Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sequence Alignment
2.2. SARS-CoV-2 YLQ Conservation
2.3. Generation of Peptide-Specific CD8+ T Cell Lines
2.4. Intracellular Cytokine Assay
2.5. Protein Refold, Purification, Crystallisation
2.6. Structure Determination
2.7. Stability Assay
2.8. Surface Plasmon Resonance (SPR)
3. Results
3.1. The YLQ Epitope Induced a Polyfunctional CD8+ T Cell Response in COVID-19 Recovered Donors
3.2. The Conserved YLQ Peptide Is Stably Presented by the HLA-A*02:01 Molecule
| Residue | Y | L | Q | P | R | T | F | L | L |
|---|---|---|---|---|---|---|---|---|---|
| H (0.006%) | V (0.005%) | K (0.005%) | S (0.012%) | K (0.014%) | S (0.004%) | L (0.002%) | |||
| mutation | D (0.005%) | E (0.003%) | L (0.431%) | M (0.009%) | I (0.014%) | ||||
| C (0.005%) | R (0.009%) | H (0.012%) | S (0.021%) | ||||||
| L (0.002%) | |||||||||
| % variant | 0.016 | 0.005 | 0.021 | 0.456 | 0.045 | 0.018 | 0.002 | 0 | 0 |
| Data Collection Statistics | HLA-A*02:01-YLQ | YLQ-SG3 TCR HLA-A*02:01-YLQ |
|---|---|---|
| Space group | P21 | C2 |
| Cell Dimensions (a,b,c) (Å) | 54.09, 80.12, 58.50 | 225.36, 49.62, 91.72, β = 91.83° |
| Resolution (Å) | 47.43–2.05 (2.11–2.05) | 48.68–2.60 (2.72–2.60) |
| Total number of observations | 92,231 (7410) | 140,103 (16,773) |
| Nb of unique observation | 27,671 (2172) | 31,710 (3800) |
| Multiplicity | 3.3 (3.4) | 4.4 (4.4) |
| Data completeness (%) | 97.5 (97.8) | 100 (99.9) |
| I/σI | 8.6 (2.1) | 7.0 (2.2) |
| Rpim a (%) | 6.7 (43.8) | 7.5 (49.3) |
| CC1/2 | 0.993 (0.593) | 0.992 (0.823) |
| Refinement Statistics | ||
| Rfactor b (%) | 19.6 | 19.1 |
| Rfree b (%) | 23.4 | 23.5 |
| rmsd from ideality | ||
| Bond lengths (Å) | 0.01 | 0.01 |
| Bond angles (°) | 1.07 | 1.18 |
| Ramachandran plot (%) | ||
| Favoured | 99.0 | 95.1 |
| Allowed | 0.01 | 4.7 |
| Disallowed | 0 | 0.2 |
| PBD code | 7RDT | 7RTR |
3.3. The Dominant YLQ Peptide Is Recognised by Public TCRs
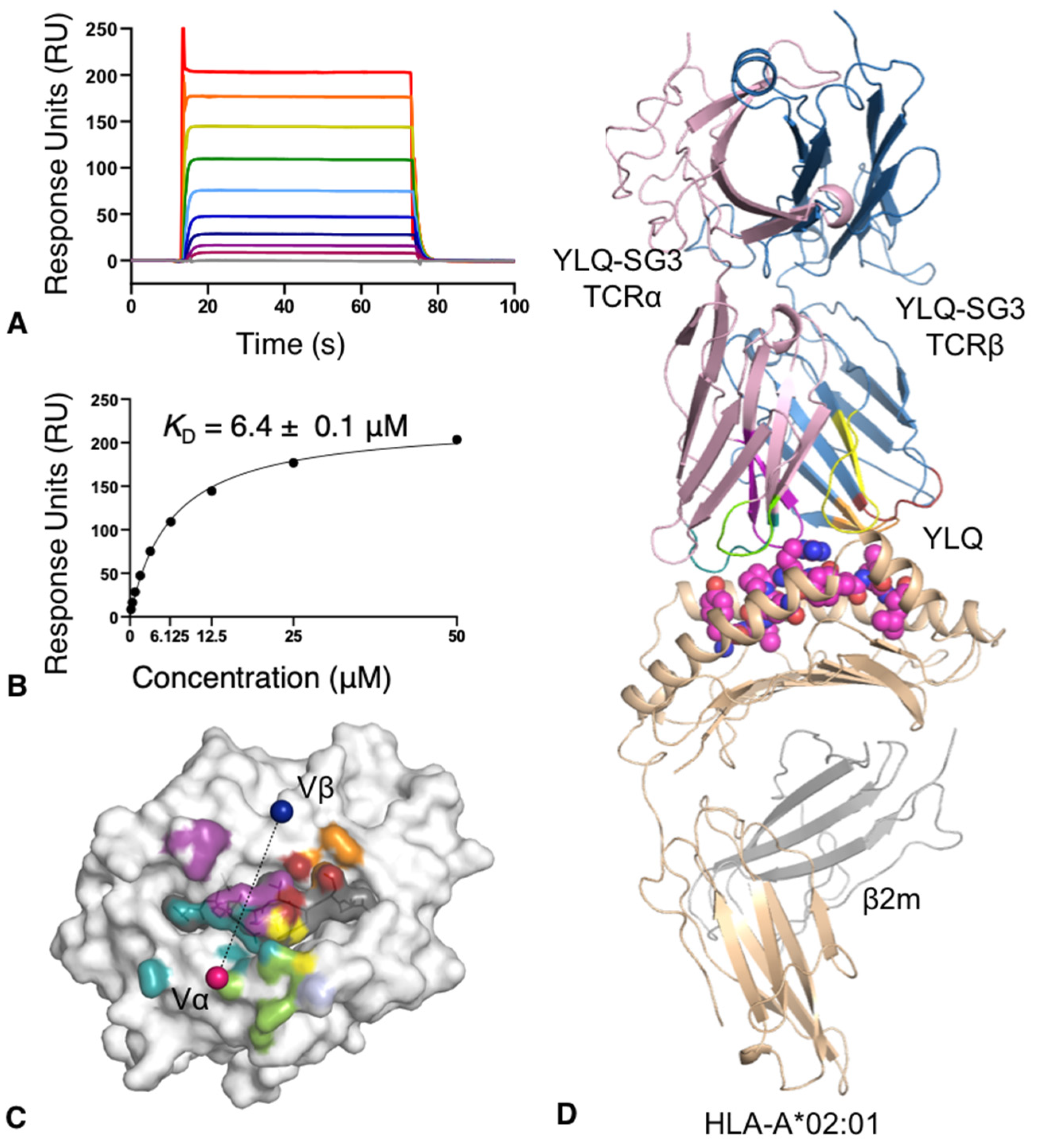
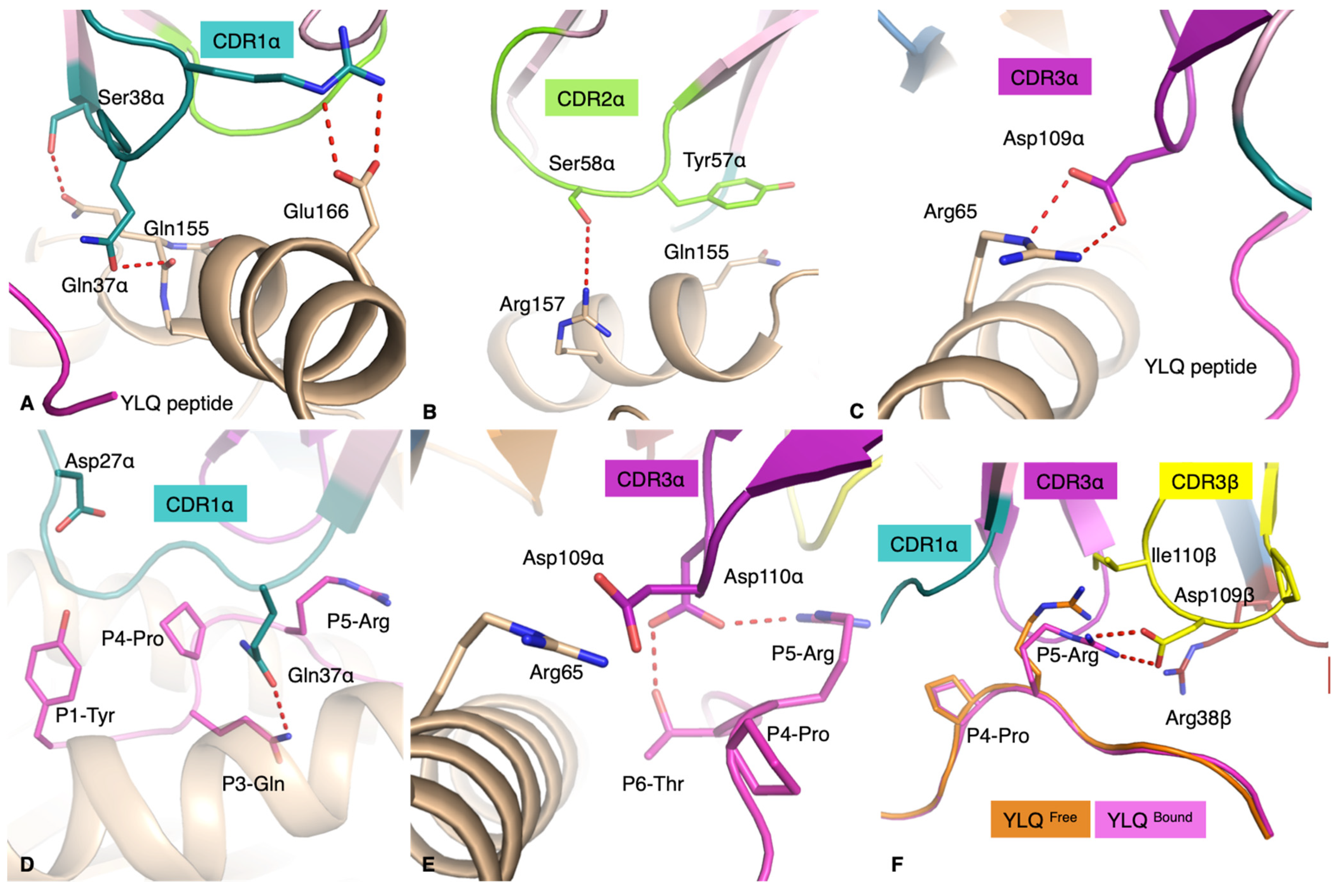
3.4. Structure of the Public YLQ-SG3 TCR Recognising the Dominant YLQ Epitope Presented by HLA-A*02:01
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- McMichael, A.J.; Gotch, F.M.; Noble, G.R.; Beare, P.A.S. Cytotoxic T-Cell Immunity to Influenza. N. Engl. J. Med. 1983, 309, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wan, Y.; Qiu, C.; Quiñones-Parra, S.; Zhu, Z.; Loh, L.; Tian, D.; Ren, Y.; Hu, Y.; Zhang, X.; et al. Recovery from severe H7N9 disease is associated with diverse response mechanisms dominated by CD8+ T cells. Nat. Commun. 2015, 6, 6833. [Google Scholar] [CrossRef] [PubMed]
- Grant, E.; Quiñones-Parra, S.M.; Clemens, E.B.; Kedzierska, K. Human influenza viruses and CD8+ T cell responses. Curr. Opin. Virol. 2016, 16, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Sridhar, S.; Begom, S.; Bermingham, A.; Hoschler, K.; Adamson, W.; Carman, W.; Bean, T.; Barclay, W.; Deeks, J.; Lalvani, A. Cellular immune correlates of protection against symptomatic pandemic influenza. Nat. Med. 2013, 19, 1305–1312. [Google Scholar] [CrossRef]
- Geers, D.; Shamier, M.C.; Bogers, S.; den Hartog, G.; Gommers, L.; Nieuwkoop, N.N.; Schmitz, K.S.; Rijsbergen, L.C.; van Osch, J.A.; Dijkhuizen, E.; et al. SARS-CoV-2 variants of concern partially escape humoral but not T-cell responses in COVID-19 convalescent donors and vaccinees. Sci. Immunol. 2021, 6, eabj1750. [Google Scholar] [CrossRef]
- Szeto, C.; Lobos, C.A.; Nguyen, A.T.; Gras, S. TCR Recognition of Peptide–MHC-I: Rule Makers and Breakers. Int. J. Mol. Sci. 2020, 22, 68. [Google Scholar] [CrossRef]
- Vita, R.; Mahajan, S.; Overton, J.A.; Dhanda, S.K.; Martini, S.; Cantrell, J.R.; Wheeler, D.K.; Sette, A.; Peters, B. The Immune Epitope Database (IEDB): 2018 update. Nucleic Acids Res. 2019, 47, D339–D343. [Google Scholar] [CrossRef] [Green Version]
- Ellis, J.M.; Henson, V.; Slack, R.; Ng, J.; Hartzman, R.J.; Hurley, C.K. Frequencies of HLA-A2 alleles in five U.S. population groups: Predominance of A∗02011 and identification of HLA-A∗0231. Hum. Immunol. 2000, 61, 334–340. [Google Scholar] [CrossRef]
- Agerer, B.; Koblischke, M.; Gudipati, V.; Montano-Gutierrez, L.F.; Smyth, M.; Popa, A.; Genger, J.W.; Endler, L.; Florian, D.M.; Mühlgrabner, V.; et al. SARS-CoV-2 mutations in MHC-I-restricted epitopes evade CD8(+) T cell responses. Sci. Immunol. 2021, 6, eabg6461. [Google Scholar] [CrossRef]
- Gallerani, E.; Proietto, D.; Dallan, B.; Campagnaro, M.; Pacifico, S.; Albanese, V.; Marzola, E.; Marconi, P.; Caputo, A.; Appay, V.; et al. Impaired Priming of SARS-CoV-2-Specific Naive CD8(+) T Cells in Older Subjects. Front. Immunol. 2021, 12, 693054. [Google Scholar] [CrossRef]
- Gangaev, A.; Ketelaars, S.L.C.; Isaeva, O.I.; Patiwael, S.; Dopler, A.; Hoefakker, K.; De Biasi, S.; Gibellini, L.; Mussini, C.; Guaraldi, G.; et al. Identification and characterization of a SARS-CoV-2 specific CD8(+) T cell response with immunodominant features. Nat. Commun. 2021, 12, 2593. [Google Scholar] [CrossRef]
- Kared, H.; Redd, A.D.; Bloch, E.M.; Bonny, T.S.; Sumatoh, H.; Kairi, F.; Carbajo, D.; Abel, B.; Newell, E.W.; Bettinotti, M.P.; et al. SARS-CoV-2-specific CD8+ T cell responses in convalescent COVID-19 individuals. J. Clin. Investig. 2021, 131, e145476. [Google Scholar] [CrossRef]
- Saini, S.K.; Hersby, D.S.; Tamhane, T.; Povlsen, H.R.; Amaya Hernandez, S.P.; Nielsen, M.; Gang, A.O.; Hadrup, S.R. SARS-CoV-2 genome-wide T cell epitope mapping reveals immunodominance and substantial CD8(+) T cell activation in COVID-19 patients. Sci. Immunol. 2021, 6, eabf7550. [Google Scholar] [CrossRef] [PubMed]
- Tarke, A.; Sidney, J.; Kidd, C.K.; Dan, J.M.; Ramirez, S.I.; Yu, E.D.; Mateus, J.; da Silva Antunes, R.; Moore, E.; Rubiro, P.; et al. Comprehensive analysis of T cell immunodominance and immunoprevalence of SARS-CoV-2 epitopes in COVID-19 cases. Cell Rep. Med. 2021, 2, 100204. [Google Scholar] [CrossRef]
- Minervina, A.A.; Komech, E.A.; Titov, A.; Koraichi, M.B.; Rosati, E.; Mamedov, I.Z.; Franke, A.; Efimov, G.A.; Chudakov, D.M.; Mora, T.; et al. Longitudinal high-throughput TCR repertoire profiling reveals the dynamics of T-cell memory formation after mild COVID-19 infection. eLife 2021, 10, e63502. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.H.O.; Rowntree, L.C.; Petersen, J.; Chua, B.Y.; Hensen, L.; Kedzierski, L.; van de Sandt, C.E.; Chaurasia, P.; Tan, H.X.; Habel, J.R.; et al. CD8+ T cells specific for an immunodominant SARS-CoV-2 nucleocapsid epitope display high naive precursor frequency and TCR promiscuity. Immunity 2021, 54, 1066–1082. [Google Scholar] [CrossRef] [PubMed]
- Shomuradova, A.S.; Vagida, M.S.; Sheetikov, S.A.; Zornikova, K.V.; Kiryukhin, D.; Titov, A.; Peshkova, I.O.; Khmelevskaya, A.; Dianov, D.V.; Malasheva, M.; et al. SARS-CoV-2 Epitopes Are Recognized by a Public and Diverse Repertoire of Human T Cell Receptors. Immunity 2020, 53, 1245–1257. [Google Scholar] [CrossRef]
- Davis, M.M.; Bjorkman, P.J. T-cell antigen receptor genes and T-cell recognition. Nat. Cell Biol. 1988, 334, 395–402. [Google Scholar] [CrossRef]
- Cabaniols, J.P.; Fazilleau, N.; Casrouge, A.; Kourilsky, P.; Kanellopoulos, J.M. Most alpha/beta T cell receptor diversity is due to terminal deoxynucleotidyl transferase. J. Exp. Med. 2001, 194, 1385–1390. [Google Scholar] [CrossRef]
- Arstila, T.P.; Casrouge, A.; Baron, V.; Even, J.; Kanellopoulos, J.; Kourilsky, P. A direct estimate of the human alphabeta T cell receptor diversity. Science 1999, 286, 958–961. [Google Scholar] [CrossRef]
- Gras, S.; Kjer-Nielsen, L.; Burrows, S.R.; McCluskey, J.; Rossjohn, J. T-cell receptor bias and immunity. Curr. Opin. Immunol. 2008, 20, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Valkenburg, S.A.; Josephs, T.M.; Clemens, E.B.; Grant, E.; Nguyen, T.H.O.; Wang, G.C.; Price, D.; Miller, A.; Tong, S.; Thomas, P.G.; et al. Molecular basis for universal HLA-A*0201–restricted CD8+ T-cell immunity against influenza viruses. Proc. Natl. Acad. Sci. USA 2016, 113, 4440–4445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Venturi, V.; Kedzierska, K.; Price, D.A.; Doherty, P.C.; Douek, D.C.; Turner, S.J.; Davenport, M.P. Sharing of T cell receptors in antigen-specific responses is driven by convergent recombination. Proc. Natl. Acad. Sci. USA 2006, 103, 18691–18696. [Google Scholar] [CrossRef] [Green Version]
- Garcia, K.C.; Adams, J.J.; Feng, D.; Ely, L.K. The molecular basis of TCR germline bias for MHC is surprisingly simple. Nat. Immunol. 2009, 10, 143–147. [Google Scholar] [CrossRef]
- Lineburg, K.E.; Grant, E.J.; Swaminathan, S.; Chatzileontiadou, D.S.M.; Szeto, C.; Sloane, H.; Panikkar, A.; Raju, J.; Crooks, P.; Rehan, S.; et al. CD8+ T cells specific for an immunodominant SARS-CoV-2 nucleocapsid epitope cross-react with selective seasonal coronaviruses. Immunity 2021, 54, 1055–1065.e5. [Google Scholar] [CrossRef]
- Grant, E.; Josephs, T.M.; Loh, L.; Clemens, E.B.; Sant, S.; Bharadwaj, M.; Chen, W.; Rossjohn, J.; Gras, S.; Kedzierska, K. Broad CD8+ T cell cross-recognition of distinct influenza A strains in humans. Nat. Commun. 2018, 9, 5427. [Google Scholar] [CrossRef] [PubMed]
- Kedzierska, K.; Turner, S.J.; Doherty, P.C. Conserved T cell receptor usage in primary and recall responses to an immunodominant influenza virus nucleoprotein epitope. Proc. Natl. Acad. Sci. USA 2004, 101, 4942–4947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Szeto, C.; Chatzileontiadou, D.S.M.; Nguyen, A.T.; Sloane, H.; Lobos, C.A.; Jayasinghe, D.; Halim, H.; Smith, C.; Riboldi-Tunnicliffe, A.; Grant, E.J.; et al. The presentation of SARS-CoV-2 peptides by the common HLA-A*02:01 molecule. iScience 2021, 24, 102096. [Google Scholar] [CrossRef]
- Chatzileontiadou, D.S.M.; Szeto, C.; Jayasinghe, D.; Gras, S. Protein purification and crystallization of HLA-A∗02:01 in complex with SARS-CoV-2 peptides. STAR Protoc. 2021, 2, 100635. [Google Scholar] [CrossRef] [PubMed]
- Aragão, D.; Aishima, J.; Cherukuvada, H.; Clarken, R.; Clift, M.; Cowieson, N.P.; Ericsson, D.J.; Gee, C.L.; Macedo, S.; Mudie, N.; et al. MX2: A high-flux undulator microfocus beamline serving both the chemical and macromolecular crystallography communities at the Australian Synchrotron. J. Synchrotron Radiat. 2018, 25, 885–891. [Google Scholar] [CrossRef] [Green Version]
- Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef] [Green Version]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser crystallographic software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef] [Green Version]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 2011, 67 Pt 4, 235–242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gras, S.; Saulquin, X.; Reiser, J.-B.; Debeaupuis, E.; Echasserieau, K.; Kissenpfennig, A.; Legoux, F.; Chouquet, A.; Le Gorrec, M.; Machillot, P.; et al. Structural Bases for the Affinity-Driven Selection of a Public TCR against a Dominant Human Cytomegalovirus Epitope. J. Immunol. 2009, 183, 430–437. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.; Cowtan, K.D. Features and development of Coot. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef] [Green Version]
- Bricogne, G.; Blanc, E.; Brandl, M.; Flensburg, C.; Keller, P.; Paciorek, W.; Roversi, P.; Sharff, A.; Smart, O.S.; Vonrhein, C.W.T.O.; et al. Buster Version 2.10; Global Phasing Ltd.: Cambridge, UK, 2011. [Google Scholar]
- Day, E.B.; Guillonneau, C.; Gras, S.; La Gruta, N.; Vignali, D.A.A.; Doherty, P.C.; Purcell, A.; Rossjohn, J.; Turner, S.J. Structural basis for enabling T-cell receptor diversity within biased virus-specific CD8+ T-cell responses. Proc. Natl. Acad. Sci. USA 2011, 108, 9536–9541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cole, D.; Yuan, F.; Rizkallah, P.J.; Miles, J.; Gostick, E.; Price, D.; Gao, G.F.; Jakobsen, B.K.; Sewell, A.K. Germ Line-governed Recognition of a Cancer Epitope by an Immunodominant Human T-cell Receptor. J. Biol. Chem. 2009, 284, 27281–27289. [Google Scholar] [CrossRef] [Green Version]
- Cole, D.; Fuller, A.; Dolton, G.; Zervoudi, E.; Legut, M.; Miles, K.; Blanchfield, L.; Madura, F.; Holland, C.J.; Bulek, A.M.; et al. Dual Molecular Mechanisms Govern Escape at Immunodominant HLA A2-Restricted HIV Epitope. Front. Immunol. 2017, 8, 1503. [Google Scholar] [CrossRef] [Green Version]
- Riley, T.P.; Hellman, L.; Gee, M.H.; Mendoza, J.L.; Alonso, J.A.; Foley, K.C.; Nishimura, M.I.; Kooi, C.W.V.; Garcia, K.C.; Baker, B.M. T cell receptor cross-reactivity expanded by dramatic peptide–MHC adaptability. Nat. Chem. Biol. 2018, 14, 934–942. [Google Scholar] [CrossRef] [PubMed]
- Coles, C.H.; Mulvaney, R.M.; Malla, S.; Walker, A.; Smith, K.J.; Lloyd, A.; Lowe, K.L.; McCully, M.L.; Hague, R.M.; Aleksic, M.; et al. TCRs with Distinct Specificity Profiles Use Different Binding Modes to Engage an Identical Peptide–HLA Complex. J. Immunol. 2020, 204, 1943–1953. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U.; Muik, A.; Vogler, I.; Derhovanessian, E.; Kranz, L.M.; Vormehr, M.; Quandt, J.; Bidmon, N.; Ulges, A.; Baum, A.; et al. BNT162b2 vaccine induces neutralizing antibodies and poly-specific T cells in humans. Nat. Cell Biol. 2021, 595, 572–577. [Google Scholar] [CrossRef]


| Virus | YLQ Homologue | Sequence Identity (%) |
|---|---|---|
| SARS-CoV-2 | YLQPRTFLL | - |
| OC43 | PLTSRQYLL | 44 |
| HKU-1 | PLSRRQYLL | 44 |
| 229E | ALPKTVREF | 11 |
| NL63 | FGPSSQPYY | 0 |
| Study | TRAV | CDR3α | TRBV | CDR3β |
|---|---|---|---|---|
| [18] | 12-1 (74.7%) | CAVNDDKIIF, CAVNxDDKIIF, CAVNxxDDKIIF (23%) | 7-9 (21.3%) | CASSPDIxxxF (32%) |
| 20-1 (13.6%) | ||||
| 12-2 (10.1%) | CAVNxDDKIIF (48.4%) | 2 (12.2%) | ||
| [17] | 12-1 (23.9%) | CVVNxD, CVVNxxD/N (65.8%) | 7-9 (12.6%) | CASSPDIEAFF (33%) |
| 12-2 (3.5%) | CAVNxDDKIIF (100%) | 20-1 (20.1%) | ||
| 2 (12.1%) | ||||
| [16] | 12-1 (58.3%) | CVVNDx, CVVNxDN, CVVNxxN (37.5%) | 7-9 (17.6%) | CASSPDIEAFF (100%) |
| 20-1 (5.8%) | ||||
| 12-2 (33.3%) | CAVNxDDKIIF (50%) | 2 (23.5%) | ||
| YLQ-SG3 [16,18] | 12-2 | CAVNRDDKIIF | 7-9 | CASSPDIEQYF |
| TCR Segment | TCR Residues | HLA-A*02:01 Residues | Type Bond |
|---|---|---|---|
| CDR1α | Arg28Nη1−Nε | Glu166Oε2−Oε1 | VDW, SB |
| CDR1α | Gln37Nε2 | Gln155O, Tyr159 | VDW, HB |
| CDR1α | Ser38Oγ | Gln155Oε1 | VDW, HB |
| FWα | Phe55 | His151 | VDW |
| CDR2α | Tyr57 | Glu154, Gln155, Ala158 | VDW |
| CDR2α | Ser58Oγ | Glu154Oε2, Arg157Nη2 | VDW, HB |
| CDR3α | Asp109Oδ1−Oδ2 | Arg65Nε−Nη2, Lys66 | VDW, SB |
| CDR1β | Arg38 | Thr73 | VDW, HB |
| CDR2β | Gln57Nε2 | Thr73Oγ1, Val76 | VDW |
| CDR2β | Asn58 | Val76 | VDW |
| CDR3β | Asp109Oδ2 | Ala150, Gln155Nε2 | VDW, HB |
| CDR3β | Ile110 | Gln115 | VDW |
| TCR Segment | TCR Residues | YLQ Peptide Residues | Type Bond |
| CDR1α | Asp27Oδ1 | Tyr1OH | HB |
| CDR1α | Gly29 | Tyr1, Pro4 | VDW |
| CDR1α | Gln37Oδ1 | Gln3Nε2, Pro4, Arg5 | VDW, HB |
| CDR1α | Ser38 | Arg5 | VDW |
| CDR3α | Asn107 | Arg5 | VDW |
| CDR3α | Asp109O−Oδ1 | Pro4, Arg5Nε−Nη1 | VDW, HB, SB |
| CDR3α | Asp110Oδ1−Oδ2 | Arg5, Thr6O−N−Oγ | VDW, HB |
| CDR1β | Asn37 | Leu8 | VDW |
| CDR1β | Arg38Nη2 | Arg5, Thr6O, Leu8 | VDW, HB |
| CDR2β | Gln57 | Leu8 | VDW |
| CDR3β | Asp109O−Oδ2−Oδ1 | Arg5Nη1−Nη2, Leu8 | VDW, HB, SB |
| CDR3β | Ile110 | Arg5 | VDW |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szeto, C.; Nguyen, A.T.; Lobos, C.A.; Chatzileontiadou, D.S.M.; Jayasinghe, D.; Grant, E.J.; Riboldi-Tunnicliffe, A.; Smith, C.; Gras, S. Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR. Cells 2021, 10, 2646. https://doi.org/10.3390/cells10102646
Szeto C, Nguyen AT, Lobos CA, Chatzileontiadou DSM, Jayasinghe D, Grant EJ, Riboldi-Tunnicliffe A, Smith C, Gras S. Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR. Cells. 2021; 10(10):2646. https://doi.org/10.3390/cells10102646
Chicago/Turabian StyleSzeto, Christopher, Andrea T. Nguyen, Christian A. Lobos, Demetra S. M. Chatzileontiadou, Dhilshan Jayasinghe, Emma J. Grant, Alan Riboldi-Tunnicliffe, Corey Smith, and Stephanie Gras. 2021. "Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR" Cells 10, no. 10: 2646. https://doi.org/10.3390/cells10102646
APA StyleSzeto, C., Nguyen, A. T., Lobos, C. A., Chatzileontiadou, D. S. M., Jayasinghe, D., Grant, E. J., Riboldi-Tunnicliffe, A., Smith, C., & Gras, S. (2021). Molecular Basis of a Dominant SARS-CoV-2 Spike-Derived Epitope Presented by HLA-A*02:01 Recognised by a Public TCR. Cells, 10(10), 2646. https://doi.org/10.3390/cells10102646








