Genotype-Environment Interaction: Trade-Offs between the Agronomic Performance and Stability of Dual-Purpose Sorghum (Sorghum bicolor L. Moench) Genotypes in Senegal
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sites
2.2. Plant Material
2.3. Trial Management
2.4. Experimental Design and Data Collection
2.5. Data Analysis
3. Results
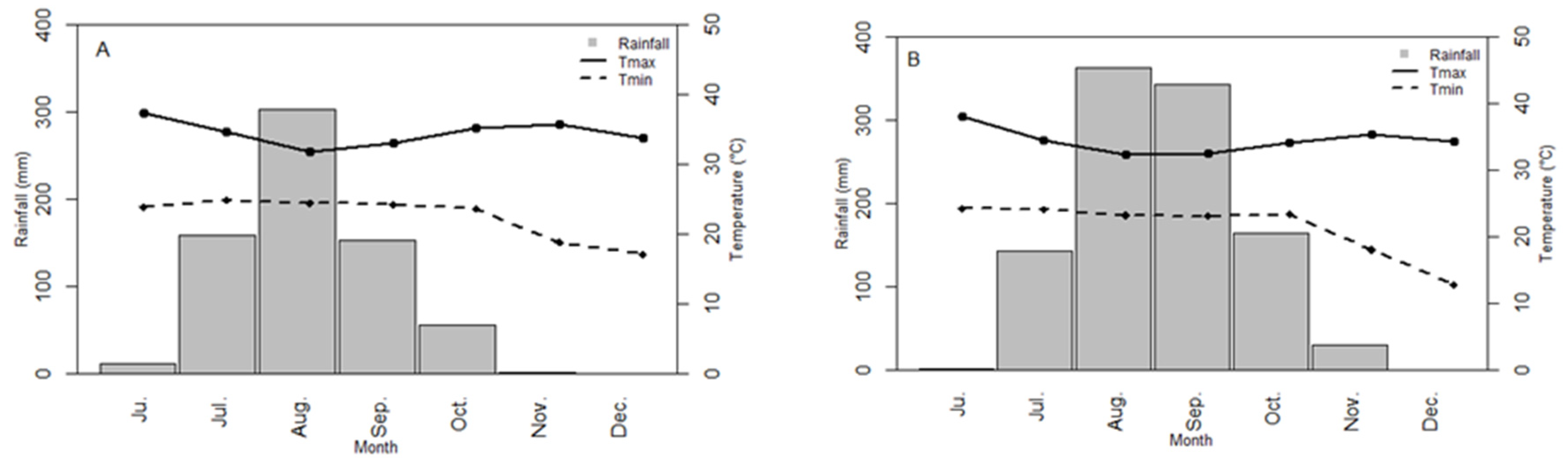
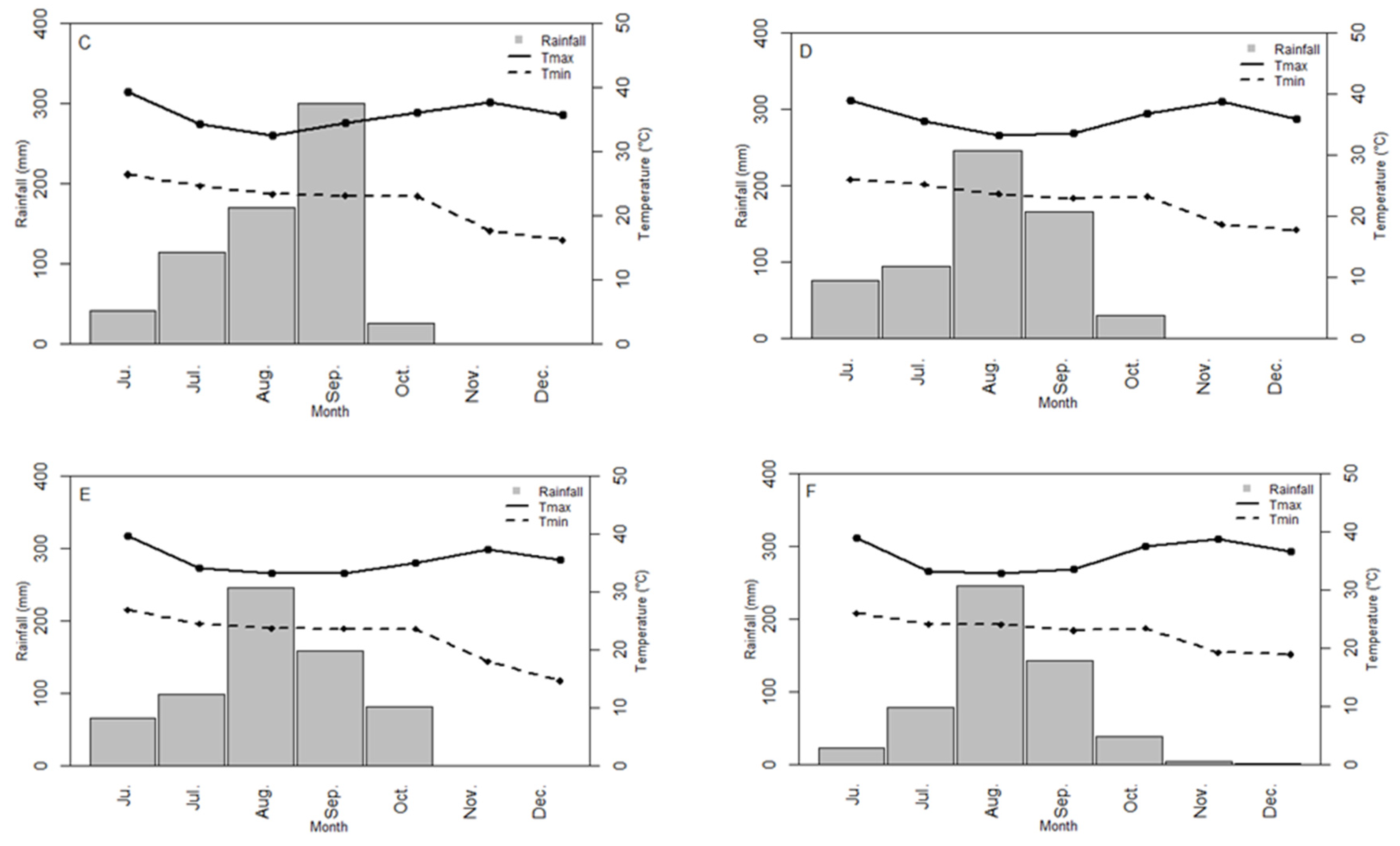
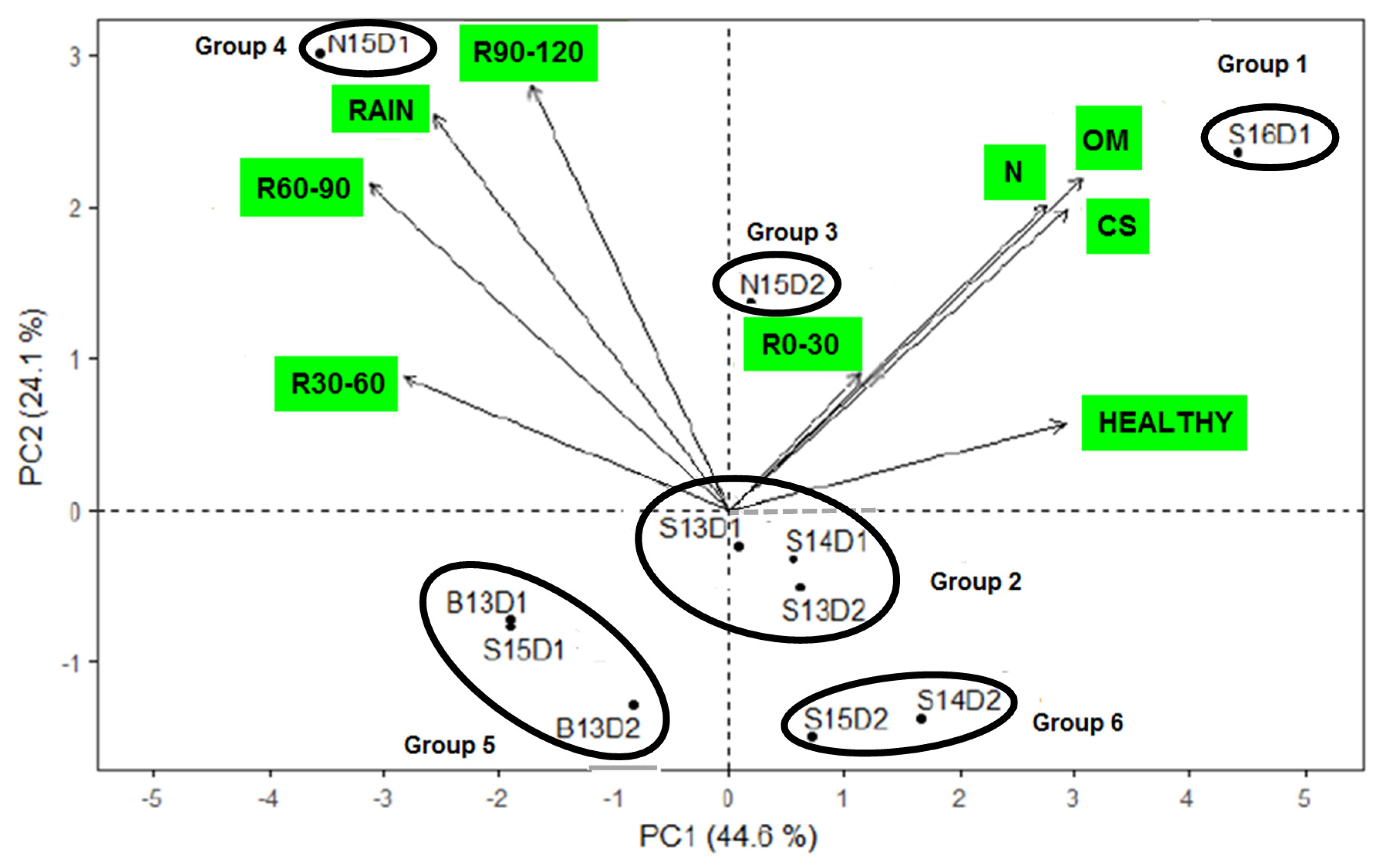
3.1. Environment Characterization
3.2. Effects of Genotypes, Environments and Genotype × Environment Interactions
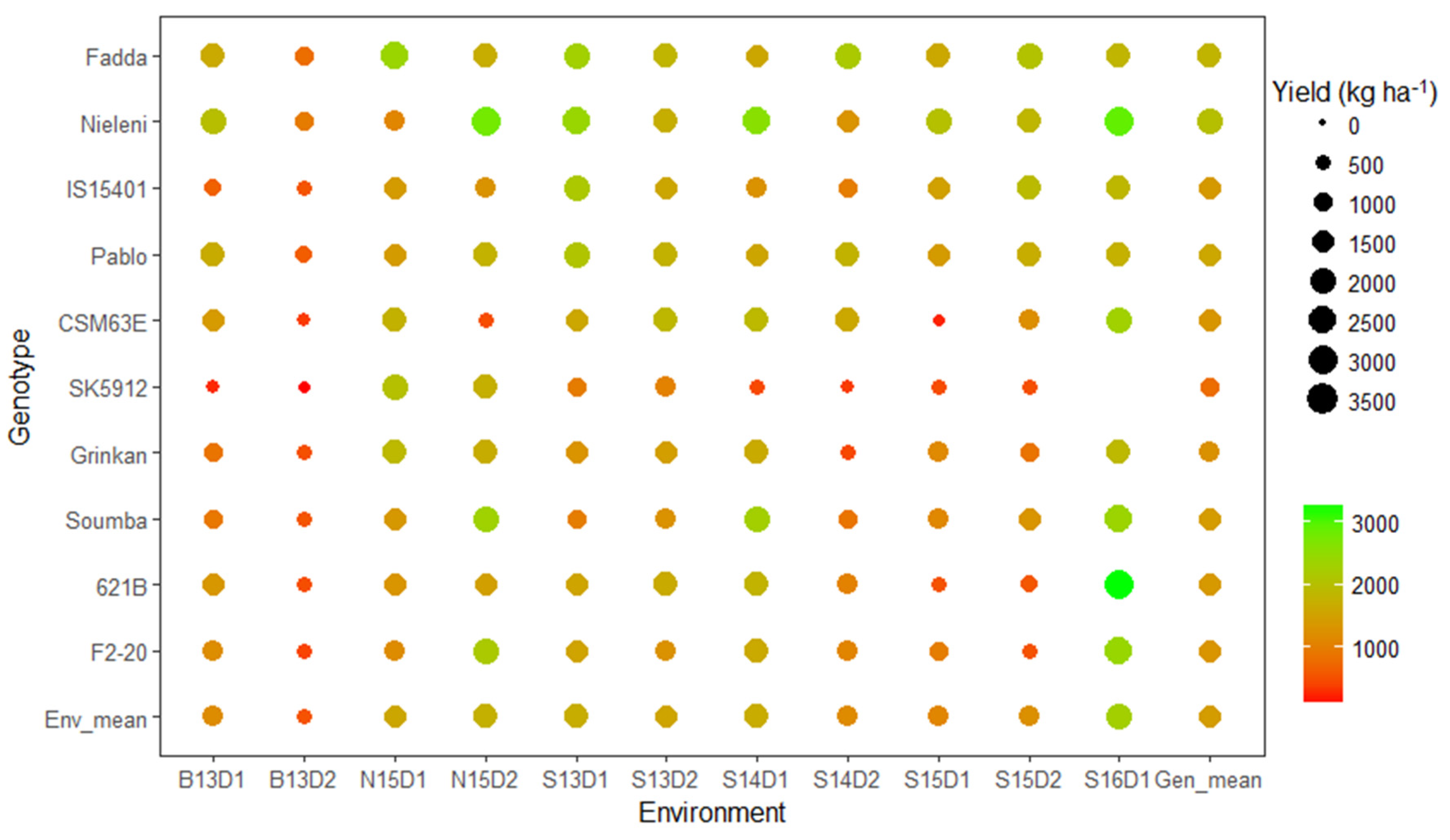
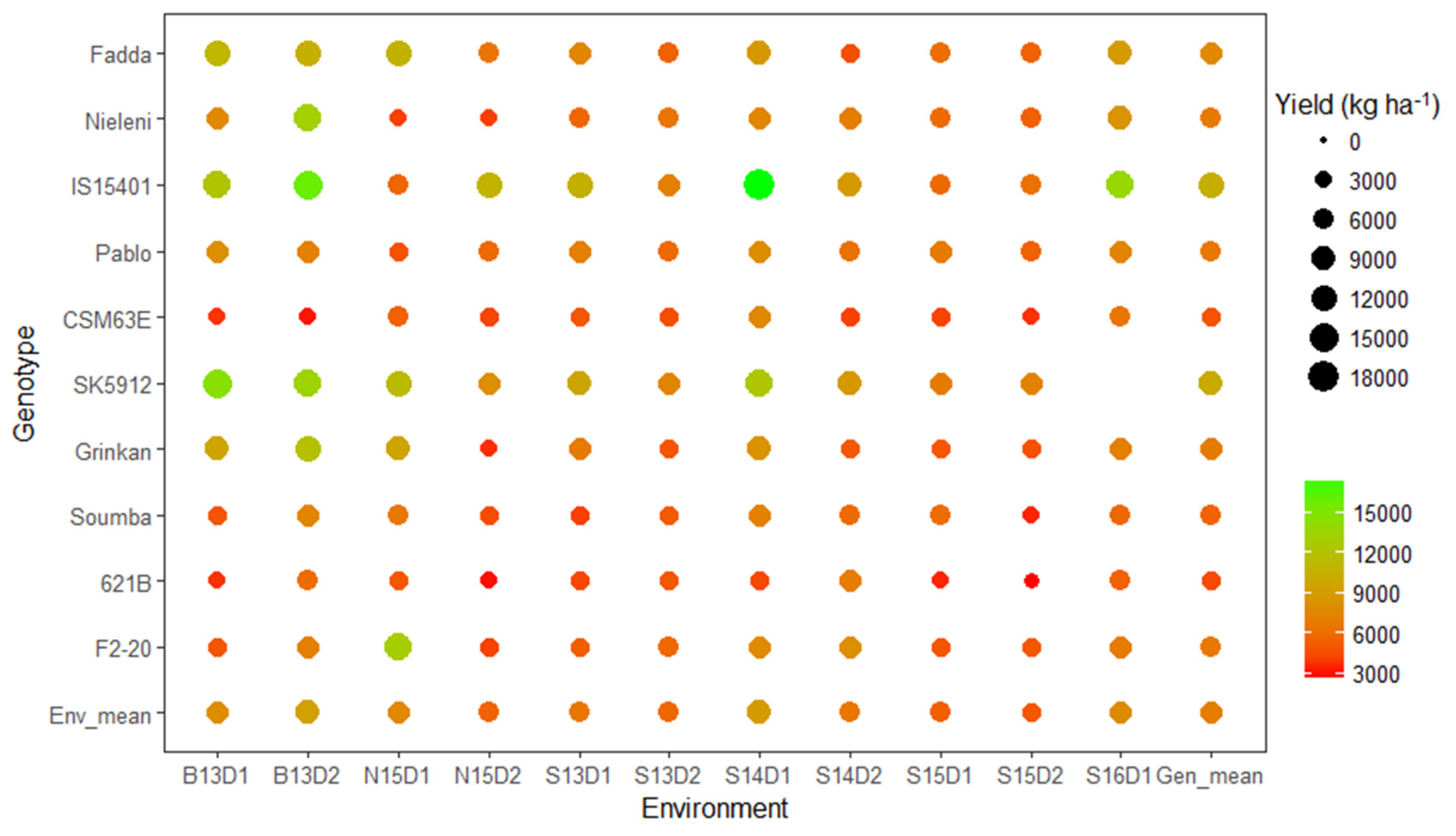
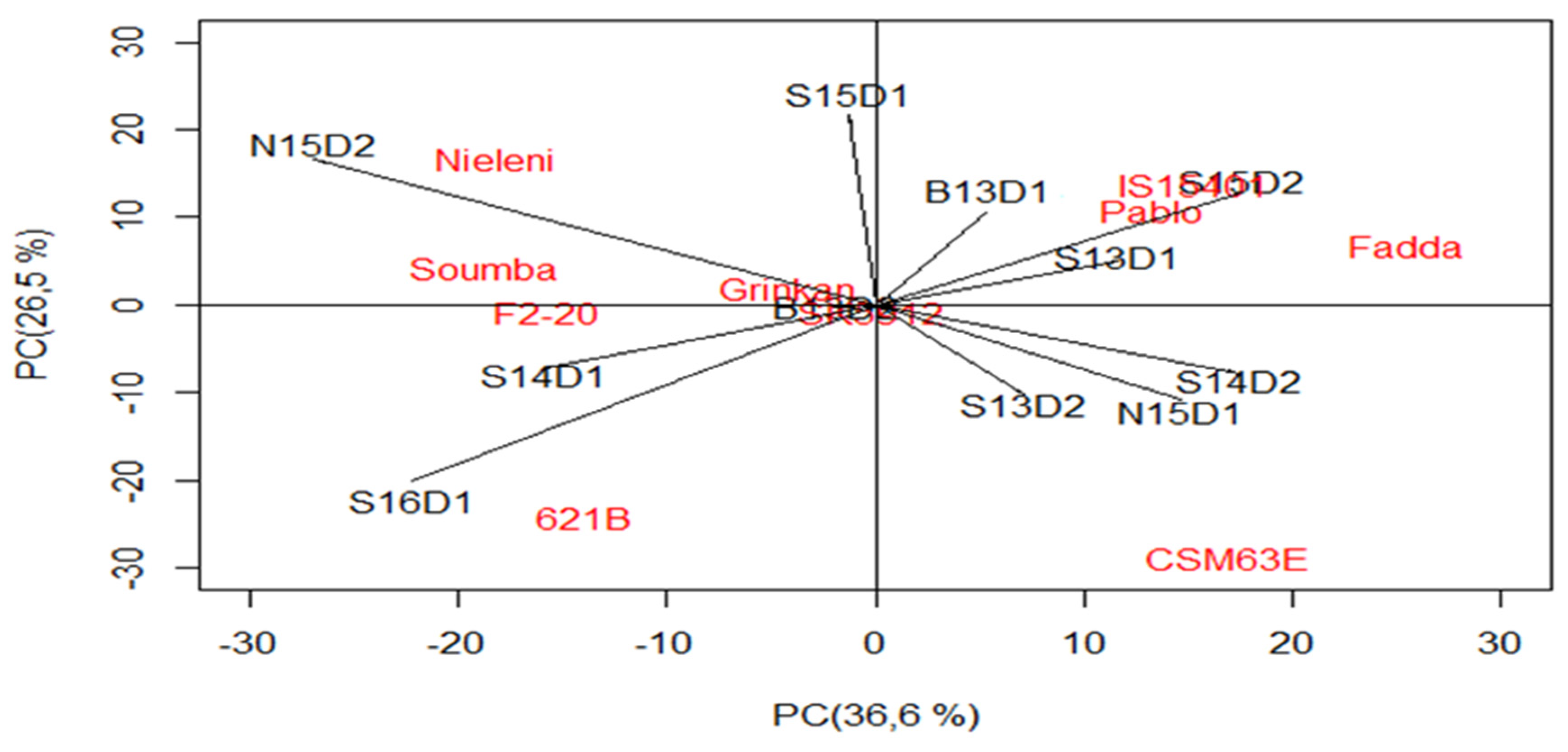
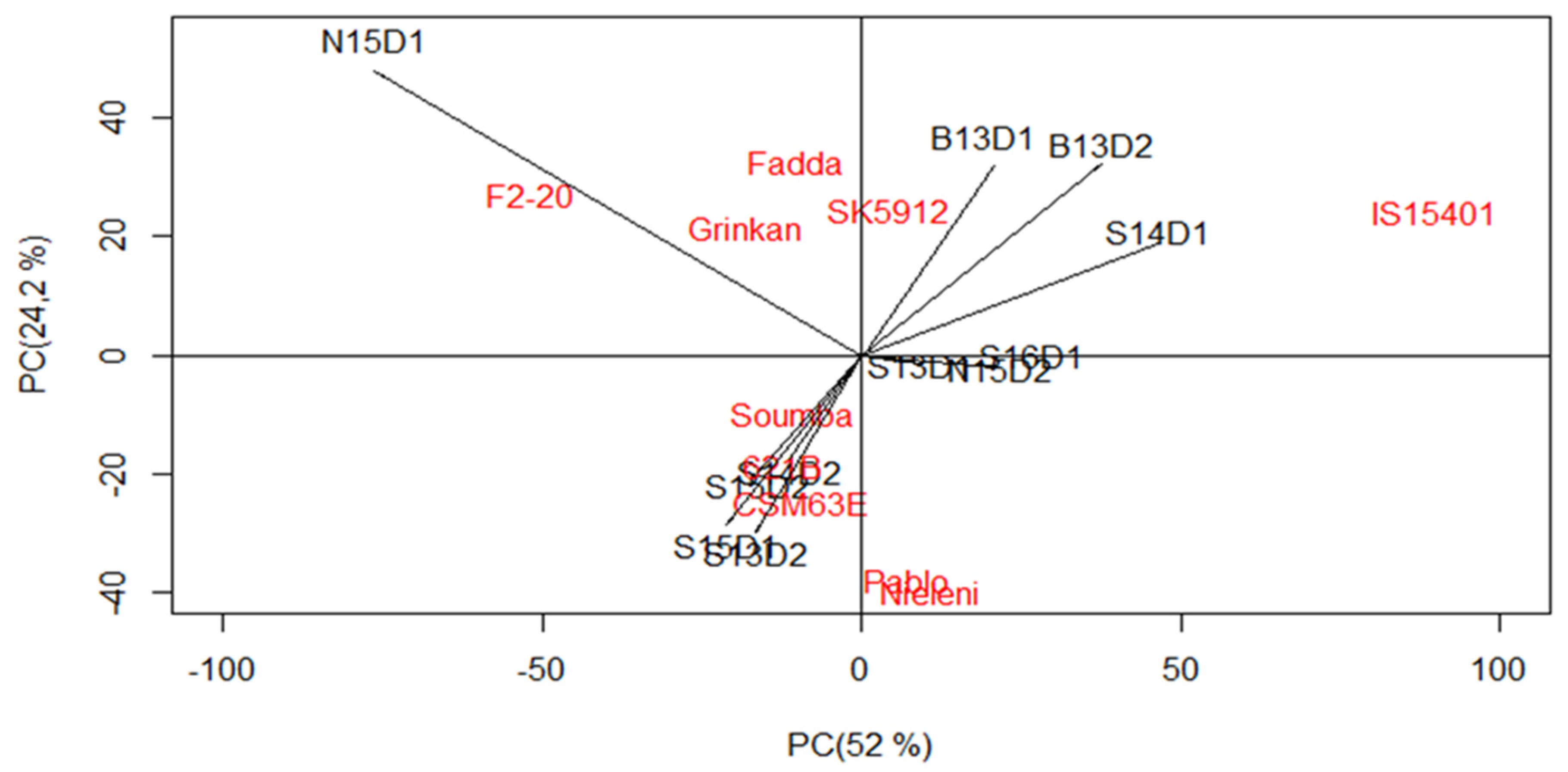
3.3. Which Genotype(s) for Which Environment(s)?
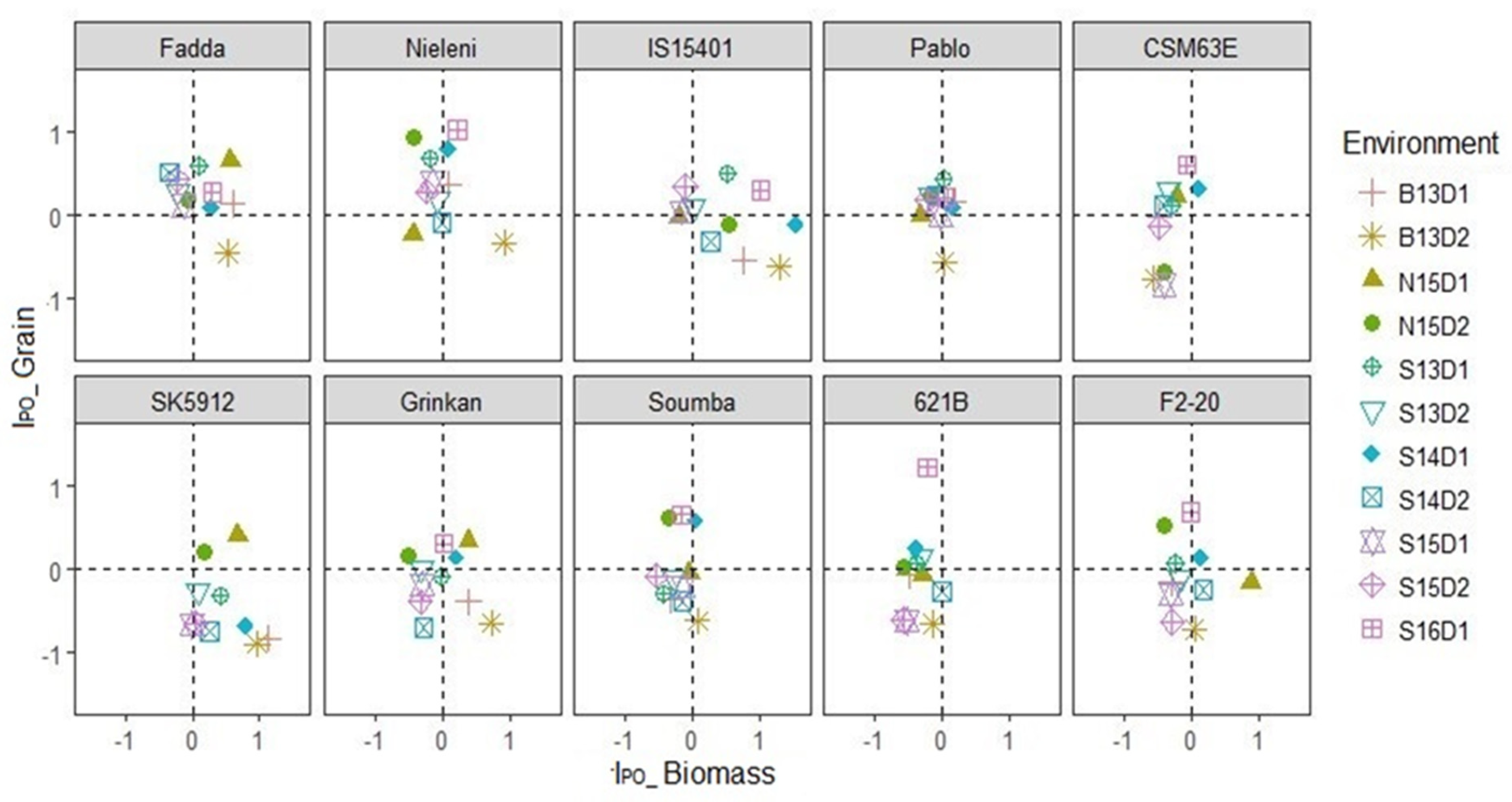
3.4. Which Genotype(s) Showed Dual–Purpose Potential?
4. Discussion
4.1. Which Genotype(s) for Which Environment(s)?
4.2. Choice of Genotypes with Dual–Purpose Potential
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Djè, Y.; Heuertz, M.; Ater, M.; Lefebvre, C.; Vekemans, X. Évaluation de la diversité morphologique des variétés traditionnelles de sorgho du Nord–ouest du Maroc. Biotechnol. Agron. Soc. Environ. 2007, 11, 30–40. [Google Scholar]
- FAOSTAT 2015. Statistiques des Données Année 2015; United Nations Food and Agriculture Organization: Rome, Italy, 2015. [Google Scholar]
- Agrama, H.A.; Tuinstra, M.R. La diversité phylogénétique et les relations entre le sorgho adhésions à l’aide SSR et RAPD. Afr. J. Biotechnol. 2003, 2, 334–340. [Google Scholar] [CrossRef]
- Reddy, B.V.S.; Kumar, A.A.; Reddy, P.S.; Elangovan, M. Sorghum germplasm: Diversity and utilization. In Sorghum Genetic Enhancement: Research Process, Dissemination and Impacts; International Crops Research Institute for the Semi-Arid Tropics: Patancheru, Andhra Pradesh, India, 2008; pp. 153–169. [Google Scholar]
- ANSD (Agence Nationale de la Statistique et de la Démographie). Bulletin Mensuel des Statistiques Economiques de 2018; Division des Statistiques Economiques Ministère de l’Economie, des Finances et du Plan: Dakar, Sénégal, 2018; p. 109. [Google Scholar]
- Ba, K.; Tine, E.; Destain, J.; Cissé, N.; Thonart, P. Étude comparative des composés phénoliques, du pouvoir antioxydant de différentes variétés de sorgho sénégalais et des enzymes amylolytiques de leur malt. Biotechnol. Agron. Soc. Environ. 2010, 14, 131–139. [Google Scholar]
- Seguin, B.; Soussana, J.F. Emissions de Gaz à Effet de Serre et Changement Climatique: Causes et Conséquences Observées Pour L’agriculture et L’élevage. Courrierde L’environnement de l’INRA 2008, 55, 79–91. [Google Scholar]
- Cruz, C.D.; Regazzi, A.J. Biometrical Models Applied to Plant Breeding; Editora UFV: Viçosa, Brazil, 1997; p. 390. [Google Scholar]
- Holhs, T. Analysis of genotype environment interactions. S. Afr. J. Sci. 1995, 91, 121–124. [Google Scholar]
- Crossa, J. Statistical analyses of multilocation trials. Adv. Agron. 1990, 44, 55–85. [Google Scholar]
- Romagosa, I.; Fox, P.N. Genotype x environment interaction and adaptation. In Plant Breeding; Springer: Dordrecht, The Netherlands, 1993; pp. 373–390. ISBN 9401046654. [Google Scholar]
- Zobel, R.W.; Wright, M.W.; Gauch, H.G. Statistical analysis of a yield trial. Agron. J. 1988, 80, 388–393. [Google Scholar] [CrossRef]
- Crossa, J.; Gauch, H.G.; Zobel, R.W. Additive main effect and multiplicative interaction analysis of two international maize cultivar trials. Crop. Sci. 1990, 30, 493–500. [Google Scholar] [CrossRef]
- Gauch, H.G.; Zobel, R.W. AMMI analysis of yield trials. In Genotype by Environment Interaction; Kang, M.S., Gauch, H.G., Eds.; CRC Press: Boca Raton, FL, USA, 1996; pp. 85–122. [Google Scholar]
- Kempton, R.A. The use of biplots in interpreting variety by environment interactions. J. Agric. Sci. 1984, 103, 123–135. [Google Scholar] [CrossRef]
- Ferreira, D.F.; Demétrio, C.G.B.; Manly, B.F.J.; Machado, A.A.; Vencovsky, R. Statistical models in agriculture: Biometrical methods for evaluating phenotypic stability in plant breeding. Cerne 2006, 12, 373–388. [Google Scholar]
- Vachaud, G.; Dancette, C.; Sonko, S.; Thony, J.L. Méthode de caractérisation hydrodynamique d’un sol non saturé: Application à deux types de sols du Sénégal. Ann. Agron. 1978, 29, 1–36. [Google Scholar]
- Hamon, G. Mise en Œuvre et Critique des Méthodes de Caractérisation Hydrodynamique de la Zone non Saturée du sol. Application Aux Sols de Culture du Sénégal. Ph.D. Thesis, Instut de Mécanique, Grenoble, France, 1980. [Google Scholar]
- Imberno, J. Variabilité Spatiale des Caractéristiques Hydrodynamiques d’un sol du Sénégal. Application au Calcul d’un Bilan Sous Culture. Ph.D. Thesis, USMG et Institut National Polytechnique de Grenoble, Grenoble, France, 1981. [Google Scholar]
- Hamon, G. Caractérisation Hydrodynamique in situ d’un sol de Culture en Moyenne Casamance; Institut Sénégalais de Recherches Agricoles: Djibélor, Sénégal, 1978. [Google Scholar]
- Baret, F. Caractérisation Hydrodynamique d’un Sol de la Région de Nioro; Institut Sénégalais de Recherches Agricoles (ISRA): Dakar, Sénégal, 1980. [Google Scholar]
- Valet, S. Bilan Hydrique sous Cultures Dans les Essais Travail du Sol en Sols Sableux (Nioro) et en Sols Sablo-Argileux (Thysse); Institut Sénégalais de Recherches Agricoles: Kaolack, Sénégal, 1984; p. 16. [Google Scholar]
- Bartlett, M.S. Properties of sufficiency and statistical tests. Proc. R. Stat. Soc. Ser. A 1937, 160, 268–282. [Google Scholar]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Almeida Filho, J.E.; Tardin, F.D.; Daher, R.F.; Barbé, T.C.; Paula, C.M.; Cardoso, M.J.; Godinho, V.P.C. Stability and adaptability of grain sorghum hybrids in the off-season. Genet. Mol. Res. 2014, 13, 7626–7635. [Google Scholar] [CrossRef] [PubMed]
- Ndiaye, M.; Adam, M.; Muller, B.; Guisse, A.; Cissé, N. Performances agronomiques et stabilité phénotypique de génotypes de Sorgho (Sorghum bicolor (L.) Moench) au Sénégal: Une étude des interactions génotypes-environnement. Appl. Biosci. 2018, 125, 12617–12629. [Google Scholar]
- Ganyo, K.K.; Muller, B.; Gaglo, E.K.; Guisse, A.; Cissé, N. Optimisation du NPK et urée basée sur les informations climatiques pour accroitre la production du sorgho en zones soudano-sahéliennes du Sénégal. Appl. Biosci. 2018, 131, 13293–13307. [Google Scholar] [CrossRef]
- Showemimo, F.A.; Echekwu, C.A.; Yeye, M.Y. Genotype x environment interaction in Sorghum trials and their implication for future variety evaluation in Sorghum growing areas of northern Nigeria. Plant Sci. 2000, 1, 24–31. [Google Scholar]
- Kouressy, M.; Dingkuhn, M.; Vaksmann, M.; Heinemann, A.B. Adaptation to diverse semi–arid environments of sorghum genotypes having different plant type and sensitivity to photoperiod. Agric. For. Meteorol. 2008, 148, 357–371. [Google Scholar] [CrossRef]
- Bazile, D.; Dembélé, S.; Soumaré, M.; Dembele, D. Utilisation de la diversité variétale du sorgho pour valoriser la diversité des sols au Mali. Cah. Agric. 2008, 17, 86–94. [Google Scholar] [CrossRef]
- Raton, F.L.; Gauch, H.G. Statistical Analysis of Regional Yield Trials. AMMI Analysis of Factorial Designs; Elsevier: New York, NY, USA, 1992. [Google Scholar]
- Raju, B.M.K. Study of AMMI model and its biplots. J. Ind. Soc. Agric. Stat. 2002, 55, 297–322. [Google Scholar]
- Zali, H.; Farshadfar, E.; Sabaghpour, S.H.; Karimizadeh, R. Evaluation of genotype × environment interaction in chickpea using measures of stability from AMMI model. Ann. Biol. Res. 2012, 3, 3126–3136. [Google Scholar]
- Silva Filho, J.L.; Morello, C.L.; Farias, F.J.C.; Lamas, F.M. Comparação de métodos para avaliar a adaptabilidade eestabilidade produtiva em algodoeiro. Pesqui. Agropecuária Bras. 2008, 43, 349–355. [Google Scholar] [CrossRef]
- Menad, A.; Meziani, N.; Bouzerzour, H.; Benmahammad, A. Analyse de l’interaction génotype x milieux du rendement de l’orge (Hordeum vulgare L.): Application des modèles AMMI et la régression conjointe. Nat. Biotechnol. 2010, 5, 99–106. [Google Scholar]
- Yan, W.; Hunt, L.A. Biplot analysis of multi-environment trial data. In Kang MS: Quantitative Genetics, Genomics and Plant Breeding; Louisiana State University: Baton Rouge, LA, USA, 2002; pp. 289–304. [Google Scholar]
- Ganyo, K.K. Etude et Modélisation des Réponses de Variétés de Sorgho (Sorghum bicolor (L.) Moench) à des Stratégies Contrastées D’apports D’intrants. Ph.D. Thesis, Université Cheikh Anta Diop, Dakar, Senegal, 2019. [Google Scholar]
- Rattunde, H.F.W.; Weltzien, E.; Diallo, B.; Diallo, A.G.; Sidibe, M.; Touré, A.O.; Rathore, A.; Das, R.R.; Leiser, W.L.; Touré, A. Yield of photoperiod–sensitive sorghum hybrids based on guinea–race germplasm under farmers’ field conditions in Mali. Crop. Sci. 2013, 53, 1–8. [Google Scholar] [CrossRef]
- Voigt, J.; Botha, P.R.; Gerber, H.S. The effect of planting date on the dry matter production of annual forage sorghum hybrids and hybrid millet cultivars. Grassroots Newsl. Grassl. Soc. S. Afr. 2008, 8, 18–24. [Google Scholar]
- Erickson, J.E.; Helsel, Z.R.; Woodard, K.R.; Vendramini, J.M.B.; Wang, Y.; Sollenberger, L.E.; Gilbert, R.A. Planting date affects biomass and Brix of sweet sorghum grown for biofuel across Florida. Agron. J. 2011, 103, 1827–1833. [Google Scholar] [CrossRef]
- Reddi, S.G.; Janawade, A.D.; Palled, Y.B. Influence of sowing dates on growth, grain and ethanol yield and economics. Int. J. Agric. Sci. Vet. Med. 2013, 1, 12–17. [Google Scholar]
- Gutjahr, S.; Vaksmann, M.; Dingkuhn, M.; Thera, K.; Trouche, G.; Braconnier, S.; Luquet, D. Grain, sugar and biomass accumulation in tropical sorghums. I. Trade-offs and effects of phenological plasticity. Funct. Plant Biol. 2013, 40, 342–354. [Google Scholar]
| Environment | Zone | Code | Coordinates | Alt (m) | Soil Type * | SAN (%) | CS (%) | N (%) | OM (%) | Rain (mm) | R0–30 (mm) | R30–60 (mm) | R60–90 (mm) | R90–120 (mm) | Tmin (°C) | Tmax (°C) | Healthy ** | Sowing Date | Previo-uscrop |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sowing 1/2013 | BBY | B13D1 | 14°42′N 16°29′W | 20 | Sandy | 94.2 | 6.6 | 0.15 | 3.1 | 644 | 180 | 352 | 110 | 3 | 23 | 33.9 | 2 | 07/17/2013 | Fallow |
| Sowing 2/2013 | BBY | B13D2 | Sandy | 94.2 | 6.6 | 0.15 | 3.1 | 566 | 253 | 256 | 56 | 1 | 22.8 | 33.9 | 3 | 07/31/2013 | Fallow | ||
| Sowing 1/2013 | SIN | S13D1 | 13°49′N 13°55′W | 23 | Sandy-silty | 89.4 | 11.6 | 0.21 | 4.3 | 575 | 146 | 365 | 59 | 6 | 21.4 | 35.3 | 5 | 07/25/2013 | Fallow |
| Sowing 2/2013 | SIN | S13D2 | Sandy-silty | 89.4 | 11.6 | 0.21 | 4.3 | 536 | 183 | 306 | 46 | 1 | 21.2 | 35.4 | 5 | 08/06/2013 | Fallow | ||
| Sowing 1/2014 | SIN | S14D1 | Sandy | 91.2 | 10.2 | 0.17 | 5.7 | 488 | 158 | 213 | 88 | 31 | 22.2 | 35.7 | 4 | 07/17/2014 | Peanut | ||
| Sowing 2/2014 | SIN | S14D2 | Sandy | 90.9 | 9.7 | 0.17 | 4.5 | 377 | 190 | 156 | 31 | 1 | 22.1 | 35.6 | 5 | 08/06/2014 | Peanut | ||
| Sowing 1/2015 | SIN | S15D1 | Sandy | 93.7 | 6.3 | 0.32 | 3.5 | 505 | 52 | 259 | 153 | 43 | 21.8 | 34.7 | 2 | 07/09/2015 | Peanut | ||
| Sowing 2/2015 | SIN | S15D2 | Sandy | 93.2 | 6.8 | 0.33 | 3.8 | 455 | 259 | 155 | 44 | 2 | 21.2 | 34.9 | 4 | 08/08/2015 | Peanut | ||
| Sowing 1/2016 | SIN | S16D1 | Sandy-silty | 84.1 | 15.9 | 0.55 | 10.6 | 447 | 230 | 155 | 24 | 38 | 22.5 | 35.6 | 5 | 07/25/2016 | Fallow | ||
| Sowing 1/2015 | NIO | N15D1 | 13°45′N 15°45′W | 45 | Sandy | 92.4 | 7.6 | 0.31 | 3.5 | 943 | 196 | 361 | 261 | 126 | 20.6 | 33.8 | 3 | 07/16/2015 | Cowpea |
| Sowing 2/2015 | NIO | N15D2 | Sandy-silty | 87.0 | 13.0 | 0.43 | 6.1 | 747 | 329 | 273 | 145 | 0 | 19.7 | 33.8 | 4 | 08/13/2015 | Fallow |
| Genotype | Code | Type | Photoperiod-Sensitivity | Cycle Duration | Isohyets | Purpose | Plant Height | Yield Potential | Panicle Shape | Others | Origin |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Fadda | G1 | Guinea (Hybride) | Moderate | 110 days | 700–1000 mm | Grain–biomass | 2–3 m | 4.5 t/ha | Semi–loose | Tolerant: mold, anthracnose | Mali, IER/ICRISAT selection, pedigree 02–SB–F5DT–12A xLata. |
| Nieleni | G2 | Caudatum (Hybride) | Low | 100 days | 700–800 mm | Grain | 3 m | 4 t/ha | Semi–compact | Tolerant: mold, anthracnose | Mali, IER/ICRISAT selection |
| IS15401 | G3 | Guinea | High | 120 days | 900–1200 mm | Biomass | 4–4.5 m | 2 t/ha | Semi–compact | Resistant: mold, striga and midges | Cameroon, IER/ICRISAT selection |
| Pablo | G4 | Guinea (Hybride) | Moderate | 110 days | 700–1000 mm | Biomass | 4 m | 4 t/ha | Loose | Tolerant: mold, anthracnose | Mali, IER/ICRISAT selection, pedigree FambeA x Lata. |
| CSM63E | G5 | Guinea | Low | 90 days | 600–1000 mm | Grain | 4 m | 2 t/ha | Loose | Tolerant: diseases and insects | Mali, traditional variety |
| SK5912 | G6 | Caudatum | High | 110 days | 700–900 mm | Biomass | 2 m | 2.5–3.5 t/ha | Semi–compact | Tolerant: mold, anthracnose | Nigeria |
| Grinkan | G7 | Caudatum | No | 110 days | 500–800 mm | Grain–biomass | 1.2 m | 4 t/ha | Semi–compact | Resistant: midges, insects | Mali, ICRISAT selection |
| Soumba | G8 | Caudatum | Low | 100 days | 600–1000 mm | Grain–biomass | 2.5 m | 2.5 t/ha | Semi–compact | Tolerant: diseases and, insects, striga | Mali |
| 621B | G9 | Caudatum | No | 90 days | 600–900 mm | Grain | 1.75 m | 2.5–3 t/ha | Semi–compact | Mold resistant | Senegal, ISRA selection, pedigree CE 151–262 xSarvato–1 |
| F2–20 | G10 | Caudatum | Low | 110 days | 600–900 mm | Grain | 2.1m | 3– 5.3 t/ha | Semi–compact | Resistant: mold, striga | Senegal, ISRA selection, pedigree (MN1056 × 68–20) x 7410–195–1 |
| Source of Variation | Grain (kg ha−1) | Biomass (kg ha−1) | ||||
|---|---|---|---|---|---|---|
| DF | Mean Square | TSS Explained (%) | DF | Mean Square | TSS Explained (%) | |
| Genotype (G) | 9 | 3,990,633 *** | 17.9 | 9 | 178,164,830 *** | 36.7 |
| Environment (E) | 10 | 7,936,033 *** | 39.6 | 10 | 92,439,498 *** | 21.2 |
| Blocks (E) | 33 | 523,880 *** | 8.6 | 33 | 17,553,265 *** | 13.3 |
| Interaction (G × E) | 89 | 759,922 *** | 33.8 | 89 | 14,146,802 *** | 28.8 |
| IPCA1 | 18 | 1,371,515 *** | 36.6 | 18 | 32,487,030 *** | 52 |
| IPCA2 | 16 | 1,117,060 *** | 26.5 | 16 | 17,004,129 *** | 24.2 |
| IPCA3 | 14 | 1,011,953 *** | 21 | 14 | 10,009,335 * | 12.5 |
| IPCA4 | 12 | 396,066 | 7 | 12 | 4,671,580 | 5 |
| 1PCA5 | 10 | 386,334 | 5.7 | 10 | 3,398,772 | 3 |
| Error | 289 | 231,846 | 287 | 4,711,029 | ||
| Genotype | Environment | Genotypic Mean | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| B13D1 | B13D2 | N15D1 | N15D2 | S13D1 | S13D2 | S14D1 | S14D2 | S15D1 | S15D2 | S16D1 | ||
| Fadda | 1662 | 804 | 2417 | 1719 | 2329 | 1855 | 1604 | 2206 | 1634 | 2077 | 1857 | 1833 |
| Nieleni | 2011 | 972 | 1122 | 2824 | 2445 | 1742 | 2626 | 1326 | 2049 | 1871 | 2946 | 2018 |
| IS15401 | 665 | 554 | 1431 | 1310 | 2182 | 1608 | 1297 | 1008 | 1524 | 1958 | 1883 | 1402 |
| Pablo | 1688 | 624 | 1432 | 1796 | 2111 | 1786 | 1592 | 1806 | 1435 | 1716 | 1780 | 1615 |
| CSM63E | 1423 | 346 | 1791 | 478 | 1628 | 1895 | 1939 | 1634 | 232 | 1247 | 2345 | 1360 |
| SK5912 | 252 | 151 | 2050 | 1754 | 992 | 1071 | 459 | 358 | 477 | 503 | – | 807 |
| Grinkan | 888 | 502 | 1929 | 1707 | 1323 | 1475 | 1677 | 441 | 1171 | 881 | 1905 | 1281 |
| Soumba | 931 | 553 | 1381 | 2365 | 1016 | 1301 | 2307 | 900 | 1149 | 1339 | 2412 | 1443 |
| 621B | 1367 | 491 | 1342 | 1503 | 1566 | 1665 | 1810 | 1064 | 533 | 572 | 3233 | 1392 |
| F2–20 | 1223 | 409 | 1207 | 2205 | 1549 | 1302 | 1658 | 1110 | 1012 | 536 | 2453 | 1333 |
| Mean | 1211 | 530 | 1610 | 1766 | 1714 | 1570 | 1697 | 1192 | 1122 | 1270 | 2313 | 1454 |
| Genotype | Environment | Genotypic Mean | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| B13D1 | B13D2 | N15D1 | N15D2 | S13D1 | S13D2 | S14D1 | S14D2 | S15D1 | S15D2 | S16D1 | ||
| Fadda | 11,111 | 10,546 | 10,857 | 6496 | 7571 | 5409 | 10,508 | 4966 | 6056 | 5447 | 8972 | 7995 |
| Nieleni | 7667 | 13,322 | 3990 | 4004 | 5617 | 6473 | 8141 | 7277 | 5895 | 5360 | 8509 | 6784 |
| IS15401 | 12,315 | 15,989 | 5655 | 10,803 | 10,611 | 7151 | 17,077 | 8211 | 5841 | 6276 | 14073 | 10364 |
| Pablo | 8137 | 7198 | 4712 | 5855 | 7109 | 5775 | 8728 | 5982 | 6822 | 5431 | 7460 | 6655 |
| CSM63E | 3643 | 3051 | 5376 | 3134 | 4926 | 4529 | 7650 | 3842 | 4129 | 3591 | 6460 | 4576 |
| SK5912 | 14,806 | 13,623 | 11,591 | 8156 | 9827 | 7480 | 11,870 | 8783 | 6867 | 7332 | – | 10,115 |
| Grinkan | 9675 | 12,020 | 9638 | 3489 | 6870 | 4812 | 8295 | 5297 | 4882 | 4682 | 7094 | 6860 |
| Soumba | 4756 | 7497 | 6669 | 4483 | 4034 | 5109 | 7196 | 5718 | 5991 | 3290 | 5820 | 5459 |
| 621B | 3581 | 5955 | 4969 | 3003 | 4300 | 4889 | 4258 | 6829 | 3308 | 2888 | 5413 | 4379 |
| F2–20 | 4863 | 7219 | 13,140 | 4061 | 5261 | 5791 | 7568 | 7602 | 4833 | 4931 | 6870 | 6558 |
| Mean | 8055 | 9536 | 7660 | 5348 | 6613 | 5742 | 9129 | 6431 | 5426 | 4923 | 7852 | 6954 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ndiaye, M.; Adam, M.; Ganyo, K.K.; Guissé, A.; Cissé, N.; Muller, B. Genotype-Environment Interaction: Trade-Offs between the Agronomic Performance and Stability of Dual-Purpose Sorghum (Sorghum bicolor L. Moench) Genotypes in Senegal. Agronomy 2019, 9, 867. https://doi.org/10.3390/agronomy9120867
Ndiaye M, Adam M, Ganyo KK, Guissé A, Cissé N, Muller B. Genotype-Environment Interaction: Trade-Offs between the Agronomic Performance and Stability of Dual-Purpose Sorghum (Sorghum bicolor L. Moench) Genotypes in Senegal. Agronomy. 2019; 9(12):867. https://doi.org/10.3390/agronomy9120867
Chicago/Turabian StyleNdiaye, Malick, Myriam Adam, Komla Kyky Ganyo, Aliou Guissé, Ndiaga Cissé, and Bertrand Muller. 2019. "Genotype-Environment Interaction: Trade-Offs between the Agronomic Performance and Stability of Dual-Purpose Sorghum (Sorghum bicolor L. Moench) Genotypes in Senegal" Agronomy 9, no. 12: 867. https://doi.org/10.3390/agronomy9120867
APA StyleNdiaye, M., Adam, M., Ganyo, K. K., Guissé, A., Cissé, N., & Muller, B. (2019). Genotype-Environment Interaction: Trade-Offs between the Agronomic Performance and Stability of Dual-Purpose Sorghum (Sorghum bicolor L. Moench) Genotypes in Senegal. Agronomy, 9(12), 867. https://doi.org/10.3390/agronomy9120867




