Effect of Constitutive miR164 Expression on Plant Morphology and Fruit Development in Arabidopsis and Tomato
Abstract
:1. Introduction
2. Results
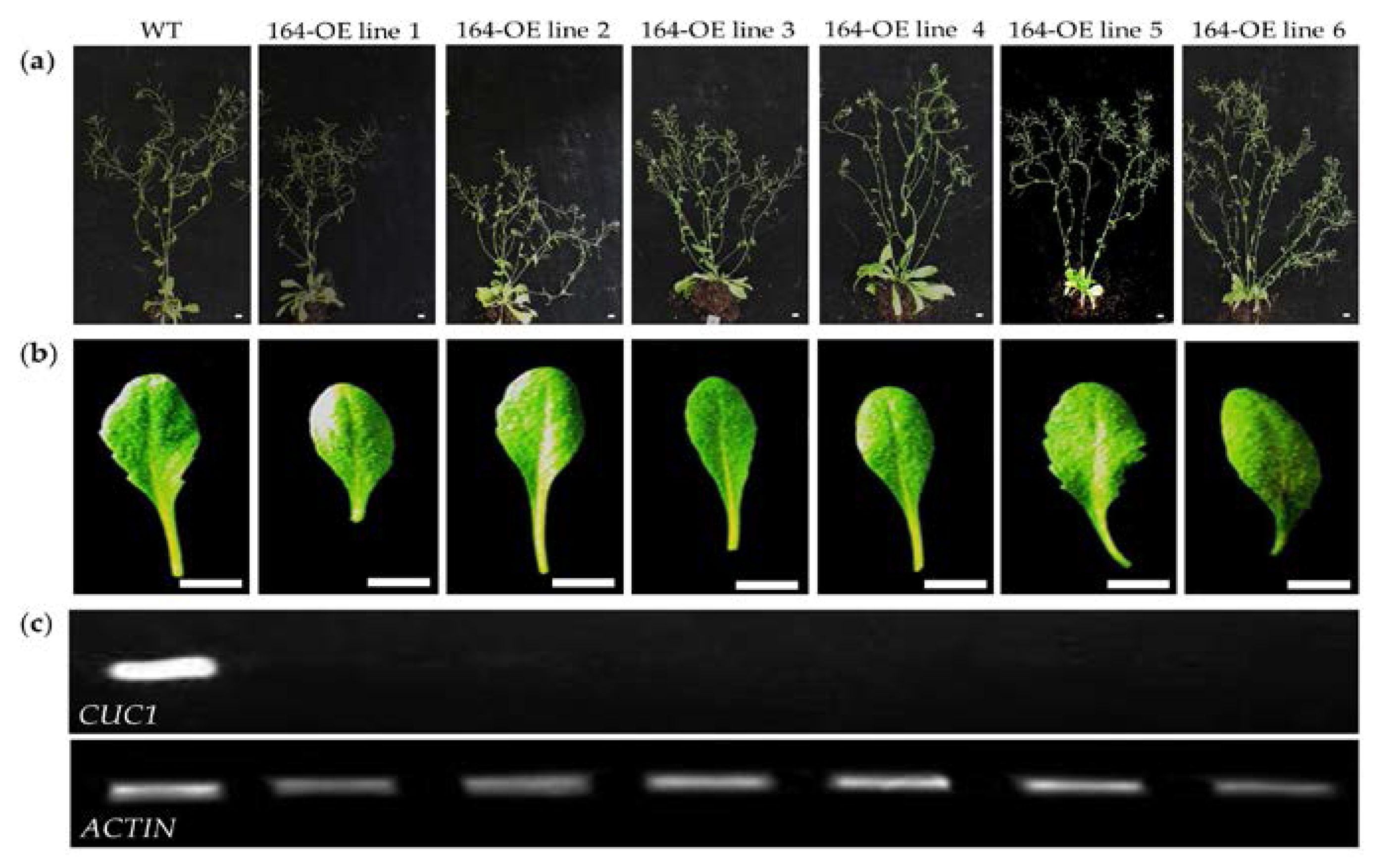
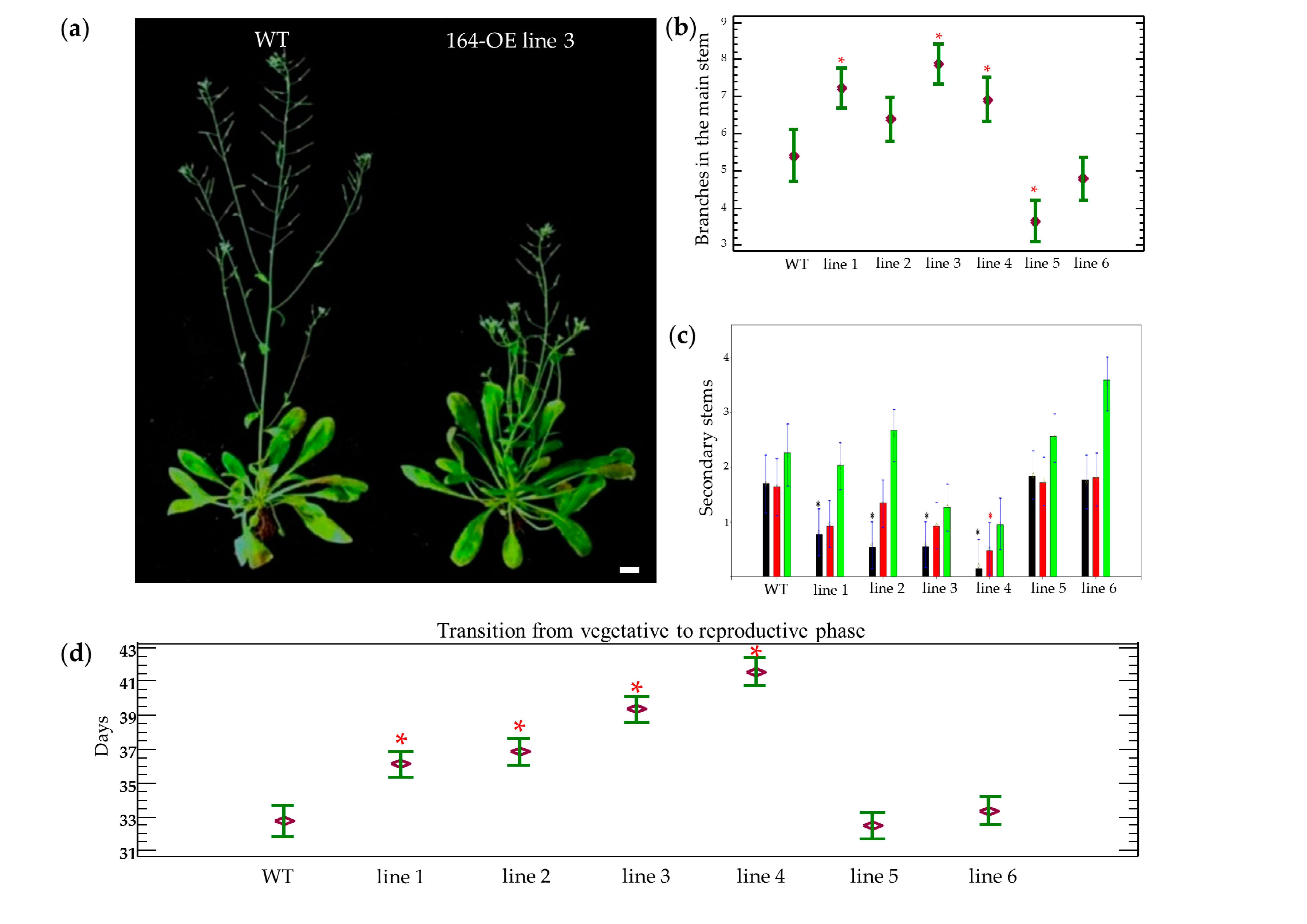
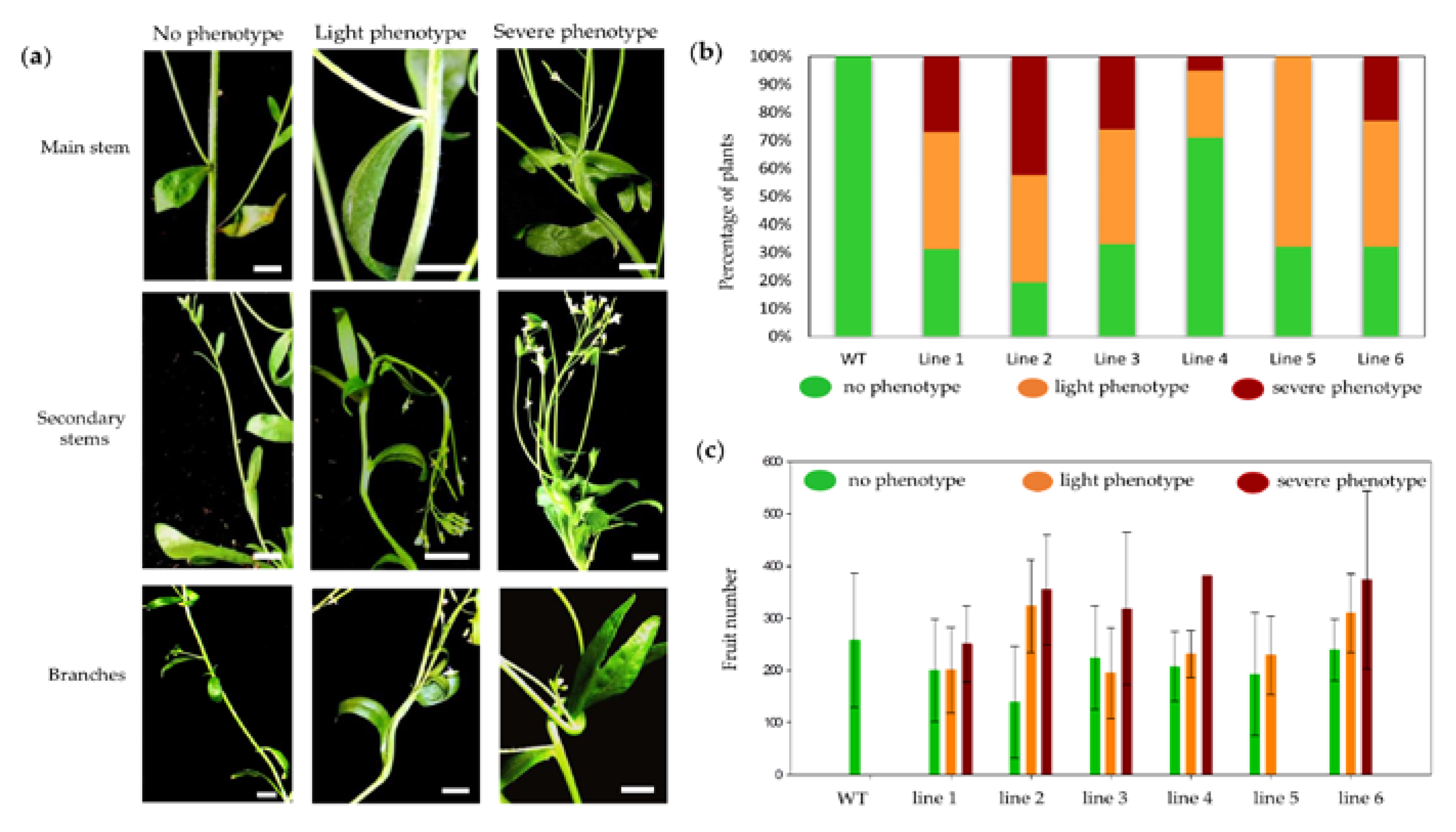
2.1. Characterization of miR164 Overexpression in Arabidopsis
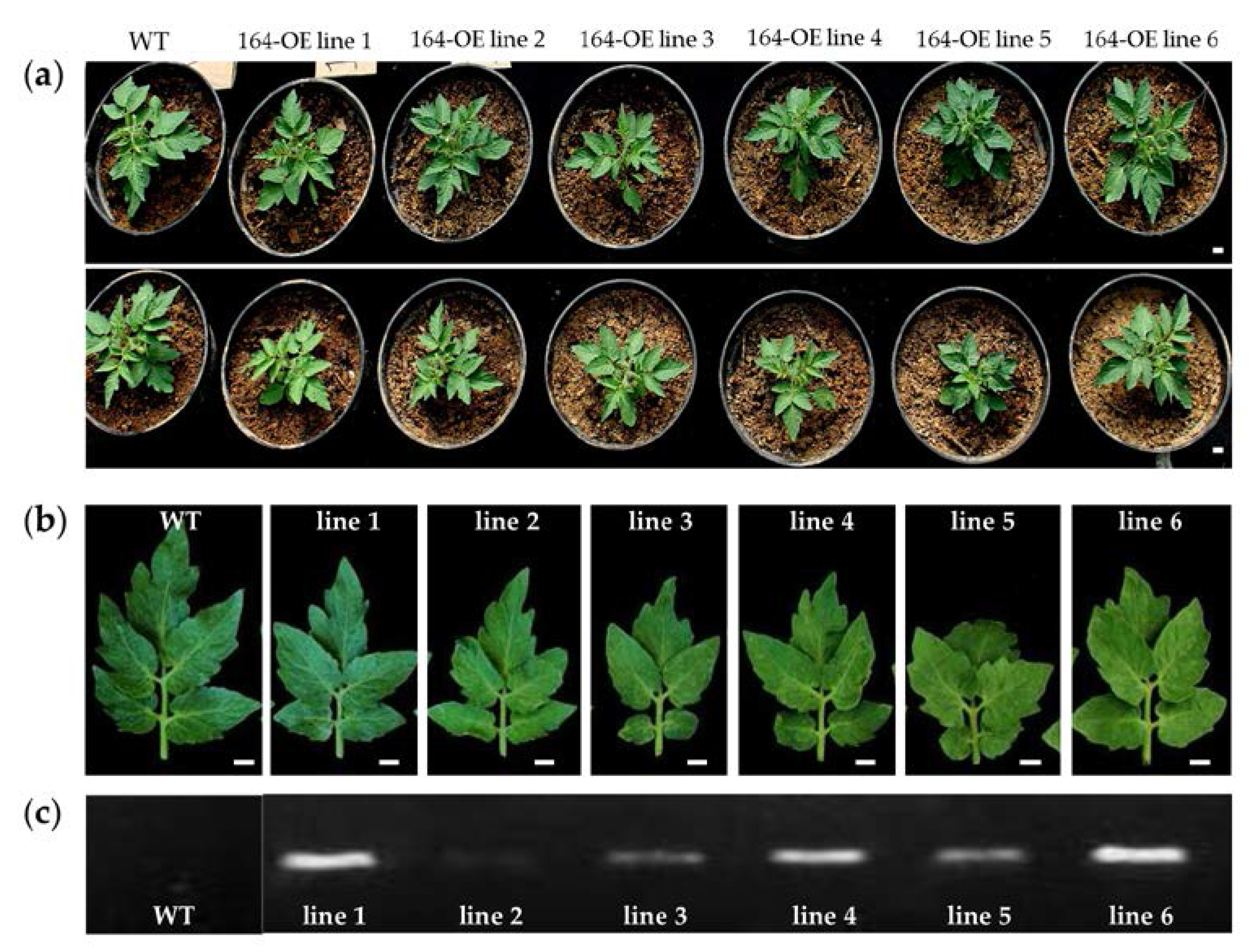
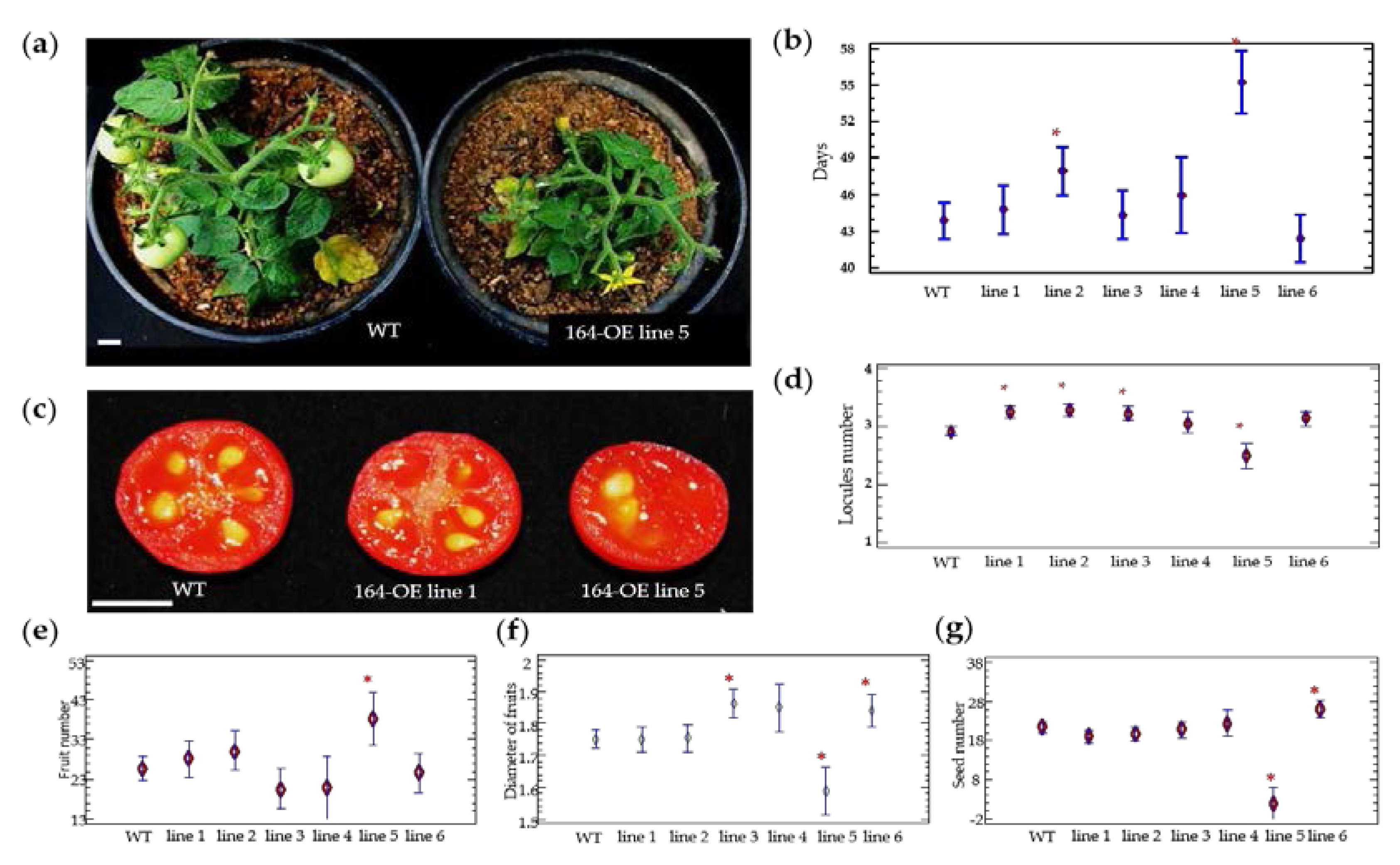
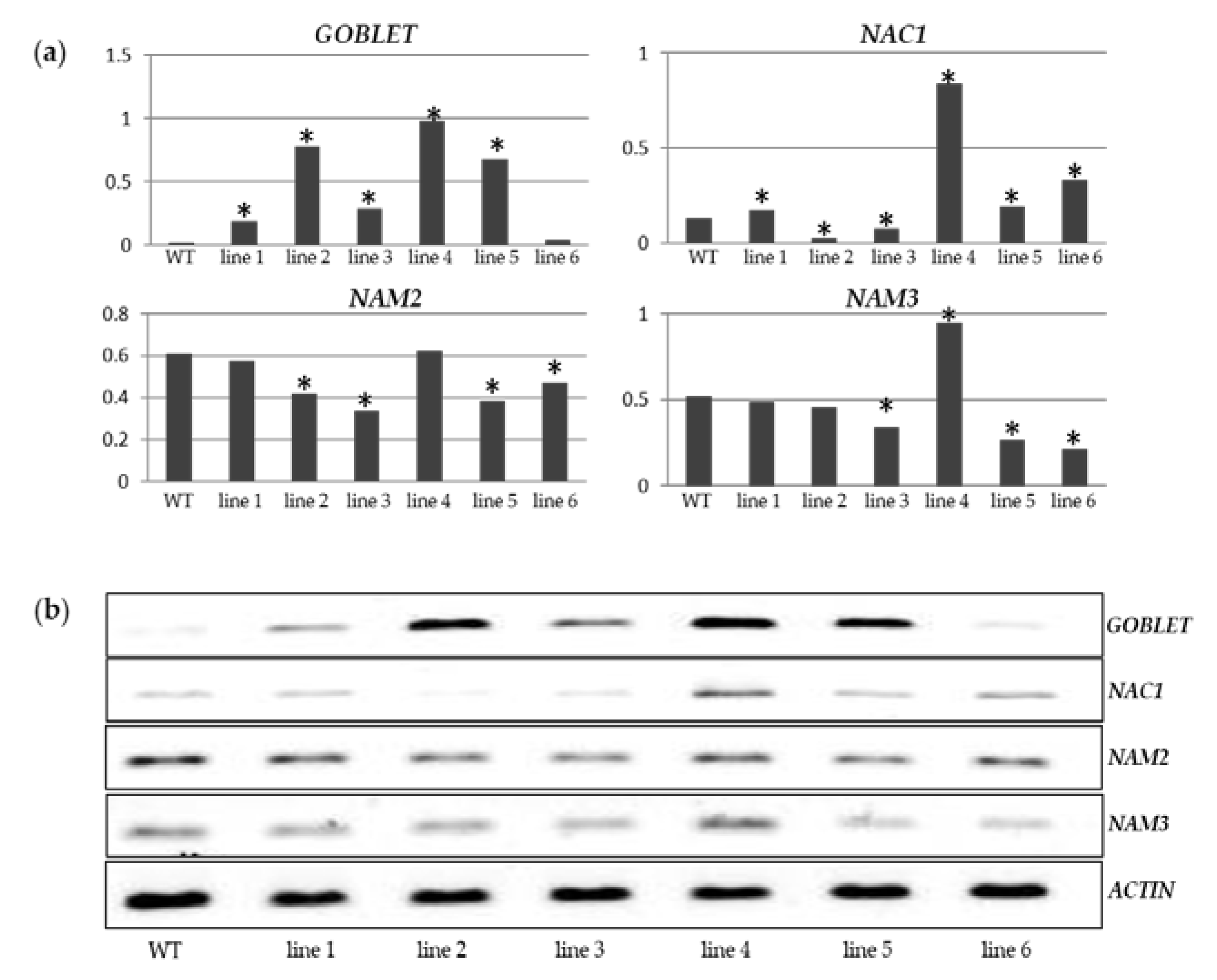
2.2. Characterization of miR164 Overexpression in Tomato
3. Discussion
3.1. Overexpression of miR164 Affects Plant and Organ Shape
3.2. Overexpression of miR164 Affects the Transition from the Vegetative to the Reproductive Phase
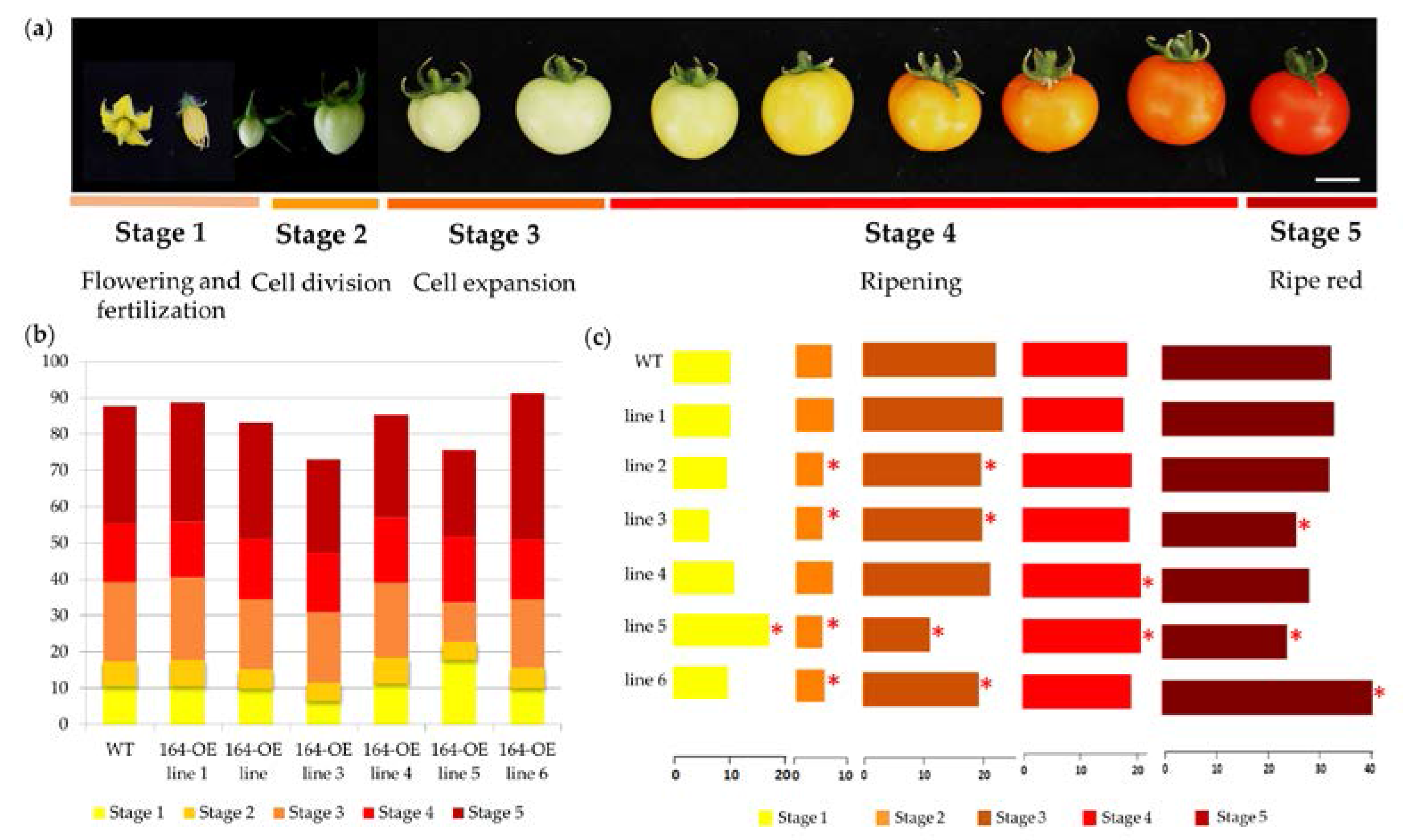
3.3. Overexpression of miR164 Affects the Time Required for Each Stage of Tomato Fruit Development
4. Materials and Methods
4.1. Plant Growth and Plant Material
4.2. Vector Construction
4.3. Arabidopsis Transformation
4.4. Tomato Transformation
4.5. DNA Extraction and PCR
4.6. RNA Extraction and RT-PCR
4.7. Phenotypic and Molecular Characterization
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhang, B.; Pan, X.; Cannon, C.H.; Cobb, G.P.; Anderson, T.A. Conservation and divergence of plant microRNA genes. Plant J. 2006, 46, 243–259. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.S.; Tarver, J.E.; Hiscock, S.J.; Donoghue, P.C.J. Evolutionary history of plant microRNAs. Trends Plant Sci. 2014, 19, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J.; Bowman, J.L. Evolution of plant microRNAs and their targets. Trends Plant Sci. 2008, 13, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Cuperus, J.T.; Fahlgren, N.; Carrington, J.C. Evolution and functional diversification of miRNA genes. Plant Cell 2011, 23, 431–442. [Google Scholar] [CrossRef] [PubMed]
- Chávez Montes, R.A.; Rosas-Cárdenas, F.D.F.; De Paoli, E.; Accerbi, M.; Rymarquis, L.A.; Mahalingam, G.; Marsch-Martínez, N.; Meyers, B.C.; Green, P.J.; de Folter, S. Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs. Nat. Commun. 2014, 5, 3722. [Google Scholar] [PubMed]
- Jones-Rhoades, M.W. Conservation and divergence in plant microRNAs. Plant Mol. Biol. 2011, 80, 3–16. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Mao, L. Evolution of plant microRNA gene families. Cell Res. 2007, 17, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Fahlgren, N.; Jogdeo, S.; Kasschau, K.D.; Sullivan, C.M.; Chapman, E.J.; Laubinger, S.; Smith, L.M.; Dasenko, M.; Givan, S.A.; Weigel, D.; et al. MicroRNA gene evolution in Arabidopsis lyrata and Arabidopsis thaliana. Plant Cell 2010, 22, 1074–1089. [Google Scholar] [CrossRef] [PubMed]
- Nozawa, M.; Miura, S.; Nei, M. Origins and evolution of microRNA genes in plant species. Genome Biol. Evol. 2012, 4, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 2009, 136, 669–687. [Google Scholar] [CrossRef] [PubMed]
- Mallory, A.C.; Bouché, N. MicroRNA-directed regulation: To cleave or not to cleave. Trends Plant Sci. 2008, 13, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R.; Steitz, J.A. Review the noncoding RNA revolution—Trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Woo, H.R.; Kim, J.; Lim, P.O.; Lee, I.C.; Choi, S.H.; Hwang, D.; Nam, H.G. Trifurcate feed-forward regulation of age-dependent cell death involving miR164 in Arabidopsis. Science 2009, 323, 1053–1057. [Google Scholar] [CrossRef] [PubMed]
- Rosas-Cárdenas, F.D.F.; De Folter, S. Conservation, divergence and abundance of miRNAs and their effect in plants. In RNA Technologies, Plant Epigenetics; Rajewsky, N., Jurga, S., Barciszewski, J., Eds.; Springer: Berlin, Germany, 2017; pp. 1–22. [Google Scholar]
- Jin, D.; Wang, Y.; Zhao, Y.; Chen, M. MicroRNAs and their cross-talks in plant development. J. Genet. Genom. 2013, 40, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.; Li, Y.; Wang, J.; Nan, H.; Wang, Y.; Lu, S.; Jiang, Q.; Li, X.; Shi, D.; Fang, C.; et al. GmmiR156b overexpression delays flowering time in soybean. Plant Mol. Biol. 2015, 89, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-S.; Lee, D.-Y.; Cho, L.-H.; An, G. Rice miR172 induces flowering by suppressing OsIDS1 and SNB, two AP2 genes that negatively regulate expression of Ehd1 and florigens. Rice 2014, 7, 31. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.-L.; Tomes, S.; Xu, J.; Gleave, A.P. How microRNA172 affects fruit growth in different species is dependent on fruit type. Plant Signal. Behav. 2016, 2324, e1156833. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.F.F.E.; Silva, E.M.; Da Silva Azevedo, M.; Guivin, M.A.C.; Ramiro, D.A.; Figueiredo, C.R.; Carrer, H.; Peres, L.E.P.; Nogueira, F.T.S. MicroRNA156-targeted SPL/SBP box transcription factors regulate tomato ovary and fruit development. Plant J. 2014, 78, 604–618. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Hu, T.; Smith, M.R.; Poethig, R.S. Epigenetic regulation of vegetative phase change in Arabidopsis. Plant Cell 2016, 28, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.; Sunkar, R.; Zhou, C.; Shen, H.; Zhang, J.-Y.; Matts, J.; Wolf, J.; Mann, D.G.J.; Stewart, C.N.; Tang, Y.; et al. Overexpression of miR156 in switchgrass (Panicum virgatum L.) results in various morphological alterations and leads to improved biomass production. Plant Biotechnol. J. 2012, 10, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Jones-Rhoades, M.W.; Bartel, D.P. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell 2004, 14, 787–799. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Weinstein, E.G.; Rhoades, M.W.; Bartel, B.; Bartel, D.P. MicroRNAs in plants. Genes Dev. 2002, 16, 1616–1626. [Google Scholar] [CrossRef] [PubMed]
- Hendelman, A.; Stav, R.; Zemach, H.; Arazi, T. The tomato NAC transcription factor SlNAM2 is involved in flower-boundary morphogenesis. J. Exp. Bot. 2013, 64, 5497–5507. [Google Scholar] [CrossRef] [PubMed]
- Mallory, A.C.; Dugas, D.V.; Bartel, D.P.; Bartel, B. MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr. Biol. 2004, 14, 1035–1046. [Google Scholar] [CrossRef] [PubMed]
- Baker, C.C.; Sieber, P.; Wellmer, F.; Meyerowitz, E.M. The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr. Biol. 2005, 15, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Xie, Q.; Fei, J.; Chua, N. MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 2005, 17, 1376–1386. [Google Scholar] [CrossRef] [PubMed]
- Laufs, P.; Peaucelle, A.; Morin, H.; Traas, J. MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 2004, 131, 4311–4322. [Google Scholar] [CrossRef] [PubMed]
- Nikovics, K.; Blein, T.; Peaucelle, A.; Ishida, T.; Morin, H.; Aida, M.; Laufs, P. The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 2006, 18, 2929–2945. [Google Scholar] [CrossRef] [PubMed]
- Sieber, P.; Wellmer, F.; Gheyselinck, J.; Riechmann, J.L.; Meyerowitz, E.M. Redundancy and specialization among plant microRNAs: Role of the MIR164 family in developmental robustness. Development 2007, 134, 1051–1060. [Google Scholar] [CrossRef] [PubMed]
- Blein, T.; Pulido, A.; Vialette-Guiraud, A.; Nikovics, K.; Morin, H.; Hay, A.; Johansen, I.E.; Tsiantis, M.; Laufs, P. A conserved molecular framework for compound leaf development. Science 2008, 322, 1835–1839. [Google Scholar] [CrossRef] [PubMed]
- Koyama, T.; Mitsuda, N.; Seki, M.; Shinozaki, K.; Ohme-Takagi, M. TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis. Plant Cell 2010, 22, 3574–3588. [Google Scholar] [CrossRef] [PubMed]
- Hasson, A.; Plessis, A.; Blein, T.; Adroher, B.; Grigg, S.; Tsiantis, M.; Boudaoud, A.; Damerval, C.; Laufs, P. Evolution and diverse roles of the CUP-SHAPED COTYLEDON genes in Arabidopsis leaf development. Plant Cell 2011, 23, 54–68. [Google Scholar] [CrossRef] [PubMed]
- Berger, Y.; Harpaz-Saad, S.; Brand, A.; Melnik, H.; Sirding, N.; Alvarez, J.P.; Zinder, M.; Samach, A.; Eshed, Y.; Ori, N. The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 2009, 136, 823–832. [Google Scholar] [CrossRef] [PubMed]
- Mallory, A.C.; Vaucheret, H. MicroRNAs: Something important between the genes. Curr. Opin. Plant Biol. 2004, 7, 120–125. [Google Scholar] [CrossRef] [PubMed]
- Raman, S.; Greb, T.; Peaucelle, A.; Blein, T.; Laufs, P.; Theres, K. Interplay of miR164, CUP-SHAPED COTYLEDON genes and LATERAL SUPPRESSOR controls axillary meristem formation in Arabidopsis thaliana. Plant J. 2008, 55, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Vialette-Guiraud, A.C.M.; Chauvet, A.; Gutierrez-Mazariegos, J.; Eschstruth, A.; Ratet, P.; Scutt, C.P. A conserved role for the NAM/miR164 developmental module reveals a common mechanism underlying carpel margin fusion in monocarpous and syncarpous eurosids. Front. Plant Sci. 2016, 6, 1239. [Google Scholar] [CrossRef] [PubMed]
- Rhoades, M.W.; Reinhart, B.J.; Lim, L.P.; Burge, C.B.; Bartel, B.; Bartel, D.P. Prediction of plant microRNA targets. Cell 2002, 110, 513–520. [Google Scholar] [CrossRef]
- Schwab, R.; Palatnik, J.F.; Riester, M.; Schommer, C.; Schmid, M.; Weigel, D. Specific effects of microRNAs on the plant transcriptome. Dev. Cell 2005, 8, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Karlova, R.; van Haarst, J.C.; Maliepaard, C.; van de Geest, H.; Bovy, A.G.; Lammers, M.; Angenent, G.C.; de Maagd, R.A. Identification of microRNA targets in tomato fruit development using high-throughput sequencing and degradome analysis. J. Exp. Bot. 2013, 64, 1863–1878. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Hong, S.H.; Kim, Y.W.; Lee, I.H.; Jun, J.H.; Phee, B.K.; Rupak, T.; Jeong, H.; Lee, Y.; Hong, B.S.; et al. Gene regulatory cascade of senescence-associated NAC transcription factors activated by ETHYLENE-INSENSITIVE2-mediated leaf senescence signalling in Arabidopsis. J. Exp. Bot. 2014, 65, 4023–4036. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, L.; Chen, D.; Wu, X.; Huang, D.; Chen, L.; Li, L.; Deng, X. Genome-wide comparison of microRNAs and their targeted transcripts among leaf, flower and fruit of sweet orange. BMC Genom. 2014, 15, 695. [Google Scholar] [CrossRef] [PubMed]
- Pantaleo, V.; Szittya, G.; Moxon, S.; Miozzi, L.; Moulton, V.; Dalmay, T.; Burgyan, J. Identification of grapevine microRNAs and their targets using high-throughput sequencing and degradome analysis. Plant J. 2010, 62, 960–976. [Google Scholar] [PubMed]
- Sun, X.; Korir, N.K.; Han, J.; Shangguan, L.F.; Kayesh, E.; Leng, X.P.; Fang, J.G. Characterization of grapevine microR164 and its target genes. Mol. Biol. Rep. 2012, 39, 9463–9472. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Gomollon, S.; Mohorianu, I.; Szittya, G.; Moulton, V.; Dalmay, T. Diverse correlation patterns between microRNAs and their targets during tomato fruit development indicates different modes of microRNA actions. Planta 2012, 236, 1875–1887. [Google Scholar] [CrossRef] [PubMed]
- Rosas-Cárdenas, F.D.F.; Caballero-Pérez, J.; Ramos, X.G.; Cruz-Hernández, A.; Marsch-Martínez, N.; de Folter, S. miRNA expression during prickly pear cactus fruit development. Planta 2014, 241, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Bilsborough, G.D.; Runions, A.; Barkoulas, M.; Jenkins, H.W.; Hasson, A.; Galinha, C.; Laufs, P.; Hay, A.; Prusinkiewicz, P.; Tsiantis, M. Model for the regulation of Arabidopsis thaliana leaf margin development. Proc. Natl. Acad. Sci. USA 2011, 108, 3424–3429. [Google Scholar] [CrossRef] [PubMed]
- Adam, H.; Marguerettaz, M.; Qadri, R.; Adroher, B.; Richaud, F.; Collin, M.; Thuillet, A.-C.; Vigouroux, Y.; Laufs, P.; Tregear, J.W.; et al. Divergent expression patterns of miR164 and CUP-SHAPED COTYLEDON genes in palms and other monocots: Implication for the evolution of meristem function in angiosperms. Mol. Biol. Evol. 2011, 28, 1439–1454. [Google Scholar] [CrossRef] [PubMed]
- Lauter, N.; Kampani, A.; Carlson, S.; Goebel, M.; Moose, S.P. microRNA172 down-regulates glossy15 to promote vegetative phase change in maize. Proc. Natl. Acad. Sci. USA 2005, 102, 9412–9417. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Park, M.Y.; Wang, L.J.; Koo, Y.; Chen, X.Y.; Weigel, D.; Poethig, R.S. MiRNA control of vegetative phase change in trees. PLoS Genet. 2011, 7, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Galbiati, F.; Sinha Roy, D.; Simonini, S.; Cucinotta, M.; Ceccato, L.; Cuesta, C.; Simaskova, M.; Benkova, E.; Kamiuchi, Y.; Aida, M.; et al. An integrative model of the control of ovule primordia formation. Plant J. 2013, 76, 446–455. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, B.; Hasson, A.; Belcram, K.; Cortizo, M.; Morin, H.; Nikovics, K.; Vialette-Guiraud, A.; Takeda, S.; Aida, M.; Laufs, P.; et al. A conserved role for CUP-SHAPED COTYLEDON genes during ovule development. Plant J. 2015, 83, 732–742. [Google Scholar] [CrossRef] [PubMed]
- Pino, L.E.; Lombardi-Crestana, S.; Azavedo, M.A.; Scotton, D.C.; Borgo, L.; Quecini, V.; Figueira, A.; Peres, L.E. The Rg1 allele as a valuable tool for genetic transformation of the tomato “Micro-Tom” model system. Plant Methods 2010, 6, 23. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosas Cárdenas, F.D.F.; Ruiz Suárez, Y.; Cano Rangel, R.M.; Luna Garcia, V.; González Aguilera, K.L.; Marsch Martínez, N.; De Folter, S. Effect of Constitutive miR164 Expression on Plant Morphology and Fruit Development in Arabidopsis and Tomato. Agronomy 2017, 7, 48. https://doi.org/10.3390/agronomy7030048
Rosas Cárdenas FDF, Ruiz Suárez Y, Cano Rangel RM, Luna Garcia V, González Aguilera KL, Marsch Martínez N, De Folter S. Effect of Constitutive miR164 Expression on Plant Morphology and Fruit Development in Arabidopsis and Tomato. Agronomy. 2017; 7(3):48. https://doi.org/10.3390/agronomy7030048
Chicago/Turabian StyleRosas Cárdenas, Flor De Fátima, Yolanda Ruiz Suárez, Rosa María Cano Rangel, Valentín Luna Garcia, Karla Lorena González Aguilera, Nayelli Marsch Martínez, and Stefan De Folter. 2017. "Effect of Constitutive miR164 Expression on Plant Morphology and Fruit Development in Arabidopsis and Tomato" Agronomy 7, no. 3: 48. https://doi.org/10.3390/agronomy7030048
APA StyleRosas Cárdenas, F. D. F., Ruiz Suárez, Y., Cano Rangel, R. M., Luna Garcia, V., González Aguilera, K. L., Marsch Martínez, N., & De Folter, S. (2017). Effect of Constitutive miR164 Expression on Plant Morphology and Fruit Development in Arabidopsis and Tomato. Agronomy, 7(3), 48. https://doi.org/10.3390/agronomy7030048




