Pseudomonas aeruginosa SG01: A Novel Polyethylene-Degrading Bacterium in Petrochemical Wastewater
Abstract
1. Introduction
2. Materials and Methods
2.1. Soil, Sewage, Sludge, and PE Plastic Products
2.2. Culture Media and Main Chemical Reagents
2.3. PE Pretreatment
2.4. Enrichment and Screening of PE-Degrading Bacteria
2.5. Identification of PE-Degrading Bacteria
2.6. Statistical Analysis
3. Results and Discussion
3.1. Isolation and Purification of Degrading Bacteria
3.2. Growth Curves of Degrading Bacteria
3.3. Degradation Efficiency of Degrading Bacteria
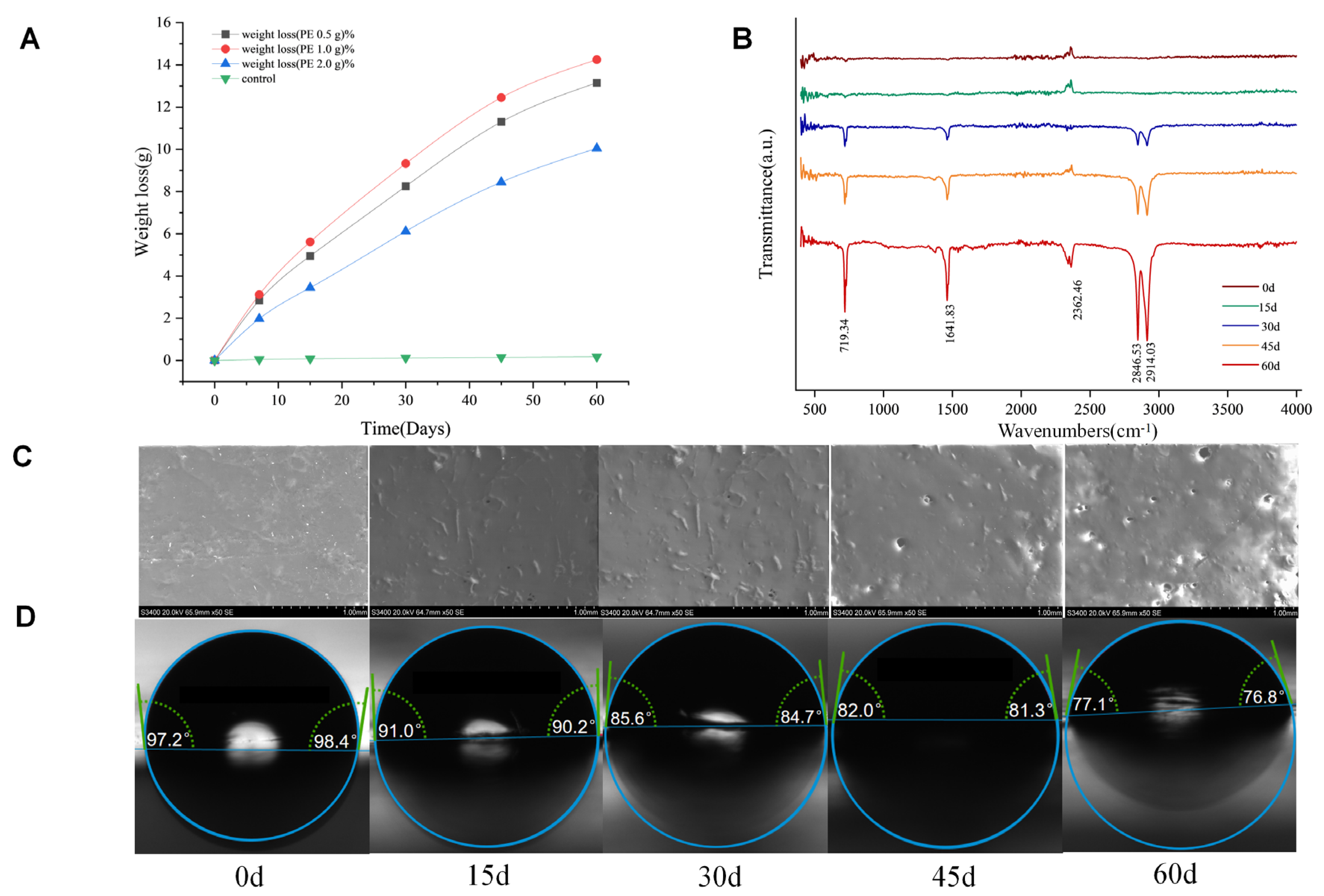
3.3.1. Effect of Degrading Bacteria on Weight Loss Rate of PE Film
3.3.2. Analysis of Surface Morphology Characteristics
3.3.3. Evolution of Functional Groups on PE Film Surface During Microbial Degradation
3.3.4. Changes in Surface Hydrophilicity of PE Films
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| PE | polyethylene |
| FTIR | Fourier transform infrared spectroscopy |
| SEM | Scanning electron microscopy |
| MPs | microplastics |
| POPs | Persistent organic pollutants |
| HDPE | High-density polyethylene |
| LDPE | Low-density polyethylene |
| PP | polypropylene |
| UV | ultraviolet |
| CFU | Colony-Forming Unit |
| NCBI | National Center for Biotechnology Information |
| OD600 | Optical Density at 600 nm |
| AlkB | Alkane Hydroxylase |
| MSM | Mineral Salt Medium |
| LB | Lysogeny Broth |
References
- Flores-Diaz, A.; Arango, J.A.; Calvo, D.C.; Rangel-Mendez, J.R.; Ontiveros-Valencia, A. Physicochemical and microbial treatments for plastics, microplastics, and nanoplastics. Environ. Dev. 2025, 54, 101181. [Google Scholar] [CrossRef]
- Hou, M.; Yang, F.; Song, L.; Hu, R.; Liu, Q.; Zhang, H.; Qi, Y. Enhanced PE degradation and diversity of bacterial biofilm by applying organic fertilizer inoculated with mixed degrading bacteria. Process Biochem. 2025, 151, 137–145. [Google Scholar] [CrossRef]
- Goukeh, M.N.; Ssekimpi, D.; Asefaw, B.K.; Zhang, Z. Occurrence of per-and polyfluorinated substances, microplastics, pharmaceuticals and personal care products as emerging contaminants in landfill leachate: A review. Total Environ. Eng. 2025, 3, 100019. [Google Scholar] [CrossRef]
- Lim, J.-H.; Kang, J.-W. Assessing biofilm formation and resistance of Vibrio parahaemolyticus on UV-aged microplastics in aquatic environments. Water Res. 2024, 254, 121379. [Google Scholar] [CrossRef]
- Feng, F.; Wang, S.; He, X.; Wang, X.; Huang, J.; Liu, G.; Rong, S.; Su, S.; Han, B.; Liu, W. Adsorption behavior and mechanism of cadmium, copper, and lead on polylactic acid microplastics exposed to Ultraviolet light. J. Environ. Chem. Eng. 2025, 13, 117033. [Google Scholar] [CrossRef]
- Dronjak, L.; Exposito, N.; Rovira, J.; Martí, E.; Raya, I.; Carabassa, V.; Domene, X.; Sierra, J. Organic amendments as vectors of micro and macro plastic pollution of terrestrial ecosystems. Sci. Total Environ. 2025, 979, 179434. [Google Scholar] [CrossRef]
- Wang, M.; Li, Q.; Shi, C.; Lv, J.; Xu, Y.; Yang, J.; Chua, S.L.; Jia, L.; Chen, H.; Liu, Q. Oligomer nanoparticle release from polylactic acid plastics catalysed by gut enzymes triggers acute inflammation. Nat. Nanotechnol. 2023, 18, 403–411. [Google Scholar] [CrossRef]
- Jahedi, F.; Fard, N.J.H. Micro-and nanoplastic toxicity in humans: Exposure pathways, cellular effects, and mitigation strategies. Toxicol. Rep. 2025, 14, 102043. [Google Scholar] [CrossRef]
- Deng, L.; Yuan, Y.; Xi, H.; Wan, C.; Yu, Y.; Wu, C. The destiny of microplastics in one typical petrochemical wastewater treatment plant. Sci. Total Environ. 2023, 896, 165274. [Google Scholar] [CrossRef]
- Deng, L.; Xi, H.; Wan, C.; Fu, L.; Wang, Y.; Wu, C. Is the petrochemical industry an overlooked critical source of environmental microplastics? J. Hazard. Mater. 2023, 451, 131199. [Google Scholar] [CrossRef]
- Guo, Y.; Wu, R.; Zhang, H.; Guo, C.; Wu, L.; Xu, J. Distribution of microplastics in the soils of a petrochemical industrial region in China: Ecological and Human Health Risks. Environ. Geochem. Health 2025, 47, 13. [Google Scholar] [CrossRef] [PubMed]
- Luomala, H.; Halme, R.; Jönkkäri, I. Reducing the carbon footprint of railway sleepers using recycled plastics. Front. Sustain. 2024, 5, 1460159. [Google Scholar] [CrossRef]
- Jeon, J.M.; Park, S.J.; Choi, T.R.; Park, J.H.; Yang, Y.H.; Yoon, J.J. Biodegradation of polyethylene and polypropylene by Lysinibacillus species JJY0216 isolated from soil grove. Polym. Degrad. Stab. 2021, 191, 109662. [Google Scholar] [CrossRef]
- Montazer, Z.; Habibi Najafi, M.B.; Levin, D.B. Microbial degradation of low-density polyethylene and synthesis of polyhydroxyalkanoate polymers. Can. J. Microbiol. 2019, 65, 224–234. [Google Scholar] [CrossRef]
- Yang, G.; Quan, X.; Shou, D.; Guo, X.; Ouyang, D.; Zhuang, L. New insights into microbial degradation of polyethylene microplastic and potential polyethylene-degrading bacteria in sediments of the Pearl River Estuary, South China. J. Hazard. Mater. 2025, 486, 137061. [Google Scholar] [CrossRef]
- Li, N.; Han, Z.; Guo, N.; Zhou, Z.; Liu, Y.; Tang, Q. Microplastics spatiotemporal distribution and plastic-degrading bacteria identification in the sanitary and non-sanitary municipal solid waste landfills. J. Hazard. Mater. 2022, 438, 129452. [Google Scholar] [CrossRef]
- Loredo-Treviño, A.; Gutiérrez-Sánchez, G.; Rodríguez-Herrera, R.; Aguilar, C.N. Microbial enzymes involved in polyurethane biodegradation: A review. J. Polym. Environ. 2012, 20, 258–265. [Google Scholar] [CrossRef]
- Roohi; Bano, K.; Kuddus, M.; Zaheer, M.R.; Zia, Q.; Khan, M.F.; Ashraf, G.M.; Gupta, A.; Aliev, G. Microbial enzymatic degradation of biodegradable plastics. Curr. Pharm. Biotechnol. 2017, 18, 429–440. [Google Scholar] [CrossRef]
- Li, Q.; Li, H.; Tian, L.; Wang, Y.; Ouyang, Z.; Li, L.; Mao, Y. Genomic insights and metabolic pathways of an enriched bacterial community capable of degrading polyethylene. Environ. Int. 2025, 197, 109334. [Google Scholar] [CrossRef]
- Tian, H.; Du, Y.; Luo, X.; Dong, J.; Chen, S.; Hu, X.; Zhang, M.; Liu, Z.; Abolfathi, S. Understanding visible light and microbe-driven degradation mechanisms of polyurethane plastics: Pathways, property changes, and product analysis. Water Res. 2024, 259, 121856. [Google Scholar] [CrossRef]
- Bombelli, P.; Howe, C.J.; Bertocchini, F. Polyethylene bio-degradation by caterpillars of the wax moth Galleria mellonella. Curr. Biol. 2017, 27, R292–R293. [Google Scholar] [CrossRef]
- Park, S.Y.; Kim, C.G. Biodegradation of micro-polyethylene particles by bacterial colonization of a mixed microbial consortium isolated from a landfill site. Chemosphere 2019, 222, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Yang, J.; Wu, W.M.; Zhao, J.; Song, Y.; Gao, L.; Yang, R.; Jiang, L. Biodegradation and mineralization of polystyrene by plastic-eating mealworms: Part 1. Chemical and physical characterization and isotopic tests. Environ. Sci. Technol. 2015, 49, 12080–12086. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Prabhu, A.; Aroney, S.T.; Rinke, C. Insights into plastic biodegradation: Community composition and functional capabilities of the superworm (Zophobas morio) microbiome in styrofoam feeding trials. Microb. Genom. 2022, 8, 000842. [Google Scholar] [CrossRef]
- Tao, X.; Ouyang, H.; Zhou, A.; Wang, D.; Matlock, H.; Morgan, J.S.; Ren, A.T.; Mu, D.; Pan, C.; Zhu, X. Polyethylene degradation by a Rhodococcous strain isolated from naturally weathered plastic waste enrichment. Environ. Sci. Technol. 2023, 57, 13901–13911. [Google Scholar] [CrossRef]
- Yun, S.D.; Lee, C.O.; Kim, H.W.; An, S.J.; Kim, S.; Seo, M.J.; Park, C.; Yun, C.H.; Chi, W.S.; Yeom, S.J. Exploring a new biocatalyst from Bacillus thuringiensis JNU01 for polyethylene biodegradation. Environ. Sci. Technol. Lett. 2023, 10, 485–492. [Google Scholar] [CrossRef]
- Yang, J.; Yang, Y.; Wu, W.M.; Zhao, J.; Jiang, L. Evidence of polyethylene biodegradation by bacterial strains from the guts of plastic-eating waxworms. Environ. Sci. Technol. 2014, 48, 13776–13784. [Google Scholar] [CrossRef]
- Howard, S.A.; de Dios, R.; Maslova, E.; Myridakis, A.; Miller, T.H.; McCarthy, R.R. Pseudomonas aeruginosa clinical isolates can encode plastic-degrading enzymes that allow survival on plastic and augment biofilm formation. Cell Rep. 2025, 44, 115650. [Google Scholar] [CrossRef]
- Zhao, K.; Tseng, B.S.; Beckerman, B.; Jin, F.; Gibiansky, M.L.; Harrison, J.J.; Luijten, E.; Parsek, M.R.; Wong, G.C. Psl trails guide exploration and microcolony formation in Pseudomonas aeruginosa biofilms. Nature 2013, 497, 388–391. [Google Scholar] [CrossRef]
- Wang, P.; Zhang, N.; Miao, T.; Chan, J.P.; Huang, H.; Lee, P.K.; Li, Y. Surface touch network structure determines bacterial contamination spread on surfaces and occupant exposure. J. Hazard. Mater. 2021, 416, 126137. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Dou, X.; Zhang, Z.; Zhang, F.; Yan, X.; Xie, Y.; Liu, J.; Zhang, S. Pseudomonas aeruginosa SG01: A Novel Polyethylene-Degrading Bacterium in Petrochemical Wastewater. Polymers 2026, 18, 519. https://doi.org/10.3390/polym18040519
Dou X, Zhang Z, Zhang F, Yan X, Xie Y, Liu J, Zhang S. Pseudomonas aeruginosa SG01: A Novel Polyethylene-Degrading Bacterium in Petrochemical Wastewater. Polymers. 2026; 18(4):519. https://doi.org/10.3390/polym18040519
Chicago/Turabian StyleDou, Xiaohan, Zhiqing Zhang, Fengyuan Zhang, Xi Yan, Yan Xie, Jingru Liu, and Shucai Zhang. 2026. "Pseudomonas aeruginosa SG01: A Novel Polyethylene-Degrading Bacterium in Petrochemical Wastewater" Polymers 18, no. 4: 519. https://doi.org/10.3390/polym18040519
APA StyleDou, X., Zhang, Z., Zhang, F., Yan, X., Xie, Y., Liu, J., & Zhang, S. (2026). Pseudomonas aeruginosa SG01: A Novel Polyethylene-Degrading Bacterium in Petrochemical Wastewater. Polymers, 18(4), 519. https://doi.org/10.3390/polym18040519




