Metabolomics: A Tool for Cultivar Phenotyping and Investigation of Grain Crops
Abstract
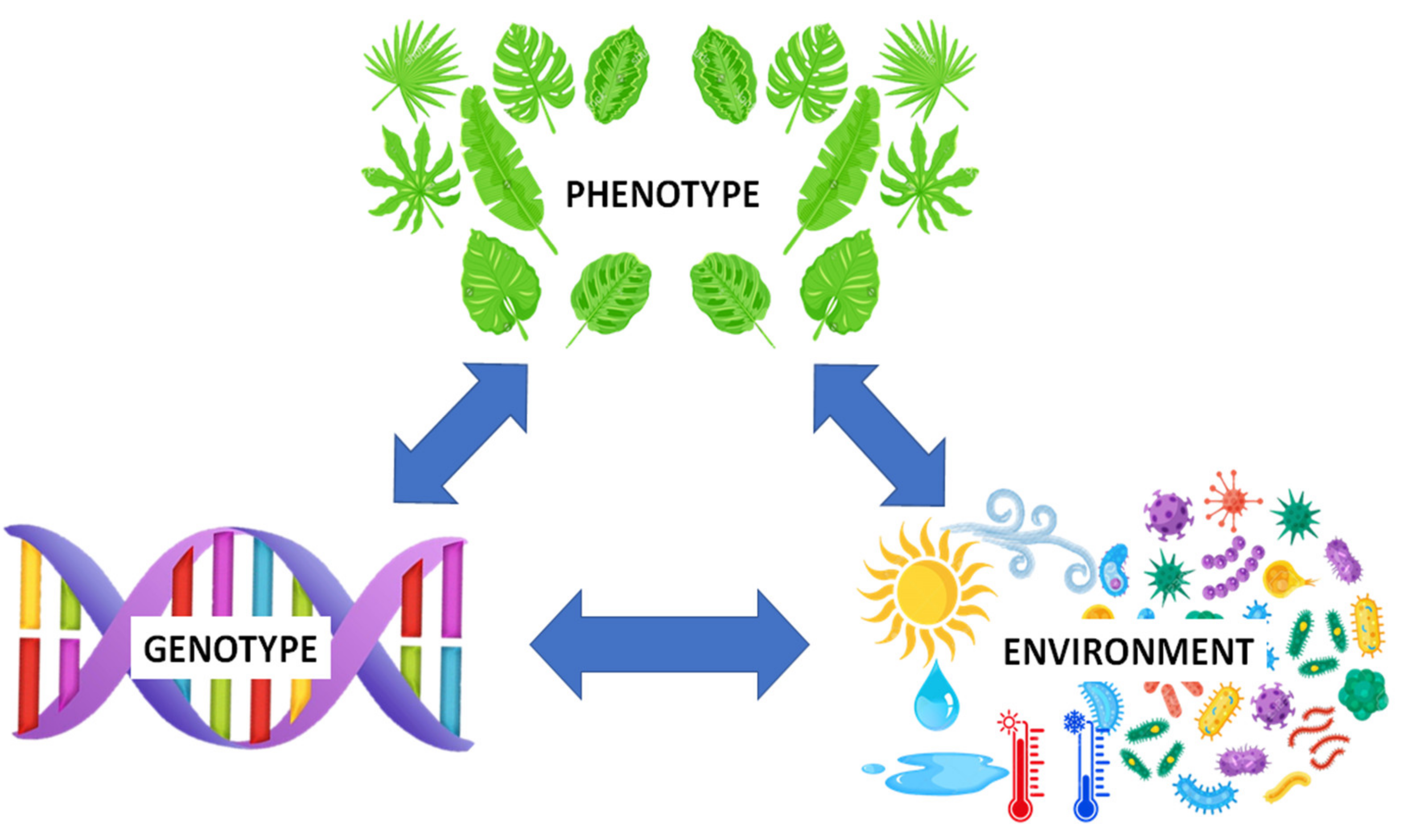
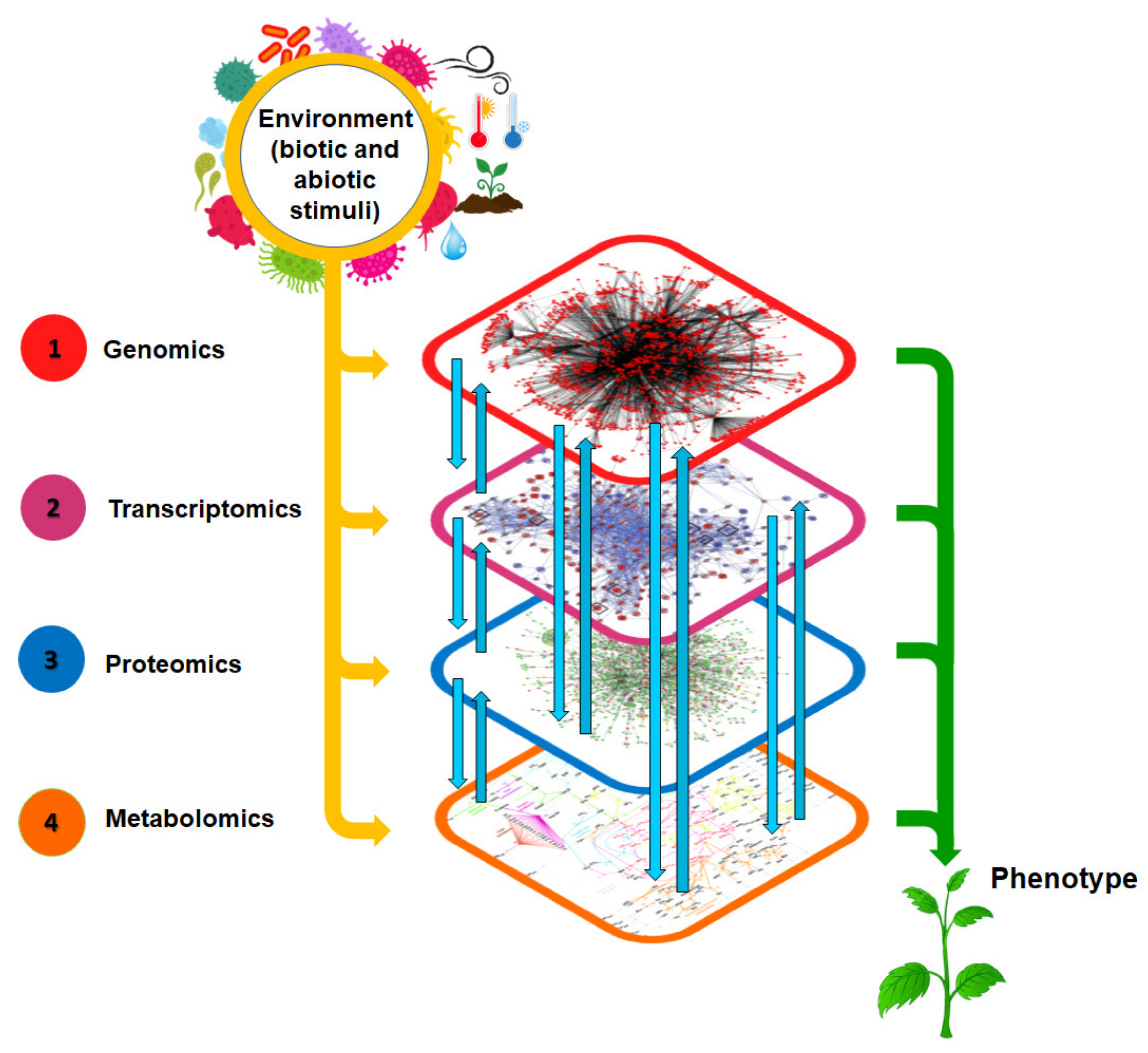
1. Introduction: The Genotype × Environment × Phenotype (G × E × P) Relationship
2. Plant Metabolomes as Responsive and Dynamic Entities
3. Potential of Metabolomics in Crop Science: Big Data, Big Expectations
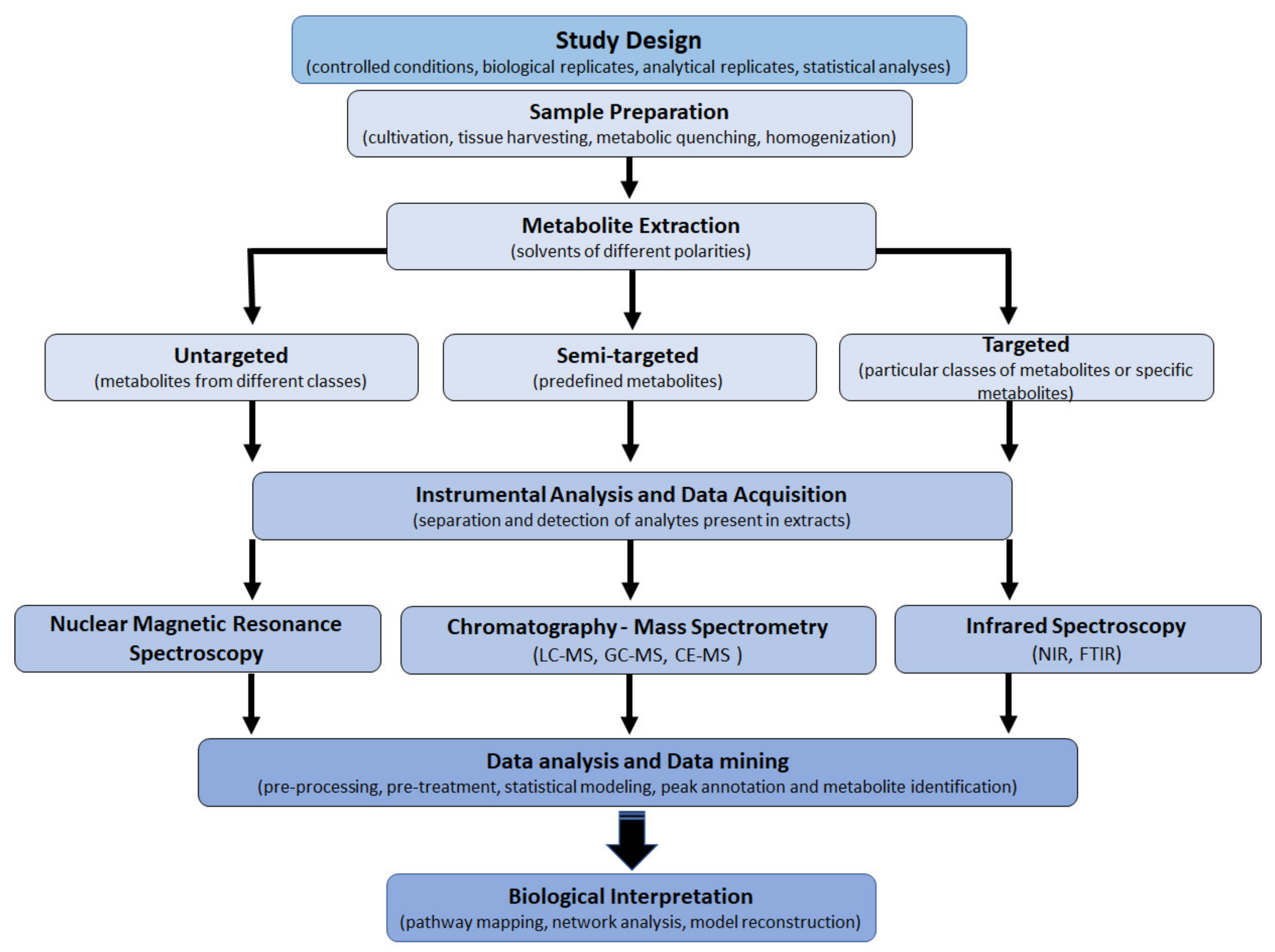
4. Experimental Designs, Workflows and Analytical Platforms Used in Plant Metabolomics
5. Handling and Mining of Metabolomic Data
6. Biological Interpretation: From Metabolite to Metabolic Pathways and Networks
7. Metabolomics Applied to Cultivar/Variety Identification and Cultivar-Specific Responses
7.1. Rice
7.2. Barley
7.3. Sorghum
7.4. Wheat
7.5. Maize
7.6. Oats
7.7. Millet
8. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gienapp, P.; Laine, V.N.; Mateman, A.C.; van Oers, K.; Visser, M.E. Environment-dependent genotype-phenotype associations in avian breeding time. Front. Genet. 2017, 8, 102. [Google Scholar] [CrossRef]
- Aslam, M.M.; Karanja, J.K. Genotype by environment interactions modulate sugarcane response to mechanical wounding stress. Physiol. Mol. Plant Pathol. 2020, 109, 101443. [Google Scholar] [CrossRef]
- Lande, R. Adaptation to an extraordinary environment by evolution of phenotypic plasticity and genetic assimilation. J. Evol. Biol. 2009, 22, 1435–1446. [Google Scholar] [CrossRef] [PubMed]
- Kusmec, A.; de Leon, N.; Schnable, P.S. Harnessing phenotypic plasticity to improve maize yields. Front. Plant Sci. 2018, 9, 1377. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Guo, T.; Mu, Q.; Li, X.; Yu, J. Genomic and environmental determinants and their interplay underlying phenotypic plasticity. Proc. Natl. Acad. Sci. USA 2018, 115, 6679–6684. [Google Scholar] [CrossRef] [PubMed]
- Weckwerth, W. Green systems biology—From single genomes, proteomes and metabolomes to ecosystems research and biotechnology. J. Proteom. 2011, 75, 284–305. [Google Scholar] [CrossRef]
- Nägler, M.; Nägele, T.; Gilli, C.; Fragner, L.; Korte, A.; Platzer, A.; Farlow, A.; Nordborg, M.; Weckwerth, W. Eco-metabolomics and metabolic modeling: Making the leap from model systems in the lab to native populations in the field. Front. Plant Sci. 2018, 9, 1556. [Google Scholar] [CrossRef]
- Liu, X.; Locasale, J.W. Metabolomics: A primer. Trends Biochem. Sci. 2017, 42, 274–284. [Google Scholar] [CrossRef]
- Mhlongo, M.I.; Piater, L.A.; Madala, N.E.; Steenkamp, P.A.; Dubery, I.A. Phenylpropanoid defences in Nicotiana tabacum cells: Overlapping metabolomes indicate common aspects to responses induced by lipopolysaccharides, chitosan and flagellin-22. PLoS ONE 2016, 11, e0151350. [Google Scholar] [CrossRef]
- Zeiss, D.; Mhlongo, M.; Tugizimana, F.; Steenkamp, P.; Dubery, I. Comparative metabolic phenotyping of tomato (Solanum lycopersicum) for the identification of metabolic signatures in cultivars differing in resistance to Ralstonia solanacearum. Int. J. Mol. Sci. 2018, 19, 2558. [Google Scholar] [CrossRef]
- Finnegan, T.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. The lipopolysaccharide-induced metabolome signature in Arabidopsis thaliana reveals dynamic reprogramming of phytoalexin and phytoanticipin pathways. PLoS ONE 2016, 11, e0163572. [Google Scholar] [CrossRef] [PubMed]
- Tugizimana, F.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. Multi-platform metabolomic analyses of ergosterol-induced dynamic changes in Nicotiana tabacum cells. PLoS ONE 2014, 9, e87846. [Google Scholar] [CrossRef] [PubMed]
- Panda, A.; Parida, A.K.; Rangani, J. Advancement of metabolomics techniques and their applications in plant science: Current scenario and future prospective. In Plant Metabolites and Regulation under Environmental Stress; Ahmad, P., Ahanger, M.A., Singh, V.P., Tripathi, D.K., Alam, P., Alyemeni, M.N., Eds.; Academic Press: London, UK, 2018; pp. 1–36. [Google Scholar]
- Tiwari, R.; Rana, C.S. Plant secondary metabolites: A review. Int. J. Eng. Res. Gen. Sic. 2015, 3, 661–670. [Google Scholar]
- Aggio, R.B.; Ruggiero, K.; Villas-Bôas, S.G. Pathway activity profiling (PAPi): From the metabolite profile to the metabolic pathway activity. Bioinformatics 2010, 26, 2969–2976. [Google Scholar] [CrossRef]
- Pandey, P.; Irulappan, V.; Bagavathiannan, M.V.; Senthil-Kumar, M. Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front. Plant Sci. 2017, 8, 537. [Google Scholar] [CrossRef]
- Bebber, D.P.; Gurr, S.J. Crop-destroying fungal and oomycete pathogens challenge food security. Fungal Genet. Biol. 2015, 74, 62–64. [Google Scholar] [CrossRef]
- Ramegowda, V.; Senthil-Kumar, M. The interactive effects of simultaneous biotic and abiotic stresses on plants: Mechanistic understanding from drought and pathogen combination. J. Plant Physiol. 2015, 176, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Kharub, A.S.; Singh, J.; Lal, C.; Kumar, V. Abiotic stress tolerance in barley. In Abiotic Stress Management for Resilient Agriculture; Minhas, P.S., Rane, J., Pasala, R.K., Eds.; Springer: Berlin/Heidelberg, Germany, 2017; pp. 363–374. [Google Scholar]
- Tugizimana, F.; Piater, L.; Dubery, I. Plant metabolomics: A new frontier in phytochemical analysis. S. Afr. J. Sci. 2013, 109, 1–11. [Google Scholar] [CrossRef]
- Djami-Tchatchou, A.T.; Ncube, E.N.; Steenkamp, P.A.; Dubery, I.A. Similar, but different: Structurally related azelaic acid and hexanoic acid trigger differential metabolomic and transcriptomic responses in tobacco cells. BMC Plant Biol. 2017, 17, 227. [Google Scholar] [CrossRef]
- Gorrochategui, E.; Jaumot, J.; Lacorte, S.; Tauler, R. Data analysis strategies for targeted and untargeted LC-MS metabolomic studies: Overview and workflow. Trends Anal. Chem. 2016, 82, 425–442. [Google Scholar] [CrossRef]
- Manzoni, C.; Kia, D.A.; Vandrovcova, J.; Hardy, J.; Wood, N.W.; Lewis, P.A.; Ferrari, R. Genome, transcriptome and proteome: The rise of omics data and their integration in biomedical sciences. Brief. Bioinform. 2016, 19, 286–302. [Google Scholar] [CrossRef] [PubMed]
- Winter, G.; Krömer, J.O. Fluxomics-connecting ‘omics analysis and phenotypes. Environ. Microbiol. 2013, 15, 1901–1916. [Google Scholar] [CrossRef] [PubMed]
- Ghatak, A.; Chaturvedi, P.; Weckwerth, W. Metabolomics in plant stress physiology. In Plant Genetics and Molecular Biology; Varshney, R.K., Pandey, M.K., Chitikineni, A., Eds.; Springer: Berlin/Heidelberg, Germany, 2018; pp. 187–236. [Google Scholar]
- Kopka, J.; Fernie, A.; Weckwerth, W.; Gibon, Y.; Stitt, M. Metabolite profiling in plant biology: Platforms and destinations. Genome Biol. 2004, 5, 109. [Google Scholar] [CrossRef] [PubMed]
- Ellis, D.I.; Dunn, W.B.; Griffin, J.L.; Allwood, J.W.; Goodacre, R. Metabolic fingerprinting as a diagnostic tool. Pharmacogenomics 2007, 8, 1243–1266. [Google Scholar] [CrossRef]
- Llanes, A.; Andrade, A.; Alemano, S.; Luna, V. Metabolomic approach to understand plant adaptations to water and salt stress. In Plant Metabolites and Regulation under Environmental Stress; Ahmad, P., Ahanger, M.A., Singh, V.P., Tripathi, D.K., Alam, P., Alyemeni, M.N., Eds.; Academic Press: London, UK, 2018; pp. 133–144. [Google Scholar]
- Wolfender, J.L.; Marti, G.; Thomas, A.; Bertrand, S. Current approaches and challenges for the metabolite profiling of complex natural extracts. J. Chromatogr. A 2015, 1382, 136–164. [Google Scholar] [CrossRef]
- Cambiaghi, A.; Ferrario, M.; Masseroli, M. Analysis of metabolomic data: Tools, current strategies and future challenges for omics data integration. Brief. Bioinform. 2017, 18, 498–510. [Google Scholar] [CrossRef]
- Ribbenstedt, A.; Ziarrusta, H.; Benskin, J.P. Development, characterization and comparisons of targeted and non-targeted metabolomics methods. PLoS ONE 2018, 13, 0207082. [Google Scholar] [CrossRef]
- Cook, D.W.; Rutan, S.C. Chemometrics for the analysis of chromatographic data in metabolomics investigations. J. Chemom. 2014, 28, 681–687. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, S.; Cui, X.; Pan, C.; Zhang, A.; Chen, F. A review of sample preparation methods for the pesticide residue analysis in foods. Cent. Eur. J. Chem. 2012, 10, 900–925. [Google Scholar] [CrossRef]
- Mushtaq, M.Y.; Choi, Y.H.; Verpoorte, R.; Wilson, E.G. Extraction for metabolomics: Access to the metabolome. Phytochem. Anal. 2014, 25, 291–306. [Google Scholar] [CrossRef]
- Moco, S.; Vervoort, J.; Bino, R.J.; De Vos, R.C.; Bino, R. Metabolomics technologies and metabolite identification. Trends Anal. Chem. 2007, 26, 855–866. [Google Scholar] [CrossRef]
- Altemimi, A.; Lakhssassi, N.; Baharlouei, A.; Watson, D.; Lightfoot, D. Phytochemicals: Extraction, isolation, and identification of bioactive compounds from plant extracts. Plants 2017, 6, 42. [Google Scholar] [CrossRef] [PubMed]
- Khoza, B.S.; Chimuka, L.; Mukwevho, E.; Steenkamp, P.A.; Madala, N.E. The effect of temperature on pressurised hot water extraction of pharmacologically important metabolites as analysed by UPLC-qTOF-MS and PCA. Evid. Based Complement. Altern. Med. 2014, 1–9. [Google Scholar] [CrossRef]
- Hamany Djande, C.Y.; Piater, L.A.; Steenkamp, P.A.; Madala, N.E.; Dubery, I.A. Differential extraction of phytochemicals from the multipurpose tree, Moringa oleifera, using green extraction solvents. S. Afr. J. Bot. 2018, 115, 81–89. [Google Scholar] [CrossRef]
- Emwas, A.-H.; Alghrably, M.; Al-Harthi, S.; Gabriel Poulson, B.; Szczepski, K.; Chandra, K.; Jaremko, M. New Advances in Fast Methods of 2D NMR Experiments; IntechOpen Press: London, UK, 2019. [Google Scholar]
- Junot, C.; Fenaille, F. Metabolomics using Fourier transform mass spectrometry. In Fundamentals and Applications of Fourier Transform Mass Spectrometry; Schmitt-Kopplin, P., Kanawati, B., Eds.; Elsevier: Amsterdam, The Netherlands, 2019; pp. 325–356. [Google Scholar]
- Wishart, D.S. NMR metabolomics: A look ahead. J. Magn. Reson. 2019, 306, 155–161. [Google Scholar] [CrossRef]
- Gouilleux, B.; Rouger, L.; Giraudeau, P. Ultrafast 2D NMR: Methods and Applications. Annu. Rep. NMR Spectrosc. 2018, 93, 75–144. [Google Scholar]
- Fortier-McGill, B.E.; Dutta Majumdar, R.; Lam, L.; Soong, R.; Liaghati-Mobarhan, Y.; Sutrisno, A.; de Visser, R.; Simpson, M.J.; Wheeler, H.L.; Campbell, M.; et al. Comprehensive multiphase (CMP) NMR monitoring of the structural changes and molecular flux within a growing seed. J. Agric. Food Chem. 2017, 65, 6779–6788. [Google Scholar] [CrossRef]
- Bastawrous, M.; Jenne, A.; Tabatabaei Anaraki, M.; Simpson, A.J. In-vivo NMR spectroscopy: A powerful and complimentary tool for understanding environmental toxicity. Metabolites 2018, 8, 35. [Google Scholar] [CrossRef]
- Kapoore, R.V.; Vaidyanathan, S. Towards quantitative mass spectrometry-based metabolomics in microbial and mammalian systems. Philos. Trans. Royal Soc. A 2016, 374, 20150363. [Google Scholar] [CrossRef]
- Paglia, G.; Astarita, G. Metabolomics and lipidomics using traveling-wave ion mobility mass spectrometry. Nat. Protoc. 2017, 12, 797. [Google Scholar] [CrossRef]
- Stoll, D.R.; Harmes, D.C.; Staples, G.O.; Potter, O.G.; Dammann, C.T.; Guillarme, D.; Beck, A. Development of comprehensive online two-dimensional liquid chromatography / mass spectrometry using hydrophilic interaction and reversed-phase separations for rapid and deep profiling of therapeutic antibodies. Anal. Chem. 2018, 90, 5923–5929. [Google Scholar] [CrossRef] [PubMed]
- Walker, L.R.; Hoyt, D.W.; Walker, S.M.; Ward, J.K.; Nicora, C.D.; Bingol, K. Unambiguous metabolite identification in high-throughput metabolomics by hybrid 1D 1H NMR/ESI MS1 approach. Magn. Reson. Chem. 2016, 54, 998–1003. [Google Scholar] [CrossRef] [PubMed]
- Aksenov, A.A.; da Silva, R.; Knight, R.; Lopes, N.P.; Dorrestein, P.C. Global chemical analysis of biology by mass spectrometry. Nat. Rev. Chem. 2017, 1, 54. [Google Scholar] [CrossRef]
- Saccenti, E.; Hoefsloot, H.C.; Smilde, A.K.; Westerhuis, J.A.; Hendriks, M.M. Reflections on univariate and multivariate analysis of metabolomics data. Metabolomics 2014, 10, 361–374. [Google Scholar] [CrossRef]
- Vargason, T.; Howsmon, D.; McGuinness, D.; Hahn, J. On the use of multivariate methods for analysis of data from biological networks. Processes 2017, 5, 36. [Google Scholar] [CrossRef] [PubMed]
- Lamichhane, S.; Sen, P.; Dickens, A.M.; Hyötyläinen, T.; Orešiè, M. An overview of metabolomics data analysis: Current tools and future perspectives. Comp. Anal. Chem. 2018, 82, 387–413. [Google Scholar]
- Madala, N.E.; Piater, L.A.; Steenkamp, P.A.; Dubery, I.A. Multivariate statistical models of metabolomic data reveals different metabolite distribution patterns in isonitrosoacetophenone-elicited Nicotiana tabacum and Sorghum bicolor cells. SpringerPlus 2014, 3, 254. [Google Scholar] [CrossRef]
- Schillemans, T.; Shi, L.; Liu, X.; Åkesson, A.; Landberg, R.; Brunius, C. Visualization and interpretation of multivariate associations with disease risk markers and disease risk—The Triplot. Metabolites 2019, 9, 133. [Google Scholar] [CrossRef]
- Cuperlovic-Culf, M. Machine learning methods for analysis of metabolic data and metabolic pathway modeling. Metabolites 2018, 8, 4. [Google Scholar] [CrossRef]
- Sathya, R.; Abraham, A. Comparison of supervised and unsupervised learning algorithms for pattern classification. Int. J. Adv. Res. Artif. Intell. 2013, 2, 34–38. [Google Scholar] [CrossRef]
- Maione, C.; Nelson, D.R.; Barbosa, R.M. Research on social data by means of cluster analysis. Appl. Comp. Inform. 2019, 15, 153–162. [Google Scholar] [CrossRef]
- Bunte, K.; Biehl, M.; Hammer, B. A general framework for dimensionality-reducing data visualization mapping. Neural Comput. 2012, 24, 771–804. [Google Scholar] [CrossRef]
- Capó, M.; Pérez, A.; Lozano, J.A. An efficient approximation to the K-means clustering for massive data. Knowl. Based Syst. 2017, 117, 56–69. [Google Scholar] [CrossRef]
- Wanichthanarak, K.; Fan, S.; Grapov, D.; Barupal, D.K.; Fiehn, O. Metabox: A toolbox for metabolomic data analysis, interpretation and integrative exploration. PLoS ONE 2017, 12, 0171046. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S. Computational approaches to metabolomics. Methods Mol. Biol. 2010, 593, 283–313. [Google Scholar]
- Sumner, L.W.; Amberg, A.; Barrett, D.A.; Beale, M.; Beger, R.; Daykin, C.; Fan, T.; Fiehn, O.; Doodacre, R.; Griffin, J.; et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef]
- MetaboAnalyst. Available online: https://www.metaboanalyst.ca/ (accessed on 4 June 2020).
- Metabox OmicsX. Available online: https://omictools.com/metabox-tool (accessed on 4 June 2020).
- MetaCore OmicsX. Available online: https://omictools.com/metabox-tool (accessed on 4 June 2020).
- InCroMAP. Available online: http://www.ra.cs.uni-tuebingen.de/software/InCroMAP/index.htm (accessed on 4 June 2020).
- SIMCA. Available online: https://umetrics.com/ (accessed on 4 June 2020).
- XCMS. Available online: https://xcmsonline.scripps.edu (accessed on 4 June 2020).
- Wolfender, J.L.; Rudaz, S.; Hae Choi, Y.; Kyong Kim, H. Plant metabolomics: From holistic data to relevant biomarkers. Curr. Med. Chem. 2013, 20, 1056–1090. [Google Scholar]
- Klassen, A.; Faccio, A.T.; Canuto, G.A.B.; da Cruz, P.L.R.; Ribeiro, H.C.; Tavares, M.F.M.; Sussulini, A. Metabolomics: Definitions and significance in systems biology. In Metabolomics: From Fundamentals to Clinical Applications; Sussulini, A., Ed.; Springer: Berlin/Heidelberg, Germany, 2017; pp. 31–37. [Google Scholar]
- Pubchem. Available online: https://pubchem.ncbi.nlm.nih.gov/ (accessed on 4 June 2020).
- MassBank. Available online: https://massbank.eu/MassBank/Search (accessed on 4 June 2020).
- Metlin. Available online: https://metlin.scripps.edu/ (accessed on 4 June 2020).
- KEGG. Available online: https://www.genome.jp/kegg/ (accessed on 4 June 2020).
- ChEBI. Available online: https://www.ebi.ac.uk/chebi/ (accessed on 4 June 2020).
- MetaboID. Available online: https://www.ebi.ac.uk/chebi/ (accessed on 4 June 2020).
- Cottret, L.; Frainay, C.; Chazalviel, M.; Cabanettes, F.; Gloaguen, Y.; Camenen, E.; Merlet, B.; Heux, S.; Portais, J.C.; Poupin, N.; et al. MetExplore: Collaborative edition and exploration of metabolic networks. Nucleic Acids Res. 2018, 46, 495–502. [Google Scholar] [CrossRef]
- Chong, J.; Wishart, D.S.; Xia, J. Using Metaboanalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr. Protoc. Bioinform. 2019, 68, e86. [Google Scholar] [CrossRef]
- Xia, J. Computational strategies for biological interpretation of metabolomics data. Adv. Exp. Med. Biol. 2017, 965, 191–206. [Google Scholar] [PubMed]
- Razzaq, A.; Sadia, B.; Raza, A.; Khalid Hameed, M.; Saleem, F. Metabolomics: A way forward for crop improvement. Metabolites 2019, 9, 303. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.; Yang, L.; Zhang, D.; Shi, J. Plant metabolomics: An indispensable system biology tool for plant science. Int. J. Mol. Sci. 2016, 17, 767. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, A. Plant Breeding: A prospect in developing world. EC Microbiol. 2017, 8, 272–278. [Google Scholar]
- McCouch, S. Diversifying selection in plant breeding. PLoS Biol. 2004, 2, 347. [Google Scholar] [CrossRef]
- Duvick, D.N. Utilization of biotechnology in plant breeding for North America: A status report. In Perspektiven Nachwachsender Rohstoffe in der Chemie; Eierdanz, H., Ed.; Wiley-VCH: Weinheim, Germany, 1996; pp. 3–10. [Google Scholar] [CrossRef]
- Pilacinski, W.; Crawford, A.; Downey, R.; Harvey, B.; Huber, S.; Hunst, P.; Lahman, L.K.; MacIntosh, S.; Pohl, M.; Rickard, C.; et al. Plants with genetically modified events combined by conventional breeding: An assessment of the need for additional regulatory data. Food Chem. Toxicol. 2011, 49, 1–7. [Google Scholar] [CrossRef][Green Version]
- Lusser, M.; Parisi, C.; Plan, D.; Rodriguez-Cerezo, E. Deployment of new biotechnologies in plant breeding. Nat. Biotech. 2012, 30, 231. [Google Scholar] [CrossRef] [PubMed]
- Simó, C.; Ibáez, C.; Valdés, A.; Cifuentes, A.; García-Cañas, V. Metabolomics of genetically modified crops. Int. J. Mol. Sci. 2014, 15, 18941–18966. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.; Sarma, S.; Bohra, A.; Mitra, A.; Sharma, N.K.; Kumar, A. Plant metabolomics: An emerging technology for crop improvement. In New Visions in Plant Science, 1st ed.; Çelik, O., Ed.; IntechOpen: London, UK, 2018; pp. 65–79. [Google Scholar]
- Zhang, H.; Mittal, N.; Leamy, L.J.; Barazani, O.; Song, B.H. Back into the wild—Apply untapped genetic diversity of wild relatives for crop improvement. Evol. Appl. 2017, 10, 5–24. [Google Scholar] [CrossRef]
- Mammadov, J.; Buyyarapu, R.; Guttikonda, S.K.; Parliament, K.; Abdurakhmonov, I.Y.; Kumpatla, S.P. Wild relatives of maize, rice, cotton, and soybean: Treasure troves for tolerance to biotic and abiotic stresses. Front. Plant Sci. 2018, 9, 886. [Google Scholar] [CrossRef]
- Dempewolf, H.; Baute, G.; Anderson, J.; Kilian, B.; Smith, C.; Guarino, L. Past and future use of wild relatives in crop breeding. Crop Sci. 2017, 57, 1070–1082. [Google Scholar] [CrossRef]
- Xu, Y.; Crouch, J.H. Marker-assisted selection in plant breeding: From publications to practice. Crop Sci. 2008, 48, 391–407. [Google Scholar] [CrossRef]
- Korir, N.K.; Han, J.; Shangguan, L.; Wang, C.; Kayesh, E.; Zhang, Y.; Fang, J. Plant variety and cultivar identification: Advances and prospects. Crit. Rev. Biotechnol. 2013, 33, 111–125. [Google Scholar] [CrossRef] [PubMed]
- Roychowdhury, R.; Taoutaou, A.; Hakeem, K.R.; Gawwad, M.R.A.; Tah, J. Molecular marker-assisted technologies for crop improvement. In Crop Improvement in the Era of Climate Change; Roychowdhury, R., Ed.; International Publishing House: New Delhi, India, 2013; pp. 241–258. [Google Scholar]
- Prohens, J. Plant breeding: A success story to be continued thanks to the advances in genomics. Front. Plant Sci. 2011, 2, 51. [Google Scholar] [CrossRef] [PubMed]
- Cobb, J.N.; Juma, R.U.; Biswas, P.S.; Arbelaez, J.D.; Rutkoski, J.; Atlin, G.; Hagen, T.; Quinn, M.; Ng, E.H. Enhancing the rate of genetic gain in public-sector plant breeding programs: Lessons from the breeder’s equation. Theor. Appl. Genet. 2019, 132, 627–645. [Google Scholar] [CrossRef] [PubMed]
- Hall, R.D. Plant metabolomics: From holistic hope, to hype, to hot topic. New Phytol. 2006, 169, 453–468. [Google Scholar] [CrossRef]
- Fernie, A.R.; Schauer, N. Metabolomics-assisted breeding: A viable option for crop improvement? Trends Genet. 2009, 25, 39–48. [Google Scholar] [CrossRef]
- Kumar, R.; Bohra, A.; Pandey, A.K.; Pandey, M.K.; Kumar, A. Metabolomics for plant improvement: Status and prospects. Front. Plant Sci. 2017, 8, 302. [Google Scholar] [CrossRef]
- Alseekh, S.; Bermudez, L.; De Haro, L.A.; Fernie, A.R.; Carrari, F. Crop metabolomics: From diagnostics to assisted breeding. Metabolomics 2018, 14, 148. [Google Scholar] [CrossRef]
- Zabotina, O.A. Metabolite-based biomarkers for plant genetics and breeding. In Diagnostics in Plant Breeding; Lübberstedt, T., Varshney, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 281–309. [Google Scholar]
- Gamboa-Becerra, R.; Hernández-Hernández, M.C.; González-Ríos, Ó.; Suárez-Quiroz, M.L.; Gálvez-Ponce, E.; Ordaz-Ortiz, J.J.; Winkler, R. Metabolomic markers for the early selection of Coffea canephora plants with desirable cup quality traits. Metabolites 2019, 9, 214. [Google Scholar] [CrossRef] [PubMed]
- Riedelsheimer, C.; Czedik-Eysenberg, A.; Grieder, C.; Lisec, J.; Technow, F.; Sulpice, R.; Altmann, T.; Stitt, M.; Willmitzer, L.; Melchinger, A.E. Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat. Genet. 2012, 44, 217–220. [Google Scholar] [CrossRef]
- Li, S.; Dong, X.; Fan, G.; Yang, Q.; Shi, J.; Wei, W.; Zhao, F.; Li, N.; Wang, X.; Wang, F.; et al. Comprehensive profiling and inheritance patterns of metabolites in foxtail millet. Front. Plant Sci. 2018, 9, 1716. [Google Scholar] [CrossRef] [PubMed]
- Cubillos, A.E.R.; Tong, H.; Alseekh, S.; Lima, F.D.A.; Yu, J.; Fernie, A.R.; Nikoloski, Z.; Laitinen, R.A. Inheritance patterns in metabolism and growth in diallel crosses of Arabidopsis thaliana from a single growth habitat. Heredity 2018, 120, 463–473. [Google Scholar] [CrossRef]
- Lisec, J.; Römisch-Margl, L.; Nikoloski, Z.; Piepho, H.P.; Giavalisco, P.; Selbig, J.; Gierl, A.; Willmitzer, L. Corn hybrids display lower metabolite variability and complex metabolite inheritance patterns. Plant J. 2011, 68, 326–336. [Google Scholar] [CrossRef] [PubMed]
- Steinfath, M.; Strehmel, N.; Peters, R.; Schauer, N.; Groth, D.; Hummel, J.; Steup, M.; Selbig, J.; Kopka, J.; Geigenberger, P.; et al. Discovering plant metabolic biomarkers for phenotype prediction using an untargeted approach. Plant Biotechnol. J. 2010, 8, 900–911. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, O.; Urrutia, M.; Bernillon, S.; Giauffret, C.; Tardieu, F.; Le Gouis, J.; Langlade, N.; Charcosset, A.; Moing, A.; Gibon, Y. Fortune telling: Metabolic markers of plant performance. Metabolomics 2016, 12, 158. [Google Scholar] [CrossRef]
- Chen, M.; Rao, R.S.P.; Zhang, Y.; Zhong, C.; Thelen, J.J. Metabolite variation in hybrid corn grain from a large-scale multisite study. Crop J. 2016, 4, 177–187. [Google Scholar] [CrossRef]
- Loskutov, I.G.; Shelenga, T.V.; Konarev, A.V.; Shavarda, A.L.; Blinova, E.V.; Dzubenko, N.I. The metabolomic approach to the comparative analysis of wild and cultivated species of oats (Avena L.). Russ. J. Genet. Appl. Res. 2017, 7, 501–508. [Google Scholar] [CrossRef]
- Song, E.H.; Jeong, J.; Park, C.Y.; Kim, H.Y.; Kim, E.H.; Bang, E.; Hong, Y.S. Metabotyping of rice (Oryza sativa L.) for understanding its intrinsic physiology and potential eating quality. Food Res. Int. 2018, 111, 20–30. [Google Scholar] [CrossRef]
- Thomason, K.; Babar, M.A.; Erickson, J.E.; Mulvaney, M.; Beecher, C.; MacDonald, G. Comparative physiological and metabolomics analysis of wheat (Triticum aestivum L.) following post-anthesis heat stress. PLoS ONE 2018, 13, 0197919. [Google Scholar] [CrossRef]
- Lawas, L.M.F.; Li, X.; Erban, A.; Kopka, J.; Jagadish, S.K.; Zuther, E.; Hincha, D.K. Metabolic responses of rice cultivars with different tolerance to combined drought and heat stress under field conditions. Gigascience 2019, 8, 50. [Google Scholar] [CrossRef]
- Mareya, C.R.; Tugizimana, F.; Piater, L.A.; Madala, N.E.; Steenkamp, P.A.; Dubery, I.A. Untargeted metabolomics reveal defensome-related metabolic reprogramming in Sorghum bicolor against infection by Burkholderia andropogonis. Metabolites 2019, 9, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zeng, X.; Xu, Q.; Mei, X.; Yuan, H.; Jiabu, D.; Sang, Z.; Nyima, T. Metabolite profiling in two contrasting Tibetan hulless barley cultivars revealed the core salt-responsive metabolome and key salt-tolerance biomarkers. AoB Plants 2019, 11, 21. [Google Scholar] [CrossRef] [PubMed]
- Gani, A.; Wani, S.M.; Masoodi, F.A.; Hameed, G. Whole-grain cereal bioactive compounds and their health benefits: A review. Int. J. Food Process Technol. 2012, 3, 146–156. [Google Scholar] [CrossRef]
- Cooper, R. Re-discovering ancient wheat varieties as functional foods. J. Tradit. Compl. Med. 2015, 5, 138–143. [Google Scholar] [CrossRef]
- Khlestkina, E. The adaptive role of flavonoids: Emphasis on cereals. Cereal Res. Commun. 2013, 41, 185–198. [Google Scholar] [CrossRef]
- Dykes, L.; Rooney, L.W. Phenolic compounds in cereal grains and their health benefits. Cereal Foods World 2007, 52, 105–111. [Google Scholar] [CrossRef]
- Joye, I. Protein digestibility of cereal products. Foods 2019, 8, 199. [Google Scholar] [CrossRef]
- Xu, M.; Rao, J.; Chen, B. Phenolic compounds in germinated cereal and pulse seeds: Classification, transformation, and metabolic process. Crit. Rev. Food Sci. Nutr. 2020, 60, 740–759. [Google Scholar] [CrossRef]
- Sidhu, J.S.; Kabir, Y.; Huffman, F.G. Functional foods from cereal grains. Int. J. Food Prop. 2007, 10, 231–244. [Google Scholar] [CrossRef]
- Leitzmann, C. Characteristics and health benefits of phytochemicals. J. Compl. Med. Res. 2016, 23, 69–74. [Google Scholar] [CrossRef]
- Berini, J.L.; Brockman, S.A.; Hegeman, A.D.; Reich, P.B.; Muthukrishnan, R.; Montgomery, R.A.; Forester, J.D. Combinations of abiotic factors differentially alter production of plant secondary metabolites in five woody plant species in the boreal-temperate transition zone. Front. Plant Sci. 2018, 9, 1257. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Huang, W.; Gao, J.; Fu, H.; Liu, J. Comparative metabolomic analysis of seed metabolites associated with seed storability in rice (Oryza sativa L.) during natural aging. Plant Physiol. Biochem. 2018, 127, 590–598. [Google Scholar] [CrossRef] [PubMed]
- Zarei, I.; Luna, E.; Leach, J.; McClung, A.; Vilchez, S.; Koita, O.; Ryan, E. Comparative rice bran metabolomics across diverse cultivars and functional rice gene-bran metabolite relationships. Metabolites 2018, 8, 63. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Shi, J.; Quan, S.; Cui, B.; Kleessen, S.; Nikoloski, Z.; Tohge, T.; Alexander, D.; Guo, L.; Lin, H.; et al. Metabolic variation between japonica and indica rice cultivars as revealed by non-targeted metabolomics. Sci. Rep. 2014, 4, 5067. [Google Scholar] [CrossRef]
- Gayen, D.; Ghosh, S.; Paul, S.; Sarkar, S.N.; Datta, S.K.; Datta, K. Metabolic regulation of carotenoid-enriched golden rice line. Front. Plant. Sci. 2016, 7, 1622. [Google Scholar] [CrossRef]
- Sana, T.R.; Fischer, S.; Wohlgemuth, G.; Katrekar, A.; Jung, K.H.; Ronald, P.C.; Fiehn, O. Metabolomic and transcriptomic analysis of the rice response to the bacterial blight pathogen Xanthomonas oryzae pv. oryzae. Metabolomics 2010, 6, 451–465. [Google Scholar] [CrossRef]
- Gupta, S.V.K.; Rupasinghe, T.W.T.; Callahan, D.L.; Natera, S.H.; Smith, P.M.; Hill, C.B.; Roessner, U.; Boughton, B.A. Spatio-temporal metabolite and elemental profiling of salt stressed barley seeds during initial stages of germination by MALDI-MSI and µ-XRF Spectrometry. Front. Plant Sci. 2019, 10, 1139. [Google Scholar] [CrossRef]
- Piasecka, A.; Sawikowska, A.; Kuczyńska, A.; Ogrodowicz, P.; Mikołajczak, K.; Krystkowiak, K.; Gudyś, K.; Guzy-Wróbelska, J.; Krajewski, P.; Kachlicki, P. Drought-related secondary metabolites of barley (Hordeum vulgare L.) leaves and their metabolomic quantitative trait loci. Plant J. 2017, 89, 898–913. [Google Scholar] [CrossRef]
- Piasecka, A.; Sawikowska, A.; Krajewski, P.; Kachlicki, P. Combined mass spectrometric and chromatographic methods for in-depth analysis of phenolic secondary metabolites in barley leaves. J. Mass Spectrom. 2015, 50, 513–532. [Google Scholar] [CrossRef]
- Khakimov, B.; Rasmussen, M.A.; Kannangara, R.M.; Jespersen, B.M.; Munck, L.; Engelsen, S.B. From metabolome to phenotype: GC-MS metabolomics of developing mutant barley seeds reveals effects of growth, temperature and genotype. Sci. Rep. 2017, 7, 8195. [Google Scholar] [CrossRef]
- Kogel, K.H.; Voll, L.M.; Schäfer, P.; Jansen, C.; Wu, Y.; Langen, G.; Imani, J.; Hofmann, J.; Schmiedl, A.; Sonnewald, S.; et al. Transcriptome and metabolome profiling of field-grown transgenic barley lack induced differences but show cultivar-specific variances. Proc. Natl. Acad. Sci. USA 2010, 107, 6198–6203. [Google Scholar] [CrossRef]
- Roessner, U.; Patterson, J.H.; Forbes, M.G.; Fincher, G.B.; Langridge, P.; Bacic, A. An investigation of boron toxicity in barley using metabolomics. Plant Physiol. 2006, 142, 1087–1101. [Google Scholar] [CrossRef] [PubMed]
- Tugizimana, F.; Djami Tchatchou, A.T.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. Metabolomic analysis of defence-related reprogramming in Sorghum bicolor in response to Colletotrichum sublineolum infection reveals a functional metabolic web of phenylpropanoid and flavonoid pathways. Front. Plant Sci. 2019, 9, 1840. [Google Scholar] [CrossRef] [PubMed]
- Ogbaga, C.C.; Stepien, P.; Dyson, B.C.; Rattray, N.J.; Ellis, D.I.; Goodacre, R.; Johnson, G.N. Biochemical analyses of sorghum varieties reveal differential responses to drought. PLoS ONE 2016, 11, e0154423. [Google Scholar] [CrossRef]
- Turner, M.F.; Heuberger, A.L.; Kirkwood, J.S.; Collins, C.C.; Wolfrum, E.J.; Broeckling, C.D.; Prenni, J.E.; Jahn, C.E. Non-targeted metabolomics in diverse sorghum breeding lines indicates primary and secondary metabolite profiles are associated with plant biomass accumulation and photosynthesis. Front. Plant Sci. 2016, 7, 953. [Google Scholar] [CrossRef]
- Michaletti, A.; Naghavi, M.R.; Toorchi, M.; Zolla, L.; Rinalducci, S. Metabolomics and proteomics reveal drought-stress responses of leaf tissues from spring-wheat. Sci. Rep. 2018, 8, 5710. [Google Scholar] [CrossRef] [PubMed]
- Shi, T.; Zhu, A.; Jia, J.; Hu, X.; Chen, J.; Liu, W.; Ren, X.; Sun, D.; Fernie, A.R.; Cui, F.; et al. Metabolomics analysis and metabolite-agronomic trait associations using kernels of wheat (Triticum aestivum) recombinant inbred lines. Plant J. 2020, (in press). [CrossRef]
- Shewry, P.R.; Corol, D.I.; Jones, H.D.; Beale, M.H.; Ward, J.L. Defining genetic and chemical diversity in wheat grain by 1H-NMR spectroscopy of polar metabolites. Mol. Nutr. Food Res. 2017, 61, 1600807. [Google Scholar] [CrossRef] [PubMed]
- Matthews, S.B.; Santra, M.; Mensack, M.M.; Wolfe, P.; Byrne, P.F.; Thompson, H.J. Metabolite profiling of a diverse collection of wheat lines using ultra performance liquid chromatography coupled with time-of-flight mass spectrometry. PLoS ONE 2012, 7, 44179. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Cheng, F.; Hu, C.; Quan, S.; Lin, H.; Wang, J.; Chen, G.; Zhao, X.; Alexander, D.; Guo, L.; et al. Metabolic map of mature maize kernels. Metabolomics 2014, 10, 775–787. [Google Scholar] [CrossRef]
- Barros, E.; Lezar, S.; Anttonen, M.J.; Van Dijk, J.P.; Röhlig, R.M.; Kok, E.J.; Engel, K.H. Comparison of two GM maize varieties with a near-isogenic non-GM variety using transcriptomics, proteomics and metabolomics. Plant Biotechnol. J. 2010, 8, 436–451. [Google Scholar] [CrossRef] [PubMed]
- Lamari, N.; Zhendre, V.; Urrutia, M.; Bernillon, S.; Maucourt, M.; Deborde, C.; Prodhomme, D.; Jacob, D.; Ballias, P.; Rolin, D.; et al. Metabotyping of 30 maize hybrids under early-sowing conditions reveals potential marker-metabolites for breeding. Metabolomics 2018, 14, 132. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Cao, J.; Wang, X.; Chen, Q.; Jin, W.; Li, Z.; Tian, F. Evolutionary metabolomics identifies substantial metabolic divergence between maize and its wild ancestor, teosinte. Plant Cell 2019, 31, 1990–2009. [Google Scholar] [CrossRef] [PubMed]
- Jin, M.; Zhang, X.; Zhao, M.; Deng, M.; Du, Y.; Zhou, Y.; Wang, S.; Tohge, T.; Fernie, A.R.; Willmitzer, L.; et al. Integrated genomics-based mapping reveals the genetics underlying maize flavonoid biosynthesis. BMC Plant Biol. 2017, 17, 17. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Martín, J.; Heald, J.; Kingston-Smith, A.; Winters, A.; Rubiales, D.; Sanz, M.; Mur, L.A.; Prats, E. A metabolomic study in oats (Avena sativa) highlights a drought tolerance mechanism based upon salicylate signalling pathways and the modulation of carbon, antioxidant and photo-oxidative metabolism. Plant Cell Environ. 2015, 38, 1434–1452. [Google Scholar] [CrossRef] [PubMed]
- Dhawi, F.; Datta, R.; Ramakrishna, W. Metabolomics, biomass and lignocellulosic total sugars analysis in foxtail millet (Setaria italica) inoculated with different combinations of plant growth promoting bacteria and mycorrhiza. Commun. Plant Sci. 2018, 8, 8–14. [Google Scholar] [CrossRef]
- Hamzelou, S.; Pascovici, D.; Kamath, K.S.; Amirkhani, A.; McKay, M.; Mirzaei, M.; Atwell, B.J.; Haynes, P.A. Proteomic responses to drought vary widely among eight diverse genotypes of rice (Oryza sativa). Int. J. Mol. Sci. 2020, 21, 363. [Google Scholar] [CrossRef]
- Khan, M.H.; Dar, Z.A.; Dar, S.A. Breeding strategies for improving rice yield-A review. Agric. Sci. 2015, 6, 467. [Google Scholar] [CrossRef]
- Baranski, M. Golden rice. In The Embryo Project Encyclopedia; Arizona State University: Tempe, AZ, USA, 2013; pp. 1940–5030. [Google Scholar]
- Matsaunyane, L.B.T.; Dubery, I.A. Genetic modification of crop plants: Molecular approaches to address intended unintended effects and substantial equivalence. In Transgenic Crops; Khan, M.S., Malik, K.A., Eds.; InTechOpen Publishers: London, UK, 2019; Chapter 7; pp. 115–131. [Google Scholar]
- Decourcelle, M.; Perez-Fons, L.; Baulande, S.; Steiger, S.; Couvelard, L.; Hem, S.; Zhu, C.; Capell, T.; Christou, P.; Fraser, P.; et al. Combined transcript, proteome, and metabolite analysis of transgenic maize seeds engineered for enhanced carotenoid synthesis reveals pleotropic effects in core metabolism. J. Exp. Bot. 2015, 66, 3141–3150. [Google Scholar] [CrossRef]
- Chen, R.; Zhao, X.; Shao, Z.; Wei, Z.; Wang, Y.; Zhu, L.; Zhao, J.; Sun, M.; He, R.; He, G. Rice UDP-glucose pyrophosphorylase1 is essential for pollen callose deposition and its consumption result in a new type of thermosensible genic male sterility. Plant Cell 2007, 19, 847–861. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Hina, A.; Song, S.; Kong, J.; Bhat, J.A.; Zhao, T. Whole-genome mapping identified novel “QTL hotspots regions” for seed storability in soybean (Glycine max L.). BMC Genom. 2019, 20, 499. [Google Scholar] [CrossRef] [PubMed]
- Tricase, C.; Amicarelli, V.; Lamonaca, E.; Rana, R.L. Economic analysis of the barley market and related uses. In Grasses as Food and Feed; Tadele, Z., Ed.; IntechOpen: London, UK, 2018; pp. 25–46. [Google Scholar]
- Zhao, Y.; Ma, Q.; Jin, X.; Peng, X.; Liu, J.; Deng, L.; Yan, H.; Sheng, L.; Jiang, H.; Cheng, B. A novel maize homeodomain–leucine zipper (HD-Zip) I gene, Zmhdz10, positively regulates drought and salt tolerance in both rice and Arabidopsis. Plant Cell Physiol. 2014, 55, 1142–1156. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, Z.; Li, Y.; Li, Z.; Liu, H.; Zhou, W. Metabolite profiling of sorghum seeds of different colors from different sweet sorghum cultivars using a widely targeted metabolomics approach. Int. J. Genom. 2020, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lamaoui, M.; Jemo, M.; Datla, R.; Bekkaoui, F. Heat and drought stresses in crops and approaches for their mitigation. Front. Chem. 2018, 6, 26. [Google Scholar] [CrossRef]
- Tesso, T.T.; Perumal, R.; Little, C.R.; Adeyanju, A.; Radwan, G.L.; Prom, L.K.; Magill, C.W. Sorghum pathology and biotechnology—A fungal disease perspective: Part II. Anthracnose, stalk rot, and downy mildew. Eur. J. Plant Sci. Biotech. 2012, 6, 31–44. [Google Scholar]
- Hill, C.B.; Taylor, J.D.; Edwards, J.; Mather, D.; Bacic, A.; Langridge, P.; Roessner, U. Whole-genome mapping of agronomic and metabolic traits to identify novel quantitative trait loci in bread wheat grown in a water-limited environment. Plant Physiol. 2013, 162, 1266–1281. [Google Scholar] [CrossRef]
- Hill, C.B.; Taylor, J.D.; Edwards, J.; Mather, D.; Langridge, P.; Bacic, A.; Roessner, U. Detection of QTL for metabolic and agronomic traits in wheat with adjustments for variation at genetic loci that affect plant phenology. Plant Sci. 2015, 233, 143–154. [Google Scholar] [CrossRef]
- Matros, A.; Liu, G.; Hartmann, A.; Jiang, Y.; Zhao, Y.; Wang, H.; Ebmeyer, E.; Korzun, V.; Schachschneider, R.; Kazman, E.; et al. Genome–metabolite associations revealed low heritability, high genetic complexity, and causal relations for leaf metabolites in winter wheat (Triticum aestivum). J. Exp. Bot. 2017, 68, 415–428. [Google Scholar] [CrossRef]
- Seki, M.; Umezawa, T.; Urano, K.; Shinozaki, K. Regulatory metabolic networks in drought stress responses. Curr. Opin. Plant Biol. 2007, 10, 296–302. [Google Scholar] [CrossRef]
- Krasensky, J.; Jonak, C. Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J. Exp. Bot. 2012, 63, 1593–1608. [Google Scholar] [CrossRef] [PubMed]
| Crops and Cvs. | Stage of Development | Plant Organ | Analytical Platforms | Data Analysis Models | Main Discriminatory Metabolites or Classes of Metabolites | References |
|---|---|---|---|---|---|---|
| Rice (Oryza sativa L.) | Flowering and early grain filling stages | Leaves, spikelets, seeds | GC-MS (Untargeted) | PCA | Primary and secondary metabolism: amino acids, succinic acid, sinapic acid, citric acid, ribitol, malic acid, glycolic acid, arabitol, putrescine, erythritol, and vanillic acid. | [113] |
| Rice (Oryza sativa L.) | Panicle formation, heading, milk ripe, dough and full ripe stages | Leaves and grains | 1H NMR and 1H HR-MAS NMR (Targeted) | PCA, OPLS-DA | Primary: saturated and unsaturated fatty acids, organic acids, amino acids. | [111] |
| Rice (Oryza sativa L.) | 24 months | Seeds | GC-MS (Untargeted) | PCA, OPLS-DA, VIP plots. | Primary: sugar-related compounds, amino acid-related compounds, free fatty acids, tricarboxylic acid (TCA) cycle intermediates. | [125] |
| Rice (Oryza sativa L.) | Not specified | Rice grain | UHPLC-MS/MS (Untargeted) | PCA | Primary: aromatic amino acids, carbohydrates, cofactors and vitamins, lipids, oxylipins, nucleotides. Secondary: benzenoids. | [126] |
| Rice (Oryza sativa L.) | Maturition | Mature seeds | UHPLC-MS-MS and GC-MS (Untargeted) | PCA, network-based analyses | Primary: amino acids and derivatives, carbohydrates, lipids, cofactors, prosthetic groups and electron carriers, nucleotides. Secondary: benzenoids. | [127] |
| (Oryza sativa L.) | Not specified | Mature Seeds | HPLC and GC-MS (Targeted and untargeted) | Not specified | Primary: carbohydrates and lipids. Secondary: α-carotene, β-carotene, and lutein. | [128] |
| Rice (Oryza sativa L.) | Six-week-old | Leaves | GC and LC-ToF-MS (Untargeted) | PCA, PLS | Primary: amino acids (arginine, ornithine, citrulline, tyrosine, phenylalanine and lysine), fatty acids and lipids, glutathione, carbohydrates. Secondary: rutin, acetophenone, alkaloids. | [129] |
| Barley (Hordeum vulgare) | Germination | Seeds | MALDI-MSI, LC-QToF-MS | Not specified | Primary: glycero(phospho)lipids, prenol lipids, sterol lipids, methylation. Secondary: polyketides. | [130] |
| Barley (Hordeum vulgare) | Two-leaf stage seedlings | Leaves | LC-ESI-MS/MS (Targeted) | PLS-DA, VIP plots, PCA, HCA | Primary: amino acids and derivatives, organic acids, nucleotides, and derivatives. Secondary: flavonoids, absiscic acid. | [115] |
| Barley (Hordeum vulgare) | Three-leaf stage | Leaves | HPLC-DAD-MSn and UPLC-PDA-MS/MS | ANOVA, Correlation networks | Secondary: flavones, chlorogenic acids, hydrocinnamic acid derivatives, and hordatines and their glycosides. | [131] |
| Barley (Hordeum vulgare) | Three-leaf stage and flag leaf stage | Leaves | HPLC-ESI-MSn; UPLC-MS/MS quadrupole-Orbitrap MS) and NMR | Not specified | Secondary: phenolic compounds, flavonoids, hydroxycinnamic acid glycosides, esters, and amides. | [132] |
| Barley (Hordeum vulgare) | During grain filling | Seeds | GC-MS (Untargeted) | ANOVA, PCA, ASCA, PLS, PLS-DA, VIP plots | Primary: TCA organic acids, aldehydes, alcohols, polyols, fatty acids, carbohydrates, mevalonate. Secondary: phenolic compounds, flavonoids. | [133] |
| Barley (Hordeum vulgare) | Four months | Leaves | ESI-MS (Targeted and untargeted) | PCA | Primary: carbohydrates, free amino acids, carboxylates, phosphorylated intermediates, antioxidants, carotenoids. | [134] |
| Barley (Hordeum vulgare) | One - three weeks old | Leaves and roots | GC-MS (Untargeted and targeted) | HCA, PCA | Primary: amino acids, organic acids, and sugars. | [135] |
| Sorghum (Sorghum bicolor) | Four-leaf stage | Leaves | UHPLC-HDMS (Untargeted) | PCA, OPLS-DA | Primary: amino acids, carboxylic acids, fatty acids, Secondary: cyanogenic glycosides, flavonoids, hydroxycinnamic acids, indoles, benzoates, phytohormones, and shikimates. | [114] |
| Sorghum (Sorghum bicolor) | Four-leaf stage | Leaves | UHPLC-HDMS (Untargeted) | PCA, HCA, OPLS-DA, VIP plots | Secondary: 3-Deoxyanthocyanidins, phenolics, flavonoids, phytohormones, luteolinidin, apigeninidin, riboflavin. | [136] |
| Sorghum (Sorghum bicolor) | Around 26 days | Roots and leaves | FT-IR spectroscopy GC-MS (Untargeted) | PCA, PC-DFA | Primary: sugars, sugar alcohols, amino acids, and organic acids. | [137] |
| Sorghum (Sorghum bicolor) | Four-weeks old | Grain and biomass | GC-MS LC-MS | Z-score, PCA, O2PLS | Primary: organic acids. Secondary: phenylpropanoids. | [138] |
| Wheat (Triticum aestivum) | Not specified | Leaves | UHPLC-MS (Untargeted) | PCA, PLS-DA | Primary: sugars, glycolysis and gluconeogenesis intermediates, amino acids, nucleic acid precursors, and intermediates. Secondary: chorismate, polyamines, L-pipecolate, aminoadipic acid, phenylpropanoids, terpene skeleton, ubiquinone. | [139] |
| Wheat (Triticum aestivum) | Physiological maturity | Leaves | LC-HRMS (Untargeted) | PLS-DA | Primary: amino acid metabolism, sugar alcohols, purine metabolism, glycerolipids, guanine. Secondary: shikimates, anthranilate, absiscic acid. | [112] |
| Wheat (Triticum aestivum) | Maturation | Matured kernels | LC-MS/MS | One-way ANOVA | Primary: fatty acids, sugar, nucleic acids and derivatives. Secondary: phenolamides, flavonoids, polyphenols, vitamins, organic acids, amino acids and derivatives, phytohormones, and derivatives. | [140] |
| Wheat (Triticum aestivum) | Not specified | Grain | 1H-NMR (Targeted) | PCA | Primary: osmolytes, glycine betaine, choline, and asparagine. | [141] |
| Wheat (Triticum aestivum) | Not specified | Seeds | UPLC-ToF-MS (Untargeted) | PCA, OPLS-DA | Primary: sterols, fatty acyls, long chain fatty acid derivatives, glycerol (phospho) lipids. Secondary: polyketides. | [142] |
| Maize (Zea mays) | R6 stage | Grains | GC-MS (Targeted) | PCA, PLS-DA, HCA, heat maps | Primary: sugars, sucrose, glucose, fructose. | [109] |
| Maize (Zea mays) | Physiological maturity | Kernels | UPLC-MS-MS GC-MS (Untargeted) | PCA, PLS-DA, heatmaps | Primary: central metabolism pathways and partial secondary pathways; glycolysis, TCA cycle, starch, amino acids. Secondary: alkaloids, benzenoids, fatty acid and sugar derivatives, flavonoids, phenylpropanoids, and terpenoids. | [143] |
| Maize (Zea mays) | Eight months | Kernels | 1H-NMR fingerprinting GC-MS (Untargeted) | PCA | Primary: glucose, fructose, sucrose, tocopherol, phytosterol, inositol, asparagine, glutamic acid, pyroglutamic acid. | [144] |
| Maize (Zea mays) | Eight-visible-leaf stage | Leaves | LC-ESI-QToF-MS and NMR (Untargeted) | PCA, OSC-PLS-DA | Primary: choline, inositol, sugars, raffinose, rhamnose, TCA cycle, amino acids, trigonelline, putrescine, quinate, shikimate, Secondary: hydroxycinnamates, flavonoids, and benzoxazinoids. | [145] |
| Maize (Zea mays) | Seedling stage | Entire seedling | UPLC-HRMS (Untargeted) | PCA | Primary: amino acids, lipids, carboxylic acid. Secondary: alkaloids, terpenoids, flavonoids, alkaloids, benzenoids. | [146] |
| Maize (Zea mays) | Physiological maturity | Kernels | LC-ESI-MS/MS (Targeted) | Not specified | Secondary: flavanones, flavones, anthocyanins, and methoxylated flavonoids. | [147] |
| Oats (Avena sativa) | Not specified | Grains | GLC-MS | Not specified | Primary metabolism: malic, gluconic, and galacturonic acids, fatty acids, palmitic acid and linoleic acid. | [110] |
| Oats (Avena sativa) | Seedling stage (three-week-old) | Leaves | DI-ESI-MS HPLC-ESI-MS/MS | PCA, DFA | Primary: malonate, hydroxypyruvate, succinate, cysteine, oxaloacetate, ornithine, and glucose. Secondary: cadaverine. | [148] |
| Foxtail millet (Setaria italica) | 60 Days | Shoots | NMR | PCA, HCA, heat maps | Primary: fructose, glucose, gluconate, formate, threonine, 4-aminobutyrate, 2-hydroxyvalerate, sarcosine, betaine, choline, isovalerate, acetate, pyruvate, TCA organic acids, and uridine. | [149] |
| Foxtail millet (Setaria italica) | Three to five leaves stages | Leaves | LC-ESI-QTRAP-MS | PCA, HCA, PLS-DA | Primary: glycerophospholipids, amino acids, organic acids. Secondary: flavonoids, hydroxycinnamic acids, phenolamides, and vitamin-related compounds. | [104] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hamany Djande, C.Y.; Pretorius, C.; Tugizimana, F.; Piater, L.A.; Dubery, I.A. Metabolomics: A Tool for Cultivar Phenotyping and Investigation of Grain Crops. Agronomy 2020, 10, 831. https://doi.org/10.3390/agronomy10060831
Hamany Djande CY, Pretorius C, Tugizimana F, Piater LA, Dubery IA. Metabolomics: A Tool for Cultivar Phenotyping and Investigation of Grain Crops. Agronomy. 2020; 10(6):831. https://doi.org/10.3390/agronomy10060831
Chicago/Turabian StyleHamany Djande, Claude Y., Chanel Pretorius, Fidele Tugizimana, Lizelle A. Piater, and Ian A. Dubery. 2020. "Metabolomics: A Tool for Cultivar Phenotyping and Investigation of Grain Crops" Agronomy 10, no. 6: 831. https://doi.org/10.3390/agronomy10060831
APA StyleHamany Djande, C. Y., Pretorius, C., Tugizimana, F., Piater, L. A., & Dubery, I. A. (2020). Metabolomics: A Tool for Cultivar Phenotyping and Investigation of Grain Crops. Agronomy, 10(6), 831. https://doi.org/10.3390/agronomy10060831







