1. Introduction
Rice production is a very important activity all around the world. Many African and Asian countries rely on rice as a source of income and daily calories [
1,
2,
3]. Bacteria leaf blight which is caused by the pathogen
Xanthomonas oryzae pv.
oryzae (
Xoo) is one of the most serious diseases responsible for significant yield reduction in rice. The disease constrains the photosynthetic area of the crop thereby causing partial grain filling and this leads to poor yield [
4]. Use of resistant varieties of rice is the best approach to guard against the disease as chemical control is not effective. To date, 42
Xoo resistance genes have been identified in rice and some of them have been incorporated into new varieties [
5]. Generally, the disease is favoured by temperatures between 25 to 34 °C, with relative humidity more than 70% and most rice growing regions in Malaysia are affected by the disease [
6]. Host plant resistance offers the most viable, low cost, effective and sustainable strategy to manage the disease compared to chemical control and biological method [
7].
Rice blast caused by the fungus
Magnaporthe oryzae (
Pyricularia oryzae (anamorph)) is one of the biotic agents responsible for severe yield loss in rice [
8]. It is among the most destructive diseases affecting rice in Malaysia and other parts of the world’s major rice growing regions. The disease causes considerably large grain yield losses in almost all major ecosystems where it is grown [
9,
10]. Blast infection could be noticed at every stage of rice growth and the symptoms occur in all above ground parts. The early symptoms look like white/gray/green spots or necrosis surrounded by dark green colour. Molecular markers that co-segregate with the gene are powerful methods to develop a resistant cultivar. The availability of different molecular markers enables an accurate detection of candidate genes of interest [
11].
Resistance genes for bacterial leaf blight and blast are usually obtained from rice varieties, land races, old cultivars and wild species. About 100 blast resistance genes have been identified, and many of them have been cloned and characterized [
12]. Variability in pathogens leading to overcoming of resistance effect of major genes after four to five years, has forced the breeder to continuously search for novel R-genes. Widespread use of resistant cultivars leads to development of selection pressure in the pathogen that resulted into breaking of resistance barrier. Majority of blast R-genes are dominant in nature, but recessive gene has also been reported [
6]. Dominant R-genes for
Xoo are
Xa-1, Xa-3, Xa-4, Xa-7, Xa-10, Xa-14, Xa-21 and
Xa-22 (
t) and some major recessive genes are
xa-5, xa-13. Pyramiding of genes is an effective strategy for enhancing the durability of R-genes over time and space. Marker assisted selection is one of the most sustainable and attractive technology that facilitates incorporation of two or more major genes in different genetic background of rice. In the presence of a number of virulent pathogens, the genes may differ in their level of resistance to such pathogens. Thus, gene pyramiding is currently being pursued in an effort to develop more durable and comprehensive resistant rice varieties to combat the effect of bacterial leaf blight and blast pathogens. Therefore, this work was carried out to introgress multiple resistance genes from two genetically diverse parents using marker-assisted selection for broad spectrum and durable resistance against bacterial leaf blight (BLB) and blast diseases.
4. Discussion
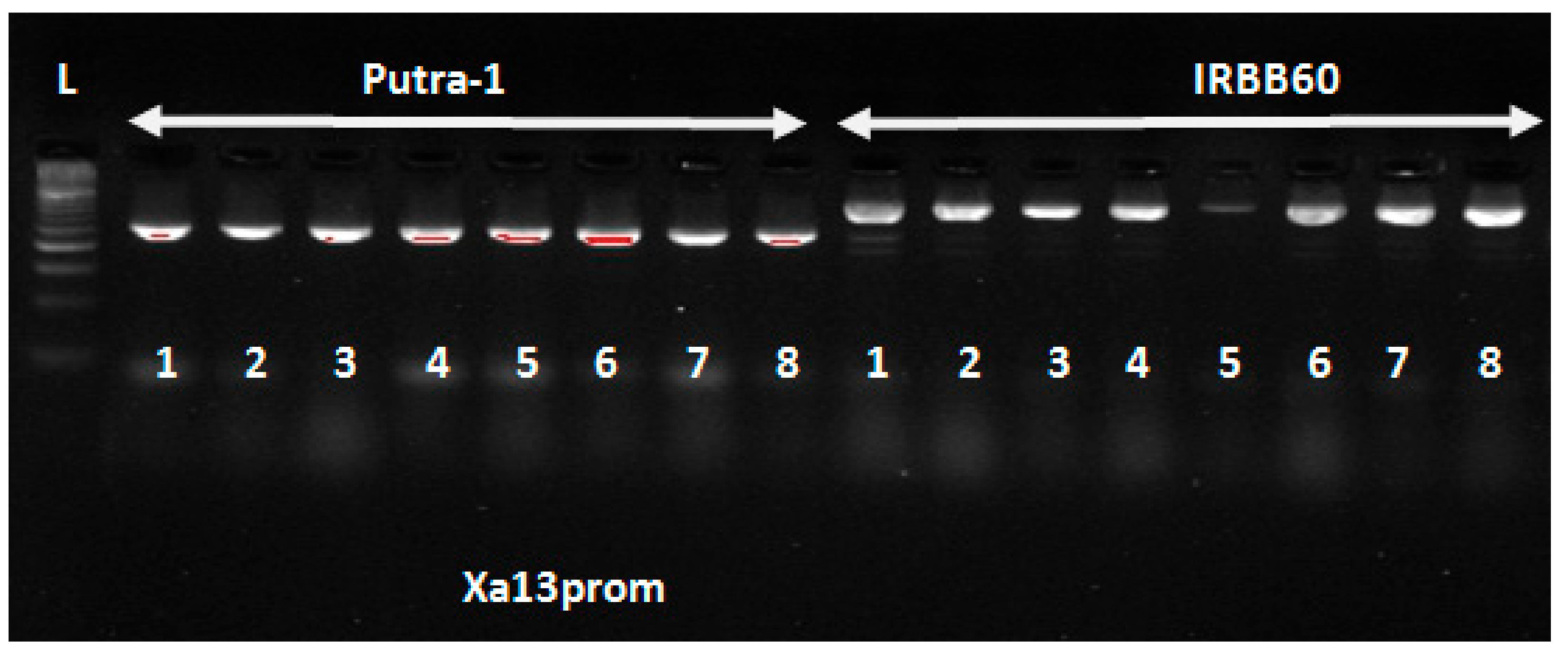
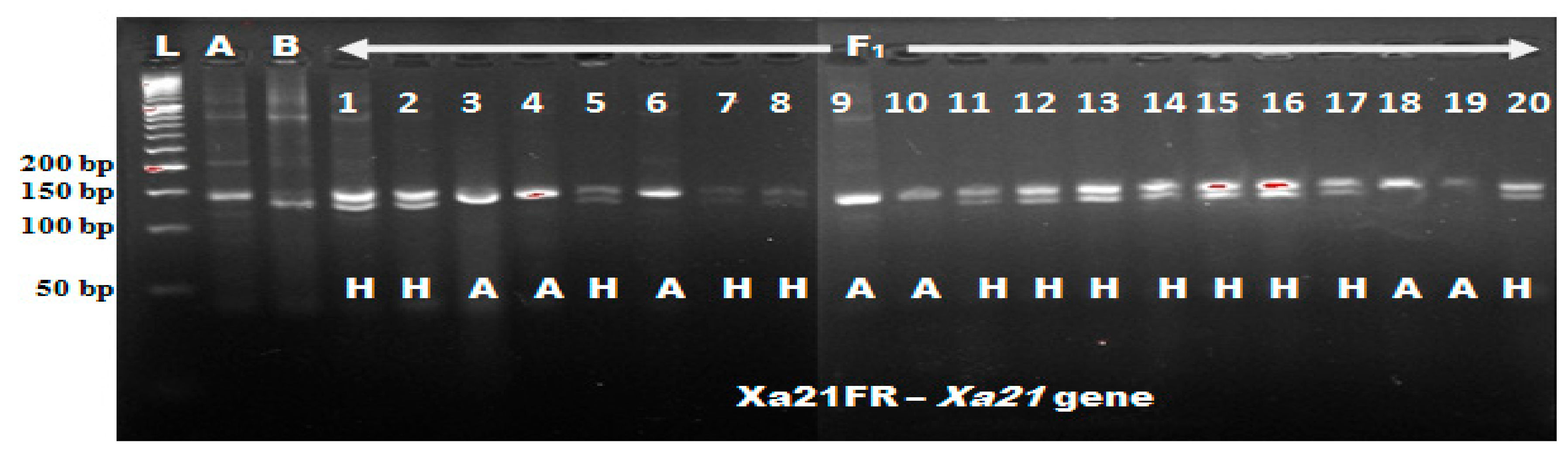
In this study, eight polymorphic tightly linked foreground markers were used to confirm the pyramiding of seven
Xoo and blast resistance genes in the newly improved lines. The four
Xoo resistance genes (
Xa4 +xa5 +
xa13 +
Xa21) were introgressed from IRBB60 variety into the Malaysia elite variety Putra-1 with genetic background of three
Pi genes, namely;
Pi9, Pi2 and
Piz leading to development of lines with seven gene pyramid (
Xa4 + xa5 +
xa13 +
Xa21 + Pi9 + Pi2 +
Piz) for broad spectrum resistance. Pyramiding may involve incorporating genes from two or more parents. For example, Pradhan et al. [
4] pyramided three bacterial leaf blight (
xa5 +
xa13 +
Xa21) resistance genes for broad-spectrum resistance in deep water rice variety, Jalmagna. Three major genes (
Pi1,
Piz-5 and
Pita) were pyramided by Hittalmani et al. [
24] using RFLP markers. Hittalmani et al. [
24] and Castro et al. [
25] combined genes originating from three parents for rice blast and stripe rust in barley, respectively. Gene pyramiding is difficult to achieve using conventional breeding alone because of linkage with some undesirable traits that is very difficult to break even after repeated backcrossings [
26]. When two or more genes are introgressed, phenotypic evaluation has been unable to distinguish the effect of individual gene precisely since each gene confers resistance to and combats multiple races of the pathogen. Moreover, in the presence of a dominant and a recessive allele, the effect of the recessive gene is masked. The advent and easy availability of molecular markers closely associated with each of the resistance genes makes identification of plants with multiple genes possible. Earlier efforts to generate three-gene pyramids in the backgrounds of rice cultivars PR106 [
27], Pusa Basmati [
28] and Samba Mahsuri by the collaborative work of the Directorate of Rice Research, Hyderabad, India (DRR) and the Centre for Cellular and Molecular Biology, Hyderabad, India (CCMB) have been successful [
29].
Studies conducted to identify the best gene combinations conferring broad-spectrum resistance showed that the four-gene combination was the most effective and did not show any sign of breakdown of resistance to various strains of
Xoo from different parts of the country [
30]. Shanty et al. [
31] worked on the pyramiding of four BLB resistance genes
Xa4, xa5,
xa13 and
Xa21 through marker-assisted selection (MAS) in the
Xoo susceptible high yielding rice cv. Mahsuri and parental lines of hybrid rice, maintainers IR58025B and Pusa 6B and the restorer lines KMR3 and PRR78 simultaneously. The results showed increased and broad spectrum resistance to the pathogen populations from different parts of India. The work was the first successful example of the use of molecular markers in foreground selection in conjunction with conventional breeding for the simultaneous introgression of genes of interest into multiple backgrounds [
31].
Popular plant varieties, adopted in large-scale production, may have some undesirable traits. These cultivars are generally high-yielding but may be highly susceptible to the attack of a certain disease or insect. In some cases, these are of poor grain quality, and thus command lower prices in the market. Also, long-term cultivation of single variety with wide adoption can lead to a shift in race or biotype frequency and ultimately cause resistance breakdown [
32].
Pyramiding of genes reduces loss of resistance [
27,
33]. Thus, to further extend the use of these popular varieties, new traits or genes are being introduced through a series of backcrossing [
34]. However, pyramiding is difficult or impossible using the conventional approach due to masking and/or epistatic effects of the genes being combined [
35]. Moreover, some genes have similar reactions to two or more races or biotypes; thus they prove rather difficult to identify and transfer through conventional approaches [
27]. Most of the rice varieties in the Philippines have
Xa4 and
xa5 genes for bacterial leaf blight resistance and some have no resistance at all [
36]. With the identification of new genes like
Xa21,
Xa22 and
Xa23 or the
Om gene from
Oryza minuta, varieties developed earlier can be improved through the incorporation of these new important genes [
37]. Several
Xoo resistance genes have been mapped and tagged using DNA markers and these can be explored in improving resistance [
38]. Among the more than two dozen genes reported [
39,
40,
41,
42,
43], eight genes (
Xa3,
Xa4,
xa5,
Xa7,
xa8,
Xa10,
xa13 and
Xa21) have been introgressed in common genetic background using marker-aided selection [
27,
33,
35,
44,
45,
46]. Lines with single or combinations of genes have been identified using different DNA markers closely linked to the genes. Classical introgression of new genes is being done through a series of backcrossing and selection, and this procedure needs a longer time and requires large resources. The use of DNA as markers for selection has streamlined and facilitated the whole process even without inoculation [
37].
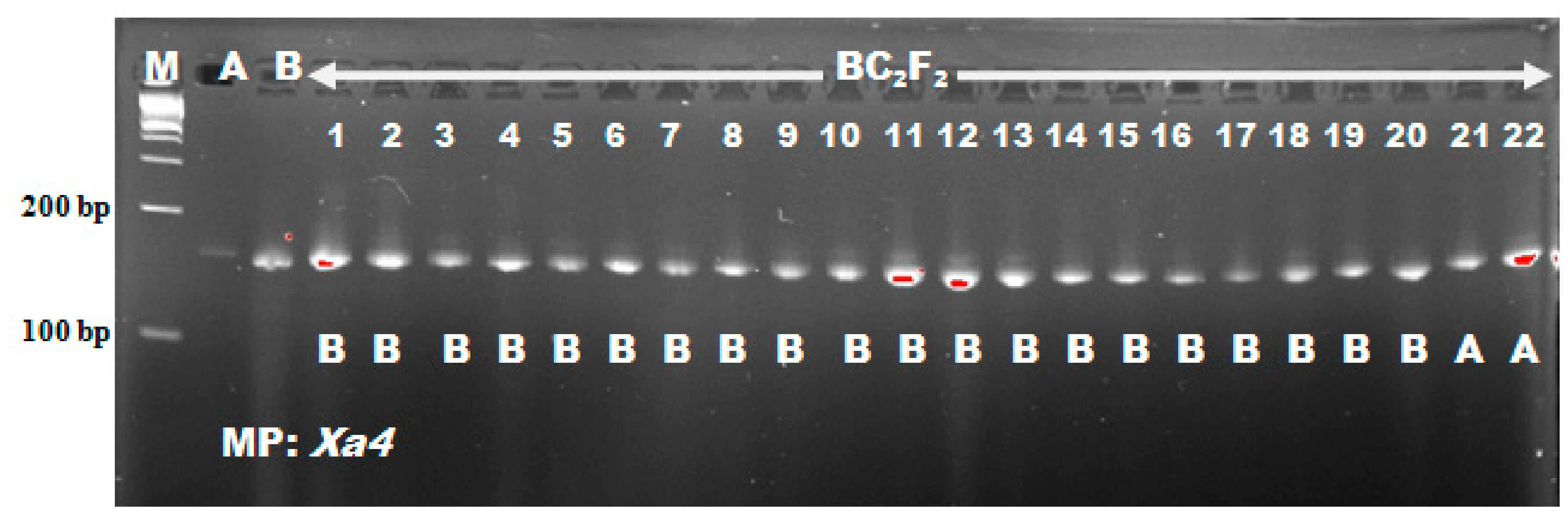
In the BC
2F
2 of this study, segregation distortion was observed among marker loci close to
Xoo resistance genes. A study conducted by Jin et al. [
47] showed that in 300 F
2 populations evaluated, the foreground marker segregation for the
BADH2 gene did not correspond with the expected 1:2:1 Mendelian segregation ratio. Also, the result reported by Amarawathi et al. [
48] did not fit into the tested ratio of 1:2:1 for nksbad2 marker, hence segregation distortion in their RIL individuals. Lau [
49] also observed a segregation ratio of 69:46:44 which failed to fit into the 1:2:1 ratio and the result was attributed to segregation distortion of foreground markers. In the same vein, Muthusamy et al. [
50] reported distortion of foreground marker segregation which was observed in all the backcross stages performed in maize and this was without regard to the size of population evaluated. Therefore, population size of BC
2F
2 progenies could not be taken as a factor responsible for the foreground markers segregation distortion observed in this study. The BC
1F
1 and BC
2F
1 both fitted into the Mendel’s single gene segregation ratio despite having varying population sizes. The distortion of foreground markers segregation observed at
Xoo resistance locus could be attributed to variation in environmental factors. Bing et al. [
51] and Xu et al. [
52] all reported that distortion of marker segregation could be as a result of environmental issues. Similarly, studies conducted by Muthusamy et al. [
50] showed that the distortion in foreground marker segregation in maize backcrossings was due to backcrossings performed during winter season. Wang et al. [
53] discovered distortion of marker segregation particularly in F
2 population which were grown at a particular altitude, but when grown at other altitudes, marker segregation distortion was not detected. There is, therefore, every likelihood that the markers segregation distortion observed in this study was due to environmental effects as the BC
2F
2 progenies were planted at varied weather conditions compared to BC
1F
1 and BC
2F
1 in 2019 and 2018 as obtainable in Malaysia’s agro-ecology. Notwithstanding, the functional markers used in this study facilitated the identification of the genotypes with the targeted
Xoo resistance genes in addition to the already established background genetic make-up of blast resistance in Putra-1.
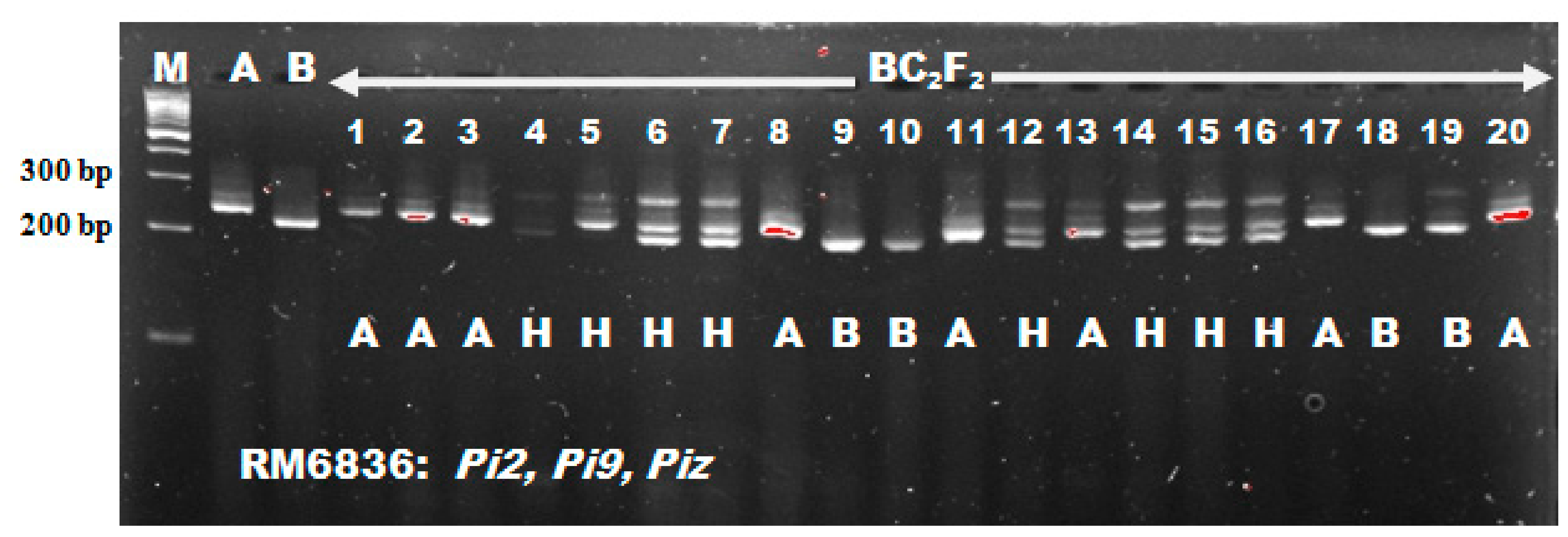
This Chi-square result obtained for blast resistance loci was in line with the result reported by Tanweer et al. [
54] who recorded foreground marker segregation ratio of 53:106:41 and 55:107:38 for RM208 and RM206 respectively which fitted into the 1:2:1 segregation ratio expected from Mendel studies. Also, Miah et al. [
15] reported a segregation ratio of 52:127:74 (RM8225) and 54:132:67 (RM6836) which did not differ significantly from the expected Mendelian ratio of 1:2:1. It is important to confirm the introgression of target gene by phenotyping in the developed cross mainly at individual level as individual phenotypic performance is a good indicator of the genotype provided that genes have major effect on phenotypic expression and the phenotyping error is less [
55]. Phenotypic selection for desirable agro-morphological traits has always been done along with backcross selection [
23]. On the other hand, Young and Tanksley [
56] proposed that the genotypic selection which was later referred to as background selection [
57] involved screening the allelic pedigree throughout the genome with the aid of markers. Singh et al. [
58] reported that foreground selection for target genes followed by stringent phenotyping gives superior yield.
The high RPGR in this study could be as a result of the stringent phenotypic selection done at every backcross generation and selfing. This strategy reduced both time and economic resources input. Singh et al. [
59] also reported that phenotypic and marker-assisted selections increase the RPGR. The result obtained in this present study was in contrast with the findings made by Sabu et al. [
60] and Martinez et al. [
61] who found no significant difference in grain yield between the parental cultivar and its progenies. Hossain et al. [
62] noted that number of panicles is the result of productive tillers which survived to produce panicles. The significant increase in yield contributing traits such as number of tillers, number of panicles, number of leaves, etc., recorded in this present study could have resulted to the significant increase in grain yield per hectare obtained. Dutta et al. [
63] and Kusutani et al. [
64] both reported that higher grain yield in rice genotypes is associated with number of productive tillers per hill and number of grains per panicle. Similarly, seed length and width are essential quantifiable characteristics that are closely related to the external physical quality of rice [
65,
66]. Juliano [
67] reported that seed length and width determines the grain/seed shape of rice. There was a significant difference in the seed length and width of the newly improved selected rice lines compared to their original recurrent parent. However, both lines have slender seed shape (seed length:width ratio >3.0). Shi and Zhu [
68] reported that seed shape is concurrently controlled by cytoplasmic and maternal genes as well as triploid endosperm.