An Analysis of the Neurological and Molecular Alterations Underlying the Pathogenesis of Alzheimer’s Disease
Abstract
1. Introduction
2. Genetic Risk Factors
2.1. APP, PSEN1, and PSEN2 Mutations
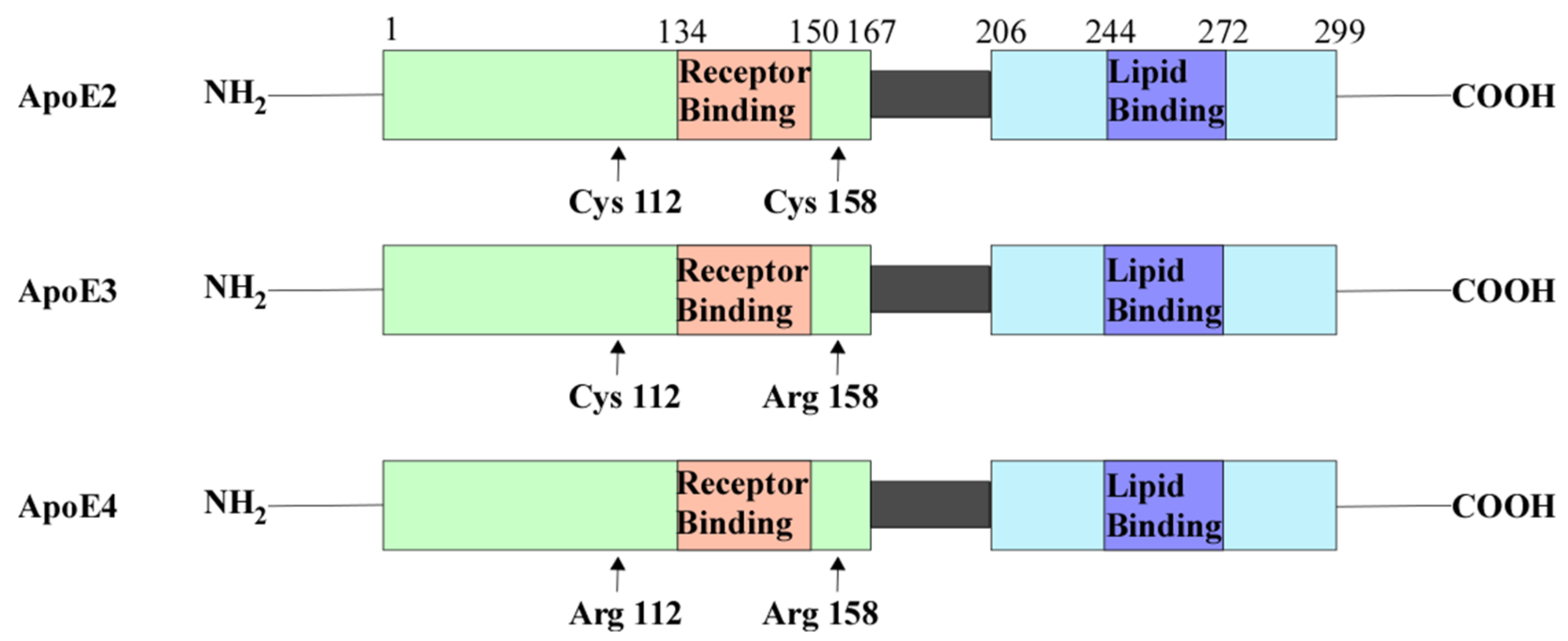
2.2. ApoE4
2.3. Other Genetic Risk Factors of AD
3. Neurological and Molecular Alterations of AD
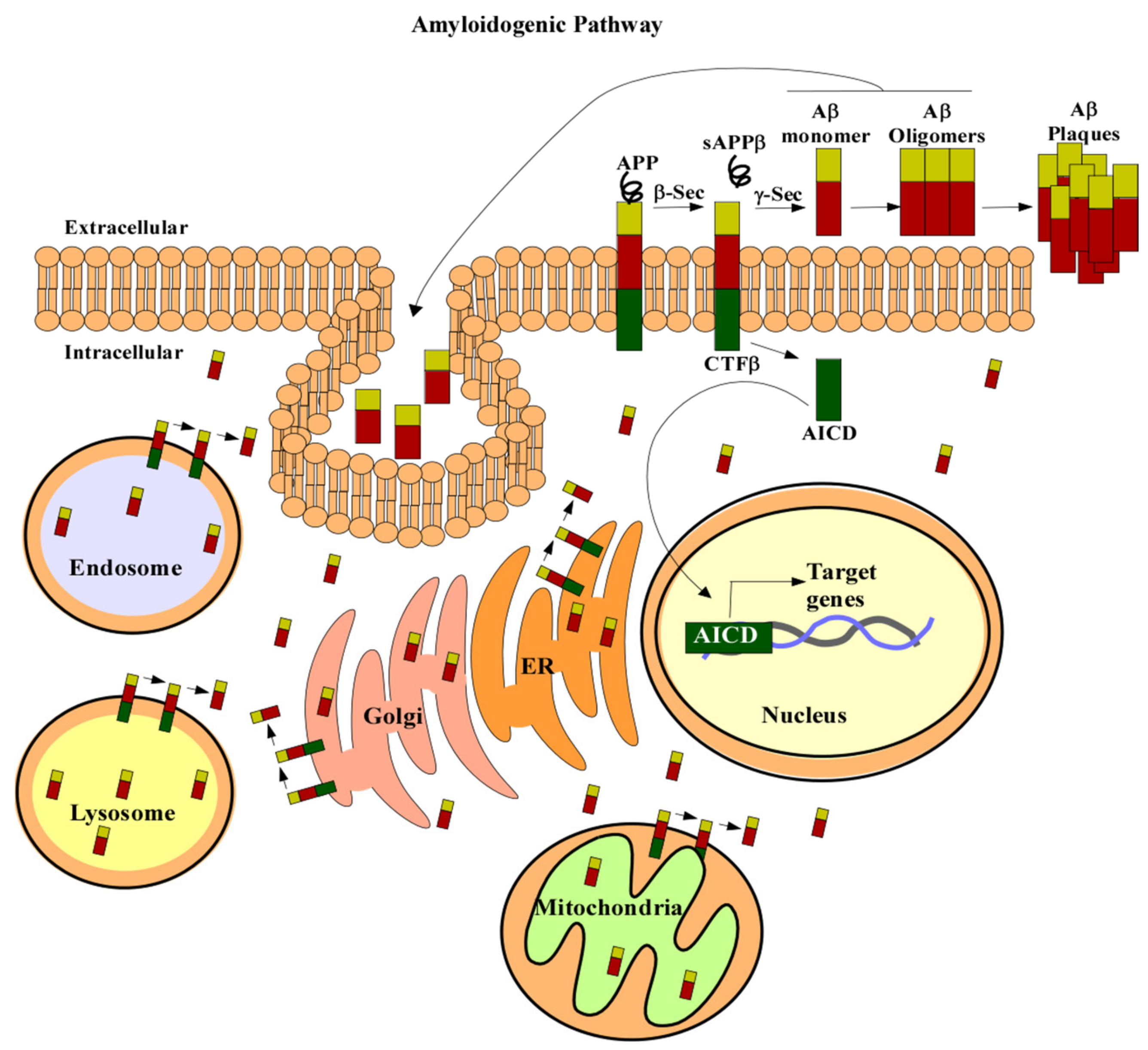
3.1. Amyloid Beta
3.2. Neurofibrillary Tangles
3.3. Neuronal Loss/Synaptic Loss
3.4. Blood–Brain Barrier Dysfunction
3.5. Inflammation
3.6. Defective Cholesterol Metabolism
3.7. Hypoxia
3.8. Mitochondrial Dysfunction
3.9. Oxidative Stress
3.10. Glucose Hypometabolism
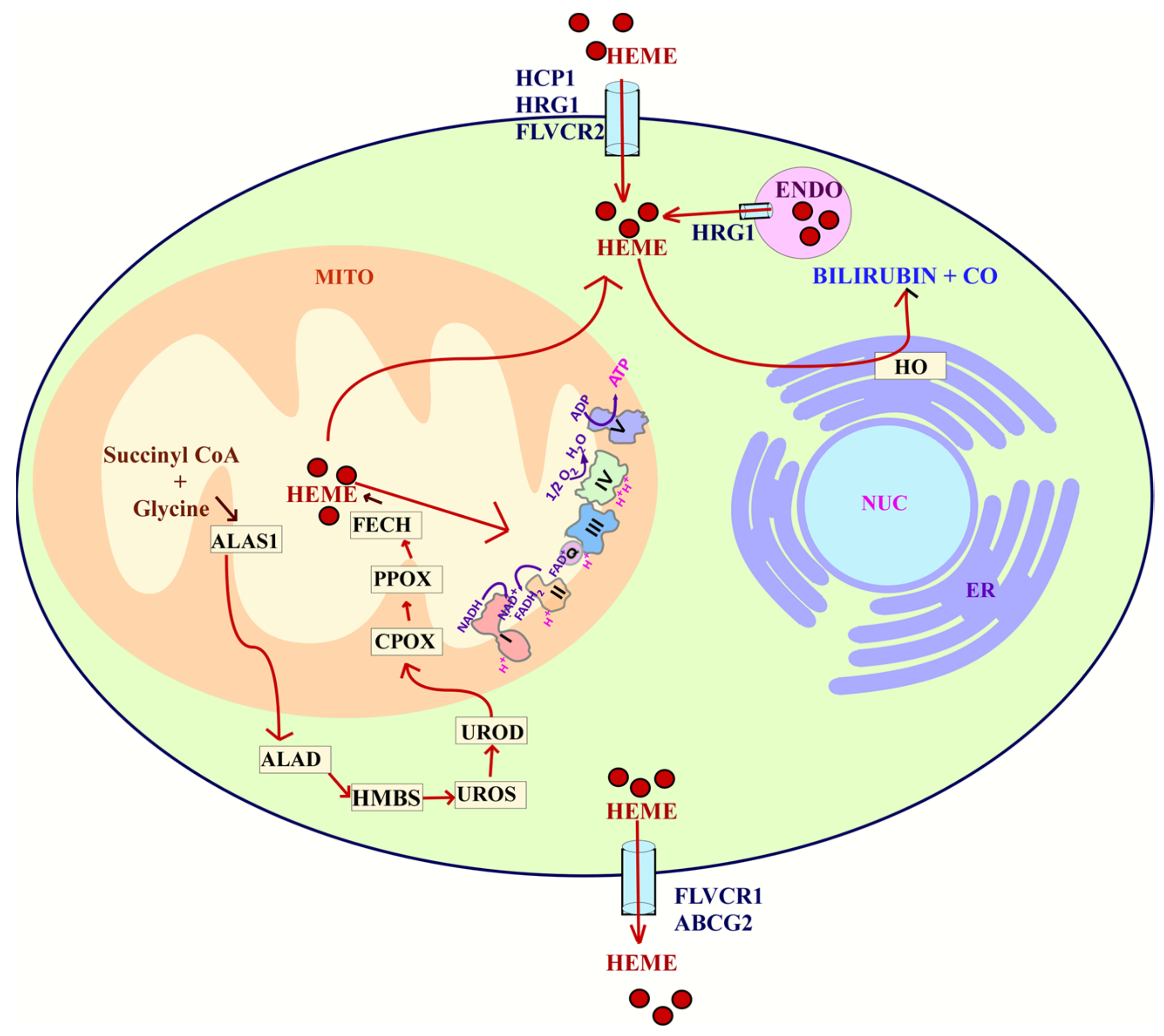
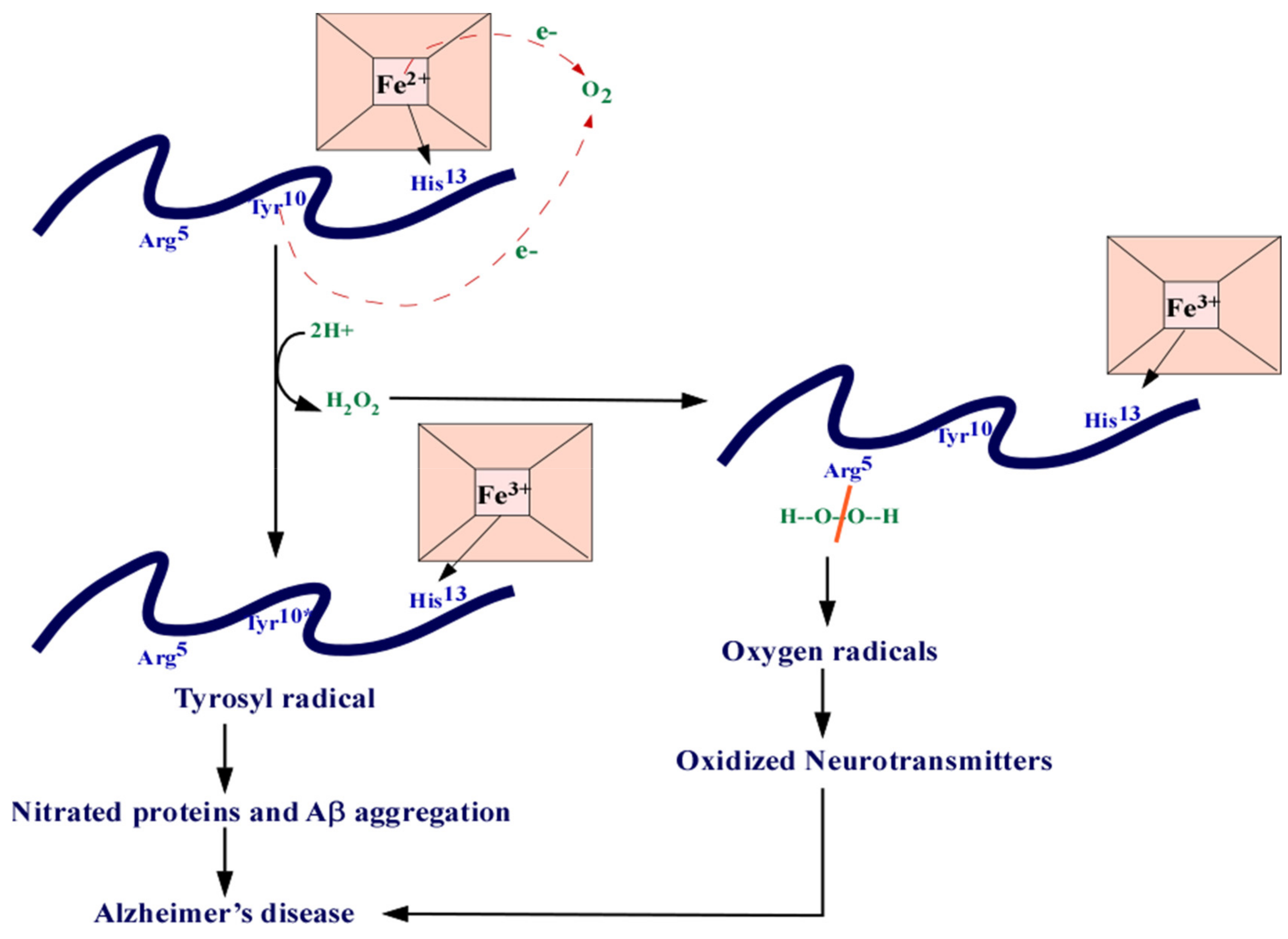
3.11. Dysregulated Homeostasis of Metals and Heme
4. Models of AD
4.1. Cell Models
4.2. Mouse Models
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ritchie, K.; Lovestone, S. The dementias. Lancet 2002, 360, 1759–1766. [Google Scholar] [CrossRef]
- Karantzoulis, S.; Galvin, J.E. Distinguishing Alzheimer’s disease from other major forms of dementia. Expert Rev. Neurother 2011, 11, 1579–1591. [Google Scholar] [CrossRef] [PubMed]
- Wolters, F.J.; Ikram, M.A. Epidemiology of Vascular Dementia. Arter. Thromb. Vasc. Biol. 2019, 39, 1542–1549. [Google Scholar] [CrossRef]
- Lobo, A.; Launer, L.J.; Fratiglioni, L.; Andersen, K.; Di Carlo, A.; Breteler, M.M.; Copeland, J.R.; Dartigues, J.F.; Jagger, C.; Martinez-Lage, J.; et al. Prevalence of dementia and major subtypes in Europe: A collaborative study of population-based cohorts. Neurol. Dis. Elder. Res. Grou Neurol. 2000, 54 (Suppl. 5), S4–S9. [Google Scholar]
- Zhang, M.Y.; Katzman, R.; Salmon, D.; Jin, H.; Cai, G.; Wang, Z.; Qu, G.; Grant, I.; Yu, E.; Levy, P.; et al. The prevalence of dementia and Alzheimer’s disease in Shanghai, China: Impact of age, gender, and education. Ann. Neurol. 1990, 27, 428–437. [Google Scholar] [CrossRef]
- 2020 Alzheimer’s disease facts and figures. Alzheimers Dement 2020, 16, 391–460. [CrossRef]
- Patterson, C. Alzheimer’s Disease International. World Alzheimer Report 2018. The State of the Art of Dementia Research: New Frontiers; Alzheimer’s Disease International: London, UK, 2018. [Google Scholar]
- Brickell, K.L.; Steinbart, E.J.; Rumbaugh, M.; Payami, H.; Schellenberg, G.D.; van Deerlin, V.; Yuan, W.; Bird, T.D. Early-onset Alzheimer disease in families with late-onset Alzheimer disease: A potential important subtype of familial Alzheimer disease. Arch. Neurol. 2006, 63, 1307–1311. [Google Scholar] [CrossRef] [PubMed]
- Van Cauwenberghe, C.; van Broeckhoven, C.; Sleegers, K. The genetic landscape of Alzheimer disease: Clinical implications and perspectives. Genet. Med. 2016, 18, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.V.F.; Loures, C.D.M.G.; Alves, L.C.V.; De Souza, L.C.; Borges, K.B.G.; Carvalho, M.D.G. Alzheimer’s disease: Risk factors and potentially protective measures. J. Biomed. Sci. 2019, 26, 33. [Google Scholar] [CrossRef] [PubMed]
- Oakley, H.; Cole, S.L.; Logan, S.; Maus, E.; Shao, P.; Craft, J.; Guillozet-Bongaarts, A.; Ohno, M.; Disterhoft, J.; van Eldik, L.; et al. Intraneuronal β-amyloid aggregates, neurodegeneration, and neuron loss in transgenic mice with five familial Alzheimer’s disease mutations: Potential factors in amyloid plaque formation. J. Neurosci. 2006, 26, 10129–10140. [Google Scholar] [CrossRef] [PubMed]
- Reitz, C.; Rogaeva, E.; Beecham, G.W. Late-onset vs nonmendelian early-onset Alzheimer disease: A distinction without a difference? Neurol. Genet. 2020, 6, e512. [Google Scholar] [CrossRef]
- Lanoiselée, H.-M.; Nicolas, G.; Wallon, D.; Rovelet-Lecrux, A.; Lacour, M.; Rousseau, S.; Richard, A.-C.; Pasquier, F.; Rollin-Sillaire, A.; Martinaud, O.; et al. APP, PSEN1, and PSEN2 mutations in early-onset Alzheimer disease: A genetic screening study of familial and sporadic cases. PLoS Med. 2017, 14, e1002270. [Google Scholar] [CrossRef]
- Van Giau, V.; Pyun, J.-M.; Suh, J.; Bagyinszky, E.; An, S.S.A.; Kim, S.Y. A pathogenic PSEN1 Trp165Cys mutation associated with early-onset Alzheimer’s disease. BMC Neurol. 2019, 19, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Veugelen, S.; Saito, T.; Saido, T.C.; Chávez-Gutiérrez, L.; de Strooper, B. Familial Alzheimer’s Disease Mutations in Presenilin Generate Amyloidogenic Aβ Peptide Seeds. Neuron 2016, 90, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Queralt, R.; Ezquerra, M.; Castellví, M.; Lleó, A.; Blesa, R.; Oliva, R. Detection of the presenilin 1 gene mutation (M139T) in early-onset familial Alzheimer disease in Spain. Neurosci. Lett. 2001, 299, 239–241. [Google Scholar] [CrossRef]
- DeTure, M.A.; Dickson, D.W. The neuropathological diagnosis of Alzheimer’s disease. Mol. Neurodegener. 2019, 14, 32. [Google Scholar] [CrossRef] [PubMed]
- Naslund, J.; Haroutunian, V.; Mohs, R.; Davis, K.L.; Davies, P.; Greengard, P.; Buxbaum, J.D. Correlation between elevated levels of amyloid β-peptide in the brain and cognitive decline. JAMA 2000, 283, 1571–1577. [Google Scholar] [CrossRef]
- Braak, H.; Braak, E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 1991, 82, 239–259. [Google Scholar] [CrossRef] [PubMed]
- Boonen, R.A.; van Tijn, P.; Zivkovic, D. Wnt signaling in Alzheimer’s disease: Up or down, that is the question. Ageing Res. Rev. 2009, 8, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Gjoneska, E.; Pfenning, A.R.; Mathys, H.; Quon, G.; Kundaje, A.; Tsai, L.-H.; Kellis, M. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer’s disease. Nature 2015, 518, 365–369. [Google Scholar] [CrossRef]
- Liang, W.S.; Reiman, E.M.; Valla, J.; Dunckley, T.; Beach, T.G.; Grover, A.; Niedzielko, T.L.; Schneider, L.E.; Mastroeni, D.; Caselli, R.; et al. Alzheimer’s disease is associated with reduced expression of energy metabolism genes in posterior cingulate neurons. Proc. Natl. Acad. Sci. USA 2008, 105, 4441–4446. [Google Scholar] [CrossRef] [PubMed]
- Bullock, R.; Bergman, H.; Touchon, J.; Gambina, G.; He, Y.; Nagel, J.; Lane, R. Effect of age on response to rivastigmine or donepezil in patients with Alzheimer’s disease. Curr. Med. Res. Opin. 2006, 22, 483–494. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.W.; Soininen, H.; Hager, K.; Aarsland, D.; Passmore, P.; Murthy, A.; Zhang, R.; Bahra, R. A multinational, randomised, 12-week study comparing the effects of donepezil and galantamine in patients with mild to moderate Alzheimer’s disease. Int. J. Geriatr. Psychiatry 2004, 19, 58–67. [Google Scholar] [CrossRef] [PubMed]
- Bullock, R.; Touchon, J.; Bergman, H.; Gambina, G.; He, Y.; Rapatz, G.; Nagel, J.; Lane, R. Rivastigmine and donepezil treatment in moderate to moderately-severe Alzheimer’s disease over a 2-year period. Curr. Med. Res. Opin. 2005, 21, 1317–1327. [Google Scholar] [CrossRef] [PubMed]
- Parsons, C.G.; Danysz, W.; Dekundy, A.; Pulte, I. Memantine and cholinesterase inhibitors: Complementary mechanisms in the treatment of Alzheimer’s disease. Neurotox. Res. 2013, 24, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Singh, A.; Ekavali, A. review on Alzheimer’s disease pathophysiology and its management: An update. Pharmacol. Rep. 2015, 67, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Atamna, H.; Frey, W.H., II. A role for heme in Alzheimer’s disease: Heme binds amyloid beta and has altered metabolism. Proc. Natl. Acad. Sci. USA 2004, 101, 11153–11158. [Google Scholar] [CrossRef] [PubMed]
- Vidal, C.; Daescu, K.; Fitzgerald, K.E.; Starokadomsk, A.; Bezprozvanny, I.; Zhang, L. Amyloid β perturbs elevated heme flux induced with neuronal development. Alzheimers Dement. 2019, 5, 27–37. [Google Scholar] [CrossRef]
- Atamna, H.; Boyle, K. Amyloid-β peptide binds with heme to form a peroxidase: Relationship to the cytopathologies of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2006, 103, 3381–3386. [Google Scholar] [CrossRef] [PubMed]
- Atamna, H.; Killilea, D.W.; Killilea, A.N.; Ames, B.N. Heme deficiency may be a factor in the mitochondrial and neuronal decay of aging. Proc. Natl. Acad. Sci. USA 2002, 99, 14807–14812. [Google Scholar] [CrossRef] [PubMed]
- Wolters, F.J.; Zonneveld, H.I.; Licher, S.; Cremers, L.G.M.; Heart Brain Connection Collaborative Research Group; Ikram, M.K.; Koudstaal, P.J.; Vernooij, M.W.; Ikram, M.A. Hemoglobin and anemia in relation to dementia risk and accompanying changes on brain MRI. Neurology 2019, 93, e917–e926. [Google Scholar] [CrossRef]
- Makin, S. The amyloid hypothesis on trial. Nature 2018, 559, S4–S7. [Google Scholar] [CrossRef] [PubMed]
- Scheuner, D.; Eckman, C.; Jensen, M.; Song, X.; Citron, M.; Suzuki, N.; Bird, T.D.; Hardy, J.; Hutton, M.; Kukull, W. Secreted amyloid β-protein similar to that in the senile plaques of Alzheimer’s disease is increased in vivo by the presenilin 1 and 2 and APP mutations linked to familial Alzheimer’s disease. Nat. Med. 1996, 2, 864–870. [Google Scholar] [CrossRef]
- Goate, A.; Chartier-Harlin, M.-C.; Mullan, M.; Brown, J.; Crawford, F.; Fidani, L.; Giuffra, L.; Haynes, A.; Irving, N.; James, L.; et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer’s disease. Nature 1991, 349, 704–706. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.-F.; Xu, T.-H.; Yan, Y.; Zhou, Y.-R.; Jiang, Y.; Melcher, K.; Xu, H.E. Amyloid beta: Structure, biology and structure-based therapeutic development. Acta Pharmacol. Sin. 2017, 38, 1205–1235. [Google Scholar] [CrossRef]
- Haass, C.; Selkoe, D.J. Soluble protein oligomers in neurodegeneration: Lessons from the Alzheimer’s amyloid β-peptide. Nat. Rev. Mol. Cell Biol. 2007, 8, 101–112. [Google Scholar] [CrossRef]
- Gandy, S. The role of cerebral amyloid β accumulation in common forms of Alzheimer disease. J. Clin. Investig. 2005, 115, 1121–1129. [Google Scholar]
- Murphy, M.P.; LeVine, H., III. Alzheimer’s disease and the amyloid-β peptide. J. Alzheimers Dis 2010, 19, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-W.; Thompson, R.; Zhang, H.; Xu, H. APP processing in Alzheimer’s disease. Mol. Brain 2011, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.; Anderson, J.P.; Barbour, R.; Basi, G.S.; Caccavello, R.; Davis, D.; Doan, M.; Dovey, H.F.; Frigon, N.; Hong, J. Purification and cloning of amyloid precursor protein β-secretase from human brain. Nature 1999, 402, 537–540. [Google Scholar] [CrossRef]
- Gravina, S.A.; Ho, L.; Eckman, C.B.; Long, K.E.; Otvos, L., Jr.; Younkin, L.H.; Suzuki, N.; Younkin, S.G. Amyloid β protein (Aβ) in Alzheimer’s disease brain. Biochemical and immunocytochemical analysis with antibodies specific for forms ending at Aβ40 or Aβ42(43). J. Biol. Chem. 1995, 270, 7013–7016. [Google Scholar] [CrossRef]
- Jakel, L.; Boche, D.; Nicoll, J.A.R.; Verbeek, M.M. Aβ43 in human Alzheimer’s disease: Effects of active Aβ42 immunization. Acta Neuropathol. Commun. 2019, 7, 141. [Google Scholar] [CrossRef]
- Zhang, X.; Li, Y.; Xu, H.; Zhang, Y.-W. The γ-secretase complex: From structure to function. Front. Cell. Neurosci. 2014, 8, 427. [Google Scholar] [CrossRef] [PubMed]
- Mucke, L.; Selkoe, D.J. Neurotoxicity of amyloid β-protein: Synaptic and network dysfunction. Cold Spring Harb. Perspect. Med. 2012, 2, a006338. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Pozo, A.; Frosch, M.P.; Masliah, E.; Hyman, B.T. Neuropathological Alterations in Alzheimer Disease. Cold Spring Harb. Perspect. Med. 2011, 1, a006189. [Google Scholar] [CrossRef]
- Citron, M.; Oltersdorf, T.; Haass, C.; McConlogue, L.; Hung, A.Y.; Seubert, P.; Vigo-Pelfrey, C.; Lieberburg, I.; Selkoe, D.J. Mutation of the β-amyloid precursor protein in familial Alzheimer’s disease increases β-protein production. Nature 1992, 360, 672–674. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.D.; Golde, T.E.; Younkin, S.G. Release of excess amyloid β protein from a mutant amyloid β protein precursor. Science 1993, 259, 514–516. [Google Scholar] [CrossRef] [PubMed]
- Karran, E.; Mercken, M.; De Strooper, B. The amyloid cascade hypothesis for Alzheimer’s disease: An appraisal for the development of therapeutics. Nat. Rev. Drug Discov. 2011, 10, 698–712. [Google Scholar] [CrossRef] [PubMed]
- Borchelt, D.R.; Thinakaran, G.; Eckman, C.B.; Lee, M.K.; Davenport, F.; Ratovitsky, T.; Prada, C.; Kim, G.; Seekins, S.; Yager, D. Familial Alzheimer’s disease-linked presenilin 1 variants elevate Aβ1-42/1-40 ratio in vitro and in vivo. Neuron 1996, 17, 1005–1013. [Google Scholar] [CrossRef]
- Wolfe, M.S. When loss is gain: Reduced presenilin proteolytic function leads to increased Aβ42/Aβ40. Talking Point on the role of presenilin mutations in Alzheimer disease. EMBO Rep. 2007, 8, 136–140. [Google Scholar] [CrossRef]
- Sun, L.; Zhou, R.; Yang, G.; Shi, Y. Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Aβ42 and Aβ40 peptides by γ-secretase. Proc. Natl. Acad. Sci. USA 2017, 114, E476–E485. [Google Scholar] [CrossRef]
- De Strooper, B. Loss-of-function presenilin mutations in Alzheimer disease. Talking Point on the role of presenilin mutations in Alzheimer disease. EMBO Rep. 2007, 8, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Kelleher, R.J., 3rd; Shen, J. Presenilin-1 mutations and Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2017, 114, 629–631. [Google Scholar] [CrossRef]
- Weggen, S.; Beher, D. Molecular consequences of amyloid precursor protein and presenilin mutations causing autosomal-dominant Alzheimer’s disease. Alzheimer’s Res. Ther. 2012, 4, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Tamaoka, A.; Odaka, A.; Ishibashi, Y.; Usami, M.; Sahara, N.; Suzuki, N.; Nukina, N.; Mizusawa, H.; Shoji, S.; Kanazawa, I. APP717 missense mutation affects the ratio of amyloid β protein species (Aβ1-42/43 and Aβ1-40) in familial Alzheimer’s disease brain. J. Biol. Chem. 1994, 269, 32721–32724. [Google Scholar] [CrossRef]
- Arber, C.; Toombs, J.; Lovejoy, C.; Ryan, N.S.; Paterson, R.W.; Willumsen, N.; Gkanatsiou, E.; Portelius, E.; Blennow, K.; Heslegrave, A.; et al. Familial Alzheimer’s disease patient-derived neurons reveal distinct mutation-specific effects on amyloid beta. Mol. Psychiatry 2020, 25, 2919–2931. [Google Scholar] [CrossRef] [PubMed]
- Fryer, J.D.; Simmons, K.; Parsadanian, M.; Bales, K.R.; Paul, S.M.; Sullivan, P.M.; Holtzman, D.M. Human apolipoprotein E4 alters the amyloid-β 40:42 ratio and promotes the formation of cerebral amyloid angiopathy in an amyloid precursor protein transgenic model. J. Neurosci. 2005, 25, 2803–2810. [Google Scholar] [CrossRef]
- Roher, A.E.; Esh, C.L.; Kokjohn, T.A.; Castaño, E.M.; Van Vickle, G.D.; Kalback, W.M.; Patton, R.L.; Luehrs, D.C.; Daugs, I.D.; Kuo, Y.-M.; et al. Amyloid beta peptides in human plasma and tissues and their significance for Alzheimer’s disease. Alzheimer’s Dement. 2009, 5, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Mahley, R.W. Apolipoprotein E: Structure and function in lipid metabolism, neurobiology, and Alzheimer’s diseases. Neurobiol. Dis. 2014, 72, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Kanekiyo, T.; Xu, H.; Bu, G. ApoE and Aβ in Alzheimer’s disease: Accidental encounters or partners? Neuron 2014, 81, 740–754. [Google Scholar] [CrossRef]
- Wu, L.; Zhao, L. ApoE2 and Alzheimer’s disease: Time to take a closer look. Neural Regen. Res. 2016, 11, 412–413. [Google Scholar]
- Mahley, R.W.; Rall, S.C., Jr. Apolipoprotein E: Far more than a lipid transport protein. Annu Rev. Genom. Hum. Genet. 2000, 1, 507–537. [Google Scholar] [CrossRef] [PubMed]
- Altenburg, M.; Arbones-Mainar, J.M.; Johnson, L.; Wilder, J.; Maeda, N. Human LDL receptor enhances sequestration of ApoE4 and VLDL remnants on the surface of hepatocytes but not their internalization in mice. Arter. Thromb. Vasc. Biol. 2008, 28, 1104–1110. [Google Scholar] [CrossRef] [PubMed]
- Heeren, J.; Grewal, T.; Laatsch, A.; Becker, N.; Rinninger, F.; Rye, K.-A.; Beisiegel, U. Impaired Recycling of Apolipoprotein E4 Is Associated with Intracellular Cholesterol Accumulation. J. Biol. Chem. 2004, 279, 55483–55492. [Google Scholar] [CrossRef] [PubMed]
- Fassbender, K.; Simons, M.; Bergmann, C.; Stroick, M.; Lütjohann, D.; Keller, P.; Runz, H.; Kühl, S.; Bertsch, T.; von Bergmann, K. Simvastatin strongly reduces levels of Alzheimer’s disease β-amyloid peptides Aβ42 and Aβ40 in vitro and in vivo. Proc. Natl. Acad. Sci. USA 2001, 98, 5856–5861. [Google Scholar] [CrossRef]
- Simons, M.; Keller, P.; de Strooper, B.; Beyreuther, K.; Dotti, C.G.; Simons, K. Cholesterol depletion inhibits the generation of β-amyloid in hippocampal neurons. Proc. Natl. Acad. Sci. USA 1998, 95, 6460–6464. [Google Scholar] [CrossRef] [PubMed]
- Sparks, D.L.; Scheff, S.W.; Hunsaker, J.C., III; Liu, H.; Landers, T.; Gross, D.R. Induction of Alzheimer-like β-amyloid immunoreactivity in the brains of rabbits with dietary cholesterol. Exp. Neurol. 1994, 126, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Refolo, L.M.; Pappolla, M.A.; Malestera, B.; La Francoisa, J.; Thomasb, T.B.-; Wangc, R.; Tintd, G.S.; Sambamurtie, K.; Duff, K. Hypercholesterolemia Accelerates the Alzheimer’s Amyloid Pathology in a Transgenic Mouse Model. Neurobiol. Dis. 2000, 7, 321–331. [Google Scholar] [CrossRef]
- Li, Z.; Shue, F.; Zhao, N.; Shinohara, M.; Bu, G. APOE2: Protective mechanism and therapeutic implications for Alzheimer’s disease. Mol. Neurodegener. 2020, 15, 1–19. [Google Scholar] [CrossRef]
- Zhao, L.; Lin, S.; Bales, K.R.; Gelfanova, V.; Koger, D.; Delong, C.; Hale, J.; Liu, F.; Hunter, J.M.; Paul, S.M. Macrophage-Mediated Degradation of β-Amyloid via an Apolipoprotein E Isoform-Dependent Mechanism. J. Neurosci. 2009, 29, 3603–3612. [Google Scholar] [CrossRef] [PubMed]
- Jansen, W.J.; Ossenkoppele, R.; Knol, D.L.; Tijms, B.M.; Scheltens, P.; Verhey, F.R.J.; Visser, P.J.; the Amyloid Biomarker Study Group. Prevalence of cerebral amyloid pathology in persons without dementia: A meta-analysis. JAMA 2015, 313, 1924–1938. [Google Scholar] [CrossRef] [PubMed]
- Corder, E.H.; Saunders, A.M.; Strittmatter, W.J.; Schmechel, D.E.; Gaskell, P.C.; Small, G.W.; Roses, A.D.; Haines, J.L.; Pericak-Vance, M.A. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 1993, 261, 921–923. [Google Scholar] [CrossRef]
- Strittmatter, W.J.; Saunders, A.M.; Schmechel, D.; Pericak-Vance, M.; Enghild, J.; Salvesen, G.S.; Roses, A.D. Apolipoprotein E: High-avidity binding to β-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc. Natl. Acad. Sci. USA 1993, 90, 1977–1981. [Google Scholar] [CrossRef]
- Saunders, A.M.; Strittmatter, W.J.; Schmechel, D.; George-Hyslop, P.H.; Pericak-Vance, M.A.; Joo, S.H.; Rosi, B.L.; Gusella, J.F.; Crapper-MacLachlan, D.R.; Alberts, M.J.; et al. Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology 1993, 43, 1467–1472. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-W.A.; Zhou, B.; Wernig, M.; Südhof, T.C. ApoE2, ApoE3, and ApoE4 Differentially Stimulate APP Transcription and Aβ Secretion. Cell 2017, 168, 427–441.e21. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-W.A.; Zhou, B.; Nabet, A.M.; Wernig, M.; Südhof, T.C. Differential Signaling Mediated by ApoE2, ApoE3, and ApoE4 in Human Neurons Parallels Alzheimer’s Disease Risk. J. Neurosci. 2019, 39, 7408–7427. [Google Scholar] [CrossRef] [PubMed]
- Nagy, Z.; Esiri, M.; Jobst, K.; Johnston, C.; Litchfield, S.; Sim, E.; Smith, A. Influence of the apolipoprotein E genotype on amyloid deposition and neurofibrillary tangle formation in Alzheimer’s disease. Neuroscience 1995, 69, 757–761. [Google Scholar] [CrossRef]
- Serrano-Pozo, A.; Qian, J.; Monsell, S.E.; Betensky, R.A.; Hyman, B.T. APOEε2 is associated with milder clinical and pathological Alzheimer disease. Ann. Neurol. 2015, 77, 917–929. [Google Scholar] [CrossRef]
- Chang, S.; Ma, T.R.; Miranda, R.D.; Balestra, M.E.; Mahley, R.W.; Huang, Y. Lipid- and receptor-binding regions of apolipoprotein E4 fragments act in concert to cause mitochondrial dysfunction and neurotoxicity. Proc. Natl. Acad. Sci. USA 2005, 102, 18694–18699. [Google Scholar] [CrossRef] [PubMed]
- Lambert, J.C.; Ibrahim-Verbaas, C.A.; Harold, D.; Naj, A.C.; Sims, R.; Bellenguez, C.; DeStafano, A.L.; Bis, J.C.; Beecham, G.W.; Grenier-Boley, B.; et al. Meta-Analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 2013, 45, 1452–1458. [Google Scholar] [CrossRef]
- Jung, Y.J.; Kim, Y.H.; Bhalla, M.; Lee, S.B.; Seo, J. Genomics: New Light on Alzheimer’s Disease Research. Int. J. Mol. Sci. 2018, 19, 3771. [Google Scholar] [CrossRef]
- Guerreiro, R.; Wojtas, A.; Bras, J.; Carrasquillo, M.M.; Rogaeva, E.; Majounie, E.; Cruchaga, C.; Sassi, C.; Kauwe, J.S.; Younkin, S.G.; et al. TREM2 Variants in Alzheimer’s Disease. N. Engl. J. Med. 2013, 368, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Parhizkar, S.; Arzberger, T.; Brendel, M.; Kleinberger, G.; Deussing, M.; Focke, C.; Nuscher, B.; Xiong, M.; Ghasemigharagoz, A.; Katzmarski, N.; et al. Loss of TREM2 function increases amyloid seeding but reduces plaque-associated ApoE. Nat. Neurosci. 2019, 22, 191–204. [Google Scholar] [CrossRef]
- Griciuc, A.; Serrano-Pozo, A.; Parrado, A.R.; Lesinski, A.N.; Asselin, C.N.; Mullin, K.; Hooli, B.; Choi, S.H.; Hyman, T.B.; Tanzi, R.E. Alzheimer’s disease risk gene CD33 inhibits microglial uptake of amyloid β. Neuron 2013, 78, 631–643. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.-S.; Yu, J.-T.; Tan, L. Bridging integrator 1 (BIN1): Form, function, and Alzheimer’s disease. Trends Mol. Med. 2013, 19, 594–603. [Google Scholar] [CrossRef] [PubMed]
- Chapuis, J.; Hansmannel, F.; Gistelinck, M.; Mounier, A.; Van Cauwenberghe, C.; Kolen, K.V.; Geller, F.; Sottejeau, Y.; Harold, D.; Dourlen, P.; et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol. Psychiatry 2013, 18, 1225–1234. [Google Scholar] [CrossRef]
- Lambert, J.-C.; the European Alzheimer’s Disease Initiative Investigators; Heath, S.; Even, G.; Campion, D.; Sleegers, K.; Hiltunen, M.O.; Combarros, O.; Zelenika, D.; Bullido, M.J.; et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat. Genet. 2009, 41, 1094–1099. [Google Scholar] [CrossRef]
- DeMattos, R.B.; Cirrito, J.R.; Parsadanian, M.; May, P.C.; O’Dell, M.A.; Taylor, J.W.; Harmony, J.A.K.; Aronow, B.J.; Bales, K.R.; Paul, S.M.; et al. ApoE and clusterin cooperatively suppress Aβ levels and deposition: Evidence that ApoE regulates extracellular Aβ metabolism in vivo. Neuron 2004, 41, 193–202. [Google Scholar] [CrossRef]
- Rogers, J.; Li, R.; Mastroeni, D.; Grover, A.; Leonard, B.W.; Ahern, G.; Cao, P.; Kolody, H.; Vedders, L.; Kolb, W.P.; et al. Peripheral clearance of amyloid β peptide by complement C3-dependent adherence to erythrocytes. Neurobiol. Aging 2006, 27, 1733–1739. [Google Scholar] [CrossRef] [PubMed]
- Naj, A.C.; Jun, G.; Beecham, G.W.; Wang, L.-S.; Vardarajan, B.N.; Buros, J.; Gallins, P.J.; Buxbaum, J.D.; Jarvik, G.P.; Crane, P.K.; et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat. Genet. 2011, 43, 436–441. [Google Scholar] [CrossRef]
- Karch, C.M.; Jeng, A.T.; Nowotny, P.; Cady, J.; Cruchaga, C.; Goate, A.M. Expression of Novel Alzheimer’s Disease Risk Genes in Control and Alzheimer’s Disease Brains. PLoS ONE 2012, 7, e50976. [Google Scholar] [CrossRef] [PubMed]
- Tao, Q.-Q.; Chen, Y.-C.; Wu, Z.-Y.; Qing-Qing, T.; Yu-Chao, C.; Zhi-Ying, W. The role of CD2AP in the Pathogenesis of Alzheimer’s Disease. Aging Dis. 2019, 10, 901–907. [Google Scholar] [CrossRef] [PubMed]
- Hollingworth, P.; Harold, D.; Sims, R.; Gerrish, A.; Lambert, J.-C.; Carrasquillo, M.M.; Abraham, R.; Hamshere, M.L.; Pahwa, J.S.; Moskvina, V.; et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat. Genet. 2011, 43, 429–435. [Google Scholar] [CrossRef]
- Cochran, J.N.; Rush, T.; Buckingham, S.C.; Roberson, E.D. The Alzheimer’s disease risk factor CD2AP maintains blood-brain barrier integrity. Hum. Mol. Genet. 2015, 24, 6667–6674. [Google Scholar] [CrossRef]
- Aikawa, T.; Holm, M.-L.; Kanekiyo, T. ABCA7 and Pathogenic Pathways of Alzheimer’s Disease. Brain Sci. 2018, 8, 27. [Google Scholar] [CrossRef] [PubMed]
- Steinberg, S.; Gene, D.; Stefansson, H.; Jonsson, T.; Johannsdottir, H.; Ingason, A.; Helgason, H.; Sulem, P.; Magnusson, O.T.; Gudjonsson, S.A.; et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat. Genet. 2015, 47, 445–447. [Google Scholar] [CrossRef] [PubMed]
- Sakae, N.; Liu, C.-C.; Shinohara, M.; Frisch-Daiello, J.; Ma, L.; Yamazaki, Y.; Tachibana, M.; Younkin, L.; Kurti, A.; Carrasquillo, M.M.; et al. ABCA7 Deficiency Accelerates Amyloid-β Generation and Alzheimer’s Neuronal Pathology. J. Neurosci. 2016, 36, 3848–3859. [Google Scholar] [CrossRef] [PubMed]
- Harold, D.; Abraham, R.; Hollingworth, P.; Sims, R.; Gerrish, A.; Hamshere, M.L.; Pahwa, J.S.; Moskvina, V.; Dowzell, K.; Williams, A.; et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat. Genet. 2009, 41, 1088–1093. [Google Scholar] [CrossRef] [PubMed]
- Seshadri, S.; Fitzpatrick, A.L.; Ikram, M.A.; DeStefano, A.L.; Gudnason, V.; Boada, M.; Bis, J.C.; Smith, A.V.; Carassquillo, M.M.; Lambert, J.C.; et al. Genome-wide Analysis of Genetic Loci Associated With Alzheimer Disease. JAMA 2010, 303, 1832–1840. [Google Scholar] [CrossRef]
- Xiao, Q.; Gil, S.-C.; Yan, P.; Wang, Y.; Han, S.; Gonzales, E.; Perez, R.; Cirrito, J.R.; Lee, J.-M. Role of Phosphatidylinositol Clathrin Assembly Lymphoid-Myeloid Leukemia (PICALM) in Intracellular Amyloid Precursor Protein (APP) Processing and Amyloid Plaque Pathogenesis. J. Biol. Chem. 2012, 287, 21279–21289. [Google Scholar] [CrossRef] [PubMed]
- Hughes, T.M.; Lopez, O.L.; Evans, R.W.; Kamboh, M.I.; Williamson, J.D.; Klunk, W.E.; Mathis, C.A.; Price, J.C.; Cohen, A.D.; Snitz, B.E.; et al. Markers of cholesterol transport are associated with amyloid deposition in the brain. Neurobiol. Aging 2014, 35, 802–807. [Google Scholar] [CrossRef] [PubMed]
- Miyashita, A.; Koike, A.; Jun, G.; Wang, L.; Takahashi, S.; Matsubara, E.; Kawarabayashi, T.; Shoji, M.; Tomita, N.; Arai, H. SORL1 is genetically associated with late-onset Alzheimer’s disease in Japanese, Koreans and Caucasians. PLoS ONE 2013, 8, e58618. [Google Scholar] [CrossRef]
- Reitz, C.; Cheng, R.; Rogaeva, E.; Lee, J.H.; Tokuhiro, S.; Zou, F.; Bettens, K.; Sleegers, K.; Tan, E.K.; Kimura, R.; et al. Meta-analysis of the association between variants in SORL1 and Alzheimer disease. Arch. Neurol. 2011, 68, 99–106. [Google Scholar] [CrossRef]
- Cuccaro, M.L.; Carney, R.M.; Zhang, Y.; Bohm, C.; Kunkle, B.W.; Vardarajan, B.N.; Whitehead, P.L.; Cukier, H.N.; Mayeux, R.; George-Hyslop, P.S.; et al. SORL1 mutations in early- and late-onset Alzheimer disease. Neurol. Genet. 2016, 2, e116. [Google Scholar] [CrossRef]
- Edwards, G.A., III; Gamez, N.; Escobedo, G., Jr.; Calderon, O.; Moreno-Gonzalez, I. Modifiable Risk Factors for Alzheimer’s Disease. Front. Aging Neurosci. 2019, 11, 146. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Zhao, F.; Ma, X.; Perry, G.; Zhu, X. Mitochondria dysfunction in the pathogenesis of Alzheimer’s disease: Recent advances. Mol. Neurodegener. 2020, 15, 1–22. [Google Scholar] [CrossRef]
- Huang, W.-J.; Zhang, X.; Chen, W.-W. Role of oxidative stress in Alzheimer’s disease. Biomed. Rep. 2016, 4, 519–522. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, H.; Barger, S.; Barnum, S.; Bradt, B.; Bauer, J.; Cole, G.M.; Cooper, N.R.; Eikelenboom, P.; Emmerling, M.; Fiebich, B.L.; et al. Inflammation and Alzheimer’s disease. Neurobiol. Aging 2000, 21, 383–421. [Google Scholar] [CrossRef]
- Leyns, C.E.G.; Holtzman, D.M. Glial contributions to neurodegeneration in tauopathies. Mol. Neurodegener. 2017, 12, 50. [Google Scholar] [CrossRef] [PubMed]
- Niikura, T.; Tajima, H.; Kita, Y. Neuronal cell death in Alzheimer’s disease and a neuroprotective factor, humanin. Curr. Neuropharmacol. 2006, 4, 139–147. [Google Scholar] [CrossRef]
- Costantini, L.C.; Barr, L.J.; Vogel, J.L.; Henderson, S.T. Hypometabolism as a therapeutic target in Alzheimer’s disease. BMC Neurosci. 2008, 9 (Suppl. 2), S16. [Google Scholar] [CrossRef]
- Hippius, H.; Neundorfer, G. The discovery of Alzheimer’s disease. Dialogues Clin. Neurosci. 2003, 5, 101–108. [Google Scholar] [PubMed]
- Alzheimer, A.; Stelzmann, R.A.; Schnitzlein, H.N.; Murtagh, F.R. An English translation of Alzheimer’s 1907 paper, “Uber eine eigenartige Erkankung der Hirnrinde”. Clin. Anat. 1995, 8, 429–431. [Google Scholar] [CrossRef]
- Cras, P.; Kawai, M.; Lowery, D.; Gonzalez-DeWhitt, P.; Greenberg, B.; Perry, G. Senile plaque neurites in Alzheimer disease accumulate amyloid precursor protein. Proc. Natl. Acad. Sci. USA 1991, 88, 7552–7556. [Google Scholar] [CrossRef] [PubMed]
- Allsop, D.; Mayes, J. Amyloid β-peptide and Alzheimer’s disease. Essays Biochem. 2014, 56, 99–110. [Google Scholar] [PubMed]
- Strodel, B.; Lee, J.W.L.; Whittleston, C.S.; Wales, D.J. Transmembrane structures for Alzheimer’s Aβ(1-42) oligomers. J. Am. Chem. Soc. 2010, 132, 13300–13312. [Google Scholar] [CrossRef] [PubMed]
- Demuro, A.; Mina, E.; Kayed, R.; Milton, S.C.; Parker, I.; Glabe, C.G. Calcium Dysregulation and Membrane Disruption as a Ubiquitous Neurotoxic Mechanism of Soluble Amyloid Oligomers. J. Biol. Chem. 2005, 280, 17294–17300. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.C. Calcium homeostasis and modulation of synaptic plasticity in the aged brain. Aging Cell 2007, 6, 319–325. [Google Scholar] [CrossRef]
- Takuma, H.; Tomiyama, T.; Kuida, K.; Mori, H. Amyloid β peptide-induced cerebral neuronal loss is mediated by caspase-3 in vivo. J. Neuropathol. Exp. Neurol. 2004, 63, 255–261. [Google Scholar] [CrossRef][Green Version]
- Han, X.J.; Hu, Y.-Y.; Yang, Z.-J.; Jiang, L.-P.; Shi, S.-L.; Li, Y.-R.; Guo, M.-Y.; Wu, H.-L.; Wan, Y.-Y. Amyloid β-42 induces neuronal apoptosis by targeting mitochondria. Mol. Med. Rep. 2017, 16, 4521–4528. [Google Scholar] [CrossRef] [PubMed]
- Musiek, E.S.; Holtzman, D.M. Three dimensions of the amyloid hypothesis: Time, space and “wingmen”. Nat. Neurosci. 2015, 18, 800–806. [Google Scholar] [CrossRef]
- Bateman, R.J.; Xiong, C.; Benzinger, T.L.; Fagan, A.M.; Goate, A.; Fox, N.C.; Marcus, D.S.; Cairns, N.J.; Xie, X.; Blazey, T.M.; et al. Clinical and biomarker changes in dominantly inherited Alzheimer’s disease. N. Engl. J. Med. 2012, 367, 795–804. [Google Scholar] [CrossRef] [PubMed]
- Esparza, T.J.; Zhao, H.; Cirrito, J.R.; Cairns, N.J.; Bateman, R.J.; Holtzman, D.M.; Brody, D.L. Amyloid-β oligomerization in Alzheimer dementia versus high-pathology controls. Ann. Neurol. 2013, 73, 104–119. [Google Scholar] [CrossRef]
- Jin, M.; Shepardson, N.; Yang, T.; Chen, G.; Walsh, D.; Selkoe, D.J. Soluble amyloid β-protein dimers isolated from Alzheimer cortex directly induce Tau hyperphosphorylation and neuritic degeneration. Proc. Natl. Acad. Sci. USA 2011, 108, 5819–5824. [Google Scholar] [CrossRef] [PubMed]
- LaFerla, F.M.; Green, K.N.; Oddo, S. Intracellular amyloid-β in Alzheimer’s disease. Nat. Rev. Neurosci. 2007, 8, 499–509. [Google Scholar] [CrossRef]
- Wesen, E.; Jeffries, G.D.M.; Dzebo, M.M.; Esbjörner, E.E. Endocytic uptake of monomeric amyloid-β peptides is clathrin- and dynamin-independent and results in selective accumulation of Aβ(1-42) compared to Aβ(1-40). Sci. Rep. 2017, 7, 2021. [Google Scholar] [CrossRef]
- Yazawa, H.; Yu, Z.-X.; Takeda, K.; Le, Y.; Gong, W.; Ferrans, V.J.; Oppenheim, J.J.; Li, C.C.H.; Wang, J.M. β amyloid peptide (Aβ42) is internalized via the G-protein-coupled receptor FPRL1 and forms fibrillar aggregates in macrophages. FASEB J. 2001, 15, 2454–2462. [Google Scholar] [CrossRef]
- Nagele, R.G.; D’Andrea, M.R.; Anderson, W.J.; Wang, H.-Y. Intracellular accumulation of β-amyloid(1-42) in neurons is facilitated by the α7 nicotinic acetylcholine receptor in Alzheimer’s disease. Neuroscience 2002, 110, 199–211. [Google Scholar] [CrossRef]
- Deane, R.; Du Yan, S.; Submamaryan, R.K.; LaRue, B.; Jovanovic, S.; Hogg, E.; Welch, D.; Manness, L.; Lin, C.; Yu, J.; et al. RAGE mediates amyloid-β peptide transport across the blood-brain barrier and accumulation in brain. Nat. Med. 2003, 9, 907–913. [Google Scholar] [CrossRef] [PubMed]
- Lai, A.Y.; McLaurin, J. Mechanisms of amyloid-β Peptide uptake by neurons: The role of lipid rafts and lipid raft-associated proteins. Int. J. Alzheimers Dis. 2010, 2011, 548380. [Google Scholar] [CrossRef]
- Takuma, K.; Fang, F.; Zhang, W.; Yan, S.; Fukuzaki, E.; Du, H.; Sosunov, A.; McKhann, G.; Funatsu, Y.; Nakamichi, N.; et al. RAGE-mediated signaling contributes to intraneuronal transport of amyloid-β and neuronal dysfunction. Proc. Natl. Acad. Sci. USA 2009, 106, 20021–20026. [Google Scholar] [CrossRef] [PubMed]
- Snyder, E.M.; Nong, Y.; Almeida, C.G.; Paul, S.; Moran, T.; Choi, E.Y.; Nairn, A.C.; Salter, M.W.; Lombroso, P.J.; Gouras, G.K.; et al. Regulation of NMDA receptor trafficking by amyloid-β. Nat. Neurosci. 2005, 8, 1051–1058. [Google Scholar] [CrossRef]
- Iribarren, P.; Zhou, Y.; Hu, J.; Le, Y.; Wang, J.M. Role of Formyl Peptide Receptor-Like 1 (FPRL1/FPR2) in Mononuclear Phagocyte Responses in Alzheimer Disease. Immunol. Res. 2005, 31, 165–176. [Google Scholar] [CrossRef]
- Takahashi, R.H.; Milner, T.A.; Li, F.; Nam, E.E.; Edgar, M.A.; Yamaguchi, H.; Beal, M.F.; Xu, H.; Greengard, P.; Gouras, G.K. Intraneuronal Alzheimer Aβ42 Accumulates in Multivesicular Bodies and Is Associated with Synaptic Pathology. Am. J. Pathol. 2002, 161, 1869–1879. [Google Scholar] [CrossRef]
- Grant, S.M.; Shankar, S.L.; Chalmers-Redman, R.M.E.; Tatton, W.G.; Szyf, M.; Cuello, A.C. Mitochondrial abnormalities in neuroectodermal cells stably expressing human amyloid precursor protein (hAPP751). NeuroReport 1999, 10, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; McLaughlin, R.; Goodyer, C.G.; Leblanc, A.C. Selective cytotoxicity of intracellular amyloid β peptide 1-42 through p53 and Bax in cultured primary human neurons. J. Cell Biol. 2002, 156, 519–529. [Google Scholar] [CrossRef]
- Binder, L.I.; Guillozet-Bongaarts, A.L.; Garcia-Sierra, F.; Berry, R.W. Tau, tangles, and Alzheimer’s disease. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2005, 1739, 216–223. [Google Scholar] [CrossRef]
- D’Errico, P.; Meyer-Luehmann, M. Mechanisms of Pathogenic Tau and Aβ Protein Spreading in Alzheimer’s Disease. Front. Aging Neurosci. 2020, 12, 265. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, D.W.; Hwo, S.-Y.; Kirschner, M.W. Purification of tau, a microtubule-associated protein that induces assembly of microtubules from purified tubulin. J. Mol. Biol. 1977, 116, 207–225. [Google Scholar] [CrossRef]
- Ramkumar, A.; Jong, B.Y.; Ori-McKenney, K.M. ReMAPping the microtubule landscape: How phosphorylation dictates the activities of microtubule-associated proteins. Dev. Dyn. 2018, 247, 138–155. [Google Scholar] [CrossRef]
- Lindwall, G.; Cole, R.D. Phosphorylation affects the ability of tau protein to promote microtubule assembly. J. Biol. Chem. 1984, 259, 5301–5305. [Google Scholar] [CrossRef]
- Ksiezak-Reding, H.; Pyo, H.K.; Feinstein, B.; Pasinetti, G.M. Akt/PKB kinase phosphorylates separately Thr212 and Ser214 of tau protein in vitro. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2003, 1639, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, A.; Kabatb, J.; Novakb, M.; Wua, Q.; Iqbala, I.G.; Iqbala, K. Phosphorylation of Tau at Both Thr 231 and Ser 262 Is Required for Maximal Inhibition of Its Binding to Microtubules. Arch. Biochem. Biophys. 1998, 357, 299–309. [Google Scholar] [CrossRef]
- Liu, F.; Li, B.; Tung, E.-J.; Grundke-Iqbal, I.; Iqbal, K.; Gong, C.-X. Site-specific effects of tau phosphorylation on its microtubule assembly activity and self-aggregation. Eur. J. Neurosci. 2007, 26, 3429–3436. [Google Scholar] [CrossRef] [PubMed]
- Johnson, G.V.; Stoothoff, W.H. Tau phosphorylation in neuronal cell function and dysfunction. J. Cell Sci. 2004, 117 Pt 24, 5721–5729. [Google Scholar] [CrossRef]
- Gendron, T.F.; Petrucelli, L. The role of tau in neurodegeneration. Mol. Neurodegener. 2009, 4, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Miao, J.; Shi, R.; Li, L.; Chen, F.; Zhou, Y.; Tung, Y.C.; Hu, W.; Gong, C.-X.; Iqbal, K.; Liu, F. Pathological Tau from Alzheimer’s Brain Induces Site-Specific Hyperphosphorylation and SDS- and Reducing Agent-Resistant Aggregation of Tau In Vivo. Front. Aging Neurosci. 2019, 11, 34. [Google Scholar] [CrossRef] [PubMed]
- Arriagada, P.V.; Growdon, J.H.; Hedley-Whyte, E.T.; Hyman, B.T. Neurofibrillary tangles but not senile plaques parallel duration and severity of Alzheimer’s disease. Neurology 1992, 42 Pt 1, 631–639. [Google Scholar] [CrossRef]
- Lin, W.-L.; Lewis, J.; Yen, S.-H.; Hutton, M.; Dickson, D.W. Ultrastructural neuronal pathology in transgenic mice expressing mutant (P301L) human tau. J. Neurocytol. 2003, 32, 1091–1105. [Google Scholar] [CrossRef] [PubMed]
- Keck, S.; Nitsch, R.; Grune, T.; Ullrich, O. Proteasome inhibition by paired helical filament-tau in brains of patients with Alzheimer’s disease. J. Neurochem. 2003, 85, 115–122. [Google Scholar] [CrossRef]
- Lasagna-Reeves, C.A.; Castillo-Carranza, D.L.; Sengupta, U.; Clos, A.L.; Jackson, G.R.; Kayed, R. Tau oligomers impair memory and induce synaptic and mitochondrial dysfunction in wild-type mice. Mol. Neurodegener. 2011, 6, 39. [Google Scholar] [CrossRef]
- Stamer, K.; Vogel, R.; Thies, E.; Mandelkow, E.-M. Tau blocks traffic of organelles, neurofilaments, and APP vesicles in neurons and enhances oxidative stress. J. Cell Biol. 2002, 156, 1051–1063. [Google Scholar] [CrossRef] [PubMed]
- Eckert, A.; Schmitt, K.; Götz, J. Mitochondrial dysfunction—The beginning of the end in Alzheimer’s disease? Separate and synergistic modes of tau and amyloid-β toxicity. Alzheimer’s Res. Ther. 2011, 3, 1–11. [Google Scholar] [CrossRef]
- Maccioni, R.B.; Perry, G. Current Hypotheses and Research Milestones in Alzheimer’s Disease; Springer: New York, NY, USA, 2009; Volume IX, 247p. [Google Scholar]
- Cowan, C.M.; Mudher, A. Are tau aggregates toxic or protective in tauopathies? Front. Neurol. 2013, 4, 114. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, A.; Hoerndli, F.J.; Baechi, T.; Nitsch, R.M.; Götz, J. β-Amyloid Induces Paired Helical Filament-like Tau Filaments in Tissue Culture. J. Biol. Chem. 2003, 278, 40162–40168. [Google Scholar] [CrossRef]
- Smith, M.A.; Casadesus, G.; Joseph, J.A.; Perry, G. Amyloid-β and tau serve antioxidant functions in the aging and Alzheimer brain. Free Radic. Biol. Med. 2002, 33, 1194–1199. [Google Scholar] [CrossRef]
- Melov, S.; Adlard, P.A.; Morten, K.; Johnson, F.; Golden, T.R.; Hinerfeld, D.; Schilling, B.; Mavros, C.; Masters, C.L.; Volitakis, I.; et al. Mitochondrial Oxidative Stress Causes Hyperphosphorylation of Tau. PLoS ONE 2007, 2, e536. [Google Scholar] [CrossRef] [PubMed]
- Nunomura, A.; Perry, G.; Aliev, G.; Hirai, K.; Takeda, A.; Balraj, E.K.; Jones, P.K.; Ghanbari, H.; Wataya, T.; Shimohama, S.; et al. Oxidative Damage Is the Earliest Event in Alzheimer Disease. J. Neuropathol. Exp. Neurol. 2001, 60, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Carrillo-Mora, P.; Luna, R.; Colin-Barenque, L. Amyloid β: Multiple mechanisms of toxicity and only some protective effects? Oxidative Med. Cell. Longev. 2014, 2014, 795375. [Google Scholar] [CrossRef]
- Colom-Cadena, M.; the Synaptic Health Endpoints Working Group; Spires-Jones, T.; Zetterberg, H.; Blennow, K.; Caggiano, A.; DeKosky, S.T.; Fillit, H.; Harrison, J.E.; Schneider, L.S.; et al. The clinical promise of biomarkers of synapse damage or loss in Alzheimer’s disease. Alzheimer Res. Ther. 2020, 12, 1–12. [Google Scholar] [CrossRef]
- DeKosky, S.T.; Scheff, S.W. Synapse loss in frontal cortex biopsies in Alzheimer’s disease: Correlation with cognitive severity. Ann. Neurol. 1990, 27, 457–464. [Google Scholar] [CrossRef] [PubMed]
- De Wilde, M.C.; Overk, C.R.; Sijben, J.W.; Masliah, E. Meta-analysis of synaptic pathology in Alzheimer’s disease reveals selective molecular vesicular machinery vulnerability. Alzheimers Dement. 2016, 12, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Małkiewicz, M.A.; Szarmach, A.; Sabisz, A.; Cubała, W.J.; Szurowska, E.; Winklewski, P.J. Blood-brain barrier permeability and physical exercise. J. Neuroinflamm. 2019, 16, 1–16. [Google Scholar] [CrossRef]
- Reese, T.S.; Karnovsky, M.J. Fine structural localization of a blood-brain barrier to exogenous peroxidase. J. Cell Biol. 1967, 34, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Daneman, R.; Prat, A. The blood-brain barrier. Cold Spring Harb. Perspect. Biol. 2015, 7, a020412. [Google Scholar] [CrossRef]
- Oldendorf, W.H.; Cornford, M.E.; Brown, W.J. The large apparent work capability of the blood-brain barrier: A study of the mitochondrial content of capillary endothelial cells in brain and other tissues of the rat. Ann. Neurol. 1977, 1, 409–417. [Google Scholar] [CrossRef]
- Zlokovic, B.V. Neurovascular mechanisms of Alzheimer’s neurodegeneration. Trends Neurosci. 2005, 28, 202–208. [Google Scholar] [CrossRef]
- Korte, N.; Nortley, R.; Attwell, D. Cerebral blood flow decrease as an early pathological mechanism in Alzheimer’s disease. Acta Neuropathol. 2020, 140, 793–810. [Google Scholar] [CrossRef]
- Attwell, D.; Laughlin, S.B. An Energy Budget for Signaling in the Grey Matter of the Brain. J. Cereb. Blood Flow Metab. 2001, 21, 1133–1145. [Google Scholar] [CrossRef] [PubMed]
- Fowler, J.C. Adenosine antagonists alter the synaptic response to in vitro ischemia in the rat hippocampus. Brain Res. 1990, 509, 331–334. [Google Scholar] [CrossRef]
- Hansra, G.K.; Popov, G.; Banaczek, P.O.; Vogiatzis, M.; Jegathees, T.; Goldsbury, C.S.; Cullen, K.M. The neuritic plaque in Alzheimer’s disease: Perivascular degeneration of neuronal processes. Neurobiol. Aging 2019, 82, 88–101. [Google Scholar] [CrossRef]
- Chow, N.; Bell, R.D.; Deane, R.; Streb, J.W.; Chen, J.; Brooks, A.; Van Nostrand, W.; Miano, J.M.; Zlokovic, B.V. Serum response factor and myocardin mediate arterial hypercontractility and cerebral blood flow dysregulation in Alzheimer’s phenotype. Proc. Natl. Acad. Sci. USA 2007, 104, 823–828. [Google Scholar] [CrossRef] [PubMed]
- Frost, G.R.; Jonas, L.A.; Li, Y.M. Friend, Foe or Both? Immune Activity in Alzheimer’s Disease. Front. Aging Neurosci. 2019, 11, 337. [Google Scholar] [CrossRef] [PubMed]
- Heneka, M.T.; Carson, M.J.; Khoury, J.E.; Landreth, G.E.; Brosseron, F.; Feinstein, D.L.; Jacobs, A.H.; Wyss-Coray, T.; Vitorica, J.; Ransohoff, R.M.; et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 2015, 14, 388–405. [Google Scholar] [CrossRef]
- Zhang, B.; Gaiteri, C.; Bodea, L.-G.; Wang, Z.; McElwee, J.; Podtelezhnikov, A.A.; Zhang, C.; Xie, T.; Tran, L.; Dobrin, R.; et al. Integrated Systems Approach Identifies Genetic Nodes and Networks in Late-Onset Alzheimer’s Disease. Cell 2013, 153, 707–720. [Google Scholar] [CrossRef]
- Cribbs, D.H.; Berchtold, N.C.; Perreau, V.; Coleman, P.D.; Rogers, J.; Tenner, A.J.; Cotman, C.W. Extensive innate immune gene activation accompanies brain aging, increasing vulnerability to cognitive decline and neurodegeneration: A microarray study. J. Neuroinflamm. 2012, 9, 179. [Google Scholar] [CrossRef] [PubMed]
- Brosseron, F.; Krauthausen, M.; Kummer, M.; Heneka, M.T. Body Fluid Cytokine Levels in Mild Cognitive Impairment and Alzheimer’s Disease: A Comparative Overview. Mol. Neurobiol. 2014, 50, 534–544. [Google Scholar] [CrossRef] [PubMed]
- Leng, F.; Edison, P. Neuroinflammation and microglial activation in Alzheimer disease: Where do we go from here? Nat. Rev. Neurol. 2020, 1–16. [Google Scholar] [CrossRef]
- Xiang, Z.; Haroutunian, V.; Ho, L.; Purohit, D.; Pasinetti, G.M. Microglia Activation in the Brain as Inflammatory Biomarker of Alzheimer’s Disease Neuropathology and Clinical Dementia. Dis. Markers 2006, 22, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Heppner, F.L.; Ransohoff, R.M.; Becher, B. Immune attack: The role of inflammation in Alzheimer disease. Nat. Rev. Neurosci. 2015, 16, 358–372. [Google Scholar] [CrossRef]
- Qin, L.; Liu, Y.; Cooper, C.; Liu, B.; Wilson, B.; Hong, J.-S. Microglia enhance β-amyloid peptide-induced toxicity in cortical and mesencephalic neurons by producing reactive oxygen species. J. Neurochem. 2002, 83, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Block, M.L.; Zecca, L.; Hong, J.-S. Microglia-mediated neurotoxicity: Uncovering the molecular mechanisms. Nat. Rev. Neurosci. 2007, 8, 57–69. [Google Scholar] [CrossRef]
- Hickman, S.E.; Allison, E.K.; El Khoury, J. Microglial dysfunction and defective β-amyloid clearance pathways in aging Alzheimer’s disease mice. J. Neurosci. 2008, 28, 8354–8360. [Google Scholar] [CrossRef]
- He, X.F.; Xu, J.-H.; Li, G.; Li, M.-Y.; Li, L.-L.; Pei, Z.; Zhang, L.-Y.; Hu, X.-Q. NLRP3-dependent microglial training impaired the clearance of amyloid-β and aggravated the cognitive decline in Alzheimer’s disease. Cell Death Dis. 2020, 11, 849. [Google Scholar] [CrossRef]
- McGeer, P.; Akiyama, H.; Itagaki, S.; McGeer, E. Activation of the classical complement pathway in brain tissue of Alzheimer patients. Neurosci. Lett. 1989, 107, 341–346. [Google Scholar] [CrossRef]
- Morgan, B.P. Complement in the pathogenesis of Alzheimer’s disease. Semin. Immunopathol. 2018, 40, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Dejanovic, B.; Gandham, V.D.; Gogineni, A.; Edmonds, R.; Schauer, S.; Srinivasan, K.; Huntley, M.A.; Wang, Y.; Wang, T.-M.; et al. Complement C3 Is Activated in Human AD Brain and Is Required for Neurodegeneration in Mouse Models of Amyloidosis and Tauopathy. Cell Rep. 2019, 28, 2111–2123.e6. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.-H.; Yu, J.-T.; Tan, L. The Role of Cholesterol Metabolism in Alzheimer’s Disease. Mol. Neurobiol. 2015, 51, 947–965. [Google Scholar] [CrossRef]
- Fünfschilling, U.; Saher, G.; Xiao, L.; Möbius, W.; Nave, K.-A. Survival of adult neurons lacking cholesterol synthesis in vivo. BMC Neurosci. 2007, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.-T.; Tan, L.; Hardy, J. Apolipoprotein E in Alzheimer’s Disease: An Update. Annu. Rev. Neurosci. 2014, 37, 79–100. [Google Scholar] [CrossRef]
- Vance, J.E.; Hayashi, H.; Karten, B. Cholesterol homeostasis in neurons and glial cells. Semin. Cell Dev. Biol. 2005, 16, 193–212. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liu, Q. Cholesterol metabolism and homeostasis in the brain. Protein Cell 2015, 6, 254–264. [Google Scholar] [CrossRef]
- Popp, J.; Meichsner, S.; Kölsch, H.; Lewczuk, P.; Maier, W.; Kornhuber, J.; Jessen, F.; Lütjohann, D. Cerebral and extracerebral cholesterol metabolism and CSF markers of Alzheimer’s disease. Biochem. Pharmacol. 2013, 86, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Kivipelto, M.; Helkala, E.-L.; Soininen, H.; Laakso, M.P.; Hänninen, T.; Hallikainen, M.; Alhainen, K.; Iivonen, S.; Mannermaa, A.; Tuomilehto, J.; et al. Apolipoprotein E ε4 Allele, Elevated Midlife Total Cholesterol Level, and High Midlife Systolic Blood Pressure Are Independent Risk Factors for Late-Life Alzheimer Disease. Ann. Intern. Med. 2002, 137, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Helzner, E.P.; Luchsinger, J.A.; Scarmeas, N.; Cosentino, S.; Brickman, A.M.; Glymour, M.M.; Stern, Y. Contribution of Vascular Risk Factors to the Progression in Alzheimer Disease. Arch. Neurol. 2009, 66, 343–348. [Google Scholar] [CrossRef]
- Chen, X.; Gawryluk, J.W.; Wagener, J.F.; Ghribi, O.; Geiger, J.D. Caffeine blocks disruption of blood brain barrier in a rabbit model of Alzheimer’s disease. J. Neuroinflammation 2008, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Beel, A.J.; Sakakura, M.; Barrett, P.J.; Sanders, C.R. Direct binding of cholesterol to the amyloid precursor protein: An important interaction in lipid-Alzheimer’s disease relationships? Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2010, 1801, 975–982. [Google Scholar] [CrossRef] [PubMed]
- Lall, R.; Mohammed, R.; Ojha, U. What are the links between hypoxia and Alzheimer’s disease? Neuropsychiatry Dis. Treat. 2019, 15, 1343–1354. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; He, G.; Qing, H.; Zhou, W.; Dobie, F.; Cai, F.; Staufenbiel, M.; Huang, L.E.; Song, W. Hypoxia facilitates Alzheimer’s disease pathogenesis by up-regulating BACE1 gene expression. Proc. Natl. Acad. Sci. USA 2006, 103, 18727–18732. [Google Scholar] [CrossRef] [PubMed]
- Kerridge, C.; Kozlova, D.I.; Nalivaeva, N.N.; Turner, A.J. Hypoxia Affects Neprilysin Expression Through Caspase Activation and an APP Intracellular Domain-dependent Mechanism. Front Neurosci 2015, 9, 426. [Google Scholar] [CrossRef]
- Peers, C.; Dallas, M.L.; Boycott, H.E.; Scragg, J.L.; Pearson, H.A.; Boyle, J.P. Hypoxia and Neurodegeneration. Ann. N. Y. Acad. Sci. 2009, 1177, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Qiu, C.; Kivipelto, M.; Von Strauss, E. Epidemiology of Alzheimer’s disease: Occurrence, determinants, and strategies toward intervention. Dialogues Clin. Neurosci. 2009, 11, 111–128. [Google Scholar] [PubMed]
- Kalaria, R.N.; Maestre, G.E.; Arizaga, R.; Friedland, R.P.; Galasko, D.; Hall, K.; Luchsinger, J.A.; Ogunniyi, A.; Perry, E.K.; Potocnik, F.; et al. Alzheimer’s disease and vascular dementia in developing countries: Prevalence, management, and risk factors. Lancet Neurol. 2008, 7, 812–826. [Google Scholar] [CrossRef]
- Launer, L.J.; Rossb, G.W.; Petrovitch, H.; Masaki, K.; Foley, D.; White, L.R.; Havlik, R.J. Midlife blood pressure and dementia: The Honolulu-Asia aging study. Neurobiol. Aging 2000, 21, 49–55. [Google Scholar] [CrossRef]
- Kivipelto, M.; Helkala, E.-L.; Laakso, M.P.; Hänninen, T.; Hallikainen, M.; Alhainen, K.; Soininen, H.; Tuomilehto, J.; Nissinen, A. Midlife vascular risk factors and Alzheimer’s disease in later life: Longitudinal, population based study. BMJ 2001, 322, 1447–1451. [Google Scholar] [CrossRef]
- Ruitenberg, A.; Heijer, T.D.; Bakker, S.L.M.; Van Swieten, J.C.; Koudstaal, P.J.; Hofman, A.; Breteler, M.M.B. Cerebral hypoperfusion and clinical onset of dementia: The Rotterdam study. Ann. Neurol. 2005, 57, 789–794. [Google Scholar] [CrossRef]
- Snowdon, D.A.; Greiner, L.H.; Mortimer, J.A.; Riley, K.P.; Greiner, P.A.; Markesbery, W.R. Brain infarction and the clinical expression of Alzheimer disease. Nun. Study JAMA 1997, 277, 813–817. [Google Scholar] [CrossRef] [PubMed]
- Picone, P.; Nuzzo, D.; Caruana, L.; Scafidi, V.; Di Carlo, M. Mitochondrial Dysfunction: Different Routes to Alzheimer’s Disease Therapy. Oxidative Med. Cell. Longev. 2014, 2014, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Devine, M.J.; Kittler, J.T. Mitochondria at the neuronal presynapse in health and disease. Nat. Rev. Neurosci. 2018, 19, 63–80. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Boyer, L.; Jin, M.; Mertens, J.; Kim, Y.; Mandel, G.; Hamm, M.; Gage, F.H.; Hunter, T. Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation. eLife 2016, 5, 5. [Google Scholar] [CrossRef] [PubMed]
- Mattson, M.P.; Partin, J. Evidence for mitochondrial control of neuronal polarity. J. Neurosci. Res. 1999, 56, 8–20. [Google Scholar] [CrossRef]
- Mattson, M.P.; Gleichmann, M.; Cheng, A. Mitochondria in Neuroplasticity and Neurological Disorders. Neuron 2008, 60, 748–766. [Google Scholar] [CrossRef]
- Billups, B.; Forsythe, I.D. Presynaptic Mitochondrial Calcium Sequestration Influences Transmission at Mammalian Central Synapses. J. Neurosci. 2002, 22, 5840–5847. [Google Scholar] [CrossRef] [PubMed]
- Cha, M.-Y.; Han, S.-H.; Son, S.M.; Hong, H.-S.; Choi, Y.-J.; Byun, J.; Mook-Jung, I. Mitochondria-Specific Accumulation of Amyloid β Induces Mitochondrial Dysfunction Leading to Apoptotic Cell Death. PLoS ONE 2012, 7, e34929. [Google Scholar] [CrossRef]
- Anandatheerthavarada, H.K.; Biswas, G.; Robin, M.-A.; Avadhani, N.G. Mitochondrial targeting and a novel transmembrane arrest of Alzheimer’s amyloid precursor protein impairs mitochondrial function in neuronal cells. J. Cell Biol. 2003, 161, 41–54. [Google Scholar] [CrossRef] [PubMed]
- Caspersen, C.; Wang, N.; Yao, J.; Sosunov, A.; Chen, X.; Lustbader, J.W.; Xu, H.W.; Stern, D.; McKhann, G.; Du Yan, S. Mitochondrial Aβ: A potential focal point for neuronal metabolic dysfunction in Alzheimer’s disease. FASEB J. 2005, 19, 2040–2041. [Google Scholar] [CrossRef] [PubMed]
- Gabuzda, D.; Busciglio, J.; Chen, L.; Matsudaira, P.; Yankner, B. Inhibition of energy metabolism alters the processing of amyloid precursor protein and induces a potentially amyloidogenic derivative. J. Biol. Chem. 1994, 269, 13623–13628. [Google Scholar] [CrossRef]
- Gasparini, L.; Racchi, M.; Benussi, L.; Curti, D.; Binetti, G.; Bianchetti, A.; Trabucchi, M.; Govoni, S. Effect of energy shortage and oxidative stress on amyloid precursor protein metabolism in COS cells. Neurosci. Lett. 1997, 231, 113–117. [Google Scholar] [CrossRef]
- Lin, M.T. High aggregate burden of somatic mtDNA point mutations in aging and Alzheimer’s disease brain. Hum. Mol. Genet. 2002, 11, 133–145. [Google Scholar] [CrossRef]
- Swerdlow, R.H.; Burns, J.M.; Khan, S.M. The Alzheimer’s disease mitochondrial cascade hypothesis: Progress and perspectives. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2014, 1842, 1219–1231. [Google Scholar] [CrossRef] [PubMed]
- Manczak, M.; Park, B.S.; Jung, Y.; Reddy, P.H. Differential Expression of Oxidative Phosphorylation Genes in Patients with Alzheimer’s Disease: Implications for Early Mitochondrial Dysfunction and Oxidative Damage. NeuroMol. Med. 2004, 5, 147–162. [Google Scholar] [CrossRef]
- Mecocci, P.; MacGarvey, U.; Beal, M.F. Oxidative damage to mitochondrial DNA is increased in Alzheimer’s disease. Ann. Neurol. 1994, 36, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Sun, L.; Chen, X.; Zhang, D. Oxidative stress, mitochondrial damage and neurodegenerative diseases. Neural Regen. Res. 2013, 8, 2003–2014. [Google Scholar] [PubMed]
- LaRosa, V.; Remacle, C. Insights into the respiratory chain and oxidative stress. Biosci. Rep. 2018, 38, 38. [Google Scholar] [CrossRef]
- Tönnies, E.; Trushina, E. Oxidative Stress, Synaptic Dysfunction, and Alzheimer’s Disease. J. Alzheimers Dis. 2017, 57, 1105–1121. [Google Scholar] [CrossRef] [PubMed]
- Murphy, M.P. How mitochondria produce reactive oxygen species. Biochem. J. 2009, 417, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Moreira, P.I.; Carvalho, C.; Zhu, X.; Smith, M.A.; Perry, G. Mitochondrial dysfunction is a trigger of Alzheimer’s disease pathophysiology. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2010, 1802, 2–10. [Google Scholar] [CrossRef]
- Mark, R.J.; Lovell, M.A.; Markesbery, W.R.; Uchida, K.; Mattson, M.P. A Role for 4-Hydroxynonenal, an Aldehydic Product of Lipid Peroxidation, in Disruption of Ion Homeostasis and Neuronal Death Induced by Amyloid β-Peptide. J. Neurochem. 1997, 68, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Ansari, M.A.; Abdul, H.M.; Joshi, G.; Opii, W.O.; Butterfield, D.A. Protective effect of quercetin in primary neurons against Aβ(1-42): Relevance to Alzheimer’s disease. J. Nutr. Biochem. 2009, 20, 269–275. [Google Scholar] [CrossRef] [PubMed]
- Butterfield, D.A.; Castegna, A.; Lauderback, C.M.; Drake, J. Evidence that amyloid β-peptide-induced lipid peroxidation and its sequelae in Alzheimer’s disease brain contribute to neuronal death. Neurobiol. Aging 2002, 23, 655–664. [Google Scholar] [CrossRef]
- Mark, R.J.; Fuson, K.S.; May, P.C. Characterization of 8-Epiprostaglandin F2α as a Marker of Amyloid β-Peptide-Induced Oxidative Damage. J. Neurochem. 1999, 72, 1146–1153. [Google Scholar] [CrossRef] [PubMed]
- Sultana, R.; Perluigi, M.; Butterfield, D.A. Oxidatively modified proteins in Alzheimer’s disease (AD), mild cognitive impairment and animal models of AD: Role of Aβ in pathogenesis. Acta Neuropathol. 2009, 118, 131–150. [Google Scholar] [CrossRef] [PubMed]
- Boyd-Kimball, D.; Sultana, R.; Poon, H.F.; Mohmmad-Abdul, H.; Lynn, B.C.; Klein, J.B.; Butterfield, D.A. γ-glutamylcysteine ethyl ester protection of proteins from Aβ(1-42)-mediated oxidative stress in neuronal cell culture: A proteomics approach. J. Neurosci. Res. 2005, 79, 707–713. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhong, C. Decoding Alzheimer’s disease from perturbed cerebral glucose metabolism: Implications for diagnostic and therapeutic strategies. Prog. Neurobiol. 2013, 108, 21–43. [Google Scholar] [CrossRef]
- Sokoloff, L. Energetics of Functional Activation in Neural Tissues. Neurochem. Res. 1999, 24, 321–329. [Google Scholar] [CrossRef] [PubMed]
- Simpson, I.A.; Carruthers, A.; Vannucci, S.J. Supply and Demand in Cerebral Energy Metabolism: The Role of Nutrient Transporters. J. Cereb. Blood Flow Metab. 2007, 27, 1766–1791. [Google Scholar] [CrossRef] [PubMed]
- Duelli, R.; Kuschinsky, W. Brain Glucose Transporters: Relationship to Local Energy Demand. Physiology 2001, 16, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Kuzuya, T. Outline of glucose metabolism and its regulations. Nihon Rinsho. Jpn. J. Clin. Med. 1990, 48, 51–59. [Google Scholar]
- Mosconi, L. Brain glucose metabolism in the early and specific diagnosis of Alzheimer’s disease. FDG-PET studies in MCI and AD. Eur. J. Nucl. Med. Mol. Imaging 2005, 32, 486–510. [Google Scholar] [CrossRef]
- Mosconi, L.; Pupi, A.; De Leon, M.J. Brain Glucose Hypometabolism and Oxidative Stress in Preclinical Alzheimer’s Disease. Ann. N. Y. Acad. Sci. 2008, 1147, 180–195. [Google Scholar] [CrossRef] [PubMed]
- Petersen, R.C.; Smith, G.E.; Waring, S.C.; Ivnik, R.J.; Tangalos, E.G.; Kokmen, E. Mild cognitive impairment: Clinical characterization and outcome. Arch. Neurol. 1999, 56, 303–308. [Google Scholar] [CrossRef]
- Lee, T.; Lee, H. Shared Blood Transcriptomic Signatures between Alzheimer’s Disease and Diabetes Mellitus. Biomedicines 2021, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Simpson, I.A.; Chundu, K.R.; Davies-Hill, T.; Honer, W.G.; Davies, P. Decreased concentrations of GLUT1 and GLUT3 glucose transporters in the brains of patients with Alzheimer’s disease. Ann. Neurol. 1994, 35, 546–551. [Google Scholar] [CrossRef]
- Liu, G.; Huang, W.; Moir, R.D.; Vanderburg, C.R.; Laif, B.; Peng, Z.; Tanzi, R.E.; Rogers, J.T.; Huang, X. Metal exposure and Alzheimer’s pathogenesis. J. Struct. Biol. 2006, 155, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Greenough, M.A.; Camakaris, J.; Bush, A.I. Metal dyshomeostasis and oxidative stress in Alzheimer’s disease. Neurochem. Int. 2013, 62, 540–555. [Google Scholar] [CrossRef]
- Lovell, M.; Robertson, J.; Teesdale, W.; Campbell, J.; Markesbery, W. Copper, iron and zinc in Alzheimer’s disease senile plaques. J. Neurol. Sci. 1998, 158, 47–52. [Google Scholar] [CrossRef]
- Huang, X.; Atwood, C.S.; Tanzi, R.E.; Bush, A.I.; Hartshorn, M.A.; Multhaup, G.; Goldstein, L.E.; Scarpa, R.C.; Cuajungco, M.P.; Gray, D.N.; et al. The Aβ Peptide of Alzheimer’s Disease Directly Produces Hydrogen Peroxide through Metal Ion Reduction. Biochemistry 1999, 38, 7609–7616. [Google Scholar] [CrossRef] [PubMed]
- Rogers, J.T.; Venkataramani, V.; Washburn, C.; Liu, Y.; Tummala, V.; Jiang, H.; Smith, A.; Cahill, C.M. A role for amyloid precursor protein translation to restore iron homeostasis and ameliorate lead (Pb) neurotoxicity. J. Neurochem. 2016, 138, 479–494. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, P.; Sahoo, A.; Anand, S.; Ganguly, A.; Righi, G.; Bovicelli, P.; Saso, L.; Chakrabarti, S. Multiple mechanisms of iron-induced amyloid β-peptide accumulation in SHSY5Y cells: Protective action of negletein. Neuromol. Med. 2014, 16, 787–798. [Google Scholar] [CrossRef]
- Hooda, J.; Shah, A.; Zhang, L. Heme, an Essential Nutrient from Dietary Proteins, Critically Impacts Diverse Physiological and Pathological Processes. Nutrients 2014, 6, 1080–1102. [Google Scholar] [CrossRef]
- Faux, N.G.; the AIBL Research Group; Rembach, A.; Wiley, J.S.; Ellis, K.A.; Ames, D.; Fowler, C.J.; Martins, R.N.; Pertile, K.K.; Rumble, R.L.; et al. An anemia of Alzheimer’s disease. Mol. Psychiatry 2014, 19, 1227–1234. [Google Scholar] [CrossRef] [PubMed]
- Hong, C.H.; Falvey, C.; Harris, T.B.; Simonsick, E.M.; Satterfield, S.; Ferrucci, L.; Metti, A.L.; Patel, K.V.; Yaffe, K. Anemia and risk of dementia in older adults: Findings from the Health ABC study. Neurology 2013, 81, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Atamna, H.; Walter, P.B.; Ames, B.N. The Role of Heme and Iron-Sulfur Clusters in Mitochondrial Biogenesis, Maintenance, and Decay with Age. Arch. Biochem. Biophys. 2002, 397, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Shetty, T.; Corson, T.W. Mitochondrial Heme Synthesis Enzymes as Therapeutic Targets in Vascular Diseases. Front. Pharmacol. 2020, 11, 1015. [Google Scholar] [CrossRef] [PubMed]
- FitzGerald, K.E.; Lal, S.; Kalainayakan, S.P.; Zhang, L. Molecular Mechanisms Underlying Heme Action in Promoting the Pathogenesis of Alzheimer’s Disease. In Alzheimer’s Disease; SMGroup: Dover, DE, USA, 2016. [Google Scholar]
- Gozzelino, R. The Pathophysiology of Heme in the Brain. Curr. Alzheimer Res. 2016, 13, 174–184. [Google Scholar] [CrossRef] [PubMed]
- Chiabrando, D.; Fiorito, V.; Petrillo, S.; Tolosano, E. Unraveling the Role of Heme in Neurodegeneration. Front. Neurosci. 2018, 12, 712. [Google Scholar] [CrossRef] [PubMed]
- Rajagopal, A.; Rao, A.U.; Krause, M.; Hamza, I.; Amigo, J.; Tian, M.; Upadhyay, S.K.; Hall, C.; Uhm, S.; Mathew, M.K.; et al. Haem homeostasis is regulated by the conserved and concerted functions of HRG-1 proteins. Nature 2008, 453, 1127–1131. [Google Scholar] [CrossRef]
- Duffy, S.P.; Shing, J.; Saraon, P.; Berger, L.C.; Eiden, M.V.; Wilde, A.; Tailor, C.S. The Fowler Syndrome-Associated Protein FLVCR2 Is an Importer of Heme. Mol. Cell. Biol. 2010, 30, 5318–5324. [Google Scholar] [CrossRef]
- Castro-Gago, M.; Alonso, A.; Pintos-Martinez, E.; Beiras-Iglesias, A.; Campos, Y.; Arenas, J.; Novo-Rodriguez, M.I.; Eiris-Punal, J. Congenital hydranencephalic-hydrocephalic syndrome associated with mitochondrial dysfunction. J. Child Neurol. 1999, 14, 131–135. [Google Scholar] [CrossRef]
- Zhu, Y.; Hon, T.; Ye, W.; Zhang, L. Heme deficiency interferes with the Ras-mitogen-activated protein kinase signaling pathway and expression of a subset of neuronal genes. Cell Growth Differ. Mol. Boil. J. Am. Assoc. Cancer Res. 2002, 13, 431–439. [Google Scholar]
- Smith, A.G.; Raven, E.L.; Chernova, T. The regulatory role of heme in neurons. Metallomics 2011, 3, 955–962. [Google Scholar] [CrossRef]
- Kim, H.J.; Khalimonchuk, O.; Smith, P.M.; Winge, D.R. Structure, function, and assembly of heme centers in mitochondrial respiratory complexes. Biochim. Biophys. Acta (BBA) Bioenerg. 2012, 1823, 1604–1616. [Google Scholar] [CrossRef] [PubMed]
- Atamna, H.; Frey, W.H. Mechanisms of mitochondrial dysfunction and energy deficiency in Alzheimer’s disease. Mitochondrion 2007, 7, 297–310. [Google Scholar] [CrossRef]
- Sankar, S.B.; Donegan, R.K.; Shah, K.J.; Reddi, A.R.; Wood, L.B. Heme and hemoglobin suppress amyloid β-mediated inflammatory activation of mouse astrocytes. J. Biol. Chem. 2018, 293, 11358–11373. [Google Scholar] [CrossRef]
- Atamna, H.; Liu, J.; Ames, B.N. Heme deficiency selectively interrupts assembly of mitochondrial complex IV in human fibroblasts: Revelance to aging. J. Biol. Chem. 2001, 276, 48410–48416. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Choi, Y.K. Regenerative Effects of Heme Oxygenase Metabolites on Neuroinflammatory Diseases. Int. J. Mol. Sci. 2018, 20, 78. [Google Scholar] [CrossRef]
- Jofre-Monseny, L.; Loboda, A.; Wagner, A.E.; Huebbe, P.; Boesch-Saadatmandi, C.; Jozkowicz, A.; Minihane, A.-M.; Dulak, J.; Rimbach, G. Effects of apoE genotype on macrophage inflammation and heme oxygenase-1 expression. Biochem. Biophys. Res. Commun. 2007, 357, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Schipper, H.M. Heme oxygenase-1 in Alzheimer disease: A tribute to Moussa Youdim. J. Neural Transm. 2011, 118, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Nitti, M.; Piras, S.; Brondolo, L.; Marinari, U.M.; Pronzato, M.A.; Furfaro, A.L. Heme Oxygenase 1 in the Nervous System: Does It Favor Neuronal Cell Survival or Induce Neurodegeneration? Int. J. Mol. Sci. 2018, 19, 2260. [Google Scholar] [CrossRef] [PubMed]
- Barone, E.; Di Domenico, F.; Cassano, T.; Arena, A.; Tramutola, A.; Lavecchia, M.A.; Coccia, R.; Butterfield, D.A.; Perluigi, M. Impairment of biliverdin reductase-A promotes brain insulin resistance in Alzheimer disease: A new paradigm. Free. Radic. Biol. Med. 2016, 91, 127–142. [Google Scholar] [CrossRef]
- Takahashi, M.; Doré, S.; Ferris, C.D.; Tomita, T.; Sawa, A.; Wolosker, H.; Borchelt, D.R.; Iwatsubo, T.; Kim, S.-H.; Thinakaran, G.; et al. Amyloid Precursor Proteins Inhibit Heme Oxygenase Activity and Augment Neurotoxicity in Alzheimer’s Disease. Neuron 2000, 28, 461–473. [Google Scholar] [CrossRef]
- Gordon, J.; Amini, S.; White, M.K. General overview of neuronal cell culture. Methods Mol. Biol. 2013, 1078, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sahu, M.P.; Nikkilä, O.; Lågas, S.; Kolehmainen, S.; Castrén, E. Culturing primary neurons from rat hippocampus and cortex. Neuronal Signal. 2019, 3, NS20180207. [Google Scholar] [CrossRef] [PubMed]
- Westerink, R.H.S.; Ewing, A.G. The PC12 cell as model for neurosecretion. Acta Physiol. 2008, 192, 273–285. [Google Scholar] [CrossRef]
- Wiatrak, B.; Kubis-Kubiak, A.; Piwowar, A.; Barg, E. PC12 Cell Line: Cell Types, Coating of Culture Vessels, Differentiation and Other Culture Conditions. Cells 2020, 9, 958. [Google Scholar] [CrossRef] [PubMed]
- Biedler, J.L.; Helson, L.; Spengler, B.A. Morphology and growth, tumorigenicity, and cytogenetics of human neuroblastoma cells in continuous culture. Cancer Res. 1973, 33, 2643–2652. [Google Scholar] [PubMed]
- Kobayashi, M.; Kurihara, K.; Matsuoka, I. Retinoic acid induces BDNF responsiveness of sympathetic neurons by alteration of Trk neurotrophin receptor expression. FEBS Lett. 1994, 356, 60–65. [Google Scholar] [CrossRef]
- Price, R.D.; Oe, T.; Yamaji, T.; Matsuoka, N. A Simple, Flexible, Nonfluorescent System for the Automated Screening of Neurite Outgrowth. J. Biomol. Screen 2006, 11, 155–164. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ambjørn, M.; Dubreuil, V.; Miozzo, F.; Nigon, F.; Møller, B.; Issazadeh-Navikas, S.; Berg, J.; Lees, M.; Sap, J. A Loss-of-Function Screen for Phosphatases that Regulate Neurite Outgrowth Identifies PTPN12 as a Negative Regulator of TrkB Tyrosine Phosphorylation. PLoS ONE 2013, 8, e65371. [Google Scholar] [CrossRef] [PubMed]
- Agholme, L.; Lindström, T.; Kågedal, K.; Marcusson, J.; Hallbeck, M. An In Vitro Model for Neuroscience: Differentiation of SH-SY5Y Cells into Cells with Morphological and Biochemical Characteristics of Mature Neurons. J. Alzheimer’s Dis. 2010, 20, 1069–1082. [Google Scholar] [CrossRef]
- Kovalevich, J.; Langford, D. Considerations for the Use of SH-SY5Y Neuroblastoma Cells in Neurobiology. Methods Mol. Biol. 2013, 1078, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Krishtal, J.; Bragina, O.; Metsla, K.; Palumaa, P.; Tõugu, V. In situ fibrillizing amyloid-β1-42 induces neurite degeneration and apoptosis of differentiated SH-SY5Y cells. PLoS ONE 2017, 12, e0186636. [Google Scholar] [CrossRef] [PubMed]
- Gilson, V.; Mbebi-Liegeois, C.; Sellal, F.; de Barry, J. Effects of Low Amyloid-β (Aβ) Concentration on Aβ1-42 Oligomers Binding and GluN2B Membrane Expression. J. Alzheimers Dis. 2015, 47, 453–466. [Google Scholar] [CrossRef] [PubMed]
- Lopez Sanchez, M.I.G.; Waugh, H.S.; Tsatsanis, A.; Wong, B.X.; Crowston, J.G.; Duce, J.A.; Trounce, I.A. Amyloid precursor protein drives down-regulation of mitochondrial oxidative phosphorylation independent of amyloid β. Sci. Rep. 2017, 7, 9835. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Jiao, B.; Zhou, M.; Zhou, T.; Shen, L. Modeling Alzheimer’s Disease with Induced Pluripotent Stem Cells: Current Challenges and Future Concerns. Stem Cells Int. 2016, 2016, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Gunhanlar, N.; Shpak, G.; Van Der Kroeg, M.; Gouty-Colomer, L.; Munshi, S.T.; Lendemeijer, B.; Ghazvini, M.; Dupont, C.; Hoogendijk, W.J.G.; Gribnau, J.; et al. A simplified protocol for differentiation of electrophysiologically mature neuronal networks from human induced pluripotent stem cells. Mol. Psychiatry 2018, 23, 1336–1344. [Google Scholar] [CrossRef] [PubMed]
- Kondo, T.; Asai, M.; Tsukita, K.; Kutoku, Y.; Ohsawa, Y.; Sunada, Y.; Imamura, K.; Egawa, N.; Yahata, N.; Okita, K.; et al. Modeling Alzheimer’s Disease with iPSCs Reveals Stress Phenotypes Associated with Intracellular Aβ and Differential Drug Responsiveness. Cell Stem Cell 2013, 12, 487–496. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Najm, R.; Xu, Q.; Jeong, D.-E.; Walker, D.; Balestra, M.E.; Yoon, S.Y.; Yuan, H.; Li, G.; Miller, Z.A.; et al. Gain of toxic apolipoprotein E4 effects in human iPSC-derived neurons is ameliorated by a small-molecule structure corrector. Nat. Med. 2018, 24, 647–657. [Google Scholar] [CrossRef]
- Díaz-Guerra, E.; Moreno-Jiménez, E.P.; De Rojas, I.; Rodríguez, C.; Rodríguez-Traver, E.; Arribas-González, E.; Orera, M.; Hernández, I.; Ruiz, A.; Vicario, C. A collection of four integration-free iPSC lines derived from diagnosed sporadic Alzheimer’s disease patients with different APOE alleles. Stem Cell Res. 2019, 39, 101522. [Google Scholar] [CrossRef]
- Sierksma, A.; Escott-Price, V.; De Strooper, B. Translating genetic risk of Alzheimer’s disease into mechanistic insight and drug targets. Science 2020, 370, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Elder, G.A.; Gama Sosa, M.A.; de Gasperi, R. Transgenic mouse models of Alzheimer’s disease. Mt. Sinai J. Med. 2010, 77, 69–81. [Google Scholar] [CrossRef] [PubMed]
- Radde, R.; Bolmont, T.; Kaeser, S.A.; Coomaraswamy, J.; Lindau, D.; Stoltze, L.; Calhoun, M.E.; Jäggi, F.; Wolburg, H.; Gengler, S.; et al. Aβ42-driven cerebral amyloidosis in transgenic mice reveals early and robust pathology. EMBO Rep. 2006, 7, 940–946. [Google Scholar] [CrossRef] [PubMed]
- Sasaguri, H.; Nilsson, P.; Hashimoto, S.; Nagata, K.; Saito, T.; De Strooper, B.; Hardy, J.; Vassar, R.; Winblad, B.; Saido, T.C. APP mouse models for Alzheimer’s disease preclinical studies. EMBO J. 2017, 36, 2473–2487. [Google Scholar] [CrossRef] [PubMed]
- Espuny-Camacho, I.M.; Arranz, A.M.; Fiers, M.M.; Snellinx, A.; Ando, K.; Munck, S.; Bonnefont, J.; Lambot, L.; Corthout, N.; Omodho, L.; et al. Hallmarks of Alzheimer’s Disease in Stem-Cell-Derived Human Neurons Transplanted into Mouse Brain. Neuron 2017, 93, 1066–1081.e8. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vidal, C.; Zhang, L. An Analysis of the Neurological and Molecular Alterations Underlying the Pathogenesis of Alzheimer’s Disease. Cells 2021, 10, 546. https://doi.org/10.3390/cells10030546
Vidal C, Zhang L. An Analysis of the Neurological and Molecular Alterations Underlying the Pathogenesis of Alzheimer’s Disease. Cells. 2021; 10(3):546. https://doi.org/10.3390/cells10030546
Chicago/Turabian StyleVidal, Chantal, and Li Zhang. 2021. "An Analysis of the Neurological and Molecular Alterations Underlying the Pathogenesis of Alzheimer’s Disease" Cells 10, no. 3: 546. https://doi.org/10.3390/cells10030546
APA StyleVidal, C., & Zhang, L. (2021). An Analysis of the Neurological and Molecular Alterations Underlying the Pathogenesis of Alzheimer’s Disease. Cells, 10(3), 546. https://doi.org/10.3390/cells10030546






