezo1 Protein Level Detected by Western Blot
2.8. Patch-Clamp Recording of Piezo1 Activity
2.9. Statistical Analyses
3. Results
3.1. Piezo1 Overexpression in HEK Cells Leads to Increased Cell Stiffness
3.2. PiCS Does Not Require Ion Flux Through the Channel, Nor Calpain Activity in HEK Cells
3.3. PiCS in HEK Cells Requires Components of the Integrin Signaling Pathway
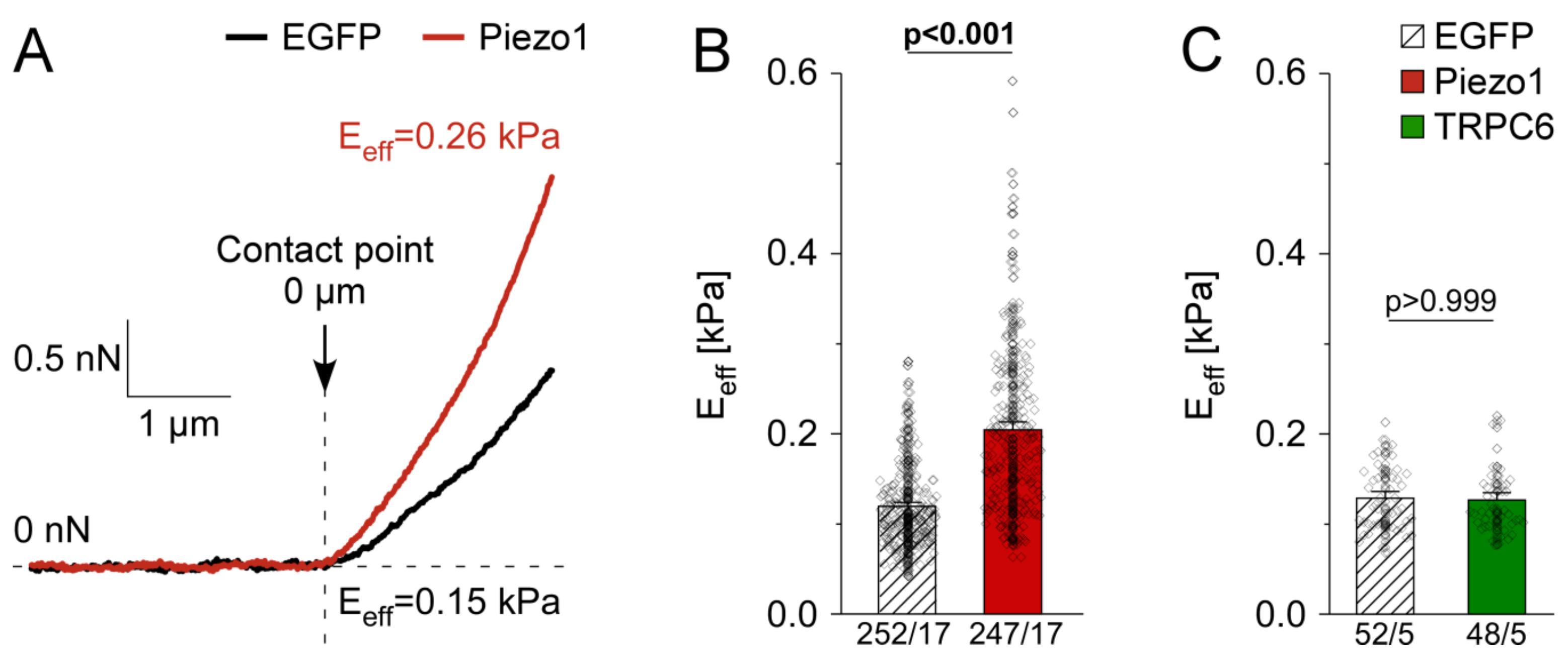
3.4. PiCS Is Conserved in HAF
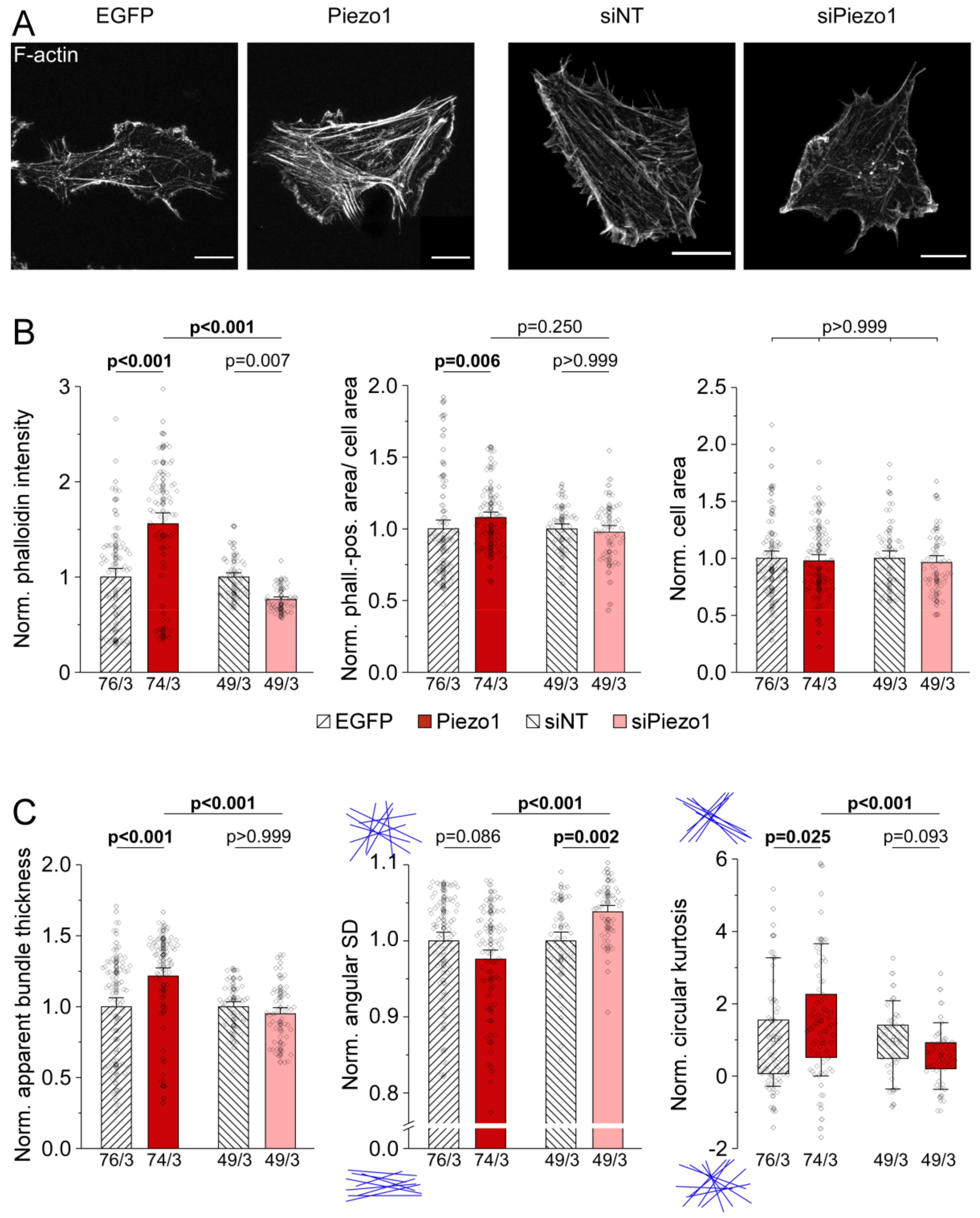
3.5. Piezo1 Expression Affects Architecture of the Actin Cytoskeleton in HAF
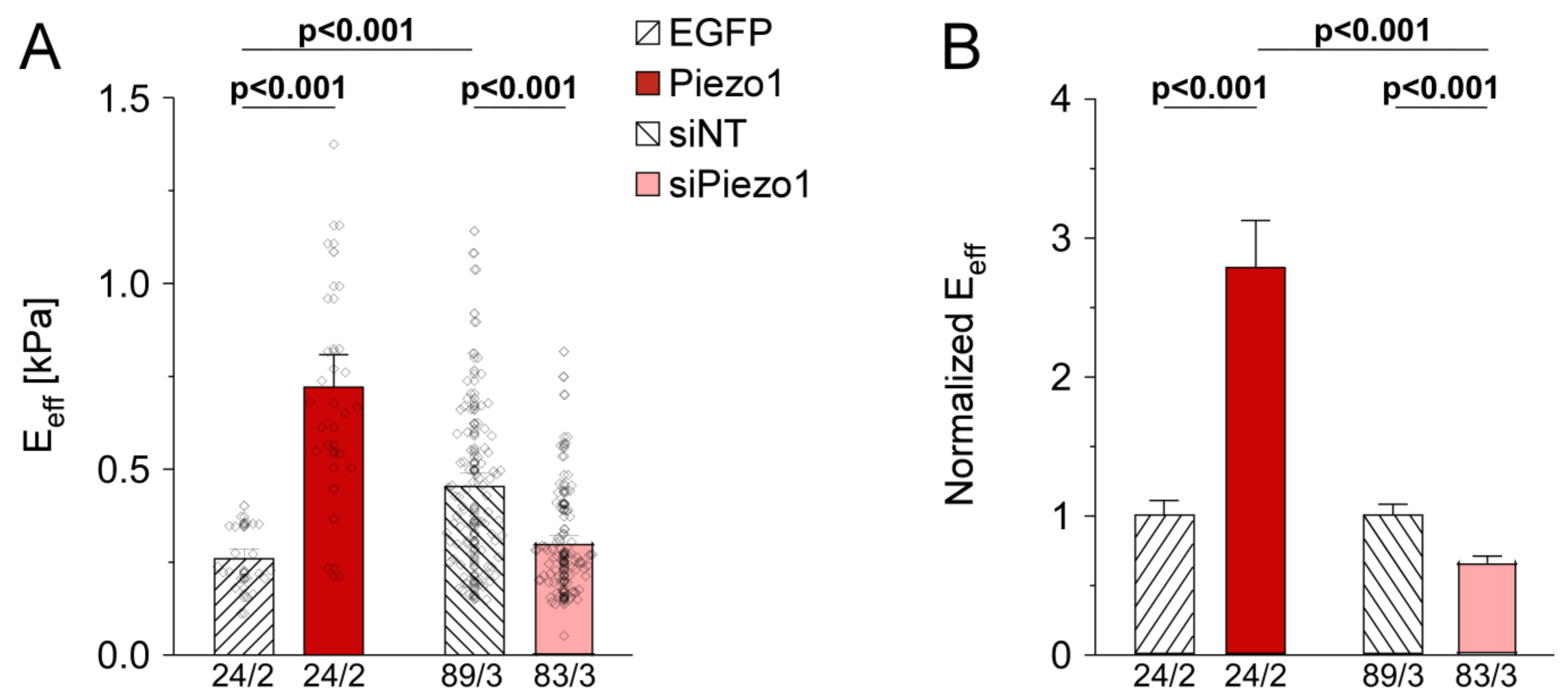
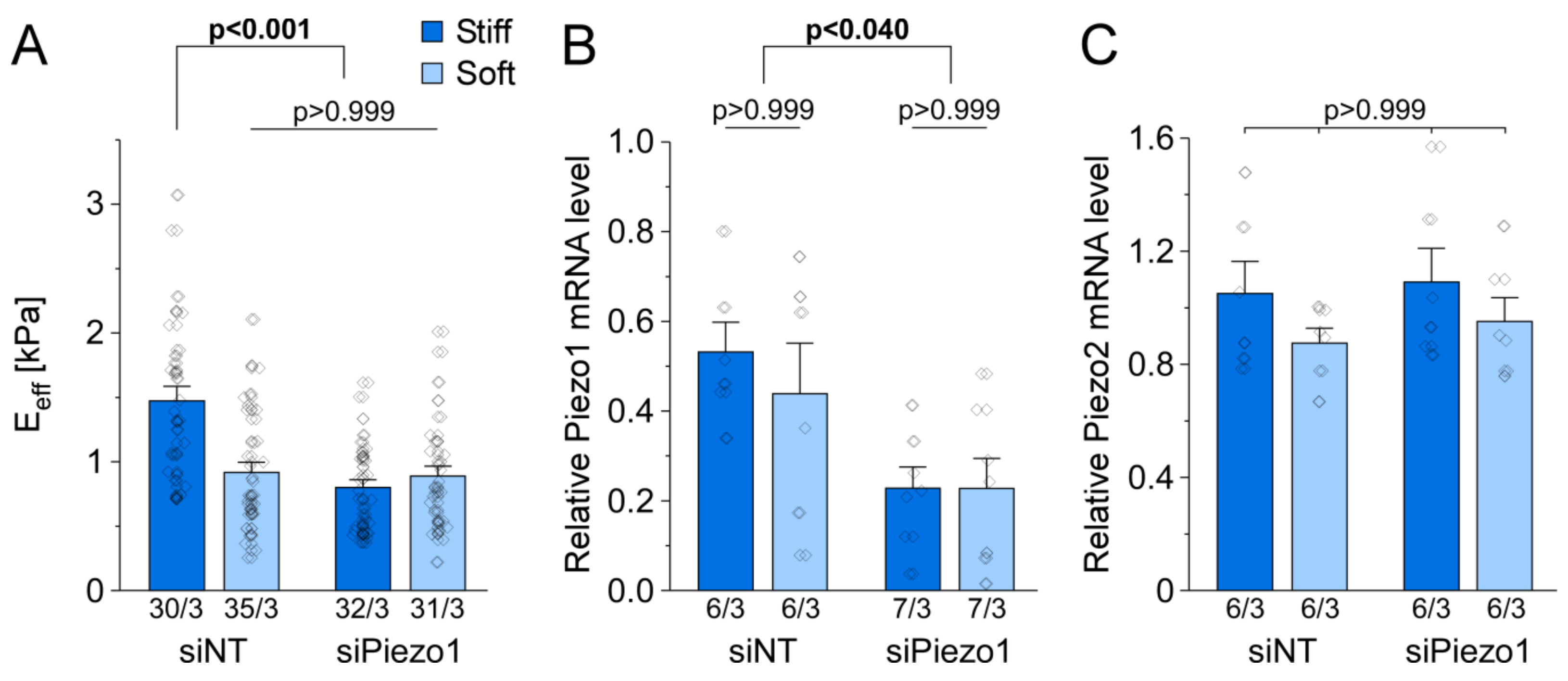
3.6. Piezo1 Is Required for Cell Stiffness Adaptation to Matrix Stiffness in HAF
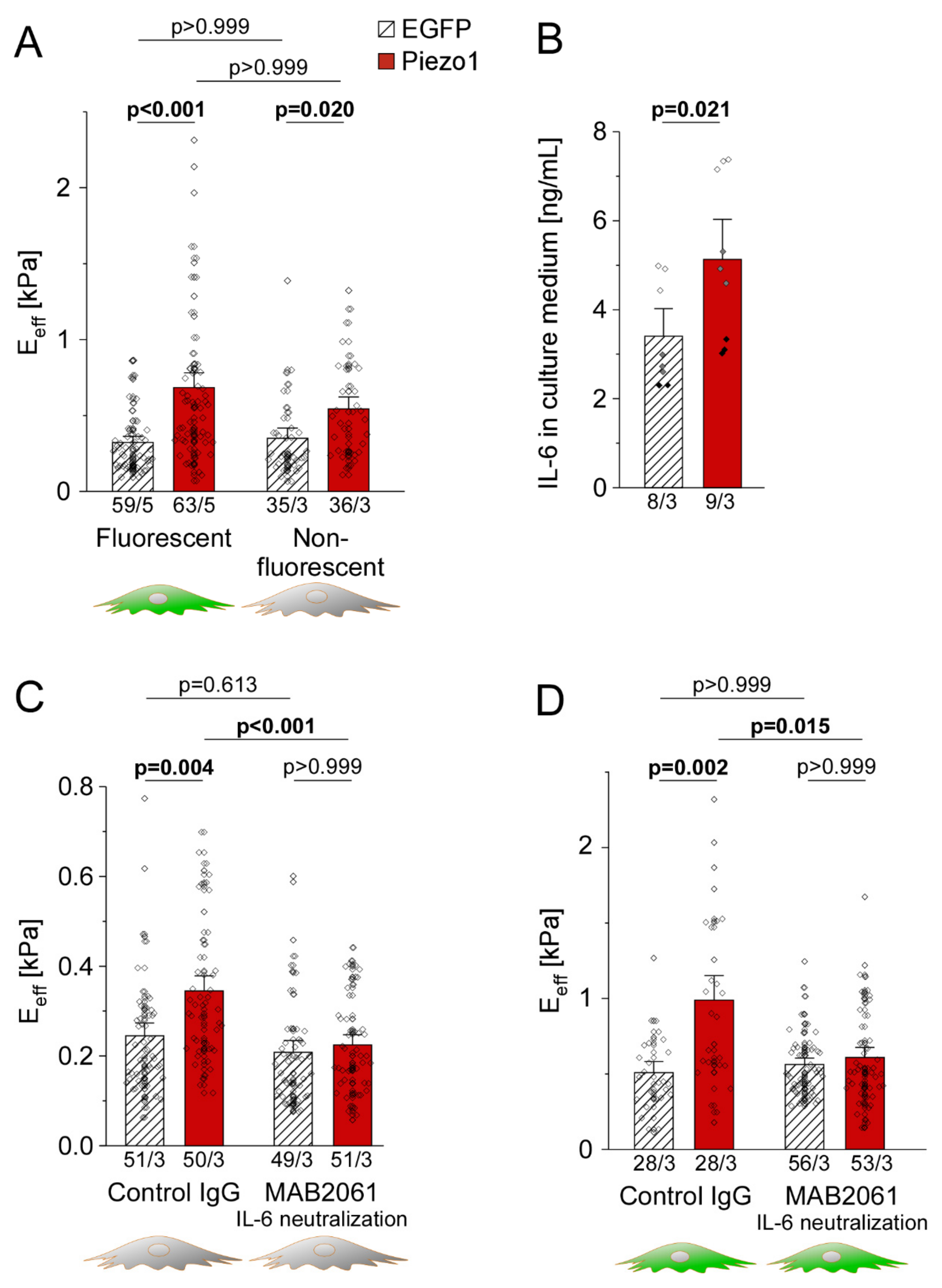
3.7. PiCS Is Transmitted to Neighboring HAF by IL-6
4. Discussion
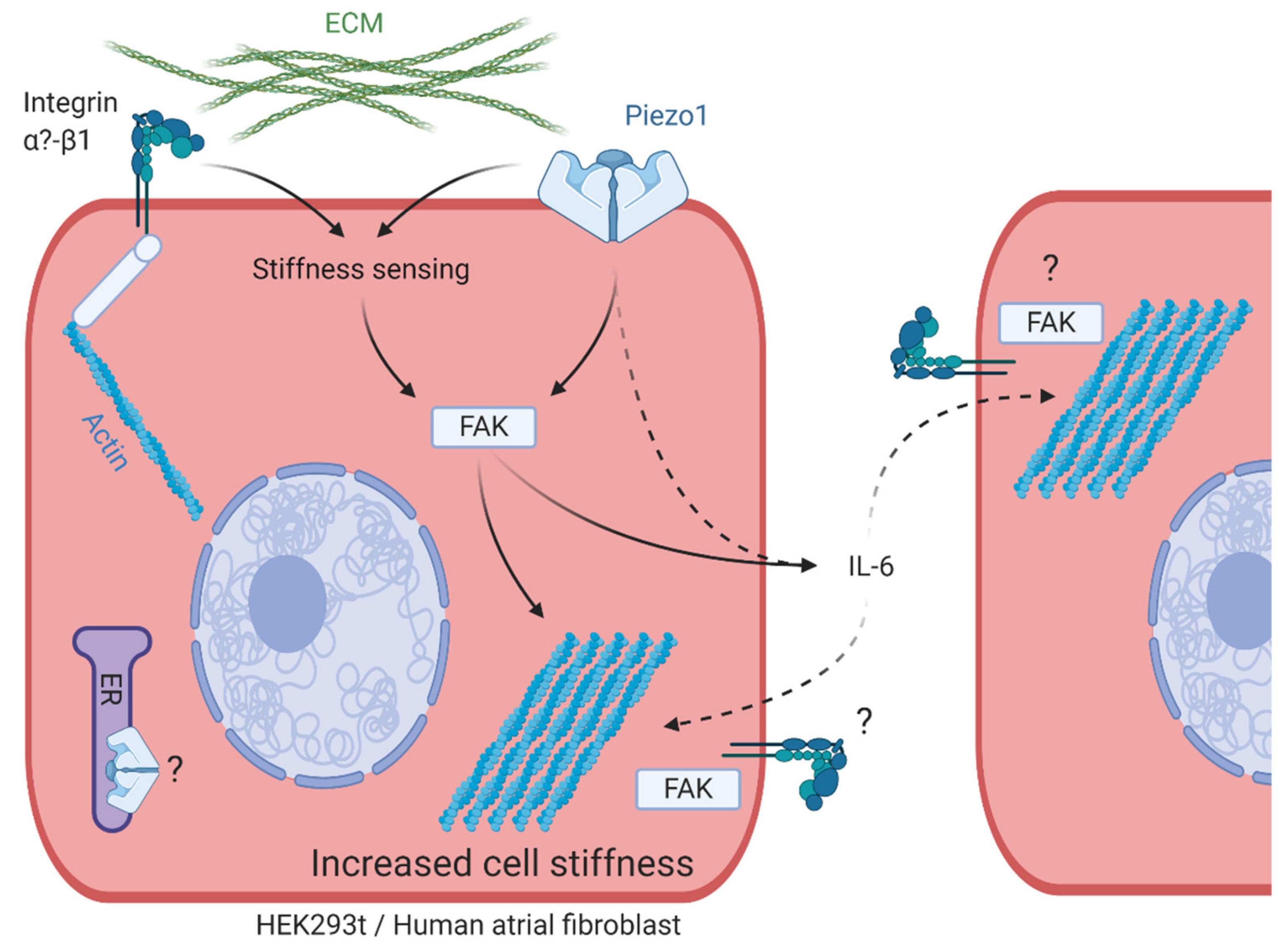
4.1. Mechanism Underlying PiCS
4.2. PiCS in HAF and Beyond
4.3. Relevance of PiCS in the Context of Altered Environmental Stiffness
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hinz, B.; McCulloch, C.A.; Coelho, N.M. Mechanical regulation of myofibroblast phenoconversion and collagen contraction. Exp. Cell Res. 2019, 379, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Wells, R.G. Tissue mechanics and fibrosis. Biochim. et Biophys. Acta (BBA) Mol. Basis Dis. 2013, 1832, 884–890. [Google Scholar] [CrossRef] [PubMed]
- Forrester, J.S.; Diamond, G.; Parmley, W.W.; Swan, H.J.C. Early Increase in Left Ventricular Compliance after Myocardial Infarction. J. Clin. Investig. 1972, 51, 598–603. [Google Scholar] [CrossRef] [PubMed]
- Holmes, J.W.; Borg, T.K.; Covell, J.W. Structure and Mechanics of Healing Myocardial Infarcts. Annu. Rev. Biomed. Eng. 2005, 7, 223–253. [Google Scholar] [CrossRef]
- Farré, N.; Jorba, I.; Torres, M.; Falcones, B.; Martí-Almor, J.; Farré, R.; Almendros, I.; Navajas, D. Passive Stiffness of Left Ventricular Myocardial Tissue Is Reduced by Ovariectomy in a Post-menopause Mouse Model. Front. Physiol. 2018, 9. [Google Scholar] [CrossRef]
- Berry, M.F.; Engler, A.J.; Woo, Y.J.; Pirolli, T.J.; Bish, L.T.; Jayasankar, V.; Morine, K.J.; Gardner, T.J.; Discher, D.E.; Sweeney, H.L. Mesenchymal stem cell injection after myocardial infarction improves myocardial compliance. Am. J. Physiol. Circ. Physiol. 2006, 290, H2196–H2203. [Google Scholar] [CrossRef]
- Gluck, J.M.; Herren, A.W.; Yechikov, S.; Kao, H.K.J.; Khan, A.; Phinney, B.S.; Chiamvimonvat, N.; Chan, J.W.; Lieu, D.K. Biochemical and biomechanical properties of the pacemaking sinoatrial node extracellular matrix are distinct from contractile left ventricular matrix. PLoS ONE 2017, 12, e0185125. [Google Scholar] [CrossRef]
- Gaetani, R.; Zizzi, E.A.; Deriu, M.A.; Morbiducci, U.; Pesce, M.; Messina, E. When Stiffness Matters: Mechanosensing in Heart Development and Disease. Front. Cell Dev. Biol. 2020, 8, 334. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, J.D.; Dufresne, E.R.; Schwartz, M.A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 2014, 15, 802–812. [Google Scholar] [CrossRef]
- Hamidi, H.; Ivaska, J. Every step of the way: Integrins in cancer progression and metastasis. Nat. Rev. Drug Discov. 2018, 17, 31–46. [Google Scholar] [CrossRef]
- Legate, K.R.; Wickström, S.A.; Fässler, R. Genetic and cell biological analysis of integrin outside-in signaling. Genes Dev. 2009, 23, 397–418. [Google Scholar] [CrossRef]
- Goult, B.T.; Yan, J.; Schwartz, M.A. Talin as a mechanosensitive signaling hub. J. Cell Biol. 2018, 217, 3776–3784. [Google Scholar] [CrossRef]
- Mitra, S.K.; Hanson, D.A.; Schlaepfer, D.D. Focal adhesion kinase: In command and control of cell motility. Nat. Rev. Mol. Cell Biol. 2005, 6, 56–68. [Google Scholar] [CrossRef]
- Campbell, I.D.; Humphries, M.J. Integrin Structure, Activation, and Interactions. Cold Spring Harb. Perspect. Biol. 2011, 3, a004994. [Google Scholar] [CrossRef]
- Sun, Z.; Guo, S.S.; Fässler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 2016, 215, 445–456. [Google Scholar] [CrossRef]
- Chen, C.; Li, R.; Ross, R.S.; Manso, A.M. Integrins and integrin-related proteins in cardiac fibrosis. J. Mol. Cell. Cardiol. 2016, 93, 162–174. [Google Scholar] [CrossRef]
- Farmer, L.K.; Rollason, R.; Whitcomb, D.J.; Ni, L.; Goodliff, A.; Lay, A.C.; Birnbaumer, L.; Heesom, K.J.; Xu, S.-Z.; Saleem, M.A.; et al. TRPC6 Binds to and Activates Calpain, Independent of Its Channel Activity, and Regulates Podocyte Cytoskeleton, Cell Adhesion, and Motility. J. Am. Soc. Nephrol. 2019, 30, 1910–1924. [Google Scholar] [CrossRef]
- Coste, B.; Mathur, J.; Schmidt, M.; Earley, T.J.; Ranade, S.; Petrus, M.J.; Dubin, A.E.; Patapoutian, A. Piezo1 and Piezo2 Are Essential Components of Distinct Mechanically Activated Cation Channels. Science 2010, 330, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Coste, B.; Xiao, B.; Santos, J.S.; Syeda, R.; Grandl, J.; Spencer, K.S.; Kim, S.E.; Schmidt, M.; Mathur, J.; Dubin, A.E.; et al. Piezo proteins are pore-forming subunits of mechanically activated channels. Nat. Cell Biol. 2012, 483, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Murthy, S.E.; Dubin, A.E.; Patapoutian, A. Piezos thrive under pressure: Mechanically activated ion channels in health and disease. Nat. Rev. Mol. Cell Biol. 2017, 18, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Douguet, D.; Patel, A.; Xu, A.; Vanhoutte, P.M.; Honoré, E. Piezo Ion Channels in Cardiovascular Mechanobiology. Trends Pharmacol. Sci. 2019, 40, 956–970. [Google Scholar] [CrossRef] [PubMed]
- Beech, D.J.; Kalli, A.C. Force Sensing by Piezo Channels in Cardiovascular Health and Disease. Arter. Thromb. Vasc. Biol. 2019, 39, 2228–2239. [Google Scholar] [CrossRef]
- Ridone, P.; Vassalli, M.; Martinac, B. Piezo1 mechanosensitive channels: What are they and why are they important. Biophys. Rev. 2019, 11, 795–805. [Google Scholar] [CrossRef]
- Blythe, N.M.; Muraki, K.; Ludlow, M.J.; Stylianidis, V.; Gilbert, H.T.; Evans, E.L.; Cuthbertson, K.; Foster, R.; Swift, J.; Li, J.; et al. Mechanically activated Piezo1 channels of cardiac fibroblasts stimulate p38 mitogen-activated protein kinase activity and interleukin-6 secretion. J. Biol. Chem. 2019, 294, 17395–17408. [Google Scholar] [CrossRef]
- Zhao, Q.; Zhou, H.; Chi, S.; Wang, Y.; Wang, J.; Geng, J.; Wu, K.; Liu, W.; Zhang, T.; Dong, M.-Q. Structure and mechanogating mechanism of the piezo1 channel. Nature 2018, 554, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wanggou, S.; Bodalia, A.; Zhu, M.; Dong, W.; Fan, J.J.; Yin, W.C.; Min, H.-K.; Hu, M.; Draghici, D.; et al. A Feedforward Mechanism Mediated by Mechanosensitive Ion Channel PIEZO1 and Tissue Mechanics Promotes Glioma Aggression. Neuron 2018, 100, 799–815.e7. [Google Scholar] [CrossRef]
- Pathak, M.M.; Nourse, J.L.; Tran, T.; Hwe, J.; Arulmoli, J.; Le, D.T.T.; Bernardis, E.; Flanagan, L.A.; Tombola, F. Stretch-activated ion channel Piezo1 directs lineage choice in human neural stem cells. Proc. Natl. Acad. Sci. USA 2014, 111, 16148–16153. [Google Scholar] [CrossRef] [PubMed]
- Ellefsen, K.L.; Holt, J.R.; Chang, A.C.; Nourse, J.L.; Arulmoli, J.; Mekhdjian, A.H.; Abuwarda, H.; Tombola, F.; Flanagan, L.A.; Dunn, A.R. Myosin-ii mediated traction forces evoke localized piezo1-dependent ca 2+ flickers. Commun. Biol. 2019, 2, 1–13. [Google Scholar] [CrossRef] [PubMed]
- McHugh, B.J.; Buttery, R.; Lad, Y.; Banks, S.; Haslett, C.; Sethi, T. Integrin activation by Fam38A uses a novel mechanism of R-Ras targeting to the endoplasmic reticulum. J. Cell Sci. 2009, 123, 51–61. [Google Scholar] [CrossRef]
- Gaub, B.M.; Müller, D.J. Mechanical Stimulation of Piezo1 Receptors Depends on Extracellular Matrix Proteins and Directionality of Force. Nano Lett. 2017, 17, 2064–2072. [Google Scholar] [CrossRef]
- Forget, A.; Gianni-Barrera, R.; Uccelli, A.; Sarem, M.; Kohler, E.; Fogli, B.; Muraro, M.G.; Bichet, S.; Aumann, K.; Banfi, A. Mechanically defined microenvironment promotes stabilization of microvasculature, which correlates with the enrichment of a novel piezo-1+ population of circulating cd11b+/cd115+ monocytes. Adv. Mater. 2019, 31, 1808050. [Google Scholar] [CrossRef]
- Velasco-Estevez, M.; Mampay, M.; Boutin, H.; Chaney, A.; Warn, P.; Sharp, A.; Burgess, E.; Moeendarbary, E.; Dev, K.K.; Sheridan, G.K. Infection Augments Expression of Mechanosensing Piezo1 Channels in Amyloid Plaque-Reactive Astrocytes. Front. Aging Neurosci. 2018, 10, 332. [Google Scholar] [CrossRef]
- Zarychanski, R.; Schulz, V.P.; Houston, B.L.; Maksimova, Y.; Houston, D.S.; Smith, B.; Rinehart, J.; Gallagher, P.G. Mutations in the mechanotransduction protein PIEZO1 are associated with hereditary xerocytosis. Blood 2012, 120, 1908–1915. [Google Scholar] [CrossRef]
- Ceccato, T.L.; Starbuck, R.B.; Hall, J.K.; Walker, C.J.; Brown, T.E.; Killgore, J.P.; Anseth, K.S.; Leinwand, L.A. Defining the Cardiac Fibroblast Secretome in a Fibrotic Microenvironment. J. Am. Hear. Assoc. 2020, 9. [Google Scholar] [CrossRef]
- Jakob, D.A.K.; Allegrini, B.; Darkow, E.; Aria, D.; Emig, R.; Chica, A.S.; Rog-Zielinska, E.A.; Guth, T.; Beyersdorf, F.; Kari, F.A.; et al. Piezo1 and BKca channels in human atrial fibroblasts: Interplay and remodelling in atrial fibrillation. bioRxiv 2021. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kant, T.A.; Emig, R.; Rausch, J.S.E.; Newe, M.; Schubert, M.; Künzel, K.; Winter, L.; Klapproth, E.; Peyronnet, R.; et al. Repurposing mesalazine against cardiac fibrosis in vitro. Naunyn-Schmiedeberg’s Arch. Pharmacol. 2021, 394, 533–543. [Google Scholar] [CrossRef]
- Peyronnet, R.; Martins, J.R.; Duprat, F.; Demolombe, S.; Arhatte, M.; Jodar, M.; Tauc, M.; Duranton, C.; Paulais, M.; Teulon, J. Piezo1-dependent stretch-activated channels are inhibited by polycystin-2 in renal tubular epithelial cells. EMBO Rep. 2013, 14, 1143–1148. [Google Scholar] [CrossRef] [PubMed]
- Okuyama, H.; Hori, R.; Nishida, T. Hydrophilic and Hydrophobic Compounds Antithetically Affect the Growth of Eicosapentaenoic Acid-Synthesizing Escherichia coli Recombinants. Open Microbiol. J. 2011, 5, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Vaudaux, P.; Waldvogel, F.A. Gentamicin antibacterial activity in the presence of human polymorphonuclear leukocytes. Antimicrob. Agents Chemother. 1979, 16, 743–749. [Google Scholar] [CrossRef] [PubMed]
- Vinitsky, A.; Michaud, C.; Powers, J.C.; Orlowski, M. Inhibition of the chymotrypsin-like activity of the pituitary multicatalytic proteinase complex. Biochemistry 1992, 31, 9421–9428. [Google Scholar] [CrossRef]
- Gahmberg, C.G.; Fagerholm, S.C.; Nurmi, S.M.; Chavakis, T.; Marchesan, S.; Grönholm, M. Regulation of integrin activity and signalling. Biochim. et Biophys. Acta (BBA) Gen. Subj. 2009, 1790, 431–444. [Google Scholar] [CrossRef] [PubMed]
- Hörner, M.; Raute, K.; Hummel, B.; Madl, J.; Creusen, G.; Thomas, O.S.; Christen, E.H.; Hotz, N.; Gübeli, R.J.; Engesser, R.; et al. Phytochrome-Based Extracellular Matrix with Reversibly Tunable Mechanical Properties. Adv. Mater. 2019, 31, e1806727. [Google Scholar] [CrossRef] [PubMed]
- Hörner, M.; Hoess, P.; Emig, R.; Rebmann, B.; Weber, W. Synthesis of a light-controlled phytochrome-based extracellular matrix with reversibly adjustable mechanical properties. In Photoswitching Proteins; Springer: Berlin/Heidelberg, Germany, 2020; pp. 217–231. [Google Scholar] [CrossRef]
- Emig, R.; Zgierski-Johnston, C.M.; Beyersdorf, F.; Rylski, B.; Ravens, U.; Weber, W.; Kohl, P.; Hörner, M.; Peyronnet, R. Human Atrial Fibroblast Adaptation to Heterogeneities in Substrate Stiffness. Front. Physiol. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Hertz, H. Ueber die Beruehrung fester elastischer Koerper. Angew. Math. 1881, 92, 156–171. [Google Scholar] [CrossRef]
- Guz, N.; Dokukin, M.; Kalaparthi, V.; Sokolov, I. If Cell Mechanics Can Be Described by Elastic Modulus: Study of Different Models and Probes Used in Indentation Experiments. Biophys. J. 2014, 107. [Google Scholar] [CrossRef]
- Wu, C.-E.; Lin, K.-H.; Juang, J.-Y. Hertzian load–displacement relation holds for spherical indentation on soft elastic solids undergoing large deformations. Tribol. Int. 2016, 97, 71–76. [Google Scholar] [CrossRef]
- Luo, Q.; Kuang, D.; Zhang, B.; Song, G. Cell stiffness determined by atomic force microscopy and its correlation with cell motility. Biochim. et Biophys. Acta (BBA) Gen. Subj. 2016, 1860, 1953–1960. [Google Scholar] [CrossRef]
- Liu, H.; Sun, Y.; Simmons, C.A. Determination of local and global elastic moduli of valve interstitial cells cultured on soft substrates. J. Biomech. 2013, 46, 1967–1971. [Google Scholar] [CrossRef]
- Nguyen, D.T.; Nagarajan, N.; Zorlutuna, P. Effect of Substrate Stiffness on Mechanical Coupling and Force Propagation at the Infarct Boundary. Biophys. J. 2018, 115, 1966–1980. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Kartasalo, K.; Pölönen, R.-P.; Ojala, M.; Rasku, J.; Lekkala, J.; Aalto-Setala, K.; Kallio, P. CytoSpectre: A tool for spectral analysis of oriented structures on cellular and subcellular levels. BMC Bioinform. 2015, 16, 344. [Google Scholar] [CrossRef] [PubMed]
- Peyronnet, R.; Nerbonne, J.M.; Kohl, P. Cardiac Mechano-Gated Ion Channels and Arrhythmias. Circ. Res. 2016, 118, 311–329. [Google Scholar] [CrossRef]
- Künzel, S.R.; Rausch, J.S.; Schäffer, C.; Hoffmann, M.; Künzel, K.; Klapproth, E.; Kant, T.; Herzog, N.; Küpper, J.H.; Lorenz, K. Modeling atrial fibrosis in vitro—generation and characterization of a novel human atrial fibroblast cell line. FEBS Open Bio. 2020, 10, 1210–1218. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Leddy, H.A.; Chen, Y.; Lee, S.H.; Zelenski, N.A.; McNulty, A.L.; Wu, J.; Beicker, K.N.; Coles, J.; Zauscher, S.; et al. Synergy between Piezo1 and Piezo2 channels confers high-strain mechanosensitivity to articular cartilage. Proc. Natl. Acad. Sci. USA 2014, 111, E5114–E5122. [Google Scholar] [CrossRef]
- Pardo-Pastor, C.; Rubio-Moscardo, F.; Vogel-González, M.; Serra, S.A.; Afthinos, A.; Mrkonjic, S.; Destaing, O.; Abenza, J.F.; Fernández-Fernández, J.M.; Trepat, X.; et al. Piezo2 channel regulates RhoA and actin cytoskeleton to promote cell mechanobiological responses. Proc. Natl. Acad. Sci. USA 2018, 115, 1925–1930. [Google Scholar] [CrossRef]
- Yao, M.; Tijore, A.S.; Cox, C.D.; Hariharan, A.; Van Nhieu, G.T.; Martinac, B.; Sheetz, M. Force-dependent piezo1 recruitment to focal adhesions regulates adhesion maturation and turnover specifically in non-transformed cells. bioRxiv 2020. [Google Scholar] [CrossRef]
- Fabry, B.; Klemm, A.H.; Kienle, S.; Schäffer, T.E.; Goldmann, W.H. Focal Adhesion Kinase Stabilizes the Cytoskeleton. Biophys. J. 2011, 101, 2131–2138. [Google Scholar] [CrossRef]
- Totsukawa, G.; Yamakita, Y.; Yamashiro, S.; Hartshorne, D.J.; Sasaki, Y.; Matsumura, F. Distinct Roles of Rock (Rho-Kinase) and Mlck in Spatial Regulation of Mlc Phosphorylation for Assembly of Stress Fibers and Focal Adhesions in 3t3 Fibroblasts. J. Cell Biol. 2000, 150, 797–806. [Google Scholar] [CrossRef] [PubMed]
- Serrels, B.; Serrels, A.; Brunton, V.G.; Holt, M.; McLean, G.W.; Gray, C.H.; Jones, G.E.; Frame, M.C. Focal adhesion kinase controls actin assembly via a ferm-mediated interaction with the arp2/3 complex. Nat. Cell Biol. 2007, 9, 1046–1056. [Google Scholar] [CrossRef]
- Zeller, K.S.; Idevall-Hagren, O.; Stefansson, A.; Velling, T.; Jackson, S.P.; Downward, J.; Tengholm, A.; Johansson, S. PI3-kinase p110α mediates β1 integrin-induced Akt activation and membrane protrusion during cell attachment and initial spreading. Cell. Signal. 2010, 22, 1838–1848. [Google Scholar] [CrossRef] [PubMed]
- Engler, A.J.; Sen, S.; Sweeney, H.L.; Discher, D.E. Matrix Elasticity Directs Stem Cell Lineage Specification. Cell 2006, 126, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Gałdyszyńska, M.; Bobrowska, J.; Lekka, M.; Radwańska, P.; Piera, L.; Szymański, J.; Drobnik, J. The stiffness-controlled release of interleukin-6 by cardiac fibroblasts is dependent on integrin α2β1. J. Cell. Mol. Med. 2020, 24, 13853–13862. [Google Scholar] [CrossRef] [PubMed]

| Purpose | Agent | Class | Catalogue No. | Supplier | Final Concentration/Dilution | Solvent (Final Concen- tration) |
|---|---|---|---|---|---|---|
| Calpain inhibition | ALLN | Peptide | 208719 | Merck Millipore | 5 or 10 µM | DMSO (0.05 or 0.1%) |
| Integrin-β1 activation | P5D2 | Mono- clonal antibody | ab24693 | Abcam | 2.5 µg/mL | Na-azide (0.04%) |
| Integrin-β1 inhibition | HMβ1.1 | Mono- clonal antibody | ab36219 | Abcam | 1.25 µg/mL | Na-azide (0.09%) |
| FAK inhibition | PF-00562271 | Small molecule | S2672 | Selleck Chemicals | 1 µM | DMSO (0.1%) |
| ROCK inhibition | Y-27632 | Small molecule | S1049 | Selleck Chemicals | 1 µM | DMSO (0.1%) |
| IL-6 neutra- lization | MAB2061 | Mono- clonal antibody | MAB2061 | R&D Systems | 0.6 µg/mL | PBS (0.1%) |
| Target Gene | TaqMan Assay Number |
|---|---|
| PIEZO1 | Hs00270203_m1 |
| PIEZO2 | Hs00401026_m1 |
| GAPDH | Hs02786624_g1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Emig, R.; Knodt, W.; Krussig, M.J.; Zgierski-Johnston, C.M.; Gorka, O.; Groß, O.; Kohl, P.; Ravens, U.; Peyronnet, R. Piezo1 Channels Contribute to the Regulation of Human Atrial Fibroblast Mechanical Properties and Matrix Stiffness Sensing. Cells 2021, 10, 663. https://doi.org/10.3390/cells10030663
Emig R, Knodt W, Krussig MJ, Zgierski-Johnston CM, Gorka O, Groß O, Kohl P, Ravens U, Peyronnet R. Piezo1 Channels Contribute to the Regulation of Human Atrial Fibroblast Mechanical Properties and Matrix Stiffness Sensing. Cells. 2021; 10(3):663. https://doi.org/10.3390/cells10030663
Chicago/Turabian StyleEmig, Ramona, Wiebke Knodt, Mario J. Krussig, Callum M. Zgierski-Johnston, Oliver Gorka, Olaf Groß, Peter Kohl, Ursula Ravens, and Rémi Peyronnet. 2021. "Piezo1 Channels Contribute to the Regulation of Human Atrial Fibroblast Mechanical Properties and Matrix Stiffness Sensing" Cells 10, no. 3: 663. https://doi.org/10.3390/cells10030663
APA StyleEmig, R., Knodt, W., Krussig, M. J., Zgierski-Johnston, C. M., Gorka, O., Groß, O., Kohl, P., Ravens, U., & Peyronnet, R. (2021). Piezo1 Channels Contribute to the Regulation of Human Atrial Fibroblast Mechanical Properties and Matrix Stiffness Sensing. Cells, 10(3), 663. https://doi.org/10.3390/cells10030663








