Overexpression of Maize Glutathione S-Transferase ZmGST26 Decreases Drought Resistance of Arabidopsis
Abstract
1. Introduction
2. Results
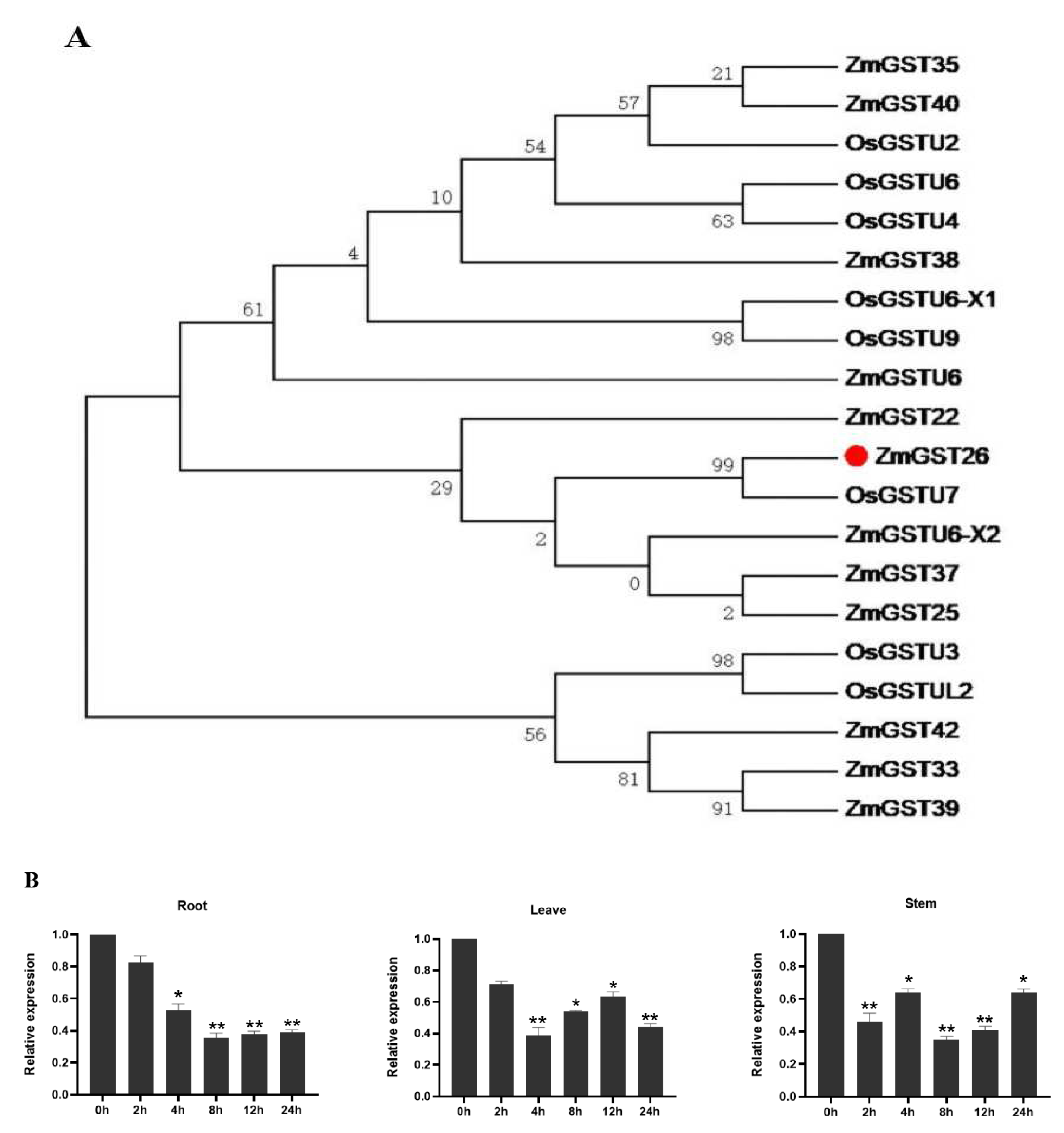
2.1. Drought Stress Induced the Downregulated Expression of ZmGST26 Gene
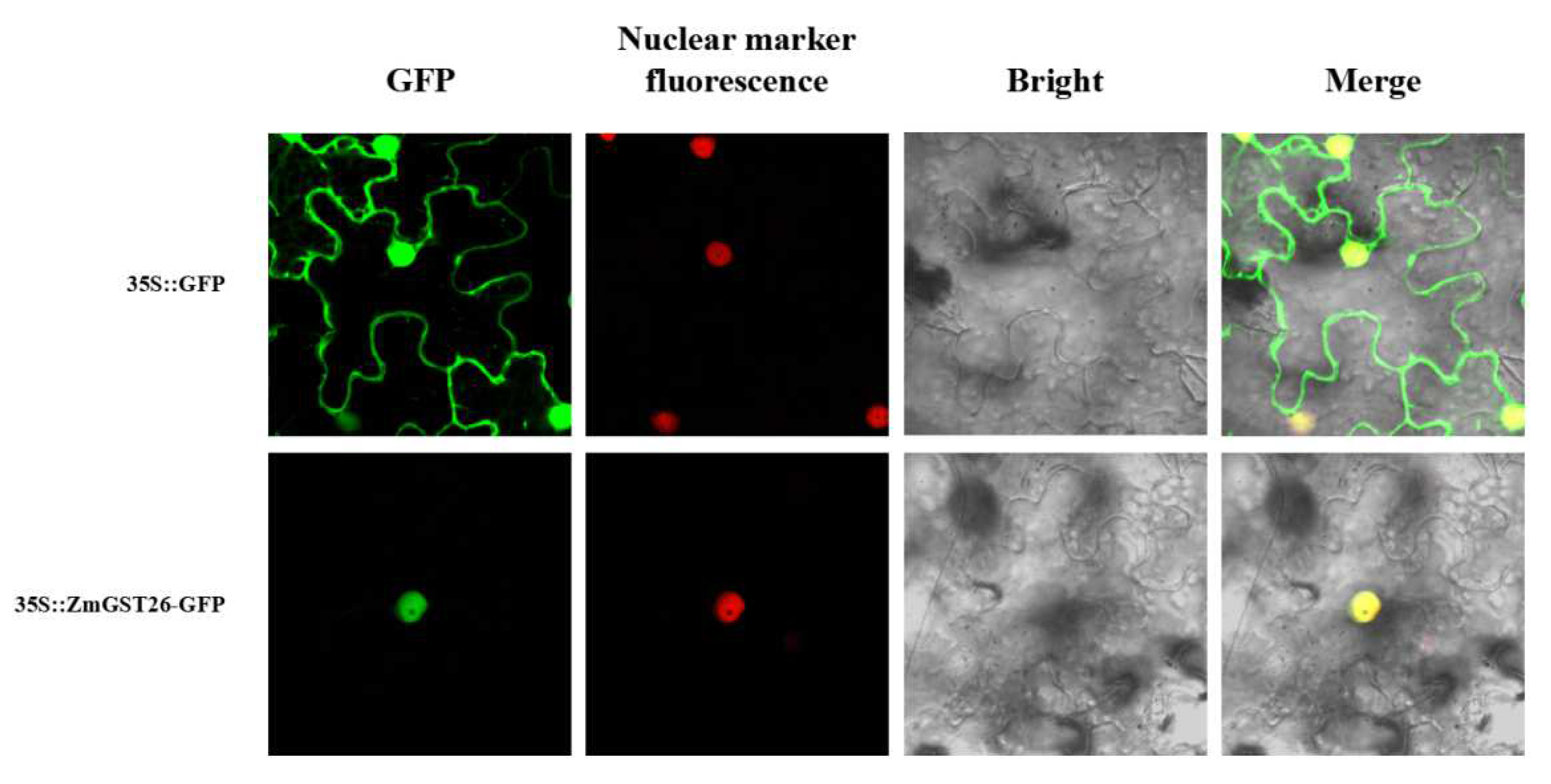
2.2. Subcellular Localization
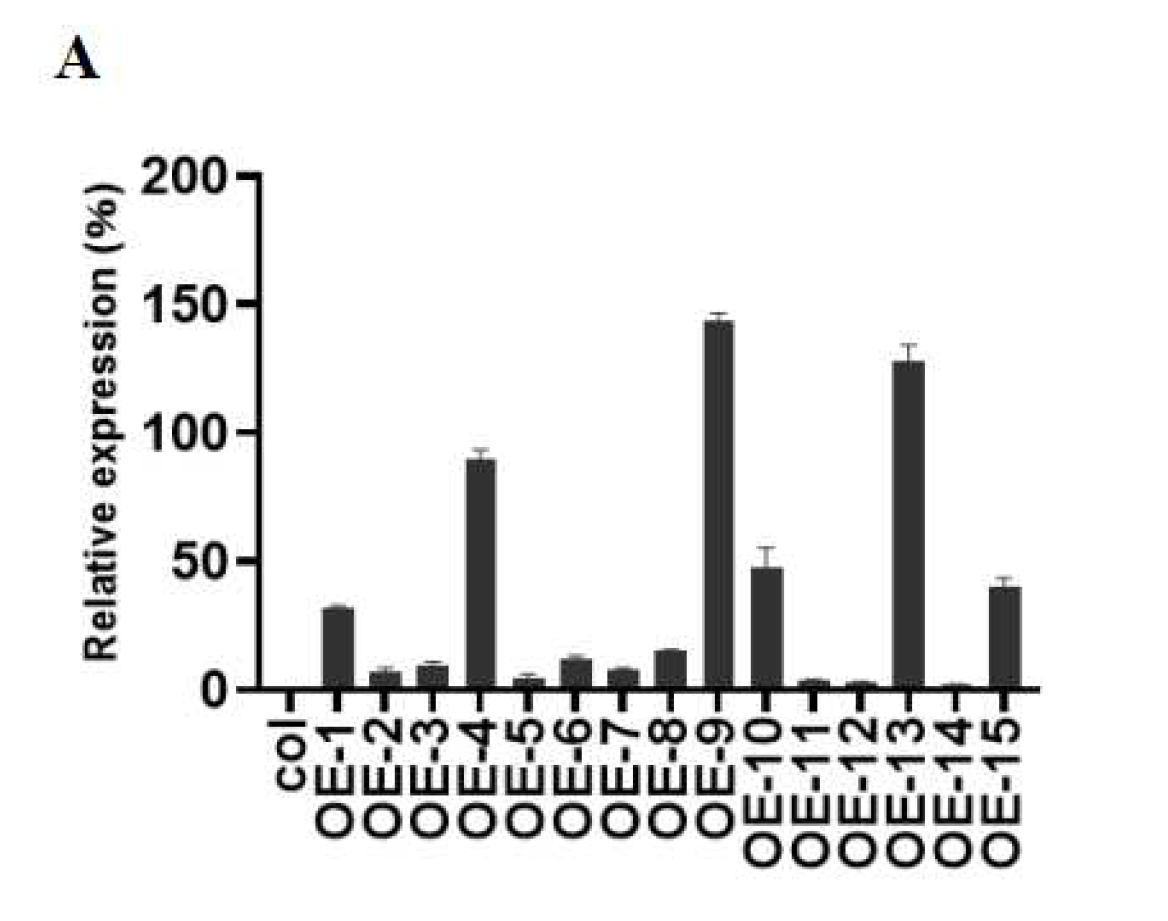
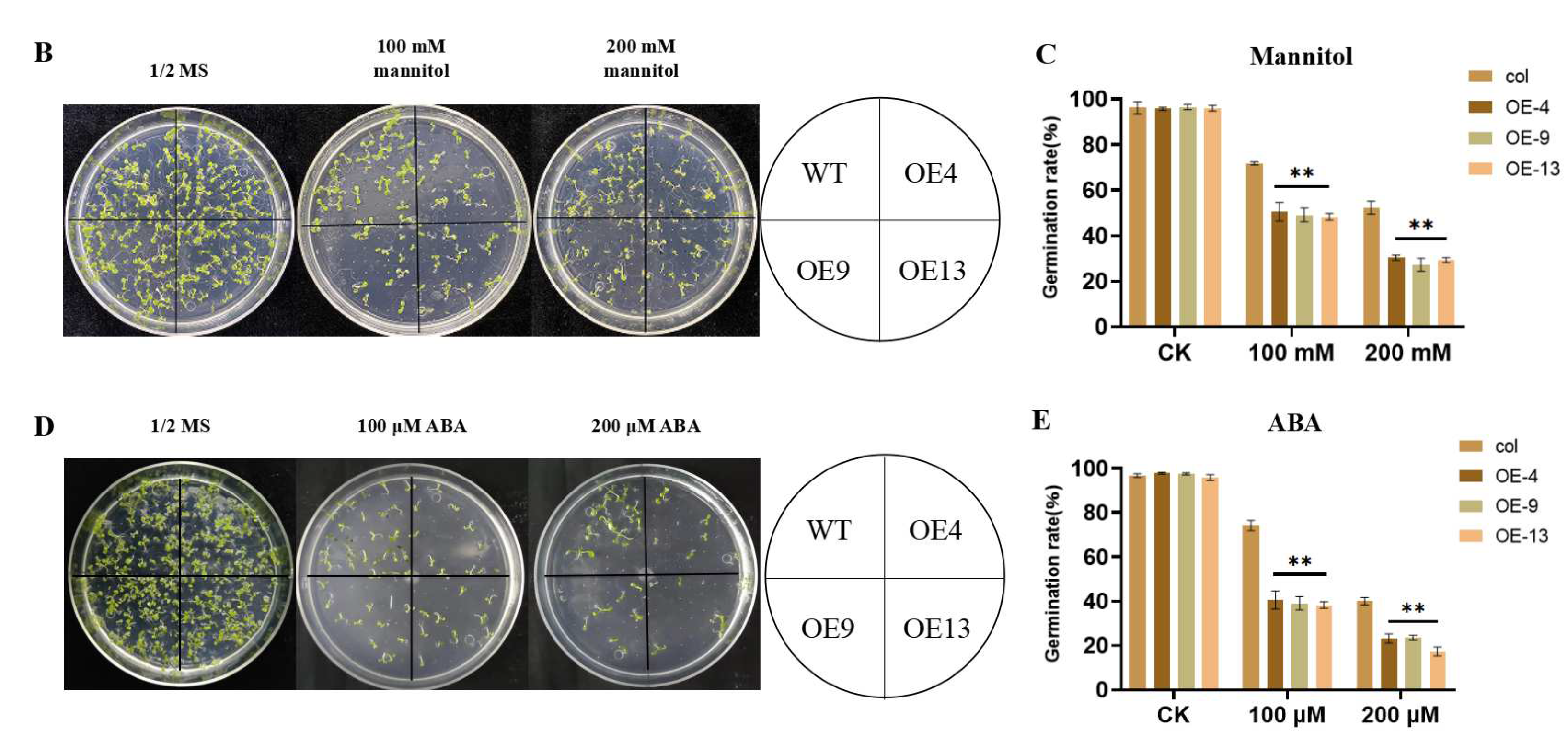
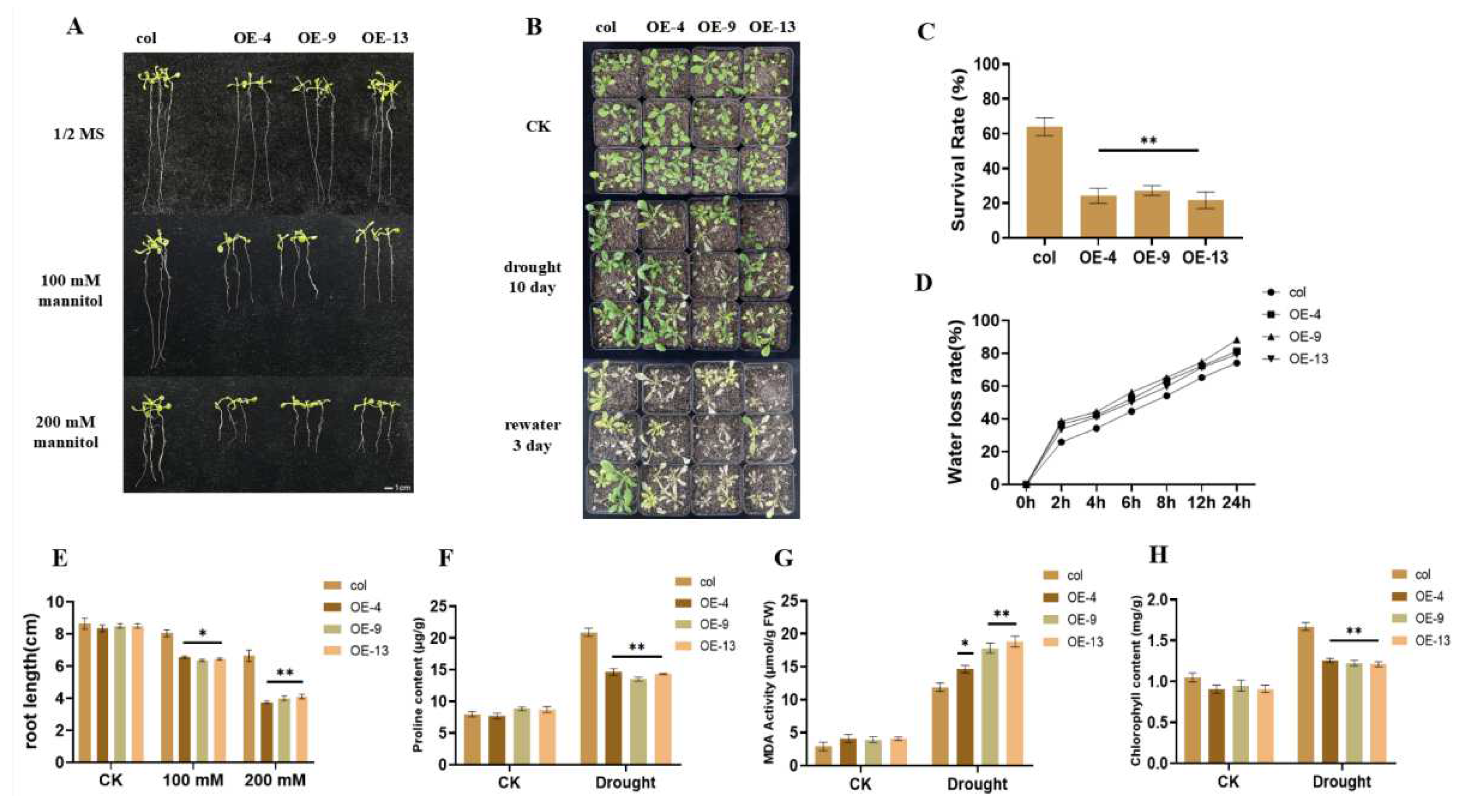
2.3. Overexpression of ZmGST26 Reduced the Drought Tolerance of Transgenic Arabidopsis
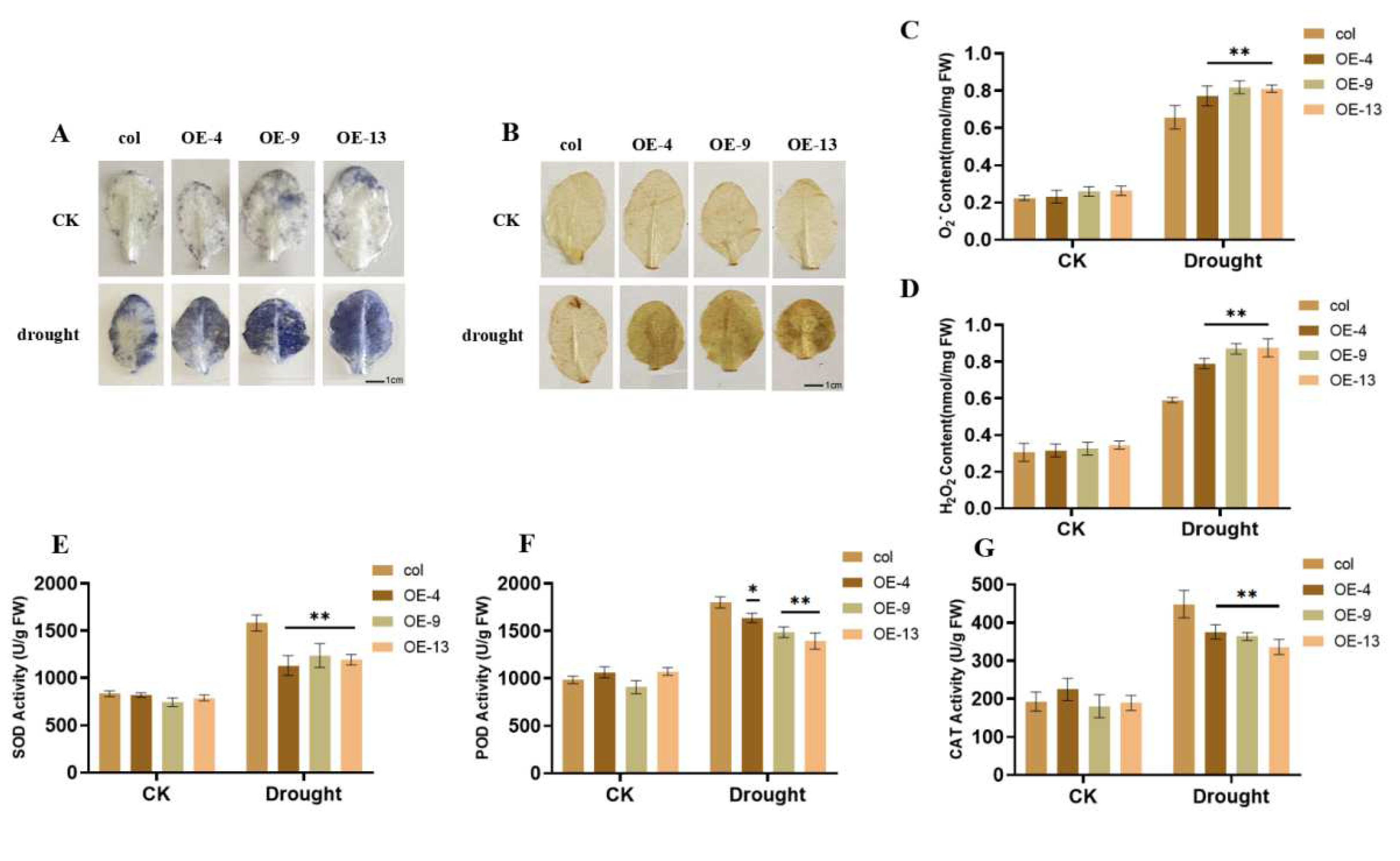
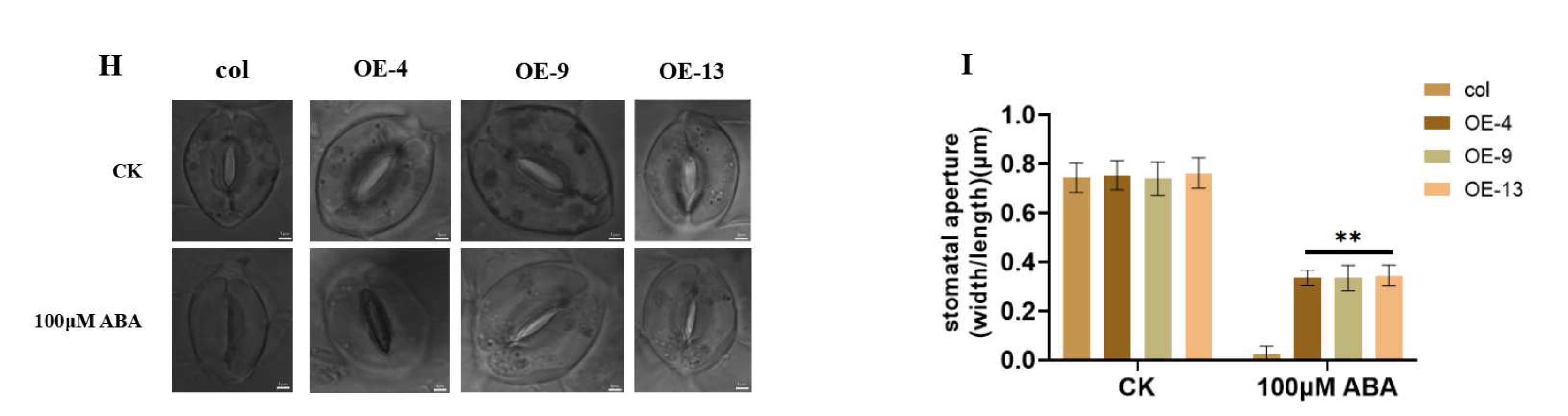
2.4. Overexpression of ZmGST26 Increased the Sensitivity of Transgenic Arabidopsis to Drought
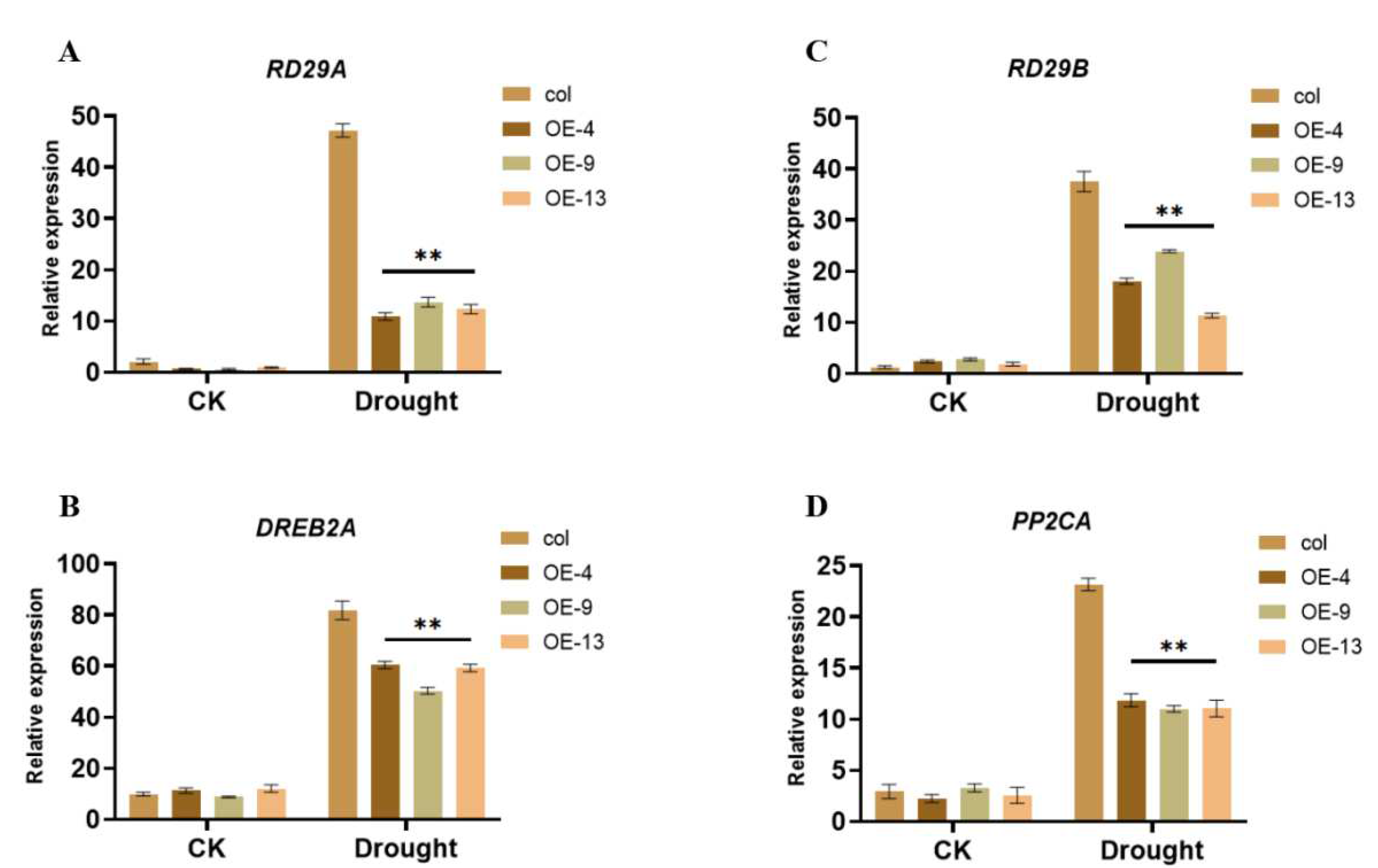
2.5. ZmGST26 Negatively Regulates the Expression of Drought-Related Genes in Overexpressed Arabidopsis
3. Materials and Methods
3.1. Plant Materials and Growth Conditions
3.2. Sequence Analysis
3.3. RNA Extraction and qRT-PCR Analysis
3.4. Subcellular Localization
3.5. Acquisition and Phenotypic Analysis of Overexpressed Arabidopsis
3.6. Determination of O2−, H2O2 and Physiological and Biochemical Indexes in Overexpressed Arabidopsis
3.7. Guard Cell Stomatal Aperture Observations
3.8. Statistical Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Manna, M.; Thakur, T.; Chirom, O.; Mandlik, R.; Deshmukh, R.; Salvi, P. Transcription factors as key molecular target to strengthen the drought stress tolerance in plants. Physiol. Plant 2020, 172, 847–868. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Zhang, W.; Zheng, D.; Xiong, H.; Feng, X.; Zhang, X.; Wang, Q.; Wu, F.; Xu, J.; Lu, Y. ZmLBD5 Increases Drought Sensitivity by Suppressing ROS Accumulation in Arabidopsis. Plants 2022, 11, 1382. [Google Scholar] [CrossRef]
- Xing, X.; Fang, C.; Li, L.; Jiang, H.; Zhou, Q.; Jiang, H.; Wang, S. Improved drought tolerance by α-naphthaleneacetic acid-induced ROS accumulation in two soybean cultivars. J. Integr. Agric. 2016, 15, 1770–1784. [Google Scholar] [CrossRef]
- Shi, Y.; Chang, Y.L.; Wu, H.T.; Shalmani, A.; Liu, W.T.; Li, W.Q.; Xu, J.W.; Chen, K.M. OsRbohB-mediated ROS production plays a crucial role in drought stress tolerance of rice. Plant Cell Rep. 2020, 39, 1767–1784. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, F.; Hong, Y.; Huang, J.; Shi, H.; Zhu, J.K. Two Chloroplast Proteins Suppress Drought Resistance by Affecting ROS Production in Guard Cells. Plant Physiol. 2016, 172, 2491–2503. [Google Scholar] [CrossRef]
- Pan, R.; Buitrago, S.; Feng, Z.; Abou-Elwafa, S.F.; Xu, L.; Li, C.; Zhang, W. HvbZIP21, a Novel Transcription Factor From Wild Barley Confers Drought Tolerance by Modulating ROS Scavenging. Front. Plant Sci. 2022, 13, 878459. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Shen, C.; Zhang, Y.; Zhou, Y.; Niu, M.; Wang, H.L.; Lian, C.; Tian, Q.; Mao, W.; Wang, X.; et al. PePYL4 enhances drought tolerance by modulating water use efficiency and ROS scavenging in Populus. Tree Physiol. 2022. Advance online publication. [Google Scholar] [CrossRef]
- Yang, L.; Lang, C.; Wu, Y.; Meng, D.; Yang, T.; Li, D.; Jin, T.; Zhou, X. ROS1-mediated decrease in DNA methylation and increase in expression of defense genes and stress response genes in Arabidopsis thaliana due to abiotic stresses. BMC Plant Biol. 2022, 22, 104. [Google Scholar] [CrossRef]
- Niu, M.; Bao, C.; Chen, J.; Zhou, W.; Zhang, Y.; Zhang, X.; Su, N.; Cui, J. RsGSTF12 Contributes to Anthocyanin Sequestration in Radish (Raphanus sativus L.). Front. Plant Sci. 2022, 13, 870202. [Google Scholar] [CrossRef]
- Naboulsi, I.; Ben Mrid, R.; Ennoury, A.; Zouaoui, Z.; Nhiri, M.; Nhiri, M.; Bakrim, B.; Yasri, A.; Aboulmouhajir, A. Crataegus oxyacantha Extract as a Biostimulant to Enhance Tolerance to Salinity in Tomato Plants. Plants 2022, 11, 1283. [Google Scholar] [CrossRef]
- Sabir, I.A.; Manzoor, M.A.; Shah, I.H.; Liu, X.; Jiu, S.; Wang, J.; Alam, P.; Abdullah, M.; Zhang, C. Identification and Comprehensive Genome-Wide Analysis of Glutathione S-Transferase Gene Family in Sweet Cherry (Prunus avium) and Their Expression Profiling Reveals a Likely Role in Anthocyanin Accumulation. Front. Plant Sci. 2022, 13, 938800. [Google Scholar] [CrossRef] [PubMed]
- Yue, Z.; Chen, Y.; Wang, Y.; Zheng, L.; Zhang, Q.; Liu, Y.; Hu, C.; Chen, C.; Ma, K.; Sun, Z. Halotolerant Bacillus altitudinis WR10 improves salt tolerance in wheat via a multi-level mechanism. Front. Plant Sci. 2022, 13, 941388. [Google Scholar] [CrossRef] [PubMed]
- Kortheerakul, C.; Kageyama, H.; Waditee-Sirisattha, R. Molecular and functional insights into glutathione S-transferase genes associated with salt stress in Halothece sp. PCC7418. Plant Cell Environ. 2021, 44, 3583–3596. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.H.; Jiang, H.W.; Hsieh, E.J.; Chen, H.Y.; Chien, C.T.; Hsieh, H.L.; Lin, T.P. Drought and salt stress tolerance of an Arabidopsis glutathione S-transferase U17 knockout mutant are attributed to the combined effect of glutathione and abscisic acid. Plant Physiol. 2012, 158, 340–351. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhang, N.; Liu, Z.; Liu, S.; Liu, C.; Lin, J.; Yang, H.; Li, S.; Yukawa, Y. The AtGSTU7 gene influences glutathione-dependent seed germination under ABA and osmotic stress in Arabidopsis. Biochem. Biophys. Res. Commun. 2020, 528, 538–544. [Google Scholar] [CrossRef]
- Li, X.J.; Wang, J.; Li, F.; Pia, B.; Zhao, T.; Yuan, B. Transcriptomic analysis of Glycine soja and G. max seedlings and functional characterization of GsGSTU24 and GsGSTU42 genes under submergence stress. Environ. Exp. Bot. 2020, 171, 103963–103974. [Google Scholar] [CrossRef]
- Ding, F.; Wang, C.; Zhang, S.; Wang, M. A jasmonate-responsive glutathione S-transferase gene SlGSTU24 mitigates cold-induced oxidative stress in tomato plants. Sci. Hortic. 2022, 303, 111231. [Google Scholar] [CrossRef]
- Srivastava, D.; Verma, G.; Chauhan, A.S.; Pande, V.; Chakrabarty, D. Rice (Oryza sativa L.) tau class glutathione S-transferase (OsGSTU30) overexpression in Arabidopsis thaliana modulates a regulatory network leading to heavy metal and drought stress tolerance. Metallomics 2019, 11, 375–389. [Google Scholar] [CrossRef]
- Shao, D.; Li, Y.; Zhu, Q.; Zhang, X.; Liu, F.; Xue, F.; Sun, J. GhGSTF12, a glutathione S-transferase gene, is essential for anthocyanin accumulation in cotton (Gossypium hirsutum L.). Plant Sci. 2021, 305, 110827. [Google Scholar] [CrossRef]
- Milligan, A.S.; Daly, A.; Parry MA, J.; Lazzeri, P.A.; Jepson, I. The expression of a maize glutathione S-transferase gene in transgenic wheat confers herbicide tolerance, both in planta and in vitro. Mol. Breed. 2021, 7, 301–315. [Google Scholar] [CrossRef]
- Christopher, D.G.; Paula, C.; Virginia, W. A Multidrug Resistance–Associated Protein Involved in Anthocyanin Transport in Zea mays, Plant Cell 2004, 16, 1812–1826. Plant Cell 2004, 16, 1812–1826. [Google Scholar]
- Peng, J.; Ruiqi, M.; Chunlai, W.; Nannan, C.; Siyan, L.; Jing, Q.; Shuyan, G.; Yiyong, M. Integration of mRNA and microRNA analysis reveals the molecular mechanisms underlying drought stress tolerance in maize (Zea mays L.). Front. Plant Sci. 2022, 13, 932667. [Google Scholar]
- Zhu, Y.; Liu, Y.; Zhou, K.; Tian, C.; Aslam, M.; Zhang, B.; Liu, W.; Zou, H. Overexpression of ZmEREBP60 enhances drought tolerance in maize. J. Plant Physiol. 2022, 275, 153763. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Li, W.; Peng, Y.; Cao, Y.; Qu, J.; Sun, F.; Yang, Q.; Lu, Y.; Zhang, X.; Zheng, L.; et al. ZmPP2C26 Alternative Splicing Variants Negatively Regulate Drought Tolerance in Maize. Front. Plant Sci. 2022, 13, 851531. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Tai, F.J.; Zheng, Y.; Luo, J.; Gong, S.Y.; Zhang, Z.T.; Li, X.B. Two cotton Cys2/His2-type zinc-fifinger proteins, GhDi19-1 and GhDi19-2, are involved in plant response to salt/drought stress and abscisic acid signaling. Plant Mol. Biol. 2010, 74, 437–452. [Google Scholar] [CrossRef]
- Li, S.; Xu, C.H.; Yang, Y.N.; Xia, G.M. Functional analysis of TaDi19A, a salt-responsive gene in wheat. Plant Cell Environ. 2010, 33, 117–129. [Google Scholar]
- Fryer, M.J.; Oxborough, K.; Mullineaux, P.M.; Baker, N.R. Imaging of photooxidative stress responses in leaves. J. Exp. Bot. 2002, 53, 1249–1254. [Google Scholar]
- Wang, C.L.; Chen, N.N.; Liu, J.Q.; Jiao, P.; Liu, S.Y.; Guan, S.Y.; Ma, Y.Y. Overexpression of ZmSAG39 in maize accelerates leaf senescence in Arabidopsis thaliana. Plant Growth Regul. 2022, 188, 104523–104533. [Google Scholar] [CrossRef]
- Negi, N.P.; Shrivastava, D.C.; Sharma, V.; Sarin, N.B. Overexpression of CuZnSOD from Arachis hypogaea alleviates salinity and drought stress in tobacco. Plant Cell Rep. 2015, 34, 1109–1126. [Google Scholar] [CrossRef]
- Fitter, D.W.; Martin, D.J.; Copley, M.J.; Scotland, R.W.; Langdale, J.A. GLK gene pairs regulate chloroplast development in diverse plant species. Plant J. 2002, 31, 713–727. [Google Scholar] [CrossRef]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of free proline for water-stress studies. Plant Soil 1979, 39, 205–207. [Google Scholar] [CrossRef]
- Xiang, Y.; Sun, X.; Bian, X.; Wei, T.; Han, T.; Yan, J.; Zhang, A. The transcription factor ZmNAC49 reduces stomatal density and improves drought tolerance in maize. J. Exp. Bot. 2021, 72, 1399–1410. [Google Scholar] [CrossRef] [PubMed]
- Stavridou, E.; Voulgari, G.; Michailidis, M.; Kostas, S.; Chronopoulou, E.G.; Labrou, N.E.; Madesis, P.; Nianiou-Obeidat, I. Overexpression of A Biotic Stress-Inducible Pvgstu Gene Activates Early Protective Responses in Tobacco under Combined Heat and Drought. Int. J. Mol. Sci. 2021, 22, 2352. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; An, Y.; Zheng, J.; Shangguan, L.; Wang, L. Genome-wide identification and comparative analysis of GST gene family in apple (Malus domestica) and their expressions under ALA treatment. 3 Biotech 2020, 10, 307. [Google Scholar] [CrossRef] [PubMed]
- Ghangal, R.; Rajkumar, M.S.; Garg, R.; Jain, M. Genome-wide analysis of glutathione S-transferase gene family in chickpea suggests its role during seed development and abiotic stress. Mol. Biol. Rep. 2020, 47, 2749–2761. [Google Scholar] [CrossRef] [PubMed]
- Gallé, Á.; Czékus, Z.; Bela, K.; Horváth, E.; Ördög, A.; Csiszár, J.; Poór, P. Plant Glutathione Transferases and Light. Front. Plant Sci. 2019, 9, 1944. [Google Scholar] [CrossRef]
- Zhang, Y.; He, J.; Xiao, Y.; Zhang, Y.; Liu, Y.; Wan, S.; Liu, L.; Dong, Y.; Liu, H.; Yu, Y. CsGSTU8, a Glutathione S-Transferase from Camellia sinensis, Is Regulated by CsWRKY48 and Plays a Positive Role in Drought Tolerance. Front. Plant Sci. 2021, 12, 795919. [Google Scholar] [CrossRef]
- Sun, H.; Li, J.; Li, X.; Lv, Q.; Chen, L.; Wang, B.; Li, L. RING E3 ubiquitin ligase TaSADR1 negatively regulates drought resistance in transgenic Arabidopsis. Plant Physiol. Biochem. 2022, 170, 255–265. [Google Scholar] [CrossRef]
- Gao, H.; Yu, C.; Liu, R.; Li, X.; Huang, H.; Wang, X.; Zhang, C.; Jiang, N.; Li, X.; Cheng, S.; et al. The Glutathione S-Transferase PtGSTF1 Improves Biomass Production and Salt Tolerance through Regulating Xylem Cell Proliferation, Ion Homeostasis and Reactive Oxygen Species Scavenging in Poplar. Int. J. Mol. Sci. 2022, 23, 11288. [Google Scholar] [CrossRef]
- Li, S.; Wu, Z.; Liu, C.; Fan, L.; He, Y.; Lu, K.; Liu, D.; Feng, G. Overexpression of CsGSH2 Alleviates Propamocarb Residues and Phytotoxicity in Cucumber by Enhancing Antioxidant and Glutathione Detoxification Properties. Agriculture 2022, 12, 1528. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, D.; Chen, L.; Wang, J.; Zhang, W.H. Genome-wide analysis of the Glutathione S-Transferase family in wild Medicago ruthenica and drought-tolerant breeding application of MruGSTU39 gene in cultivated alfalfa. TAG. Theoretical and applied genetics. Theor. Appl. Genet. 2022, 135, 853–864. [Google Scholar] [CrossRef] [PubMed]
- Jing, X.Q.; Zhou, M.R.; Nie, X.M.; Zhang, L.; Shi, P.T.; Abdullah, S.; Miao, H.; Li, W.Q.; Liu, W.T.; Chen, K.M. OsGSTU6 Contributes to Cadmium Stress Tolerance in Rice by Involving in Intracellular ROS Homeostasis. J. Plant Growth Regul. 2021, 40, 945–961. [Google Scholar] [CrossRef]
- Gao, C.; Yang, G.; Guo, Y.; Zhao, Y.; Yang, C. Overexpression of ThGSTZ1 from Tamarix hispida improves tolerance to exogenous ABA and methyl viologen. Trees 2016, 30, 1935–1944. [Google Scholar] [CrossRef]
- Adler, G.; Mishra, A.K.; Maymon, T.; Raveh, D.; Bar-Zvi, D. Overexpression of Arabidopsis ubiquitin ligase AtPUB46 enhances tolerance to drought and oxidative stress. Plant Sci. 2018, 276, 220–228. [Google Scholar] [CrossRef]
- Yunhua, Y.; Adhimoolam, K.; Jinlong, Y.; Tongtong, J.; Rui, R.; Fei, F.; Han, C.; Mengzhuo, L.; Dagang, W.; Kai, L.; et al. The E3 Ligase GmPUB21 Negatively Regulates Drought and Salinity Stress Response in Soybean. Int. J. Mol. Sci. 2022, 23, 6893. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, Y.; Zhang, Y.; Duan, R.; Fan, J.; Jiao, P.; Sun, H.; Guan, S.; Liu, S. Overexpression of Maize Glutathione S-Transferase ZmGST26 Decreases Drought Resistance of Arabidopsis. Agronomy 2022, 12, 2948. https://doi.org/10.3390/agronomy12122948
Jiang Y, Zhang Y, Duan R, Fan J, Jiao P, Sun H, Guan S, Liu S. Overexpression of Maize Glutathione S-Transferase ZmGST26 Decreases Drought Resistance of Arabidopsis. Agronomy. 2022; 12(12):2948. https://doi.org/10.3390/agronomy12122948
Chicago/Turabian StyleJiang, Yushi, Yuzhe Zhang, Ruijie Duan, Jiayi Fan, Peng Jiao, Hongji Sun, Shuyan Guan, and Siyan Liu. 2022. "Overexpression of Maize Glutathione S-Transferase ZmGST26 Decreases Drought Resistance of Arabidopsis" Agronomy 12, no. 12: 2948. https://doi.org/10.3390/agronomy12122948
APA StyleJiang, Y., Zhang, Y., Duan, R., Fan, J., Jiao, P., Sun, H., Guan, S., & Liu, S. (2022). Overexpression of Maize Glutathione S-Transferase ZmGST26 Decreases Drought Resistance of Arabidopsis. Agronomy, 12(12), 2948. https://doi.org/10.3390/agronomy12122948






