Mycorrhizal Fungi Enhance Resistance to Herbivores in Tomato Plants with Reduced Jasmonic Acid Production
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Growth and Mycorrhizal Inoculation
2.2. Plant Growth and Resistance Bioassay
2.3. Statistical Analyses
2.3.1. Plant Traits
2.3.2. Resistance
2.3.3. AM-Colonization
2.3.4. Multivariate Analysis
3. Results
3.1. Plant Functional Traits
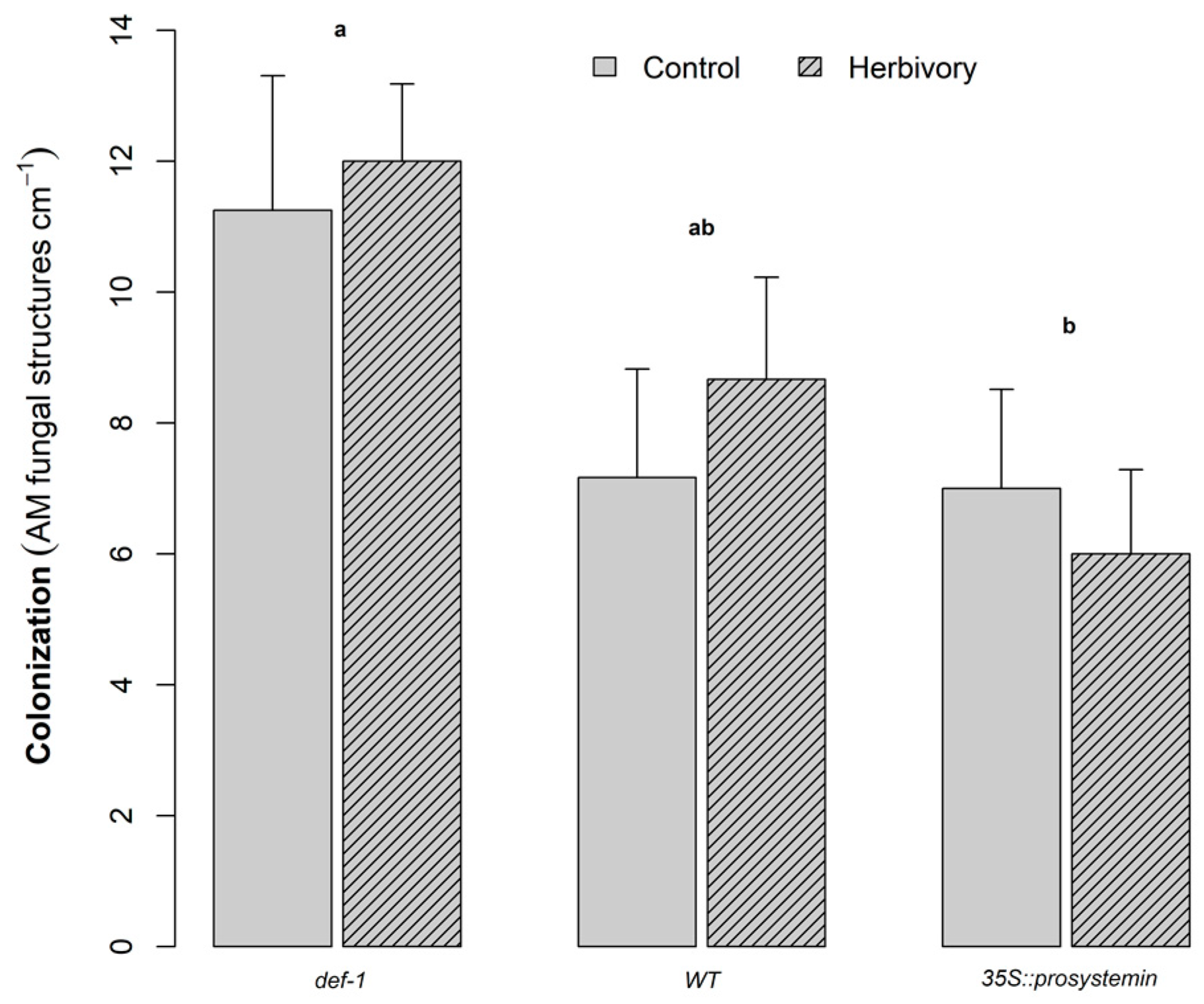
3.2. AMF Colonization
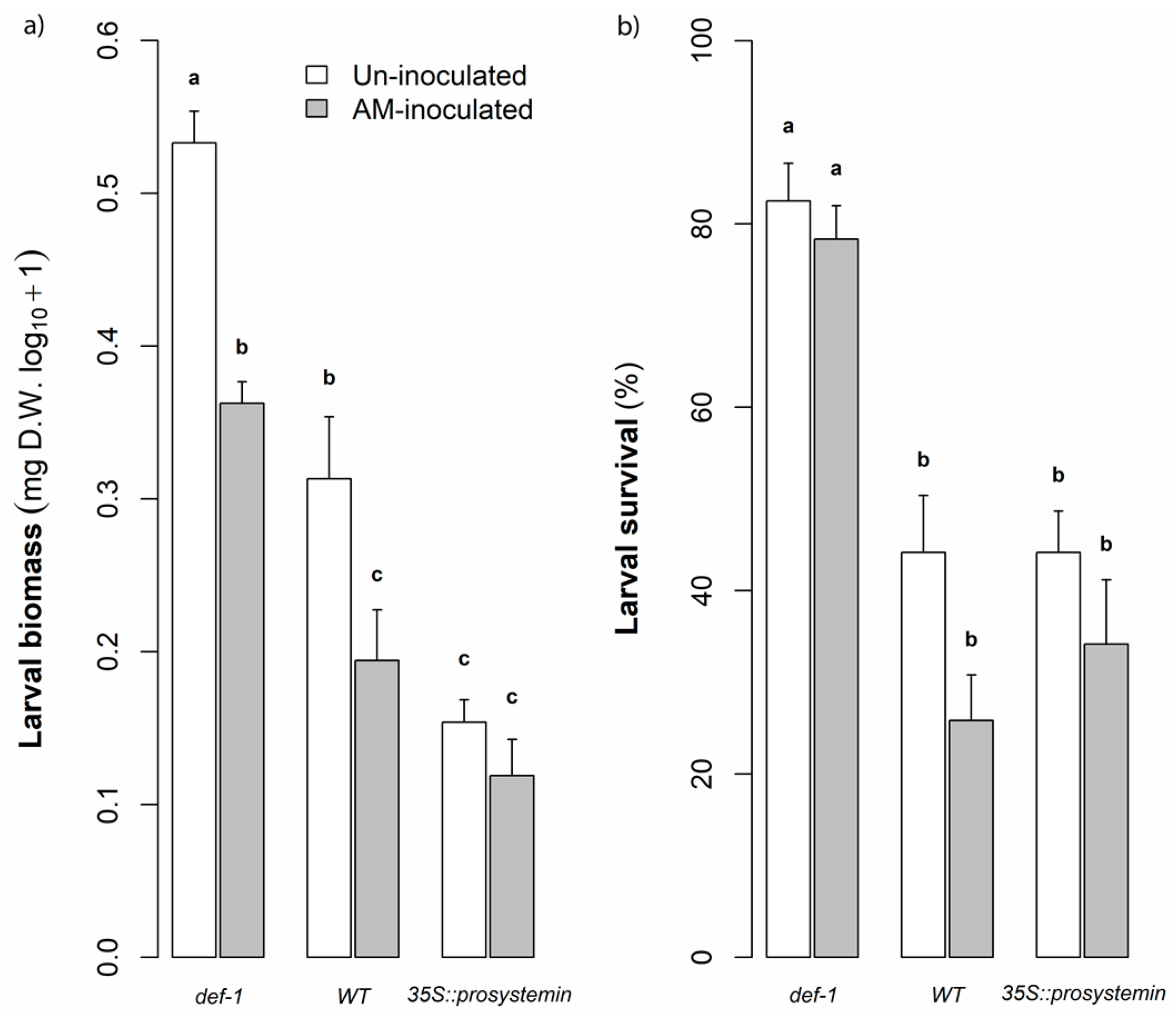
3.3. Defense and Resistance Traits
3.4. Multivariate Analyses
4. Discussion
4.1. Effect of JA on Plant Functional Traits
4.2. Effect of JA on AM Fungal Colonization
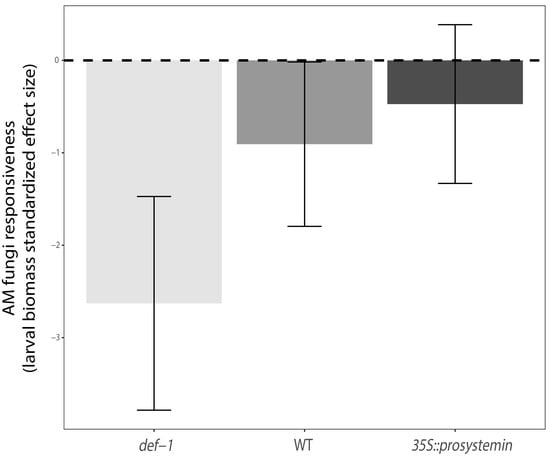
4.3. Interactive Effects of JA and AM Fungi on Plant Growth and Resistance
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Strong, D.R.; Lawton, J.H.; Southwood, R. Insects on Plants: Community Patterns and Mechanisms; Blackwell Scientific: London, UK, 1984. [Google Scholar]
- Schoonhoven, L.M.; van Loon, J.J.A.; Dicke, M. Insect-Plant Biology; Oxford University Press: Oxford, UK, 2005. [Google Scholar]
- Farmer, E.E. Leaf DefenceDefense; Oxford University Press: Oxford, UK, 2014. [Google Scholar]
- War, A.R.; Paulraj, M.G.; Ahmad, T.; Buhroo, A.A.; Hussain, B.; Ignacimuthu, S.; Sharma, H.C. Mechanisms of plant defense against insect herbivores. Plant Signal Behav. 2012, 7, 1306–1320. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Zamioudis, C.; Does, D.V.; Van Wees, S.C.M. Signalling Networks Involved in Induced Resistance. In Induced Resistance for Plant Defense; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2014; pp. 58–80. [Google Scholar] [CrossRef]
- Giron, D.; Frago, E.; Glevarec, G.; Pieterse, C.M.J.; Dicke, M. Cytokinins as key regulators in plant–microbe–insect interactions: Connecting plant growth and defencedefense. Funct. Ecol. 2013, 27, 599–609. [Google Scholar] [CrossRef]
- Robert-Seilaniantz, A.; Grant, M.; Jones, J.D.G. Hormone Crosstalk in Plant Disease and Defense: More Than Just JASMONATE-SALICYLATE Antagonism. Annu. Rev. Phytopathol. 2011, 49, 317–343. [Google Scholar] [CrossRef] [PubMed]
- Farmer, E.E.; Alméras, E.; Krishnamurthy, V. Jasmonates and related oxylipins in plant responses to pathogenesis and herbivory. Curr. Opin. Plant Biol. 2003, 6, 372–378. [Google Scholar] [CrossRef]
- Wasternack, C.; Stenzel, I.; Hause, B.; Hause, G.; Kutter, C.; Maucher, H.; Neumerkel, J.; Feussner, I.; Miersch, O. The wound response in tomato--role of jasmonic acid. J. Plant Physiol. 2006, 163, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Conrath, U.; Beckers, G.J.M.; Flors, V.; García-Agustín, P.; Jakab, G.; Mauch, F.; Newman, M.-A.; Pieterse, C.M.J.; Poinssot, B.; Pozo, M.J.; et al. Priming: Getting Ready for Battle. Mol. Plant-Microbe Interact. 2006, 19, 1062–1071. [Google Scholar] [CrossRef] [PubMed]
- Frost, C.J.; Mescher, M.C.; Carlson, J.E.; De Moraes, C.M. Plant Defense Priming against Herbivores: Getting Ready for a Different Battle. Plant Physiol. 2008, 146, 818–824. [Google Scholar] [CrossRef]
- Ton, J.; De Vos, M.; Robben, C.; Buchala, A.; Metraux, J.P.; Van Loon, L.C.; Pieterse, C.M.J. Characterization of Arabidopsis enhanced disease susceptibility mutants that are affected in systemically induced resistance. Plant J. 2002, 29, 11–21. [Google Scholar] [CrossRef]
- Pozo, M.J.; Azcon-Aguilar, C. Unraveling mycorrhiza-induced resistance. Curr. Opin. Plant Biol. 2007, 10, 393–398. [Google Scholar] [CrossRef]
- Song, Y.Y.; Ye, M.; Li, C.Y.; Wang, R.L.; Wei, X.C.; Luo, S.M.; Zeng, R.S. Priming of anti-herbivore defense in tomato by arbuscular mycorrhizal fungus and involvement of the jasmonate pathway. J. Chem. Ecol. 2013, 39, 1036–1044. [Google Scholar] [CrossRef]
- Song, Y.; Chen, D.; Lu, K.; Sun, Z.; Zeng, R. Enhanced tomato disease resistance primed by arbuscular mycorrhizal fungus. Front. Plant Sci. 2015, 6, 786. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.E.; Read, D.R. Mycorrhizal Symbiosis; Academic Press: Amsterdam, The Netherlands; New York, NY, USA; Boston, MA, USA, 2008. [Google Scholar]
- Cameron, D.D.; Neal, A.L.; van Wees, S.C.; Ton, J. Mycorrhiza-induced resistance: More than the sum of its parts? Trends Plant Sci. 2013, 18, 539–545. [Google Scholar] [CrossRef]
- Fritz, M.; Jakobsen, I.; Lyngkjaer, M.F.; Thordal-Christensen, H.; Pons-Kuhnemann, J. Arbuscular mycorrhiza reduces susceptibility of tomato to Alternaria solani. Mycorrhiza 2006, 16, 413–419. [Google Scholar] [CrossRef] [PubMed]
- Bennett, A.E.; Alers-Garcia, J.; Bever, J.D. Three-way interactions among mutualistic mycorrhizal fungi, plants, and plant enemies: Hypotheses and synthesis. Am. Nat. 2006, 167, 141–152. [Google Scholar] [PubMed]
- Gange, A.C.; West, H.M. Interactions between arbuscular mycorrhizal fungi and foliar-feeding insects in Plantago lanceolata L. New Phytol. 1994, 128, 79–87. [Google Scholar] [CrossRef]
- Koricheva, J.; Gange, A.C.; Jones, T. Effects of mycorrhizal fungi on insect herbivores: A meta-analysis. Ecology 2009, 90, 2088–2097. [Google Scholar] [CrossRef]
- Kempel, A.; Schmidt, A.K.; Brandl, R.; Schädler, M. Support from the underground: Induced plant resistance depends on arbuscular mycorrhizal fungi. Funct. Ecol. 2010, 24, 293–300. [Google Scholar] [CrossRef]
- Vannette, R.L.; Rasmann, S. Arbuscular mycorrhizal fungi mediate below-ground plant–herbivore interactions: A phylogenetic study. Funct. Ecol. 2012, 26, 1033–1042. [Google Scholar] [CrossRef]
- Vannette, R.L.; Hunter, M.D. Plant defencedefense theory re-examined: Nonlinear expectations based on the costs and benefits of resource mutualisms. J. Ecol. 2011, 99, 66–76. [Google Scholar] [CrossRef]
- Jung, S.C.; Martinez-Medina, A.; Lopez-Raez, J.A.; Pozo, M.J. Mycorrhiza-induced resistance and priming of plant defenses. J. Chem. Ecol. 2012, 38, 651–664. [Google Scholar] [CrossRef]
- Pozo, M.J.; Lopez-Raez, J.A.; Azcon-Aguilar, C.; Garcia-Garrido, J.M. Phytohormones as integrators of environmental signals in the regulation of mycorrhizal symbioses. New Phytol. 2015, 205, 1431–1436. [Google Scholar] [CrossRef]
- Tao, L.; Ahmad, A.; de Roode, J.C.; Hunter, M.D.; van der Heijden, M. Arbuscular mycorrhizal fungi affect plant tolerance and chemical defencedefenses to herbivory through different mechanisms. J. Ecol. 2016, 104, 561–571. [Google Scholar] [CrossRef]
- Vannette, R.; Hunter, M.; Rasmann, S. Arbuscular mycorrhizal fungi alter above- and below-ground chemical defense expression differentially among Asclepias species. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Herrera Medina, M.J.; Gagnon, H.; Piché, Y.; Ocampo, J.A.; García-Garrido, J.M.; Vierheilig, H. Root colonization by arbuscular mycorrhizal fungi is affected by the salicylic acid content of the plant. Plant Sci. 2003, 164, 993–998. [Google Scholar] [CrossRef]
- Vannette, R.L.; Hunter, M.D. Mycorrhizal fungi as mediators of defencedefense against insect pests in agricultural systems. Agric. For. Entomol. 2009, 11, 351–358. [Google Scholar] [CrossRef]
- Fernandez, I.; Merlos, M.; Lopez-Raez, J.A.; Martinez-Medina, A.; Ferrol, N.; Azcon, C.; Bonfante, P.; Flors, V.; Pozo, M.J. Defense related phytohormones regulation in arbuscular mycorrhizal symbioses depends on the partner genotypes. J. Chem. Ecol. 2014, 40, 791–803. [Google Scholar] [CrossRef]
- Howe, G.A.; Lightner, J.; Browse, J.; Ryan, C.A. An octadecanoid pathway mutant (JL5) of tomato is compromised in signaling for defense against insect attack. Plant Cell 1996, 8, 2067–2077. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Romera, B.; Calvo-Polanco, M.; Ruiz-Lozano, J.M.; Zamarreno, A.M.; Arbona, V.; Garcia-Mina, J.M.; Gomez-Cadenas, A.; Aroca, R. Involvement of the def-1 Mutation in the Response of Tomato Plants to Arbuscular Mycorrhizal Symbiosis Under Well-Watered and Drought Conditions. Plant Cell Physiol. 2018, 59, 248–261. [Google Scholar] [CrossRef]
- Li, C.; Williams, M.M.; Loh, Y.T.; Lee, G.I.; Howe, G.A. Resistance of cultivated tomato to cell content-feeding herbivores is regulated by the octadecanoid-signaling pathway. Plant Physiol. 2002, 130, 494–503. [Google Scholar] [CrossRef]
- Abouelsaad, I.; Renault, S. Enhanced oxidative stress in the jasmonic acid-deficient tomato mutant def-1 exposed to NaCl stress. J. Plant Physiol. 2018, 226, 136–144. [Google Scholar] [CrossRef]
- Thaler, J.S.; Owen, B.; Higgins, V.J. The role of the jasmonate response in plant susceptibility to diverse pathogens with a range of lifestyles. Plant Physiol. 2004, 135, 530–538. [Google Scholar] [CrossRef]
- Howe, G.A.; Ryan, C.A. Suppressors of Systemin Signaling Identify Genes in the Tomato Wound Response Pathway. Genetics 1999, 153, 1411–1421. [Google Scholar]
- Sun, J.-Q.; Jiang, H.-L.; Li, C.-Y. Systemin/Jasmonate-Mediated Systemic Defense Signaling in Tomato. Mol. Plant 2011, 4, 607–615. [Google Scholar] [CrossRef]
- McGurl, B.; Orozco-Cardenas, M.; Pearce, G.; Ryan, C.A. Overexpression of the prosystemin gene in transgenic tomato plants generates a systemic signal that constitutively induces proteinase inhibitor synthesis. Proc. Natl. Acad. Sci. USA 1994, 91, 9799–9802. [Google Scholar] [CrossRef] [PubMed]
- Coppola, M.; Corrado, G.; Coppola, V.; Cascone, P.; Martinelli, R.; Digilio, M.C.; Pennacchio, F.; Rao, R. Prosystemin Overexpression in Tomato Enhances Resistance to Different Biotic Stresses by Activating Genes of Multiple Signaling Pathways. Plant Mol. Biol. Report. Ispmb 2015, 33, 1270–1285. [Google Scholar] [CrossRef]
- Diaz, J.; ten Have, A.; van Kan, J.A. The role of ethylene and wound signaling in resistance of tomato to Botrytis cinerea. Plant Physiol. 2002, 129, 1341–1351. [Google Scholar] [CrossRef]
- Kaplan, I.; Thaler, J.S. Plant resistance attenuates the consumptive and non-consumptive impacts of predators on prey. Oikos 2010, 119, 1105–1113. [Google Scholar] [CrossRef]
- Thaler, J.S. Jasmonate-inducible plant defencedefenses cause increased parasitism of herbivores. Nature 1999, 399, 686. [Google Scholar] [CrossRef]
- Brown, E.S.; Dewhurst, C.F. The genus Spodoptera (Lepidoptera, Noctuidae) in Africa and the Near East. Bull. Entomol. Res. 1975, 65, 221–262. [Google Scholar] [CrossRef]
- Tian, D.; Tooker, J.; Peiffer, M.; Chung, S.H.; Felton, G.W. Role of trichomes in defense against herbivores: Comparison of herbivore response to woolly and hairless trichome mutants in tomato (Solanum lycopersicum). Planta 2012, 236, 1053–1066. [Google Scholar] [CrossRef]
- Díaz, S.; Kattge, J.; Cornelissen, J.H.C.; Wright, I.J.; Lavorel, S.; Dray, S.; Reu, B.; Kleyer, M.; Wirth, C.; Colin Prentice, I.; et al. The global spectrum of plant form and function. Nature 2016, 529, 167–171. [Google Scholar] [CrossRef] [PubMed]
- Wright, I.J.; Reich, P.B.; Westoby, M.; Ackerly, D.D.; Baruch, Z.; Bongers, F.; Cavender-Bares, J.; Chapin, T.; Cornelissen, J.H.C.; Diemer, M.; et al. The worldwide leaf economics spectrum. Nature 2004, 428, 821–827. [Google Scholar] [CrossRef] [PubMed]
- Escobar-Bravo, R.; Klinkhamer, P.G.L.; Leiss, K.A. Induction of Jasmonic Acid-Associated Defenses by Thrips Alters Host Suitability for Conspecifics and Correlates with Increased Trichome Densities in Tomato. Plant Cell Physiol. 2017, 58, 622–634. [Google Scholar] [CrossRef] [PubMed]
- Ataide, L.M.; Pappas, M.L.; Schimmel, B.C.; Lopez-Orenes, A.; Alba, J.M.; Duarte, M.V.; Pallini, A.; Schuurink, R.C.; Kant, M.R. Induced plant-defenses suppress herbivore reproduction but also constrain predation of their offspring. Plant Sci. 2016, 252, 300–310. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. R: A Language and Environement for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Torchiano, M. Effsize: Efficient Effect Size Computation, 0.7.4. 2018. Available online: http://github.com/mtorchiano/effsize/ (accessed on 21 December 2018).
- Nakagawa, S.; Cuthill, I.C. Effect size, confidence interval and statistical significance: A practical guide for biologists. Biol. Rev. 2007, 82, 591–605. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package, 2.0.10. 2013. Available online: http://vegan.r-forge.r-project.org/ (accessed on 4 January 2013).
- Agrawal, A.A.; Fishbein, M. Plant defense syndromes. Ecology 2006, 87, S132–S149. [Google Scholar] [CrossRef]
- Traw, M.B.; Bergelson, J. Interactive Effects of Jasmonic Acid, Salicylic Acid, and Gibberellin on Induction of Trichomes in Arabidopsis. Plant Physiol. 2003, 133, 1367–1375. [Google Scholar] [CrossRef]
- Yoshida, Y.; Sano, R.; Wada, T.; Takabayashi, J.; Okada, K. Jasmonic acid control of GLABRA3 links inducible defense and trichome patterning in Arabidopsis. Development 2009, 136, 1039–1048. [Google Scholar] [CrossRef]
- Li, L.; Zhao, Y.; McCaig, B.C.; Wingerd, B.A.; Wang, J.; Whalon, M.E.; Pichersky, E.; Howe, G.A. The Tomato Homolog of CORONATINE-INSENSITIVE1 Is Required for the Maternal Control of Seed Maturation, Jasmonate-Signaled Defense Responses, and Glandular Trichome Development. Plant Cell 2004, 16, 126–143. [Google Scholar] [CrossRef]
- Boughton, A.J.; Hoover, K.; Felton, G.W. Methyl Jasmonate Application Induces Increased Densities of Glandular Trichomes on Tomato, Lycopersicon esculentum. J. Chem. Ecol. 2005, 31, 2211–2216. [Google Scholar] [CrossRef]
- Fine, P.V.A.; Mesones, I.; Coley, P.D. Herbivores Promote Habitat Specialization by Trees in Amazonian Forests. Science 2004, 305, 663–665. [Google Scholar] [CrossRef] [PubMed]
- Thaler, J.S. Induced Resistance in Agricultural Crops: Effects of Jasmonic Acid on Herbivory and Yield in Tomato Plants. Environ. Entomol. 1999, 28, 30–37. [Google Scholar] [CrossRef]
- Redman, A.M.; Cipollini, D.F., Jr.; Schultz, J.C. Fitness costs of jasmonic acid-induced defense in tomato, Lycopersicon esculentum. Oecologia 2001, 126, 380–385. [Google Scholar] [CrossRef] [PubMed]
- Bubici, G.; Carluccio, A.V.; Stavolone, L.; Cillo, F. Prosystemin overexpression induces transcriptional modifications of defense-related and receptor-like kinase genes and reduces the susceptibility to Cucumber mosaic virus and its satellite RNAs in transgenic tomato plants. PLoS ONE 2017, 12, e0171902. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yu, P.; Zhao, J.; Jiang, H.; Wang, H.; Zhu, Y.; Botella, M.A.; Šamaj, J.; Li, C.; Lin, J. Expression of tomato prosystemin gene in Arabidopsis reveals systemic translocation of its mRNA and confers necrotrophic fungal resistance. New Phytol. 2018, 217, 799–812. [Google Scholar] [CrossRef] [PubMed]
- Tejeda-Sartorius, M.; Martinez de la Vega, O.; Delano-Frier, J.P. Jasmonic acid influences mycorrhizal colonization in tomato plants by modifying the expression of genes involved in carbohydrate partitioning. Plant Physiol. 2008, 133, 339–353. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Medina, M.J.; Tamayo, M.I.; Vierheilig, H.; Ocampo, J.A.; García-Garrido, J.M. The Jasmonic Acid Signalling Pathway Restricts the Development of the Arbuscular Mycorrhizal Association in Tomato. J. Plant Growth Regul. 2008, 27, 221–230. [Google Scholar] [CrossRef]
- Leon-Morcillo, R.J.; Angel, J.; Martin, R.; Vierheilig, H.; Ocampo, J.A.; Garcia-Garrido, J.M. Late activation of the 9-oxylipin pathway during arbuscular mycorrhiza formation in tomato and its regulation by jasmonate signalling. J. Exp. Bot. 2012, 63, 3545–3558. [Google Scholar] [CrossRef] [PubMed]
- Gutjahr, C.; Siegler, H.; Haga, K.; Iino, M.; Paszkowski, U. Full establishment of arbuscular mycorrhizal symbiosis in rice occurs independently of enzymatic jasmonate biosynthesis. PLoS ONE 2015, 10, e0123422. [Google Scholar] [CrossRef]
- Ludwig-Müller, J.; Bennett, R.N.; García-Garrido, J.M.; Piché, Y.; Vierheilig, H. Reduced arbuscular mycorrhizal root colonization in Tropaeolum majus and Carica papaya after jasmonic acid application can not be attributed to increased glucosinolate levels. J. Plant Physiol. 2002, 159, 517–523. [Google Scholar] [CrossRef]
- Regvar, M.; Gogala, N.; Zalar, P. Effects of jasmonic acid on mycorrhizal Allium sativum. New Phytol. 1996, 134, 703–707. [Google Scholar] [CrossRef]
- Isayenkov, S.; Mrosk, C.; Stenzel, I.; Strack, D.; Hause, B. Suppression of allene oxide cyclase in hairy roots of Medicago truncatula reduces jasmonate levels and the degree of mycorrhization with Glomus intraradices. Plant Physiol. 2005, 139, 1401–1410. [Google Scholar] [CrossRef] [PubMed]
- Ligaba, A.; Maron, L.; Shaff, J.; Kochian, L.; Piñeros, M. Maize ZmALMT2 is a root anion transporter that mediates constitutive root malate efflux. Plant Cell Environ. 2012, 35, 1185–1200. [Google Scholar] [CrossRef] [PubMed]
- Paszkowski, U. Mutualism and parasitism: The yin and yang of plant symbioses. Curr. Opin. Plant Biol. 2006, 9, 364–370. [Google Scholar] [CrossRef] [PubMed]
- Güimil, S.; Chang, H.-S.; Zhu, T.; Sesma, A.; Osbourn, A.; Roux, C.; Ioannidis, V.; Oakeley, E.J.; Docquier, M.; Descombes, P.; et al. Comparative transcriptomics of rice reveals an ancient pattern of response to microbial colonization. Proc. Natl. Acad. Sci. USA 2005, 102, 8066–8070. [Google Scholar] [CrossRef] [PubMed]
- Hause, B.; Mrosk, C.; Isayenkov, S.; Strack, D. Jasmonates in arbuscular mycorrhizal interactions. Phytochemistry 2007, 68, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Hause, B.; Schaarschmidt, S. The role of jasmonates in mutualistic symbioses between plants and soil-born microorganisms. Phytochemistry 2009, 70, 1589–1599. [Google Scholar] [CrossRef] [PubMed]
- López-Ráez, J.A.; Verhage, A.; Fernández, I.; García, J.M.; Azcón-Aguilar, C.; Flors, V.; Pozo, M.J. Hormonal and transcriptional profiles highlight common and differential host responses to arbuscular mycorrhizal fungi and the regulation of the oxylipin pathway. J. Exp. Bot. 2010, 61, 2589–2601. [Google Scholar] [CrossRef] [PubMed]
- Thaler, J.S.; Humphrey, P.T.; Whiteman, N.K. Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci. 2012, 17, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Werner, G.D.A.; Kiers, E.T. Partner selection in the mycorrhizal mutualism. New Phytol. 2015, 205, 1437–1442. [Google Scholar] [CrossRef] [PubMed]
- Kiers, E.T.; van der Heijden, M.G.A. Mutualistic Stability in the Arbuscular Mycorrhizal Symbiosis: Exploring Hypotheses of Evolutionary Cooperation. Ecology 2006, 87, 1627–1636. [Google Scholar] [CrossRef]
- Kiers, E.T.; Duhamel, M.; Beesetty, Y.; Mensah, J.A.; Franken, O.; Verbruggen, E.; Fellbaum, C.R.; Kowalchuk, G.A.; Hart, M.M.; Bago, A.; et al. Reciprocal Rewards Stabilize Cooperation in the Mycorrhizal Symbiosis. Science 2011, 333, 880–882. [Google Scholar] [CrossRef]
- Fellbaum, C.R.; Mensah, J.A.; Pfeffer, P.E.; Kiers, E.T.; Bücking, H. The role of carbon in fungal nutrient uptake and transport. Plant Signal. Behav. 2012, 7, 1509–1512. [Google Scholar] [CrossRef]
- Johnson, N.C.; Graham, J.-H.; Smith, F.A. Functioning of mycorrhizal associations along the mutualism–parasitism continuum. New Phytol. 1997, 135, 575–585. [Google Scholar] [CrossRef]
- Roger, A.; GÉTaz, M.; Rasmann, S.; Sanders, I.R. Identity and combinations of arbuscular mycorrhizal fungal isolates influence plant resistance and insect preference. Ecol. Entomol. 2013, 38, 330–338. [Google Scholar] [CrossRef]
- Bennett, A.E.; Bever, J.D. Mycorrhizal species differentially alter plant growth and response to herbivory. Ecology 2007, 88, 210–218. [Google Scholar] [CrossRef]
- Klironomos, J.N. Variation in plant response to native and exotic arbuscular mycorrhizal fungi. Ecology 2003, 84, 2292–2301. [Google Scholar] [CrossRef]
- Vannette, R.L.; Hunter, M.D.; van der Heijden, M. Mycorrhizal abundance affects the expression of plant resistance traits and herbivore performance. J. Ecol. 2013, 101, 1019–1029. [Google Scholar] [CrossRef]
- Mathur, S.; Sharma, M.P.; Jajoo, A. Improved photosynthetic efficacy of maize (Zea mays) plants with arbuscular mycorrhizal fungi (AMF) under high temperature stress. J. Photochem. Photobiol. B Biol. 2018, 180, 149–154. [Google Scholar] [CrossRef]
- Allen, M.F.; Smith, W.K.; Moore, T.S.; Christensen, M. Comparative Water Relations and Photosynthesis of Mycorrhizal and non-Mycorrhizal Bouteloua gracilis h.b.k. lag ex steud. New Phytol. 1981, 88, 683–693. [Google Scholar] [CrossRef]
- Reich, P.B.; Ellsworth, D.S.; Walters, M.B. Leaf structure (specific leaf area) modulates photosynthesis–nitrogen relations: Evidence from within and across species and functional groups. Funct. Ecol. 1998, 12, 948–958. [Google Scholar] [CrossRef]
- Reich, P.B.; Walters, M.B.; Ellsworth, D.S. From tropics to tundra: Global convergence in plant functioning. Proc. Natl. Acad. Sci. USA 1997, 94, 13730–13734. [Google Scholar] [CrossRef] [PubMed]
- Hartley, S.E.; Gange, A.C. Impacts of plant symbiotic fungi on insect herbivores: Mutualism in a multitrophic context. Annu. Rev. Entomol. 2009, 54, 323–342. [Google Scholar] [CrossRef] [PubMed]
- Pineda, A.; Zheng, S.J.; van Loon, J.J.; Pieterse, C.M.; Dicke, M. Helping plants to deal with insects: The role of beneficial soil-borne microbes. Trends Plant Sci. 2010, 15, 507–514. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Medina, A.; Fernandez, I.; Lok, G.B.; Pozo, M.J.; Pieterse, C.M.J.; Wees, S.C.M.V. Shifting from priming of salicylic acid- to jasmonic acid-regulated defencedefenses by Trichoderma protects tomato against the root knot nematode Meloidogyne incognita. New Phytol. 2017, 213, 1363–1377. [Google Scholar] [CrossRef] [PubMed]
- Cornelissen, J.H.C.; Lavorel, S.; Garnier, E.; Díaz, S.; Buchmann, N.; Gurvich, D.E.; Reich, P.B.; ter Steege, H.; Morgan, H.D.; van der Heijden, M.G.A.; et al. A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Aust. J. Bot. 2003, 51, 335–380. [Google Scholar] [CrossRef]
- Züst, T.; Rasmann, S.; Agrawal, A.A. Growth–defense tradeoffs for two major anti-herbivore traits of the common milkweed Asclepias syriaca. Oikos 2015, 124, 1404–1415. [Google Scholar] [CrossRef]
- Viani, R.A.G.; Rodrigues, R.R.; Dawson, T.E.; Lambers, H.; Oliveira, R.S. Soil pH accounts for differences in species distribution and leaf nutrient concentrations of Brazilian woodland savannah and seasonally dry forest species. Perspect. Plant Ecol. Evol. Syst. 2014, 16, 64–74. [Google Scholar] [CrossRef]
- Schädler, M.; Jung, G.; Auge, H.; Brandl, R. Palatability, decomposition and insect herbivory: Patterns in a successional old-field plant community. Oikos 2003, 103, 121–132. [Google Scholar] [CrossRef]
- Estrada-Luna, A.A.; Davies, F.T., Jr.; Egilla, J.N. Mycorrhizal fungi enhancement of growth and gas exchange of micropropagated guava plantlets (Psidium guajava L.) during ex vitro acclimatization and plant establishment. Mycorrhiza 2000, 10, 1–8. [Google Scholar] [CrossRef]
- Gross, N.; Le Bagousse-Pinguet, Y.; Liancourt, P.; Urcelay, C.; Catherine, R.; Lavorel, S. Trait-mediated effect of arbuscular mycorrhiza on the competitive effect and response of a monopolistic species. Funct. Ecol. 2010, 24, 1122–1132. [Google Scholar] [CrossRef]
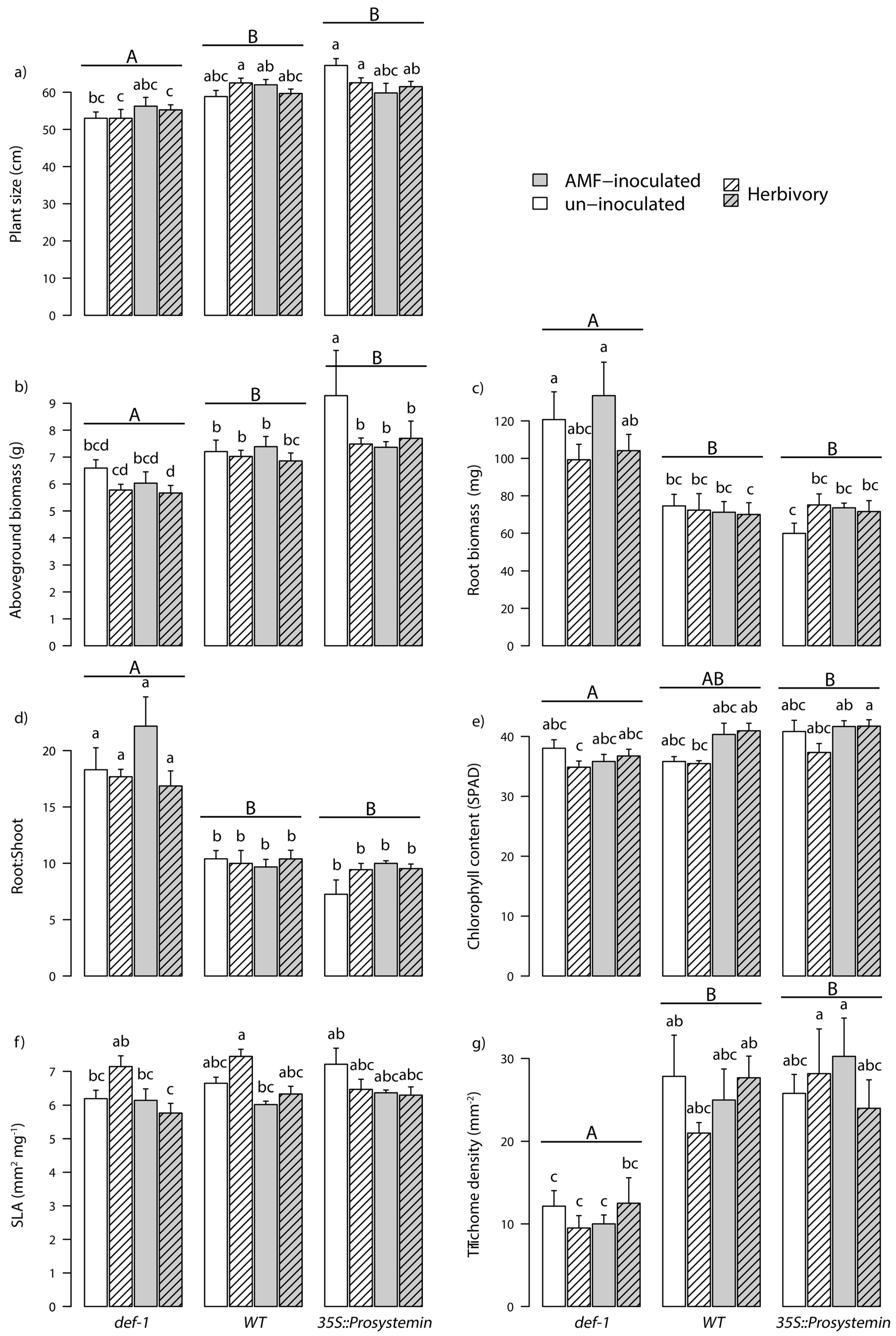
| Response Variable | Factor | d.f. | Fx,y | p |
|---|---|---|---|---|
| Plant size (cm) | Genotype (G) | 2 | 23.33 | <0.001 |
| AM fungi (M) | 1 | 0.14 | 0.71 | |
| Herbivore (H) | 1 | 0.29 | 0.59 | |
| G × M | 2 | 4.17 | <0.05 | |
| G × H | 2 | 0.10 | 0.91 | |
| M × H | 1 | 0.04 | 0.84 | |
| G × M × H | 2 | 2.75 | 0.07 | |
| Residuals (R) | 53 | |||
| AG biomass (g) | G | 2 | 10.7 | <0.001 |
| M | 1 | 1.85 | 0.18 | |
| H | 1 | 1.41 | 0.24 | |
| G × M | 2 | 1.71 | 0.19 | |
| G × H | 2 | 0.05 | 0.95 | |
| M × H | 1 | 0.72 | 0.40 | |
| G × M × H | 2 | 0.94 | 0.40 | |
| R | 53 | |||
| Root biomass (mg) | G | 2 | 30.86 | <0.001 |
| M | 1 | 0.03 | 0.88 | |
| H | 1 | 1.05 | 0.31 | |
| G × M | 2 | 0.22 | 0.83 | |
| G × H | 2 | 3.99 | <0.05 | |
| M × H | 1 | 0.66 | 0.42 | |
| G × M × H | 2 | 0.37 | 0.69 | |
| R | 53 | |||
| Root:shoot | G | 2 | 80.00 | <0.001 |
| M | 1 | 0.65 | 0.42 | |
| H | 1 | 0.33 | 0.57 | |
| G × M | 2 | 0.42 | 0.66 | |
| G × H | 2 | 2.56 | 0.09 | |
| M × H | 1 | 2.18 | 0.15 | |
| G × M × H | 2 | 1.83 | 0.17 | |
| R | 53 | |||
| Chlorophyll content | G | 2 | 4.81 | <0.05 |
| M | 1 | 5.93 | <0.05 | |
| H | 1 | 2.32 | 0.13 | |
| G × M | 2 | 1.62 | 0.21 | |
| G × H | 2 | 0.09 | 0.91 | |
| M × H | 1 | 0.75 | 0.39 | |
| G × M × H | 2 | 1.36 | 0.27 | |
| R | 53 | |||
| SLA (mm2/mg) | G | 2 | 1.26 | 0.29 |
| M | 1 | 19.55 | <0.001 | |
| H | 1 | 1.58 | 0.21 | |
| G × M | 2 | 0.76 | 0.47 | |
| G × H | 2 | 4.55 | <0.05 | |
| M × H | 1 | 0.94 | 0.34 | |
| G × M × H | 2 | 3.74 | <0.05 | |
| R | 53 | |||
| Trichome density | G | 2 | 23.44 | <0.001 |
| M | 1 | 0.11 | 0.74 | |
| H | 1 | 0.63 | 0.43 | |
| G × M | 2 | 0.12 | 0.88 | |
| G × H | 2 | 0.03 | 0.97 | |
| M × H | 1 | 0.38 | 0.54 | |
| G × M × H | 2 | 1.88 | 0.16 | |
| R | 53 | |||
| AMF colonization | G | 2 | 5.96 | <0.01 |
| H | 1 | 0.12 | 0.73 | |
| G × H | 2 | 0.36 | 0.70 | |
| R | 27 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Formenti, L.; Rasmann, S. Mycorrhizal Fungi Enhance Resistance to Herbivores in Tomato Plants with Reduced Jasmonic Acid Production. Agronomy 2019, 9, 131. https://doi.org/10.3390/agronomy9030131
Formenti L, Rasmann S. Mycorrhizal Fungi Enhance Resistance to Herbivores in Tomato Plants with Reduced Jasmonic Acid Production. Agronomy. 2019; 9(3):131. https://doi.org/10.3390/agronomy9030131
Chicago/Turabian StyleFormenti, Ludovico, and Sergio Rasmann. 2019. "Mycorrhizal Fungi Enhance Resistance to Herbivores in Tomato Plants with Reduced Jasmonic Acid Production" Agronomy 9, no. 3: 131. https://doi.org/10.3390/agronomy9030131
APA StyleFormenti, L., & Rasmann, S. (2019). Mycorrhizal Fungi Enhance Resistance to Herbivores in Tomato Plants with Reduced Jasmonic Acid Production. Agronomy, 9(3), 131. https://doi.org/10.3390/agronomy9030131