Chemical and Microbial Characterization of Washed Rice Water Waste to Assess Its Potential as Plant Fertilizer and for Increasing Soil Health
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals and Media
2.2. Sample Preparation
2.3. Chemical Analyses
2.4. Culture Media and Bacterial Growth
2.5. Bacterial Isolations
2.6. N2 Fixation, Phosphate Solubilization, and Potassium Solubilization
2.7. Bacterial Identification Using 16S rRNA Gene Sequence
2.8. Phylogenetic Analysis
2.9. Data Analysis
3. Results
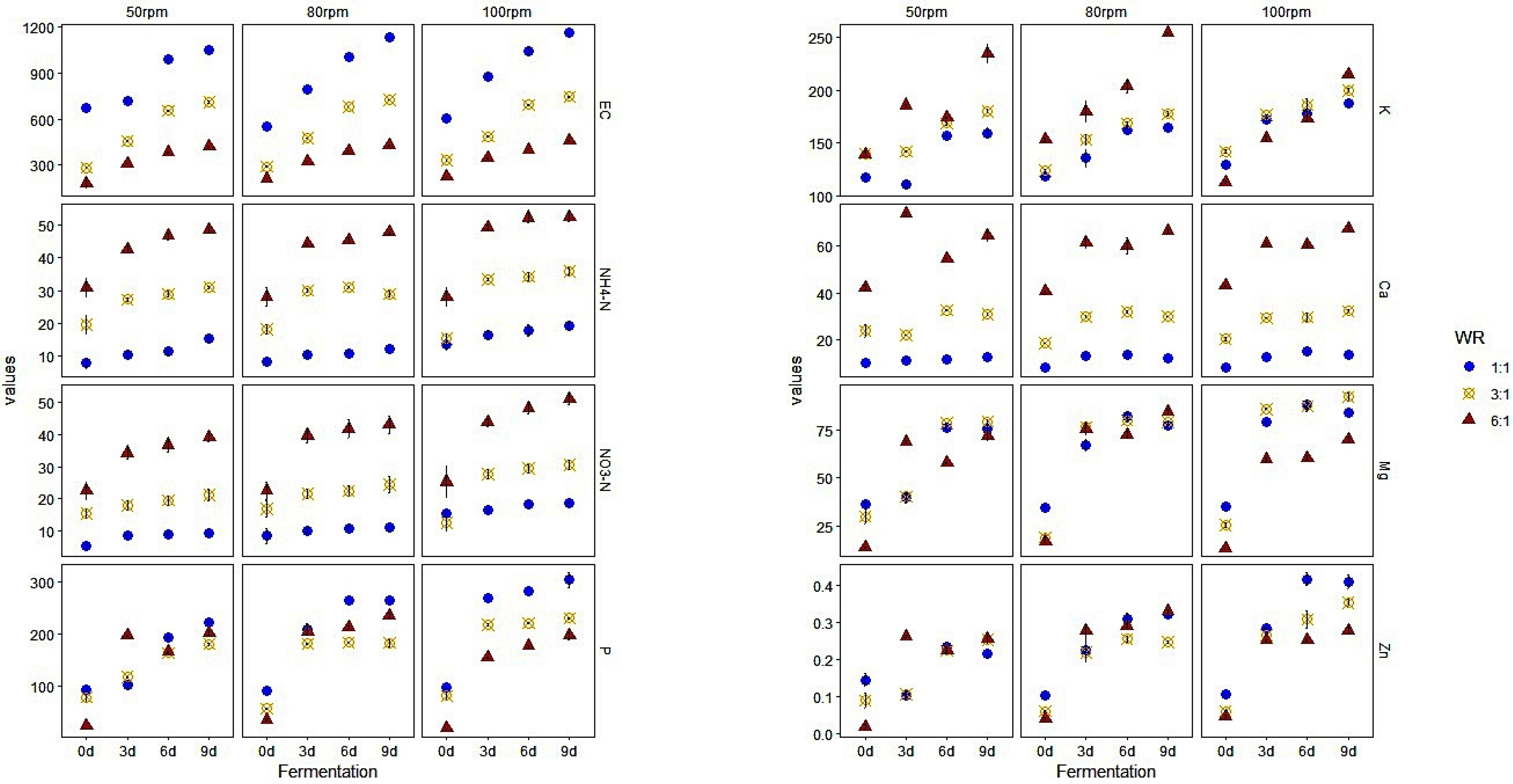
3.1. Chemical Properties and Elemental Concentrations in WRW
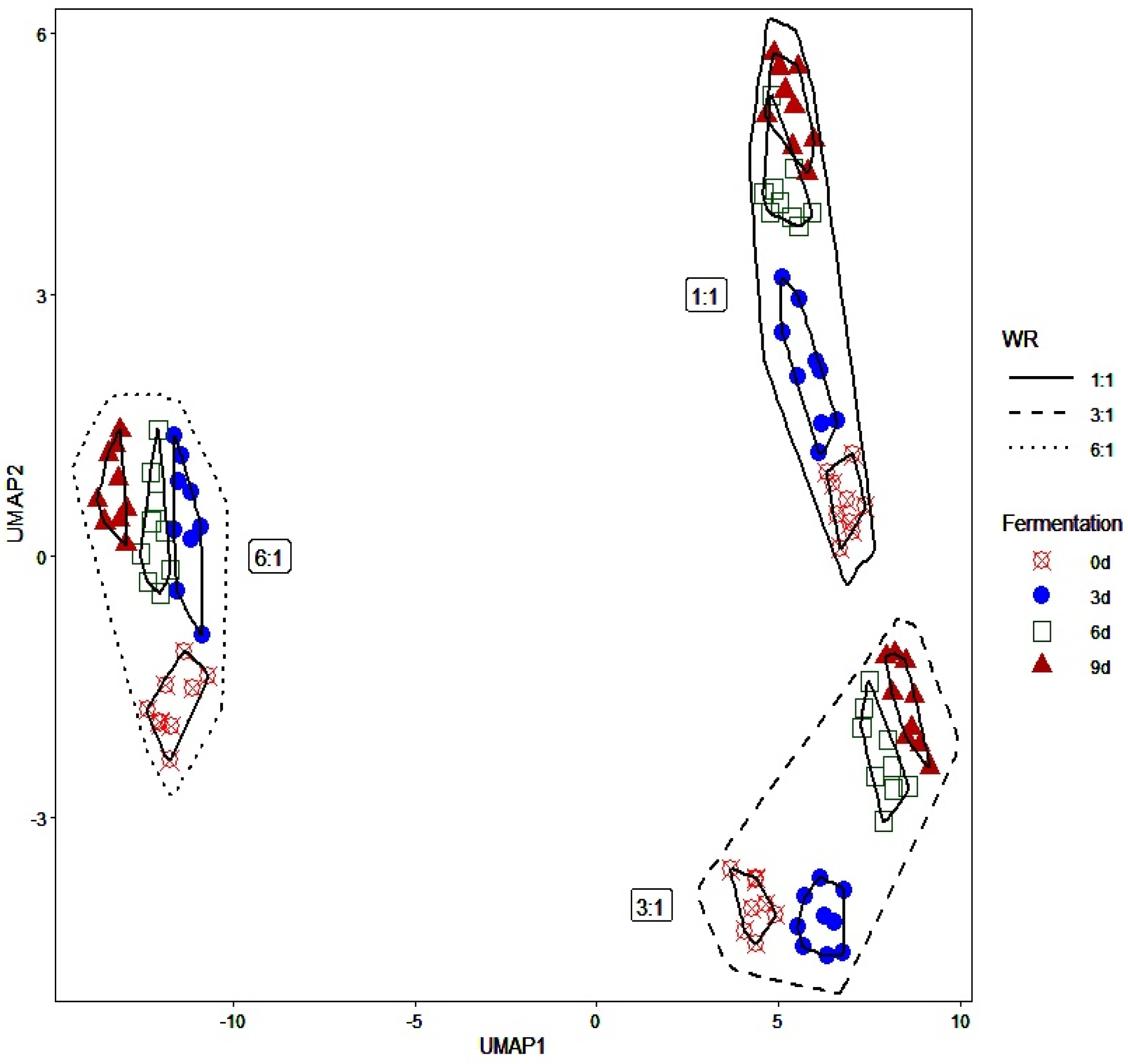
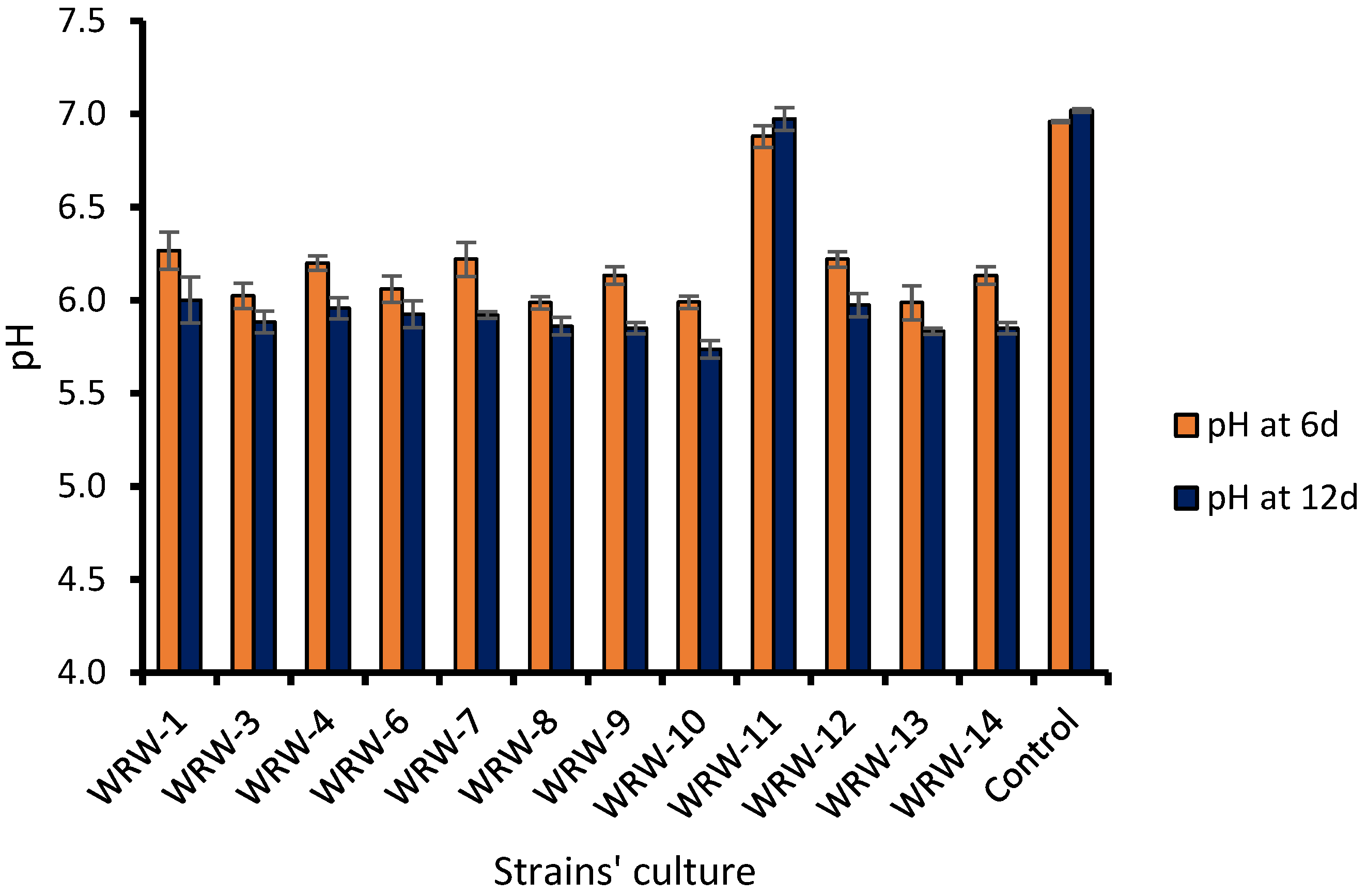
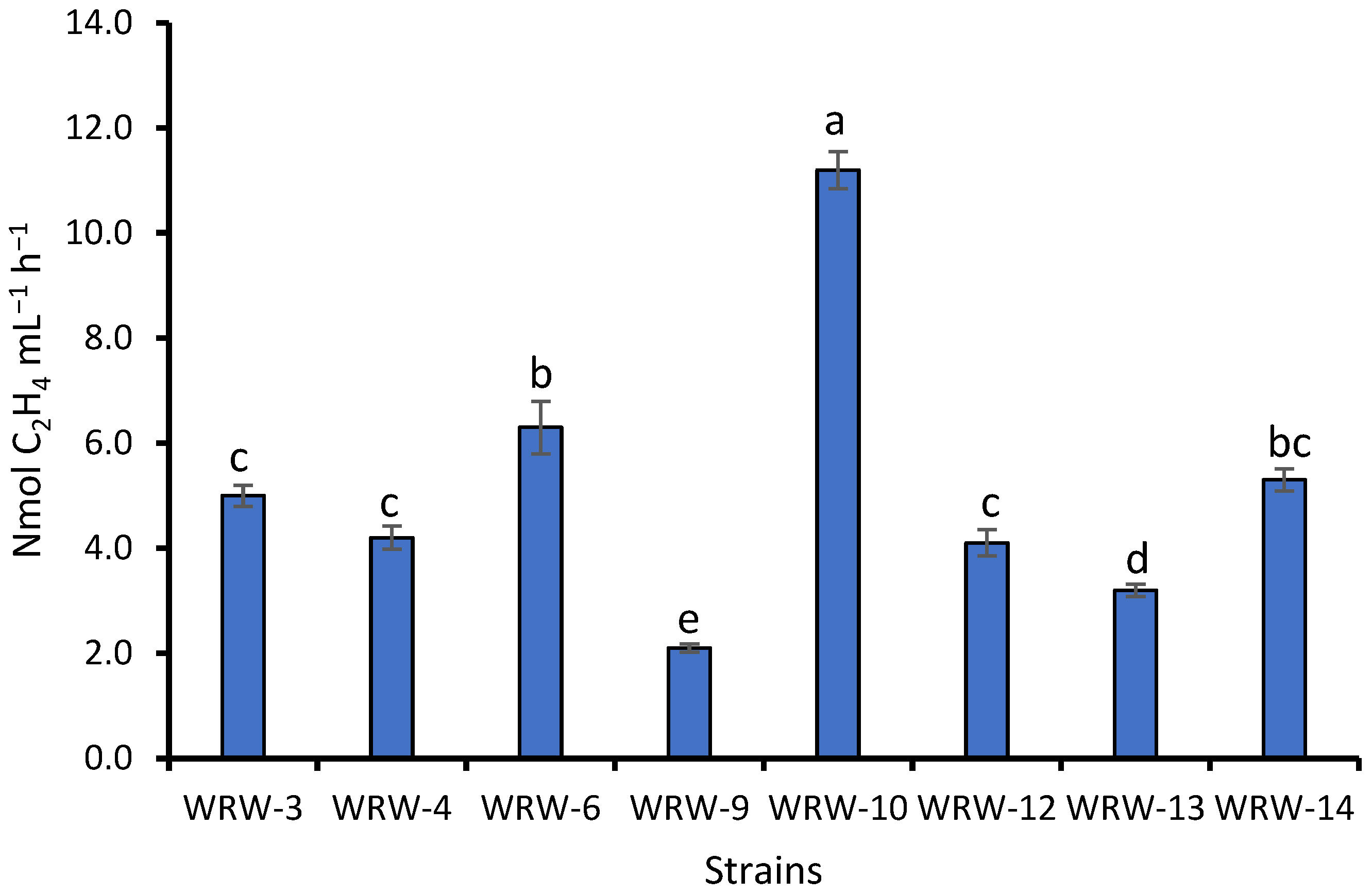
3.2. Bacterial Population and Identification of the WRW
4. Discussion
4.1. Chemical Characteristics of WRW
4.2. Identification and Characterization of the Bacterial Strains
4.3. Potential Use of WRW for Agriculture
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Juliano, B.O. Rice in Human Nutrition; FAO Food and Nutrition Series (26); FAO: Rome, Italy, 1993. [Google Scholar]
- Dini, I.R.; Salbiah, D. Growth and pepper yields (Capsicum annuum L.) by giving a formulation of biological fertilizer of cellulolytic bacteria based on organic liquid waste. J. Phys. Conf.Ser. 2019, 1351, 12097. [Google Scholar]
- He, Q.; Feng, C.; Peng, T.; Chen, N.; Hu, Q.; Hao, C. Denitrification of synthetic nitrate-contaminated groundwater combined with rice washing drainage treatment. Ecol. Eng. 2016, 95, 152–159. [Google Scholar] [CrossRef]
- Malakar, M.C.; Banerjee, S.N. Effect of cooking rice with different volumes of water on the loss of nutrients and on digestibility of rice in vitro. Food Res. 1959, 24, 751–756. [Google Scholar] [CrossRef]
- Nabayi, A.; The, C.B.S.; Ngai, T.P.; Tan, A.K.Z. Wastewater from washed rice water as plant nutrient source: Current understanding and knowledge gaps. Pertanika J. Sci. Technol. 2021, 29, 1347–1369. [Google Scholar] [CrossRef]
- Wulandari, C.G.M.; Muhartini, S.; Trisnowati, S. Pengaruh air cucian beras merah dan beras putih terhadap pertumbuhan dan hasil selada (Lactuca sativa L.). Vegetalika 2012, 1, 24–35. [Google Scholar]
- Ariwibowo, F. Utilization of Chicken Eggshells and Rice Washed Water on the Growth of Tomato Plants (Solonum lycopercium, L.) with Hydroponic Growing Media. Undergraduate Thesis, Universitas Muhammadiah Surakarta, Jakarta, Indonesia, 2012. Unpublished. [Google Scholar]
- Bahar, A. Pengaruh pemberian limbah air cucian beras terhadap pertumbuhan tanaman kangkung darat (Ipomoea reptans Poir). J. Ilm. Mhs. Fak. Pertan. 2016, 3, 1–9. [Google Scholar]
- Handiyanto, S.; Hastuti, U.S.; Prabaningtyas, S. The effect of rice water washing medium on the speed of mycelium growth of white oyster mushrooms (Pleurotus ostreatus var. Florida). Semin. Nas. X Pendidik. Biol. FKIP UN. 2013, 10, 1–6. [Google Scholar]
- Hariyadi, S.P. The Effect of watering on rice washing water, MSG water, and AC (Air Conditioner) wastewater on yields components and tomato yields (Lycopersicon esculentum L.) In the Deep Peatland. Univers. J. 2020, 7, 1–12. [Google Scholar]
- Sairi, F.; Ismail, N.; Ibrahim, N. The effect of FRAW towards the growth of chilli seedlings and its associated microorganisms. Malays. J. Microbiol. 2018, 14, 606–610. [Google Scholar]
- Siagian, A.S. Response of Organic Fertilizer to Liquid water Wash on Growth and Production of Lettuce (Lactuca sativa L.). Undergraduate Thesis, University of Medan Area Faculty of Agricultural Studies Program of Agrotechnology, Medan, Indonesia, 2018. Unpublished. Available online: http://repository.uma.ac.id/handle/123456789/10578 (accessed on 26 May 2021).
- Wardiah, L.; Hafnati, L.R. Potensi limbah air cucian beras sebagai pupuk rganic cair pada pertumbuhan pakchoy (Brassica rapa L.). J. Biol. Edukasi 2014, 6, 34–38. [Google Scholar]
- Yulianingsih, R. Pengaruh air cucian beras terhadap pertumbuhan dan hasil terung ungu (Solanum melongena L.). Publ. Inf. Pertan. 2017, 13, 61–68. [Google Scholar] [CrossRef]
- Beneduzi, A.; Ambrosini, A.; Passaglia, L.M.P. Plant growth-promoting rhizobacteria (PGPR): Their potential as antagonists and biocontrol agents. Genetics Molecular Biol. 2012, 4, 1044–1051. [Google Scholar] [CrossRef] [Green Version]
- Bhattacharyya, P.N.; Jha, D.K. Plant growth-promoting rhizobacteria (PGPR): Emergence in agriculture. World J. Microbiol. Biotechnol. 2012, 28, 1327–1350. [Google Scholar] [CrossRef]
- The Jakarta Post. Village in Bogor uses leftover rice water to grow plants. The Jakarta Post News Desk. 2017. Available online: https://www.thejakartapost.com/news/2017/10/07/village-in-bogor-uses-leftover-rice-water-to-grow-plants.html (accessed on 26 May 2021).
- Supraptiningsih, L.K.; Nuriyanti, R. Processing household waste (washed rice water) into liquid organic fertilizer in Probolinggo Regency Empowering. J. Pengabdi. Masy. Fak. Psikol. 2019, 3, 12–20. [Google Scholar]
- GriSP. Global Rice Science Partnership Rice Almanac, 4th ed.; International Rice Research Institute: Los Baños, Philippines, 2013. [Google Scholar]
- Shahbandeh, M. Total Global Rice Consumption 2008–2021. STATISTA. 2021. Available online: https://www.statista.com/statistics/255977/total-global-rice-consumption (accessed on 5 October 2021).
- OECD. Environmental Outlook to 2050: The Consequences of Iinaction; OECD: Paris, France, 2012. [Google Scholar]
- WWAP. The United Nations World Water Development Report UNESCO; WWAP: Paris, France, 2017. [Google Scholar]
- Nelson, D.W.; Sommers, L. Total carbon, organic carbon, and organic matter. Methods Soil Anal. Part 2 Chem. Microbiol. Prop. 1983, 9, 539–579. [Google Scholar]
- McLean, E.O. Soil pH and lime requirement. Methods Soil Anal. Part 2 Chem. Microbiol. Prop. 1983, 9, 199–224. [Google Scholar]
- Tan, K.Z.; Radziah, O.; Halimi, M.S.; Khairuddin, A.R.; Habib, S.H.; Shamsuddin, Z.H. Isolation and characterization of rhizobia and plant growth-promoting rhizobacteria and their effects on growth of rice seedlings. Am. J. Agric. Biol. Sci. 2014, 9, 342–360. [Google Scholar] [CrossRef] [Green Version]
- Thomas, P.; Sekhar, A.C.; Upreti, R.; Mujawar, M.M.; Pasha, S.S. Optimization of single plate-serial dilution spotting (SP-SDS) with sample anchoring as an assured method for bacterial and yeast cfu enumeration and single colony isolation from diverse samples. Biotechnol. Rep. 2015, 8, 45–55. [Google Scholar] [CrossRef] [Green Version]
- Döbereiner, J.; Day, J.M. Associative symbioses in tropical grasses: Characterization of microorganisms and dinitrogen-fixing sites. In Proceedings of the 1st International Symposium on Nitrogen Fixation; Washington State University Press: Pullman, WD, USA, 1976; Volume 2, pp. 518–538. [Google Scholar]
- Pikovskaya, R.I. Mobilization of phosphorus in soil in connection with vital activity of some microbial species. Mikrobiologiya 1948, 17, 362–370. [Google Scholar]
- Hu, X.; Chen, J.; Guo, J. Two phosphate-and potassium-solubilizing bacteria isolated from Tianmu Mountain, Zhejiang, China. World J. Microbiol. Biotechnol. 2006, 22, 983–990. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T.A. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. Molecular evolutionary genetics analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- RStudio. Rstudio Team Rstudio: Integrated Development Environment for R 2021. 2021. Available online: https://www.rstudio.com/products/rstudio/ (accessed on 9 May 2021).
- DOE. National Water Quality Standards for Malaysia; Department of Environment: Putrajaya, Malaysia, 2017.
- McInnes, L.; Healy, J.; Saul, N.; Großberger, L. UMAP: Uniform manifold approximation and projection. J. Open Source Softw. 2018, 3, 861. [Google Scholar] [CrossRef]
- Atique-ur, R.; Farooq, M.; Rashid, A.; Nadeem, F.; Stuerz, S.; Asch, F.; Bell, R.W.; Siddique, K.H.M. Boron nutrition of rice in different production systems. A review. Agron. Sustain. Dev. 2018, 38, 1–25. [Google Scholar] [CrossRef] [Green Version]
- Petrucci; Ralph, H.; Geoffrey, H.F.; Jeffrey, D.M.; Bissonnette, C. General Chemistry: Principles and Modern Applications, 10th ed.; Pearson Education: Upper Saddle River, NJ, USA, 2011. [Google Scholar]
- Deepa, C.K.; Dastager, S.G.; Pandey, A. Isolation and characterization of plant growth promoting bacteria from non-rhizospheric soil and their effect on cowpea (Vigna unguiculata (L.) Walp.) seedling growth. World J. Microbiol. Biotechnol. 2010, 26, 1233–1240. [Google Scholar] [CrossRef]
- Pranoto, Y.; Anggrahini, S.; Efendi, Z. Effect of natural and Lactobacillus plantarum fermentation on in-vitro protein and starch digestibilities of sorghum flour. Food Biosci. 2013, 2, 46–52. [Google Scholar] [CrossRef]
- Sripriya, G.; Antony, U.; Chandra, T.S. Changes in carbohydrate, free amino acids, organic acids, phytate and HCl extractability of minerals during germination and fermentation of finger millet (Eleusine coracana). Food Chem. 1997, 58, 345–350. [Google Scholar] [CrossRef]
- Blandino, A.; Al-Aseeri, M.E.; Pandiella, S.S.; Cantero, D.; Webb, C. Cereal-based fermented foods and beverages. Food Res. Int. 2003, 36, 527–543. [Google Scholar] [CrossRef]
- Diaz, O.A.; Reddy, K.R.; Moore, P.A. Solubility of inorganic phosphorus in stream water as influenced by pH and calcium concentration. Water Res. 1994, 28, 1755–1763. [Google Scholar] [CrossRef]
- Kleinman, P.J.A.; Wolf, A.M.; Sharpley, A.N.; Beegle, D.B.; Saporito, L.S. Survey of water-extractable phosphorus in livestock manures. Soil Sci. Soc. Am. J. 2005, 69, 701–708. [Google Scholar] [CrossRef] [Green Version]
- Tan, S.L. Utilization of Graywater in Formulating Liquid Organo-Mineral Fertilizer and Effects on Growth and Yield Performance of Chili (Capsicum Annuum, L.). Master’s Thesis, Universiti Putra Malaysia, Selangor, Malaysia, 2021. Unpublished. [Google Scholar]
- Nkhata, S.G.; Ayua, E.; Kamau, E.H.; Shingiro, J.B. Fermentation and germination improve nutritional value of cereals and legumes through activation of endogenous enzymes. Food Sci. Nutr. 2018, 6, 2446–2458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bohn, L.; Meyer, A.S.; Rasmussen, S.K. Phytate: Impact on environment and human nutrition. A challenge for molecular breeding. J. Zhejiang Univ. Sci. B 2008, 9, 165–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Osman, M.A. Effect of traditional fermentation process on the nutrient and antinutrient contents of pearl millet during preparation of Lohoh. J. Saudi Soc. Agric. Sci. 2011, 10, 1–6. [Google Scholar] [CrossRef] [Green Version]
- He, Q.; Feng, C.; Chen, N.; Zhang, D.; Hou, T.; Dai, J.; Hao, C.; Mao, B. Characterizations of dissolved organic matter and bacterial community structures in rice washing drainage (RWD)-based synthetic groundwater denitrification. Chemosphere 2019, 215, 142–152. [Google Scholar] [CrossRef]
- Neina, D. The role of soil pH in plant nutrition and soil remediation. Appl. Environ. Soil Sci. 2019, 2019, 1–9. [Google Scholar] [CrossRef]
- Musa, A.M.; Ishak, C.F.; Karam, D.S.; Jaafar, N.M. Effects of fruit and vegetable wastes and biodegradable municipal wastes co-mixed composts on nitrogen dynamics in an Oxisol. Agronomy 2020, 10, 1609. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, W.; Duan, P.; Cong, Y.; An, T.; Yu, N.; Zou, H.; Dang, X.; An, J.; Fan, Q.; et al. Evaluation and simulation of nitrogen mineralization of paddy soils in Mollisols area of Northeast China under waterlogged incubation. PLoS ONE 2017, 12, 1–19. [Google Scholar] [CrossRef]
- Rousk, J.; Brookes, P.C.; Bååth, E. Contrasting soil pH effects on fungal and bacterial growth suggest functional redundancy in carbon mineralization. Appl. Environ. Microbiol. 2009, 75, 1589–1596. [Google Scholar] [CrossRef] [Green Version]
- Gómez-Brandón, M.; Lazcano, C.; Domínguez, J. The evaluation of stability and maturity during the composting of cattle manure. Chemosphere 2008, 70, 436–444. [Google Scholar] [CrossRef]
- Gao, M.; Li, B.; Yu, A.; Liang, F.; Yang, L.; Sun, Y. The effect of aeration rate on forced-aeration composting of chicken manure and sawdust. Bioresour. Technol. 2010, 101, 1899–1903. [Google Scholar] [CrossRef]
- Ding, X.; Jiang, Y.; Zhao, H.; Guo, D.; He, L.; Liu, F.; Zhou, Q.; Nandwani, D.; Hui, D.; Yu, J. Electrical conductivity of nutrient solution influenced photosynthesis, quality, and antioxidant enzyme activity of pak choi (Brassica campestris L. sp. Chinensis) in a hydroponic system. PLoS ONE 2018, 13, e202090. [Google Scholar] [CrossRef] [PubMed]
- Nemali, K.S.; Van Iersel, M.W. Light intensity and fertilizer concentration: I. Estimating optimal fertilizer concentrations from water-use efficiency of wax begonia. HortScience 2004, 39, 1287–1292. [Google Scholar] [CrossRef] [Green Version]
- Adugna, G. A review on impact of compost on soil properties, water use and crop productivity. Agric. Sci. Res. J. 2016, 4, 93–104. [Google Scholar] [CrossRef]
- Dunlap, C.A.; Kim, S.J.; Kwon, S.W.; Rooney, A.P. Bacillus velezensis is not a later heterotypic synonym of Bacillus amyloliquefaciens; Bacillus methylotrophicus, Bacillus amyloliquefaciens subsp. Plantarum and ‘Bacillus oryzicola’ are later heterotypic synonyms of Bacillus velezensis based on phylogenom. Int. J. Syst. Evol. Microbiol. 2016, 66, 1212–1217. [Google Scholar] [CrossRef] [PubMed]
- Aeron, A.; Kumar, D.; Gurukula, M.; Vishwavidyalaya, K.; Pgpr, Z.S.; View, I.; Jha, C.K. Enterobacter: Role in plant growth promotion. In Bacteria in Agrobiology; Springer: Berlin/Heidelberge, Germany, 2011; pp. 159–182. [Google Scholar]
- Chen, X.H.; Koumoutsi, A.; Scholz, R.; Eisenreich, A.; Schneider, K.; Heinemeyer, I.; Morgenstern, B.; Voss, B.; Hess, W.R.; Reva, O.; et al. Comparative analysis of the complete genome sequence of the plant growth-promoting bacterium Bacillus amyloliquefaciens FZB42. Nat. Biotechnol. 2007, 25, 1007–1014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iniguez, A.L.; Dong, Y.; Triplett, E.W. Nitrogen fixation in wheat provided by Klebsiella pneumoniae 342. Mol. Plant-Microbe Interact. 2004, 17, 1078–1085. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shariati, V.J.; Malboobi, M.A.; Tabrizi, Z.; Tavakol, E.; Owilia, P.; Safari, M. Comprehensive genomic analysis of a plant growth-promoting rhizobacterium Pantoea agglomerans strain P5 /631/208/212/748 /631/208/464 /38/43 /45 article. Sci. Rep. 2017, 7, 15820. [Google Scholar] [CrossRef]
- Messiha, N.A.S.; van Diepeningen, A.D.; Farag, N.S.; Abdallah, S.A.; Janse, J.D.; van Bruggen, A.H.C. Stenotrophomonas maltophilia: A new potential biocontrol agent of Ralstonia solanacearum, causal agent of potato brown rot. Eur. J. Plant Pathol. 2007, 118, 211–225. [Google Scholar] [CrossRef] [Green Version]
- Suhaimee, S.; Ibrahim, I.Z.; Abdul Wahab, M.A.M. Organic agriculture in Malaysia. Available online: https://ap.fftc.org.tw/article/1010 (accessed on 27 September 2021).
- Park, H.; DuPonte, M.W. How to Cultivate Indigenous Microorganisms. Trop. Agric. Univ. Hawaii 2008, 9, 1–7. [Google Scholar]
- Yadav, A.N.; Kumar, R.; Kumar, S.; Kumar, V.; Sugitha, T.C.K.; Singh, B.; Chauahan, V.S.; Dhaliwal, H.S.; Saxena, A.K. Beneficial microbiomes: Biodiversity and potential biotechnological applications for sustainable agriculture and human health. J. Appl. Biol. Biotechnol. 2017, 5, 45–57. [Google Scholar] [CrossRef] [Green Version]
- Zakry, F.A.A.; Shamsuddin, Z.H.; Khairuddin, A.R.; Zin, Z.Z.; Anuar, A.R. Inoculation of Bacillus sphaericus UPMB-10 to young oil palm and measurement of its uptake of fixed nitrogen using the 15N isotope dilution technique. Microbes Environ. 2012, 27, 257–262. [Google Scholar] [CrossRef] [Green Version]
- Katupitiya, S.; Millet, J.; Vesk, M.; Viccars, L.; Zeman, A.; Lidong, Z.; Elmerich, C.; Kennedy, I.R. A mutant of Azospirillum brasilense Sp7 impaired in flocculation with a modified colonization pattern and superior nitrogen fixation in association with wheat. Appl. Environ. Microbiol. 1995, 61, 1987–1995. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naher, U.A.; Radziah, O.; Shamsuddin, Z.H.; Halimi, M.S.; Mohd Razi, I. Isolation of diazotrophs from different soils of Tanjong Karang rice growing area in Malaysia. Int. J. Agric. Biol. 2009, 11, 547–552. [Google Scholar]
- Tang, A.; Haruna, A.O.; Majid, N.M.A.; Jalloh, M.B. Potential PGPR properties of cellulolytic, nitrogen-fixing, and phosphate-solubilizing bacteria of a rehabilitated tropical forest soil. Microorganisms 2020, 8, 442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, A.; Haruna, A.O.; Majid, N.M.A.; Jalloh, M.B. Effects of selected functional bacteria on maize growth and nutrient use efficiency. Microorganisms 2020, 8, 854. [Google Scholar] [CrossRef]
- Talaat, N.B.; Ghoniem, A.E.; Abdelhamid, M.T.; Shawky, B.T. Effective microorganisms improve growth performance, alter nutrients acquisition and induce compatible solutes accumulation in common bean (Phaseolus vulgaris L.) plants subjected to salinity stress. Plant Growth Regul. 2015, 75, 281–295. [Google Scholar] [CrossRef]
- Dastager, S.G.; Deepa, C.K.; Puneet, S.C.; Nautiyal, C.S.; Pandey, A. Isolation and characterization of plant growth-promoting strain Pantoea NII-186. from Western Ghat forest soil, India. Lett. Appl. Microbiol. 2009, 49, 20–25. [Google Scholar] [CrossRef]
- Rodríguez, H.; Fraga, R. Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol. Adv. 1999, 17, 319–339. [Google Scholar] [CrossRef]
- Pérez, E.; Sulbarán, M.; Ball, M.M.; Yarzábal, L.A. Isolation and characterization of mineral phosphate-solubilizing bacteria naturally colonizing a limonitic crust in the south-eastern Venezuelan region. Soil Biol. Biochem. 2007, 39, 2905–2914. [Google Scholar] [CrossRef] [Green Version]
- Sugumaran, P.; Janarthanum, B. Solubilization of potassium containing minerals by bacteria and their effect of plant growth. World J. Agric. Sci. 2007, 3, 350–355. [Google Scholar]
- We, A.C.E.; Aris, A.; Zain, N.A.M.; Muda, K.; Sulaiman, S. Influence of static mixer on the development of aerobic granules for the treatment of low-medium strength domestic wastewater. Chemosphere 2021, 263, 128209. [Google Scholar] [CrossRef]
- Karakas, I.; Sam, S.B.; Cetin, E.; Dulekgurgen, E.; Yilmaz, G. Resource recovery from an aerobic granular sludge process treating domestic wastewater. J. Water Process Eng. 2020, 34, 101148. [Google Scholar] [CrossRef]
- Sancho, I.; Licon, E.; Valderrama, C.; de Arespacochaga, N.; López-Palau, S.; Cortina, J.L. Recovery of ammonia from domestic wastewater effluents as liquid fertilizers by integration of natural zeolites and hollow fibre membrane contactors. Sci. Total Environ. 2017, 584–585, 244–251. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; An, S.; Choi, Y. Ammonia harvesting via membrane gas extraction at moderately alkaline pH: A step toward net-profitable nitrogen recovery from domestic wastewater. Chem. Eng. J. 2021, 405, 126662. [Google Scholar] [CrossRef]
- Sahu, O.; Rao, D.G.; Gopal, R.; Tiwari, A.; Pal, D. Treatment of wastewater from sugarcane process industry by electrochemical and chemical process: Aluminum (metal and salt). J. Water Process Eng. 2017, 17, 50–62. [Google Scholar] [CrossRef]
- Guerreiro, R.C.S.; Jerónimo, E.; Luz, S.; Pinheiro, H.M.; Prazeres, A.R. Cheese manufacturing wastewater treatment by combined physicochemical processes for reuse and fertilizer production. J. Environ. Manage. 2020, 264, 110470. [Google Scholar] [CrossRef]
- Gonçalves, R.F.; Assis, T.I.; Maciel, G.B.; Borges, R.M.; Cassini, S.T.A. Co-digestion of municipal wastewater and microalgae biomass in an up flow anaerobic sludge blanket reactor. Algal Resour. 2020, 52, 102117. [Google Scholar] [CrossRef]
- Cainglet, A.; Tesfamariam, A.; Heiderscheidt, E. Organic polyelectrolytes as the sole precipitation agent in municipal wastewater treatment. J. Environ. Manag. 2020, 271, 111002. [Google Scholar] [CrossRef] [PubMed]
- Lahav, O.; Telzhensky, M.; Zewuhn, A.; Gendel, Y.; Gerth, J.; Calmano, W.; Birnhack, L. Struvite recovery from municipal-wastewater sludge centrifuge supernatant using seawater NF concentrate as a cheap Mg(II) source. Sep. Purif. Technol. 2013, 108, 103–110. [Google Scholar] [CrossRef]
- Sakiah; Wahyuni, M. Analysis of C-organic, nitrogen, phosphorus, and potassium in application areas and without application of palm oil mill effluent. J. Agric. Vet. Sci. 2018, 11, 23–27. [Google Scholar]
- Muyang, K.S.T.; Ahmed, O.H.; Khew, C.Y.; Zehnder, J.A.M.; Jalloh, M.B.; Musah, A.A.; Abdu, A. Chemical and biological characteristics of organic amendments produced from selected agro-wastes with potential for sustaining soil health: A laboratory assessment. Sustainability 2021, 13, 4919. [Google Scholar] [CrossRef]




| Parameters | Rice Grain | WRW * | Tap Water † |
|---|---|---|---|
| pH | – | 6.53 ± 0.02 | 6.58 ± 0.02 |
| EC (µS cm−1) | – | 372.83 ± 34.53 | 125.36 ± 28.21 |
| Ash (%) | 0.95 ± 0.04 | – | – |
| TOC (%) | 30.30 ± 0.21 | 2.64 ± 0.72 | Trace |
| Moisture (%) | 14.39 ± 0.06 | 99.32 ± 0.31 | – |
| Total C (%) | 40.30 ± 0.01 | 3.87 ± 0.24 | 0.03 ± 0.002 |
| Total N (mg kg−1) | 12,500 ± 100.70 | 80.50 ± 5.20 | 30.20 ± 4.12 |
| -N (mg kg−1) | 215.45 ± 4.41 | 18.88 ± 1.68 | 1.44 ± 0.04 |
| (mg kg−1) | 100.82 ± 8.53 | 16.02 ± 1.41 | 1.45 ± 0.03 |
| C:N | 32.24 ± 0.02 | 48.3 ± 5.64 | 0.50 ± 0.001 |
| S (mg kg−1) | 1000 ± 38.12 | 452 ± 62.15 | 100 ± 9.64 |
| P (mg kg−1) | 1320.83 ± 34.04 | 64.64 ± 5.76 | 0.05 ± 0.02 |
| K (mg kg−1) | 1130.83 ± 22.64 | 130.66 ± 2.55 | 5.74 ± 0.15 |
| Ca (mg kg−1) | 427.08 ± 5.72 | 23.97 ± 2.68 | 10.95 ± 0.06 |
| Mg (mg kg−1) | 244.93 ± 10.26 | 25.23 ± 1.78 | 0.97 ± 0.06 |
| Cu (µg kg−1) | 5250 ± 120.40 | 188.52 ± 11.50 | 2.4 ± 1.02 |
| Zn (µg kg−1) | 5020 ± 97.04 | 73.77 ± 7.73 | 5.3 ± 1.30 |
| B (µg kg−1) | 1400 ± 96.45 | 121.18 ± 23.93 | 1.2 ± 1.02 |
| (a) | ||||||||
| W:R | pH | C | C:N | -N | Ca | S | ||
| % | mg kg−1 | |||||||
| 1:1 | 6.48b | 2.48c | 35.17b | 9.94c | 9.78b | 9.05c | 110.68c | |
| 3:1 | 6.47b | 3.76b | 38.68b | 17.74b | 14.94b | 21.02b | 449.33b | |
| 6:1 | 6.63a | 5.37a | 68.05a | 28.95a | 23.35a | 41.84a | 797.13a | |
| SE (±) | 0.04 | 0.13 | 5.35 | 1.23 | 1.48 | 0.55 | 49.80 | |
| (b) | ||||||||
| Washing Intensity (R) | W:R | EC | TN | P | K | Mg | Cu | Zn |
| rpm | µS cm−1 | mg kg−1 | ||||||
| 100 | 1:1 | 607.0b | 115.0a | 98.0a | 129.7c | 35.4a | 0.111e | 0.105ab |
| 80 | 551.3c | 67.0bc | 90.8ab | 118.1de | 34.9a | 0.112e | 0.101ab | |
| 50 | 671.7a | 56.6c | 93.3ab | 117.5de | 36.5a | 0.133de | 0.144a | |
| 100 | 3:1 | 332.0d | 101.0ab | 82.0b | 141.9b | 25.7bc | 0.243a | 0.060bcd |
| 80 | 291.1d | 93.0ab | 57.7c | 123.8cd | 19.3cd | 0.193bc | 0.060bcd | |
| 50 | 283.4d | 100.6ab | 79.2b | 139.6b | 30.0ab | 0.161cd | 0.089bc | |
| 100 | 6:1 | 223.5e | 50.0c | 19.6e | 112.9e | 13.5d | 0.244a | 0.046cd |
| 80 | 212.7e | 47.0c | 35.4d | 153.1a | 17.2d | 0.260a | 0.040d | |
| 50 | 182.7e | 35.4c | 25.3de | 138.9b | 14.2d | 0.240ab | 0.018d | |
| W:R | *** | *** | ns | * | ns | * | ns | |
| R | *** | *** | *** | *** | *** | *** | *** | |
| R × (W:R) | *** | *** | ** | *** | ** | *** | * | |
| SE (±) | 10.90 | 7.14 | 3.13 | 1.76 | 1.47 | 0.0098 | 0.0095 | |
| Molecular Identification | ||||
|---|---|---|---|---|
| Strain | Fermentation Period | Accession Number | Close Relatives in NCBI | Similarity (%) |
| WRW-1 | 3d | MW365554.1 | Bacillus velezensis strain HSB1 | 99.35 |
| WRW-3 | 3d | MW365555.1 | Enterobacter ludwigii strain SDI-19 | 98.75 |
| WRW-4 | 3d | MW365556.1 | Enterobacter sp. Strain LSB19 | 99.10 |
| WRW-6 | 0d | MW365557.1 | Enterobacter sp. Strain LSB3 | 97.59 |
| WRW-7 | 6d | MW365558.1 | Enterobacter mori strain BC1 | 98.51 |
| WRW-8 | 6d | MW365559.1 | Bacillus velezensis strain 2656 | 99.70 |
| WRW-9 | 6d | MW365560.1 | Stenotrophomonas maltophilia strain JM11 | 99.87 |
| WRW-10 | 9d | MW365561.1 | Enterobacter sp. Strain LSB10 | 99.49 |
| WRW-11 | 9d | MW365562.1 | Enterobacter mori strain BC1 | 99.19 |
| WRW-12 | 3d | MW365563.1 | Klebsiella pneumoniae strain LB-AMP3KSU | 99.87 |
| WRW-13 | 3d | MW365564.1 | Pantoea agglomerans stain SVMR | 97.92 |
| WRW-14 | 9d | MW365565.1 | Stenotrophomonas maltophilia strain F41 | 99.47 |
| Strains | N2 Fixation | PS | KS | PSI | KSI | Gram Stain |
|---|---|---|---|---|---|---|
| Bacillus velezensis strain WRW-1 | − | + | + | 1.3 | 1.2 | − |
| Enterobacter ludwigii strain WRW-3 | ++ | + | − | 1.4 | 0 | + |
| Enterobacter sp. strain WRW-4 | ++ | + | − | 1.2 | 0 | − |
| Enterobacter sp. strain WRW-6 | + | + | − | 1.2 | 0 | − |
| Enterobacter mori strain WRW-7 | − | ++ | − | 1.2 | 0 | + |
| Bacillus velezensis strain WRW-8 | − | + | + | 1.4 | 1.3 | + |
| Stenotrophomonas maltophilia strain WRW-9 | ++ | + | ++ | 1.5 | 1.3 | − |
| Enterobacter sp. strain 10 | ++ | + | − | 1.6 | 0 | − |
| Enterobacter mori strain WRW-11 | − | + | − | 1.16 | 0 | − |
| Klebsiella pneumoniae strain WRW-12 | ++ | ++ | ++ | 1.50 | 2.3 | − |
| Pantoea agglomerans strain 13 | ++ | ++ | ++ | 1.53 | 1.2 | − |
| Stenotrophomonas maltophilia strain WRW-14 | + | + | + | 1.2 | 1.1 | − |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nabayi, A.; Sung, C.T.B.; Zuan, A.T.K.; Paing, T.N.; Akhir, N.I.M. Chemical and Microbial Characterization of Washed Rice Water Waste to Assess Its Potential as Plant Fertilizer and for Increasing Soil Health. Agronomy 2021, 11, 2391. https://doi.org/10.3390/agronomy11122391
Nabayi A, Sung CTB, Zuan ATK, Paing TN, Akhir NIM. Chemical and Microbial Characterization of Washed Rice Water Waste to Assess Its Potential as Plant Fertilizer and for Increasing Soil Health. Agronomy. 2021; 11(12):2391. https://doi.org/10.3390/agronomy11122391
Chicago/Turabian StyleNabayi, Abba, Christopher Teh Boon Sung, Ali Tan Kee Zuan, Tan Ngai Paing, and Nurul Izzati Mat Akhir. 2021. "Chemical and Microbial Characterization of Washed Rice Water Waste to Assess Its Potential as Plant Fertilizer and for Increasing Soil Health" Agronomy 11, no. 12: 2391. https://doi.org/10.3390/agronomy11122391
APA StyleNabayi, A., Sung, C. T. B., Zuan, A. T. K., Paing, T. N., & Akhir, N. I. M. (2021). Chemical and Microbial Characterization of Washed Rice Water Waste to Assess Its Potential as Plant Fertilizer and for Increasing Soil Health. Agronomy, 11(12), 2391. https://doi.org/10.3390/agronomy11122391






