Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges
Abstract
1. Introduction
2. Approaches for Phenotyping
3. Proximal and Remote Sensing Technologies in Horticulture
3.1. Definition and Reference Characteristics
3.2. Radiation Interception
3.3. Radiation Reflectance, Absorbance, and Transmittance
3.4. Fluorescence
3.5. Radiation Emission
3.6. Magnetic Resonance
3.7. Performance Indices
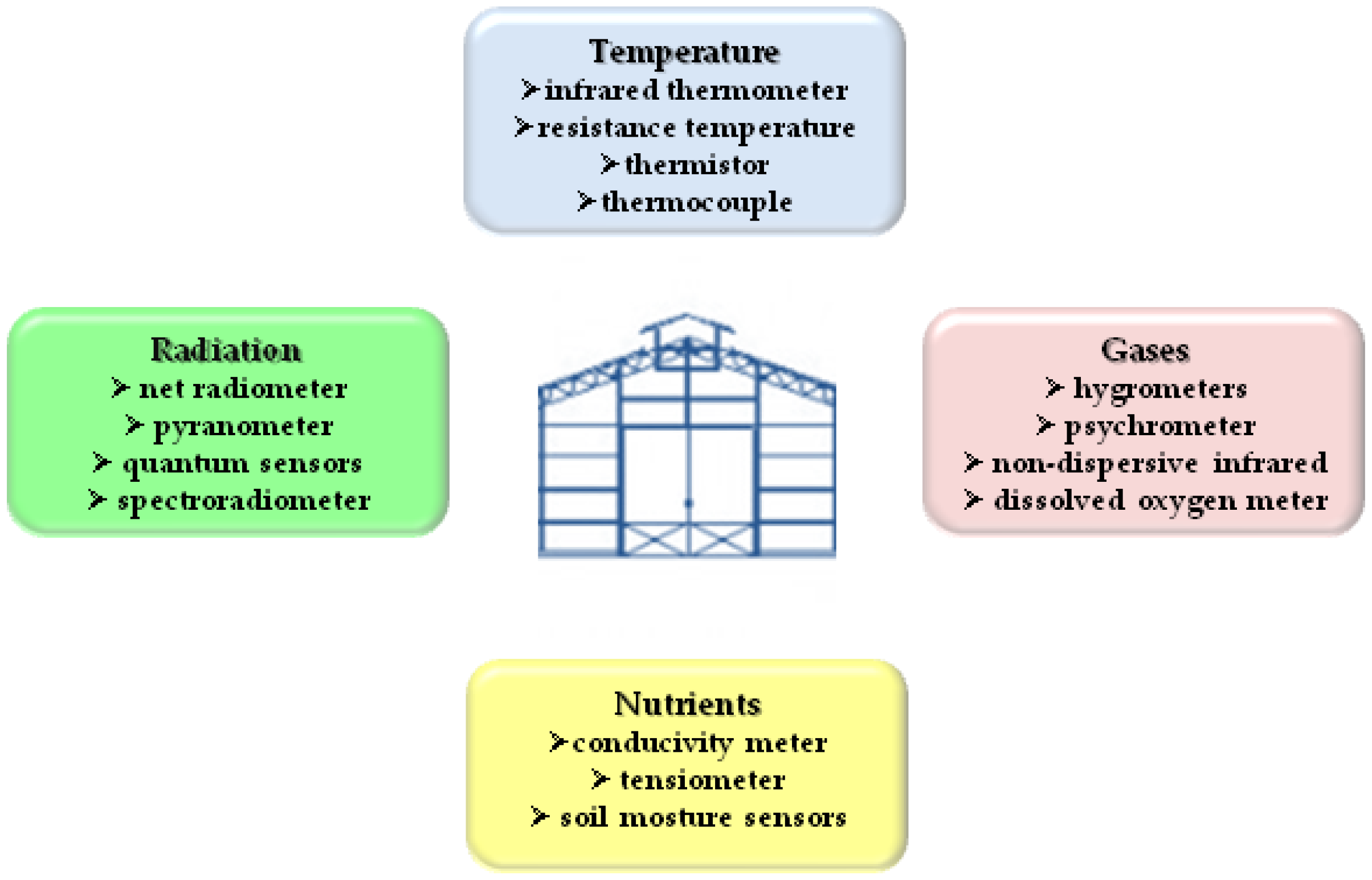
4. Controlled Indoor Environments: From Cultivation to Phenotyping
5. Next Generation Phenotyping Studies Using Automated Devices in Vegetable Crops
5.1. Hyperspectral Sensing Studies
5.2. Magnetic Resonance Studies
5.3. VIS/NIR Studies
5.4. Chlorophyll Fluorescence and Content Studies
5.5. RGB and LIDAR for Morphology and Canopy Assessment
6. Conclusions and Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Furbank, R.T.; Tester, M. Phenomics-technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 2011, 16, 635–644. [Google Scholar] [CrossRef] [PubMed]
- Collard, B.C.Y.; Mackill, D.J. Marker-assisted selection: An approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. B 2008. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.K.; Rustgi, S.; Kulwal, P.L. Linkage disequilibrium and association studies in higher plants: Present status and future prospects. Plant Mol. Biol. 2005, 57, 461–485. [Google Scholar] [CrossRef] [PubMed]
- Cobb, J.N.; DeClerck, G.; Greenberg, A.; Clark, R.; McCouch, S. Next-generation phenotyping: Requirements and strategies for enhancing our understanding of genotype-phenotype relationships and its relevance to crop improvement. Theor. Appl. Genet. 2013, 126, 867–887. [Google Scholar] [CrossRef] [PubMed]
- Dhondt, S.; Wuyts, N.; Inzé, D. Cell to whole-plant phenotyping: The best is yet to come. Trends Plant Sci. 2013, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fiorani, F.; Schurr, U. Future scenarios for plant phenotyping. Annu. Rev. Plant Biol. 2013, 64, 267–291. [Google Scholar] [CrossRef] [PubMed]
- Araus, J.L.; Cairns, J.E. Field high-throughput phenotyping: The new crop breeding frontier. Trends Plant Sci. 2014, 19, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Montepellier Plant Phenotyping Network. Available online: https://www6.montpellier.inra.fr/lepse_eng/M3P (accessed on 28 February 2018).
- JPPC-the Jülich Plant Phenotyping Centre. Available online: http://www.fz-juelich.de/ibg/ibg-2/EN/_organisation/JPPC/JPPC_node.html (accessed on 28 February 2018).
- IPK Gaterbsleben Phenotyping. Available online: http://www.ipk-gatersleben.de/en/phenotyping/ (accessed on 28 February 2018).
- APPS. Available online: https://www.plantphenomics.org.au/ (accessed on 28 February 2018).
- Humplık, J.F.; Lazar, D.; Husickova, A.; Spıchal, L. Automated phenotyping of plant shoots using imaging methods for analysis of plant stress responses—A review. Plant Methods 2015, 11, 29. [Google Scholar] [CrossRef] [PubMed]
- Shakoor, N.; Lee, S.; Mockler, T.C. High throughput phenotyping to accelerate crop breeding and monitoring of diseases in the field. Curr. Opin. Plant Biol. 2017, 38, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhang, Q.; Huang, D. A review of imaging techniques for plant phenotyping. Sensors 2014, 14, 20078–20111. [Google Scholar] [CrossRef] [PubMed]
- Usha, K.; Singh, B. Potential applications of remote sensing in horticulture-A review. Sci. Hortic. (Amsterdam) 2013, 153, 71–83. [Google Scholar] [CrossRef]
- Ruiz-Garcia, L.; Lunadei, L.; Barreiro, P.; Robla, J.I. A review of wireless sensor technologies and applications in agriculture and food industry: State of the art and current trends. Sensors 2009, 9, 4728–4750. [Google Scholar] [CrossRef] [PubMed]
- Nishina, H. Development of speaking plant approach technique for intelligent greenhouse. Agric. Agric. Sci. Procedia 2015, 3, 9–13. [Google Scholar] [CrossRef]
- Incrocci, L.; Massa, D.; Pardossi, A. New trends in the fertigation management of irrigated vegetable crops. Horticulturae 2017, 3, 37. [Google Scholar] [CrossRef]
- Muñoz-Huerta, R.F.; Guevara-Gonzalez, R.G.; Contreras-Medina, L.M.; Torres-Pacheco, I.; Prado-Olivarez, J.; Ocampo-Velazquez, R.V. A review of methods for sensing the nitrogen status in plants: Advantages, disadvantages and recent advances. Sensors 2013, 13, 10823–10843. [Google Scholar] [CrossRef] [PubMed]
- Basu, S.; Rabara, R. Abscisic acid—An enigma in the abiotic stress tolerance of crop plants. Plant Gene 2017, 11, 90–98. [Google Scholar] [CrossRef]
- Marschner, H. Marschner’s Mineral Nutrition of Higher Plants, 3rd ed.; Accademic Press: London, UK, 2011. [Google Scholar]
- Deery, D.; Jimenez-Berni, J.; Jones, H.; Sirault, X.; Furbank, R. Proximal Remote Sensing Buggies and Potential Applications for Field-Based Phenotyping. Agronomy 2014, 4, 349–379. [Google Scholar] [CrossRef]
- Fahlgren, N.; Gehan, M.A.; Baxter, I. Lights, camera, action: High-throughput plant phenotyping is ready for a close-up. Curr. Opin. Plant Biol. 2015, 24, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Pound, M.P.; French, A.P.; Murchie, E.H.; Pridmore, T.P. Automated recovery of three-dimensional models of plant shoots from multiple color images. Plant Physiol. 2014, 166, 1688–1698. [Google Scholar] [CrossRef] [PubMed]
- Paulus, S.; Dupuis, J.; Mahlein, A.-K.; Kuhlmann, H. Surface feature based classification of plant organs from 3D laserscanned point clouds for plant phenotyping. BMC Bioinform. 2013, 14, 238. [Google Scholar] [CrossRef] [PubMed]
- Chaudhury, A.; Ward, C.; Talasaz, A.; Ivanov, A.G.; Brophy, M.; Grodzinski, B.; Huner, N.P.A.; Patel, R.V.; Barron, J.L. Machine Vision System for 3D Plant Phenotyping. Available online: http://arxiv.org/abs/1705.00540 (accessed on 15 April 2018).
- Gitelson, A.A.; Gritz, Y.; Merzlyak, M.N. Relationships between leaf chlorophyll content and spectral reflectance and algorithms for non-destructive chlorophyll assessment in higher plant leaves. J. Plant Physiol. 2003, 160, 271–282. [Google Scholar] [CrossRef] [PubMed]
- Francone, C.; Pagani, V.; Foi, M.; Cappelli, G.; Confalonieri, R. Comparison of leaf area index estimates by ceptometer and PocketLAI smart app in canopies with different structures. Field Crop Res. 2014, 155, 38–41. [Google Scholar] [CrossRef]
- Golzarian, M.R.; Frick, R.A.; Rajendran, K.; Berger, B.; Roy, S.; Tester, M.; Lun, D.S. Accurate inference of shoot biomass from high-throughput images of cereal plants. Plant Methods 2011, 7. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Pattey, E. Retrieval of leaf area index from top-of-canopy digital photography over agricultural crops. Agric. For. Meteorol. 2010, 150, 1485–1490. [Google Scholar] [CrossRef]
- Arvidsson, S.; Pérez-Rodríguez, P.; Mueller-Roeber, B. A growth phenotyping pipeline for Arabidopsis thaliana integrating image analysis and rosette area modeling for robust quantification of genotype effects. New Phytol. 2011, 191, 895–907. [Google Scholar] [CrossRef] [PubMed]
- Iyer-Pascuzzi, A.S.; Symonova, O.; Mileyko, Y.; Hao, Y.; Belcher, H.; Harer, J.; Weitz, J.S.; Benfey, P.N. Imaging and analysis platform for automatic phenotyping and trait ranking of plant root systems. Plant Physiol. 2010, 152, 1148–1157. [Google Scholar] [CrossRef] [PubMed]
- Hoyos-Villegas, V.; Houx, J.H.; Singh, S.K.; Fritschi, F.B. Ground-based digital imaging as a tool to assess soybean growth and yield. Crop Sci. 2014, 54, 1756–1768. [Google Scholar] [CrossRef]
- Takizawa, H.; Ezaki, N.; Mizuno, S.; Yamamoto, S. Measurement of plants by stereo vision for agricultural applications. In Proceedings of the Seventh IASTED International Conference on Signal and Image Processing, SIP 2005, Honolulu, HI, USA, 15–17 August 2005; pp. 359–364. [Google Scholar]
- Omasa, K.; Hosoi, F.; Konishi, A. 3D lidar imaging for detecting and understanding plant responses and canopy structure. J. Exp. Bot. 2007, 58, 881–898. [Google Scholar] [CrossRef] [PubMed]
- Llop, J.; Gil, E.; Llorens, J.; Miranda-Fuentes, A.; Gallart, M. Testing the suitability of a terrestrial 2D LiDAR scanner for canopy characterization of greenhouse tomato crops. Sensors 2016, 16, 1435. [Google Scholar] [CrossRef] [PubMed]
- Hosoi, F.; Nakabayashi, K.; Omasa, K. 3-D modeling of tomato canopies using a high-resolution portable scanning lidar for extracting structural information. Sensors 2011, 11, 2166–2174. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhu, H.; Ozkan, H.E. Real-time tree foliage density estimation with laser scanning sensor for variable-rate tree sprayer development. In Proceedings of the American Society of Agricultural and Biological Engineers Annual International Meeting 2013, ASABE 2013, Kansas City, MO, USA, 21–24 July 2013; Volume 3, pp. 2274–2285. [Google Scholar]
- Dornbusch, T.; Lorrain, S.; Kuznetsov, D.; Fortier, A.; Liechti, R.; Xenarios, I.; Fankhauser, C. Measuring the diurnal pattern of leaf hyponasty and growth in Arabidopsis–a novel phenotyping approach using laser scanning. Funct. Plant Biol. 2012, 39, 860–869. [Google Scholar] [CrossRef]
- Li, Y.; Fan, X.; Mitra, N.J.; Chamovitz, D.; Cohen-Or, D.; Chen, B. Analyzing growing plants from 4D point cloud data. ACM Trans. Graph. 2013, 32. [Google Scholar] [CrossRef]
- Maguire, M.S.; Woldt, W.E.; Frew, E.W.; Smith, J.; Elston, J. Thermal infrared and multi-spectral dual sensor integration for unmanned aircraft systems. In Proceedings of the 2017 ASABE Annual International Meeting, Spokane, WA, USA, 16–19 July 2017. [Google Scholar]
- Corti, M.; Gallina, P.M.; Cavalli, D.; Cabassi, G. Hyperspectral imaging of spinach canopy under combined water and nitrogen stress to estimate biomass, water, and nitrogen content. Biosyst. Eng. 2017, 158, 38–50. [Google Scholar] [CrossRef]
- Thorp, K.R.; Gore, M.A.; Andrade-Sanchez, P.; Carmo-Silva, A.E.; Welch, S.M.; White, J.W.; French, A.N. Proximal hyperspectral sensing and data analysis approaches for field-based plant phenomics. Comput. Electron. Agric. 2015, 118, 225–236. [Google Scholar] [CrossRef]
- Misra, A.N.; Misra, M.; Singh, R. Chlorophyll Fluorescence in Plant Biology. In Biophysics; Misra, A.N., Ed.; InTech: London, UK, 2012; p. 220. ISBN 978-953-51-0376-9. [Google Scholar]
- Belasque, J.; Gasparoto, M.C.G.; Marcassa, L.G. Detection of mechanical and disease stresses in citrus plants by fluorescence spectroscopy. Appl. Opt. 2008, 47, 1922–1926. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Sun, H.; Li, M.Z.; Yang, W. Development and application of crop monitoring system for detecting chlorophyll content of tomato seedlings. Int. J. Agric. Biol. Eng. 2014, 7, 138–145. [Google Scholar] [CrossRef]
- Ding, Y.; Li, M.; Sun, H.; Li, X.; Zhao, R. Diagnosis model of tomato nutrient content based on multispectral images. Nongye Gongcheng Xuebao/Trans. Chin. Soc. Agric. Eng. 2012, 28, 175–180. [Google Scholar] [CrossRef]
- Elvanidi, A.; Katsoulas, N.; Bartzanas, T.; Ferentinos, K.P.; Kittas, C. Crop water status assessment in controlled environment using crop reflectance and temperature measurements. Precis. Agric. 2017, 18, 332–349. [Google Scholar] [CrossRef]
- Li, B.; Emr, N.; Malling, E.; Me, K. Advances in Non-Destructive Early Assessment of Fruit Ripeness towards Defining Optimal Time of Harvest and Yield Prediction—A Review. Plants 2018, 7, 3. [Google Scholar] [CrossRef]
- Durmus, H.; Gunes, E.O.; Kirci, M. Disease detection on the leaves of the tomato plants by using deep learning. In Proceedings of the 6th International Conference on Agro-Geoinformatics, Agro-Geoinformatics 2017, Fairfax, VA, USA, 7–10 August 2017. [Google Scholar]
- Candiago, S.; Remondino, F.; De Giglio, M.; Dubbini, M.; Gattelli, M. Evaluating multispectral images and vegetation indices for precision farming applications from UAV images. Remote Sens. 2015, 7, 4026–4047. [Google Scholar] [CrossRef]
- Rouse, J.W.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring Vegetation Systems in the Great Plains with ERTS. In Proceedings of the Earth Resources Technology Satellite Symposium, Washington, DC, USA, 10–14 December 1973; p. 371. [Google Scholar]
- Di Gennaro, S.F.; Rizza, F.; Badeck, F.W.; Berton, A.; Delbono, S.; Gioli, B.; Toscano, P.; Zaldei, A.; Matese, A. UAV-based high-throughput phenotyping to discriminate barley vigour with visible and near-infrared vegetation indices. Int. J. Remote Sens. 2017, 1–15. [Google Scholar] [CrossRef]
- Agati, G.; Tuccio, L.; Kusznierewicz, B.; Chmiel, T.; Bartoszek, A.; Kowalski, A.; Grzegorzewska, M.; Kosson, R.; Kaniszewski, S. Nondestructive optical sensing of flavonols and chlorophyll in white head cabbage (Brassica oleracea L. var. capitata subvar. alba) grown under different nitrogen regimens. J. Agric. Food Chem. 2016, 64, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhang, F.; Ma, Q.; An, D.; Li, L.; Zhang, X.; Zhu, D.; Li, S. Advances in crop phenotyping and multi-environment trials. Front. Agric. Sci. Eng. 2015, 2, 28–37. [Google Scholar] [CrossRef]
- Lenk, S.; Chaerle, L.; Pfündel, E.E.; Langsdorf, G.; Hagenbeek, D.; Lichtenthaler, H.K.; Van Der Straeten, D.; Buschmann, C.; van der Straeten, D.; Buschmann, C.; et al. Multispectral fluorescence and reflectance imaging at the leaf level and its possible applications. J. Exp. Bot. 2007, 58, 807–814. [Google Scholar] [CrossRef] [PubMed]
- Jones, H.G. Application of thermal imaging and infrared sensing in plant physiology and ecophysiology. In Incorporating Advances in Plant Pathology; Advances in Botanical Research; Academic Press: Cambridge, MA, USA, 2004; Volume 41, pp. 107–163. [Google Scholar]
- Nilsson, H.E. Remote sensing and image analysis in plant pathology. Annu. Rev. Phytopathol. 1995, 33, 489–527. [Google Scholar] [CrossRef] [PubMed]
- Chaerle, L.; Van Der Straeten, D. Imaging techniques and the early detection of plant stress. Trends Plant Sci. 2000, 5, 495–501. [Google Scholar] [CrossRef]
- Prashar, A.; Yildiz, J.; McNicol, J.W.; Bryan, G.J.; Jones, H.G. Infra-red thermography for high throughput field phenotyping in Solanum tuberosum. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Moradi, A.B.; Oswald, S.E.; Nordmeyer-Massner, J.A.; Pruessmann, K.P.; Robinson, B.H.; Schulin, R. Analysis of nickel concentration profiles around the roots of the hyperaccumulator plant Berkheya coddii using MRI and numerical simulations. Plant Soil 2010, 328, 291–302. [Google Scholar] [CrossRef]
- Van As, H. Intact plant MRI for the study of cell water relations, membrane permeability, cell-to-cell and long distance water transport. J. Exp. Bot. 2007, 58, 743–756. [Google Scholar] [CrossRef] [PubMed]
- Windt, C.W.; Vergeldt, F.J.; De Jager, P.A.; Van As, H. MRI of long-distance water transport: A comparison of the phloem and xylem flow characteristics and dynamics in poplar, castor bean, tomato and tobacco. Plant Cell Environ. 2006, 29, 1715–1729. [Google Scholar] [CrossRef] [PubMed]
- Melkus, G.; Rolletschek, H.; Fuchs, J.; Radchuk, V.; Grafahrend-Belau, E.; Sreenivasulu, N.; Rutten, T.; Weier, D.; Heinzel, N.; Schreiber, F.; et al. Dynamic 13C/1H NMR imaging uncovers sugar allocation in the living seed. Plant Biotechnol. J. 2011, 9, 1022–1037. [Google Scholar] [CrossRef] [PubMed]
- Bühler, J.; Huber, G.; Schmid, F.; Blümler, P. Analytical model for long-distance tracer-transport in plants. J. Theor. Biol. 2011, 270, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Kiyomiya, S.; Nakanishi, H.; Uchida, H.; Tsuji, A.; Nishiyama, S.; Futatsubashi, M.; Tsukada, H.; Ishioka, N.S.; Watanabe, S.; Ito, T.; et al. Real time visualization of 13N-translocation in rice under different environmental conditions using positron emitting tracer imaging system. Plant Physiol. 2001, 125, 1743–1754. [Google Scholar] [CrossRef] [PubMed]
- Padilla, F.M.; Peña-Fleitas, M.T.; Gallardo, M.; Thompson, R.B. Determination of sufficiency values of canopy reflectance vegetation indices for maximum growth and yield of cucumber. Eur. J. Agron. 2017, 84, 1–15. [Google Scholar] [CrossRef]
- Hunink, J.E.; Contreras, S.; Soto-García, M.; Martin-Gorriz, B.; Martinez-Álvarez, V.; Baille, A. Estimating groundwater use patterns of perennial and seasonal crops in a Mediterranean irrigation scheme, using remote sensing. Agric. Water Manag. 2015, 162, 47–56. [Google Scholar] [CrossRef]
- Alsiņa, I.; Dūma, M.; Dubova, L.; Šenberga, A.; Daģis, S. Comparison of different chlorophylls determination methods for leafy vegetables. Agron. Res. 2016, 14, 309–316. [Google Scholar]
- Galieni, A.; Stagnari, F.; Speca, S.; Pisante, M. Leaf traits as indicators of limiting growing conditions for lettuce (Lactuca sativa). Ann. Appl. Biol. 2016, 169, 342–356. [Google Scholar] [CrossRef]
- Kizil, U.; Genç, L.; Inalpulat, M.; Şapolyo, D.; Mirik, M. Lettuce (Lactuca sativa L.) yield prediction under water stress using artificial neural network (ANN) model and vegetation indices [Sėjamosios salotos (Lactuca sativa L.) derliaus prognozavimas vandens streso sa{ogonek}lygomis, taikant dirbtinio neurotink]. Zemdirbyste 2012, 99, 409–418. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Merzlyak, M.N. Use of a green channel in remote sensing of global vegetation from EOS-MODIS. Remote Sens. Environ. 1996, 58, 289–298. [Google Scholar] [CrossRef]
- Gitelson, A.; Merzlyak, M.N. Spectral reflectance changes associated with autumn senescence of Aesculus hippocastanum L. and Acer platanoides L. leaves. Spectral features and relation to chlorophyll estimation. J. Plant Physiol. 1994, 143, 286–292. [Google Scholar] [CrossRef]
- Vogelmann, J.E.; Rock, B.N.; Moss, D.M. Red edge spectral measurements from sugar maple leaves. Int. J. Remote Sens. 1993, 14, 1563–1575. [Google Scholar] [CrossRef]
- Birth, G.S.; McVey, G.R. Measuring the color of growing turf with a reflectance spectrophotometer1. Agron. J. 1968, 60, 640–643. [Google Scholar] [CrossRef]
- Penuelas, J.; Filella, I.; Biel, C.; Serrano, L.; Save, R. The reflectance at the 950–970 nm region as an indicator of plant water status. Int. J. Remote Sens. 1993, 14, 1887–1905. [Google Scholar] [CrossRef]
- Dash, J.; Curran, P.J. The MERIS terrestrial chlorophyll index. Int. J. Remote Sens. 2004, 25, 5403–5413. [Google Scholar] [CrossRef]
- Xue, L.; Yang, L. Deriving leaf chlorophyll content of green-leafy vegetables from hyperspectral reflectance. ISPRS J. Photogramm. Remote Sens. 2009, 64, 97–106. [Google Scholar] [CrossRef]
- Marino, S.; Basso, B.; Leone, A.P.; Alvino, A. Agronomic traits and vegetation indices of two onion hybrids. Sci. Hortic. (Amsterdam) 2013, 155, 56–64. [Google Scholar] [CrossRef]
- Kabakeris, T.; Poth, A.; Intreß, J.; Schmidt, U.; Geyer, M. Detection of postharvest quality loss in broccoli by means of non-colorimetric reflection spectroscopy and hyperspectral imaging. Comput. Electron. Agric. 2015, 118, 322–331. [Google Scholar] [CrossRef]
- Padilla, F.M.; Peña-Fleitas, M.T.; Gallardo, M.; Thompson, R.B. Threshold values of canopy reflectance indices and chlorophyll meter readings for optimal nitrogen nutrition of tomato. Ann. Appl. Biol. 2015, 166, 271–285. [Google Scholar] [CrossRef]
- De Benedetto, D.; Castrignanò, A.; Diacono, M.; Rinaldi, M.; Ruggieri, S.; Tamborrino, R. Field partition by proximal and remote sensing data fusion. Biosyst. Eng. 2013, 114, 372–383. [Google Scholar] [CrossRef]
- Soto, F.; Gallardo, M.; Thompson, R.B.; Peña-Fleitas, M.T.; Padilla, F.M. Consideration of total available N supply reduces N fertilizer requirement and potential for nitrate leaching loss in tomato production. Agric. Ecosyst. Environ. 2015, 200, 62–70. [Google Scholar] [CrossRef]
- Yang, I.-C.; Chen, S. Precision cultivation system for greenhouse production. Smart Sens. Meas. Instrum. 2015, 13, 191–211. [Google Scholar] [CrossRef]
- Sridhar, B.B.M.; Witter, J.D.; Wu, C.; Spongberg, A.L.; Vincent, R.K. Effect of biosolid amendments on the metal and nutrient uptake and spectral characteristics of five vegetable plants. Water Air Soil Pollut. 2014, 225. [Google Scholar] [CrossRef]
- Marino, S.; Alvino, A. Proximal sensing and vegetation indices for site-specific evaluation on an irrigated crop tomato. Eur. J. Remote Sens. 2014, 47, 271–283. [Google Scholar] [CrossRef]
- Caramante, M.; Oliva, M.; Ricci, S.; Ruggiero, A.; D’Agostino, N.; Venezia, A.; Mennella, G.; Albrizio, R.; Giorio, P.; Grillo, S. Pepper response to salt stress at physiological, molecular and biochemical level. In Proceedings of the 3rd Spot-ITN Conference, Stress Biology and Crop Fertility, Sorrento, Italy, 18–22 March 2015; p. 50. [Google Scholar]
- Both, A.J.; Benjamin, L.; Franklin, J.; Holroyd, G.; Incoll, L.D.; Lefsrud, M.G.; Pitkin, G. Guidelines for measuring and reporting environmental parameters for experiments in greenhouses. Plant Methods 2015, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Apan, A.; Datt, B.; Kelly, R. Detection of pests and diseases in vegetable crops using hyperspectral sensing: A comparison of reflectance data for different sets of symptoms. In Proceedings of the SSC 2005 Spatial Intelligence, Innovation and Praxis: The National biennial Conference of the Spatial Sciences Institute, Melbourne, Australia, 12–16 September 2005. [Google Scholar]
- Hahn, F. Spectral bandwidth effect on a Rhizopus stolonifer spores detector and its on-line behavior using red tomato fruits. Can. Biosyst. Eng. 2004, 46, 349–354. [Google Scholar]
- Xu, H.R.; Ying, Y.B.; Fu, X.P.; Zhu, S.P. Near-infrared spectroscopy in detecting leaf miner damage on tomato leaf. Biosyst. Eng. 2007, 96, 447–454. [Google Scholar] [CrossRef]
- Xing, J.; Ngadi, M.; Wang, N.; De Baerdemaeker, J. Wavelength selection for surface defects detection on tomatoes by means of a hyperspectral imaging system. In Proceedings of the ASAE Annual International Meeting, Portland, OR, USA, 9–12 July 2006. [Google Scholar]
- Polder, G.; van der Heijden, G.W.A.M.; Young, I.T. Hyperspectral image analysis for measuring ripeness of tomatoes. In Proceedings of the ASAE Annual International Meeting, Milwaukee, WI, USA, 9–12 July 2000. [Google Scholar]
- Diezma, B.; Lleó, L.; Roger, J.M.; Herrero-Langreo, A.; Lunadei, L.; Ruiz-Altisent, M. Examination of the quality of spinach leaves using hyperspectral imaging. Postharv. Biol. Technol. 2013, 85, 8–17. [Google Scholar] [CrossRef]
- Tamburini, E.; Costa, S.; Rugiero, I.; Pedrini, P.; Marchetti, M.G. Quantification of Lycopene, -Carotene, and Total Soluble Solids in Intact Red-Flesh Watermelon (Citrullus lanatus) Using On-Line Near-Infrared Spectroscopy. Sensors 2017, 17, 746. [Google Scholar] [CrossRef] [PubMed]
- Simko, I.; Hayes, R.J.; Furbank, R.T. Non-destructive Phenotyping of Lettuce Plants in Early Stages of Development with Optical Sensors. Front. Plant Sci. 2016, 7, 1985. [Google Scholar] [CrossRef] [PubMed]
- Windt, C.W.; Gerkema, E.; Van As, H. Most water in the tomato truss is imported through the xylem, not the phloem: A nuclear magnetic resonance flow imaging study. Plant Physiol. 2009, 151, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; McCarthy, M.J. Measurement and evaluation of tomato maturity using magnetic resonance imaging. Postharvest Biol. Technol. 2012, 67, 37–43. [Google Scholar] [CrossRef]
- Kotwaliwale, N.; Curtis, E.; Othman, S.; Naganathan, G.K.; Subbiah, J. Magnetic resonance imaging and relaxometry to visualize internal freeze damage to pickling cucumber. Postharvest Biol. Technol. 2012, 68, 22–31. [Google Scholar] [CrossRef]
- Rascher, U.; Blossfeld, S.; Fiorani, F.; Jahnke, S.; Jansen, M.; Kuhn, A.J.; Matsubara, S.; Märtin, L.L.A.; Merchant, A.; Metzner, R.; et al. Non-invasive approaches for phenotyping of enhanced performance traits in bean. Funct. Plant Biol. 2011, 38, 968. [Google Scholar] [CrossRef]
- Ecarnot, M.; Baczyk, P.; Tessarotto, L.; Chervin, C. Rapid phenotyping of the tomato fruit model, Micro-Tom, with a portable VIS-NIR spectrometer. Plant Physiol. Biochem. 2013, 70, 159–163. [Google Scholar] [CrossRef] [PubMed]
- Szuvandzsiev, P.; Helyes, L.; Lugasi, A.; Szántó, C.; Baranowski, P.; Pék, Z. Estimation of antioxidant components of tomato using VIS-NIR reflectance data by handheld portable spectrometer. Int. Agrophys. 2014, 28, 521. [Google Scholar]
- Clément, A.; Dorais, M.; Vernon, M. Multivariate approach to the measurement of tomato maturity and gustatory attributes and their rapid assessment by vis-NIR spectroscopy. J. Agric. Food Chem. 2008, 56, 1538–1544. [Google Scholar] [PubMed]
- Xu, H.R.; Yu, P.; Fu, X.P.; Ying, Y.B. On-site variety discrimination of tomato plant using visible-near infrared reflectance spectroscopy. J. Zhejiang Univ. Sci. B 2009, 10, 126–132. [Google Scholar] [PubMed]
- Xie, L.; Ying, Y.; Ying, T. Quantification of chlorophyll content and classification of nontransgenic and transgenic tomato leaves using visible/near-infrared diffuse reflectance spectroscopy. J. Agric. Food Chem. 2007, 55, 4645–4650. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.Q. Nondestructive Prediction of Optimal Harvest Time of Cherry Tomatoes Using VIS-NIR Spectroscopy and PLSR Calibration. Adv. Eng. Forum 2011, 1, 92–96. [Google Scholar] [CrossRef]
- Mishra, Y.; Jankanpaa, H.J.; Kiss, A.Z.; Funk, C.; Schroder, W.P.; Jansson, S. Arabidopsis plants grown in the field and climate chambers significantly differ in leaf morphology and photosystem components. BMC Plant Biol. 2012, 12, 6. [Google Scholar] [CrossRef] [PubMed]
- Devacht, S.; Lootens, P.; Baert, J.; Van Waes, J.; Van Bockstaele, E.; Roldán-Ruiz, I. Evaluation of cold stress of young industrial chicory (Cichorium intybus L.) plants by chlorofyll a fluorescence imaging. I. Light induction curve. Photosynthetica 2011, 49, 161–171. [Google Scholar] [CrossRef]
- Lootens, P.; Devacht, S.; Baert, J.; Van Waes, J.; Van Bockstaele, E.; Roldán-Ruiz, I. Evaluation of cold stress of young industrial chicory (Cichorium intybus L.) plants byvchlororphyll a fluorescence imaging. II. Dark relaxation kinetics. Photosynthetica 2011, 49, 185–194. [Google Scholar] [CrossRef]
- Rodriguez-Moreno, L.; Pineda, M.; Soukupova, J.; Macho, A.P.; Beuzon, C.R.; Baron, M. Early detection of bean infection by Pseudomonas syringae in asymptomatic leaf areas using chlorophyll fluorescence imaging. Photosynth. Res. 2008, 96, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Chaerle, L.; Hagenbeek, D.; Vanrobaeys, X.; Van Der Straeten, D. Jones and Schofield early detection of nutrient and biotic stress in Phaseolus vulgaris. Int. J. Remote Sens. 2007, 28, 3479–3492. [Google Scholar] [CrossRef]
- Ptushenko, V.V.; Avercheva, O.V.; Bassarskaya, E.M.; Berkovich, Y.A.; Erokhin, A.N.; Smolyanina, S.O.; Zhigalova, T.V. Possible reasons of a decline in growth of Chinese cabbage under a combined narrowband red and blue light in comparison with illumination by high pressure sodium lamp. Sci. Hortic. 2015. [Google Scholar] [CrossRef]
- Calatayud, Á.; San Bautista, A.; Pascual, B.; Maroto, J.V.; López-Galarza, S. Use of chlorophyll fluorescence imaging as diagnostic technique to predict compatibility in melon graft. Sci. Hortic. 2013, 149, 13–18. [Google Scholar] [CrossRef]
- Hoffmann, A.M.; Noga, G.; Hunsche, M. Fluorescence indices for monitoring the ripening of tomatoes in pre- and postharvest phases. Sci. Hortic. 2015, 191, 74–81. [Google Scholar] [CrossRef]
- Limantara, L.; Dettling, M.; Indrawati, R.; Indriatmoko, T.; Brotosudarmoa, P. Analysis on the chlorophyll content of commercial green leafy vegetables. Procedia Chem. 2015, 14, 225–231. [Google Scholar] [CrossRef]
- Van der Heijden, G.; Song, Y.; Horgan, G.; Polder, G.; Dieleman, A.; Bink, M.; Palloix, A.; van Eeuwijk, F.; Glasbey, C. Spicy: Towards automated phenotyping of large pepper plants in the greenhouse. Funct. Plant Biol. 2012, 39, 870–877. [Google Scholar] [CrossRef]
- Aguilar, M.A.; Pozo, J.L.; Aguilar, F.J.; Sánchez-Hermosilla, J.; Páez, F.C.; Negreiros, J.G. 3D surface modelling of tomato plants using close-range photogrammetry. Int. Arch. Photogram. Remote Sens. Spat. Inf. Sci. 2008, XXXVII, 139–144. [Google Scholar]
- Gonzalo, M.J.; Brewer, M.T.; Anderson, C.; Sullivan, D.; Gray, S.; van der Knaap, E. Tomato fruit shape analysis using morphometric and morphology attributes implemented in Tomato Analyzer software program. J. Am. Soc. Hortic. Sci. 2009, 134, 77–87. [Google Scholar]
- Hurtado, M.; Vilanova, S.; Plazas, M.; Gramazio, P.; Herraiz, F.J.; Andújar, I. Phenomics of fruit shape in eggplant (Solanum melongena L.) using tomato Analyzer software. Sci. Hortic. 2013, 164, 625–632. [Google Scholar] [CrossRef]
- Figàs, M.R.; Prohens, J.; Raigón, M.D.; Fernández-de-Córdova, P.; Fita, A.; Soler, S. Characterization of a collection of local varieties of tomato (Solanum lycopersicum L.) using conventional descriptors and the high-throughput phenomics tool Tomato Analyzer. Genet. Res. Crop Evol. 2015, 62, 189–204. [Google Scholar] [CrossRef]
- Knoblauch, M.; Peters, W.S.; Ehlers, K.; van Bel, A.J.E. Reversible calcium-regulated stopcocks in legume sieve tubes. Plant Cell 2001, 13, 1221–1230. [Google Scholar] [CrossRef] [PubMed]
- Windt, C.W.; Soltner, H.; Dusschoten, D.; Blümler, P. A portable Halbach magnet that can be opened and closed without force: The NMR-CUFF. J. Magn. Reson. 2011, 208, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.R.; Rosenqvist, E. Applications of chlorophyll fluorescence can improve crop production strategies: An examination of future possibilities. J. Exp. Bot. 2004, 55, 1607–1621. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef] [PubMed]
- Ehlert, B.; Hincha, D.K. Chlorophyll fluorescence imaging accurately quantifies freezing damage and cold acclimation responses in Arabidopsis leaves. Plant Methods 2008, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- IMAGE J. Available online: https://imagej.nih.gov/ij/ (accessed on 28 February 2018).
- Lobet, G.; Draye, X.; Perilleux, C. An online database for plant image analysis software tools. Plant Methods 2013, 9, 38. [Google Scholar] [CrossRef] [PubMed]
- Brewer, M.T.; Lang, L.; Fujimura, K.; Dujmovic, N.; Gray, S.; van der Knaap, E. Development of a controlled vocabulary and software application to analyse fruit shape variation in tomato and other plant species. Plant Physiol. 2006, 141, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Kamilaris, A.; Kartakoullis, A.; Prenafeta-Boldú, F.X. A review on the practice of big data analysis in agriculture. Comput. Electron. Agric. 2017, 143, 23–37. [Google Scholar] [CrossRef]
| Index | Common Abbreviation | Formula a | Crop | Reference |
|---|---|---|---|---|
| Green Normalized Difference Vegetation Index | GNDVI | cucumber, lettuce, tomato, onion | [70,71,77,80,81] | |
| Green Vegetation Index | GVI | cucumber, lettuce, tomato | [67,73,81,82,83] | |
| MERIS Terrestrial Chlorophyll Index | MTCI | cucumber, lettuce | [67,71] | |
| Normalized Difference Vegetation Index | NDVI | broccoli, cabbage, cucumber, leaf mustard, lettuce, onion, pepper, radish, spinach, tomato | [42,46,48,67,69,70,71,79,80,81,82,84,85] | |
| Red-Edge Index | REI | broccoli, onion | [79,80] | |
| Red-Edge Normalized Difference Vegetation Index | RENDVI | cucumber, tomato | [67,82] | |
| Red Vegetation Index | RVI | cucumber, lettuce, tomato | [46,67,71,81] | |
| Water Index | WI | lettuce, onion, tomato | [70,79,86] |
| Plant Species | Traits | Type of Analysis a | Instrument | Reference |
|---|---|---|---|---|
| Tomato and Eggplant | Alternaria solani and Epilachna vigintioctopunctata | NIR Hyperspectral imaging | ASD FieldSpec Pro FR spectrometer | [89] |
| Tomato | Rhizopus stolonifer spores | NIR Hyperspectral imaging | Homemade built | [90] |
| Tomato | Leaves damaged by leaf miner | NIR Hyperspectral imaging | Nexus FT-NIR spectrometer | [91] |
| Tomato | Surface defects detection | NIR Hyperspectral imaging | ImSpector V10 | [92] |
| Tomato | Ripeness | NIR Hyperspectral imaging | ImSpector V9 | [93] |
| Leafy vegetables | Chlorophyll content | NIR Hyperspectral imaging | ASD Fieldspec FR spectroradiometer | [78] |
| Spinach | Quality during storage | NIR Hyperspectral imaging | EMCCD Luca-R camera-Hyperspec® VNIR | [94] |
| Spinach | Crop canopy under water | NIR Hyperspectral imaging | Specim V10 spectrometer | [42] |
| Watermelon | Lycopene, -Carotene, and Total Soluble Solids | NIR Hyperspectral imaging | NIR On-Line® X-One | [95] |
| Lettuce | Plant traits under extreme temperature and salinity stress treatments | Hyperspectral and Fluorecence imaging | Series VNIR Micro-Hyperspec Sensor; Fluor Cam 800 MF | [96] |
| Tomato | Growing of truss | MRI, NMR | Homemade built | [97] |
| Tomato | Maturity | MRI | 1 T MR system | [98] |
| Cucumber | Internal freeze damage | MRI | 9.4 T MR system | [99] |
| Bean | Pod water content | NMR | Homemade built | [100] |
| Tomato | Qualitative | VIS/NIR | LabSpec 5000 | [101] |
| Tomato | Antioxidants | VIS/NIR | HandHeld 2™ | [102] |
| Tomato | Lycopene and physicochemical parameters | VIS/NIR | Varian Cary 500 | [103] |
| Tomato | Varietal discrimination | VIS/NIR | USB2000 spectrometer | [104] |
| Tomato | Transgenic lines discrimination | VIS/NIR | FT-NIR spectrometer | [105] |
| Tomato | Harvest time | VIS/NIR | AgroSpec VIS-NIR spectrophotometer | [106] |
| Tomato | Drought stress | Chlorophyll fluorescence | Handy FluorCam FC 1000-H system | [107] |
| Chicory | Cold stress | Chlorophyll fluorescence | CF Imager | [108,109] |
| Bean | Pseudomonas syringae infection | Chlorophyll fluorescence | Fluorcam | [110] |
| Bean | Botrytis infection, magnesium deficiency | Chlorophyll fluorescence and thermoimaging | Homemade built | [111] |
| Bean | Photosyntetic traits, morphological parameters and shoot architecture | PAM fluorimetry | Growscreen Fluoro | [100] |
| Chinese cabbage | Quality | PAM fluorimetry | FluorPenFP 100 fluorimeter | [112] |
| Melon | Grafting compatibility | PAM fluorimetry | Imaging-PAM fluorometer | [113] |
| Tomato | Fruit ripening | Chlorophyll and polyphenol content | Multiplex sensor | [114] |
| Leafy vegetables | quality in post-harvest storage | Chlorophyll content | SPAD-502; Agriexpert CCN 6000 | [115] |
| Pepper | Canopy and plant architecture | RGB | Homemade built | [116] |
| Tomato | Canopy | 3D imaging | Digital camera | [117] |
| Tomato, Eggplant | Fruit morphology and shape | Tomato analyzer | Scanner imaging | [118,119,120] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tripodi, P.; Massa, D.; Venezia, A.; Cardi, T. Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges. Agronomy 2018, 8, 57. https://doi.org/10.3390/agronomy8040057
Tripodi P, Massa D, Venezia A, Cardi T. Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges. Agronomy. 2018; 8(4):57. https://doi.org/10.3390/agronomy8040057
Chicago/Turabian StyleTripodi, Pasquale, Daniele Massa, Accursio Venezia, and Teodoro Cardi. 2018. "Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges" Agronomy 8, no. 4: 57. https://doi.org/10.3390/agronomy8040057
APA StyleTripodi, P., Massa, D., Venezia, A., & Cardi, T. (2018). Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges. Agronomy, 8(4), 57. https://doi.org/10.3390/agronomy8040057







