Genome-Wide Analysis of the HD-Zip Gene Family in Chinese Cabbage (Brassica rapa subsp. pekinensis) and the Expression Pattern at High Temperatures and in Carotenoids Regulation
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials, Heat Stress Treatment, and Carotenoid Content Measurement
2.2. GenomeWide Identification of HD-Zip Genes
2.3. Multiple Sequence Alignment and Phylogenetic Analysis
2.4. Gene Structure, Motif, and Cis-Regulatory Elements Analysis
2.5. Synteny Analysis of HD-Zip Genes
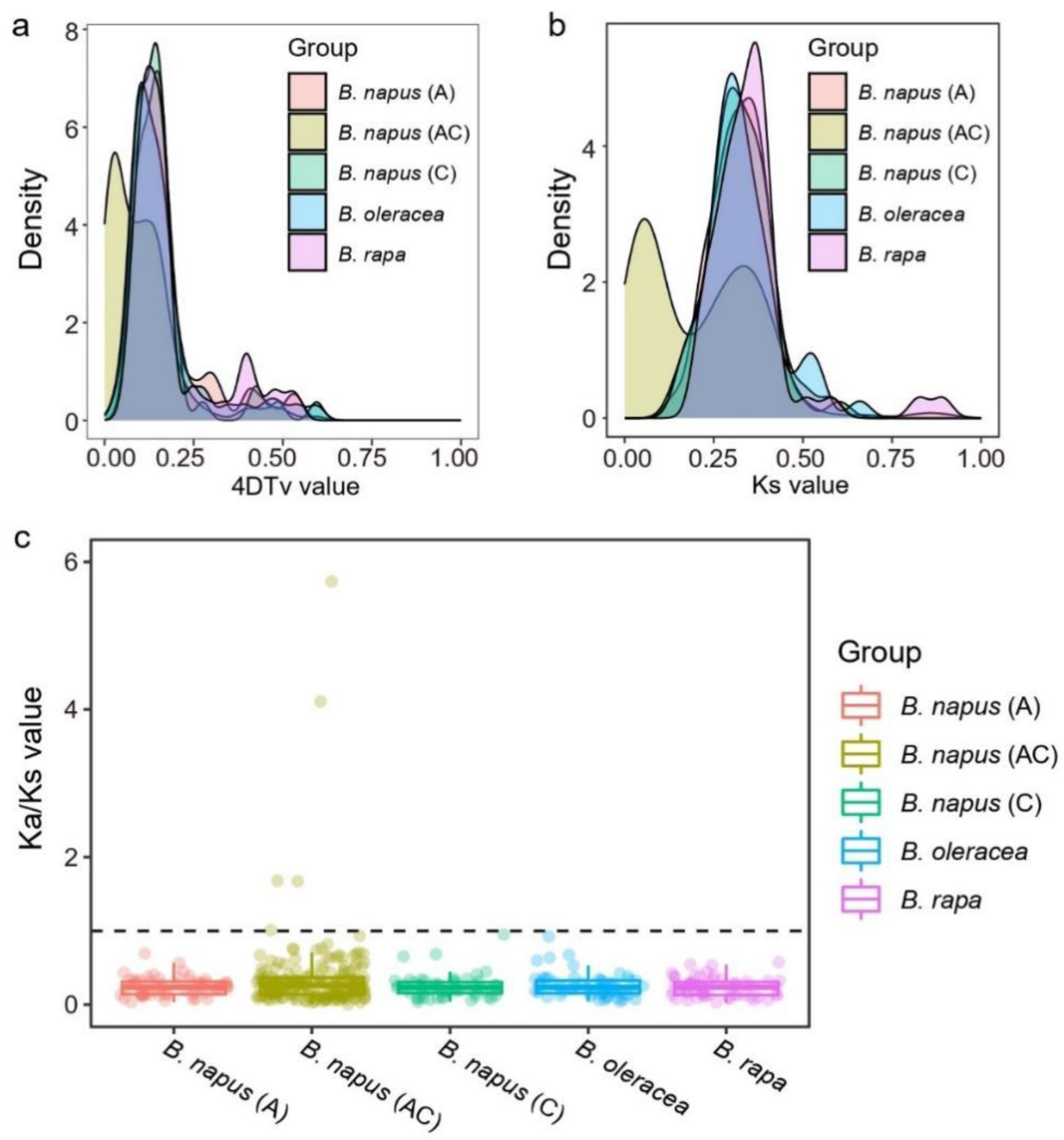
2.6. Calculating the Ka, Ks, and 4DTv of HD-Zip Paralogs
2.7. Expression Pattern Analysis of HD-Zip Genes
3. Results
3.1. Whole-Genome Identification of HD-Zip Genes in Brassicaceae Plants
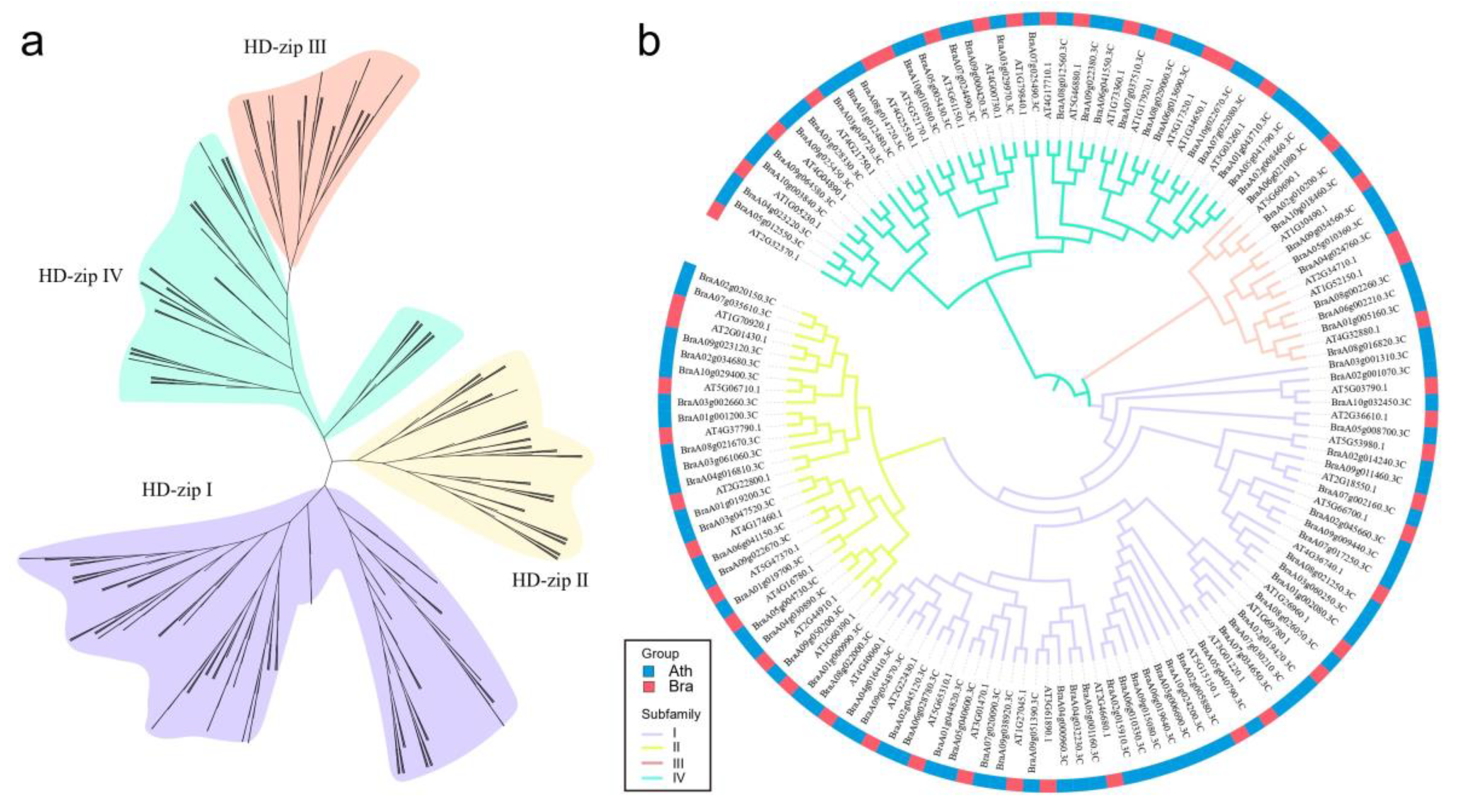
3.2. Phylogenetic Analysis of the HD-Zip Genes
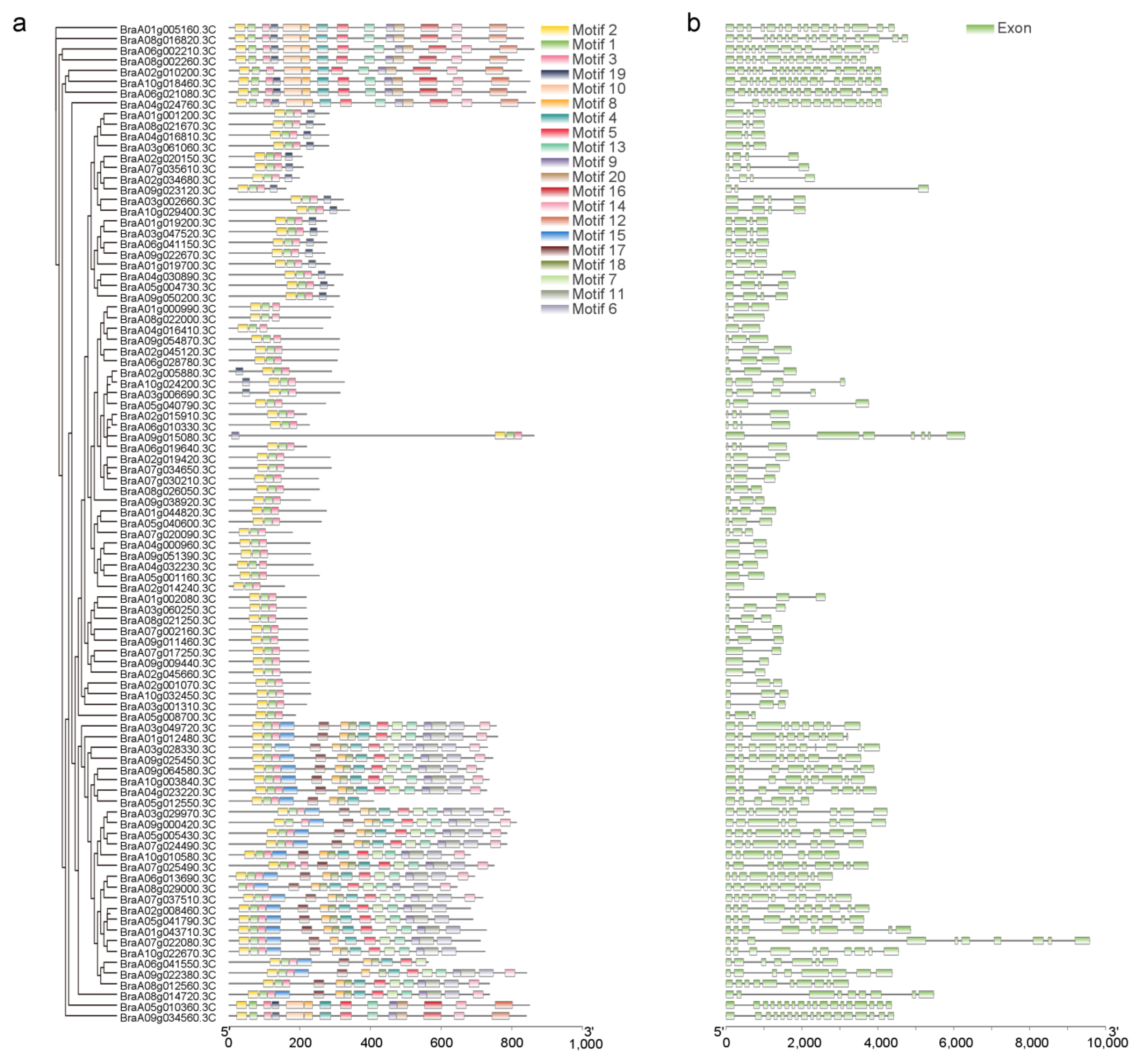
3.3. Conserved Motif Analysis and Gene Structural Analysis of HD-Zip Genes
3.4. Cis-Acting Elements Analysis in the Putative Promoter of HD-Zip Genes
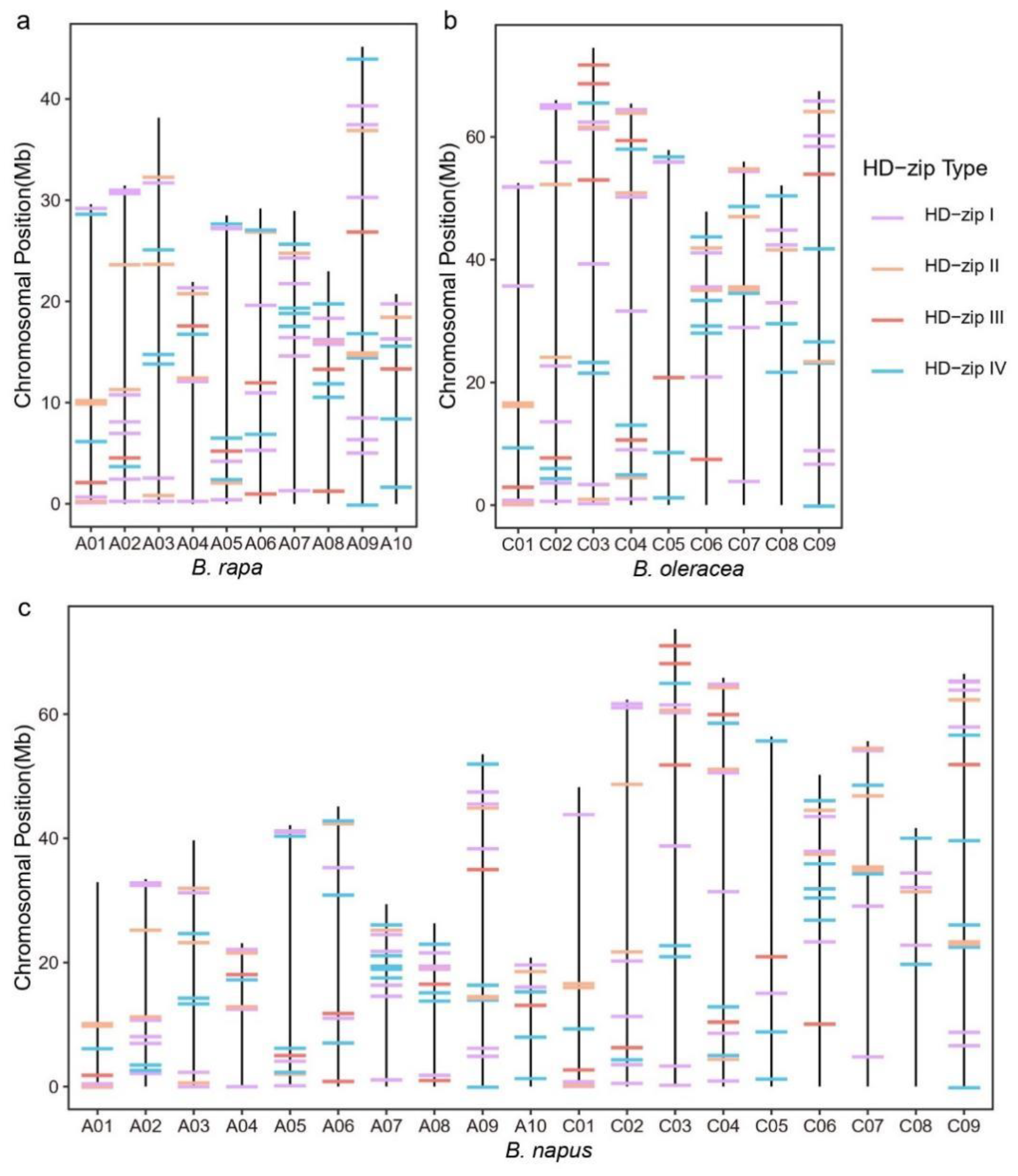
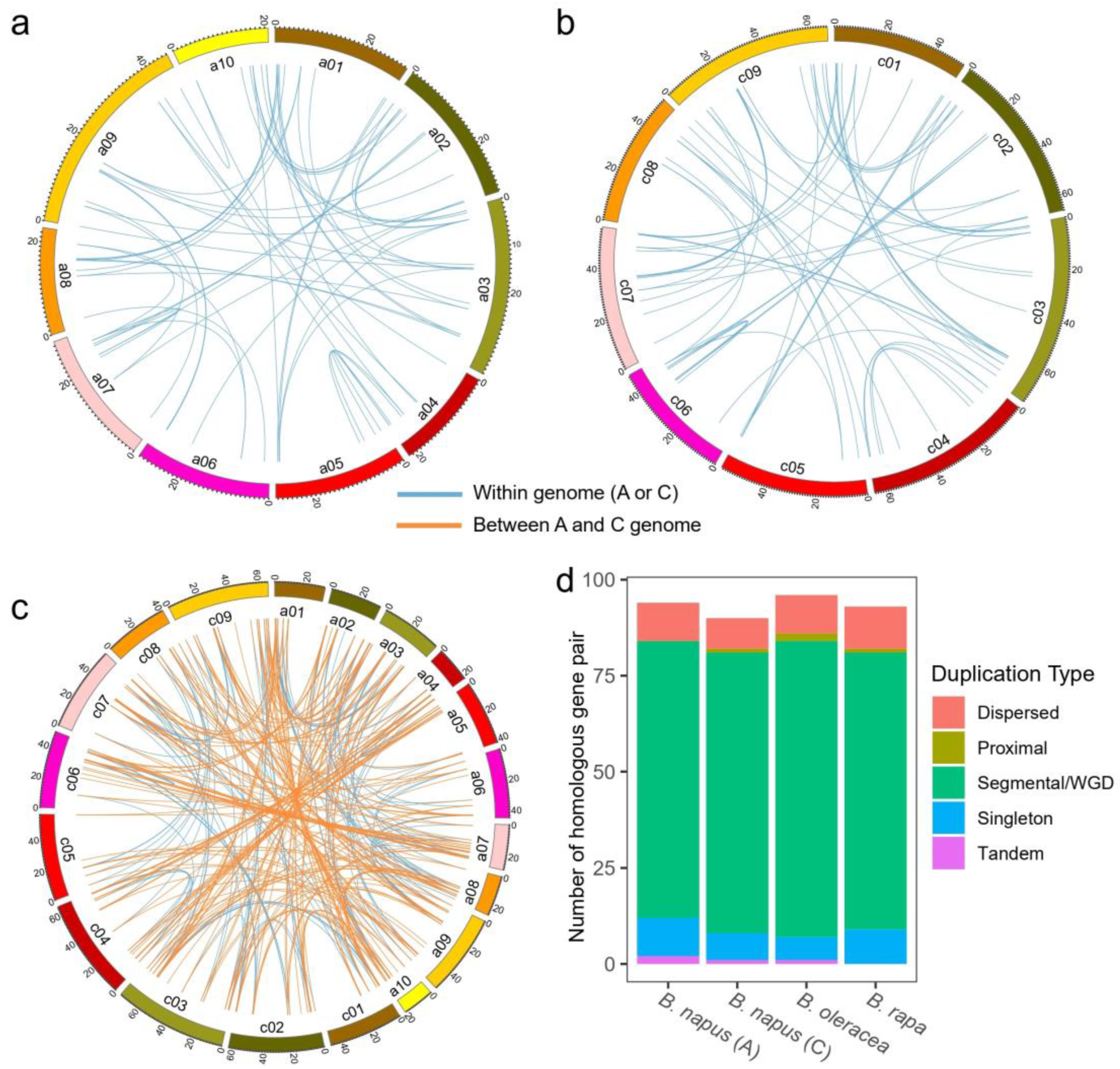
3.5. Chromosome Location and Gene Family Expansion Analysis of HD-Zip Genes in Brassicaceae Plants
3.6. Estimating Dates and Driving Forces for the Evolution of the HD-Zip Gene Family
3.7. Expression Patterns of HD-Zip Genes in Different Chinese Cabbage Varieties
4. Discussion
4.1. Whole-Genome Identification and Phylogenetic Analysis of HD-Zip Genes in Chinese Cabbage
4.2. The Evolution History of the HD-Zip Gene Family
4.3. The Potential Roles of Chinese Cabbage HD-Zip Transcription Factors
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Aso, K.; Kato, M.; Banks, J.A.; Hasebe, M. Characterization of homeodomain-leucine zipper genes in the fern Ceratopteris richardii and the evolution of the homeodomain-leucine zipper gene family in vascular plants. Mol. Biol. Evol. 1999, 16, 544–552. [Google Scholar] [CrossRef]
- Sakakibara, K.; Nishiyama, T.; Sumikawa, N.; Kofuji, R.; Murata, T.; Hasebe, M. Involvement of auxin and a homeodomain-leucine zipper I gene in rhizoid development of the moss Physcomitrella patens. Development 2003, 130, 4835–4846. [Google Scholar] [CrossRef] [PubMed]
- Kamata, N.; Okada, H.; Komeda, Y.; Takahashi, T. Mutations in epidermis-specific HD-ZIP IV genes affect floral organ identity in Arabidopsis thaliana. Plant J. 2013, 75, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Fu, L.; Yan, Y.; Tie, W.; Xia, Z.; Wang, W.; Peng, M.; Hu, W.; Zhang, J.; Zhang, J. Genome-wide characterization and expression profiling of HD-Zip gene family related to abiotic stress in cassava. PLoS ONE 2017, 12, e0173043. [Google Scholar] [CrossRef]
- Vernoud, V.; Laigle, G.; Rozier, F.; Meeley, R.B.; Perez, P.; Rogowsky, P.M. The HD-ZIP IV transcription factor OCL4 is necessary for trichome patterning and anther development in maize. Plant J. 2009, 59, 883–894. [Google Scholar] [CrossRef] [PubMed]
- Sharif, R.; Raza, A.; Chen, P.; Li, Y.; El-Ballat, E.M.; Rauf, A.; Hano, C.; El-Esawi, M.A. HD-ZIP Gene Family: Potential roles in improving plant growth and regulating stress-responsive mechanisms in plants. Genes 2021, 12, 1256. [Google Scholar] [CrossRef]
- Elhiti, M.; Stasolla, C. Structure and function of homodomain-leucine zipper (HD-Zip) proteins. Plant Signal Behav. 2009, 4, 86–88. [Google Scholar] [CrossRef]
- Harris, J.C.; Hrmova, M.; Lopato, S.; Langridge, P. Modulation of plant growth by HD-Zip class I and II transcription factors in response to environmental stimuli. New Phytol. 2011, 190, 823–837. [Google Scholar] [CrossRef]
- Zhang, R.X.; Zhu, W.C.; Cheng, G.X.; Yu, Y.N.; Li, Q.H.; ul Haq, S.; Said, F.; Gong, Z.H. A novel gene, CaATHB-12, negatively regulates fruit carotenoid content under cold stress in Capsicum annuum. Food Nutr. Res. 2020, 64, 3729. [Google Scholar] [CrossRef]
- Roodbarkelari, F.; Groot, E.P. Regulatory function of homeodomain-leucine zipper (HD-ZIP) family proteins during embryogenesis. New Phytol. 2017, 213, 95–104. [Google Scholar] [CrossRef]
- Turchi, L.; Baima, S.; Morelli, G.; Ruberti, I. Interplay of HD-Zip II and III transcription factors in auxin-regulated plant development. J. Exp. Bot. 2015, 66, 5043–5053. [Google Scholar] [CrossRef]
- Takada, S.; Takada, N.; Yoshida, A. ATML1 promotes epidermal cell differentiation in Arabidopsis shoots. Development 2013, 140, 1919–1923. [Google Scholar] [CrossRef]
- Ursache, R.; Miyashima, S.; Chen, Q.; Vatén, A.; Nakajima, K.; Carlsbecker, A.; Zhao, Y.; Helariutta, Y.; Dettmer, J. Tryptophan-dependent auxin biosynthesis is required for HD-ZIP III-mediated xylem patterning. Development 2014, 141, 1250–1260. [Google Scholar] [CrossRef]
- Sun, X.X.; Feng, D.; Liu, M.; Qin, R.; Li, Y.; Lu, Y.; Zhang, X.; Wang, Y.; Shen, S.; Ma, W.; et al. Single-cell transcriptome reveals dominant subgenome expression and transcriptional response to heat stress in Chinese cabbage. Genome. Biol. 2022, 23, 1–19. [Google Scholar] [CrossRef]
- Zhang, L.; Dai, Y.; Yue, L.; Chen, G.; Yuan, L.; Zhang, S.; Li, F.; Zhang, H.; Li, G.; Zhu, S.; et al. Heat stress response in Chinese cabbage (Brassica rapa L.) revealed by transcriptome and physiological analysis. Peerj 2022, 10, e13427. [Google Scholar] [CrossRef]
- Quan, J.; Zheng, W.; Wu, M.; Shen, Z.; Tan, J.; Li, Z.; Zhu, B.; Hong, S.B.; Zhao, Y.; Zhu, Z.; et al. Glycine betaine and beta-aminobutyric acid mitigate the detrimental effects of heat stress on Chinese cabbage (Brassica rapa L. ssp. pekinensis) Seedlings with Improved Photosynthetic Performance and Antioxidant System. Plants 2022, 11, 1213. [Google Scholar] [CrossRef]
- Song, X.; Liu, G.; Duan, W.; Liu, T.; Huang, Z.; Ren, J.; Hou, X. Genome-wide identification, classification and expression analysis of the heat shock transcription factor family in Chinese cabbage. Mol. Genet. Genomics. 2014, 289, 541–551. [Google Scholar] [CrossRef]
- Nakashima, K.; Ito, Y.; Yamaguchi-Shinozaki, K. Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol. 2009, 149, 88–95. [Google Scholar] [CrossRef]
- Wu, Z.; Li, T.; Zhang, D.; Teng, N. Lily HD-Zip I transcription factor LlHB16 promotes thermotolerance by activating LlHSFA2 and LlMBF1c. Plant Cell Physiol. 2022, 63, 1729–1744. [Google Scholar] [CrossRef]
- Li, W.; Dong, J.; Cao, M.; Gao, X.; Wang, D.; Liu, B.; Chen, Q. Genome-wide identification and characterization of HD-ZIP genes in potato. Gene 2019, 697, 103–117. [Google Scholar] [CrossRef]
- Wang, K.; Xu, L.; Wang, Y.; Ying, J.; Li, J.; Dong, J.; Li, C.; Zhang, X.; Liu, L. Genome-wide characterization of homeodomain-leucine zipper genes reveals RsHDZ17 enhances the heat tolerance in radish (Raphanus sativus L.). Physiol. Plantarum. 2022, 174, e13789. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhuang, L.; Zhang, J.; Yu, J.; Yang, Z.; Huang, B. Identification and characterization of novel homeodomain leucine zipper (HD-Zip) transcription factors associated with heat tolerance in perennial ryegrass. Environ. Exp. Bot. 2019, 160, 1–11. [Google Scholar] [CrossRef]
- Zhao, J.G.; Lu, Z.G.; Wang, L.; Jin, B. Plant responses to heat stress: Physiology, transcription, noncoding RNAs, and epigenetics. Int. J. Mol. Sci. 2020, 22, 117. [Google Scholar] [CrossRef]
- Inbaraj, B.S.; Lu, H.; Hung, C.F.; Wu, W.B.; Lin, C.L.; Chen, B.H. Determination of carotenoids and their esters in fruits of Lycium barbarum Linnaeus by HPLC-DAD-APCI-MS. J. Pharm. Biomed. 2008, 47, 812–818. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic. Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Johnson, L.S.; Eddy, S.R.; Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinform. 2010, 11, 431. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic. Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Zhang, Z.; Xiao, J.; Wu, J.; Zhang, H.; Liu, G.; Wang, X.; Dai, L. ParaAT: A parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 2012, 419, 779–781. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A toolkit incorporating gamma-series methods and sliding window strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, J.; Yu, J. Computing Ka and Ks with a consideration of unequal transitional substitutions. BMC Evol. Biol. 2006, 6, 44. [Google Scholar] [CrossRef]
- Yue, L.X.; Li, G.; Dai, Y.; Sun, X.; Li, F.; Zhang, S.; Zhang, H.; Sun, R.; Zhang, S. Gene co-expression network analysis of the heat-responsive core transcriptome identifies hub genes in Brassica rapa. Planta 2021, 253, 1–23. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Zhang, Z.; Guo, J.; Cai, X.; Li, Y.; Xi, X.; Lin, R.; Liang, J.; Wang, X.; Wu, J. Improved reference genome annotation of Brassica rapa by pacific biosciences RNA sequencing. Front. Plant Sci. 2022, 13, 841618. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Cai, X.; Chang, L.; Zhang, T.; Chen, H.; Zhang, L.; Lin, R.; Liang, J.; Wu, J.; Freeling, M.; Wang, X. Impacts of allopolyploidization and structural variation on intraspecific diversification in Brassica rapa. Genome. Biol. 2021, 22, 166. [Google Scholar] [CrossRef]
- Zhao, J.; Zhai, Z.; Li, Y.; Geng, S.; Song, G.; Guan, J.; Jia, M.; Wang, F.; Sun, G.; Feng, N. Genome-wide identification and expression profiling of the TCP family genes in spike and grain development of wheat (Triticum aestivum L.). Front. Plant Sci. 2018, 9, 1282. [Google Scholar] [CrossRef]
- He, G.H.; Liu, P.; Zhao, H.X.; Sun, J.Q. The HD-ZIP II transcription factors regulate plant architecture through the auxin pathway. Int. J. Mol. Sci. 2020, 21, 3250. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Sun, R.; Hou, X.; Zheng, H.; Zhang, F.; Zhang, Y.; Liu, B.; Liang, J.; Zhuang, M.; Liu, Y.; et al. Subgenome parallel selection is associated with morphotype diversification and convergent crop domestication in Brassica rapa and Brassica oleracea. Nat. Genet. 2016, 48, 1218–1224. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Song, B.; Dai, Y.; Zhang, S.; Huang, X. Genome-wide identification and functional analysis of U-box E3 ubiquitin ligases gene family related to drought stress response in Chinese white pear (Pyrus bretschneideri). Bmc Plant Biol. 2021, 21, 235. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.X.; Haider, I.; Kohlen, W.; Jiang, L.; Bouwmeester, H.; Meijer, A.H.; Schluepmann, H.; Liu, C.M.; Ouwerkerk, P.B. Function of the HD-Zip I gene Oshox22 in ABA-mediated drought and salt tolerances in rice. Plant Mol. Biol. 2012, 80, 571–585. [Google Scholar] [CrossRef]
- Zhang, J.S.; Wu, J.; Guo, M.; Aslam, M.; Wang, Q.; Ma, H.; Li, S.; Zhang, X.; Cao, S. Genome-wide characterization and expression profiling of Eucalyptus grandis HD-Zip gene family in response to salt and temperature stress. Bmc Plant Biol. 2020, 20, 1–15. [Google Scholar] [CrossRef]
- Wang, D.; Gong, Y.; Li, Y.; Nie, S.M. Genome-wide analysis of the homeodomain-leucine zipper family in Lotus japonicus and the overexpression of LjHDZ7 in Arabidopsis for salt tolerance. Front. Plant Sci. 2022, 13, 955199. [Google Scholar] [CrossRef]
- Gonzalez-Grandio, E.; Pajoro, A.; Franco-Zorrilla, J.M.; Tarancón, C.; Immink, R.G.; Cubas, P. Abscisic acid signaling is controlled by a BRANCHED1/HD-ZIP I cascade in Arabidopsis axillary buds. Proc. Natl. Acad. Sci. USA 2017, 114, E245–E254. [Google Scholar] [CrossRef]
- Sun, Y.; Bai, P.P.; Gu, K.J.; Yang, S.Z.; Lin, H.Y.; Shi, C.G.; Zhao, Y.P. Dynamic transcriptome and network-based analysis of yellow leaf mutant Ginkgo biloba. Bmc Plant Biol. 2022, 22, 465. [Google Scholar] [CrossRef]
- Zhou, H.; Wang, Y.; Zhang, Y.; Xiao, Y.; Liu, X.; Deng, H.; Lu, X.; Tang, W.; Zhang, G. Comparative analysis of heat-tolerant and heat-susceptible rice highlights the role of OsNCED1 gene in heat stress tolerance. Plants 2022, 11, 1062. [Google Scholar] [CrossRef]
- Li, Y.; Yang, Z.; Zhang, Y.; Guo, J.; Liu, L.; Wang, C.; Wang, B.; Han, G. The roles of HD-ZIP proteins in plant abiotic stress tolerance. Front. Plant Sci. 2022, 13, 1027071. [Google Scholar] [CrossRef]
- Tang, Y.; Wang, J.; Bao, X.; Liang, M.; Lou, H.; Zhao, J.; Sun, M.; Liang, J.; Jin, L.; Li, G.; et al. Genome-wide identification and expression profile of HD-ZIP genes in physic nut and functional analysis of the JcHDZ16 gene in transgenic rice. Bmc Plant Biol. 2019, 19, 298. [Google Scholar] [CrossRef]
- Wang, Z.; Wu, X.; Zhang, B.; Xiao, Y.; Guo, J.; Liu, J.; Chen, Q.; Peng, F. Genome-wide identification, bioinformatics and expression analysis of HD-Zip gene family in peach. Bmc Plant Biol. 2023, 23, 122. [Google Scholar] [CrossRef]
- Sharif, R.; Xie, C.; Wang, J.; Cao, Z.; Zhang, H.; Chen, P.; Li, Y.H. Genome wide identification, characterization and expression analysis of HD-ZIP gene family in Cucumis sativus L. under biotic and various abiotic stresses. Int. J. Biol. Macromol. 2020, 158, 502–520. [Google Scholar] [CrossRef]
- Song, J.M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef]
- Zhao, S.; Wang, H.; Jia, X.; Gao, H.; Mao, K.; Ma, F. The HD-Zip I transcription factor MdHB7-like confers tolerance to salinity in transgenic apple (Malus domestica). Physiol. Plant 2021, 172, 1452–1464. [Google Scholar] [CrossRef]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R.; Zhang, S.; Paterson, A.H. Gene duplication and evolution in recurring polyploidization-diploidization cycles in plants. Genome. Biol. 2019, 20, 38. [Google Scholar] [CrossRef]
- Yu, J.Y.; Tehrim, S.; Zhang, F.; Tong, C.; Huang, J.; Cheng, X.; Dong, C.; Zhou, Y.; Qin, R.; Hua, W.; et al. Genome-wide comparative analysis of NBS-encoding genes between Brassica species and Arabidopsis thaliana. Bmc Genom. 2014, 15, 3. [Google Scholar] [CrossRef]
- Yang, Z.H.; Nielsen, R. Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol. Biol. Evol. 2002, 19, 908–917. [Google Scholar] [CrossRef]
- Sabeti, P.C.; Schaffner, S.F.; Fry, B.; Lohmueller, J.; Varilly, P.; Shamovsky, O.; Palma, A.; Mikkelsen, T.S.; Altshuler, D.; Lander, E.S. Positive natural selection in the human lineage. Science 2006, 312, 1614–1620. [Google Scholar] [CrossRef]
- Umezawa, T.; Fujita, M.; Fujita, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Engineering drought tolerance in plants: Discovering and tailoring genes to unlock the future. Curr. Opin. Biotechnol. 2006, 17, 113–122. [Google Scholar] [CrossRef]
- Valliyodan, B.; Nguyen, H.T. Understanding regulatory networks and engineering for enhanced drought tolerance in plants. Curr. Opin. Plant Biol. 2006, 9, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Ma, Q.; Jin, X.; Peng, X.; Liu, J.; Deng, L.; Yan, H.; Sheng, L.; Jiang, H.; Cheng, B. A novel maize homeodomain-leucine zipper (HD-Zip) I gene, Zmhdz10, positively regulates drought and salt tolerance in both rice and Arabidopsis. Plant Cell Physiol. 2014, 55, 1142–1156. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Fan, Y.; Jiao, Y.; Wu, J.; Zhang, Z.; Yu, X.; Ma, Y. Transcriptome profiling of yellow leafy head development during the heading stage in Chinese cabbage (Brassica rapa subsp. pekinensis). Physiol. Plant 2019, 165, 800–813. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.J.; Manoharan, R.K.; Park, J.I.; Chung, M.Y.; Lee, J.; Lim, Y.P.; Hur, Y.; Nou, I.S. Identification of yellow pigmentation genes in Brassica rapa ssp. pekinensis using Br300 microarray. Int. J. Genom. 2014, 2014, 204969. [Google Scholar] [CrossRef]
- Yuan, P.; Umer, M.J.; He, N.; Zhao, S.; Lu, X.; Zhu, H.; Gong, C.; Diao, W.; Gebremeskel, H.; Kuang, H.; et al. Transcriptome regulation of carotenoids in five flesh-colored watermelons (Citrullus lanatus). Bmc Plant Biol. 2021, 21, 203. [Google Scholar] [CrossRef]
- Huang, Q.; Liu, J.; Hu, C.; Wang, N.; Zhang, L.; Mo, X.; Li, G.; Liao, H.; Huang, H.; Ji, S.; et al. Integrative analyses of transcriptome and carotenoids profiling revealed molecular insight into variations in fruits color of Citrus Reticulata Blanco induced by transplantation. Genomics 2022, 114, 110291. [Google Scholar] [CrossRef]
- Jiang, F.; Zhang, J.; Wang, S.; Yang, L.; Luo, Y.; Gao, S.; Zhang, M.; Wu, S.; Hu, S.; Sun, H.; et al. The apricot (Prunus armeniaca L.) genome elucidates Rosaceae evolution and beta-carotenoid synthesis. Hortic. Res. 2019, 6, 128. [Google Scholar] [CrossRef]



| Species | Genome Size | Chromosome Number (2n) | Whole Gene Number | HD-Zip Gene Number | |||
|---|---|---|---|---|---|---|---|
| I | II | III | IV | ||||
| Brassica rapa | 351.06 | 20 | 46,250 | 39 | 18 | 10 | 26 |
| Brassica oleracea | 561.16 | 18 | 59,064 | 41 | 20 | 10 | 25 |
| Brassica napus | 924 | 38 | 108,190 | 74 | 38 | 19 | 53 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yin, L.; Sun, Y.; Chen, X.; Liu, J.; Feng, K.; Luo, D.; Sun, M.; Wang, L.; Xu, W.; Liu, L.; et al. Genome-Wide Analysis of the HD-Zip Gene Family in Chinese Cabbage (Brassica rapa subsp. pekinensis) and the Expression Pattern at High Temperatures and in Carotenoids Regulation. Agronomy 2023, 13, 1324. https://doi.org/10.3390/agronomy13051324
Yin L, Sun Y, Chen X, Liu J, Feng K, Luo D, Sun M, Wang L, Xu W, Liu L, et al. Genome-Wide Analysis of the HD-Zip Gene Family in Chinese Cabbage (Brassica rapa subsp. pekinensis) and the Expression Pattern at High Temperatures and in Carotenoids Regulation. Agronomy. 2023; 13(5):1324. https://doi.org/10.3390/agronomy13051324
Chicago/Turabian StyleYin, Lian, Yudong Sun, Xuehao Chen, Jiexia Liu, Kai Feng, Dexu Luo, Manyi Sun, Linchuang Wang, Wenzhao Xu, Lu Liu, and et al. 2023. "Genome-Wide Analysis of the HD-Zip Gene Family in Chinese Cabbage (Brassica rapa subsp. pekinensis) and the Expression Pattern at High Temperatures and in Carotenoids Regulation" Agronomy 13, no. 5: 1324. https://doi.org/10.3390/agronomy13051324
APA StyleYin, L., Sun, Y., Chen, X., Liu, J., Feng, K., Luo, D., Sun, M., Wang, L., Xu, W., Liu, L., & Zhao, J. (2023). Genome-Wide Analysis of the HD-Zip Gene Family in Chinese Cabbage (Brassica rapa subsp. pekinensis) and the Expression Pattern at High Temperatures and in Carotenoids Regulation. Agronomy, 13(5), 1324. https://doi.org/10.3390/agronomy13051324






