Hyperspectral Sensing of Plant Diseases: Principle and Methods
Abstract
:1. Introduction
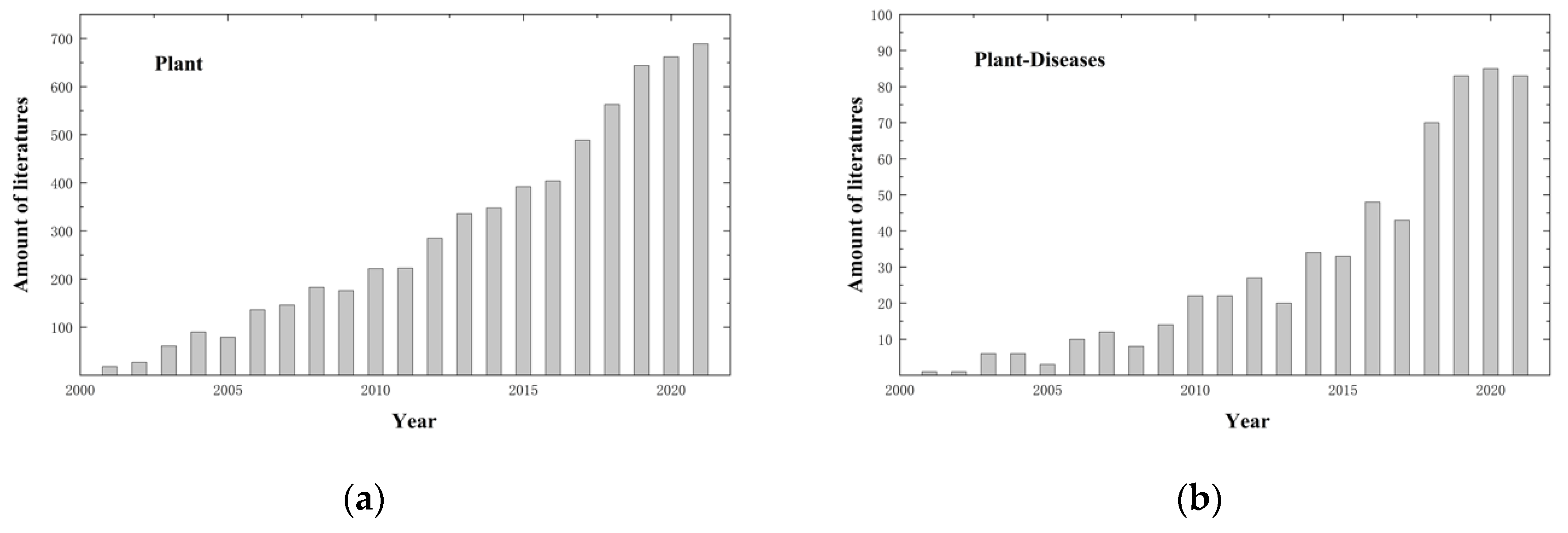
2. Literature Overview
3. Variation Mechanism of Spectral Information and Photo Response of Plant Diseases
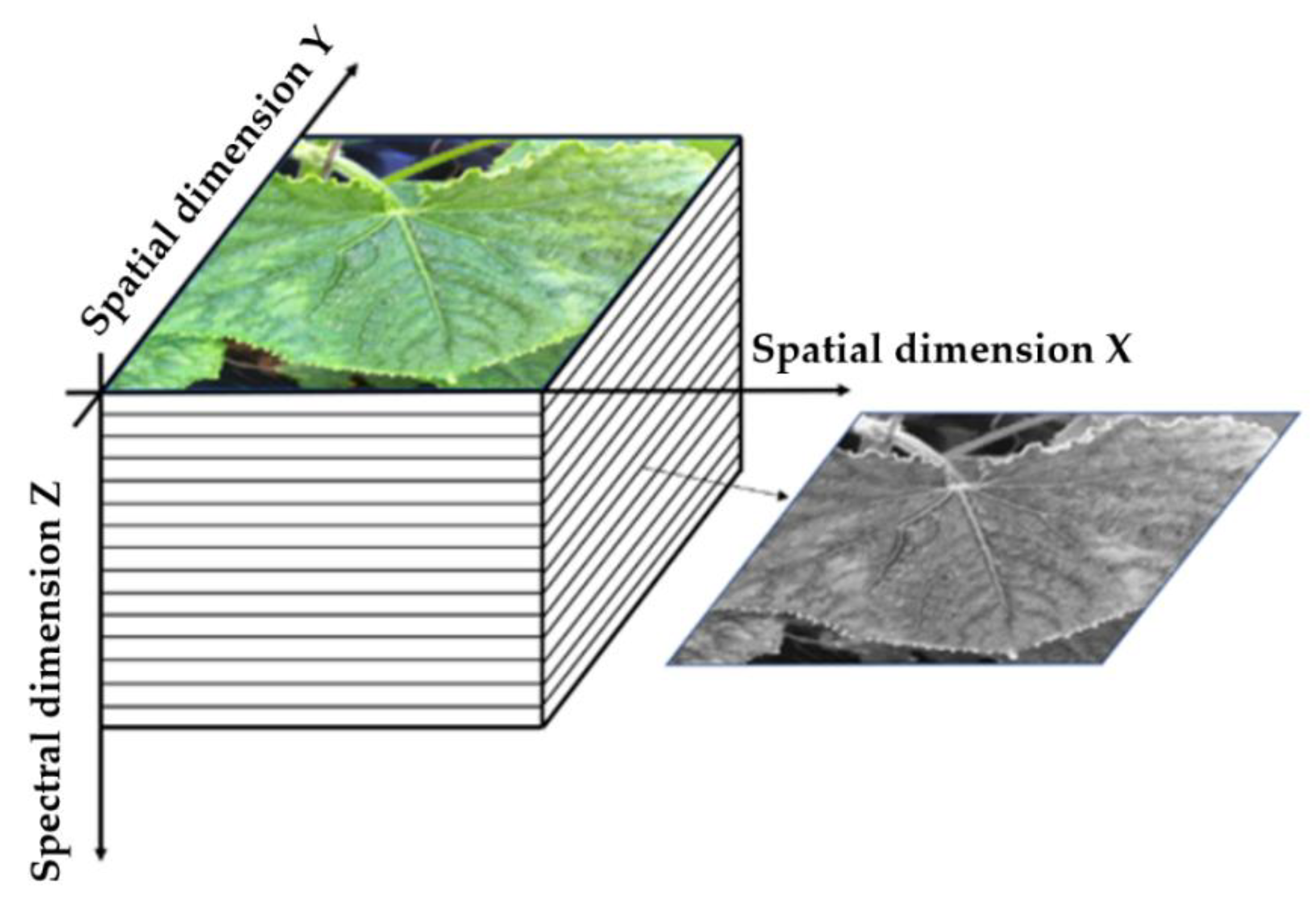
3.1. Principles of Hyperspectral Imaging Technology and Sensors
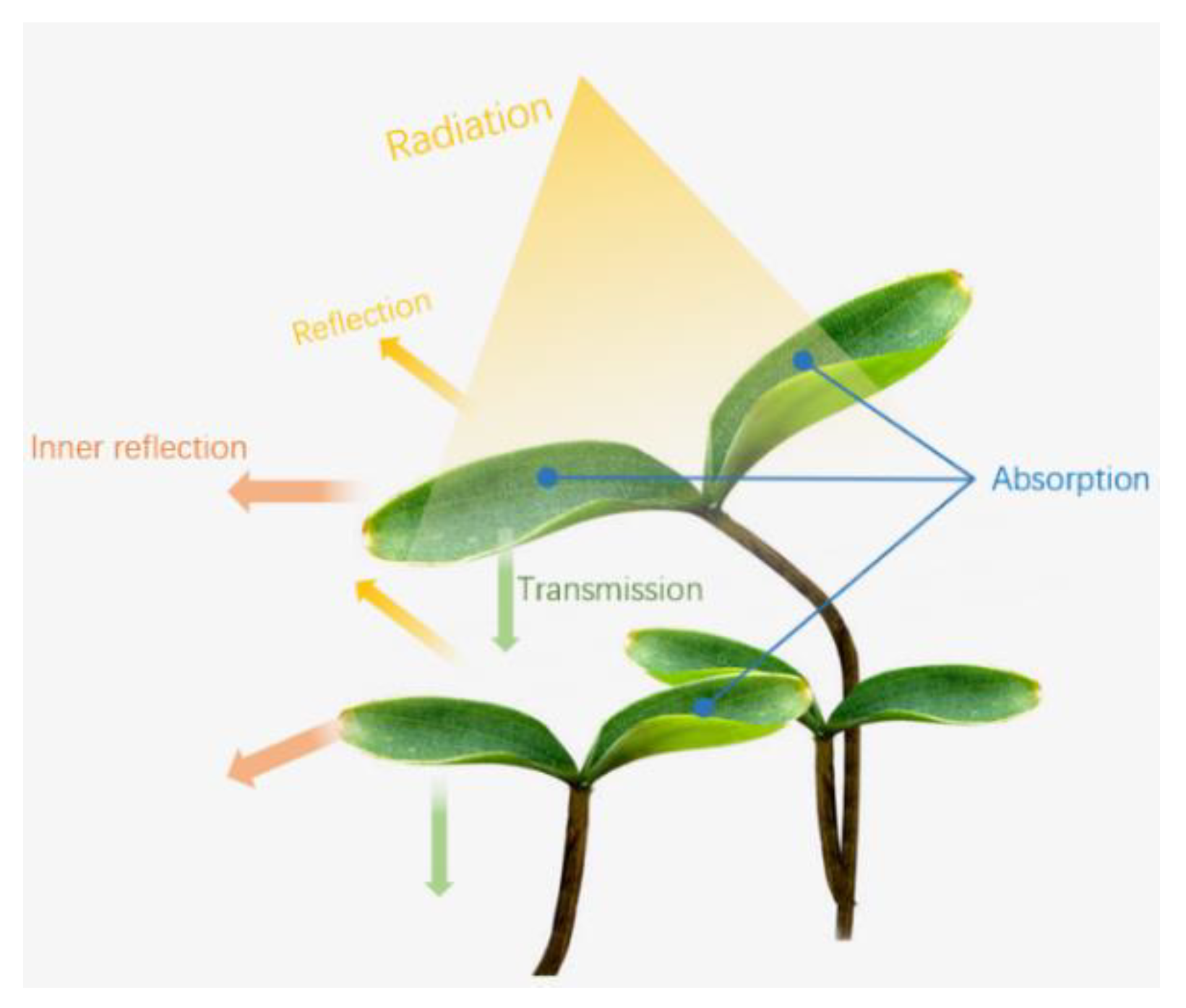
3.2. Photo Response of Pathogen-Infected Plants
4. Analysis Hyperspectral Imaging Information for Plant Disease Identification
4.1. Preprocessing of Hyperspectral Images
4.1.1. Image Mosaic
4.1.2. Image Segmentation
4.2. Vegetation Index
4.3. Machine Learning
4.4. Deep Learning
5. Summary
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ANN | Artificial Neural Network |
| ARI | Anthocyanin Reflectance Index |
| ASSDN | Architecture Self-Search Deep Network |
| BRT | Boosted Regression Tree |
| BPNN | Back Propagation Neural Network |
| CARI | Carotenoid Reflectance Index |
| CNN | Convolutional Neural Networks |
| CWA | Continuous Wavelet Analysis |
| DBN | Deep Belief Network |
| DCNN | Dynamic Convolution Neural Network) |
| ELM | Extreme Learning Machine |
| FDA | Fisher Discriminant Analysis |
| FLDA | Fisher’s Linear Discrimination Analysis |
| FR | Features Ranking |
| GA | Genetic Algorithm |
| GLCM | Gray-level Co-occurrence Matrix |
| HFPSSD | Hyperspectral Feature Profile Scanning-based Scab Detection |
| HLB | HuangLongBing |
| ISODATA | Interactive Self-Organizing Data Analysis |
| KNN | K-Nearest Neighbor |
| LDA | Linear Discriminant Analysis |
| LR | Logistic Regression |
| LS | Least Squares |
| MLE | Maximum Likelihood Estimate |
| MSC | Multiplicative Scatter Correction |
| NB | Naive Bayes |
| NVDI | Normalized Difference Vegetation Index |
| PCA | Principal Component Analysis |
| PRI | Photochemical Reflectance Index |
| PSRI | Plant Senescence Reflectance Index |
| RBF | Radial Basis Function |
| RGB | Red, Green, Blue |
| RVI | Ratio Vegetation Index |
| SAE | Stacked Auto-Encoder |
| SDA | Stepwise Discriminant Analysis |
| SFDI | Spectral Fractal Dimension Index |
| SIPI | Structure Insensitive Pigment Index |
| SPA | Successive Projections Algorithm |
| SPAD | Soil and Plant Analyzer Development |
| SRC | Sparse Representation-based Classifier |
| SVM | Support Vector Machine |
| UAV | Unmanned Aerial Vehicle |
| VIS | Vegetation Indices |
| WI | Water Index |
References
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Fisher, M.C.; Henk, D.A.; Briggs, C.J.; Brownstein, J.S.; Madoff, L.C.; McCraw, S.L.; Gurr, S.J. Emerging fungal threats to animal, plant and ecosystem health. Nature 2012, 484, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Boyd, L.A.; Ridout, C.; O’Sullivan, D.M.; Leach, J.E.; Leung, H. Plant–pathogen interactions: Disease resistance in modern agriculture. Trends. Genet. 2013, 29, 233–240. [Google Scholar] [CrossRef]
- Peralta, A.L.; Sun, Y.; McDaniel, M.D.; Lennon, J.T. Crop rotational diversity increases disease suppressive capacity of soil microbiomes. Ecosphere 2018, 9, e02235. [Google Scholar] [CrossRef]
- Ristaino, J.B.; Anderson, P.K.; Bebber, D.P.; Brauman, K.A.; Wei, Q. The persistent threat of emerging plant disease pandemics to global food security. Proc. Natl. Acad. Sci. USA 2021, 118, e2022239118. [Google Scholar] [CrossRef]
- Dan, E.; Ho, T.; Rwahnih, M.A.; Martin, R.R.; Tzanetakis, I. High throughput sequencing for plant virus detection and discovery. Phytopathology 2019, 109, 716–725. [Google Scholar]
- Ma, Z.; Luo, Y.; Michailides, T.J. Nested pcr assays for detection of monilinia fructicola in stone fruit orchards and botryosphaeria dothidea from pistachios in california. J. Phytopathol. 2010, 151, 312–322. [Google Scholar] [CrossRef]
- Singh, V.; Sharma, N.; Singh, S. A review of imaging techniques for plant disease detection. Artif. Intell. Agric. 2020, 4, 229–242. [Google Scholar] [CrossRef]
- Zhu, W.; Chen, H.; Ciechanowska, I.; Spaner, D. Application of infrared thermal imaging for the rapid diagnosis of crop disease. IFAC-PapersOnLine 2018, 51, 424–430. [Google Scholar] [CrossRef]
- Cen, H.; Weng, H.; Yao, J.; He, M.; Lv, J.; Hua, S.; Li, H.; He, Y. Chlorophyll fluorescence imaging uncovers photosynthetic fingerprint of citrus Huanglongbing. Front. Plant Sci. 2017, 8, 1509. [Google Scholar] [CrossRef] [Green Version]
- Mahlein, A.K.; Kuska, M.T.; Behmann, J.; Polder, G.; Walter, A. Hyperspectral sensors and imaging technologies in phytopathology: State of the art. Annu. Rev. Phytopathol. 2018, 56, 535–558. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Huang, Y.; Pu, R.; Gonzalez-Moreno, P.; Yuan, L.; Wu, K.; Huang, W. Monitoring plant diseases and pests through remote sensing technology: A review. Comput. Electron. Agric. 2019, 165, 104943. [Google Scholar] [CrossRef]
- Zaneti, R.N.; Girardi, V.; Spilki, F.R.; Mena, K.; Etchepare, R. Quantitative microbial risk assessment of Sars-CoV-2 for workers in wastewater treatment plants. Sci. Total Environ. 2020, 754, 142163. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, B.; Zhang, X.; Liu, P.; Huang, W. Impact of spectral interval on wavelet features for detecting wheat yellow rust with hyperspectral data. Int. J. Agr. Biolog. Eng. 2018, 11, 138–144. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Yang, Y.; Feng, X.; Xu, H.; He, Y. Identification of bacterial blight resistant rice seeds using terahertz imaging and hyperspectral imaging combined with convolutional neural network. Front. Plant Sci. 2020, 11, 821. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Wang, L.; Yang, F.; Yang, L. Spring corn leaf blight monitoring based on hyperspectral derivative index. Chin. Agric. Sci. Bull. 2019, 35, 143–150. [Google Scholar]
- Lu, J.; Zhou, M.; Gao, Y.; Jiang, H. Using hyperspectral imaging to discriminate yellow leaf curl disease in tomato leaves. Precis. Agric. 2018, 19, 379–394. [Google Scholar] [CrossRef]
- Nouri, M.; Gorretta, N.; Vaysse, P.; Giraud, M.; Germain, C.; Keresztes, B.; Roger, J.M. Near infrared hyperspectral dataset of healthy and infected apple tree leaves images for the early detection of apple scab disease. Data Brief 2018, 16, 967–971. [Google Scholar] [CrossRef]
- Sandasi, M.; Vermaak, I.; Chen, W.; Viljoen, A. Skullcap and germander: Preventing potential toxicity through the application of hyperspectral imaging and multivariate image analysis as a novel quality control method. Planta Med. 2014, 80, 1329–1339. [Google Scholar] [CrossRef] [Green Version]
- Zhu, H.; Chu, B.; Zhang, C.; Liu, F.; Jiang, L.; He, Y. Hyperspectral imaging for presymptomatic detection of tobacco disease with successive projections algorithm and machine-learning classifiers. Sci. Rep. 2017, 7, 4125. [Google Scholar] [CrossRef] [Green Version]
- Hovmller, M.S.; Walter, S.; Bayles, R.A.; Hubbard, A.; Flath, K.; Sommerfeldt, N.; Leconte, M.; Czembor, P.; Rodriguez-Algaba, J.; Thach, T. Replacement of the European wheat yellow rust population by new races from the centre of diversity in the near-himalayan region. Plant Pathol. 2016, 65, 402–411. [Google Scholar] [CrossRef] [Green Version]
- Huang, L.; Zhang, H.; Chao, R.; Huang, W.; Hu, T.; Zhao, J. Detection of scab in wheat ears using in situ hyperspectral data and support vector machine optimized by genetic algorithm. Int. J. Agr. Biol. Eng. 2020, 13, 7. [Google Scholar] [CrossRef]
- Shi, Y.; Huang, W.; Zhou, X. Evaluation of wavelet spectral features in pathological detection and discrimination of yellow rust and powdery mildew in winter wheat with hyperspectral reflectance data. J. Appl. Remote Sens. 2017, 11, 026025. [Google Scholar] [CrossRef]
- Mahlein, A.K.; Alisaac, E.; Masri, A.A.; Behmann, J.; Oerke, E.C. Comparison and combination of thermal, fluorescence, and hyperspectral imaging for monitoring fusarium head blight of wheat on spikelet scale. Sensors 2019, 19, 2281. [Google Scholar] [CrossRef] [Green Version]
- Leucker, M.; Wahabzada, M.; Kersting, K.; Peter, M.; Beyer, W.; Steiner, U.; Mahlein, A.K.; Oerke, E.C. Hyperspectral imaging reveals the effect of sugar beet quantitative trait loci on cercospora leaf spot resistance. Funct. Plant Bio. 2016, 44, 1. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Qi, L.; Xue, K.; Wang, W.; Zhu, X. Hyperspectral image analysis based on bosw model for rice panicle blast grading. Comput. Electron. Agric. 2015, 118, 167–178. [Google Scholar] [CrossRef]
- Shuaibu, M.; Lee, W.S.; Schueller, J.; Gader, P.; Hong, Y.; Kim, S. Unsupervised hyperspectral band selection for apple marssonina blotch detection. Comput. Electron. Agric. 2018, 12, 28. [Google Scholar] [CrossRef]
- Ruszczak, B.; Smykaa, K. The detection of alternaria solani infection on tomatoes using ensemble learning. J. Amb. Intel. Smart Environ. 2020, 12, 407–418. [Google Scholar] [CrossRef]
- Liang, K.; Huang, J.; He, R.; Wang, Q.; Chai, Y.; Shen, M. Comparison of Vis-NIR and SWIR hyperspectral imaging for the non-destructive detection of DON levels in Fusarium head blight wheat kernels and wheat flour. Infrared Phys. Techn. 2020, 106, 103281. [Google Scholar] [CrossRef]
- Sun, Y.; Wang, K.; Liu, Q.; Pan, L.; Tu, K. Classification and discrimination of different fungal diseases of three infection levels on peaches using hyperspectral reflectance imaging analysis. Sensors 2018, 18, 1295. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Liu, B.; Liu, L.; Zhu, Y.; Li, X. A review of deep learning used in the hyperspectral image analysis for agriculture. Artif. Intell. Rev. 2021, 54, 5205–5253. [Google Scholar] [CrossRef]
- Xing, H.; Feng, H.; Fu, J.; Xu, X.; Yang, G. Development and Application of Hyperspectral Remote Sensing; Springer: Berlin, Germany, 2017; Volume 546, pp. 271–282. [Google Scholar]
- Telmo, A.O.; Joná, H.; Luís, P.; José, B.; Emanuel, P.; Raul, M.; Joaquim, S. Hyperspectral imaging: A review on UAV-based sensors, data processing and applications for agriculture and forestry. Remote Sens. 2017, 9, 1110. [Google Scholar]
- Thomas, S.; Kuska, M.T.; Bohnenkamp, D.; Brugger, A.; Alisaac, E.; Wahabzada, M.; Behmann, J.; Mahlein, A.K. Benefits of hyperspectral imaging for plant disease detection and plant protection: A technical perspective. J. Plant. Dis. Protect. 2018, 125, 5–20. [Google Scholar] [CrossRef]
- Gates, D.M.; Keegan, H.J.; Schleter, J.C. Sensorik für einen präzisierten Pflanzenschutz. Gesunde Pflanz. 2008, 60, 131–141. [Google Scholar]
- Wang, L.; Jia, M.; Yin, D.; Tian, J. A review of remote sensing for mangrove forests: 1956–2018. Remote Sens. Environ. 2019, 231, 111223. [Google Scholar] [CrossRef]
- Hennessy, A.; Clarke, K.; Lewis, M. Hyperspectral classification of plants: A review of waveband selection generalisability. Remote Sens. 2020, 12, 113. [Google Scholar] [CrossRef] [Green Version]
- Olli, N.; Eija, H.; Sakari, T.; Niko, V.; Teemu, H.; Xiaowei, Y.; Juha, H.; Heikki, S.; Ilkka, P.L.N.; Nilton, I. Individual tree detection and classification with UAV-based photogrammetric point clouds and hyperspectral imaging. Remote Sens. 2017, 9, 185. [Google Scholar]
- René, H.; Norbert, J.; André, G.; Jens, O. The effect of epidermal structures on leaf spectral signatures of ice plants (Aizoaceae). Remote Sens. 2015, 7, 16901. [Google Scholar]
- Liu, Y.; Zhang, G.W.; Liu, D. Simultaneous measurement of chlorophyll and water content in navel orange leaves based on hyperspectral imaging. Spectroscopy 2014, 29, 40, 42–46. [Google Scholar]
- Mutanga, O.; Van Aardt, J.; Kumar, L. Imaging spectroscopy (hyperspectral remote sensing) in Southern Africa: An overview. S. Afr. J. Sci. 2010, 105, 193–198. [Google Scholar] [CrossRef]
- Wei, A.A.; Tian, L.; Chen, X.; Yu, Y. Retrieval and application of chlorophyll-a concentration in the Poyang Lake based on exhaustion method: A case study of Chinese Gaofen-5Satellitc AHSI data. J. Huazhong Normal Univ. 2020, 54, 447–453. [Google Scholar]
- Qu, J.; Sun, D.; Cheng, J.; Pu, H. Mapping moisture contents in grass carp (ctenopharyngodon idella) slices under different freeze drying periods by vis-nir hyperspectral imaging. LWT-Food Sci. Technol. 2017, 75, 529–536. [Google Scholar] [CrossRef]
- Oerke, E. Remote sensing of diseases. Annu. Rev. Phytopathol. 2020, 58, 225–252. [Google Scholar] [CrossRef] [PubMed]
- Bonants, P.; Schoen, C.; Wolf, J.; Zijlstra, C. Developments in detection of plant pathogens and other plant-related organisms: Detection in the past towards detection in the future. Mededelingen 2001, 66, 25–37. [Google Scholar] [PubMed]
- Pu, H.; Lin, L.; Sun, D. Principles of hyperspectral microscope imaging techniques and their applications in food quality and safety detection: A review. Compr. Rev. Food Sci. F. 2019, 18, 853–866. [Google Scholar] [CrossRef] [Green Version]
- Gary, A.; Roth, S.; Tahiliani, N.M.; Neu-Baker, S.A. Hyperspectral microscopy as an analytical tool for nanomaterials. Wires. Nanomed. Nanobi. 2015, 7, 565–579. [Google Scholar]
- Mahlein, A.K.; Rumpf, T.; Welke, P.; Dehne, H.W.; Plümer, L.; Steiner, U.; Oerke, E.C. Development of spectral indices for detecting and identifying plant diseases. Remote Sens. Environ. 2013, 128, 21–30. [Google Scholar] [CrossRef]
- Mahlein, A.K.; Steiner, U.; Hillnhütter, C.; Dehne, H.W.; Oerke, E.C. Hyperspectral imaging for small-scale analysis of symptoms caused by different sugar beet diseases. Plant Methods 2012, 8, 3. [Google Scholar] [CrossRef] [Green Version]
- Wahabzada, M.; Mahlein, A.K.; Bauckhage, C.; Steiner, U.; Oerke, E.C.; Kersting, K. Plant Phenotyping using Probabilistic Topic Models: Uncovering the Hyperspectral Language of Plants. Sci. Rep. 2016, 6, 22482. [Google Scholar] [CrossRef] [Green Version]
- Behmann, J.; Mahlein, A.K.; Rumpf, T.; Romer, C.; Plumer, L. A review of advanced machine learning methods for the detection of biotic stress in precision crop protection. Precis. Agric. 2015, 16, 239–260. [Google Scholar] [CrossRef]
- Lowe, A.; Harrison, N.; French, A.P. Hyperspectral image analysis techniques for the detection and classification of the early onset of plant disease and stress. Plant Methods 2017, 13, 80. [Google Scholar] [CrossRef] [PubMed]
- Mahlein, A.K. Plant Disease Detection by Imaging Sensors-Parallels and Specific Demands for Precision Agriculture and Plant Phenotyping. Plant Dis. 2016, 100, 241–251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, H.; Bian, L.; Cao, X.; Zhang, J. Hyperspectral imaging from a raw mosaic image with end-to-end learning. Opt. Express 2020, 28, 314–324. [Google Scholar] [CrossRef]
- Liu, S.; Yang, G.; Zhou, H.; Jing, H.; Feng, H.; Xu, B.; Yang, H. Extraction of maize seedling number information based on UAV imagery. Trans. CSAE 2018, 34, 9. [Google Scholar]
- Zhang, J.; Yang, C.; Zhao, B.; Song, H.; Hoffmann, W.C.; Shi, Y.; Zhang, D.; Zhang, G. Crop Classification and LAI Estimation Using Original and Resolution-Reduced Images from Two Consumer-Grade Cameras. Remote Sens. 2017, 9, 1054. [Google Scholar] [CrossRef] [Green Version]
- Dai, J.; Zhang, G.; Guo, P.; Zeng, Y.; Cui, M.; Xue, J. Classification method of main crops in northern Xinjiang based on UAV visible waveband images. Trans. CSAE 2018, 34, 8. [Google Scholar]
- He, Y.; Zhang, Y.; Li, J.; Wang, J. Estimation of stem biomass of individual Abies faxoniana through unmanned aerial vehicle remote sensing. J. Beijing For. Univ. 2016, 38, 8. [Google Scholar]
- Nalepa, J.; Myller, M.; Kawulok, M. Transfer learning for segmenting dimensionally-reduced hyperspectral images. IEEE Geosci. Remote Sens. 2019, 17, 1228–1232. [Google Scholar] [CrossRef] [Green Version]
- Cui, B.; Ma, X.; Xie, X.; Ren, G.; Ma, Y. Classification of visible and infrared hyperspectral images based on image segmentation and edge-preserving filtering. Infrared Phys. Techn. 2017, 81, 79–88. [Google Scholar] [CrossRef]
- Ishiyama, T.; Tanaka, S.; Uchida, K.; Fujikawa, S.; Yamashita, Y.; Kato, M. Relationship among vegetation variables and vegetation features of arid lands derived from satellite data. Adv. Space Res. 2001, 28, 183–188. [Google Scholar] [CrossRef]
- Vicente-Serrano, S.M. Evaluating the impact of drought using remote sensing in a mediterranean, semi-arid region. Nat. Hazards 2007, 40, 173–208. [Google Scholar] [CrossRef]
- Zhao, X.; Zhou, D.; Fang, J. Satellite-based studies on large-scale vegetation changes in China. J. Integr. Plant Bio. 2012, 54, 713–728. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J. Research on Hyperspectral Remote Sensing Images Classification Based on K-means Clustering. Geospat. Inform. 2016, 14, 4. [Google Scholar]
- Zhang, H.; Li, Y.; Jiang, H. Research status and Prospect of deep learning in hyperspectral image classification. Acta Autom. Sin. 2018, 44, 17. [Google Scholar]
- Szulczewski, W.; Zyromski, A.; Jakubowski, W.; Biniak-Pierog, M. A new method for the estimation of biomass yield of giant miscanthus (Miscanthus giganteus) in the course of vegetation. Renew. Sust. Energ. Rev. 2018, 82, 1787–1795. [Google Scholar] [CrossRef]
- Barriguinha, A.; Neto, M.D.; Gil, A. Vineyard yield estimation, prediction, and forecasting: A systematic literature review. Agronomy 2021, 11, 1789. [Google Scholar] [CrossRef]
- Zou, L.; Cao, S.; Sanchez-Azofeifa, A. Evaluating the utility of various drought indices to monitor meteorological drought in tropical dry forests. Int. J. Biometeorol. 2020, 64, 701–711. [Google Scholar] [CrossRef]
- Calzarano, F.; Pagnani, G.; Pisante, M.; Bellocci, M.; Cillo, G.; Metruccio, E.G.; Di Marco, S. Factors involved on tiger-stripe foliar symptom expression of esca of grapevine. Plants 2021, 10, 1041. [Google Scholar] [CrossRef]
- Rumpf, T.; Mahlein, A.K.; Steiner, U.; Oerke, E.C.; De Hne, H.W.; Plümer, L. Early detection and classification of plant diseases with support vector machines based on hyperspectral reflectance. Comput. Electron. Agr. 2010, 74, 91–99. [Google Scholar] [CrossRef]
- Cao, Y.F.; Xu, H.L.; Song, J.; Yang, Y.; Hu, X.H.; Wiyao, K.T.; Zhai, Z.Y. Applying spectral fractal dimension index to predict the spad value of rice leaves under bacterial blight disease stress. Plant Methods 2022, 18, 67. [Google Scholar] [CrossRef]
- Sims, D.A.; Gamon, J.A. Relationships between leaf pigment content and spectral reflectance across a wide range of species, leaf structures and developmental stages. Remote Sens. Environ. 2002, 81, 337–354. [Google Scholar] [CrossRef]
- Zhang, J.C.; Tian, Y.Y.; Yan, L.J.; Wang, B.; Wang, L.; Xu, J.F.; Wu, K.H. Diagnosing the symptoms of sheath blight disease on rice stalk with an in-situ hyperspectral imaging technique. Biosyst. Eng. 2021, 209, 94–105. [Google Scholar] [CrossRef]
- Yuan, L.; Yan, P.; Han, W.; Huang, Y.; Bao, Z. Detection of anthracnose in tea plants based on hyperspectral imaging. Comput. Electron. Agr. 2019, 167, 105039. [Google Scholar] [CrossRef]
- Calderon, R.; Navas-Cortes, J.A.; Zarco-Tejada, P.J. Early detection and quantification of verticillium wilt in olive using hyperspectral and thermal imagery over large areas. Remote Sens. 2015, 7, 5584–5610. [Google Scholar] [CrossRef] [Green Version]
- Huang, W.; Lu, J.; Ye, H.; Kong, W.; Yue, S. Quantitative identification of crop disease and nitrogen-water stress in winter wheat using continuous wavelet analysis. Int. J. Agr. Biol. Eng. 2018, 11, 8. [Google Scholar] [CrossRef] [Green Version]
- Karadag, K.; Tenekeci, M.E.; Tasaltin, R.; Bilgili, A. Detection of pepper fusarium disease using machine learning algorithms based on spectral reflectance. Sustain. Comput Inform. Syst. 2020, 28, 100299. [Google Scholar] [CrossRef]
- Xie, C.; Yang, C.; He, Y. Hyperspectral imaging for classification of healthy and gray mold diseased tomato leaves with different infection severities. Comput. Electron. Agr. 2017, 135, 154–162. [Google Scholar] [CrossRef]
- Lu, J.; Ehsani, R.; Shi, Y.; Abdulridha, J.; Castro, A.D.; Xu, Y. Field detection of anthracnose crown rot in strawberry using spectroscopy technology. Comput. Electron. Agric. 2017, 135, 289–299. [Google Scholar] [CrossRef]
- Weng, H.Y.; Lv, J.W.; Cen, H.Y.; He, M.B.; Zeng, Y.B.; Hua, S.J.; Li, H.Y.; Meng, Y.Q.; Fang, H.; He, Y. Hyperspectral reflectance imaging combined with carbohydrate metabolism analysis for diagnosis of citrus Huanglongbing in different seasons and cultivars. Sens. Actuators B-Chem. 2018, 275, 50–60. [Google Scholar] [CrossRef]
- Nagasubramanian, K.; Jones, S.; Sarkar, S.; Singh, A.K.; Singh, A.; Ganapathysubramanian, B. Hyperspectral band selection using genetic algorithm and support vector machines for early identification of charcoal rot disease in soybean stems. Plant Methods 2018, 14, 86. [Google Scholar] [CrossRef]
- Abdulridha, J.; Batuman, O.; Ampatzidis, Y. Uav-based remote sensing technique to detect citrus canker disease utilizing hyperspectral imaging and machine learning. Remote Sens. 2019, 11, 1373. [Google Scholar] [CrossRef] [Green Version]
- Deng, X.; Huang, Z.; Zheng, Z.; Lan, Y.; Dai, F. Field detection and classification of citrus huanglongbing based on hyperspectral reflectance. Comput. Electron. Agr. 2019, 167, 105006. [Google Scholar] [CrossRef]
- Lin, F.; Guo, S.; Tan, C.; Zhou, X.; Zhang, D. Identification of rice sheath blight through spectral responses using hyperspectral images. Sensors 2020, 20, 6243. [Google Scholar] [CrossRef] [PubMed]
- Vijver, R.; Mertens, K.; Heungens, K.; Somers, B.; Saeys, W. In-field detection of Alternaria solani in potato crops using hyperspectral imaging. Comput. Electron. Agr. 2019, 168, 105106. [Google Scholar] [CrossRef]
- Zhang, D.; Chen, G.; Zhang, H.; Jin, N.; Chen, Y. Integration of spectroscopy and image for identifying fusarium damage in wheat kernels using hyperspectral imaging. Spectrochim. Acta A 2020, 236, 118344. [Google Scholar] [CrossRef] [PubMed]
- Gu, Q.; Sheng, L.; Zhang, T.H.; Lu, Y.W.; Zhang, Z.J.; Zheng, K.F.; Hu, H.; Zhou, H.K. Early detection of tomato spotted wilt virus infection in tobacco using the hyperspectral imaging technique and machine learning algorithms. Comput. Electron. Agr. 2019, 167, 105066. [Google Scholar] [CrossRef]
- Calamita, F.; Imran, H.A.; Vescovo, L.; Mekhalfi, M.L.; Porta, N.L. Early identification of root rot disease by using hyperspectral reflectance: The case of pathosystem grapevine/armillaria. Remote Sens. 2021, 13, 2436. [Google Scholar] [CrossRef]
- Zhao, X.H.; Zhang, J.C.; Huang, Y.B.; Tian, Y.Y.; Yuan, L. Detection and discrimination of disease and insect stress of tea plants using hyperspectral imaging combined with wavelet analysis. Comput. Electron. Agr. 2022, 193, 106717. [Google Scholar] [CrossRef]
- Du, P.; Xia, J.; Xue, Z.; Tan, K.; Su, H.; Bao, R. Research progress of hyperspectral remote sensing image classification. Remote Sens. Bull. 2016, 20, 21. [Google Scholar]
- Hou, P.; Yao, M.; Jia, W.; Zhang, F.; Wang, D. Spatial spectrum discriminant analysis for hyperspectral image classification. Opt. Precis. Eng. 2018, 26, 450–460. [Google Scholar]
- Ghasimi, P.; Benediktsson, J.A.; Ulfarsson, M.O. The spectral-spatial classification of hyperspectral images based on Hidden Markov Random Field. IEEE T. Geosci. Remote Sens. 2014, 52, 2565–2574. [Google Scholar]
- Hinton, G.E.; Salakhutdinov, R.R. Reducing the Dimensionality of Data with Neural Networks. Science 2006, 313, 504–507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hinton, G.E.; Osindero, S.; Teh, Y.W. A Fast Learning Algorithm for Deep Belief Nets. Neural Comput. 2014, 18, 1527–1554. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2012, 25, 1097–1105. [Google Scholar] [CrossRef]
- Wei, F.; Li, S.; Fang, L. Spectral-spatial hyperspectral image classification via superpixel merging and sparse representation. IEEE Geosci. Remote. Sens. 2015, 18, 861–865. [Google Scholar]
- Zhao, X.; Chen, Y.; Jia, X. Spectral-Spatial Classification of Hyperspectral Data Based on Deep Belief Network. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 2381–2392. [Google Scholar]
- Deng, X.L.; Zhu, Z.H.; Yang, J.C.; Zheng, Z.; Huang, Z.X.; Yin, X.B.; Wei, S.J.; Lan, Y.B. Detection of citrus huanglongbing based on multi-input neural network model of uav hyperspectral remote sensing. Remote Sens. 2020, 12, 2678. [Google Scholar] [CrossRef]
- Zhong, P.; Gong, Z.; Li, S.; Schonlieb, C.B. Learning to Diversify Deep Belief Networks for Hyperspectral Image Classification. IEEE T. Geosci. Remote 2017, 55, 3516–3530. [Google Scholar] [CrossRef]
- Fazari, A.; Pellicer-Valero, O.J.; Gomez-SancHs, J.; Bernardi, B.; Blasco, J. Application of deep convolutional neural networks for the detection of anthracnose in olives using vis/nir hyperspectral images. Comput. Electron. Agr. 2021, 187, 106252. [Google Scholar] [CrossRef]
- Polder, G.; Blok, P.M.; Villiers, H.; Wolf, J.; Kamp, J. Potato virus detection in seed potatoes using deep learning on hyperspectral images. Front. Plant Sci. 2019, 10, 209. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Han, L.; Dong, Y.; Shi, Y.; Sobeih, T. A deep learning-based approach for automated yellow rust disease detection from high-resolution hyperspectral uav images. Remote Sens. 2019, 11, 1554. [Google Scholar] [CrossRef] [Green Version]
- Xiu, J.; Lu, J.; Shuai, W.; Hai, Q.; Shao, L. Classifying wheat hyperspectral pixels of healthy heads and fusarium head blight disease using a deep neural network in the wild field. Remote Sens. 2018, 10, 395. [Google Scholar]
- Feng, L.; Wu, B.; Zhu, S.; Wang, J.; Zhang, C. Investigation on data fusion of multisource spectral data for rice leaf diseases identification using machine learning methods. Front. Plant Sci. 2020, 11, 577063. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, I.; Gutierrez, S.; Ceballos, S.; Iniguez, R.; Barrio, I.; Tardaguila, J. Artificial intelligence and novel sensing technologies for assessing downy mildew in grapevine. Horticulturae 2021, 7, 103. [Google Scholar] [CrossRef]
- Nguyen, C.; Sagan, V.; Maimaitiyiming, M.; Maimaitijiang, M.; Bhadra, S.; Kwasniewski, M.T. Early detection of plant viral disease using hyperspectral imaging and deep learning. Sensors 2021, 21, 742. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.P.; Lv, W.B.; Han, K.X.; Tao, W.T.; Zheng, L.; Weng, S.Z.; Huang, L.S. Determination of wheat kernels damaged by fusarium head blight using monochromatic images of effective wavelengths from hyperspectral imaging coupled with an architecture self-search deep network. Food Control 2022, 135, 108819. [Google Scholar]

| Vegetation Index | Formula | Application |
|---|---|---|
| Normalized difference vegetation index (NDVI) | (R800 − R670)/(R800 + R670) | Detecting vegetation coverage, related to chlorophyll content. |
| Green normalized difference vegetation index (GNDVI) | (NIR − GREEN)/(NIR + GREEN) | Detecting withered or aged crops and measuring the nitrogen content in leaves in the absence of red bands. |
| Water index (WI) | R900/R970 | Estimating the water stress. |
| Ratio vegetation index (RVI) | R800/R670 | Detecting and estimating plant biomass, related to chlorophyll content. |
| Plant senescence reflectance index (PSRI) | (R680 − R500)/R750 | Detecting plant senescence, related to pigment content. |
| Carotenoid reflectance index (CARI) | 1/R510 − 1/R550 | Related to carotenoid content. |
| Anthocyanin reflectance index (ARI) | 1/R510 − 1/R700 | Detecting yellow rust of wheat, related to anthocyanin content. |
| Photochemical reflectance index (PRI) | (R570 − R531)/(R570 + R531) | Detecting yellow rust of wheat, related to photosynthesis and carotenoid content. |
| Structure insensitive pigment index (SIPI) | (R800 − R445)/R800 + R680) | Related to the ratio of carotinoids and chlorophylla. |
| Method | Advantage | Disadvantage |
|---|---|---|
| K-NN | (1) The theory has matured and easy application. (2) Less training time. | (1) High computational complexity and spatial complexity. (2) Sample imbalance. |
| NBM | (1) Performs well for small-scale data and is suitable for multi-classification. (2) Simple computation. | (1) Due to the emphasis on conditional independence, the accuracy is reduced. (2) Sensitive to the expression of input data. |
| A priori probability needs to be calculated. | ||
| MLE | It can quickly specify the pixels to be classified into one of several classes. | When there are many types of hyperspectral data, the operation speed slows down significantly, and more training samples are needed. |
| Decision Tree | (1) Low computing cost. (2) Able to handle irrelevant features. (3) Suitable for datasets with many attributes and has high scalability. | (1) Easy to cause over-fitting. (2) When all kinds of samples are unbalanced, the result of information gain will be biased to the characteristics with more data values. |
| ELM | Has fast learning speed and good generalization ability. | The parameters of the model are randomly selected, which causes ELM instability. |
| SRC | Low computing cost. | Various sub-dictionaries have high atomic correlation, and similar samples may be linearly represented by different sub-dictionaries, resulting in poor classification accuracy. |
| SVM | (1) Low generalization error rate, low computing cost. (2) It can solve the classification problem of small-sample and high-dimensional data. | It is sensitive to parameter adjustment and function selection. |
| Has good classification effect. |
| Reference | Pathosystem | Scale | Methods | Detection | Early Detection | Precision |
|---|---|---|---|---|---|---|
| Calderon et al. (2015) [75] | Olive—Verticillium wilt | Canopy | LDA, SVM | ✓ | Overall: LDA 59.0%, SVM 79.2% Early detection: LDA 75.0%, SVM 40.6%. | |
| Xie et al. (2017) [78] | Tomato—Gray mold disease | Leaf | KNN, C5.0, FR-KNN | ✓ | Detection: 94.44%, 94.44%, 97.22% (KNN, C5.0, KNN) Early detection: 66.67%, 66.67%, 41.67% | |
| Lu et al. (2017) [79] | Strawberry—Anthracnose crown rot | Leaf | SDA, FDA, KNN | ✓ | ✓ | SDA: 71.3%, FDA: 70.05%, KNN: 73.6% |
| Zhu et al. (2017) [20] | Tobacco—Mosaic virus | Leaf | SPA–GLCM–BPNN/ELM/ SVM | ✓ | 95.00% | |
| Huang et al. (2018) [76] | Wheat—Powdery mildew, stripe rust | Canopy | FLDA, SVM | ✓ | FLDA: Powdery mildew 78.1%, stripe rust 95.6% | |
| Weng et al. (2018) [80] | Citrus—Huanglongbing, Fe deficient | Leaf | LS–SVM | 93.50% | ||
| Nagasubramanian et al. (2018) [81] | Sugar beet—Cercospora leaf spot | stem | GA–SVM | ✓ | Detection: 97% Early detection: 90.91% | |
| Abdulridha et al. (2019) [82] | Citrus—Canker | Leaf/canopy | RBF, KNN | ✓ | Leaf: 94%, 95%, 96% (asymptomatic, early, and late symptoms) Canopy: 94%, 96%, 100% | |
| Mahlein et al. (2019) [24] | Wheat—Fusarium head blight | Spikelet | SVM | ✓ | Detection: 89.0% Early detection: 78.0% | |
| Deng et al. (2019) [83] | Citrus—Huanglongbing | Leaf | SVM | ✓ | ✓ | Detection: 96% Early Detection: 90.8% |
| Lin et al. (2020) [84] | Rice—Sheath blight | Leaf | Relief decision tree | 95.50% | ||
| Van de Vijver et al. (2020) [85] | Potato—Early blight | Leaf | PLS–DA, PCA–SVM, decision tree | ✓ | 92.78% | |
| Karadag et al. (2020) [77] | Pepper—Fusarium disease, mycorrhizal fungus | Leaf | KNN, ANN, NB | KNN: 100%, ANN: 88.125%, NB: 82% | ||
| Zhang et al. (2020) [86] | Wheat—Fusarium head blight | Leaf | SPA-RF | 96.44% | ||
| Gu et al. (2020) [87] | Tobacco—Tomato spotted wilt virus | Leaf | SPA–BRT | ✓ | 85.20% | |
| Yuan et al. (2020) [74] | Tea—Anthracnose | Leaf | ISODATA | Pixel level: 98% Patch level: 94% | ||
| Calamita et al. (2021) [88] | Vitis vinifera—Fungi infection | Leaf | NB | ✓ | Detection: 90% Early detection: 75% | |
| Zhang et al. (2021) [73] | Rice—Sheath blight | Leaf | K-means–FLDA–HFPSSD | Pixel level: 98.42% Patch level: 95.92% | ||
| Zhao et al. (2022) [89] | Tea—Green leafhopper, anthracnose, sunburn | Leaf | CWA–K-means, SVM–RF | Green leafhopper: 93.99–94.20%, Anthracnose 94.12–94.28%, Sunburn 82.50–83.91% |
| Method | Advantages | Disadvantages |
|---|---|---|
| ANN | Able to learn advanced features of data automatically and has excellent classification performance. | (1) The model requires huge parameters and high computational force. (2) The input is a one-dimensional vector, which cannot make good use of map spatial information. (3) Difficult training, easy to produce gradient disappearance. |
| CNN | (1) Able to learn advanced features of data automatically and has excellent classification performance. (2) Compared to the ANN, the CNN model requires significantly fewer data parameters and trains faster. | The existence of a pooling layer may lead to the loss of valuable information while ignoring the relationship between the global and local scales. |
| Reference | Host–Pathogen System | Scale | Methods | Field Capable | Early Detection | Precision |
|---|---|---|---|---|---|---|
| Sun et al. (2018) [30] | Peach—Botrytis cinerea, Rhizopus stolonifera, Colletotrichum acutatum | Flesh | PCA–DBN | ✓ | Detection: 92.5% Early Detection: 82.5% | |
| Jin et al. (2018) [103] | Wheat—Fusarium head blight | Kernels | CNN | ✓ | 74.30% | |
| Polder et al. (2019) [101] | Potato—Viruses | Leaf | CNN | ✓ | ✓ | 78.00% |
| Zhang et al. (2019) [102] | Wheat—Yellow rust | canopy | DCNN | ✓ | 85% | |
| Deng et al. (2020) [98] | Citrus—Huanglongbing | canopy | SAE | ✓ | 99.72% | |
| Liang et al. (2020) [29] | Wheat—Fusarium head blight | Kernels/Flour | MSC–GA–SAE | Kernels: 100% Flour: 96% | ||
| Zhang et al. (2020) [15] | Rice—Bacterial blight | Leaf | CNN | 82.60% | ||
| Feng et al. (2020) [104] | Rice—Leaf blight, rice blast, rice sheath blight | Leaf | SVM, LR, CNN | 93.00% | ||
| Fazari et al. (2021) [100] | Olive—Anthracnose | Leaf | CNN | ✓ | Detection: 100% Early Detection: 85% | |
| Hernandez et al. (2021) [105] | Grape—Downy mildew | Leaf | KNN, CNN | ✓ | CNN: 82% KNN: 66% | |
| Nguyen et al. (2021) [106] | Grape—Grapevine vein-clearing virus | vine | 2D CNN, 3D CNN | ✓ | 2D CNN:71% 3D CNN: 75% | |
| Liu et al. (2022) [107] | Wheat—Fusarium head blight | kernels | ASSDN | 98.31% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wan, L.; Li, H.; Li, C.; Wang, A.; Yang, Y.; Wang, P. Hyperspectral Sensing of Plant Diseases: Principle and Methods. Agronomy 2022, 12, 1451. https://doi.org/10.3390/agronomy12061451
Wan L, Li H, Li C, Wang A, Yang Y, Wang P. Hyperspectral Sensing of Plant Diseases: Principle and Methods. Agronomy. 2022; 12(6):1451. https://doi.org/10.3390/agronomy12061451
Chicago/Turabian StyleWan, Long, Hui Li, Chengsong Li, Aichen Wang, Yuheng Yang, and Pei Wang. 2022. "Hyperspectral Sensing of Plant Diseases: Principle and Methods" Agronomy 12, no. 6: 1451. https://doi.org/10.3390/agronomy12061451
APA StyleWan, L., Li, H., Li, C., Wang, A., Yang, Y., & Wang, P. (2022). Hyperspectral Sensing of Plant Diseases: Principle and Methods. Agronomy, 12(6), 1451. https://doi.org/10.3390/agronomy12061451








