Camelina sativa (L.) Crantz as a Promising Cover Crop Species with Allelopathic Potential
Abstract
:1. Introduction
2. Camelina sativa: A Promising Cover Crop
3. Camelina sativa: A Potential Allelopathic Crop
4. Sulfur Availability and Glucosinolates Production
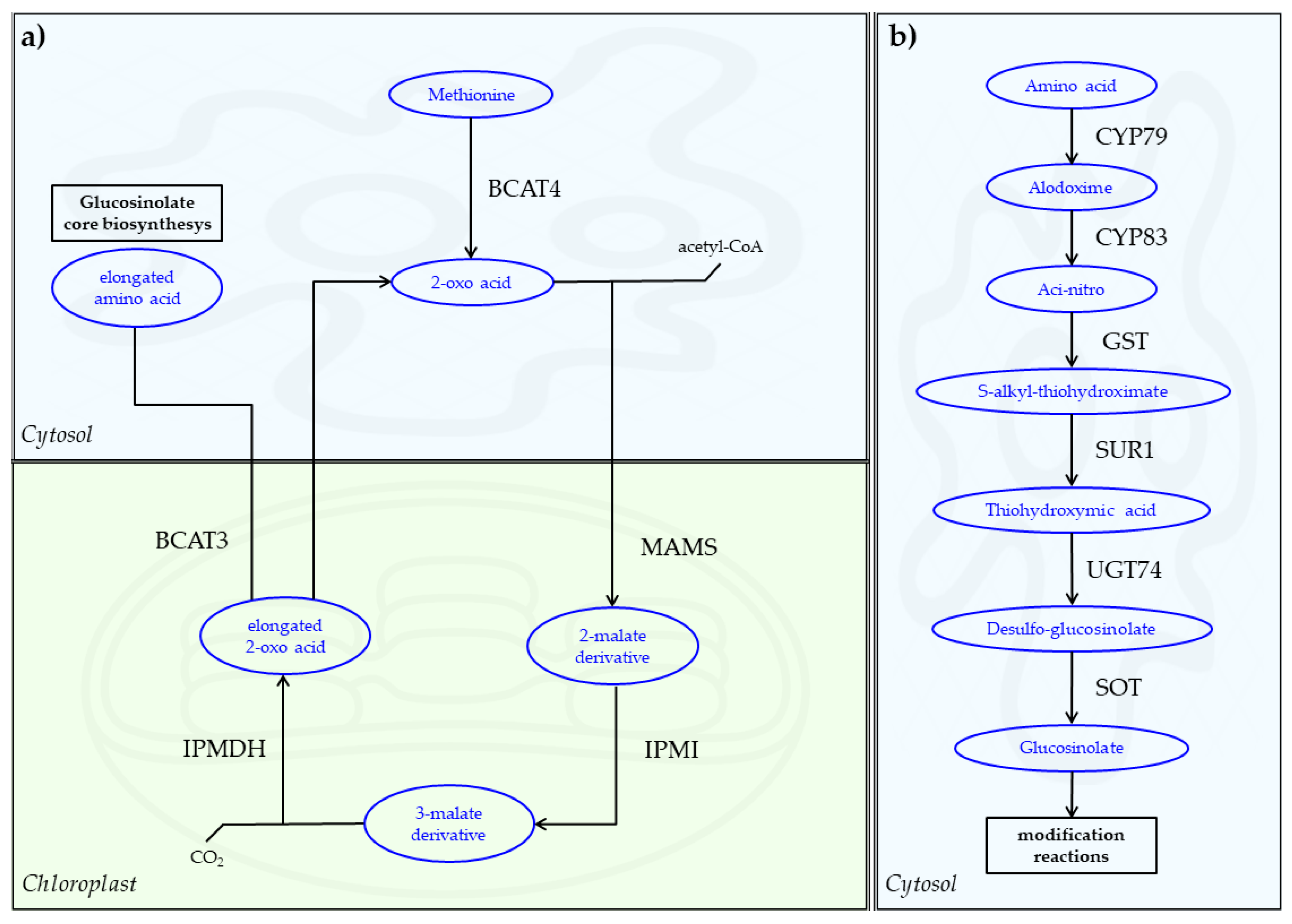
5. Biosynthesis of Glucosinolates
6. Glucosinolate Transport
7. Glucosinolates Breakdown Product
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Aci, M.M.; Sidari, R.; Araniti, F.; Lupini, A. Emerging Trends in Allelopathy: A Genetic Perspective for Sustainable Agriculture. Agronomy 2022, 12, 2043. [Google Scholar] [CrossRef]
- Choudhary, C.S.; Behera, B.; Raza, M.B.; Mrunalini, K.; Bhoi, T.K.; Lal, M.K.; Nongmaithem, D.; Pradhan, S.; Song, B.; Das, T.K. Mechanisms of Allelopathic Interactions for Sustainable Weed Management. Rhizosphere 2023, 25, 100667. [Google Scholar] [CrossRef]
- Hickman, D.T.; Comont, D.; Rasmussen, A.; Birkett, M.A. Novel and Holistic Approaches Are Required to Realize Allelopathic Potential for Weed Management. Ecol. Evol. 2023, 13, e10018. [Google Scholar] [CrossRef] [PubMed]
- Gealy, D.R.; Dilday, R.H.; Rutger, J.N. Interaction of Flush Irrigation Timing and Suppression of Barnyardgrass with Potentially Allelopathic Rice Lines. Res. Ser.-Ark. Agric. Exp. Stn. 1998, 460, 49–55. [Google Scholar]
- He, H.-Q.; Shen, L.-H.; Xiong, J.; Jia, X.-L.; Lin, W.-X.; Wu, H. Conditional Genetic Effect of Allelopathy in Rice (Oryza sativa L.) under Different Environmental Conditions. Plant Growth Regul. 2004, 44, 211–218. [Google Scholar] [CrossRef]
- Scavo, A.; Abbate, C.; Mauromicale, G. Plant Allelochemicals: Agronomic, Nutritional and Ecological Relevance in the Soil System. Plant Soil 2019, 442, 23–48. [Google Scholar] [CrossRef]
- Scavo, A.; Mauromicale, G. Crop Allelopathy for Sustainable Weed Management in Agroecosystems: Knowing the Present with a View to the Future. Agronomy 2021, 11, 2104. [Google Scholar] [CrossRef]
- Kong, C.-H.; Chen, X.-H.; Hu, F.; Zhang, S.-Z. Breeding of Commercially Acceptable Allelopathic Rice Cultivars in China. Pest Manag. Sci. 2011, 67, 1100–1106. [Google Scholar] [CrossRef]
- Worthington, M.; Reberg-Horton, C. Breeding Cereal Crops for Enhanced Weed Suppression: Optimizing Allelopathy and Competitive Ability. J. Chem. Ecol. 2013, 39, 213–231. [Google Scholar] [CrossRef]
- Shehzad, T.; Okuno, K. Genetic Analysis of QTLs Controlling Allelopathic Characteristics in Sorghum. PLoS ONE 2020, 15, e0235896. [Google Scholar] [CrossRef]
- Pan, Z.; Baerson, S.R.; Wang, M.; Bajsa-Hirschel, J.; Rimando, A.M.; Wang, X.; Nanayakkara, N.P.D.; Noonan, B.P.; Fromm, M.E.; Dayan, F.E.; et al. A Cytochrome P450 CYP71 Enzyme Expressed in Sorghum Bicolor Root Hair Cells Participates in the Biosynthesis of the Benzoquinone Allelochemical Sorgoleone. New Phytol. 2018, 218, 616–629. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.I.; Araniti, F.; Schulz, M.; Baerson, S.; Vieites-Álvarez, Y.; Rempelos, L.; Bilsborrow, P.; Chinchilla, N.; Macías, F.A.; Weston, L.A.; et al. Benzoxazinoids in Wheat Allelopathy—From Discovery to Application for Sustainable Weed Management. Environ. Exp. Bot. 2022, 202, 104997. [Google Scholar] [CrossRef]
- Jabran, K.; Mahajan, G.; Sardana, V.; Chauhan, B.S. Allelopathy for Weed Control in Agricultural Systems. Crop Prot. 2015, 72, 57–65. [Google Scholar] [CrossRef]
- Haramoto, E.R.; Gallandt, E.R. Brassica Cover Cropping: I. Effects on Weed and Crop Establishment. Weed Sci. 2005, 53, 695–701. [Google Scholar] [CrossRef]
- Haramoto, E.R.; Gallandt, E.R. Brassica Cover Cropping for Weed Management: A Review. Renew. Agric. Food Syst. 2004, 19, 187–198. [Google Scholar] [CrossRef]
- Kruger, D.H.M.; Fourie, J.C.; Malan, A.P. Cover Crops with Biofumigation Properties for the Suppression of Plant-Parasitic Nematodes: A Review. S. Afr. J. Enol. Vitic. 2013, 34, 287–295. [Google Scholar] [CrossRef]
- Rehman, S.; Shahzad, B.; Bajwa, A.A.; Hussain, S.; Rehman, A.; Cheema, S.A.; Abbas, T.; Ali, A.; Shah, L.; Adkins, S.; et al. Utilizing the Allelopathic Potential of Brassica Species for Sustainable Crop Production: A Review. J. Plant Growth Regul. 2019, 38, 343–356. [Google Scholar] [CrossRef]
- Francisco, M.; Tortosa, M.; Martínez-Ballesta, M.d.C.; Velasco, P.; García-Viguera, C.; Moreno, D.A. Nutritional and Phytochemical Value of Brassica Crops from the Agri-Food Perspective. Ann. Appl. Biol. 2017, 170, 273–285. [Google Scholar] [CrossRef]
- Rakow, G. Species Origin and Economic Importance of Brassica. In Brassica; Pua, E.-C., Douglas, C.J., Eds.; Biotechnology in Agriculture and Forestry; Springer: Berlin/Heidelberg, Germany, 2004; pp. 3–11. ISBN 978-3-662-06164-0. [Google Scholar]
- Soodabeh Saeidnia Importance of Brassica napus as a Medicinal Food Plant. J. Med. Plants Res. 2012, 6, 2700–2703. [CrossRef]
- Halkier, B.A.; Gershenzon, J. Biology and Biochemistry of Glucosinolates. Annu. Rev. Plant Biol. 2006, 57, 303–333. [Google Scholar] [CrossRef]
- Bialy, Z.; Oleszek, W.; Lewis, J.; Fenwick, G.R. Allelopathic Potential of Glucosinolates (Mustard Oil Glycosides) and Their Degradation Products against Wheat. Plant Soil 1990, 129, 277–281. [Google Scholar] [CrossRef]
- Rivera-Vega, L.J.; Krosse, S.; de Graaf, R.M.; Garvi, J.; Garvi-Bode, R.D.; van Dam, N.M. Allelopathic Effects of Glucosinolate Breakdown Products in Hanza [Boscia senegalensis (Pers.) Lam.] Processing Waste Water. Front. Plant Sci. 2015, 6, 532. [Google Scholar] [CrossRef] [PubMed]
- Intanon, S.; Reed, R.L.; Stevens, J.F.; Hulting, A.G.; Mallory-Smith, C.A. Identification and Phytotoxicity of a New Glucosinolate Breakdown Product from Meadowfoam (Limnanthes alba) Seed Meal. J. Agric. Food Chem. 2014, 62, 7423–7429. [Google Scholar] [CrossRef] [PubMed]
- Wolf, R.B.; Spencer, G.F.; Kwolek, W.F. Inhibition of Velvetleaf (Abutilon theophrasti) Germination and Growth by Benzyl Isothiocyanate, a Natural Toxicant. Weed Sci. 1984, 32, 612–615. [Google Scholar] [CrossRef]
- Yamane, A.; Fujikura, J.; Ogawa, H.; Mizutani, J. Isothiocyanates as Alleopathic Compounds from Rorippa indica Hiern. (Cruciferae) Roots. J. Chem. Ecol. 1992, 18, 1941–1954. [Google Scholar] [CrossRef]
- Baysal-Gurel, F.; Liyanapathiranage, P.; Addesso, K.M. Effect of Brassica Crop-Based Biofumigation on Soilborne Disease Suppression in Woody Ornamentals. Can. J. Plant Pathol. 2020, 42, 94–106. [Google Scholar] [CrossRef]
- Harvey, S.G.; Hannahan, H.N.; Sams, C.E. Indian Mustard and Allyl Isothiocyanate Inhibit Sclerotium rolfsii. J. Am. Soc. Hortic. Sci. 2002, 127, 27–31. [Google Scholar] [CrossRef]
- Wang, T.; Li, Y.; Bi, Y.; Zhang, M.; Zhang, T.; Zheng, X.; Dong, Y.; Huang, Y. Benzyl Isothiocyanate Fumigation Inhibits Growth, Membrane Integrity and Mycotoxin Production in Alternaria alternata. RSC Adv. 2020, 10, 1829–1837. [Google Scholar] [CrossRef]
- Barani, E.; Shafaat, G. Allelopathic Effect of Brassica napus Residues and Etalfluraline Herbicide on Germination and Some Cotton Characteristics of Bakhtegan Cultivar. J. Plant Prod. Sci. 2022, 12, 47–61. [Google Scholar] [CrossRef]
- Rehman, S.U. Allelopathic Potential of Sinapis alba L. Residues in Weeds Management System. J. Arable Crops Mark. 2021, 3, 39–43. [Google Scholar] [CrossRef]
- Toosi, F.; Baki, B.B. Allelopathic Potential of Brassica juncea (L.) Czern. Var. Ensabi. In Proceedings of the 23rd Asian-Pacific Weed Science Society Conference, Cairns, QLD, Australia, 26–29 September 2011; Volume 1, pp. 555–558. [Google Scholar]
- Zhou, X.; Xing, C.; Jiang, B.; Li, C.; Liu, X. Allelopathic effects of water extracts of Brassica juncea var. tumida leaf on seed germination of three species of crops. J. Henan Agric. Sci. 2015, 44, 117–121. [Google Scholar]
- Sainger, M.; Jaiwal, A.; Sainger, P.A.; Chaudhary, D.; Jaiwal, R.; Jaiwal, P.K. Advances in Genetic Improvement of Camelina sativa for Biofuel and Industrial Bio-Products. Renew. Sustain. Energy Rev. 2017, 68, 623–637. [Google Scholar] [CrossRef]
- Francis, A.; Warwick, S.I. The Biology of Canadian Weeds. 142. Camelina alyssum (Mill.) Thell.; C. microcarpa Andrz. Ex DC.; C. sativa (L.) Crantz. Can. J. Plant Sci. 2009, 89, 791–810. [Google Scholar] [CrossRef]
- Gehringer, A.; Friedt, W.; Lühs, W.; Snowdon, R.J. Genetic Mapping of Agronomic Traits in False Flax (Camelina sativa Subsp. sativa). Genome 2006, 49, 1555–1563. [Google Scholar] [CrossRef]
- Liu, X.; Brost, J.; Hutcheon, C.; Guilfoil, R.; Wilson, A.K.; Leung, S.; Shewmaker, C.K.; Rooke, S.; Nguyen, T.; Kiser, J.; et al. Transformation of the Oilseed Crop Camelina sativa by Agrobacterium-Mediated Floral Dip and Simple Large-Scale Screening of Transformants. Vitro Cell. Dev. Biol.-Plant 2012, 48, 462–468. [Google Scholar] [CrossRef]
- Liu, J.; Rice, A.; McGlew, K.; Shaw, V.; Park, H.; Clemente, T.; Pollard, M.; Ohlrogge, J.; Durrett, T.P. Metabolic Engineering of Oilseed Crops to Produce High Levels of Novel Acetyl Glyceride Oils with Reduced Viscosity, Freezing Point and Calorific Value. Plant Biotechnol. J. 2015, 13, 858–865. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Madsen, S.R.; Engelen, S.; Jørgensen, M.E.; Olsen, C.E.; Andersen, J.S.; Seynnaeve, D.; Verhoye, T.; Fulawka, R.; Denolf, P.; et al. Reduction of Antinutritional Glucosinolates in Brassica Oilseeds by Mutation of Genes Encoding Transporters. Nat. Biotechnol. 2017, 35, 377–382. [Google Scholar] [CrossRef]
- Amyot, L.; McDowell, T.; Martin, S.L.; Renaud, J.; Gruber, M.Y.; Hannoufa, A. Assessment of Antinutritional Compounds and Chemotaxonomic Relationships between Camelina sativa and Its Wild Relatives. J. Agric. Food Chem. 2019, 67, 796–806. [Google Scholar] [CrossRef]
- Hu, P.; Wu, L.; Hollister, E.B.; Wang, A.S.; Somenahally, A.C.; Hons, F.M.; Gentry, T.J. Fungal Community Structural and Microbial Functional Pattern Changes after Soil Amendments by Oilseed Meals of Jatropha curcas and Camelina sativa: A Microcosm Study. Front. Microbiol. 2019, 10, 537. [Google Scholar] [CrossRef]
- Arora, C.; Kaushik, R. Fungicidal Activity of Plants Extracts from Uttaranchal Hills against Soybean Fungal Pathogens. Allelopath. J. 2003, 11, 217–228. [Google Scholar]
- Acharya, K.; Yan, G.; Berti, M. Can Winter Camelina, Crambe, and Brown Mustard Reduce Soybean Cyst Nematode Populations? Ind. Crops Prod. 2019, 140, 111637. [Google Scholar] [CrossRef]
- Acharya, J.; Moorman, T.B.; Kaspar, T.C.; Lenssen, A.W.; Robertson, A.E. Cover Crop Rotation Effects on Growth and Development, Seedling Disease, and Yield of Corn and Soybean. Plant Dis. 2020, 104, 677–687. [Google Scholar] [CrossRef]
- Powell, S.; Mcphee, J.; Dean, G.; Hinton, S.; Sparrow, L.; Wilson, C.; Tegg, R. Managing Soil Health and Crop Productivity in Potato: A Challenging Test System. Soil Res. 2020, 58, 697–712. [Google Scholar] [CrossRef]
- Walker, B.A.R.; Powell, S.M.; Tegg, R.S.; Doyle, R.B.; Hunt, I.G.; Wilson, C.R. Soil Microbial Community Dynamics during Ryegrass Green Manuring and Brassica Biofumigation. Appl. Soil Ecol. 2022, 179, 104600. [Google Scholar] [CrossRef]
- Morra, M.J.; Kirkegaard, J.A. Isothiocyanate Release from Soil-Incorporated Brassica Tissues. Soil Biol. Biochem. 2002, 34, 1683–1690. [Google Scholar] [CrossRef]
- Fierer, N. Embracing the Unknown: Disentangling the Complexities of the Soil Microbiome. Nat. Rev. Microbiol. 2017, 15, 579–590. [Google Scholar] [CrossRef]
- Trivedi, P.; Delgado-Baquerizo, M.; Anderson, I.C.; Singh, B.K. Response of Soil Properties and Microbial Communities to Agriculture: Implications for Primary Productivity and Soil Health Indicators. Front. Plant Sci. 2016, 7, 990. [Google Scholar] [CrossRef]
- Hanschen, F.S.; Yim, B.; Winkelmann, T.; Smalla, K.; Schreiner, M. Degradation of Biofumigant Isothiocyanates and Allyl Glucosinolate in Soil and Their Effects on the Microbial Community Composition. PLoS ONE 2015, 10, e0132931. [Google Scholar] [CrossRef]
- Petersen, J.; Belz, R.; Walker, F.; Hurle, K. Weed Suppression by Release of Isothiocyanates from Turnip-Rape Mulch. Agron. J. 2001, 93, 37–43. [Google Scholar] [CrossRef]
- Mohler, C.L. Ecological Bases for the Cultural Control of Annual Weeds. J. Prod. Agric. 1996, 9, 468–474. [Google Scholar] [CrossRef]
- Berti, M.; Samarappuli, D.; Johnson, B.L.; Gesch, R.W. Integrating Winter Camelina into Maize and Soybean Cropping Systems. Ind. Crops Prod. 2017, 107, 595–601. [Google Scholar] [CrossRef]
- Šćepanović, M.; Sarić-Krsmanović, M.; Šoštarčić, V.; Brijačak, E.; Lakić, J.; Špirović Trifunović, B.; Gajić Umiljendić, J.; Radivojević, L. Inhibitory Effects of Brassicaceae Cover Crop on Ambrosia artemisiifolia Germination and Early Growth. Plants 2021, 10, 794. [Google Scholar] [CrossRef] [PubMed]
- Grummer, G.; Beyer, H. The influence exerted by species of Camelina on flax by means of toxic substances. Biol. Weeds Symp. Brit. Ecol. Soc. 1960, 153–157. [Google Scholar]
- Lovett, J. Defensive Stratagems of Plants, with Special Reference to Allelopathy. Pap. Proc. R. Soc. Tasman. 1985, 119, 31–37. [Google Scholar] [CrossRef]
- Lovett, J.V.; Sagar, G.R. Influence of Bacteria in the Phyllosphere of Camelina sativa (L.) Crantz on Germination of Linum usitatissimum L. New Phytol. 1978, 81, 617–625. [Google Scholar] [CrossRef]
- Lovett, J.V.; Jackson, H.F. Allelopathic Activity of Camelina sativa (l.) Crantz in Relation to Its Phyllosphere Bacteria. New Phytol. 1980, 86, 273–277. [Google Scholar] [CrossRef]
- Lovett, J.V. The Science of Allelopathy. In Allelopathy: The Australian Experience; Putnam, A.R., Tang, C.S., Eds.; John Wiley & Sons Inc.: New York, NY, USA, 1986; pp. 75–99. [Google Scholar]
- Lovett, J.V.; Duffield, A.M. Allelochemicals of Camelina sativa. J. Appl. Ecol. 1981, 18, 283–290. [Google Scholar] [CrossRef]
- Lovett, J.V. Allelopathy and Self-Defence in Plants. Aust. Weeds 1982, 2, 33–36. [Google Scholar]
- Lovett, J.V. The Effects of Allelochemicals on Crop Growth and Development. In Chemical Manipulation of Crop Growth and Development; McLaren, J.S., Ed.; Butterworths: London, UK, 1982; pp. 93–110. [Google Scholar]
- Saucke, H.; Ackermann, K. Weed Suppression in Mixed Cropped Grain Peas and False Flax (Camelina sativa). Weed Res. 2006, 46, 453–461. [Google Scholar] [CrossRef]
- Walsh, K.; Sanderson, D.; Hall, L.; Mugo, S.; Hills, M. Allelopathic Effects of Camelina (Camelina sativa) and Canola (Brassica napus) on Wild Oat, Flax and Radish. Allelopathy J. 2014, 33, 83. [Google Scholar]
- Jabran, K. Brassicaceae Allelopathy for Weed Control. In Manipulation of Allelopathic Crops for Weed Control; Jabran, K., Ed.; Springer Briefs in Plant Science; Springer International Publishing: Cham, Switzerland, 2017; pp. 21–27. ISBN 978-3-319-53186-1. [Google Scholar]
- Czerniawski, P.; Piasecka, A.; Bednarek, P. Evolutionary Changes in the Glucosinolate Biosynthetic Capacity in Species Representing Capsella, Camelina and Neslia Genera. Phytochemistry 2021, 181, 112571. [Google Scholar] [CrossRef]
- Berhow, M.A.; Vaughn, S.F.; Moser, B.R.; Belenli, D.; Polat, U. Evaluating the Phytochemical Potential of Camelina: An Emerging New Crop of Old World Origin. In Phytochemicals—Biosynthesis, Function and Application: Volume 44; Jetter, R., Ed.; Recent Advances in Phytochemistry; Springer International Publishing: Cham, Switzerland, 2014; pp. 129–148. ISBN 978-3-319-04045-5. [Google Scholar]
- Russo, R.; Reggiani, R. Glucosinolates and Sinapine in Camelina Meal. Food Nutr. Sci. 2017, 8, 1063–1073. [Google Scholar] [CrossRef]
- Hofmann, D.; Thiele, B.; Siebers, M.; Rahmati, M.; Schütz, V.; Jeong, S.; Cui, J.; Bigler, L.; Held, F.; Wu, B.; et al. Implications of Below-Ground Allelopathic Interactions of Camelina sativa and Microorganisms for Phosphate Availability and Habitat Maintenance. Plants 2023, 12, 2815. [Google Scholar] [CrossRef]
- Quéro, A.; Molinié, R.; Mathiron, D.; Thiombiano, B.; Fontaine, J.-X.; Brancourt, D.; Van Wuytswinkel, O.; Petit, E.; Demailly, H.; Mongelard, G.; et al. Metabolite Profiling of Developing Camelina sativa Seeds. Metabolomics 2016, 12, 186. [Google Scholar] [CrossRef]
- Bäuerle, R.; Wagner, H.; Schraudolf, H. Distribution of 4-Methoxy-3-Indolylmethyl-Glucosinolate (4-Methoxy-Glucobrassicin) in Brassicaceae. Experientia 1986, 42, 86. [Google Scholar] [CrossRef]
- Droux, M. Sulfur Assimilation and the Role of Sulfur in Plant Metabolism: A Survey. Photosynth. Res. 2004, 79, 331–348. [Google Scholar] [CrossRef] [PubMed]
- Saito, K. Sulfur Assimilatory Metabolism. The Long and Smelling Road. Plant Physiol. 2004, 136, 2443–2450. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Kopriva, S.; Giordano, M.; Saito, K.; Hell, R. Sulfur Assimilation in Photosynthetic Organisms: Molecular Functions and Regulations of Transporters and Assimilatory Enzymes. Annu. Rev. Plant Biol. 2011, 62, 157–184. [Google Scholar] [CrossRef]
- Sugiyama, R.; Li, R.; Kuwahara, A.; Nakabayashi, R.; Sotta, N.; Mori, T.; Ito, T.; Ohkama-Ohtsu, N.; Fujiwara, T.; Saito, K.; et al. Retrograde Sulfur Flow from Glucosinolates to Cysteine in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2021, 118, e2017890118. [Google Scholar] [CrossRef]
- Abdel-Massih, R.M.; Debs, E.; Othman, L.; Attieh, J.; Cabrerizo, F.M. Glucosinolates, a Natural Chemical Arsenal: More to Tell than the Myrosinase Story. Front. Microbiol. 2023, 14, 1130208. [Google Scholar] [CrossRef]
- Eugui, D.; Escobar, C.; Velasco, P.; Poveda, J. Glucosinolates as an Effective Tool in Plant-Parasitic Nematodes Control: Exploiting Natural Plant Defenses. Appl. Soil Ecol. 2022, 176, 104497. [Google Scholar] [CrossRef]
- Miękus, N.; Marszałek, K.; Podlacha, M.; Iqbal, A.; Puchalski, C.; Świergiel, A.H. Health Benefits of Plant-Derived Sulfur Compounds, Glucosinolates, and Organosulfur Compounds. Molecules 2020, 25, 3804. [Google Scholar] [CrossRef] [PubMed]
- Aarabi, F.; Kusajima, M.; Tohge, T.; Konishi, T.; Gigolashvili, T.; Takamune, M.; Sasazaki, Y.; Watanabe, M.; Nakashita, H.; Fernie, A.R.; et al. Sulfur Deficiency–Induced Repressor Proteins Optimize Glucosinolate Biosynthesis in Plants. Sci. Adv. 2016, 2, e1601087. [Google Scholar] [CrossRef] [PubMed]
- Wittstock, U.; Halkier, B.A. Cytochrome P450 CYP79A2 from Arabidopsis thaliana L. Catalyzes the Conversion of l-Phenylalanine to Phenylacetaldoxime in the Biosynthesis of Benzylglucosinolate. J. Biol. Chem. 2000, 275, 14659–14666. [Google Scholar] [CrossRef]
- Diebold, R.; Schuster, J.; Däschner, K.; Binder, S. The Branched-Chain Amino Acid Transaminase Gene Family in Arabidopsis Encodes Plastid and Mitochondrial Proteins. Plant Physiol. 2002, 129, 540–550. [Google Scholar] [CrossRef]
- Falk, K.L.; Vogel, C.; Textor, S.; Bartram, S.; Hick, A.; Pickett, J.A.; Gershenzon, J. Glucosinolate Biosynthesis: Demonstration and Characterization of the Condensing Enzyme of the Chain Elongation Cycle in Eruca sativa. Phytochemistry 2004, 65, 1073–1084. [Google Scholar] [CrossRef]
- Gigolashvili, T.; Yatusevich, R.; Rollwitz, I.; Humphry, M.; Gershenzon, J.; Flügge, U.-I. The Plastidic Bile Acid Transporter 5 Is Required for the Biosynthesis of Methionine-Derived Glucosinolates in Arabidopsis thaliana. Plant Cell 2009, 21, 1813–1829. [Google Scholar] [CrossRef]
- Knill, T.; Reichelt, M.; Paetz, C.; Gershenzon, J.; Binder, S. Arabidopsis thaliana Encodes a Bacterial-Type Heterodimeric Isopropylmalate Isomerase Involved in Both Leu Biosynthesis and the Met Chain Elongation Pathway of Glucosinolate Formation. Plant Mol. Biol. 2009, 71, 227–239. [Google Scholar] [CrossRef]
- Sawada, Y.; Toyooka, K.; Kuwahara, A.; Sakata, A.; Nagano, M.; Saito, K.; Hirai, M.Y. Arabidopsis Bile Acid:Sodium Symporter Family Protein 5 Is Involved in Methionine-Derived Glucosinolate Biosynthesis. Plant Cell Physiol. 2009, 50, 1579–1586. [Google Scholar] [CrossRef]
- Schuster, J.; Knill, T.; Reichelt, M.; Gershenzon, J.; Binder, S. Branched-Chain Aminotransferase4 Is Part of the Chain Elongation Pathway in the Biosynthesis of Methionine-Derived Glucosinolates in Arabidopsis. Plant Cell 2006, 18, 2664–2679. [Google Scholar] [CrossRef]
- Textor, S.; Bartram, S.; Kroymann, J.; Falk, K.L.; Hick, A.; Pickett, J.A.; Gershenzon, J. Biosynthesis of Methionine-Derived Glucosinolates in Arabidopsis thaliana: Recombinant Expression and Characterization of Methylthioalkylmalate Synthase, the Condensing Enzyme of the Chain-Elongation Cycle. Planta 2004, 218, 1026–1035. [Google Scholar] [CrossRef] [PubMed]
- Bednarek, P.; Piślewska-Bednarek, M.; Svatoš, A.; Schneider, B.; Doubský, J.; Mansurova, M.; Humphry, M.; Consonni, C.; Panstruga, R.; Sanchez-Vallet, A.; et al. A Glucosinolate Metabolism Pathway in Living Plant Cells Mediates Broad-Spectrum Antifungal Defense. Science 2009, 323, 101–106. [Google Scholar] [CrossRef] [PubMed]
- Schlaeppi, K.; Bodenhausen, N.; Buchala, A.; Mauch, F.; Reymond, P. The Glutathione-Deficient Mutant Pad2-1 Accumulates Lower Amounts of Glucosinolates and Is More Susceptible to the Insect Herbivore Spodoptera littoralis. Plant J. 2008, 55, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, M.D.; Naur, P.; Halkier, B.A. Arabidopsis Mutants in the C–S Lyase of Glucosinolate Biosynthesis Establish a Critical Role for Indole-3-Acetaldoxime in Auxin Homeostasis. Plant J. 2004, 37, 770–777. [Google Scholar] [CrossRef] [PubMed]
- Ravilious, G.E.; Jez, J.M. Structural Biology of Plant Sulfur Metabolism: From Assimilation to Biosynthesis. Nat. Prod. Rep. 2012, 29, 1138–1152. [Google Scholar] [CrossRef] [PubMed]
- Jez, J.M.; Ravilious, G.E.; Herrmann, J. Structural Biology and Regulation of the Plant Sulfation Pathway. Chem. Biol. Interact. 2016, 259, 31–38. [Google Scholar] [CrossRef]
- Mugford, S.G.; Lee, B.-R.; Koprivova, A.; Matthewman, C.; Kopriva, S. Control of Sulfur Partitioning between Primary and Secondary Metabolism. Plant J. 2011, 65, 96–105. [Google Scholar] [CrossRef]
- Mugford, S.G.; Matthewman, C.A.; Hill, L.; Kopriva, S. Adenosine-5′-Phosphosulfate Kinase Is Essential for Arabidopsis Viability. FEBS Lett. 2010, 584, 119–123. [Google Scholar] [CrossRef]
- Mugford, S.G.; Yoshimoto, N.; Reichelt, M.; Wirtz, M.; Hill, L.; Mugford, S.T.; Nakazato, Y.; Noji, M.; Takahashi, H.; Kramell, R.; et al. Disruption of Adenosine-5′-Phosphosulfate Kinase in Arabidopsis Reduces Levels of Sulfated Secondary Metabolites. Plant Cell 2009, 21, 910–927. [Google Scholar] [CrossRef]
- Ravilious, G.E.; Nguyen, A.; Francois, J.A.; Jez, J.M. Structural Basis and Evolution of Redox Regulation in Plant Adenosine-5′-Phosphosulfate Kinase. Proc. Natl. Acad. Sci. USA 2012, 109, 309–314. [Google Scholar] [CrossRef]
- Ravilious, G.E.; Jez, J.M. Nucleotide Binding Site Communication in Arabidopsis thaliana Adenosine 5′-Phosphosulfate Kinase. J. Biol. Chem. 2012, 287, 30385–30394. [Google Scholar] [CrossRef]
- Yatusevich, R.; Mugford, S.G.; Matthewman, C.; Gigolashvili, T.; Frerigmann, H.; Delaney, S.; Koprivova, A.; Flügge, U.-I.; Kopriva, S. Genes of Primary Sulfate Assimilation Are Part of the Glucosinolate Biosynthetic Network in Arabidopsis thaliana. Plant J. 2010, 62, 1–11. [Google Scholar] [CrossRef]
- Hirschmann, F.; Papenbrock, J. The Fusion of Genomes Leads to More Options: A Comparative Investigation on the Desulfo-Glucosinolate Sulfotransferases of Brassica Napus and Homologous Proteins of Arabidopsis thaliana. Plant Physiol. Biochem. 2015, 91, 10–19. [Google Scholar] [CrossRef]
- Klein, M.; Reichelt, M.; Gershenzon, J.; Papenbrock, J. The Three Desulfoglucosinolate Sulfotransferase Proteins in Arabidopsis Have Different Substrate Specificities and Are Differentially Expressed. FEBS J. 2006, 273, 122–136. [Google Scholar] [CrossRef]
- Klein, M.; Papenbrock, J. Kinetics and Substrate Specificities of Desulfo-Glucosinolate Sulfotransferases in Arabidopsis thaliana. Physiol. Plant. 2009, 135, 140–149. [Google Scholar] [CrossRef]
- Klein, M.; Papenbrock, J. The Multi-Protein Family of Arabidopsis Sulphotransferases and Their Relatives in Other Plant Species. J. Exp. Bot. 2004, 55, 1809–1820. [Google Scholar] [CrossRef]
- Piotrowski, M.; Schemenewitz, A.; Lopukhina, A.; Müller, A.; Janowitz, T.; Weiler, E.W.; Oecking, C. Desulfoglucosinolate Sulfotransferases from Arabidopsis thaliana Catalyze the Final Step in the Biosynthesis of the Glucosinolate Core Structure. J. Biol. Chem. 2004, 279, 50717–50725. [Google Scholar] [CrossRef]
- Hopkins, R.J.; van Dam, N.M.; van Loon, J.J.A. Role of Glucosinolates in Insect-Plant Relationships and Multitrophic Interactions. Annu. Rev. Entomol. 2009, 54, 57–83. [Google Scholar] [CrossRef]
- Ouassou, M.; Mukhaimar, M.; El Amrani, A.; Kroymann, J.; Chauveau, O. Biosynthesis of indole glucosinolates and ecological role of secondary modification pathways. Comptes Rendus Biol. 2019, 342, 58–80. [Google Scholar] [CrossRef]
- Sønderby, I.E.; Geu-Flores, F.; Halkier, B.A. Biosynthesis of Glucosinolates—Gene Discovery and Beyond. Trends Plant Sci. 2010, 15, 283–290. [Google Scholar] [CrossRef]
- Brown, P.D.; Tokuhisa, J.G.; Reichelt, M.; Gershenzon, J. Variation of Glucosinolate Accumulation among Different Organs and Developmental Stages of Arabidopsis thaliana. Phytochemistry 2003, 62, 471–481. [Google Scholar] [CrossRef]
- Petersen, B.; Chen, S.; Hansen, C.; Olsen, C.; Halkier, B. Composition and Content of Glucosinolates in Developing Arabidopsis thaliana. Planta 2002, 214, 562–571. [Google Scholar] [CrossRef]
- Giamoustaris, A.; Mithen, R. Genetics of Aliphatic Glucosinolates. IV. Side-Chain Modification in Brassica oleracea. Theor. Appl. Genet. 1996, 93, 1006–1010. [Google Scholar] [CrossRef]
- Hansen, B.G.; Kliebenstein, D.J.; Halkier, B.A. Identification of a Flavin-Monooxygenase as the S-Oxygenating Enzyme in Aliphatic Glucosinolate Biosynthesis in Arabidopsis. Plant J. 2007, 50, 902–910. [Google Scholar] [CrossRef] [PubMed]
- Kliebenstein, D.J.; Kroymann, J.; Brown, P.; Figuth, A.; Pedersen, D.; Gershenzon, J.; Mitchell-Olds, T. Genetic Control of Natural Variation in Arabidopsis Glucosinolate Accumulation. Plant Physiol. 2001, 126, 811–825. [Google Scholar] [CrossRef]
- Uda, Y.; Kurata, T.; Arakawa, N. Effects of PH and Ferrous Ion on the Degradation of Glucosinolates by Myrosinase. Agric. Biol. Chem. 1986, 50, 2735–2740. [Google Scholar] [CrossRef]
- Chen, S.; Petersen, B.L.; Olsen, C.E.; Schulz, A.; Halkier, B.A. Long-Distance Phloem Transport of Glucosinolates in Arabidopsis. Plant Physiol. 2001, 127, 194–201. [Google Scholar] [CrossRef]
- Ellerbrock, B.L.J.; Kim, J.H.; Jander, G. Contribution of Glucosinolate Transport to Arabidopsis Defense Responses. Plant Signal. Behav. 2007, 2, 282–283. [Google Scholar] [CrossRef]
- Du, L.; Ann Halkier, B. Biosynthesis of Glucosinolates in the Developing Silique Walls and Seeds of Sinapis alba. Phytochemistry 1998, 48, 1145–1150. [Google Scholar] [CrossRef]
- Léran, S.; Varala, K.; Boyer, J.-C.; Chiurazzi, M.; Crawford, N.; Daniel-Vedele, F.; David, L.; Dickstein, R.; Fernandez, E.; Forde, B.; et al. A Unified Nomenclature of Nitrate Transporter 1/Peptide Transporter Family Members in Plants. Trends Plant Sci. 2014, 19, 5–9. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Andersen, T.G.; Burow, M.; Madsen, S.R.; Jørgensen, M.E.; Olsen, C.E.; Dreyer, I.; Hedrich, R.; Geiger, D.; Halkier, B.A. NRT/PTR Transporters Are Essential for Translocation of Glucosinolate Defence Compounds to Seeds. Nature 2012, 488, 531–534. [Google Scholar] [CrossRef]
- Nambiar, D.M.; Kumari, J.; Augustine, R.; Kumar, P.; Bajpai, P.K.; Bisht, N.C. GTR1 and GTR2 Transporters Differentially Regulate Tissue-Specific Glucosinolate Contents and Defence Responses in the Oilseed Crop Brassica juncea. Plant Cell Environ. 2021, 44, 2729–2743. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Halkier, B.A. The Emerging Field of Transport Engineering of Plant Specialized Metabolites. Curr. Opin. Biotechnol. 2013, 24, 263–270. [Google Scholar] [CrossRef]
- Xu, D.; Hunziker, P.; Koroleva, O.; Blennow, A.; Crocoll, C.; Schulz, A.; Nour-Eldin, H.H.; Halkier, B.A. GTR-Mediated Radial Import Directs Accumulation of Defensive Glucosinolates to Sulfur-Rich Cells in the Phloem Cap of Arabidopsis Inflorescence Stem. Mol. Plant 2019, 12, 1474–1484. [Google Scholar] [CrossRef]
- Andersen, T.G.; Nour-Eldin, H.H.; Fuller, V.L.; Olsen, C.E.; Burow, M.; Halkier, B.A. Integration of Biosynthesis and Long-Distance Transport Establish Organ-Specific Glucosinolate Profiles in Vegetative Arabidopsis. Plant Cell 2013, 25, 3133–3145. [Google Scholar] [CrossRef]
- Jørgensen, M.E.; Nour-Eldin, H.H.; Halkier, B.A. Transport of Defense Compounds from Source to Sink: Lessons Learned from Glucosinolates. Trends Plant Sci. 2015, 20, 508–514. [Google Scholar] [CrossRef]
- Wittstock, U.; Kurzbach, E.; Herfurth, A.-M.; Stauber, E.J. Chapter Six—Glucosinolate Breakdown. In Advances in Botanical Research; Kopriva, S., Ed.; Glucosinolates; Academic Press: Cambridge, MA, USA, 2016; Volume 80, pp. 125–169. [Google Scholar]
- Poveda, J.; Eugui, D.; Velasco, P. Natural Control of Plant Pathogens through Glucosinolates: An Effective Strategy against Fungi and Oomycetes. Phytochem. Rev. 2020, 19, 104497. [Google Scholar] [CrossRef]
- Shofran, B.G.; Purrington, S.T.; Breidt, F.; Fleming, H.P. Antimicrobial Properties of Sinigrin and Its Hydrolysis Products. J. Food Sci. 1998, 63, 621–624. [Google Scholar] [CrossRef]
- Agrawal, A.A.; Kurashige, N.S. A Role for Isothiocyanates in Plant Resistance against the Specialist Herbivore Pieris Rapae. J. Chem. Ecol. 2003, 29, 1403–1415. [Google Scholar] [CrossRef]
- Aissani, N.; Tedeschi, P.; Maietti, A.; Brandolini, V.; Garau, V.L.; Caboni, P. Nematicidal Activity of Allylisothiocyanate from Horseradish (Armoracia rusticana) Roots against Meloidogyne incognita. J. Agric. Food Chem. 2013, 61, 4723–4727. [Google Scholar] [CrossRef]
- Jeschke, V.; Gershenzon, J.; Vassão, D.G. A Mode of Action of Glucosinolate-Derived Isothiocyanates: Detoxification Depletes Glutathione and Cysteine Levels with Ramifications on Protein Metabolism in Spodoptera littoralis. Insect Biochem. Mol. Biol. 2016, 71, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Tookey, H.L. Crambe Thioglucoside Glucohydrolase (EC 3.2.3.1): Separation of a Protein Required for Epithiobutane Formation. Can. J. Biochem. 1973, 51, 1654–1660. [Google Scholar] [CrossRef] [PubMed]
- Lambrix, V.; Reichelt, M.; Mitchell-Olds, T.; Kliebenstein, D.J.; Gershenzon, J. The Arabidopsis Epithiospecifier Protein Promotes the Hydrolysis of Glucosinolates to Nitriles and Influences Trichoplusia Ni Herbivory. Plant Cell 2001, 13, 2793–2807. [Google Scholar] [CrossRef] [PubMed]
- Wittstock, U.; Burow, M. Tipping the Scales--Specifier Proteins in Glucosinolate Hydrolysis. IUBMB Life 2007, 59, 744–751. [Google Scholar] [CrossRef]
- Kuchernig, J.C.; Burow, M.; Wittstock, U. Evolution of Specifier Proteins in Glucosinolate-Containing Plants. BMC Evol. Biol. 2012, 12, 127. [Google Scholar] [CrossRef]
- Jander, G.; Cui, J.; Nhan, B.; Pierce, N.E.; Ausubel, F.M. The TASTY Locus on Chromosome 1 of Arabidopsis Affects Feeding of the Insect Herbivore Trichoplusia Ni. Plant Physiol. 2001, 126, 890–898. [Google Scholar] [CrossRef]
- Mumm, R.; Burow, M.; Bukovinszkine’kiss, G.; Kazantzidou, E.; Wittstock, U.; Dicke, M.; Gershenzon, J. Formation of Simple Nitriles upon Glucosinolate Hydrolysis Affects Direct and Indirect Defense against the Specialist Herbivore, Pieris Rapae. J. Chem. Ecol. 2008, 34, 1311–1321. [Google Scholar] [CrossRef]
- de Vos, M.; Kriksunov, K.L.; Jander, G. Indole-3-Acetonitrile Production from Indole Glucosinolates Deters Oviposition by Pieris rapae. Plant Physiol. 2008, 146, 916–926. [Google Scholar] [CrossRef]


| Classes | Iupac Name | Tissue | Bibliography |
|---|---|---|---|
| Aliphatic glucosinolates | 3-(methylthio)propyl glucosinolate | Whole plant | [66] |
| 9-(methylthio)nonyl glucosinolate | Whole plant | [66] | |
| 3-(methylsulfinyl)propyl glucosinolate | Whole plant | [66] | |
| 4-(methylsulfinyl)butyl glucosinolate | Whole plant | [66] | |
| 8-(methylsulfinyl)octyl glucosinolate | Whole plant | [66] | |
| 9-(methylsulfinyl)nonyl glucosinolate * | Whole plant, seeds, root exudates | [66,67,70] | |
| 10-(methylsulfinyl)decyl glucosinolate * | Whole plant, seeds, root exudates | [66,67,70] | |
| 11-(methylsulfinyl)undecyl-glucosinolate * | seed, root exudates | [66,67,70] | |
| Indole glucosinolates | 3-indolylmethyl glucosinolate | Whole plant | [66,67,70] |
| 4-hydroxy-3-indolylmethyl glucosinolate | Whole plant | [66] | |
| 4-methoxy-3-indolylmethyl glucosinolate | Whole plant | [66] | |
| ds-glucobrassicin | Seedlings | [71] | |
| ds-4-methoxy-glucobrassicin | Seedlings | [71] | |
| ds-neoglucobrassicin | Seedlings | [71] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghidoli, M.; Pesenti, M.; Colombo, F.; Nocito, F.F.; Pilu, R.; Araniti, F. Camelina sativa (L.) Crantz as a Promising Cover Crop Species with Allelopathic Potential. Agronomy 2023, 13, 2187. https://doi.org/10.3390/agronomy13082187
Ghidoli M, Pesenti M, Colombo F, Nocito FF, Pilu R, Araniti F. Camelina sativa (L.) Crantz as a Promising Cover Crop Species with Allelopathic Potential. Agronomy. 2023; 13(8):2187. https://doi.org/10.3390/agronomy13082187
Chicago/Turabian StyleGhidoli, Martina, Michele Pesenti, Federico Colombo, Fabio Francesco Nocito, Roberto Pilu, and Fabrizio Araniti. 2023. "Camelina sativa (L.) Crantz as a Promising Cover Crop Species with Allelopathic Potential" Agronomy 13, no. 8: 2187. https://doi.org/10.3390/agronomy13082187
APA StyleGhidoli, M., Pesenti, M., Colombo, F., Nocito, F. F., Pilu, R., & Araniti, F. (2023). Camelina sativa (L.) Crantz as a Promising Cover Crop Species with Allelopathic Potential. Agronomy, 13(8), 2187. https://doi.org/10.3390/agronomy13082187










