An Evaluation of the Variation in the Morphometric Parameters of Grain of Six Triticum Species with the Use of Digital Image Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Image Analysis
2.2. Shape Analysis
2.3. Color Analysis
2.4. Fungal Colonization of Grain
2.5. Statistical Analysis
3. Results
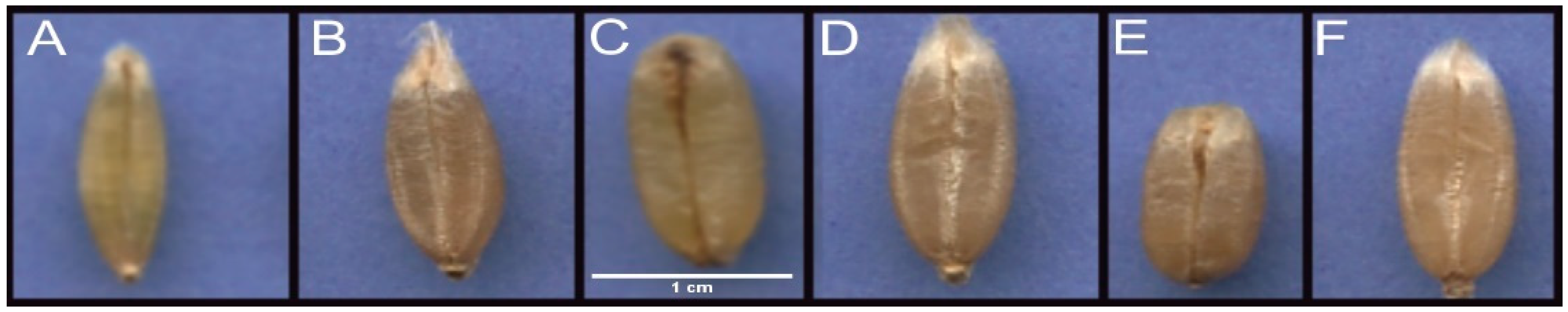
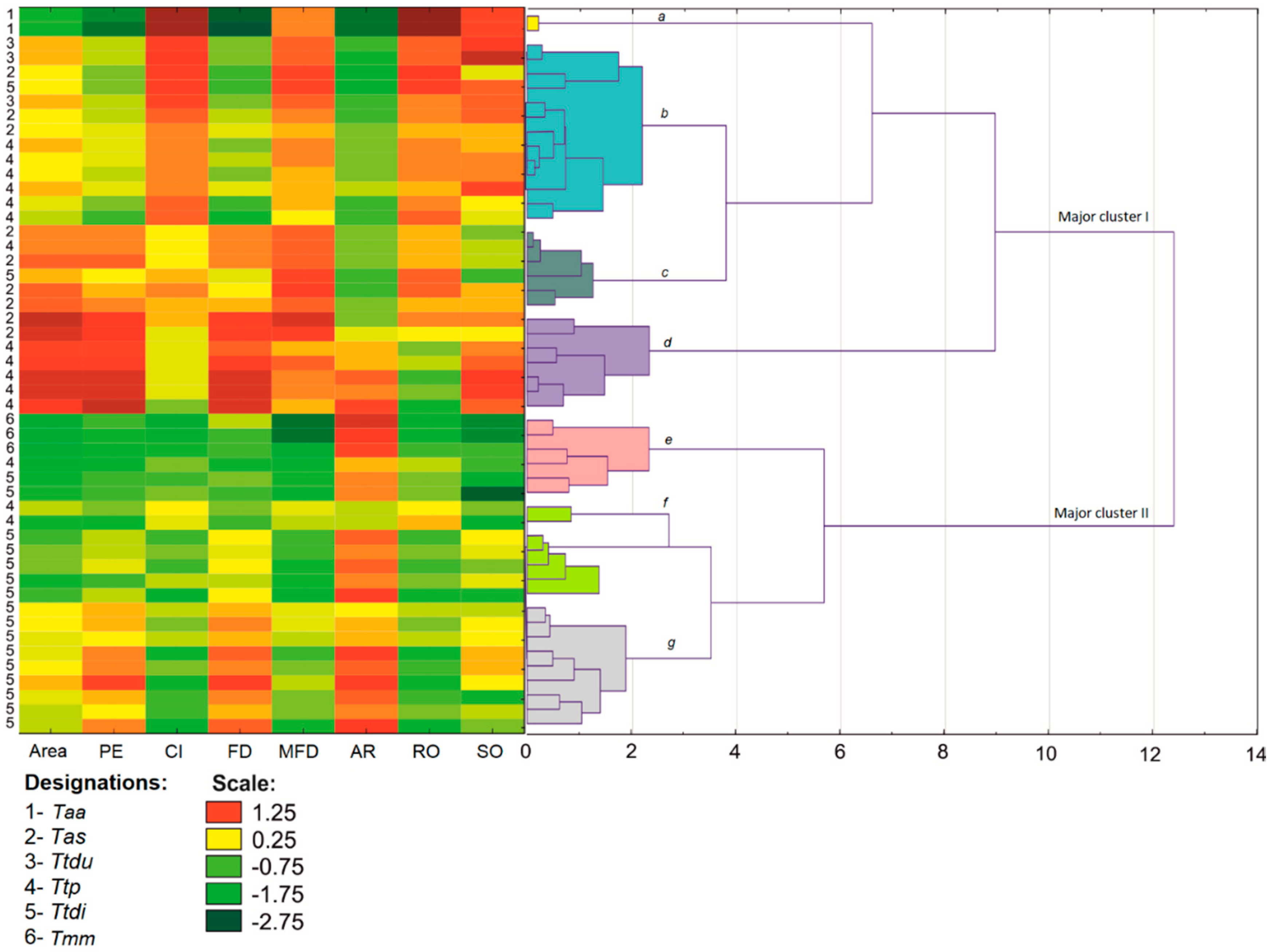
3.1. Shape Analysis
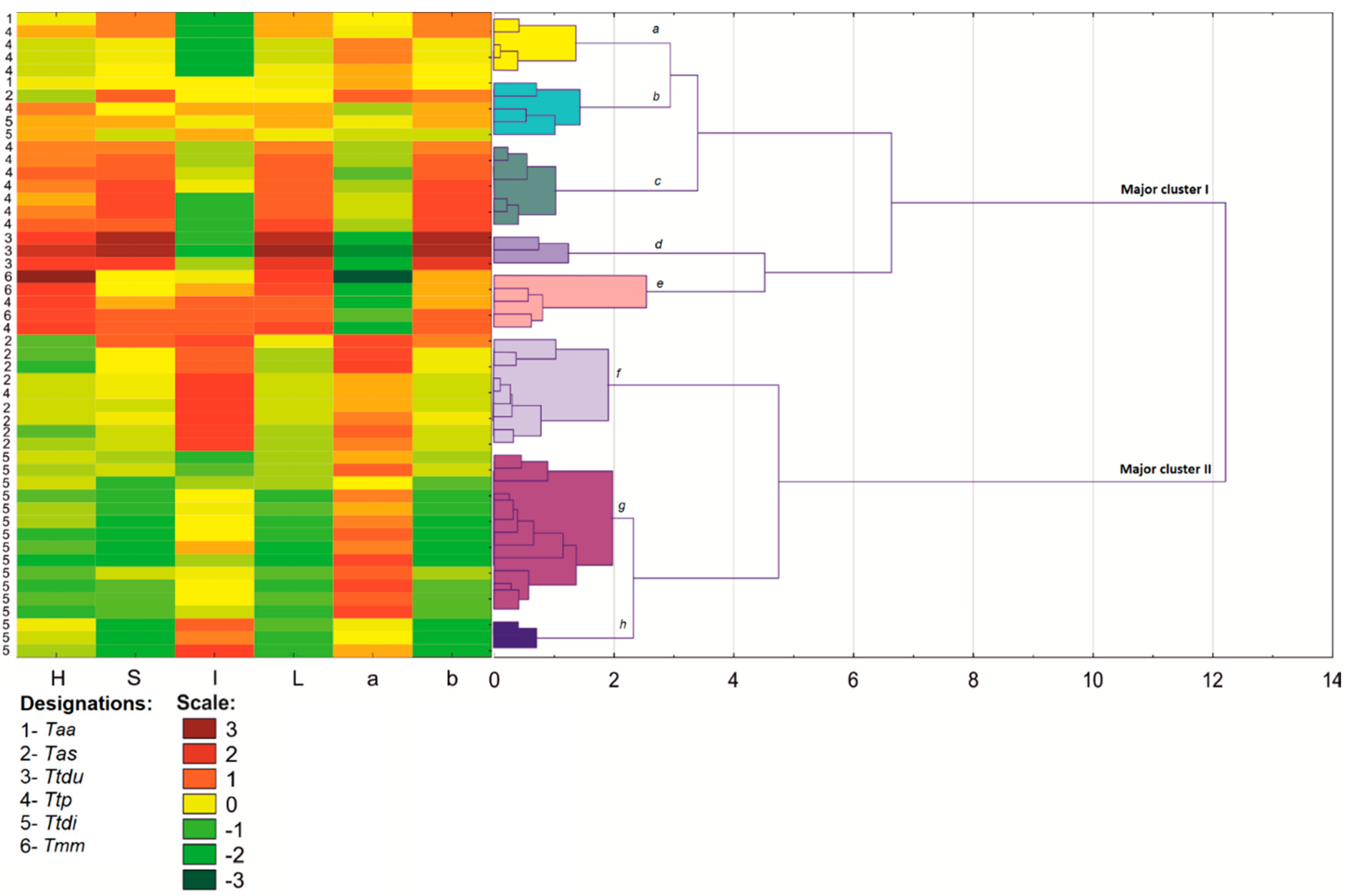
3.2. Color Analysis
3.3. Correlation Analysis
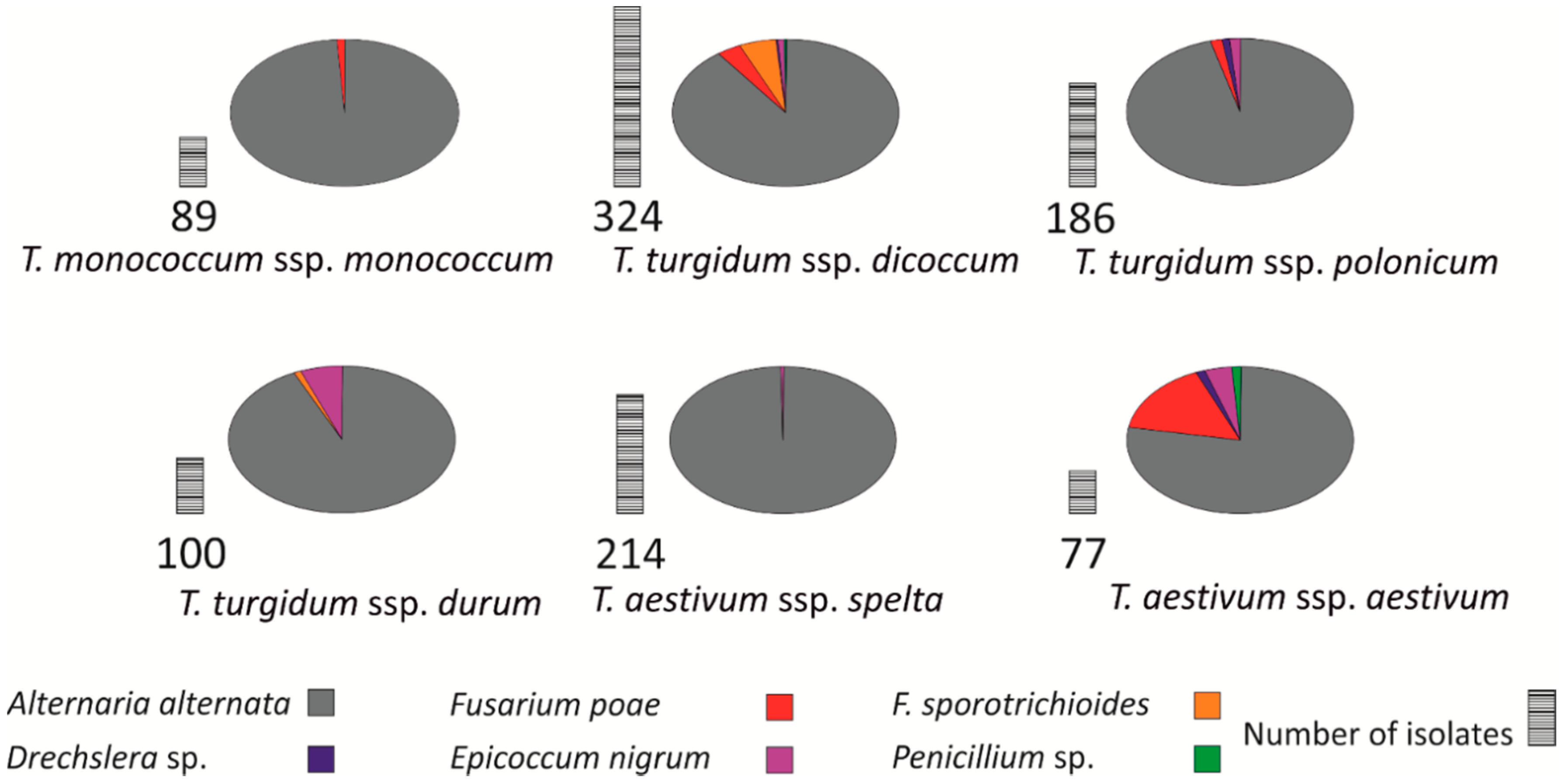
3.4. Identification of Fungi
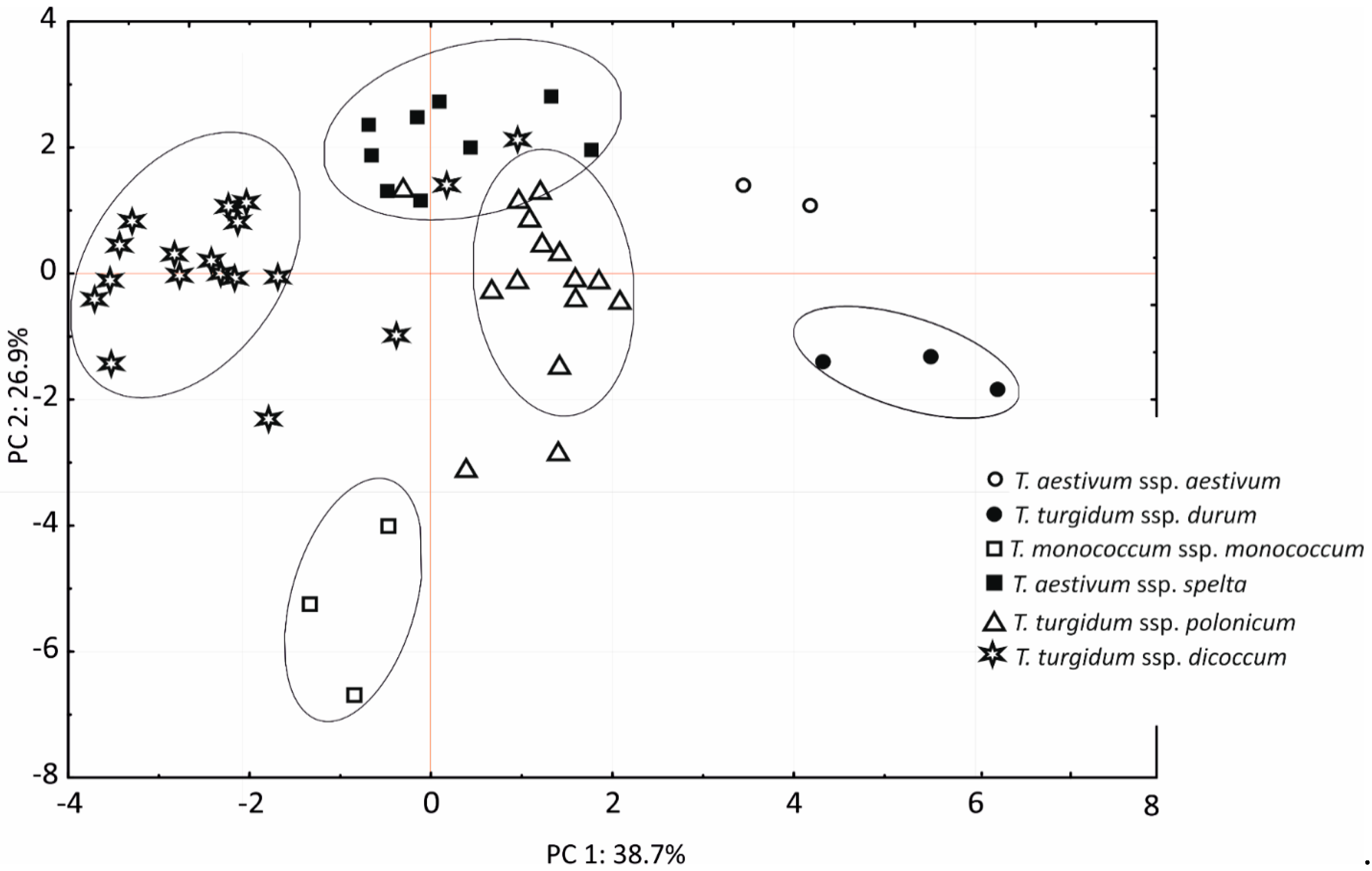
3.5. Principal Component Analysis
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Maccaferri, M.; Mantovani, P.; Tuberosa, R.; DeAmbrogio, E.; Giuliani Demontis, A.; Massi, A.; Sanguineti, M.C. A major QTL for durable leaf rust resistance widely exploited in durum wheat breeding programs maps on the distal region of chromosome arm 7BL. Theor. Appl. Genet. 2008, 117, 1225–1240. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, C.; Bolla, M.; Chignola, R.; Senna, G.; Rossin, G.; Caruso, B.; Tomelleri, C.; Cecconi, D.; Brandolini, A.; Zoccatelli, G. Study on the immunoreactivity of Triticum monococcum (Einkorn) wheat in patients with wheat-dependent exercise-induced anaphylaxis for the production of hypoallergenic foods. J. Agric. Food Chem. 2015, 63, 8299–8306. [Google Scholar] [CrossRef] [PubMed]
- Henkrar, F.; El-Haddoury, J.; Ouabbou, H.; Nsarellah, N.; Iraqi, D.; Bendaou, N.; Udupa, S.M. Genetic diversity and its temporal changes in improved bread wheat cultivars of Morocco. Rom. Agric. Res. 2015, 32, 19–25. [Google Scholar]
- Moragues, M.; Moralejo, M.; Sorrells, M.E.; Royo, C. Dispersal of durum wheat [Triticum turgidum L. ssp. turgidum convar. durum (Desf.) MacKey] landraces across the Mediterranean basin assessed by AFLPs and microsatellites. Genet. Resour. Crop Evol. 2007, 54, 1133–1144. [Google Scholar] [CrossRef]
- Figliuolo, G.; Perrino, P. Genetic diversity and intra-specific phylogeny of Triticum turgidum L. subsp. dicoccon (Schrank) Thell. revealed by RFLPs and SSRs. Genet. Resour. Crop Evol. 2004, 51, 519–527. [Google Scholar] [CrossRef]
- Jing, H.C.; Bayon, C.; Kanyuka, K.; Berry, S.; Wenzl, P.; Huttner, E.; Kilia, A.; Hammond-Kosack, K.E. DArT markers: Diversity analyses, genomes comparison, mapping and integration with SSR markers in Triticum monococcum. BMC Genom. 2009, 10, 458. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, M.R.; Sissons, M.; Conner, R.L.; Wang, H.; Clarke, J.M. Influence of biotic and abiotic factors on dark discoloration of durum wheat kernels. Crop Sci. 2011, 51, 1205–1214. [Google Scholar] [CrossRef]
- Wang, H.; Fernandez, M.R.; McCaig, T.N.; Gan, Y.T.; DePauw, R.M.; Clarke, J.M. Kernel discoloration and downgrading in spring wheat varieties in western Canada. Can. J. Plant Pathol. 2003, 25, 350–361. [Google Scholar] [CrossRef]
- Fares, C.; Codianni, P.; Nigro, F.; Platani, C.; Scazzina, F.; Pellegrini, N. Processing and cooking effects on chemical, nutritional and functional properties of pasta obtained from selected emmer genotypes. J. Sci. Food. Agric. 2008, 88, 2435–2444. [Google Scholar] [CrossRef]
- Hidalgo, A.; Brandolini, A.; Gazza, L. Influence of steaming treatment on chemical and technological characteristics of einkorn (Triticum monococcum L. ssp. monococcum) wholemeal flour. Food Chem. 2008, 111, 549–555. [Google Scholar] [CrossRef]
- Kucek, L.K.; Dyck, E.; Russell, J.; Clark, L.; Hamelman, J.; Burns-Leader, S.; Senders, S.; Jones, J.; Benscher, D.; Davis, M.; et al. Evaluation of wheat and emmer varieties for artisanal baking, pasta making, and sensory quality. J. Cereal Sci. 2017, 74, 19–27. [Google Scholar] [CrossRef]
- European Commission. Commission Regulation (EC) No 2073/2005 on microbiological criteria for foodstuffs. Off. J. Eur. Union 2005, 50, 1–26. [Google Scholar]
- Khatri, Y.; Collins, R. Impact and status of HACCP in the Australian meat industry. Br. Food J. 2007, 109, 343–354. [Google Scholar] [CrossRef]
- Narendra, V.G.; Hareesh, K. Prospects of computer vision automated grading and sorting systems in agricultural and food products for quality evaluation. Int. J. Comput. Appl. 2010, 1, 651–657. [Google Scholar] [CrossRef]
- CBH Group. 2018. Available online: https://cbh.com.au/other%20information/quality%20services/eyefoss (accessed on 1 July 2018).
- Sun, Q.; Wu, L.; Ni, Z.; Meng, F.; Wang, Z.; Lin, Z. Differential gene expression patterns in leaves between hybrids and their parental inbreds are correlated with heterosis in a wheat diallel cross. Plant Sci. 2004, 166, 651–657. [Google Scholar] [CrossRef]
- Scanalyzer HTS: LemnaTec GmbH, Germany. 2018. Available online: https://www.lemnatec.com/products/laboratory/lab-scanalyzer-hts/ (accessed on 1 July 2018).
- Neuman, M.; Sapirstein, H.D.; Shwedyk, E.; Bushuk, W. Discrimination of wheat class and variety by digital image analysis of whole grain samples. J. Cereal Sci. 1987, 6, 125–132. [Google Scholar] [CrossRef]
- Neuman, M.R.; Sapirstein, H.D.; Shwedyk, E.; Bushuk, W. Wheat grain colour analysis by digital image processing II. Wheat class discrimination. J. Cereal Sci. 1989, 10, 183–188. [Google Scholar] [CrossRef]
- Zayas, I.; Pomeranz, Y.; Lai, F.S. Discrimination of wheat and nonwheat components in grain samples by image analysis. Cereal Chem. 1989, 66, 233–237. [Google Scholar]
- Mirik, M.; Michels, G.J., Jr.; Kassymzhanova-Mirik, S.; Elliott, N.C.; Catana, V.; Jones, D.B.; Bowling, R. Using digital image analysis and spectral reflectance data to quantify damage by greenbug (Hemitera: Aphididae) in winter wheat. Comput. Electron. Agric. 2006, 51, 86–98. [Google Scholar] [CrossRef]
- Yan, X.; Wang, J.; Liu, S.; Zhang, C. Purity Identification of Maize Seed Based on Color Characteristics. In Computer and Computing Technologies in Agriculture IV, Proceedings of the International Conference on Computer and Computing Technologies in Agriculture, Nanchang, China, 22–25 October 2010; Li, D., Liu, Y., Chen, Y., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; Volume 346, ISBN 978-3-642-18354-6. [Google Scholar]
- Diéguez-Uribeondo, J.; Förster, H.; Adaskaveg, J.E. Digital image analysis of internal light spots of appressoria of Colletotrichum acutatum. Phytopathology 2003, 93, 923–930. [Google Scholar] [CrossRef]
- Wijekoon, C.P.; Goodwin, P.H.; Hsiang, T. Quantifying fungal infection of plant leaves by digital image analysis using Scion Image software. J. Microbiol. Meth. 2008, 74, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Suchowilska, E.; Wiwart, M. Multivariate analysis of image descriptors of common wheat (Triticum aestivum) and spelt (T. spelta) grain infected by Fusarium culmorum. Int. Agrophys. 2006, 20, 345. [Google Scholar]
- Bauriegel, E.; Giebel, A.; Geyer, M.; Schmidt, U.; Herppich, W.B. Early detection of Fusarium infection in wheat using hyper-spectral imaging. Comput. Electron. Agric. 2011, 75, 304–312. [Google Scholar] [CrossRef]
- Ahmad, I.S.; Reid, J.F.; Paulsen, M.R.; Sinclair, J.B. Color classifier for symptomatic soybean seeds using image processing. Plant Dis. 1999, 83, 320–327. [Google Scholar] [CrossRef]
- Leplat, J.; Mangin, P.; Falchetto, L.; Heraud, C.; Gautheron, E.; Steinberg, C. Visual assessment and computer-assisted image analysis of Fusarium head blight in the field to predict mycotoxin accumulation in wheat grains. Eur. J. Plant Pathol. 2018, 150, 1065–1081. [Google Scholar] [CrossRef]
- Zapotoczny, P. Discrimination of wheat grain varieties using image analysis and neural networks. Part I. Single kernel texture. J. Cereal Sci. 2011, 54, 60–68. [Google Scholar] [CrossRef]
- Wiwart, M.; Suchowilska, E.; Lajszner, W.; Graban, L. Identification of hybrids of spelt and wheat and their parental forms using shape and color descriptors. Comput. Electron. Agric. 2012, 83, 68–76. [Google Scholar] [CrossRef]
- Zielinska, M.; Zapotoczny, P.; Białobrzewski, I.; Zuk-Gołaszewska, K.; Markowski, M. Engineering properties of red clover (Trifolium pratense L.) seeds. Ind. Crop. Prod. 2012, 37, 69–75. [Google Scholar] [CrossRef]
- Ropelewska, E.; Zapotoczny, P.; Budzyński, W.S.; Jankowski, K.J. Discriminating power of selected physical properties of seeds of various rapeseed (Brassica napus L.) cultivars. J. Cereal Sci. 2017, 73, 62–67. [Google Scholar] [CrossRef]
- Chaugule, A.A.; Mali, S.N. Identification of paddy varieties based on novel seed angle features. Comput. Electron. Agric. 2016, 123, 415–422. [Google Scholar] [CrossRef]
- Cervantes, E.; Martín, J.J.; Saadaoui, E. Updated Methods for Seed Shape Analysis. Scientifica 2016. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y. Hopfield neural network based algorithms for image restoration and reconstruction. I. Algorithms and simulations. IEEE Trans. Signal Process. 2000, 48, 2105–2118. [Google Scholar] [CrossRef]
- Russ, J.C. Segmentation and thresholding. In The Image Processing Handbook, 6th ed.; CRC, Taylor & Francis Group: New York, NY, USA, 2016; pp. 395–443. ISBN 9781439840634. [Google Scholar]
- Szczypiński, P.M.; Klepaczko, A.; Kociołek, M. QMaZda—Software tools for image analysis and pattern recognition. In Proceedings of the 2017 Signal Processing: Algorithms, Architectures, Arrangements, and Applications (SPA), Poznan, Poland, 20–22 October 2017; pp. 217–221. [Google Scholar]
- Bock, C.H.; Poole, G.H.; Parker, P.E.; Gottwald, T.R. Plant Disease Severity Estimated Visually, by Digital Photography and Image Analysis, and by Hyperspectral Imaging. Crit. Rev. Plant Sci. 2010, 29, 59–107. [Google Scholar] [CrossRef]
- Cope, P. The Digital Photographer’s Pocket Encyclopedia: 3000 Terms Explained; Silver Pixel Press: Rochester, NY, USA, 2002; ISBN 1883403901. [Google Scholar]
- Steddom, K.; Bredehoeft, M.W.; Khan, M.; Rush, C.M. Comparison of visual and multispectral radiometric disease evaluations of Cercospora leaf spot of sugar beet. Plant Dis. 2005, 89, 153–158. [Google Scholar] [CrossRef]
- Rasband, W.S. ImageJ; U.S. National Institutes of Health: Bethesda, MD, USA, 2016. Available online: https://imagej.nih.gov/ij/ (accessed on 1 July 2018).
- Wiwart, M.; Fordoński, G.; Żuk-Gołaszewska, K.; Suchowilska, E. Early diagnostics of macronutrient deficiencies in three legume species by color image analysis. Comput. Electron. Agric. 2009, 65, 125–132. [Google Scholar] [CrossRef]
- Follstad, M.N.; Christensen, C.M. Microflora of barley kernels. Appl. Microbiol. 1962, 10, 331–336. [Google Scholar] [PubMed]
- Ellis, M.B.; Ellis, J.P. (Eds.) Microfungi on Land Plants: An Identification Handbook; Richmond Publishing: Slough, UK, 1987. [Google Scholar]
- Leslie, J.F.; Summerell, B.A. (Eds.) The Fusarium Laboratory Manual; Wiley-Blackwell Publishing: Ames, IA, USA, 2006. [Google Scholar]
- StatSoft. STATISTICA (Data Analysis Software System); Version 12; StatSoft, Inc.: Tulsa, OK, USA, 2014; Available online: www.statsoft.com (accessed on 1 July 2018).
- Fuller, D.Q. Contrasting patterns in crop domestication and domestication rates: Recent archaeobotanical insights from the Old World. Ann. Bot. 2007, 100, 903–924. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, Y. Evolution of polyploid Triticum wheats under cultivation: The role of domestication, natural hybridization and allopolyploid speciation in their diversification. Plant Cell Physiol. 2011, 52, 750–764. [Google Scholar] [CrossRef] [PubMed]
- Bakhteyev, F.K.; Yanushevich, Z.V. Discoveries of cultivated plants in the early farming settlements of Yarym-Tepe I and Yarym-Tepe II in northern Iraq. J. Archaeol. Sci. 1980, 7, 167–178. [Google Scholar] [CrossRef]
- Jacomet, S. Identification of Cereal Remains from Archaeological Sites. 2006. Available online: http://arkeobotanika.pbworks.com/f/Jacomet+cereal+ID.pdf (accessed on 1 July 2018).
- Okamoto, Y.; Takumi, S. Pleiotropic effects of the elongated glume gene P1 on grain and spikelet shape-related traits in tetraploid wheat. Euphytica 2013, 194, 207–218. [Google Scholar] [CrossRef]
- Gegas, V.C.; Nazari, A.; Griffiths, S.; Simmonds, J.; Fish, L.; Orford, S.; Sayers, L.; Doonan, J.H.; Snape, J.W. A genetic framework for grain size and shape variation in wheat. Plant Cell 2010, 22, 1046–1056. [Google Scholar] [CrossRef] [PubMed]
- Eticha, F.; Belay, G.; Bekele, E. Species diversity in wheat landrace populations from two regions of Ethiopia. Genet. Resour. Crop. Evol. 2006, 53, 387–393. [Google Scholar] [CrossRef]
- Cifci, E.A.; Yagdi, K. Study of genetic diversity in wheat (Triticum aestivum) varieties using random amplified polymorphic DNA (RAPD) analysis. Turk. J. Field Crops 2012, 17, 91–95. [Google Scholar]
- Evers, A.D.; Cox, R.I.; Shaheedullah, M.Z.; Withey, R.P. Predicting milling extraction rate by image analysis of wheat grains. Asp. Appl. Biol. 1990, 25, 417–426. [Google Scholar]
- Zhang, C.; Si, Y.; Lamkey, J.; Boydston, R.A.; Garland-Campbell, K.A.; Sankaran, S. High-Throughput Phenotyping of Seed/Seedling Evaluation Using Digital Image Analysis. Agronomy 2018, 8, 63. [Google Scholar] [CrossRef]
- Williams, K.; Munkvold, J.; Sorrells, M. Comparison of digital image analysis using elliptic Fourier descriptors and major dimensions to phenotype seed shape in hexaploid wheat (Triticum aestivum L.). Euphytica 2013, 190, 99–116. [Google Scholar] [CrossRef]
- Breseghello, F.; Sorrells, M.E. Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 2006, 172, 1165–1177. [Google Scholar] [CrossRef]
- Breseghello, F.; Sorrells, M.E. QTL analysis of kernel size and shape in two hexaploid wheat mapping populations. Field Crops Res. 2007, 101, 172–179. [Google Scholar] [CrossRef]
- Dana, W.; Ivo, W. Computer image analysis of seed shape and seed color for flax cultivar description. Comput. Electron. Agric. 2008, 61, 126–135. [Google Scholar] [CrossRef]
- Jamil, M.; Ali, A.; Ghafoor, A.; Akbar, K.F.; Napar, A.A.; Naveed, N.H.; Yasin, N.A.; Gul, A.; Mujeeb-Kazi, A. Digital image analysis of seed shape influenced by heat stress in diverse bread wheat germplasm. Pak. J. Bot. 2017, 49, 1279–1284. [Google Scholar]
- Suchowilska, E.; Kandler, W.; Sulyok, M.; Wiwart, M.; Krska, R. Mycotoxin profiles in the grain of Triticum monococcum, Triticum dicoccum and Triticum spelta after head infection with Fusarium culmorum. J. Sci. Food Agric. 2010, 90, 556–565. [Google Scholar] [PubMed]
- Oliver, R.E.; Cai, X.; Friesen, T.L.; Halley, S.; Stack, R.W.; Xu, S.S. Evaluation of Fusarium head blight resistance in tetraploid wheat (Triticum turgidum L.). Crops Sci. 2008, 48, 213–222. [Google Scholar] [CrossRef]
- Shen, Y.; Jin, L.; Xiao, P.; Lu, Y.; Bao, J. Total phenolics, flavonoids, antioxidant capacity in rice grain and their relations to grain color, size and weight. J. Cereal Sci. 2009, 49, 106–111. [Google Scholar] [CrossRef]
- Burešová, V.; Kopecký, D.; Bartoš, J.; Martinek, P.; Watanabe, N.; Vyhnánek, T.; Doležel, J. Variation in genome composition of blue-aleurone wheat. Theor. Appl. Genet. 2015, 128, 273–282. [Google Scholar] [CrossRef]

| Descriptor | Characteristics | Equation |
|---|---|---|
| Shape | ||
| Area | The quantity that expresses the extent of a two-dimensional figure or shape | n/a |
| Perimeter (PE) | A path that surrounds a two-dimensional shape | n/a |
| Circularity (CI) | Shape factor, takes on values in a range from 0 (elongated shape) to 1 (perfect circle) | |
| Feret Diameter (FD) | The longest distance between any two pixels along the selection boundary | n/a |
| Minimal Feret diameter (MFD) | The shortest distance between any two pixels along the selection boundary | n/a |
| Aspect ratio (AR) | Aspect ratio of the blob’s fitted ellipse. Major and minor axis refer to the ellipse fitted to region of interest (ROI) | |
| Roundness (RO) | Describes shapes conversely to AR | |
| Solidity (SO) | Describes ROI with regard to shape regularity. The Convex Area refers to the area of the convex hull of the region (the smallest region that is convex and that contains the original ROI). The value of this descriptor is lower for objects with an irregular shape, and higher for objects with a regular shape | |
| Color | ||
| Hue (H) | Refers to the attribute of visible light due to that is differentiated from or similar to red, green or blue | Formulas proposed by Wiwart et al. [42] |
| Saturation (S) | Proportion of pure chromatic color in the total color sensation | |
| Intensity (I) | Refers to the amount of light or the numerical value of a pixel | |
| Luminance (L*) | Measures perceived “gray-level” of pixel | |
| a* | Denotes redness–greenness | |
| b* | Denotes yellowness–blueness | |
| SKW (mg) | |||||
|---|---|---|---|---|---|
| Taa | Tas | Ttdu | Ttp | Ttdi | Tmm |
| 42.13 bc | 49.69 b | 61.00 a | 46.92 b | 39.84 c | 27.29 d |
| MV | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Area (mm2) | PE (mm) | CI | ||||||||||||||||
| Mean | 15.24 bc | 20.54 a | 19.55 a | 19.68 a | 17.26 b | 12.89 c | 15.80 c | 20.06 a | 18.81 ab | 19.93 a | 19.26 a | 17.52 b c | 0.76 a | 0.64 c | 0.69 b | 0.62 c | 0.58 d | 0.53 e |
| SD | 0.77 | 2.02 | 0.93 | 3.04 | 1.56 | 0.39 | 0.37 | 1.22 | 0.36 | 1.77 | 0.90 | 0.38 | 0.01 | 0.02 | 0.01 | 0.03 | 0.04 | 0.02 |
| RSD (%) | 5.04 | 9.84 | 4.75 | 15.42 | 9.07 | 3.06 | 2.37 | 6.10 | 1.91 | 8.89 | 4.66 | 2.15 | 0.83 | 4.15 | 1.92 | 4.86 | 6.64 | 4.23 |
| Min | 7.58 | 11.42 | 9.50 | 9.22 | 7.74 | 8.73 | 11.73 | 14.30 | 14.28 | 13.65 | 11.97 | 14.06 | 0.56 | 0.36 | 0.49 | 0.34 | 0.32 | 0.24 |
| Max | 20.29 | 31.10 | 26.80 | 35.50 | 50.67 | 20.07 | 18.34 | 31.20 | 22.18 | 32.63 | 41.74 | 27.62 | 0.82 | 0.80 | 0.76 | 0.76 | 1.00 | 0.66 |
| FD (mm) | MFD (mm) | AR | ||||||||||||||||
| Mean | 6.12 d | 8.37 a | 7.66 bc | 8.26 ab | 8.27 a | 7.6 c | 3.24 ab | 3.35 a | 3.32 ab | 3.16 b | 2.89 c | 2.43 d | 1.88 e | 2.42 d | 2.35 d | 2.62 c | 2.86 b | 3.16 a |
| SD | 0.14 | 0.54 | 0.13 | 0.83 | 0.48 | 0.20 | 0.08 | 0.14 | 0.09 | 0.16 | 0.23 | 0.07 | 0.02 | 0.13 | 0.05 | 0.23 | 0.26 | 0.14 |
| RSD (%) | 2.34 | 6.50 | 1.67 | 10.00 | 5.85 | 2.61 | 2.39 | 4.11 | 2.71 | 5.08 | 7.81 | 2.78 | 1.07 | 5.43 | 2.30 | 8.74 | 9.22 | 4.52 |
| Min | 4.76 | 5.35 | 5.95 | 5.65 | 4.64 | 5.92 | 1.89 | 2.17 | 1.87 | 1.90 | 1.69 | 1.89 | 1.41 | 1.73 | 1.91 | 1.05 | 1.43 | 2.26 |
| Max | 7.25 | 10.51 | 8.84 | 12.71 | 11.18 | 9.18 | 4.22 | 4.58 | 4.33 | 6.71 | 8.16 | 3.42 | 3.19 | 3.46 | 4.04 | 5.24 | 4.84 | 3.93 |
| RO | SO | |||||||||||||||||
| Mean | 0.54 a | 0.41 b | 0.43 b | 0.39 c | 0.36 d | 0.32 e | 0.963 ab | 0.958 b c | 0.970 a | 0.959 b | 0.950 c | 0.947 d | ||||||
| SD | 0.01 | 0.02 | 0.01 | 0.03 | 0.04 | 0.02 | <0.01 | <0.01 | <0.01 | 0.01 | 0.01 | <0.01 | ||||||
| RSD (%) | 1.04 | 5.78 | 1.97 | 8.34 | 10.28 | 4.88 | 0.21 | 0.30 | 0.37 | 0.55 | 0.53 | 0.43 | ||||||
| Min | 0.31 | 0.29 | 0.25 | 0.19 | 0.21 | 0.25 | 0.92 | 0.88 | 0.92 | 0.75 | 0.80 | 0.82 | ||||||
| Max | 0.71 | 0.58 | 0.52 | 0.96 | 1.00 | 0.44 | 0.97 | 0.97 | 0.98 | 0.98 | 1.00 | 0.96 | ||||||
| MV | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm | Taa | Tas | Ttdu | Ttp | Ttdi | Tmm |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| H | S | I | ||||||||||||||||
| Mean | 35.37 bc | 33.94 c | 40.36 a | 36.79 b | 34.05 c | 40.71 a | 0.21 bc | 0.20 c | 0.25 a | 0.21 b | 0.18 d | 0.21 bc | 0.48 cd | 0.52 a | 0.47 d | 0.48 c | 0.50 b | 0.50 b |
| SD | 1.09 | 0.96 | 1.70 | 1.86 | 1.11 | 1.85 | 0.02 | 0.01 | 0.02 | 0.04 | 0.01 | 0.02 | 0.03 | 0.02 | 0.02 | 0.03 | 0.02 | 0.02 |
| RSD (%) | 3.08 | 2.82 | 4.21 | 5.00 | 3.26 | 4.56 | 10.82 | 6.97 | 7.06 | 15.09 | 7.79 | 8.89 | 5.81 | 3.98 | 4.88 | 5.47 | 4.09 | 4.05 |
| Min | 29.29 | 28.96 | 33.58 | 13.29 | 5.53 | 34.87 | 0.13 | 0.13 | 0.19 | 0.13 | 0.07 | 0.15 | 0.40 | 0.42 | 0.38 | 0.36 | 0.39 | 0.43 |
| Max | 38.28 | 88.76 | 52.73 | 64.93 | 118.6 | 50.32 | 0.27 | 0.27 | 0.33 | 0.30 | 0.28 | 0.27 | 0.59 | 0.61 | 0.54 | 0.62 | 0.62 | 0.58 |
| L* | a* | b* | ||||||||||||||||
| Mean | 65.50 cd | 65.39 d | 65.99 a | 65.59 c | 65.3 e | 65.75 b | −4.82 ab | −4.52 a | −6.03 c | −5.16 b | −4.69 a | −6.04 c | 6.96 bc | 6.38 c | 9.51 a | 7.19 b | 5.26 d | 7.20 b |
| SD | 0.11 | 0.08 | 0.15 | 0.39 | 0.08 | 0.12 | 0.32 | 0.25 | 0.45 | 0.71 | 0.25 | 0.39 | 1.22 | 0.75 | 1.04 | 0.91 | 0.75 | 1.02 |
| RSD (%) | 0.17 | 0.12 | 0.23 | 0.60 | 0.12 | 0.19 | 6.55 | 5.56 | 7.47 | 14.94 | 5.38 | 6.53 | 17.49 | 11.72 | 10.9 | 13.68 | 14.34 | 14.21 |
| Min | 65.10 | 65.13 | 65.51 | 61.2 | 61.58 | 65.32 | −5.59 | −12.18 | −8.73 | −12.00 | −10.26 | −7.82 | 2.80 | 3.74 | 6.03 | −9.42 | −16.36 | 3.91 |
| Max | 65.85 | 66.53 | 66.58 | 67.25 | 66.74 | 66.16 | −3.17 | −3.10 | −4.25 | 9.06 | 2.34 | −4.55 | 9.96 | 10.20 | 14.11 | 11.66 | 10.61 | 10.18 |
| DS | SKW | Area | PE | CI | FD | MFD | AR | RO | SO | H | S | I | L* | a* |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Area | 0.537 ** | |||||||||||||
| PE | 0.341 | 0.862 ** | ||||||||||||
| CI | 0.376 ** | 0.250 | −0.269 | |||||||||||
| FD | 0.192 | 0.696 ** | 0.960 ** | −0.510 ** | ||||||||||
| MFD | 0.583 ** | 0.793 ** | 0.401 ** | 0.743 ** | 0.152 | |||||||||
| AR | −0.364 ** | −0.180 | 0.320 * | −0.958 ** | 0.546 ** | −0.734 ** | ||||||||
| RO | 0.306 | 0.106 | −0.390 ** | 0.959 ** | −0.610 ** | 0.674 ** | −0.985 ** | |||||||
| SO | 0.415 ** | 0.640 ** | 0.324 * | 0.629 ** | 0.143 | 0.694 ** | −0.459 ** | 0.437 ** | ||||||
| H | −0.300 | −0.113 | −0.161 | 0.016 | −0.179 | −0.134 | 0.085 | −0.052 | −0.015 | |||||
| S | 0.053 | 0.259 | −0.030 | 0.475 ** | −0.164 | 0.366 ** | −0.354 ** | 0.342 * | 0.464 ** | 0.679 ** | ||||
| I | −0.094 | −0.040 | 0.084 | −0.234 | 0.151 | −0.072 | 0.107 | −0.121 | −0.364 ** | −0.225 | −0.361 ** | |||
| L* | −0.147 | 0.080 | −0.098 | 0.256 | −0.176 | 0.118 | −0.131 | 0.141 | 0.250 | 0.920 ** | 0.909 ** | −0.328 * | ||
| a* | 0.356 ** | 0.158 | 0.169 | 0.047 | 0.166 | 0.197 | −0.143 | 0.111 | 0.059 | −0.989 ** | −0.601 ** | 0.208 | −0.879 ** | |
| b* | 0.016 | 0.228 | −0.041 | 0.433 ** | −0.165 | 0.318 * | −0.307 * | 0.300 * | 0.428 ** | 0.743 ** | 0.995 ** | −0.367 ** | 0.944 ** | −0.673 ** |
| Variable | r | Variable Contribution | ||
|---|---|---|---|---|
| PC 1 | PC 2 | PC 1 | PC 2 | |
| Area | 0.279 * | 0.611 ** | 0.014 | 0.099 |
| PE | −0.166 | 0.402 ** | 0.005 | 0.043 |
| CI | 0.805 ** | 0.457 ** | 0.120 | 0.056 |
| FD | −0.378 ** | 0.252 | 0.026 | 0.017 |
| MFD | 0.603 ** | 0.731 ** | 0.067 | 0.142 |
| AR | −0.703 ** | −0.501 ** | 0.091 | 0.067 |
| RO | 0.702 ** | 0.449 ** | 0.091 | 0.053 |
| SO | 0.613 ** | 0.501 ** | 0.069 | 0.067 |
| H | 0.542 ** | −0.760 ** | 0.054 | 0.153 |
| S | 0.877 ** | −0.264 | 0.142 | 0.019 |
| I | −0.408 ** | 0.122 | 0.031 | 0.004 |
| L* | 0.768 ** | −0.567 ** | 0.109 | 0.085 |
| a* | −0.472 ** | 0.790 ** | 0.041 | 0.166 |
| b* | 0.867 ** | −0.335 * | 0.139 | 0.030 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goriewa-Duba, K.; Duba, A.; Wachowska, U.; Wiwart, M. An Evaluation of the Variation in the Morphometric Parameters of Grain of Six Triticum Species with the Use of Digital Image Analysis. Agronomy 2018, 8, 296. https://doi.org/10.3390/agronomy8120296
Goriewa-Duba K, Duba A, Wachowska U, Wiwart M. An Evaluation of the Variation in the Morphometric Parameters of Grain of Six Triticum Species with the Use of Digital Image Analysis. Agronomy. 2018; 8(12):296. https://doi.org/10.3390/agronomy8120296
Chicago/Turabian StyleGoriewa-Duba, Klaudia, Adrian Duba, Urszula Wachowska, and Marian Wiwart. 2018. "An Evaluation of the Variation in the Morphometric Parameters of Grain of Six Triticum Species with the Use of Digital Image Analysis" Agronomy 8, no. 12: 296. https://doi.org/10.3390/agronomy8120296
APA StyleGoriewa-Duba, K., Duba, A., Wachowska, U., & Wiwart, M. (2018). An Evaluation of the Variation in the Morphometric Parameters of Grain of Six Triticum Species with the Use of Digital Image Analysis. Agronomy, 8(12), 296. https://doi.org/10.3390/agronomy8120296




