Detection and Diagnosis of Xylella fastidiosa by Specific Monoclonal Antibodies
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
2.2. Plant Material
2.3. Production and Characterization of Monoclonal and Polyclonal Antibodies against X. fastidiosa
2.4. Ethics Statement
2.5. Reactivity and Specificity
2.5.1. Double Antibody Sandwich Enzyme Linked Immunosorbent Assay (DAS-ELISA)
2.5.2. Double Antibody Sandwich Indirect Enzyme Linked Immunosorbent Assay (DASI-ELISA)
2.5.3. Direct Tissue Print-ELISA or Direct Tissue Blot Immunoassay (DTBIA) in Naturally Infected Samples
2.6. Sensitivity in DAS-ELISA and DASI-ELISA
2.7. Detection of X. fastidiosaby DAS-ELISA Mab in Naturally Infected Samples
2.8. Comparison of DAS-ELISA MAb and Real-Time PCR for Detection of X. fastidiosa
3. Results
3.1. Production of Monoclonal Antibodies (MAb) and Polyclonal Antibodies
3.2. Reactivity in Several Techniques and Specificity
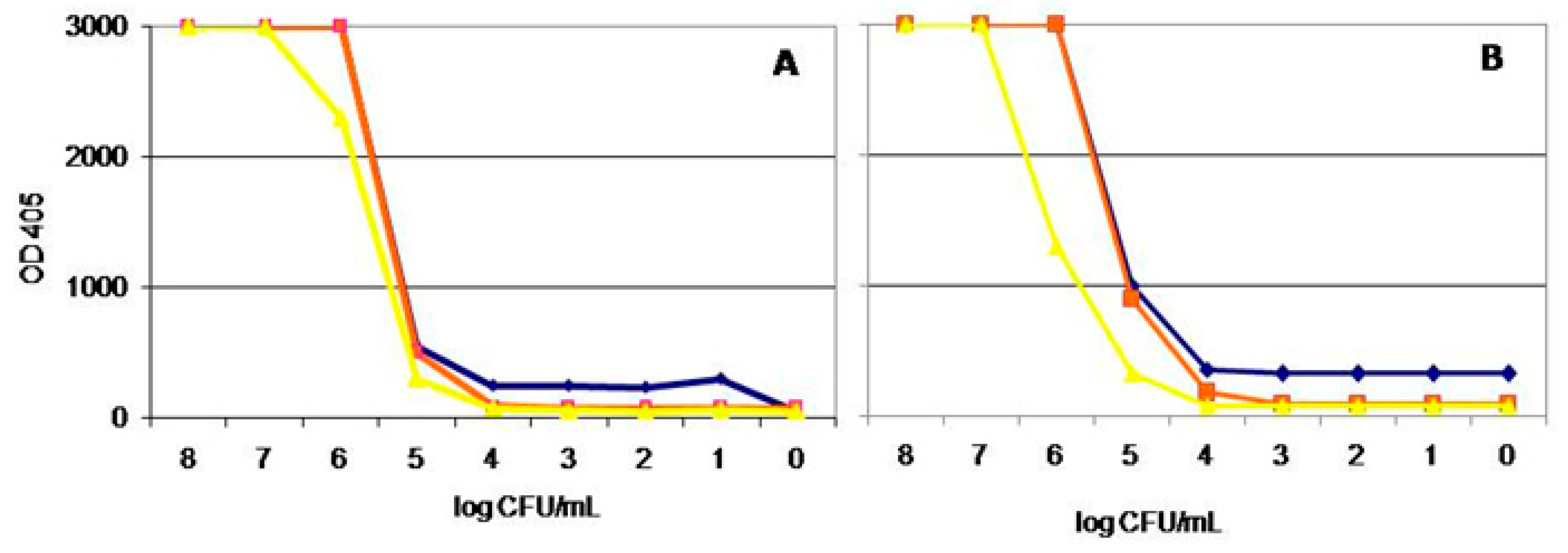
3.3. Sensitivity
3.4. Comparison of DAS-ELISA MAb with the Gold Standard Real-Time PCR for Detection of X. fastidiosa in Field Samples
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- European Union. Commission implementing regulation (EU) 2019/2072 of 28 November 2019 establishing uniform conditions for the implementation of Regulation (EU) 2016/2031 of the European Parliament and the Council, as regards protective measures against pests of plants, and repealing Commission Regulation (EC) No 690/2008 and amending Commission Implementing Regulation (EU) 2018/2019. Off. J. Eur. Union 2019, L319, 1–279. [Google Scholar]
- EFSA (European Food Safety Authority). Update of the Xylella spp. host plant database—Systematic literature search up to 30 June 2019. EFSA J. 2020, 18, e06114. [Google Scholar]
- Purcell, A.H. Paradigms: Examples from the bacterium Xylellafastidiosa. Ann. Rev. Phytopathol. 2013, 51, 339–356. [Google Scholar] [CrossRef] [PubMed]
- Landa, B.B.; Marco-Noales, E.; López, M.M. EnfermedadesCausadaspor la BacteriaXylellafastidiosa; CajamarCaja Rural: Almería, Spain, 2017; 320p. [Google Scholar]
- Sicard, A.; Zeilinger, A.R.; Vanhove, M.; Schartel, T.E.; Beal, D.J.; Daugherty, M.P.; Almeida, R.P.P. Xylellafastidiosa: Insights into an emerging plant pathogen. Ann. Rev. Phytopathol. 2018, 56, 181–202. [Google Scholar] [CrossRef]
- EPPO. PM 7/24 (4) Xylellafastidiosa. Bull. OEPP/EPPO Bull. 2019, 49, 175–227. [Google Scholar] [CrossRef]
- Landa, B.B.; Castillo, A.I.; Giampetruzzi, A.; Kahn, A.; Román-Écija, M.; Velasco-Amo, M.P.; Navas, J.A.; Marco-Noales, E.; Barbé, S.; Moralejo, E.; et al. Emergence of a plant pathogen in Europe associated with multiple intercontinental introductions. Appl. Environ. Microbiol. 2020, 86, e01521-19. [Google Scholar] [CrossRef]
- European Union. Commission Implementing Regulation (EU) 2020/1201 of 14 August 2020 as regards measures to prevent the introduction into and the spread within the Union of Xylellafastidiosa (Wells et al.). Off. J. Eur. Union 2020, L269, 2–39. [Google Scholar]
- IPPC-FAO. Diagnostic Protocols for Regulated Pests: Xylellafastidiosa. Int. Stand. Phytosanit. Meas. ISPM 27 2018, DP25, 1–32. [Google Scholar]
- EFSA (EuropeanFood Safety Authority); Lázaro, E.; Parnell, S.; Vicent Civera, A.; Schans, J.; Schenk, M.; Schrader, G.; Cortiñas Abrahantes, J.; Zancanaro, G.; Vos, S. Guidelines for statistically sound and risk-based surveys of Xylellafastidiosa. EFSA Support. Publ. 2020, 17, 1873E. [Google Scholar] [CrossRef]
- De Boer, S.H.; López, M.M. New grower-friendly methods for plant pathogen monitoring. Ann. Rev. Phytopathol. 2012, 50, 197–218. [Google Scholar] [CrossRef]
- Boscia, D.; Myrta, A. Serological detection of viruses included in certification protocols for stone fruits. Options Méditerr. 1998, 19, 171–190. [Google Scholar]
- Cambra, M.; Boscia, D.; Gil, M.; Bertolini, E.; Olmos, A. Immunology and immunological assays applied to the detection, diagnosis and control of fruit tree viruses. In Virus and Virus-Like Disease of Pome and Stone Fruits; Hadidi, A., Barba, M., Candresse, T., Jelkmann, W., Eds.; APS Press: St. Paul, MN, USA, 2011; pp. 303–313. [Google Scholar]
- López, M.M.; Bertolini, E.; Caruso, P.; Penyalver, R.; Marco-Noales, E.; Gorris, M.T.; Morente, C.; Salcedo, C.; Cambra, M.; Llop, P. Advantages of an integrated approach for diagnosis of quarantine pathogenic bacteria in plant material. Phytopathol. Pol. 2005, 35, 49–56. [Google Scholar]
- Sherald, J.L.; Lei, J.D. Evaluation of a rapid ELISA test kit for detection of Xylellafastidiosa in landscaping trees. Plant Dis. 1991, 75, 200–203. [Google Scholar] [CrossRef]
- Hartung, J.S.; Beretta, J.; Brlansky, R.H.; Spisso, J.; Lee, R. Citrus variegated chlorosis bacterium: Axenic culture, pathogenicity, and serological relationships with other strains of Xylellafastidiosa. Phytopathology 1994, 84, 591–597. [Google Scholar] [CrossRef]
- Lee, R.F.; Beretta, M.J.G.; Derrick, K.S.; Hooker, M.E. Development of a serological assay for citrus variegated chlorosis—A new disease of citrus in Brazil. Proc. Fla. State Hortic. Soc. 1992, 105, 32–35. [Google Scholar]
- Chang, C.J.; Garnier, M.; Zreik, L.; Rossetti, V.; Bove, J.M. Culture and serological detection of the xylem-limited bacterium causing citrus variegated chlorosis and its identification as a strain of Xylellafastidiosa. Curr. Microbiol. 1993, 27, 137–142. [Google Scholar] [CrossRef]
- Carbajal, D.; Morano, K.A.; Morano, L.D. Indirect immunofluorescence microscopy for direct detection of Xylellafastidiosa in xylem sap. Curr. Microbiol. 2004, 49, 372–375. [Google Scholar] [CrossRef]
- Djelouah, K.; Frasheri, D.; Valentini, F.; D’Onghia, A.M.; Digiaro, M. Direct tissue blot immunoassay for detection of Xylellafastidiosa in olive trees. Phytopathol. Mediterr. 2014, 53, 559–564. [Google Scholar]
- Harlow, E.; Lane, D. Antibodies: A Laboratory Manual; Cold Spring Harbor Laboratory: New York, NY, USA, 1988; 726p. [Google Scholar]
- Koller, G.; Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 1975, 256, 495–499. [Google Scholar] [CrossRef]
- Garnier, M.; Chang, C.J.; Zreik, L.; Rossetti, V.; Bové, J.M. Citrus variegated chlorosis: Serological detection of Xylellafastidiosa, the bacterium associated with the disease. Int. Organ. Citrus Virol.Conf. Proc. (1957–2010) 1993, 12, 301–305. [Google Scholar]
- López, M.M.; LLop, P.; Olmos, A.; Marco-Noales, E.; Cambra, M.; Bertolini, E. Are molecular tools solving the challenges posed by detection of plant pathogenic bacteria and viruses? Curr. Issues Mol. Biol. 2008, 11, 13–46. [Google Scholar] [PubMed]
- Davis, M.J.; Purcell, A.H.; Thomson, S.V. Isolation media for the Pierce’s disease bacterium. Phytopathology 1980, 70, 425–429. [Google Scholar] [CrossRef]
- Ridé, M. Bactéries phytopathogènes et maladies bactériennes des végétaux; Ponsot: Paris, France, 1969. [Google Scholar]
- Vela, C.; Cambra, M.; Cortés, E.; Moreno, P.; Miguet, J.G.; Pérez de San Román, C.; Sanz, A. Production and characterization of monoclonal antibodies specific for citrus tristeza virus and their use for diagnosis. J. Gen. Virol. 1986, 67, 91–96. [Google Scholar] [CrossRef]
- EPPO. PM 7/101 (1): ELISA tests for pathogenic bacteria. Bull. OEPP/EPPO Bull. 2010, 40, 369–372. [Google Scholar] [CrossRef]
- European Union. Council Directive of 24 November 1986 on the approximation of laws, regulations and administrative provisions of the Member States regarding the protection of animals used for experimental and other scientific purposes. Off. J. Eur. Union 1986, L358, 1–28. [Google Scholar]
- IPPC-FAO. Diagnostic protocols for regulated pests: Citrus tristeza virus. Int. Stand. Phytosanit. Meas. ISPM27 2016, DP15, 1–21. [Google Scholar]
- Harper, S.J.; Ward, L.I.; Clover, G.R.G. Development of LAMP and Real-Time PCR methods for the rapid detection of Xylellafastidiosa for quarantine and Field Applications. Phytopathology 2010, 100, 1282–1288, Erratum in Phytopathology 2013, 103, 762. [Google Scholar] [CrossRef]
- Francis, M.; Lin, H.; Cabrera-La Rosa, J.; Doddapaneni, H.; Civerolo, E.L. Genome-based PCR for specific and sensitive detection and quantification of Xylella fastidiosa. Eur. J. Plant Pathol. 2006, 115, 203–213. [Google Scholar] [CrossRef]
- Olmos, A.; Capote, N.; Bertolini, E.; Cambra, M. Molecular diagnostic methods for plant viruses. In Biotechnology and Plant Disease Management; Punja, Z.K., De Boer, S.K., Sanfaçon, H., Eds.; CABI Press: Oxfordshire, UK, 2007; pp. 227–249. [Google Scholar]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Altman, D.G.; Machin, D.; Bryant, T.N.; Gardner, M.J. Statistics with Confidence, 2nd ed.; British Medical Journal: London, UK, 2000; pp. 116–118. [Google Scholar]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef]
- McNemar, Q. Note on the sampling error of the difference between correlated proportions or percentages. Psychometrika 1947, 12, 153–157. [Google Scholar] [CrossRef] [PubMed]
- Feinstein, A.R.; Cicchetti, D.V. High agreement but low kappa: I. The problems with two paradoxes. J. Clin. Epidemiol. 1990, 43, 543–549. [Google Scholar] [CrossRef]
- Marco-Noales, E.; Barbé, S.; Monterde, A.; Navarro-Herrero, I.; Ferrer, A.; Dalmau, V.; Aure, C.M.; Domingo-Calap, M.L.; Landa, B.B.; Roselló, M. Evidence that Xylella fastidiosa is the Causal Agent of Almond Leaf Scorch Disease in Alicante, Mainland Spain (Iberian Pen-insula). Plant Dis. 2021, 105, 3349–3352. [Google Scholar] [CrossRef]
- Denancé, N.; Legendre, B.; Briand, M.; Olivier, V.; de Boisseson, C.; Poliakoff, F.; Jacques, M.-A. Several subspecies and sequence types are associated with the emergence of Xylellafastidiosa in natural settings in France. Plant Pathol. 2017, 66, 1054–1064. [Google Scholar] [CrossRef]
- Minsavage, G.V.; Thompson, C.M.; Hopkins, D.L.; Leite, R.M.V.B.C.; Stall, R.E. Development of a polymerase chain reaction protocol for detection of Xylellafastidiosa in plant tissue. Phytopathology 1994, 84, 456–461. [Google Scholar] [CrossRef]
- Waliullah, S.; Hudson, O.; Oliver, J.E.; Brannen, P.M.; Ji, P.; Ali, M.E. Comparative analysis of different molecular and serological methods for detection of Xylellafastidiosa in blueberry. PLoS ONE 2019, 14, e0221903. [Google Scholar] [CrossRef]
- Loconsole, G.; Potere, O.; Boscia, D.; Altamura, G.; Djelouah, K.; Elbeaino, T.; Frasheri, D.; Lorusso, D.; Palmisano, F.; Pollastro, P.; et al. Detection of Xylellafastidiosa in olive trees by molecular and serological methods. J. Plant Pathol. 2014, 96, 7–14. [Google Scholar]
- Xiao, Y.; Isaacs, S.N. Enzyme-linked immunosorbent assay (ELISA) and blocking with bovine serum albumin (BSA)—Not all BSAs are alike. J. Immunol. Methods 2012, 384, 148–151. [Google Scholar] [CrossRef]
- Hilton, A.; Wang, X.; Zhang, M.; Cervantes, K.; French, J.; Randall, J.J.; Bock, C.H.; Grauke, L.J.; Jo, Y. Improved methods for detecting Xylella fastidiosa in pecan and related Carya species. Eur. J. Plant Pathol. 2020, 157, 899–918. [Google Scholar] [CrossRef]
- Chiriacò, M.S.; Luvisi, A.; Primiceri, E.; Sabella, E.; De Bellis, L.; Maruccio, G. Development of a lab-on-a-chip method for rapid assay of Xylellafastidiosa subsp. pauca strain CoDiRO. Sci. Rep. 2018, 8, 7376. [Google Scholar]
- Baldi, P.; La Porta, N. Xylellafastidiosa: Host range and advances in molecular identification techniques. Front. Plant Sci. 2017, 8, 944. [Google Scholar] [CrossRef]
| Strain | Host | Geographical Origin | Subspecies | ST |
|---|---|---|---|---|
| CoDiRo | Olive tree | Puglia (Italy) | pauca | 53 |
| Conn Creek | Grapevine | Napa (CA, USA) | fastidiosa | 1 |
| Fetzer | Grapevine | Napa (CA, USA) | fastidiosa | 4 |
| Stag’s Leap | Grapevine | Napa (CA, USA) | fastidiosa | - |
| Temecula | Grapevine | Temecula (CA, USA) | fastidiosa | 1 |
| LMG 17159 | Grapevine | Florida (USA) | fastidiosa | 2 |
| LMG 15099 | Almond tree | California (USA) | fastidiosa | - |
| IVIA 5235 | Cherry | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 5387 | Almond tree | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 5388 | Almond tree | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 5770 | Grapevine | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 5772 | Grapevine | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 5773 | Grapevine | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 6015 | Rhamnusalaternus | Balearic Islands (Spain) | fastidiosa | 1 |
| IVIA 6035 | Calicotomespinosa | Balearic Islands (Spain) | fastidiosa | 1 |
| CFBP 8430 | Polygala myrtifolia | PACA (France) | multiplex | 6 |
| IVIA 5908 | Almond tree | Valencian Community (Spain) | multiplex | 6 |
| IVIA 5946 | Almond tree | Valencian Community (Spain) | multiplex | 6 |
| IVIA 5947 | Almond tree | Valencian Community (Spain) | multiplex | 6 |
| CFBP 8072 | Coffeaarabica | Ecuador | pauca | - |
| CFBP 8419 | Coffeaarabica | Costa Rica | sandyi | - |
| Bacterial Species | Strain | Host | Geographical Origin |
|---|---|---|---|
| Agrobacterium tumefaciens | C58 | Cherry tree | USA |
| A. vitis | IVIA 339-26 | Vitis vinifera | Spain |
| Breneriaquercina | IVIA 2803-2 | Quercus sp. | Spain |
| Clavibactermichiganensis subsp. michiganensis | IVIA 5153 | Solanum lycopersicum | Spain |
| Erwiniaamylovora | CFBP 1430 | Crataegus | France |
| Xanthomonasarboricola pv. pruni | IVIA 3161-2 | Prunus dulcis | Spain |
| X. arboricola pv. pruni | CFBP 5229 | Prunus spp. | Argentina |
| X. arboricola pv. pruni | IVIA 3978-2 | Corylus avellana | Spain |
| X. arboricola pv. pruni | CFBP 5562 | P. persica | France |
| X. arboricola pv. pruni | DAR 56679 | P. armeniaca | Australia |
| X. arboricola pv. fragarie | CFBP 6771 | Fragaria sp. | Italy |
| X. arboricola pv. juglandis | RIPF XO4 | Juglans regia | Poland |
| X. arboricola pv. juglandis | IVIA 1317-1a | J. regia | Spain |
| X. axonopodis pv. phaseoli | CECT 914 | Phaseolus vulgaris | Hungary |
| X. campestris pv. campestris | IVIA 2734-2 | Brasica oleracea | Spain |
| Xanthomonas spp. | IVIA 3080 | Capsicum annuum | Spain |
| Pseudomonas syringae | IVIA 2627 | Pyrus comunis | Spain |
| P. syringae | IVIA 2141 | Olea sp. | Spain |
| Host | DAS-ELISA; MAb2G1/PPD | Real-Time PCR; Harper et al. [31] | Real-Time PCR; Francis et al. [32] |
|---|---|---|---|
| Almond tree | 150/165 | 165/165 | 163/165 |
| Olive tree | 40/40 | 40/40 | 40/40 |
| Prunus spp. | 7/7 | 7/7 | 7/7 |
| Citrus tree | 5/5 | 5/5 | 5/5 |
| Calicotome sp. | 3/3 | 3/3 | 3/3 |
| Fig tree | 3/3 | 3/3 | 3/3 |
| Rosemary | 3/3 | 3/3 | 3/3 |
| Grapevine | 2/2 | 2/2 | 2/2 |
| Polygala | 1/1 | 1/1 | 1/1 |
| Laurus sp. | 1/1 | 1/1 | 1/1 |
| Cistus sp. | 1/1 | 1/1 | 1/1 |
| Helychrisum sp. | 1/1 | 1/1 | 1/1 |
| Phagnalon sp. | 1/1 | 1/1 | 1/1 |
| 218/233 | 233/233 | 231/233 |
| Harper’s Real-Time PCR | ||||
|---|---|---|---|---|
| DAS-ELISA | Positive | Negative | Total | |
| Positive | 116 | 0 | 116 | |
| Negative | 15 | 102 | 117 | |
| Total | 131 | 102 | 233 | |
| Diagnostic parameters | Diagnostic sensitivity | 88.5% | ||
| Diagnostic specificity | 100% | |||
| Positive predictive value | 100% | |||
| Negative predictive value | 87.1% | |||
| False positive rate | - | |||
| False negative rate | 6.4% | |||
| Prevalence rate | 56.2% | |||
| Likelihood ratio for positive results | - | |||
| Likelihood ratio for negative results | 0.115 | |||
| Relative accuracy | 93.5% | |||
| Francis’ Real-Time PCR | ||||
|---|---|---|---|---|
| DAS-ELISA | Positive | Negative | Total | |
| Positive | 116 | 0 | 116 | |
| Negative | 13 | 104 | 117 | |
| Total | 129 | 104 | 233 | |
| Diagnostic parameters | Diagnostic sensitivity | 89.9% | ||
| Diagnostic specificity | 100% | |||
| Positive predictive value | 100% | |||
| Negative predictive value | 88.8% | |||
| False positive rate | - | |||
| False negative rate | 5.6% | |||
| Prevalence rate | 55.3% | |||
| Likelihood ratio for positive results | - | |||
| Likelihood ratio for negative results | 0.101 | |||
| Relative accuracy | 94.4% | |||
| MAb2G1/PPD DAS-ELISA vs. Harper’s Real-Time PCR | MAb2G1/PPD DAS-ELISA vs. Francis’ Real-Time PCR | |
|---|---|---|
| Agreement | 0.93 | 0.94 |
| Cohen’s Kappa (95% CI) | 0.87 (0.81–1.0) | 0.89 (0.81–1.0) |
| McNemar’s test; p-value | 12; p-value < 0.0005 | 10; p-value < 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gorris, M.T.; Sanz, A.; Peñalver, J.; López, M.M.; Colomer, M.; Marco-Noales, E. Detection and Diagnosis of Xylella fastidiosa by Specific Monoclonal Antibodies. Agronomy 2021, 11, 48. https://doi.org/10.3390/agronomy11010048
Gorris MT, Sanz A, Peñalver J, López MM, Colomer M, Marco-Noales E. Detection and Diagnosis of Xylella fastidiosa by Specific Monoclonal Antibodies. Agronomy. 2021; 11(1):48. https://doi.org/10.3390/agronomy11010048
Chicago/Turabian StyleGorris, María Teresa, Antonio Sanz, Javier Peñalver, María M. López, Mario Colomer, and Ester Marco-Noales. 2021. "Detection and Diagnosis of Xylella fastidiosa by Specific Monoclonal Antibodies" Agronomy 11, no. 1: 48. https://doi.org/10.3390/agronomy11010048
APA StyleGorris, M. T., Sanz, A., Peñalver, J., López, M. M., Colomer, M., & Marco-Noales, E. (2021). Detection and Diagnosis of Xylella fastidiosa by Specific Monoclonal Antibodies. Agronomy, 11(1), 48. https://doi.org/10.3390/agronomy11010048





