Reproductive Cold Stress Tolerance in Sorghum F1 Hybrids is a Heterotic Trait
Abstract
:1. Introduction
2. Material and Methods
2.1. Germplasm
2.2. Field Trials
2.3. Climate Chamber Experiments
2.4. Pollen Analyses via Amphasys® Impedance Flow Cytometry
2.5. Statistical Analyses
3. Results
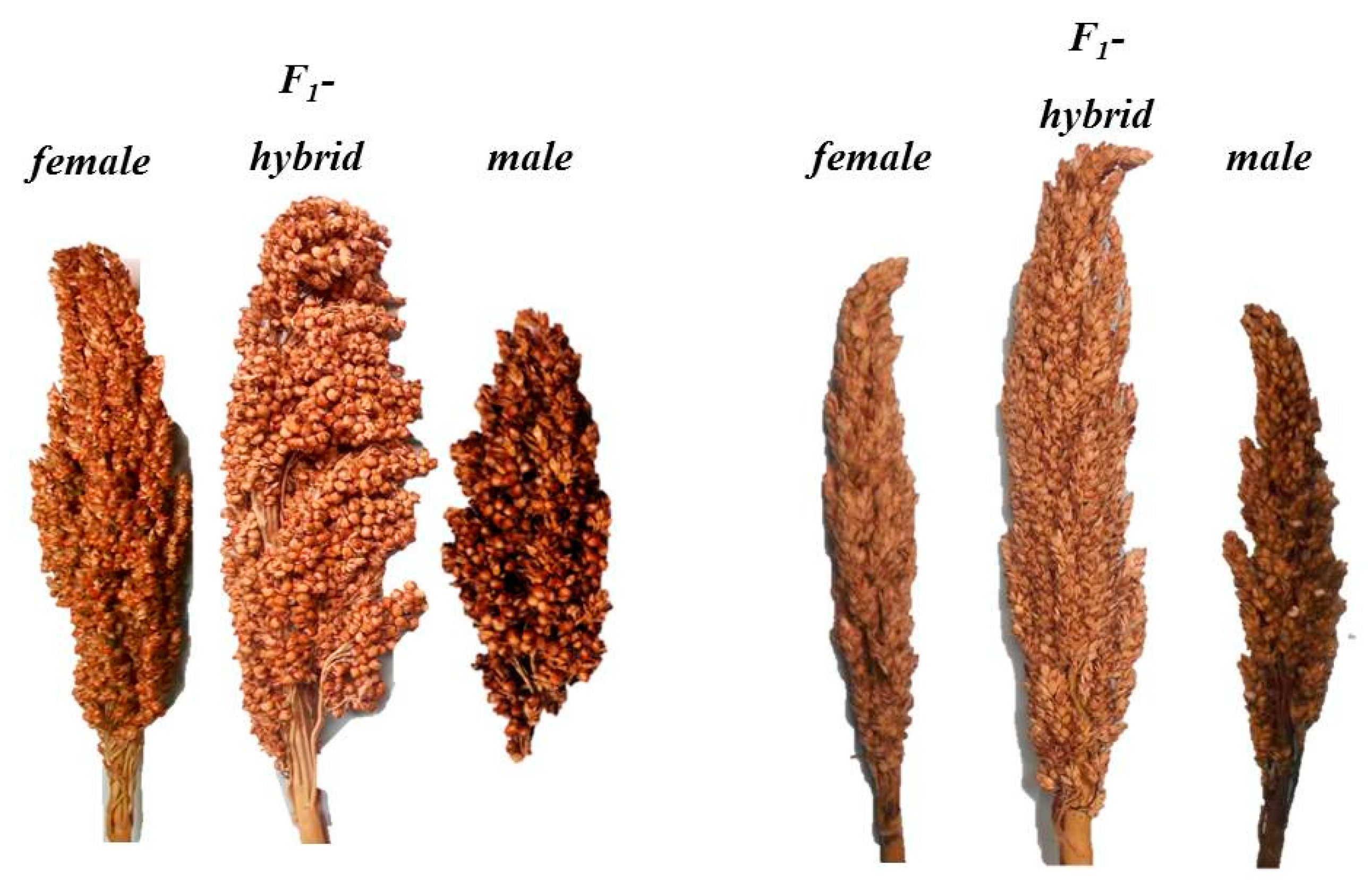
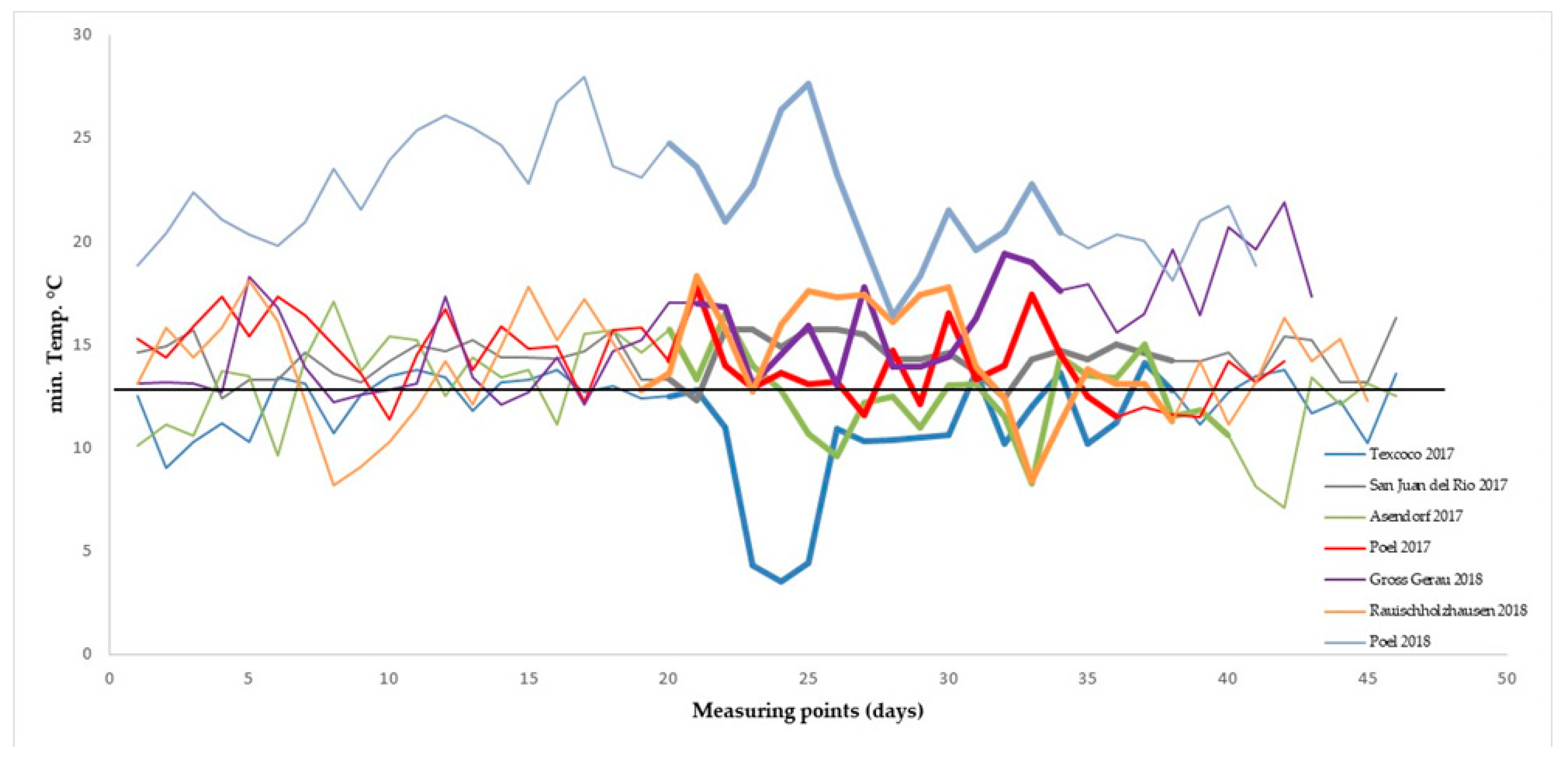
3.1. Variation for SeedSet in Field Trials
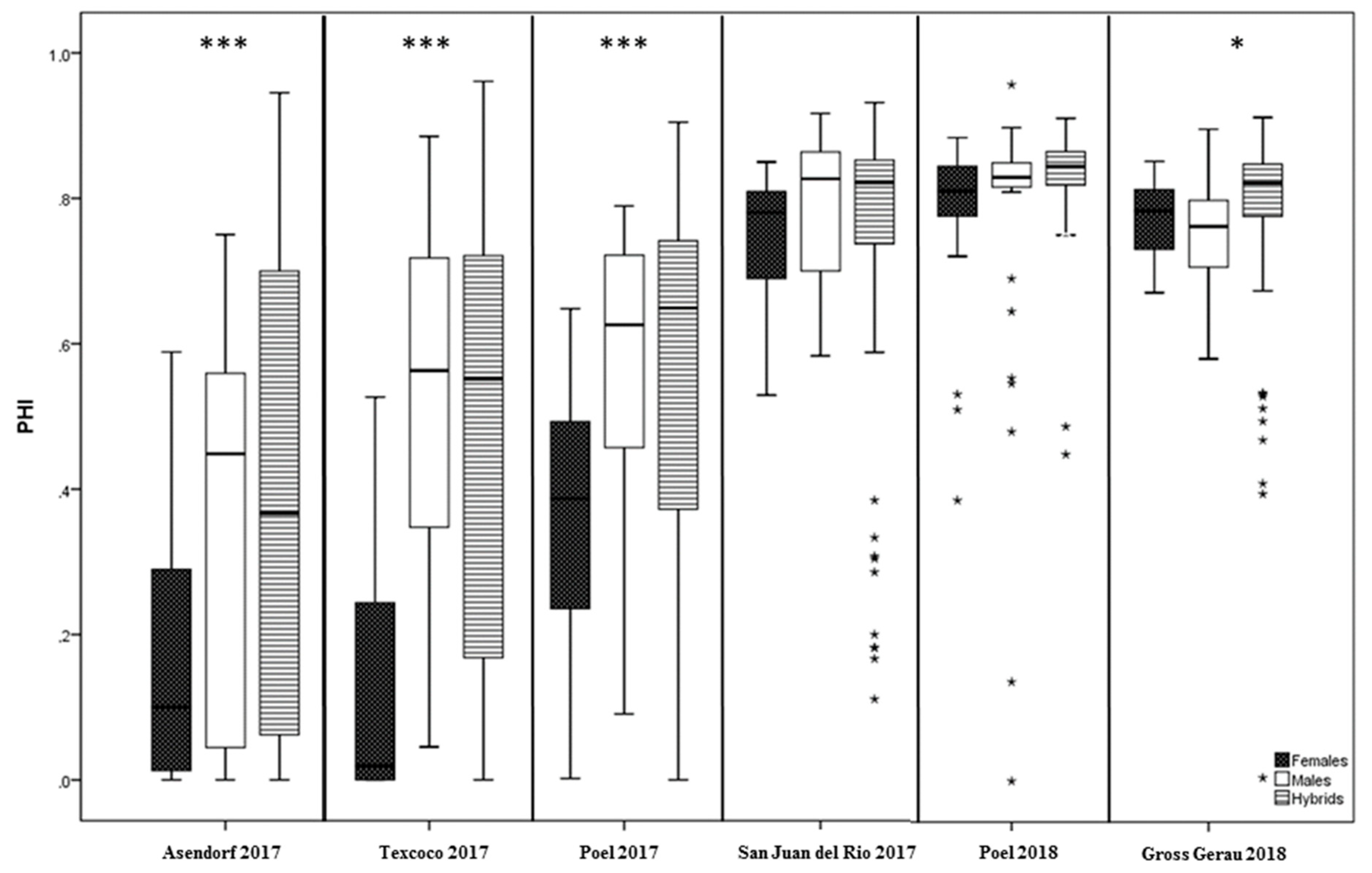
3.2. Variation for PHI and Pollen Traits in Climate Chamber Experiments
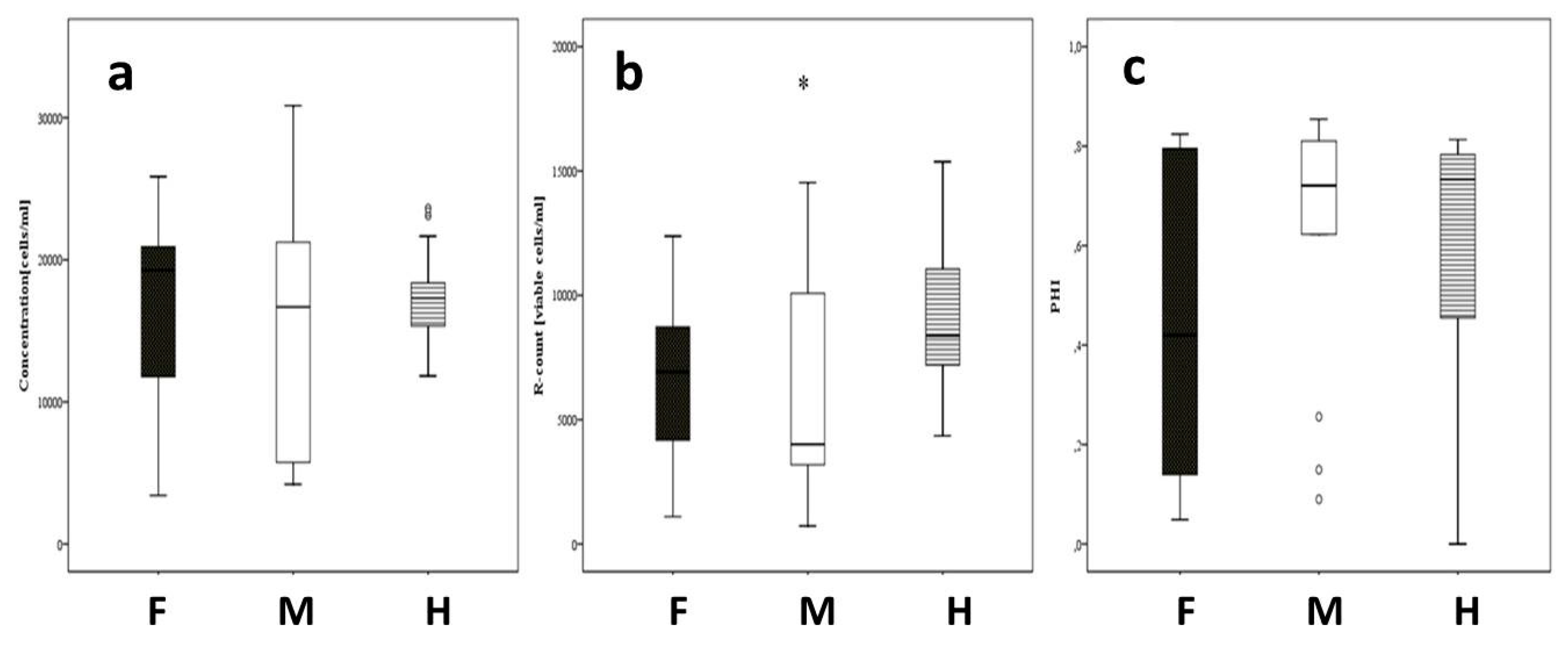
3.3. Analysis of Pollen Traits via IFC for the Field Trials in Gross-Gerau and Rauischholzhausen
3.4. Heterosis
3.5. Combining Ability for Seed Set Traits
3.6. Combining Ability for Pollen Traits
3.7. Relationship between Lineper se and Hybrid Performance
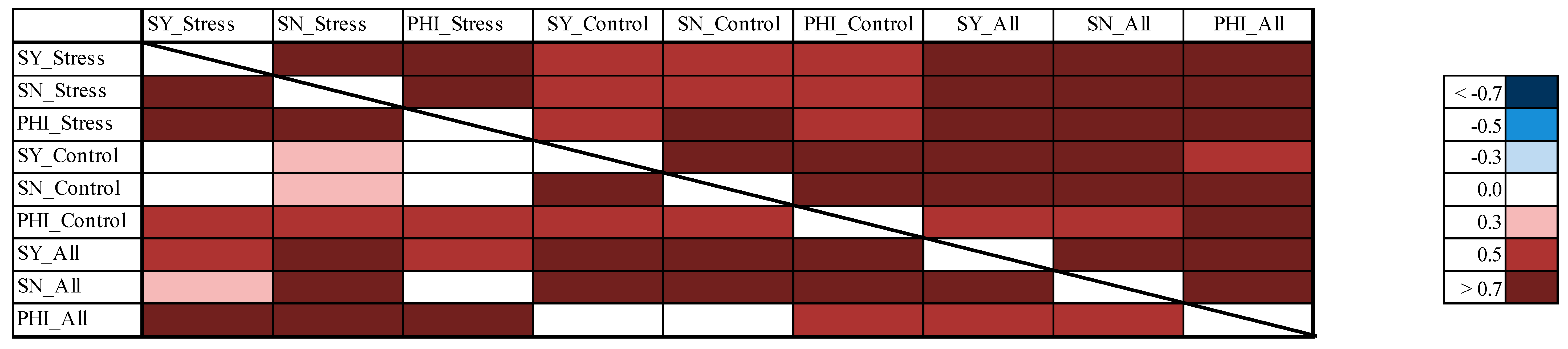
3.8. Correlations between the Different Experiments
4. Discussion
4.1. Heterosis for Reproductive Cold Tolerance
4.2. Combining Ability for Reproductive Cold Tolerance
4.3. General Mode of Inheritance for Reproductive Cold Tolerance
4.4. Implications for Hybrid Breeding on Reproductive Cold Tolerance
4.5. Suitable ScreeningMethods for Reproductive Cold Tolerance
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Singh, S.P. Sources of cold tolerance in grain sorghum. Can. J. Plant Sci. 1985, 65, 251–257. [Google Scholar] [CrossRef]
- Maulana, F.; Weerasooriya, D.; Tesso, T. Sorghum landrace collections from cooler regions of the world exhibit magnificent genetic differentiation and early season cold tolerance. Front.Plant Sci. 2017, 8, 756. [Google Scholar] [CrossRef]
- Brooking, I. Male Sterility in Sorghum bicolor (L.) Moench Induced by Low Night Temperature. I. Timing of the Stage of Sensitivity. Funct. Plant Boil. 1976, 3, 589. [Google Scholar] [CrossRef]
- Bekele, W.A.; Fiedler, K.; Shiringani, A.; Schnaubelt, D.; Windpassinger, S.; Uptmoor, R.; Friedt, W.; Snowdon, R.J. Unravelling the genetic complexity of sorghum seedling development under low-temperature conditions. Plant. Cell Environ. 2014, 37, 707–723. [Google Scholar] [CrossRef]
- Yu, J.; Tuinstra, M.R. Genetic Analysis of Seedling Growth under Cold Temperature Stress in Grain Sorghum. Crop. Sci. 2001, 41, 1438. [Google Scholar] [CrossRef]
- Windpassinger, S.; Friedt, W.; Deppé, I.; Werner, C.; Snowdon, R.; Wittkop, B. Towards Enhancement of Early-Stage Chilling Tolerance and Root Development in Sorghum F1 Hybrids. J. Agron. Crop. Sci. 2017, 203, 146–160. [Google Scholar] [CrossRef]
- Windpassinger, S.; Friedt, W.; Frauen, M.; Snowdon, R.; Wittkop, B. Designing adapted sorghum silage types with an enhanced energy density for biogas generation in temperate Europe. Biomass-Bioenerg. 2015, 81, 496–504. [Google Scholar] [CrossRef]
- Downes, R.; Marshall, D. Low temperature induced male sterility in Sorghum bicolor. Aust. J. Exp. Agric. 1971, 11, 352–356. [Google Scholar] [CrossRef]
- Mendoza-Onofre, L.E. Formación de híbridos de sorgo para grano. II. Comportamiento per se de las líneas y su aptitud combinatoria general. Revis. Fitotec. Mex. 1988, 11, 39–47. [Google Scholar]
- Osuna-Ortega, J.; Mendoza-Onofre, L.E.; González-Hernández, V.A.; Castillo-González, F.; Mendoza-Castillo, M.; Williams-Alanís, H. Potential of cold tolerant germplasm in the adaptation and adaptability of sorghum in México: I. High Valleys. Agrociencia 2000, 34, 561–572. [Google Scholar]
- Osuna-Ortega, J.; Mendoza-Castillo, M.D.C.; Mendoza-Onofre, L. Sorghum cold tolerance, pollen production, and seed yield in the central high valleys of Mexico. Maydica 2003, 48, 125–132. [Google Scholar]
- León-Velasco, H.; Mendoza-Onofre, L.E.; Castillo-González, F.; Cervantes-Santana, T.; Martínez-Garza, Á. Evaluación de dos generaciones de híbridos y progenitores de sorgo tolerantes al frío. II: Aptitude combinatoria, heterosis y heterobeltiosis. Agrociencia 2009, 43, 609–623. [Google Scholar]
- Cisneros-López, M.E.; Mendoza-Onofre, L.E.; Zavaleta-Mancera, H.A.; González-Hernández, V.A.; Mora-Aguilera, G.; Córdova-Téllez, L.; Hernández-Martínez, M. Pollen–pistil interaction, pistil histology and seed production in A× B grain sorghum crosses under chilling field temperatures. J. Agric. Sci. 2010, 148, 73–82. [Google Scholar] [CrossRef]
- Krishnamurthy, L.; Dinakaran, E.; Kumar, A.A.; Reddy, B.V.S. Field Technique and Traits to Assess Reproductive Stage Cold Tolerance in Sorghum (Sorghum bicolor (L.)Moench). Plant. Prod. Sci. 2014, 17, 218–227. [Google Scholar] [CrossRef]
- Srinivasan, A.; Saxena, N.P.; Johansen, C. Cold tolerance during early reproductive growth of chickpea (Cicer arietinum L.): Genetic variation in gamete development and function. Field Crops Res. 1999, 60, 209–222. [Google Scholar] [CrossRef]
- Shinada, H.; Iwata, N.; Sato, T.; Fujino, K. Genetical and morphological characterization of cold tolerance at fertilization stage in rice. Breed.Sci. 2013, 63, 197–204. [Google Scholar] [CrossRef] [Green Version]
- Mitchell, J.H.; Zulkafli, S.L.; Bosse, J.; Campbell, B.; Snell, P.; Mace, E.S.; Godwin, I.D.; Fukai, S. Rice-cold tolerance across reproductive stages. Crop. Pasture Sci. 2016, 67, 823–833. [Google Scholar] [CrossRef]
- Heidmann, I.; Schade-Kampmann, G.; Lambalk, J.; Ottiger, M.; Di Berardino, M. Impedance flow cytometry: A novel technique in pollen analysis. PLoS ONE 2016, 11, e0165531. [Google Scholar] [CrossRef]
- Stephens, J.C.; Holland, R.F. Cytoplasmic Male-Sterility for Hybrid Sorghum Seed Production1. Agron. J. 1954, 46, 20–23. [Google Scholar] [CrossRef]
- Kirby, J.S.; Atkins, R.E. Heterotic Response for Vegetative and Mature Plant Characters in Grain Sorghum, Sorghum bicolor (L.) Moench1. Crop. Sci. 1968, 8, 335–339. [Google Scholar] [CrossRef]
- Pinthus, M.J.; Rosenblum, J. Germination and Seedling Emergence of Sorghum at Low Temperatures. Crop. Sci. 1961, 1, 293–296. [Google Scholar] [CrossRef]
- Piepho, H.-P.; Möhring, J. Computing Heritability and Selection Response from Unbalanced Plant Breeding Trials. Genetics 2007, 177, 1881–1888. [Google Scholar] [CrossRef]
- Hallauer, A.R.; Miranda Fo, J.B. Quantitative Genetics in Plant Breeding, 2nd ed.; Iowa State University Press: Ames, IA, USA, 1988. [Google Scholar]
- Mühleisen, J.; Maurer, H.P.; Stiewe, G.; Bury, P.; Reif, J.C. Hybrid breeding in barley. Crop. Sci. 2013, 53, 819–824. [Google Scholar] [CrossRef]
- Kaw, R.N.; Khush, G.S. Heterosis in traits related to low temperature tolerance in rice. Philipp. J. Crop. Sci. 1985, 10, 93–105. [Google Scholar]
- Goff, S.A. A unifying theory for general multigenic heterosis: Energy efficiency, protein metabolism, and implications for molecular breeding. N. Phytol. 2011, 189, 923–937. [Google Scholar] [CrossRef]
- Duvick, D.H. Heterosis: Feeding people and protecting naturalresources. In The Genetics and Exploitation of Heterosis in Crops; Coors, J.G., Pandey, S., Eds.; American Society of Agronomy, Inc.; Crop Science Society of America, Inc.: Madison, WI, USA, 1999; pp. 19–29. [Google Scholar]
- Flint-Garcia, S.A.; Buckler, E.S.; Tiffin, P.; Ersoz, E.; Springer, N.M. Heterosis is prevalent for multiple traits in diverse maize germplasm. PLoS ONE 2009, 4, e7433. [Google Scholar] [CrossRef]
- Dane, F.; Hunter, A.G.; Chambliss, O.L. Fruit Set, Pollen Fertility, and Combining Ability of Selected Tomato Genotypes under High-temperature Field Conditions. J. Am. Soc. Hortic. Sci. 1991, 116, 906–910. [Google Scholar] [CrossRef]
- Singh, S.; Sahu, P.; Sharma, D.; Ojha, G.C. Combining ability analysis to identify suitable parents for heterotic rice hybrid breeding. Ecoscan 2015, 7, 361–369. [Google Scholar]
- Reif, J.C.; Hallauer, A.R.; Melchinger, A.E. Heterosis and heterotic pattern in maize. Maydica 2005, 50, 215–223. [Google Scholar]
- Ram, N.K.; Moon, H.P.; Yae, J.D.; Visperas, R.M. Estimates of combining ability for cold tolerance at reproductive stage in rice. Korean J. Breed. 1989, 21, 188–195. [Google Scholar]
- Jordan, D.R.; Klein, R.R.; Sakrewski, K.G.; Henzell, R.G.; Klein, P.E.; Mace, E.S. Mapping and characterization of Rf 5: A new gene conditioning pollen fertility restoration in A1 and A2 cytoplasm in sorghum (Sorghum bicolor (L.) Moench). Theor. Appl. Genet. 2011, 123, 383–396. [Google Scholar] [CrossRef]
- Menz, M.A.; Unruh, N.C.; Rooney, W.L.; Klein, P.E.; Mullet, J.E.; Klein, R.R. Genetic Diversity of Public Inbreds of Sorghum Determined by Mapped AFLP and SSR Markers. Crop. Sci. 2004, 44, 1236–1244. [Google Scholar] [CrossRef] [Green Version]
- Marla, S.R.; Burow, G.; Chopra, R.; Hayes, C.; Olatoye, M.; Felderhoff, T.; Hu, Z.; Raymundo, R.; Perumal, R.; Morris, G.P. Genetic architecture of chilling tolerance in sorghum dissected with a nested assocation mapping population. bioRxiv 2019, 622894. [Google Scholar] [CrossRef]
- Dhopte, A.M.; Eastin, J.D. Response of sorghum to elevated night temperature imposed during floret differentiation under field conditions. In Proceedings of the International Congress of Plant Physiology, New Delhi, India, 15–20 February 1988; Volume 2. [Google Scholar]
- Dhopte, A.M.; Eastin, J.D. Influence of night temperature during floret differentiation on microsporogenesis and seedsetting in sorghum (Sorghum bicolor (L.) Moench). Role Biotechnol. Agric. 1992, 193–200. [Google Scholar]
- Brooking, I. Male Sterility in Sorghum bicolor (L.) Moench Induced by Low Night Temperature. II. Genotypic Differences in Sensitivity. Funct. Plant Boil. 1979, 6, 14. [Google Scholar] [CrossRef]

| Site | Coordinates | Altitude | Soil Type | Year | Mean Temp. (°C) | Mean Max. Temp. (°C) | Mean Min. Temp. (°C) | Absolute Max. and Min. Temp. (°C) | Precipitation (mm) |
|---|---|---|---|---|---|---|---|---|---|
| Poel (PO) | 53°99′ N, 11°47′ E | 19 m | Loamy sand | 2017 | 16.7 | 20.2 | 13.1 | 29.5/6.6 | 301 |
| 2018 | 18.7 | 23.2 | 14.2 | 35.6/6.4 | 118 | ||||
| Rauischholz-Hausen (RH) | 50°46′ N, 8°53′ E | 270 m | Loam | 2018 | 18.7 | 26.5 | 10.9 | 37.9/−0.3 | 174 |
| Asendorf (AS) | 52°46′ N, 9°01′ E | 49 m | Loamy sand | 2017 | 16.1 | 21.0 | 11.6 | 31.7/5.1 | 611 |
| Gross-Gerau (GG) | 49°55′ N, 8°29′ E | 90 m | Sand | 2018 | 21.3 | 28.9 | 13.7 | 38.8/5.3 | 94 (+150 irrigation) |
| San Juan del Rio (SJR) | 20°25′ N, 99°56′ W | 1920 m | Loam | 2017 | 23.0 | 31.4 | 14.7 | 36.6/11.6 | 326 |
| Texcoco (TEX) | 19°31′ N, 98°51′ W | 2250 m | Loam | 2017 | 16.4 | 24 | 8.7 | 27.0/−2.4 | 360 |
| Temperature in Celsius | Day/Night Cycle in Hours | Rel. Air Humidity | |
|---|---|---|---|
| Control Treatment | 30/24 | 16/8 | 60% |
| Stress Treatment | 24/7 | 13/11 | 60% |
| Items | All Environments | Stress Environments | Control Environments | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| d.f. | Seed Yield Per Panicle (g) | Seed Number | PHI | d.f. | Seed Yield Per Panicle (g) | Seed Number | PHI | d.f. | Seed Yield Per Panicle (g) | Seed Number | PHI | |
| Entries | 61 | 4263.81 *** | 5,520,078.53 *** | 0.42 *** | 54 | 1995.35 *** | 3,475,582.26 *** | 0.70 *** | 61 | 3534.64 *** | 3,700,130.29 *** | 0.06 *** |
| CV Entries | 2.18 | 1.93 | 1.05 | 3.15 | 2.80 | 1.95 | 1.37 | 1.15 | 0.31 | |||
| Environments (Env) | 5 | 59,090.35 *** | 79,477,724.73 *** | 9.26 *** | 2 | 7634.70 *** | 21,494,186.41 *** | 1.91 *** | 2 | 2724.82 *** | 16,577,269.0 *** | 0.23 *** |
| Entries × Env | 259 | 689.00 *** | 859,084.66 *** | 0.1 *** | 100 | 234.20 *** | 459,181.84 *** | 0.10 *** | 110 | 950.32 *** | 1,032,507.28 *** | 0.03 *** |
| Error | 1158 | 123.6 | 157,059.12 | 0.01 | 521 | 46.44 | 77,405.71 | 0.01 | 661 | 179.93 | 213,954.88 | 0.01 |
| Heritability (H2) | 0.83 | 0.84 | 0.75 | 0.87 | 0.86 | 0.85 | 0.72 | 0.7 | 0.57 | |||
| Lines | 13 | 1159.35 *** | 1,972,584.96 *** | 0.36 *** | 13 | 492.79 *** | 943,314.11 *** | 0.66 *** | 13 | 1415.42 *** | 1,970,893.33 *** | 0.06 *** |
| CV Lines | 1.72 | 1.57 | 1.07 | 2.97 | 2.38 | 2.32 | 1.20 | 1.05 | 0.32 | |||
| Environments (Env) | 5 | 11,348.65 *** | 18,701,493.15 *** | 3.04 *** | 2 | 927.90 *** | 3,878,005.31 *** | 0.69 *** | 2 | 2658.51 *** | 75,731,121.65 *** | 0.01 |
| Lines × Env | 61 | 384.38 *** | 572,938.83 *** | 0.08 *** | 25 | 65.17 *** | 170,723.20 *** | 0.09 *** | 26 | 519.92 *** | 813,648.57 *** | 0.03 *** |
| Error | 295 | 64.69 | 118,969.66 | 0.01 | 148 | 14.05 | 33,708.51 | 0.01 | 163 | 104.32 | 184,820.95 | 0.01 |
| H2 | 0.65 | 0.71 | 0.77 | 0.85 | 0.8 | 0.85 | 0.85 | 0.8 | 0.85 | |||
| Males | 6 | 1501.12 *** | 2,098,936.97 *** | 0.49 *** | 6 | 633.35 *** | 1,132,878.41 *** | 0.62 *** | 6 | 1457.84 *** | 1,646,223.05 *** | 0.08 *** |
| CV Males | 1.94 | 1.63 | 1.13 | 2.41 | 2.10 | 1.64 | 1.30 | 1.02 | 0.37 | |||
| Environments (Env) | 5 | 4172.49 *** | 7,700,134.83 *** | 0.92 *** | 2 | 802.86 *** | 2,288,173.25 *** | 0.29 *** | 2 | 513.53 ** | 3,346,600.43 *** | 0.02 |
| Males × Env | 30 | 418.34 *** | 587,861.50 *** | 0.08 *** | 12 | 85.62 *** | 218,893.87 *** | 0.08 *** | 12 | 678.15 *** | 933,770.85 *** | 0.04 *** |
| Error | 157 | 63.49 | 87,720.17 | 0.01 | 78 | 17.82 | 34,379.41 | 0.01 | 81 | 105.9 | 136,920.52 | 0.01 |
| H2 | 0.73 | 0.73 | 0.81 | 0.88 | 0.79 | 0.87 | 0.58 | 0.47 | 0.53 | |||
| Females | 6 | 964.95 *** | 2,102,749.73 *** | 0.17 *** | 6 | 134.0 *** | 607,000.85 *** | 0.32 *** | 6 | 1485.72 *** | 2,377,031.89 *** | 0.04 *** |
| CV Females | 1.60 | 1.61 | 0.81 | 2.72 | 2.62 | 2.57 | 1.16 | 1.08 | 0.26 | |||
| Environments (Env) | 5 | 7849.99 *** | 12,241,455.85 *** | 2.31 *** | 2 | 230.84 *** | 1,659,332.93 *** | 0.51 *** | 2 | 3022.14 *** | 6,286,193.59 *** | 0.01 |
| Females × Env | 26 | 283.93 *** | 427,133.83 *** | 0.06 *** | 11 | 36.99 *** | 131,383.11 *** | 0.08 *** | 12 | 310.17 ** | 494,212.78 * | 0.02 *** |
| Error | 138 | 66.059 | 154,521.623 | 0.007 | 70 | 9.86 | 32,960.94 | 0.01 | 82 | 102.76 | 232,137.23 | 0.003 |
| H2 | 0.65 | 0.71 | 0.77 | 0.47 | 0.76 | 0.66 | 0.79 | 0.8 | 0.57 | |||
| F vs. M | 17.330 | 14,409.571 | 1.232 *** | 1 | 1885.67 *** | 2,060,061.57 *** | 3.19 *** | 1 | 747.27 | 1,455,133.58 | 0 | |
| Hybrids (H) | 47 | 4312.99 *** | 5,845,056.24 *** | 0.44 *** | 40 | 2228.03 *** | 3,977,574.78 *** | 0.70 *** | 47 | 3411.13 *** | 3,723,235.56 *** | 0.06 *** |
| CV Hybrids | 1.96 | 1.82 | 1.02 | 2.83 | 2.61 | 1.81 | 1.24 | 1.09 | 0.31 | |||
| Environments (Env) | 5 | 49,111.22 *** | 62,037,528.31 *** | 6.30 *** | 2 | 6901.22 *** | 17,764,234.16 *** | 1.27 *** | 2 | 2522.91 *** | 12,040,802.69 *** | 0.25 *** |
| Hybrids × Env | 193 | 760.60 *** | 938,533.24 *** | 0.10 *** | 73 | 290.83 *** | 562,860.35 *** | 0.11 *** | 82 | 1047.37 *** | 1,057,291.96 *** | 0.03 *** |
| GCA (F) | 6 | 12,602.34 *** | 23,111,079.88 *** | 1.89 *** | 6 | 4359.36 *** | 10,771,964.45 *** | 2.62 *** | 6 | 14,651.5 *** | 19,470,675.5 *** | 0.34 *** |
| GCA (M) | 6 | 8273.90 *** | 7,254,811.86 *** | 0.30 *** | 6 | 5752.50 *** | 7,628,863.68 *** | 0.64 *** | 6 | 4299.45 *** | 1,739,162.29 *** | 0.02 *** |
| Ratio GCA F: M | 1.52 | 3.18 | 6.3 | 0.76 | 1.41 | 4.09 | 3.41 | 11.20 | 14.78 | |||
| SCA (F × M) | 35 | 1015.06 *** | 1,152,137.36 *** | 0.07 *** | 28 | 380.65 *** | 613,548.49 *** | 0.11 *** | 35 | 1284.52 *** | 1,277,323.82 *** | 0.02 *** |
| Env × GCA (F) | 30 | 1862.58 *** | 1,507,934.05 *** | 0.28 *** | 12 | 356.82 *** | 455,922.27 *** | 0.18 *** | 12 | 2804.65 *** | 1,886,401.36 *** | 0.09 *** |
| Env × GCA (M) | 30 | 424.13 *** | 1,101,327.01 *** | 0.11 *** | 12 | 232.56 *** | 731,861.38 *** | 0.11 *** | 12 | 576.51 *** | 836,901.45 *** | 0.01 |
| Env × SCA | 133 | 554.90 *** | 729,432.51 *** | 0.05 *** | 49 | 276.75 *** | 498,152.1 *** | 0.08 *** | 58 | 733.09 *** | 886,435.04 *** | 0.02 *** |
| Error | 863 | 143.74 | 170,079.27 | 0.01 | 373 | 59.29 | 94,744.02 | 0.01 | 498 | 204.68 | 223,490.68 | 0.01 |
| H2 | 0.8 | 0.81 | 0.74 | 0.83 | 0.81 | 0.85 | 0.67 | 0.69 | 0.58 | |||
| L vs. H | 1 | 51,085.44 *** | 46,330,361.32 *** | 1.1 *** | 1 | 12,047.69 *** | 17,365,106.10 *** | 1.53 *** | 1 | 38,709.27 *** | 28,897,161.25 *** | 0.15 *** |
| Both Treatments | Stress Treatment | Control Treatment | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Items | d.f | Pollen Amount (Cells/mL) | Viable Pollen (Cells/mL) | PHI | d.f. | Pollen Amount (Cells/mL) | Viable Pollen (Cells/mL) | PHI | d.f. | Pollen Amount (Cells/mL) | Viable Pollen (Cells/mL) | PHI |
| Entries | 11 | 189 mio *** | 54 mio *** | 0.21 *** | 11 | 137 mio *** | 41 mio *** | 0.294 *** | 11 | 87 mio *** | 55 mio *** | 0.034 *** |
| Treatment | 1 | 70 mio * | 2 mio | 1.63 *** | ||||||||
| Entry × Treatment | 11 | 34 mio * | 36 mio *** | 0.124 *** | ||||||||
| Error | 89 | 15 mio | 7 mio | 0.016 | 42 | 10 mio | 6 mio | 0.026 | 47 | 20 mio | 8 mio | 0.007 |
| H2 | 0.83 | 0.25 | 0.40 | |||||||||
| Lines | 6 | 340 mio *** | 83 mio *** | 0.23 *** | 6 | 232 mio *** | 57 mio *** | 0.27 *** | 6 | 154 mio *** | 92 mio *** | 0.06 *** |
| Lines × Treatment | 6 | 58 mio * | 54 mio *** | 0.072 ** | ||||||||
| Error | 49 | 20 mio | 7 mio | 0.022 | 21 | 114 mio | 4 mio | 0.029 | 27 | 27 mio | 10 mio | 0.01 |
| H2 | 0.83 | 0.27 | 0.66 | |||||||||
| Males | 3 | 519 mio *** | 91 mio *** | 0.104 ** | 3 | 318 mio *** | 83 mio *** | 0.17 * | 3 | 270 mio ** | 92 mio ** | 0.003 |
| Males × Treatment | 3 | 81 mio * | 75 mio ** | 0.1 ** | ||||||||
| Error | 26 | 26 mio | 11 mio | 0.016 | 10 | 12 mio | 4 mio | 0.037 | 16 | 35 mio | 16 mio | 0.002 |
| Females | 2 | 242 mio *** | 77 mio *** | 0.41 *** | 2 | 2158 mio | 48 mio | 0.484 | 2 | 43 mio | 32 mio *** | 0.09 |
| Females × Treatment | 2 | 42 mio | 3 mio | 0.062 | ||||||||
| Error | 23 | 13 mio | 3 mio | 0.029 | 11 | 9 mio | 5 mio | 0.021 | 11 | 17 mio | 1 mio | 0.02 |
| F vs. M | 1 | 157,570 | 129 mio ** | 0.44 ** | 1 | 7 mio | 257,664 | 0.16 | 1 | 26 mio | 216 mio ** | 0.185 *** |
| Hybrids | 4 | 8 mio | 12 mio | 0.22 *** | 4 | 5 mio *** | 9 mio ** | 0.41 *** | 4 | 8 mio | 13 mio | 0.001 |
| Hybrids × Treatment | 1 | 69 mio * | 31 mio | 0.23 *** | ||||||||
| Error | 40 | 10 mio | 8 mio | 0.009 | 19 | 11 mio | 9 mio | 0.013 | 20 | 9 mio | 6 mio | 0.005 |
| H2 | 0.83 | 0.25 | 0.40 | |||||||||
| L vs. H | 1 | 2 mio | 23 mio | 0.004 | 1 | 17 mio | 70 mio * | 0 | 1 | 6 mio | 2 mio | 0.001 |
| Items | d.f. | Pollen Amount (Cells/mL) | Viable Pollen (Cells/mL) |
|---|---|---|---|
| Genotype | 30 | 57 mio *** | 24 mio *** |
| Environment (Env) | 1 | 82 mio *** | 25 mio *** |
| Genotype × Environment | 27 | 31 mio *** | 9.4 mio *** |
| Error | 144 | 7.8 mio | 3.6 mio |
| Heritability (H2) | 0.41 | 0.6 | |
| Lines | 9 | 22 mio *** | 19 mio *** |
| Environment (Env) | 1 | 96 mio *** | 23 mio ** |
| Genotype × Environment | 9 | 14 mio ** | 9.5 mio *** |
| Error | 45 | 4.5 mio | 2.5 mio |
| Heritability (H2) | 0.31 | 0.47 | |
| Males | 6 | 25 mio ** | 26 mio *** |
| Environment | 1 | 49 mio ** | 21 mio * |
| Males × Env | 6 | 8.9 mio | 10 mio * |
| Error | 30 | 5.8 mio | 3.2 mio |
| Females | 2 | 10 mio * | 7.6 mio ** |
| Environment | 1 | 55 mio *** | 2.7 mio |
| Females × Env | 2 | 36 mio *** | 8.7 mio ** |
| Error | 15 | 2 mio | 1 mio |
| F vs. M | 1 | 6.7 m | 0.3 m |
| Hybrids | 20 | 70 mio *** | 27 mio *** |
| Environment (Env) | 1 | 836 mio *** | 6.7 mio |
| Genotype × Env | 17 | 37 mio *** | 9.1 mio ** |
| GCA (F) | 2 | 350 mio *** | 176 mio *** |
| GCA (M) | 6 | 49 mio *** | 10 mio * |
| Ratio GCA F: M | 7.1 | 17.2 | |
| Environment (Env) | 1 | 845 mio * | 6.5 mio |
| SCA (F *M) | 12 | 46 mio *** | 11 mio ** |
| GCA (F) *Env | 2 | 176 mio *** | 27 mio ** |
| GCA (M) *Env | 6 | 22 mio * | 6 mio |
| SCA × Env | 10 | 19 mio * | 5 mio |
| Error | 99 | 9.3 mio | 4 mio |
| Heritability (H2) | 0.41 | 0.6 | |
| L vs. H | 1 | 125 mio * | 17 mio |
| Items | All Environments | Stress Environments | Control Environments | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| n | Seed Yield Per Panicle | Seed Number | PHI | Seed Yield Per Panicle | Seed Number | PHI | Seed Yield Per Panicle | Seed Number | PHI | |
| [%] of Hybrids with Sign. HPH | 47 | 37.5 | 35.4 | 0 | 18.3 | 14.3 | 4.1 | 18.3 | 6.1 | 0 |
| Average MPH [%] (all MPV-Hybrid Comparison) | 47 | 79.0 *** | 55.7 *** | 10.9 *** | 123.3 *** | 77.0 *** | 31.4 *** | 50.4 *** | 32.2 *** | 3.9 *** |
| GCA Prediction Accuracy (r2) | 47 | 0.71 *** | 0.70 *** | 0.73 *** | 0.85 *** | 0.86 *** | 0.86 *** | 0.75 ** | 0.74 *** | 0.8 *** |
| Correlation(r) per se M: GCA M | 7 | 0.57 | 0.36 | 0.66 | 0.83 * | 0.69 | 0.73 | 0.14 | −0.3 | 0.35 |
| Correlation (r) per se F: GCA F | 7 | 0.67 | 0.71 | 0.6 | 0.27 | 0.54 | 0.24 | 0.87 ** | 0.89 ** | 0.035 |
| Correlation (r) MPV: Hybrid Performance | 47 | 0.52 *** | 0.47 *** | 0.49 *** | 0.65 *** | 0.61 *** | 0.45 ** | 0.50 *** | 0.51 *** | 0.075 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schaffasz, A.; Windpassinger, S.; Snowdon, R.; Wittkop, B. Reproductive Cold Stress Tolerance in Sorghum F1 Hybrids is a Heterotic Trait. Agronomy 2019, 9, 508. https://doi.org/10.3390/agronomy9090508
Schaffasz A, Windpassinger S, Snowdon R, Wittkop B. Reproductive Cold Stress Tolerance in Sorghum F1 Hybrids is a Heterotic Trait. Agronomy. 2019; 9(9):508. https://doi.org/10.3390/agronomy9090508
Chicago/Turabian StyleSchaffasz, André, Steffen Windpassinger, Rod Snowdon, and Benjamin Wittkop. 2019. "Reproductive Cold Stress Tolerance in Sorghum F1 Hybrids is a Heterotic Trait" Agronomy 9, no. 9: 508. https://doi.org/10.3390/agronomy9090508
APA StyleSchaffasz, A., Windpassinger, S., Snowdon, R., & Wittkop, B. (2019). Reproductive Cold Stress Tolerance in Sorghum F1 Hybrids is a Heterotic Trait. Agronomy, 9(9), 508. https://doi.org/10.3390/agronomy9090508




