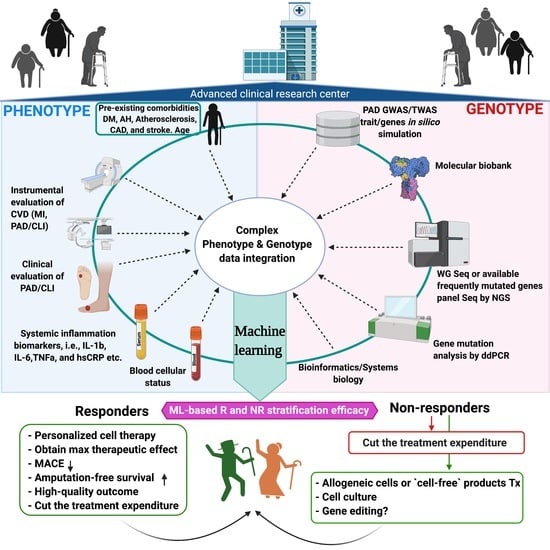
Personalized Cell Therapy for Patients with Peripheral Arterial Diseases in the Context of Genetic Alterations: Artificial Intelligence-Based Responder and Non-Responder Prediction
Abstract
1. Introduction
2. Personalized Stem Cell Therapy for Patients with PAD Based on Phenotype and Genotype Findings
2.1. Characteristic Features of the R and NR Groups
2.2. BMNC Transplantation Alleviates Thromboangiitis Obliterans (TAO), and Purified CD34/CD133+ Stem Cell Therapy Alleviates ASO
3. CLTI/PAD in Patients with Diabetes Mellitus
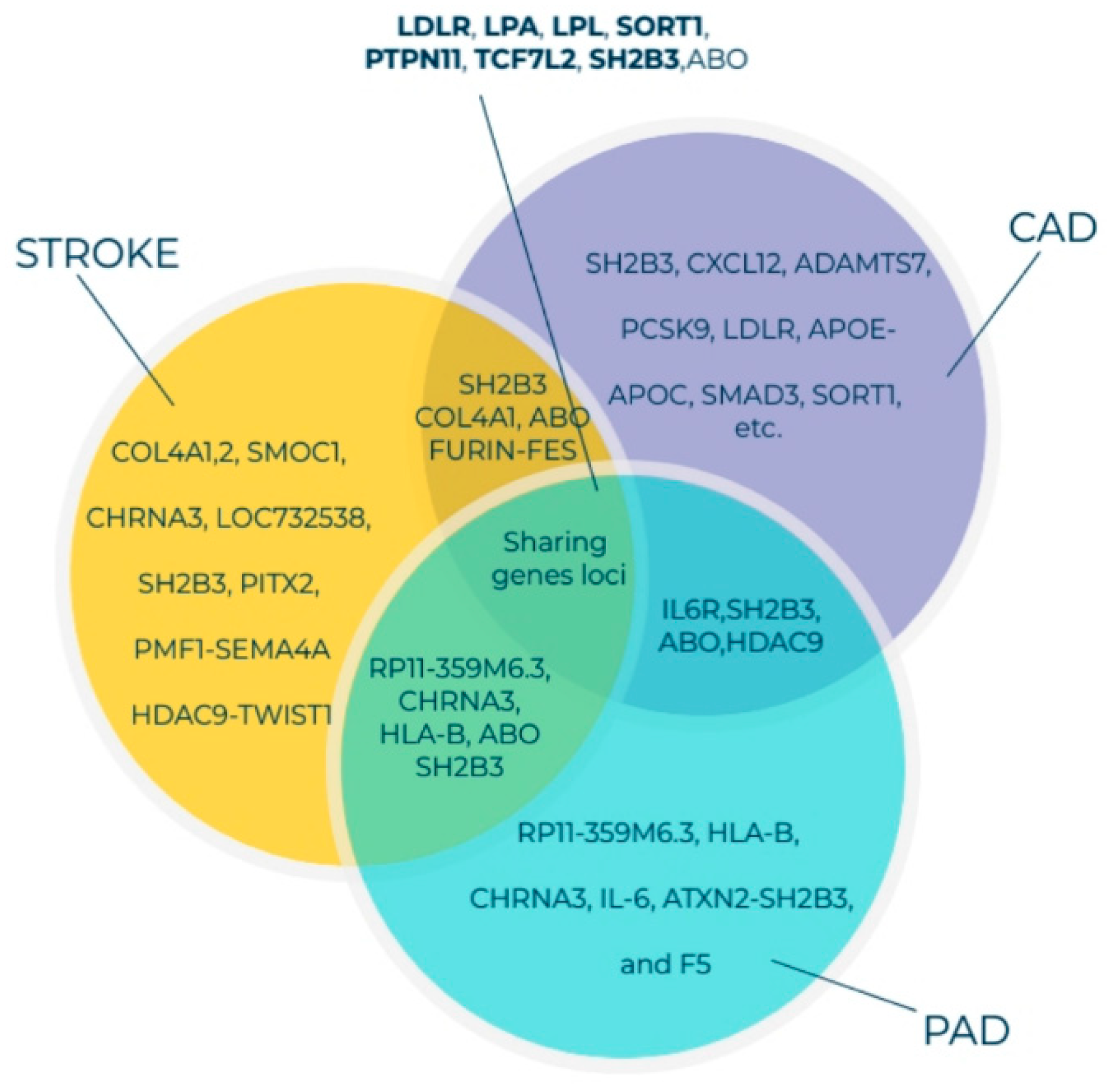
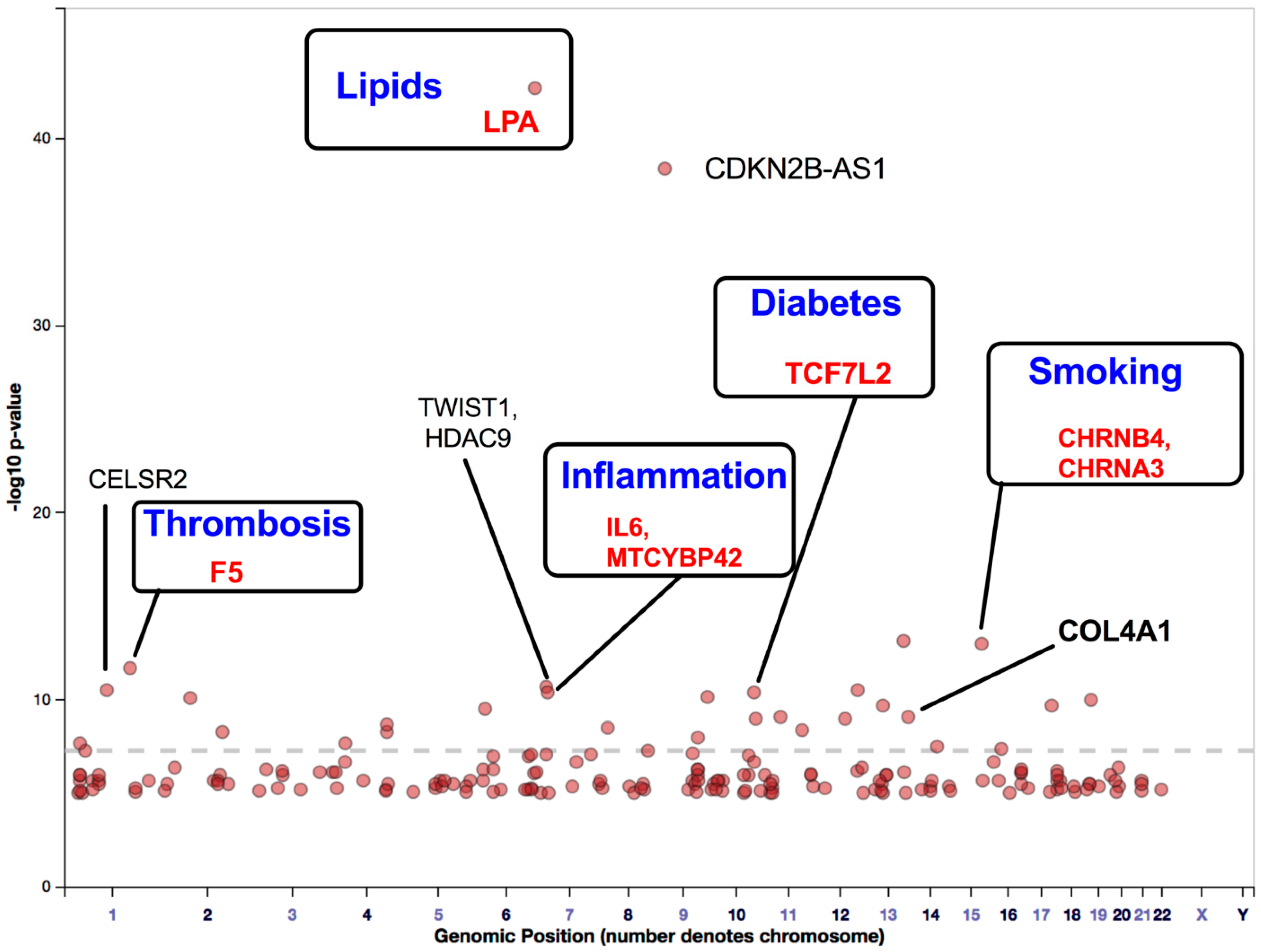
4. Findings of GWAS on Variations in PAD-Associated Gene Loci Involved in Inflammation, Thrombosis, and Lipid Imbalance
Findings of GWAS on ASO
5. Genetic Alterations in PAD
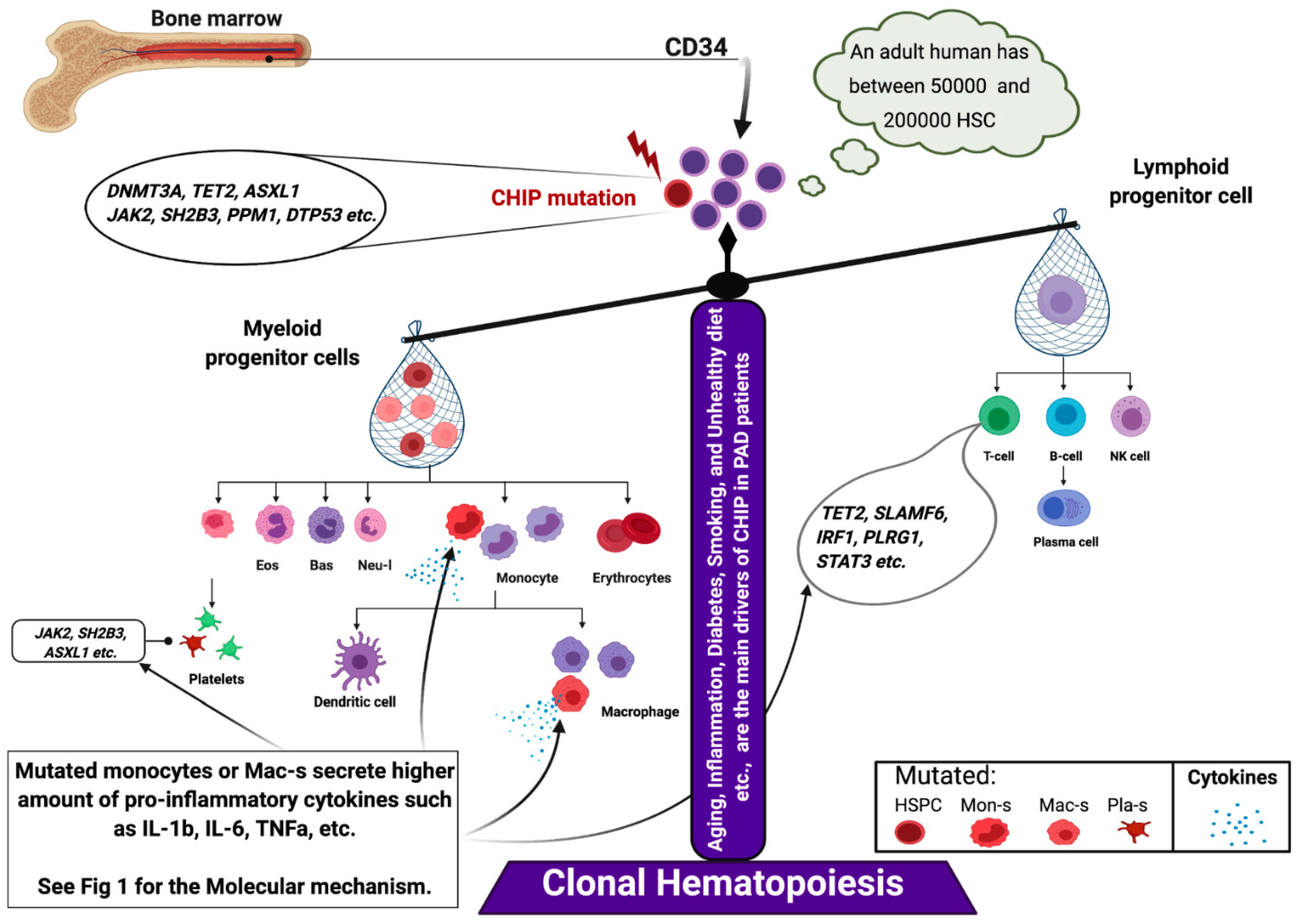
5.1. Somatic Mutation in PAD
5.2. Inflammaging and PAD
5.3. SH2B3/Lnk Mutations Interfere with Key Regeneration Pathways
6. Patient Screening
- (1)
- (2)
- (3)
- (4)
- (5)
- Patients with pre-oncology-related factors in the blood may need additional screening for CVD diseases, such as CAD, PAD, and stroke [29].
7. AI-Based R and NR Stratification of Patients with PAD Undergoing Stem Cell Transplantation
7.1. High Data Quality Is Required for Accurate Prediction Models
7.2. Supporting Decision Making through AI-Based Complex Data Analysis
7.3. Identification of Responsive Patients for Cell Therapy
7.4. A New Diagnostic Tool Supporting Individual Disease Characterization
8. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Michael, H.; Criqui, V.A. Epidemiology of Peripheral Artery Disease. Circ. Res. 2015, 116, 1509–1526. [Google Scholar]
- Howard, D.P.; Banerjee, A.; Fairhead, J.F.; Hands, L.; Silver, L.E.; Rothwell, P.M.; Oxford Vascular, S. Population-Based Study of Incidence, Risk Factors, Outcome, and Prognosis of Ischemic Peripheral Arterial Events: Implications for Prevention. Circulation 2015, 132, 1805–1815. [Google Scholar] [CrossRef]
- Fowkes, F.G.; Rudan, D.; Rudan, I.; Aboyans, V.; Denenberg, J.O.; McDermott, M.M.; Norman, P.E.; Sampson, U.K.; Williams, L.J.; Mensah, G.A.; et al. Comparison of global estimates of prevalence and risk factors for peripheral artery disease in 2000 and 2010: A systematic review and analysis. Lancet 2013, 382, 1329–1340. [Google Scholar] [CrossRef]
- Joosten, M.M.; Pai, J.K.; Bertoia, M.L.; Rimm, E.B.; Spiegelman, D.; Mittleman, M.A.; Mukamal, K.J. Associations between conventional cardiovascular risk factors and risk of peripheral artery disease in men. JAMA 2012, 308, 1660–1667. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, S.; Natarajan, P.; Silver, A.J.; Gibson, C.J.; Bick, A.G.; Shvartz, E.; McConkey, M.; Gupta, N.; Gabriel, S.; Ardissino, D.; et al. Clonal Hematopoiesis and Risk of Atherosclerotic Cardiovascular Disease. N. Engl. J. Med. 2017, 377, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Fuster, J.J.; Walsh, K. Somatic Mutations and Clonal Hematopoiesis: Unexpected Potential New Drivers of Age-Related Cardiovascular Disease. Circ. Res. 2018, 122, 523–532. [Google Scholar] [CrossRef] [PubMed]
- Muendlein, A.; Kinz, E.; Gasser, K.; Leiherer, A.; Rein, P.; Saely, C.H.; Grallert, H.; Peters, A.; Fraunberger, P.; Drexel, H.; et al. Occurrence of the JAK2 V617F mutation in patients with peripheral arterial disease. Am. J. Hematol. 2015, 90, E17–E21. [Google Scholar] [CrossRef]
- Jaiswal, S.; Ebert, B.L. Clonal Hematopoiesis in Human Aging and Disease. Science 2019, 366. [Google Scholar] [CrossRef] [PubMed]
- Gibson, C.J.; Lindsley, R.C.; Tchekmedyian, V.; Mar, B.G.; Shi, J.; Jaiswal, S.; Bosworth, A.; Francisco, L.; He, J.; Bansal, A.; et al. Clonal Hematopoiesis Associated With Adverse Outcomes After Autologous Stem-Cell Transplantation for Lymphoma. J. Clin. Oncol. 2017, 35, 1598–1605. [Google Scholar] [CrossRef]
- Wolfien, M.; Klatt, D.; Salybekov, A.A.; Ii, M.; Komatsu-Horii, M.; Gaebel, R.; Philippou-Massier, J.; Schrinner, E.; Akimaru, H.; Akimaru, E.; et al. Hematopoietic stem-cell senescence and myocardial repair—Coronary artery disease genotype/phenotype analysis of post-MI myocardial regeneration response induced by CABG/CD133+ bone marrow hematopoietic stem cell treatment in RCT PERFECT Phase 3. EBioMedicine 2020, 57, 102862. [Google Scholar] [CrossRef]
- Steensma, D.P.; Bejar, R.; Jaiswal, S.; Lindsley, R.C.; Sekeres, M.A.; Hasserjian, R.P.; Ebert, B.L. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 2015, 126, 9–16. [Google Scholar] [CrossRef]
- Falconi, G.; Fabiani, E.; Piciocchi, A.; Criscuolo, M.; Fianchi, L.; Rossi, E.L.L.; Finelli, C.; Cerqui, E.; Ottone, T.; Molteni, A.; et al. Somatic mutations as markers of outcome after azacitidine and allogeneic stem cell transplantation in higher-risk myelodysplastic syndromes. Leukemia 2019, 33, 785–790. [Google Scholar] [CrossRef] [PubMed]
- Young, A.L.; Challen, G.A.; Birmann, B.M.; Druley, T.E. Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults. Nat. Commun. 2016, 7, 12484. [Google Scholar] [CrossRef]
- Asahara, T.; Masuda, H.; Takahashi, T.; Kalka, C.; Pastore, C.; Silver, M.; Kearne, M.; Magner, M.; Isner, J.M. Bone marrow origin of endothelial progenitor cells responsible for postnatal vasculogenesis in physiological and pathological neovascularization. Circ. Res. 1999, 85, 221–228. [Google Scholar] [CrossRef]
- Steinhoff, G.; Nesteruk, J.; Wolfien, M.; Kundt, G.; Börgermann, J.; David, R.; Garbade, J.; Große, J.; Haverich, A.; Hennig, H.; et al. Cardiac Function Improvement and Bone Marrow Response -: Outcome Analysis of the Randomized PERFECT Phase III Clinical Trial of Intramyocardial CD133(+) Application After Myocardial Infarction. EBioMedicine 2017, 22, 208–224. [Google Scholar] [CrossRef]
- Tateishi-Yuyama, E.; Matsubara, H.; Murohara, T.; Ikeda, U.; Shintani, S.; Masaki, H.; Amano, K.; Kishimoto, Y.; Yoshimoto, K.; Akashi, H.; et al. Therapeutic angiogenesis for patients with limb ischaemia by autologous transplantation of bone-marrow cells: A pilot study and a randomised controlled trial. Lancet 2002, 360, 427–435. [Google Scholar] [CrossRef]
- Molavi, B.; Zafarghandi, M.R.; Aminizadeh, E.; Hosseini, S.E.; Mirzayi, H.; Arab, L.; Baharvand, H.; Aghdami, N. Safety and Efficacy of Repeated Bone Marrow Mononuclear Cell Therapy in Patients with Critical Limb Ischemia in a Pilot Randomized Controlled Trial. Arch. Iran. Med. 2016, 19, 388–396. [Google Scholar] [PubMed]
- Kawamoto, A.; Katayama, M.; Handa, N.; Kinoshita, M.; Takano, H.; Horii, M.; Sadamoto, K.; Yokoyama, A.; Yamanaka, T.; Onodera, R.; et al. Intramuscular transplantation of G-CSF-mobilized CD34(+) cells in patients with critical limb ischemia: A phase I/IIa, multicenter, single-blinded, dose-escalation clinical trial. Stem Cells 2009, 27, 2857–2864. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Kinoshita, M.; Furukawa, Y.; Nagano, T.; Hashimoto, H.; Hirami, Y.; Kurimoto, Y.; Arakawa, K.; Yamazaki, K.; Okada, Y.; et al. Phase II clinical trial of CD34+ cell therapy to explore endpoint selection and timing in patients with critical limb ischemia. Circ. J. 2014, 78, 490–501. [Google Scholar] [CrossRef]
- Barć, P.; Skóra, J.; Pupka, A.; Turkiewicz, D.; Dorobisz, A.T.; Garcarek, J.; Tomasiewicz, B.; Szyber, P. Bone-marrow cells in therapy of critical limb ischemia of lower extremities - own experience. Acta Angiol. 2006, 12, 155–166. Available online: https://journals.viamedica.pl/acta_angiologica/article/view/9862 (accessed on 13 May 2021).
- Miyamoto, K.; Nishigami, K.; Nagaya, N.; Akutsu, K.; Chiku, M.; Kamei, M.; Soma, T.; Miyata, S.; Higashi, M.; Tanaka, R.; et al. Unblinded pilot study of autologous transplantation of bone marrow mononuclear cells in patients with thromboangiitis obliterans. Circulation 2006, 114, 2679–2684. [Google Scholar] [CrossRef]
- Benoit, E.; O’Donnell, T.F., Jr.; Iafrati, M.D.; Asher, E.; Bandyk, D.F.; Hallett, J.W.; Lumsden, A.B.; Pearl, G.J.; Roddy, S.P.; Vijayaraghavan, K.; et al. The role of amputation as an outcome measure in cellular therapy for critical limb ischemia: Implications for clinical trial design. J. Transl. Med. 2011, 9, 165. [Google Scholar] [CrossRef] [PubMed]
- Teraa, M.; Sprengers, R.W.; Schutgens, R.E.; Slaper-Cortenbach, I.C.; van der Graaf, Y.; Algra, A.; van der Tweel, I.; Doevendans, P.A.; Mali, W.P.; Moll, F.L.; et al. Effect of repetitive intra-arterial infusion of bone marrow mononuclear cells in patients with no-option limb ischemia: The randomized, double-blind, placebo-controlled Rejuvenating Endothelial Progenitor Cells via Transcutaneous Intra-arterial Supplementation (JUVENTAS) trial. Circulation 2015, 131, 851–860. [Google Scholar] [CrossRef]
- Losordo, D.W.; Kibbe, M.R.; Mendelsohn, F.; Marston, W.; Driver, V.R.; Sharafuddin, M.; Teodorescu, V.; Wiechmann, B.N.; Thompson, C.; Kraiss, L.; et al. A randomized, controlled pilot study of autologous CD34+ cell therapy for critical limb ischemia. Circ. Cardiovasc. Interv. 2012, 5, 821–830. [Google Scholar] [CrossRef] [PubMed]
- Klepanec, A.; Mistrik, M.; Altaner, C.; Valachovicova, M.; Olejarova, I.; Slysko, R.; Balazs, T.; Urlandova, T.; Hladikova, D.; Liska, B.; et al. No difference in intra-arterial and intramuscular delivery of autologous bone marrow cells in patients with advanced critical limb ischemia. Cell Transpl. 2012, 21, 1909–1918. [Google Scholar] [CrossRef] [PubMed]
- Madaric, J.; Klepanec, A.; Valachovicova, M.; Mistrik, M.; Bucova, M.; Olejarova, I.; Necpal, R.; Madaricova, T.; Paulis, L.; Vulev, I. Characteristics of responders to autologous bone marrow cell therapy for no-option critical limb ischemia. Stem Cell Res. Ther. 2016, 7, 116. [Google Scholar] [CrossRef] [PubMed]
- Pan, T.; Liu, H.; Fang, Y.; Wei, Z.; Gu, S.; Fang, G.; Liu, Y.; Luo, Y.; Guo, D.; Xu, X.; et al. Predictors of responders to mononuclear stem cell-based therapeutic angiogenesis for no-option critical limb ischemia. Stem Cell Res. Ther. 2019, 10, 15. [Google Scholar] [CrossRef]
- Jaiswal, S.; Fontanillas, P.; Flannick, J.; Manning, A.; Grauman, P.V.; Mar, B.G.; Lindsley, R.C.; Mermel, C.H.; Burtt, N.; Chavez, A.; et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 2014, 371, 2488–2498. [Google Scholar] [CrossRef]
- Rivero, G.A.; Perli, E.; Moreno, S.; Salemi, J.L. Excess in Atherosclerotic and Inflammametabolic Diseases Are Differentially Expressed in Myelodysplasia and Are Highly Dependent on Age, R-IPSS and Ethnicity. Blood 2018, 132. [Google Scholar] [CrossRef]
- Cosgrove, M.E.; Suman, R.; Harrison, H.J.; Jackson, G.E.; Howard, M.R.; Hitchcock, I.S. Endothelial JAK2V617F Expression Drives Inflammation and Cellular Senescence; New Evidence for the Roles of Endothelial Cells in MPN-Related Clotting Abnormalities? Blood 2016, 128. [Google Scholar] [CrossRef]
- Malyar, N.M.; Radtke, S.; Malyar, K.; Arjumand, J.; Horn, P.A.; Kroger, K.; Freisinger, E.; Reinecke, H.; Giebel, B.; Brock, F.E. Autologous bone marrow mononuclear cell therapy improves symptoms in patients with end-stage peripheral arterial disease and reduces inflammation-associated parameters. Cytotherapy 2014, 16, 1270–1279. [Google Scholar] [CrossRef]
- Arai, M.; Misao, Y.; Nagai, H.; Kawasaki, M.; Nagashima, K.; Suzuki, K.; Tsuchiya, K.; Otsuka, S.; Uno, Y.; Takemura, G.; et al. Granulocyte Colony-Stimulating Factor A Noninvasive Regeneration Therapy for Treating Atherosclerotic Peripheral Artery Disease. Circ. J. 2006, 70, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Pan, T.; Fang, Y.; Wei, Z.; Gu, S.; Fang, G.; Liu, Y.; Luo, Y.; Liu, H.; Zhang, T.; et al. Purified CD34(+) cells versus peripheral blood mononuclear cells in the treatment of angiitis-induced no-option critical limb ischaemia: 12-Month results of a prospective randomised single-blinded non-inferiority trial. EBioMedicine 2018, 35, 46–57. [Google Scholar] [CrossRef]
- Idei, N.; Soga, J.; Hata, T.; Fujii, Y.; Fujimura, N.; Mikami, S.; Maruhashi, T.; Nishioka, K.; Hidaka, T.; Kihara, Y.; et al. Autologous bone-marrow mononuclear cell implantation reduces long-term major amputation risk in patients with critical limb ischemia: A comparison of atherosclerotic peripheral arterial disease and Buerger disease. Circ. Cardiovasc. Interv. 2011, 4, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Guo, L.; Cui, S.; Tong, Z.; Dardik, A.; Gu, Y. Autologous bone marrow-derived mononuclear cell therapy in Chinese patients with critical limb ischemia due to thromboangiitis obliterans: 10-year results. Stem Cell Res. Ther. 2018, 9, 43. [Google Scholar] [CrossRef] [PubMed]
- Ortmann, C.A.; Dorsheimer, L.; Abou-El-Ardat, K.; Hoffrichter, J.; Assmus, B.; Bonig, H.; Scholz, A.; Pfeifer, H.; Martin, H.; Schmid, T.; et al. Functional Dominance of CHIP-Mutated Hematopoietic Stem Cells in Patients Undergoing Autologous Transplantation. Cell Rep. 2019, 27, 2022–2028.e3. [Google Scholar] [CrossRef]
- Piazza, G.; Creager, M.A. Thromboangiitis obliterans. Circulation 2010, 121, 1858–1861. [Google Scholar] [CrossRef]
- Rivera-Chavarria, I.J.; Brenes-Gutierrez, J.D. Thromboangiitis obliterans (Buerger’s disease). Ann. Med. Surg. 2016, 7, 79–82. [Google Scholar] [CrossRef]
- Shu, J.; Santulli, G. Update on peripheral artery disease: Epidemiology and evidence-based facts. Atherosclerosis 2018, 275, 379–381. [Google Scholar] [CrossRef]
- Fazeli, B.; Dadgar Moghadam, M.; Niroumand, S. How to Treat a Patient with Thromboangiitis Obliterans: A Systematic Review. Ann. Vasc. Surg. 2018, 49, 219–228. [Google Scholar] [CrossRef]
- Cooper, L.T.; Henderson, S.S.; Ballman, K.V.; Offord, K.P.; Tse, T.S.; Holmes, D.R.; Hurt, R.D. A prospective, case-control study of tobacco dependence in thromboangiitis obliterans (Buerger’s Disease). Angiology 2006, 57, 73–78. [Google Scholar] [CrossRef]
- Song, P.; Rudan, D.; Zhu, Y.; Fowkes, F.J.I.; Rahimi, K.; Fowkes, F.G.R.; Rudan, I. Global, regional, and national prevalence and risk factors for peripheral artery disease in 2015: An updated systematic review and analysis. Lancet Glob. Health 2019, 7, e1020–e1030. [Google Scholar] [CrossRef]
- Thiruvoipati, T.; Kielhorn, C.E.; Armstrong, E.J. Peripheral artery disease in patients with diabetes: Epidemiology, mechanisms, and outcomes. World J. Diabetes 2015, 6, 961–969. [Google Scholar] [CrossRef] [PubMed]
- Adar, R.; Papa, M.Z.; Halpern, Z.; Mozes, M.; Shoshan, S.; Sofer, B.; Zinger, H.; Dayan, M.; Mozes, E. Cellular sensitivity to collagen in thromboangiitis obliterans. N. Engl. J. Med. 1983, 308, 1113–1116. [Google Scholar] [CrossRef]
- Wei, Z.; Jiang, W.; Wang, H.; Li, H.; Tang, B.; Liu, B.; Jiang, H.; Sun, X. The IL-6/STAT3 pathway regulates adhesion molecules and cytoskeleton of endothelial cells in thromboangiitis obliterans. Cell. Signal. 2018, 44, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Fadini, G.P.; Albiero, M.; Vigili de Kreutzenberg, S.; Boscaro, E.; Cappellari, R.; Marescotti, M.; Poncina, N.; Agostini, C.; Avogaro, A. Diabetes impairs stem cell and proangiogenic cell mobilization in humans. Diabetes Care 2013, 36, 943–949. [Google Scholar] [CrossRef]
- Fadini, G.P.; Fiala, M.; Cappellari, R.; Danna, M.; Park, S.; Poncina, N.; Menegazzo, L.; Albiero, M.; DiPersio, J.; Stockerl-Goldstein, K.; et al. Diabetes Limits Stem Cell Mobilization Following G-CSF but Not Plerixafor. Diabetes 2015, 64, 2969–2977. [Google Scholar] [CrossRef]
- Albiero, M.; Poncina, N.; Tjwa, M.; Ciciliot, S.; Menegazzo, L.; Ceolotto, G.; de Kreutzenberg, S.V.; Moura, R.; Giorgio, M.; Pelicci, P.; et al. Diabetes causes bone marrow autonomic neuropathy and impairs stem cell mobilization via dysregulated p66Shc and Sirt1. Diabetes 2014, 63, 1353–1365. [Google Scholar] [CrossRef]
- Teraa, M.; Fledderus, J.O.; Rozbeh, R.I.; Leguit, R.J.; Verhaar, M.C.; Juventas Study, G. Bone marrow microvascular and neuropathic alterations in patients with critical limb ischemia. Circ. Res. 2014, 114, 311–314. [Google Scholar] [CrossRef]
- Bonnefond, A.; Skrobek, B.; Lobbens, S.; Eury, E.; Thuillier, D.; Cauchi, S.; Lantieri, O.; Balkau, B.; Riboli, E.; Marre, M.; et al. Association between large detectable clonal mosaicism and type 2 diabetes with vascular complications. Nat. Genet. 2013, 45, 1040–1043. [Google Scholar] [CrossRef]
- Fadini, G.P.; Ciciliot, S.; Albiero, M. Concise Review: Perspectives and Clinical Implications of Bone Marrow and Circulating Stem Cell Defects in Diabetes. Stem Cells 2017, 35, 106–116. [Google Scholar] [CrossRef]
- Kojima, H.; Kim, J.; Chan, L. Emerging roles of hematopoietic cells in the pathobiology of diabetic complications. Trends Endocrinol. Metab. 2014, 25, 178–187. [Google Scholar] [CrossRef]
- Orlandi, A.; Chavakis, E.; Seeger, F.; Tjwa, M.; Zeiher, A.M.; Dimmeler, S. Long-term diabetes impairs repopulation of hematopoietic progenitor cells and dysregulates the cytokine expression in the bone marrow microenvironment in mice. Basic Res. Cardiol. 2010, 105, 703–712. [Google Scholar] [CrossRef]
- Tanaka, R.; Vaynrub, M.; Masuda, H.; Ito, R.; Kobori, M.; Miyasaka, M.; Mizuno, H.; Warren, S.M.; Asahara, T. Quality-control culture system restores diabetic endothelial progenitor cell vasculogenesis and accelerates wound closure. Diabetes 2013, 62, 3207–3217. [Google Scholar] [CrossRef] [PubMed]
- Salybekov, A.A.; Masuda, H.; Miyazaki, K.; Sheng, Y.; Sato, A.; Shizuno, T.; Iida, Y.; Okada, Y.; Asahara, T. Dipeptidyl dipeptidase-4 inhibitor recovered ischemia through an increase in vasculogenic endothelial progenitor cells and regeneration-associated cells in diet-induced obese mice. PLoS ONE 2019, 14, e0205477. [Google Scholar] [CrossRef]
- Salybekov, A.A.; Kawaguchi, A.T.; Masuda, H.; Vorateera, K.; Okada, C.; Asahara, T. Regeneration-associated cells improve recovery from myocardial infarction through enhanced vasculogenesis, anti-inflammation, and cardiomyogenesis. PLoS ONE 2018, 13, e0203244. [Google Scholar] [CrossRef] [PubMed]
- Salybekov, A.A.; Salybekova, A.; Sheng, Y.; Shinozaki, Y.; Yokoyama, K.; Kobayashi, S.; Asahara, T. Extracellular Vesicles Derived From Regeneration Associated Cells Preserve Heart Function After Ischemia-Induced Injury. Front. Cardiovasc. Med. 2021, 8, 754254. [Google Scholar] [CrossRef] [PubMed]
- Perttila, J.; Merikanto, K.; Naukkarinen, J.; Surakka, I.; Martin, N.W.; Tanhuanpaa, K.; Grimard, V.; Taskinen, M.R.; Thiele, C.; Salomaa, V.; et al. OSBPL10, a novel candidate gene for high triglyceride trait in dyslipidemic Finnish subjects, regulates cellular lipid metabolism. J. Mol. Med. 2009, 87, 825–835. [Google Scholar] [CrossRef]
- Koriyama, H.; Nakagami, H.; Katsuya, T.; Sugimoto, K.; Yamashita, H.; Takami, Y.; Maeda, S.; Kubo, M.; Takahashi, A.; Nakamura, Y.; et al. Identification of evidence suggestive of an association with peripheral arterial disease at the OSBPL10 locus by genome-wide investigation in the Japanese population. J. Atheroscler. Thromb. 2010, 17. [Google Scholar] [CrossRef]
- Maslah, N.; Cassinat, B.; Verger, E.; Kiladjian, J.J.; Velazquez, L. The role of LNK/SH2B3 genetic alterations in myeloproliferative neoplasms and other hematological disorders. Leukemia 2017, 31, 1661–1670. [Google Scholar] [CrossRef]
- Klarin, D.; Lynch, J.; Aragam, K.; Chaffin, M.; Assimes, T.L.; Huang, J.; Lee, K.M.; Shao, Q.; Huffman, J.E.; Natarajan, P.; et al. Genome-wide Association Study of Peripheral Artery Disease in the Million Veteran Program. Nat. Med. 2019, 25, 1274–1279. [Google Scholar] [CrossRef]
- Nikpay, M.; Goel, A.; Won, H.H.; Hall, L.M.; Willenborg, C.; Kanoni, S.; Saleheen, D.; Kyriakou, T.; Nelson, C.P.; Hopewell, J.C.; et al. A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 2015, 47, 1121–1130. [Google Scholar] [CrossRef] [PubMed]
- Malik, R.; Chauhan, G.; Traylor, M.; Sargurupremraj, M.; Okada, Y.; Mishra, A.; Rutten-Jacobs, L.; Giese, A.K.; van der Laan, S.W.; Gretarsdottir, S.; et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 2018, 50, 524–537. [Google Scholar] [CrossRef] [PubMed]
- Han, Z.; Dong, X.; Zhang, C.; Wu, Y.; Yuan, Z.; Wang, X. Polymorphism of HDAC9 Gene Is Associated with Increased Risk of Acute Coronary Syndrome in Chinese Han Population. Biomed. Res. Int. 2016, 2016, 3746276. [Google Scholar] [CrossRef]
- Markus, H.S.; Makela, K.M.; Bevan, S.; Raitoharju, E.; Oksala, N.; Bis, J.C.; O’Donnell, C.; Hainsworth, A.; Lehtimaki, T. Evidence HDAC9 genetic variant associated with ischemic stroke increases risk via promoting carotid atherosclerosis. Stroke 2013, 44, 1220–1225. [Google Scholar] [CrossRef] [PubMed]
- Sofer, T.; Emery, L.; Jain, D.; Ellis, A.M.; Laurie, C.C.; Allison, M.A.; Lee, J.; Kurniansyah, N.; Kerr, K.F.; Gonzalez, H.M.; et al. Variants Associated with the Ankle Brachial Index Differ by Hispanic/Latino Ethnic Group: A genome-wide association study in the Hispanic Community Health Study/Study of Latinos. Sci. Rep. 2019, 9, 11410. [Google Scholar] [CrossRef] [PubMed]
- Ou, M.; Li, X.; Zhao, S.; Cui, S.; Tu, J. Long non-coding RNA CDKN2B-AS1 contributes to atherosclerotic plaque formation by forming RNA-DNA triplex in the CDKN2B promoter. EBioMedicine 2020, 55. [Google Scholar] [CrossRef]
- Wainberg, M.; Sinnott-Armstrong, N.; Mancuso, N.; Barbeira, A.N.; Knowles, D.A.; Golan, D.; Ermel, R.; Ruusalepp, A.; Quertermous, T.; Hao, K.; et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 2019, 51, 592–599. [Google Scholar] [CrossRef]
- Savji, N.; Rockman, C.B.; Skolnick, A.H.; Guo, Y.; Adelman, M.A.; Riles, T.; Berger, J.S. Association between advanced age and vascular disease in different arterial territories: A population database of over 3.6 million subjects. J. Am. Coll. Cardiol. 2013, 61, 1736–1743. [Google Scholar] [CrossRef]
- Fuster, J.J.; MacLauchlan, S.; Zuriaga, M.A.; Polackal, M.N.; Ostriker, A.C.; Chakraborty, R.; Wu, C.L.; Sano, S.; Muralidharan, S.; Rius, C.; et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 2017, 355, 842–847. [Google Scholar] [CrossRef]
- Shah, A.D.; Langenberg, C.; Rapsomaniki, E.; Denaxas, S.; Pujades-Rodriguez, M.; Gale, C.P.; Deanfield, J.; Smeeth, L.; Timmis, A.; Hemingway, H. Type 2 diabetes and incidence of cardiovascular diseases: A cohort study in 1.9 million people. Lancet Diabetes Endocrinol. 2015, 3, 105–113. [Google Scholar] [CrossRef]
- Yoshida, K.; Gowers, K.H.C.; Lee-Six, H.; Chandrasekharan, D.P.; Coorens, T.; Maughan, E.F.; Beal, K.; Menzies, A.; Millar, F.R.; Anderson, E.; et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 2020, 578, 266–272. [Google Scholar] [CrossRef] [PubMed]
- Perner, F.; Perner, C.; Ernst, T.; Heidel, F.H. Roles of JAK2 in Aging, Inflammation, Hematopoiesis and Malignant Transformation. Cells 2019, 8, 854. [Google Scholar] [CrossRef]
- Sano, S.; Wang, Y.; Yura, Y.; Sano, M.; Oshima, K.; Yang, Y.; Katanasaka, Y.; Min, K.D.; Matsuura, S.; Ravid, K.; et al. JAK2 (V617F) -Mediated Clonal Hematopoiesis Accelerates Pathological Remodeling in Murine Heart Failure. JACC Basic Transl. Sci. 2019, 4, 684–697. [Google Scholar] [CrossRef] [PubMed]
- Losordo, D.W.; Henry, T.D.; Davidson, C.; Sup Lee, J.; Costa, M.A.; Bass, T.; Mendelsohn, F.; Fortuin, F.D.; Pepine, C.J.; Traverse, J.H.; et al. Intramyocardial, autologous CD34+ cell therapy for refractory angina. Circ. Res. 2011, 109, 428–436. [Google Scholar] [CrossRef]
- Farina, M.; Bernardi, S.; Polverelli, N.; Zanaglio, C.; Malagola, M.; Re, F.; Cattina, F.; D’Adda, M.; Rossi, G.; Zollner, T.; et al. Comparative Somatic Mutational Profiling of CD34+ Hematopoietic Precursors (HSC) and Circulating Endothelial Cells (CEC) in Patients with Primary Myelofibrosis (PMF). Blood 2019, 134, 1684. [Google Scholar] [CrossRef]
- Franceschi, C.; Capri, M.; Monti, D.; Giunta, S.; Olivieri, F.; Sevini, F.; Panourgia, M.P.; Invidia, L.; Celani, L.; Scurti, M.; et al. Inflammaging and anti-inflammaging: A systemic perspective on aging and longevity emerged from studies in humans. Mech. Ageing Dev. 2007, 128, 92–105. [Google Scholar] [CrossRef]
- Cull, A.H.; Snetsinger, B.; Buckstein, R.; Wells, R.A.; Rauh, M.J. Tet2 restrains inflammatory gene expression in macrophages. Exp. Hematol. 2017, 55, 56–70. [Google Scholar] [CrossRef] [PubMed]
- Abegunde, S.O.; Buckstein, R.; Wells, R.A.; Rauh, M.J. An inflammatory environment containing TNFalpha favors Tet2-mutant clonal hematopoiesis. Exp. Hematol. 2018, 59, 60–65. [Google Scholar] [CrossRef]
- Dorsheimer, L.; Assmus, B.; Rasper, T.; Ortmann, C.A.; Ecke, A.; Abou-El-Ardat, K.; Schmid, T.; Brüne, B.; Wagner, S.; Hoffmann, J.; et al. Association of Mutations Contributing to Clonal Hematopoiesis With Prognosis in Chronic Ischemic Heart Failure. JAMA Cardiol. 2020, 4, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Busque, L.; Sun, M.; Buscarlet, M.; Ayachi, S.; Feroz Zada, Y.; Provost, S.; Bourgoin, V.; Mollica, L.; Meisel, M.; Hinterleitner, R.; et al. High-sensitivity C-reactive protein is associated with clonal hematopoiesis of indeterminate potential. Blood Adv. 2020, 4, 2430–2438. [Google Scholar] [CrossRef]
- King, K.Y.; Huang, Y.; Nakada, D.; Goodell, M.A. Environmental influences on clonal hematopoiesis. Exp. Hematol. 2020, 83, 66–73. [Google Scholar] [CrossRef]
- Zhang, Q.; Zhao, K.; Shen, Q.; Han, Y.; Gu, Y.; Li, X.; Zhao, D.; Liu, Y.; Wang, C.; Zhang, X.; et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 2015, 525, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Elias, H.K.; Bryder, D.; Park, C.Y. Molecular mechanisms underlying lineage bias in aging hematopoiesis. Semin. Hematol. 2017, 54, 4–11. [Google Scholar] [CrossRef]
- Baughn, L.B.; Meredith, M.M.; Oseth, L.; Smolarek, T.A.; Hirsch, B. SH2B3 aberrations enriched in iAMP21 B lymphoblastic leukemia. Cancer Genet. 2018, 226-227. [Google Scholar] [CrossRef] [PubMed]
- Takaki, S.; Morita, H.; Tezuka, Y.; Takatsu, K. Enhanced hematopoiesis by hematopoietic progenitor cells lacking intracellular adaptor protein, Lnk. J. Exp. Med. 2002, 195. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Tang, Y.; Wang, Y.; Tascau, L.; Balcerek, J.; Tong, W.; Levine, R.L.; Welch, C.; Tall, A.R.; Wang, N. LNK/SH2B3 Loss of Function Promotes Atherosclerosis and Thrombosis. Circ. Res. 2016, 119. [Google Scholar] [CrossRef]
- Snetsinger, B.; Ferrone, C.K.; Rauh, M.J. Targeted, Amplicon-Based, Next-Generation Sequencing to Detect Age-Related Clonal Hematopoiesis. Methods Mol. Biol. 2019, 2045, 167–180. [Google Scholar] [CrossRef]
- Buckstein, R.; Jang, K.; Friedlich, J.; Zhang, L.; Reis, M.; Chesney, A.; Wells, R.A. Estimating the prevalence of myelodysplastic syndromes in patients with unexplained cytopenias: A retrospective study of 322 bone marrows. Leuk. Res. 2009, 33. [Google Scholar] [CrossRef]
- Rauw, J.; Wells, R.A.; Chesney, A.; Reis, M.; Zhang, L.; Buckstein, R. Validation of a scoring system to establish the probability of myelodysplastic syndrome in patients with unexplained cytopenias or macrocytosis. Leuk. Res. 2011, 35. [Google Scholar] [CrossRef]
- Svensson, E.C.; Madar, A.; Campbell, C.D.; He, Y.; Sultan, M.; Healey, M.L.; D’Aco, K.; Fernandez, A.; Wache-Mainier, C.; Ridker, P.M.; et al. TET2-Driven Clonal Hematopoiesis Predicts Enhanced Response to Canakinumab in the CANTOS Trial: An Exploratory Analysis. Circulation 2018, 138, A15111. [Google Scholar]
- Obermeyer, Z.; Emanuel, E.J. Predicting the Future—Big Data, Machine Learning, and Clinical Medicine. N. Engl. J. Med. 2016, 375, 1216–1219. [Google Scholar] [CrossRef] [PubMed]
- Müller, R.U.; Skorska, A.; Lemcke, H.; Steinhoff, G.; David, R. GLP: A requirement in cell therapies—Perspectives for the cardiovascular field. Adv. Drug Deliv. Rev. 2020. [Google Scholar] [CrossRef]
- Uellendahl-Werth, F.; Wolfien, M.; Franke, A.; Wolkenhauer, O.; Ellinghaus, D. A benchmark of hemoglobin blocking during library preparation for mRNA-Sequencing of human blood samples. Sci. Rep. 2020, 10, 5630. [Google Scholar] [CrossRef] [PubMed]
- Yue, L.; Tian, D.; Chen, W.; Han, X.; Yin, M. Deep learning for heterogeneous medical data analysis. World Wide Web 2020, 23, 2715–2737. [Google Scholar] [CrossRef]
- Zhang, D.; Yin, C.; Zeng, J.; Yuan, X.; Zhang, P. Combining structured and unstructured data for predictive models: A deep learning approach. BMC Med. Inform. Decis. Mak. 2020, 20, 280. [Google Scholar] [CrossRef]
- Lambin, P.; Leijenaar, R.T.H.; Deist, T.M.; Peerlings, J.; de Jong, E.E.C.; van Timmeren, J.; Sanduleanu, S.; Larue, R.; Even, A.J.G.; Jochems, A.; et al. Radiomics: The bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 2017, 14, 749–762. [Google Scholar] [CrossRef]
- Del Mazo-Barbara, A.; Nieto, V.; Mirabel, C.; Reyes, B.; García-López, J.; Oliver-Vila, I.; Vives, J. Streamlining the qualification of computerized systems in GxP-compliant academic cell therapy facilities. Cytotherapy 2016, 18, 1237–1239. [Google Scholar] [CrossRef]
- Shah, R.U. We Don’t Need More Data, We Need the Right Data. Circulation 2020, 142, 197–198. [Google Scholar] [CrossRef]
- Begoli, E.; Bhattacharya, T.; Kusnezov, D. The need for uncertainty quantification in machine-assisted medical decision making. Nat. Mach. Intell. 2019, 1, 20–23. [Google Scholar] [CrossRef]
- Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 2019, 1, 206–215. [Google Scholar] [CrossRef]
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Chiuve, S.E.; Cushman, M.; Delling, F.N.; Deo, R.; et al. Heart Disease and Stroke Statistics—2018 Update: A Report From the American Heart Association. Circulation 2018, 137, e67–e492. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.M.; Lee, Y.; Lee, Y.; Lee, S.; Park, J. An Algorithm Based on Deep Learning for Predicting In-Hospital Cardiac Arrest. J. Am. Hear. Assoc. 2018, 7, e008678. [Google Scholar] [CrossRef]
- Weng, S.F.; Reps, J.; Kai, J.; Garibaldi, J.M.; Qureshi, N. Can machine-learning improve cardiovascular risk prediction using routine clinical data? PLoS ONE 2017, 12, e0174944. [Google Scholar] [CrossRef] [PubMed]
- Hastie, T.; Tibshirani, R.; Friedman, J. Random Forests; Springer: New York, NY, USA, 2009; pp. 587–624. [Google Scholar]
- Dilsizian, S.E.; Siegel, E.L. Artificial intelligence in medicine and cardiac imaging: Harnessing big data and advanced computing to provide personalized medical diagnosis and treatment. Curr. Cardiol. Rep. 2014, 16, 441. [Google Scholar] [CrossRef]
- Karamperis, K.; Wadge, S.; Koromina, M.; Patrinos, G.P. Applied Genomics and Public Health; Academic Press: London, UK, 2020; pp. 189–207. Available online: https://www.sciencedirect.com/book/9780128136959/applied-genomics-and-public-health (accessed on 13 May 2021).
- Braverman, G.; Shapiro, Z.E.; Bernstein, J.A. Ethical Issues in Contemporary Clinical Genetics. Mayo Clin. Proc. Innov. Qual. Outcomes 2018, 2, e1918962. [Google Scholar] [CrossRef][Green Version]
- Desai, R.J.; Wang, S.V.; Vaduganathan, M.; Evers, T.; Schneeweiss, S. Comparison of Machine Learning Methods With Traditional Models for Use of Administrative Claims With Electronic Medical Records to Predict Heart Failure Outcomes. JAMA Netw. Open 2020, 3, e1918962. [Google Scholar] [CrossRef]
- Firouzi, F.; Sussman, M.A. Blood speaks: Personalised medicine profiling for heart failure patients. EBioMedicine 2020, 58, 102900. [Google Scholar] [CrossRef]
- Zhao, M.T.; Shao, N.Y.; Garg, V. Subtype-specific cardiomyocytes for precision medicine: Where are we now? Stem Cells 2020, 38, 822–833. [Google Scholar] [CrossRef]
- Sánchez-Rico, M.; Alvarado, J.M. A Machine Learning Approach for Studying the Comorbidities of Complex Diagnoses. Behav. Sci. 2019, 9, 122. [Google Scholar] [CrossRef]
- Komura, D.; Ishikawa, S. Machine Learning Methods for Histopathological Image Analysis. Comput. Struct. Biotechnol. J. 2018, 16, 34–42. [Google Scholar] [CrossRef]
- Siontis, K.C.; Noseworthy, P.A.; Attia, Z.I.; Friedman, P.A. Artificial intelligence-enhanced electrocardiography in cardiovascular disease management. Nat. Rev. Cardiol. 2021, 18, 465–478. [Google Scholar] [CrossRef]
- Dai, L.; Zhou, Q.; Zhou, H.; Zhang, H.; Cheng, P.; Ding, M.; Xu, X.; Zhang, X. Deep learning-based classification of lower extremity arterial stenosis in computed tomography angiography. Eur. J. Radiol. 2021, 136, 109528. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.L.; Conlin, C.C.; Li, X.; Layec, G.; Chang, K.; Kalpathy-Cramer, J.; Lee, V.S. Exercise-induced calf muscle hyperemia: Rapid mapping of magnetic resonance imaging using deep learning approach. Physiol. Rep. 2020, 8, e14563. [Google Scholar] [CrossRef]
- Veit-Haibach, P.; Huellner, M.W.; Banyai, M.; Mafeld, S.; Heverhagen, J.; Strobel, K.; Sah, B.R. CT perfusion in peripheral arterial disease-hemodynamic differences before and after revascularisation. Eur. Radiol. 2021, 31, 5507–5513. [Google Scholar] [CrossRef] [PubMed]
- Flores, A.M.; Demsas, F.; Leeper, N.J.; Ross, E.G. Leveraging Machine Learning and Artificial Intelligence to Improve Peripheral Artery Disease Detection, Treatment, and Outcomes. Circ. Res. 2021, 128, 1833–1850. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Hahn, J.-O.; Youn, B.D. Detection and Severity Assessment of Peripheral Occlusive Artery Disease via Deep Learning Analysis of Arterial Pulse Waveforms: Proof-of-Concept and Potential Challenges. Front. Bioeng. Biotechnol. 2020, 8, 720. [Google Scholar] [CrossRef]
- Fu, Y.; Jung, A.W.; Torne, R.V.; Gonzalez, S.; Vöhringer, H.; Shmatko, A.; Yates, L.R.; Jimenez-Linan, M.; Moore, L.; Gerstung, M. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Rev. Cancer 2020, 1, 800–810. [Google Scholar] [CrossRef]
- Gao, W.; Chen, D.; Liu, G.; Ran, X. Autologous stem cell therapy for peripheral arterial disease: A systematic review and meta-analysis of randomized controlled trials. Stem Cell Res. Ther. 2019, 10, 140. [Google Scholar] [CrossRef]
- Rigato, M.; Monami, M.; Fadini, G.P. Autologous Cell Therapy for Peripheral Arterial Disease: Systematic Review and Meta-Analysis of Randomized, Nonrandomized, and Noncontrolled Studies. Circ. Res. 2017, 120, 1326–1340. [Google Scholar] [CrossRef]
- Johnson, K.B.; Wei, W.; Weeraratne, D.; Frisse, M.E.; Misulis, K.; Rhee, K.; Zhao, J.; Snowdon, J.L. Precision Medicine, AI, and the Future of Personalized Health Care. Care. Clin. Transl. Sci. 2021, 14, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Kelly, C.J.; Karthikesalingam, A.; Suleyman, M.; Corrado, G.; King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 2019, 17, 195. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Preininger, A. Artificial intelligence in Health: State of the Art, Challenges, and Future Directions. Yearb. Med. Inform. 2019, 28, 16–26. [Google Scholar] [CrossRef]
- Salybekov, A.A.; Kunikeyev, A.D.; Kobayashi, S.; Asahara, T. Latest Advances in Endothelial Progenitor Cell-Derived Extracellular Vesicles Translation to the Clinic. Front. Cardiovasc. Med. 2021, 8, 734562. [Google Scholar] [CrossRef] [PubMed]

| Author | Year | Cell Type | Target Disease | Route of Tx | n = | Outcome | Follow-Up |
|---|---|---|---|---|---|---|---|
| Tateishi-Yuyama et al. [16] | 2002 | BMMC and PBMC | CLTI ASO | IM | 45 | ↑ ABI, ↑ TcO2, complete ulcer healing; 6/10 (60%) major amputation rates; not shown | 4 and 24 weeks |
| Molavi et al. [17] | 2016 | BMMC | CLTI ASO | IM rep. | 22 | ↑ SPP, ↑ TcPO2, no ABI, reduction in ulcer healing; no adverse effect | 24 weeks |
| Kawamoto et al. [18] | 2009 | CD34+ | CLTI TAO | IM | 17 | ↑ TBI, ↑ TcPO2, no ABI, complete ulcer healing; did not affect major amputation rates; 0 | 3 months |
| Fujita et al. [19] | 2014 | CD34+ | CLTI TAO | IM | 11 | ↑ ABI, ↑ LDP, ↑ SPP, ↑ TcPO2, ↑ pain-free walking distance | 12 months |
| Author | Year | Cell Type | Target Disease | Route of Tx | n = | Outcome | Follow-Up |
|---|---|---|---|---|---|---|---|
| Barc et al. [20] | 2006 | BMMC ASO | CLTI | IM | 29 | No differences in ABI and pain; major amputation rates not shown | 6 months |
| Miyamoto et al. [21] | 2006 | BMMC TAO | CLTI | IM | 8 | Half of the cell-transplanted patients had long-term adverse events, including death and unfavorable angiogenesis. | 24 to 48 months |
| Benoit et al. [22] | 2011 | BMMC ASO | CLTI | IM | 48 | Non-significant difference in amputation rate when compared with placebo | 6 months |
| Teraa et al. [23] | 2015 | BMMC ASO | CLTI | IA | 160 | No significant differences were observed for the primary outcome, i.e., major amputation at 6 months, BMMC treated versus placebo groups | 6 months |
| Losordo et al. [24] | 2011 | CD34 ASO | CLTI | IM | 28 | Favorable (but non-significant) trend for decreased amputation rate with high-dose administration when compared with placebo | 6 months |
| Clinical Trial | Cell Type | Target Disease | Route of Tx | R vs. NR and p-Value |
|---|---|---|---|---|
| Klepanec [25] et al., 2012 | BMMC | CLTI | IM and IA |
|
| Pan [27] et al., 2019 | PBMC and CD34+ | CLTI | IM |
|
| Madaric [26] et al., 2016 | BMMC | CLTI | IM and IA |
|
| Malyar [31] et al., 2014 | BMMC | CLTI/PAD | IM and IA |
|
| Steinhoff [15] et al., 2017 | CD133+ | CAD/MI | IMyo |
|
| Wolfien [10] et al., 2020 | CD133+ | CAD/MI | IMyo |
|
| TAO | ASO |
|---|---|
| TAO is an inflammatory vascular disease that predominantly affects small-sized and medium-sized blood vessels of extremities [37]. | ASO affects medium-sized or large-sized blood vessels of extremities based on atherosclerotic pathologies [1]. |
| Epidemiology: The prevalence of the PAD varies (0.5 to 5.6% in western Europe, 45 to 63% in India, 16 to 66% in Korea and Japan, and 80% in Israel among Jews of Ashkenazi ancestry) [38]. | Epidemiology: The prevalence of ASO was 10.69%, while that of critical limb ischemia was 1.33%. ASO increased by 28.7% in low-income and middle-income countries and by 13.1% in high-income countries [3,39]. |
| Onset: before 45 years [37]. | Onset: ASO prevalence and incidence are both age-related and increase by > 10% among patients aged > 60 and 70 years [1]. |
| Risk factors: Tobacco smoking is the major risk factor for the initiation, maintenance, and progression of TAO [38,40,41] | Risk factors: Genetic background, age, cigarette smoking, diabetes, dyslipidemia, and hypertension [1,4,42,43]. |
| In TAO, cellular immunity increased against Collagen type I and III. For example, anti-collagen antibody activity in TAO is higher than that in ASO [44]. | |
| Biomarker: TAO cases exhibit significantly upregulated plasma levels of IL-6, sICAM-1), and sVCAM-1, and the involved arterial tissues express upregulated levels of p-STAT3, ICAM-1, and VCAM-1 [45]. | Biomarker: CRP, IL-6, IL-1b, and fibrinogen levels are upregulated in ASO. Macrophages and CRP were detected in atherosclerotic plaque [25,26]. |
| PAD | Share Point | Somatic Mutation |
|---|---|---|
| The risk of PAD markedly increases with age. Prevalence of PAD among individuals aged 80–100 years is 22 to 33% [69]. | Age | Somatic mutation incidence increases in the aging population by 10 to 20 % at the age of 70 years [6,28]. |
| Atherosclerosis incidence among PAD cases is more than 90% [39]. | Atherosclerosis | Presence of somatic mutation in TET2 increases atherosclerotic lesions of vessels by 3 to 4-fold [5,70]. |
| Diabetic patients have two to four-fold increased risk of developing PAD, CAD, and ischemic stroke [39]. | Diabetes mellitus | Clonal mosaic event carriers with type 2 diabetes mellitus were associated with increased (2-fold) prevalence of vascular complications compared to non-carriers with type 2 diabetes mellitus (p = 7.7 × 10−4) [50,71]. |
| Smoking is the most common risk factor for PAD occurrence, with a population attributable fraction of 44% [1,39]. | Tobacco smoking | Tobacco smoking significantly increased the occurrence of somatic mutation to 1000–10,000 mutations per cell [72]. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salybekov, A.A.; Wolfien, M.; Kobayashi, S.; Steinhoff, G.; Asahara, T. Personalized Cell Therapy for Patients with Peripheral Arterial Diseases in the Context of Genetic Alterations: Artificial Intelligence-Based Responder and Non-Responder Prediction. Cells 2021, 10, 3266. https://doi.org/10.3390/cells10123266
Salybekov AA, Wolfien M, Kobayashi S, Steinhoff G, Asahara T. Personalized Cell Therapy for Patients with Peripheral Arterial Diseases in the Context of Genetic Alterations: Artificial Intelligence-Based Responder and Non-Responder Prediction. Cells. 2021; 10(12):3266. https://doi.org/10.3390/cells10123266
Chicago/Turabian StyleSalybekov, Amankeldi A., Markus Wolfien, Shuzo Kobayashi, Gustav Steinhoff, and Takayuki Asahara. 2021. "Personalized Cell Therapy for Patients with Peripheral Arterial Diseases in the Context of Genetic Alterations: Artificial Intelligence-Based Responder and Non-Responder Prediction" Cells 10, no. 12: 3266. https://doi.org/10.3390/cells10123266
APA StyleSalybekov, A. A., Wolfien, M., Kobayashi, S., Steinhoff, G., & Asahara, T. (2021). Personalized Cell Therapy for Patients with Peripheral Arterial Diseases in the Context of Genetic Alterations: Artificial Intelligence-Based Responder and Non-Responder Prediction. Cells, 10(12), 3266. https://doi.org/10.3390/cells10123266