Liquid Crystal Peptide/DNA Coacervates in the Context of Prebiotic Molecular Evolution
Abstract
1. Introduction
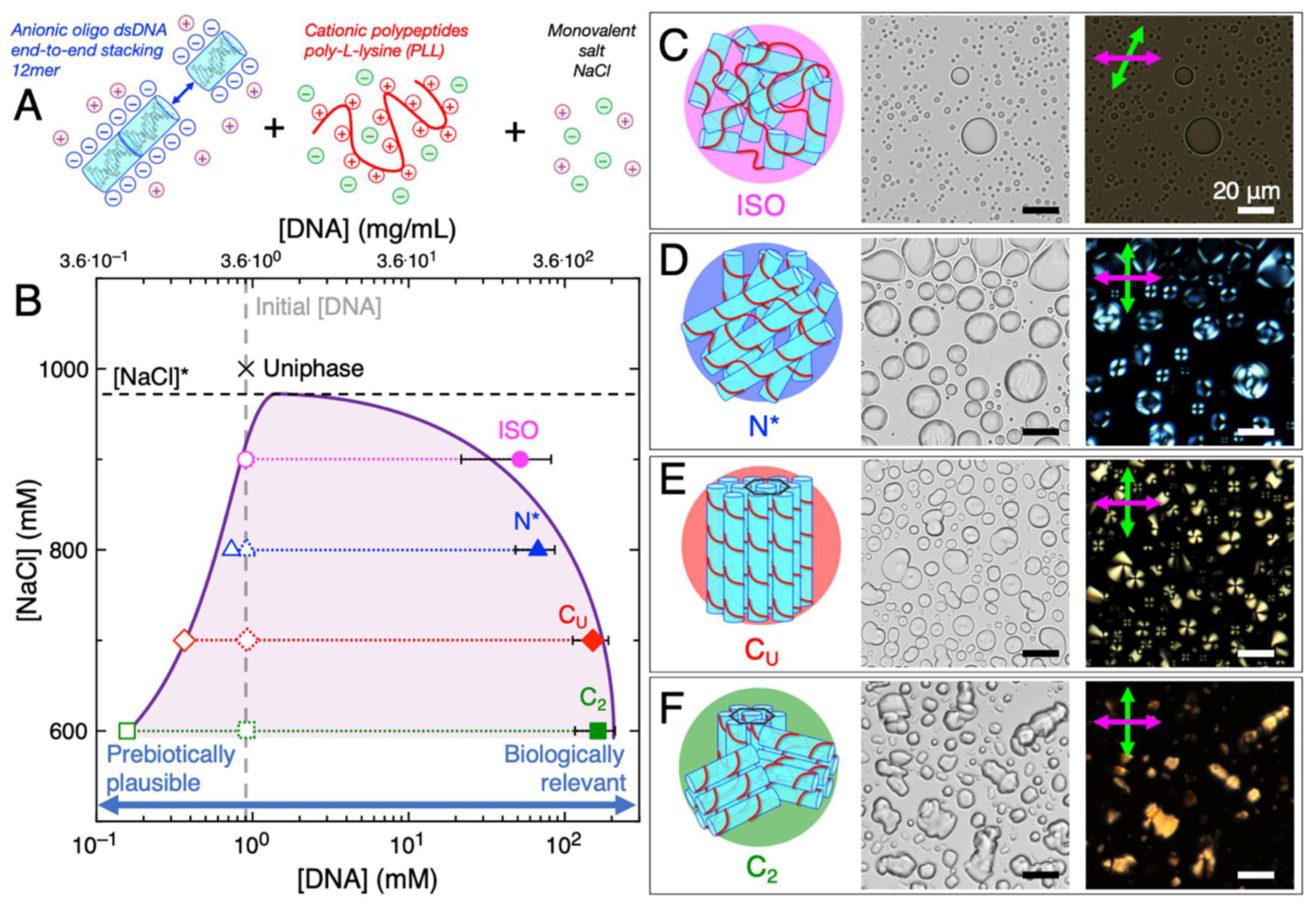
2. LC-Coacervate Structure and Local Nucleic Acid Concentration Increases
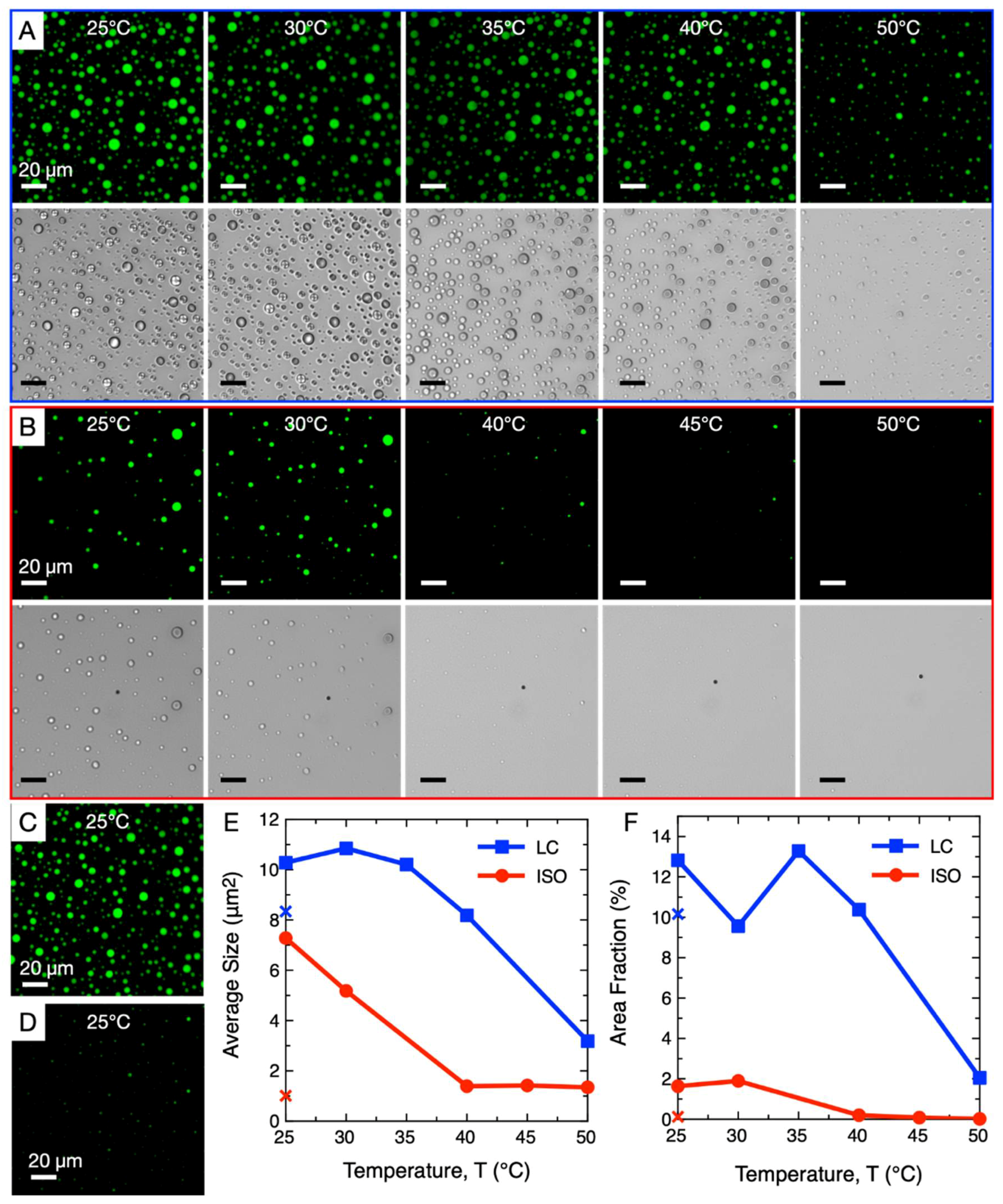
3. Comparison of ISO and LC-Coacervate Stability versus Temperature
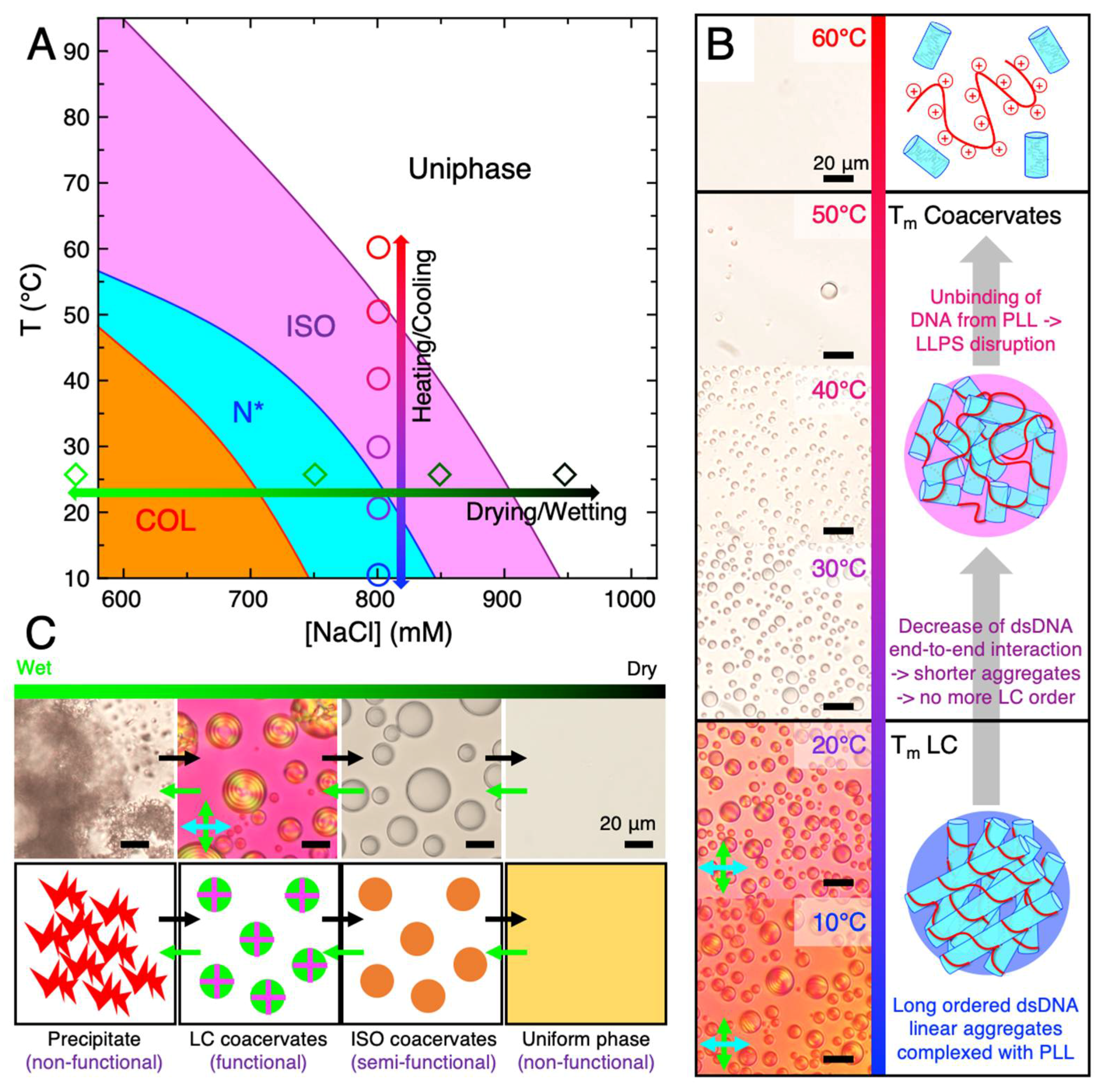
4. Thermal Stability of LC-Coacervates
5. Relevance of LC-Coacervates in Wet–Dry Cycles
6. Relevance of LC-Coacervates to Prebiotic Oceans
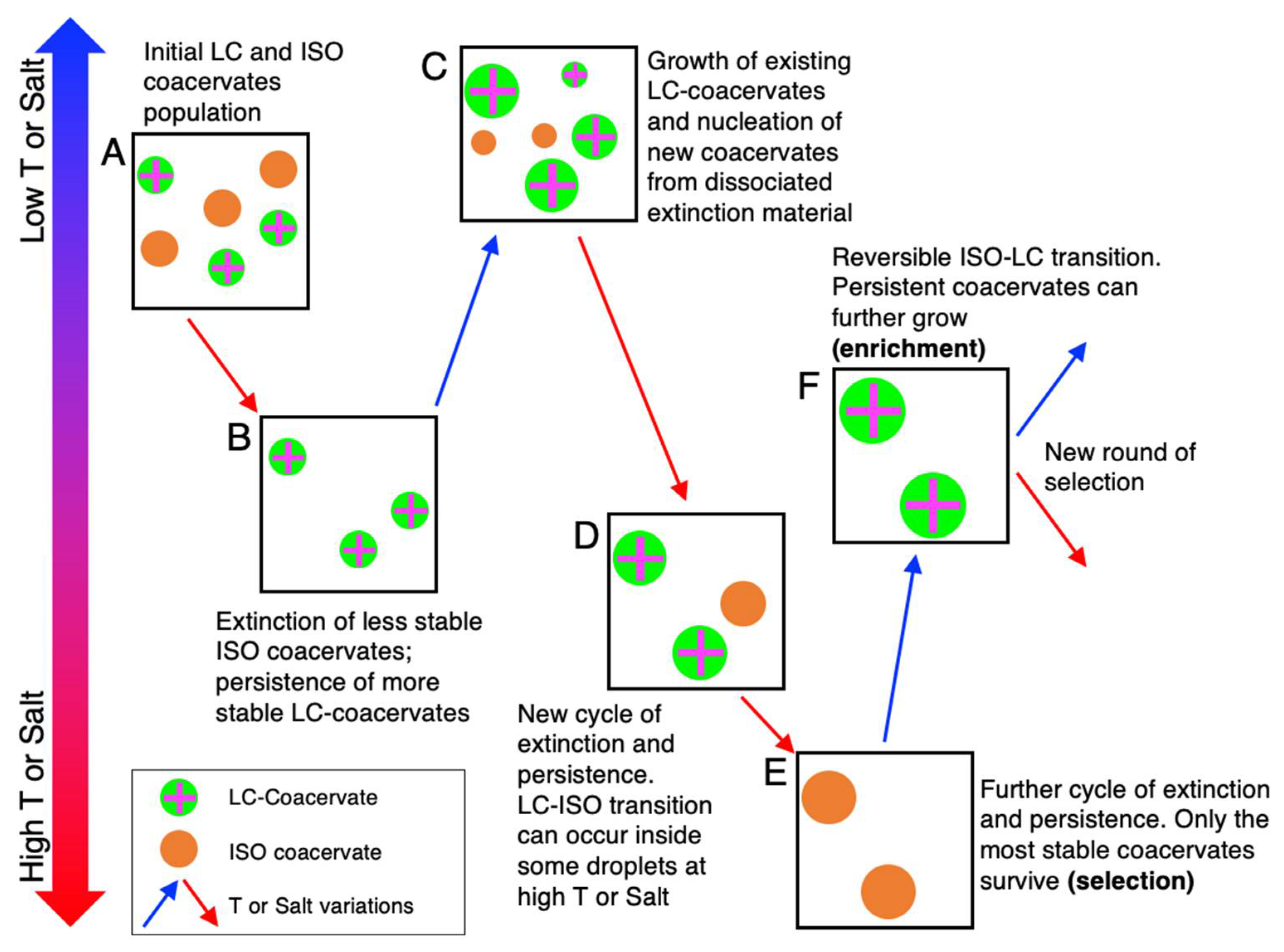
7. Stability-Based Evolutionary Scenarios in LC-Coacervate Systems
8. Relevance of Duplex DNA Structures over Single-Stranded DNA (ssDNA) Structures
9. Conclusions
10. Methods
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Szostak, J.W.; Bartel, D.P.; Luigi Luisi, P. Synthesizing life. Nature 2001, 409, 387–390. [Google Scholar] [CrossRef] [PubMed]
- Bansho, Y.; Furubayashi, T.; Ichihashi, N.; Yomo, T. Host–parasite oscillation dynamics and evolution in a compartmentalized RNA replication system. Proc. Natl. Acad. Sci. USA 2016, 113, 4045–4050. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.A.; Walde, P. From self-assembled vesicles to protocells. Cold Spring Harb. Perspect. Biol. 2010, 2, a002170. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Song, S.; van Hest, J.; Abdelmohsen, L.K.E.A.; Huang, X.; Sánchez, S. Biomimicry of Cellular Motility and Communication Based on Synthetic Soft-Architectures. Small 2020, 16, e1907680. [Google Scholar] [CrossRef] [PubMed]
- Dzieciol, A.J.; Mann, S. Designs for life: Protocell models in the laboratory. Chem. Soc. Rev. 2012, 41, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Yoshizawa, T.; Nozawa, R.-S.; Jia, T.Z.; Saio, T.; Mori, E. Biological phase separation: Cell biology meets biophysics. Biophys. Rev. 2020, 12, 519–539. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.; Brangwynne, C.P. Liquid phase condensation in cell physiology and disease. Science 2017, 357, eaaf4382. [Google Scholar] [CrossRef]
- Keating, C.D. Aqueous phase separation as a possible route to compartmentalization of biological molecules. Acc. Chem. Res. 2012, 45, 2114–2124. [Google Scholar] [CrossRef]
- Mann, S. Systems of Creation: The Emergence of Life from Nonliving Matter. Acc. Chem. Res. 2012, 45, 2131–2141. [Google Scholar] [CrossRef]
- Nakashima, K.K.; Vibhute, M.A.; Spruijt, E. Biomolecular Chemistry in Liquid Phase Separated Compartments. Front. Mol. Biosci. 2019, 6, 21. [Google Scholar] [CrossRef]
- Sing, C.E.; Perry, S.L. Recent progress in the science of complex coacervation. Soft Matter 2020, 16, 2885–2914. [Google Scholar] [CrossRef] [PubMed]
- Aumiller, W.M., Jr.; Pir Cakmak, F.; Davis, B.W.; Keating, C.D. RNA-Based Coacervates as a Model for Membraneless Organelles: Formation, Properties, and Interfacial Liposome Assembly. Langmuir 2016, 32, 10042–10053. [Google Scholar] [CrossRef]
- Jia, T.Z.; Hentrich, C.; Szostak, J.W. Rapid RNA Exchange in Aqueous Two-Phase System and Coacervate Droplets. Orig. Life Evol. Biosph. 2014, 44, 1–12. [Google Scholar] [CrossRef]
- Koga, S.; Williams, D.S.; Perriman, A.W.; Mann, S. Peptide-nucleotide microdroplets as a step towards a membrane-free protocell model. Nat. Chem. 2011, 3, 720–724. [Google Scholar] [CrossRef]
- Jia, T.Z.; Chandru, K.; Hongo, Y.; Afrin, R.; Usui, T.; Myojo, K.; Cleaves, H.J., 2nd. Membraneless polyester microdroplets as primordial compartments at the origins of life. Proc. Natl. Acad. Sci. USA 2019, 116, 15830–15835. [Google Scholar] [CrossRef]
- Frankel, E.A.; Bevilacqua, P.C.; Keating, C.D. Polyamine/Nucleotide Coacervates Provide Strong Compartmentalization of Mg2+, Nucleotides, and RNA. Langmuir 2016, 32, 2041–2049. [Google Scholar] [CrossRef] [PubMed]
- Tang, T.-Y.D.; Rohaida Che Hak, C.; Thompson, A.J.; Kuimova, M.K.; Williams, D.S.; Perriman, A.W.; Mann, S. Fatty acid membrane assembly on coacervate microdroplets as a step towards a hybrid protocell model. Nat. Chem. 2014, 6, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Drobot, B.; Iglesias-Artola, J.M.; Le Vay, K.; Mayr, V.; Kar, M.; Kreysing, M.; Mutschler, H.; Tang, T.-Y.D. Compartmentalised RNA catalysis in membrane-free coacervate protocells. Nat. Commun. 2018, 9, 3643. [Google Scholar] [CrossRef]
- Poudyal, R.R.; Guth-Metzler, R.M.; Veenis, A.J.; Frankel, E.A.; Keating, C.D.; Bevilacqua, P.C. Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates. Nat. Commun. 2019, 10, 490. [Google Scholar] [CrossRef]
- Perry, S.L.; Leon, L.; Hoffmann, K.Q.; Kade, M.J.; Priftis, D.; Black, K.A.; Wong, D.; Klein, R.A.; Pierce, C.F., 3rd; Margossian, K.O.; et al. Chirality-selected phase behaviour in ionic polypeptide complexes. Nat. Commun. 2015, 6, 6052. [Google Scholar] [CrossRef]
- Perry, S.; Li, Y.; Priftis, D.; Leon, L.; Tirrell, M. The Effect of Salt on the Complex Coacervation of Vinyl Polyelectrolytes. Polymers 2014, 6, 1756–1772. [Google Scholar] [CrossRef]
- Priftis, D.; Tirrell, M. Phase behaviour and complex coacervation of aqueous polypeptide solutions. Soft Matter 2012, 8, 9396–9405. [Google Scholar] [CrossRef]
- Mulkidjanian, A.Y.; Bychkov, A.Y.; Dibrova, D.V.; Galperin, M.Y.; Koonin, E.V. Origin of first cells at terrestrial, anoxic geothermal fields. Proc. Natl. Acad. Sci. USA 2012, 109, E821–E830. [Google Scholar] [CrossRef]
- Fares, H.; Marras, A.; Ting, J.; Tirrell, M.; Keating, C. Impact of Wet-Dry Cycling on the Phase Behavior and Compartmentalization Properties of Complex Coacervates. ChemRXiv 2020. [Google Scholar] [CrossRef]
- Parker, E.T.; Cleaves, H.J.; Dworkin, J.P.; Glavin, D.P.; Callahan, M.; Aubrey, A.; Lazcano, A.; Bada, J.L. Primordial synthesis of amines and amino acids in a 1958 Miller H2S-rich spark discharge experiment. Proc. Natl. Acad. Sci. USA 2011, 108, 5526–5531. [Google Scholar] [CrossRef]
- Koga, T.; Naraoka, H. A new family of extraterrestrial amino acids in the Murchison meteorite. Sci. Rep. 2017, 7, 636. [Google Scholar] [CrossRef] [PubMed]
- Orgel, L.E. Polymerization on the rocks: Theoretical introduction. Orig. Life Evol. Biosph. 1998, 28, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Forsythe, J.G.; Yu, S.-S.; Mamajanov, I.; Grover, M.A.; Krishnamurthy, R.; Fernández, F.M.; Hud, N.V. Ester-Mediated Amide Bond Formation Driven by Wet-Dry Cycles: A Possible Path to Polypeptides on the Prebiotic Earth. Angew. Chem. Int. Ed Engl. 2015, 54, 9871–9875. [Google Scholar] [CrossRef]
- Furukawa, Y.; Chikaraishi, Y.; Ohkouchi, N.; Ogawa, N.O.; Glavin, D.P.; Dworkin, J.P.; Abe, C.; Nakamura, T. Extraterrestrial ribose and other sugars in primitive meteorites. Proc. Natl. Acad. Sci. USA 2019, 116, 24440–24445. [Google Scholar] [CrossRef]
- Becker, S.; Feldmann, J.; Wiedemann, S.; Okamura, H.; Schneider, C.; Iwan, K.; Crisp, A.; Rossa, M.; Amatov, T.; Carell, T. Unified prebiotically plausible synthesis of pyrimidine and purine RNA ribonucleotides. Science 2019, 366, 76–82. [Google Scholar] [CrossRef]
- Higgs, P.G. The Effect of Limited Diffusion and Wet-Dry Cycling on Reversible Polymerization Reactions: Implications for Prebiotic Synthesis of Nucleic Acids. Life 2016, 6, 24. [Google Scholar] [CrossRef]
- Szostak, J.W. The Narrow Road to the Deep Past: In Search of the Chemistry of the Origin of Life. Angew. Chem. Int. Ed. Engl. 2017, 56, 11037–11043. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, W. Origin of life: The RNA world. Nature 1986, 319, 618. [Google Scholar] [CrossRef]
- Bhowmik, S.; Krishnamurthy, R. The role of sugar-backbone heterogeneity and chimeras in the simultaneous emergence of RNA and DNA. Nat. Chem. 2019, 11, 1009–1018. [Google Scholar] [CrossRef]
- Hong, F.; Zhang, F.; Liu, Y.; Yan, H. DNA Origami: Scaffolds for Creating Higher Order Structures. Chem. Rev. 2017, 117, 12584–12640. [Google Scholar] [CrossRef]
- Han, D.; Qi, X.; Myhrvold, C.; Wang, B.; Dai, M.; Jiang, S.; Bates, M.; Liu, Y.; An, B.; Zhang, F.; et al. Single-stranded DNA and RNA origami. Science 2017, 358, eaao2648. [Google Scholar] [CrossRef]
- Anastassacos, F.M.; Zhao, Z.; Zeng, Y.; Shih, W.M. Glutaraldehyde Cross-Linking of Oligolysines Coating DNA Origami Greatly Reduces Susceptibility to Nuclease Degradation. J. Am. Chem. Soc. 2020, 142, 3311–3315. [Google Scholar] [CrossRef]
- Bellini, T.; Zanchetta, G.; Fraccia, T.P.; Cerbino, R.; Tsai, E.; Smith, G.P.; Moran, M.J.; Walba, D.M.; Clark, N.A. Liquid crystal self-assembly of random-sequence DNA oligomers. Proc. Natl. Acad. Sci. USA 2012, 109, 1110–1115. [Google Scholar] [CrossRef]
- Fraccia, T.P.; Smith, G.P.; Bethge, L.; Zanchetta, G.; Nava, G.; Klussmann, S.; Clark, N.A.; Bellini, T. Liquid Crystal Ordering and Isotropic Gelation in Solutions of Four-Base-Long DNA Oligomers. ACS Nano 2016, 10, 8508–8516. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.P.; Fraccia, T.P.; Todisco, M.; Zanchetta, G.; Zhu, C.; Hayden, E.; Bellini, T.; Clark, N.A. Backbone-free duplex-stacked monomer nucleic acids exhibiting Watson-Crick selectivity. Proc. Natl. Acad. Sci. USA 2018, 115, E7658–E7664. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, S.; Jo, K.; Kinoshita, H.; Esumi, M.; Tanaka, M. Guanine damage by singlet oxygen from SYBR Green I in liquid crystalline DNA. Org. Biomol. Chem. 2020. [Google Scholar] [CrossRef]
- Livolant, F.; Levelut, A.M.; Doucet, J.; Benoit, J.P. The highly concentrated liquid-crystalline phase of DNA is columnar hexagonal. Nature 1989, 339, 724–726. [Google Scholar] [CrossRef]
- Nakata, M.; Zanchetta, G.; Chapman, B.D.; Jones, C.D.; Cross, J.O.; Pindak, R.; Bellini, T.; Clark, N.A. End-to-end stacking and liquid crystal condensation of 6 to 20 base pair DNA duplexes. Science 2007, 318, 1276–1279. [Google Scholar] [CrossRef] [PubMed]
- Zanchetta, G.; Bellini, T.; Nakata, M.; Clark, N.A. Physical polymerization and liquid crystallization of RNA oligomers. J. Am. Chem. Soc. 2008, 130, 12864–12865. [Google Scholar] [CrossRef]
- Di Leo, S.; Todisco, M.; Bellini, T.; Fraccia, T.P. Phase separations, liquid crystal ordering and molecular partitioning in mixtures of PEG and DNA oligomers. Liq. Cryst. 2018, 45, 2306–2318. [Google Scholar] [CrossRef]
- Fraccia, T.; Smith, G.; Clark, N.; Bellini, T. Liquid Crystal Ordering of Four-Base-Long DNA Oligomers with Both G–C and A–T Pairing. Crystals 2017, 8, 5. [Google Scholar] [CrossRef]
- Leforestier, A.; Livolant, F. The bacteriophage genome undergoes a succession of intracapsid phase transitions upon DNA ejection. J. Mol. Biol. 2010, 396, 384–395. [Google Scholar] [CrossRef]
- Manning, G.S. The persistence length of DNA is reached from the persistence length of its null isomer through an internal electrostatic stretching force. Biophys. J. 2006, 91, 3607–3616. [Google Scholar] [CrossRef] [PubMed]
- Matczyszyn, K.; Olesiak-Banska, J. DNA as scaffolding for nanophotonic structures. J. Nanophotonics 2012, 6, 064505. [Google Scholar] [CrossRef]
- Sung, B.; Kim, M.-H. Liquid-crystalline nanoarchitectures for tissue engineering. Beilstein J. Nanotechnol. 2018, 9, 205–215. [Google Scholar] [CrossRef]
- Kansui, H.; Hiraoka, S.; Kunieda, T. Liquid Crystal Control of Bimolecular Thermal Reactions. Highly Regioselective Pericycloaddition of Fumarates to 2,6-Dialkoxyanthracenes in Liquid−Crystalline Media. J. Am. Chem. Soc. 1996, 118, 5346–5352. [Google Scholar] [CrossRef]
- Rusling, J.F. Liquid crystal surfactant films for electrochemical catalysis. Microporous Mater. 1994, 3, 1–16. [Google Scholar] [CrossRef]
- Lucchetti, L.; Fraccia, T.P.; Ciciulla, F.; Bellini, T. Non-linear optical measurement of the twist elastic constant in thermotropic and DNA lyotropic chiral nematics. Sci. Rep. 2017, 7, 4959. [Google Scholar] [CrossRef] [PubMed]
- Lucchetti, L.; Fraccia, T.P.; Ciciulla, F.; Simoni, F.; Bellini, T. Giant optical nonlinearity in DNA lyotropic liquid crystals. Opt. Express 2017, 25, 25951–25959. [Google Scholar] [CrossRef]
- Lucchetti, L.; Fraccia, T.P.; Nava, G.; Turiv, T.; Ciciulla, F.; Bethge, L.; Klussmann, S.; Lavrentovich, O.D.; Bellini, T. Elasticity and Viscosity of DNA Liquid Crystals. ACS Macro Lett. 2020, 9, 1034–1039. [Google Scholar] [CrossRef]
- Fraccia, T.P.; Smith, G.P.; Zanchetta, G.; Paraboschi, E.; Yi, Y.; Walba, D.M.; Dieci, G.; Clark, N.A.; Bellini, T. Abiotic ligation of DNA oligomers templated by their liquid crystal ordering. Nat. Commun. 2015, 6, 6424. [Google Scholar] [CrossRef]
- Fraccia, T.P.; Zanchetta, G.; Rimoldi, V.; Clark, N.A.; Bellini, T. Evidence of Liquid Crystal–Assisted Abiotic Ligation of Nucleic Acids. Orig. Life Evol. Biosph. 2015, 45, 51–68. [Google Scholar] [CrossRef]
- Todisco, M.; Fraccia, T.P.; Smith, G.P.; Corno, A.; Bethge, L.; Klussmann, S.; Paraboschi, E.M.; Asselta, R.; Colombo, D.; Zanchetta, G.; et al. Nonenzymatic Polymerization into Long Linear RNA Templated by Liquid Crystal Self-Assembly. ACS Nano 2018, 12, 9750–9762. [Google Scholar] [CrossRef]
- Mountain, G.A.; Keating, C.D. Formation of Multiphase Complex Coacervates and Partitioning of Biomolecules within them. Biomacromolecules 2019, 21, 630–640. [Google Scholar] [CrossRef]
- Lu, T.; Spruijt, E. Multiphase Complex Coacervate Droplets. J. Am. Chem. Soc. 2020, 142, 2905–2914. [Google Scholar] [CrossRef]
- Sato, Y.; Sakamoto, T.; Takinoue, M. Sequence-based engineering of dynamic functions of micrometer-sized DNA droplets. Sci. Adv. 2020, 6, eaba3471. [Google Scholar] [CrossRef] [PubMed]
- Rumyantsev, A.M.; de Pablo, J.J. Liquid Crystalline and Isotropic Coacervates of Semiflexible Polyanions and Flexible Polycations. Macromolecules 2019, 52, 5140–5156. [Google Scholar] [CrossRef]
- Fraccia, T.P.; Jia, T.Z. Liquid Crystal Coacervates Composed of Short Double-Stranded DNA and Cationic Peptides. ACS Nano 2020. [Google Scholar] [CrossRef] [PubMed]
- Shakya, A.; King, J.T. DNA Local-Flexibility-Dependent Assembly of Phase-Separated Liquid Droplets. Biophys. J. 2018, 115, 1840–1847. [Google Scholar] [CrossRef] [PubMed]
- Priftis, D.; Megley, K.; Laugel, N.; Tirrell, M. Complex coacervation of poly(ethylene-imine)/polypeptide aqueous solutions: Thermodynamic and rheological characterization. J. Colloid Interface Sci. 2013, 398, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Pearce, B.K.D.; Pudritz, R.E.; Semenov, D.A.; Henning, T.K. Origin of the RNA world: The fate of nucleobases in warm little ponds. Proc. Natl. Acad. Sci. USA 2017, 114, 11327–11332. [Google Scholar] [CrossRef]
- Todisco, M.; Smith, G.P.; Fraccia, T.P. Liquid Crystal ordering of DNA Dickerson Dodecamer duplexes with different 5’- Phosphate terminations. Mol. Cryst. Liq. Cryst. 2019, 683, 69–80. [Google Scholar] [CrossRef]
- Livolant, F.; Maestre, M.F. Circular dichroism microscopy of compact forms of DNA and chromatin in vivo and in vitro: Cholesteric liquid-crystalline phases of DNA and single dinoflagellate nuclei. Biochemistry 1988, 27, 3056–3068. [Google Scholar] [CrossRef]
- Owczarzy, R.; You, Y.; Moreira, B.G.; Manthey, J.A.; Huang, L.; Behlke, M.A.; Walder, J.A. Effects of sodium ions on DNA duplex oligomers: Improved predictions of melting temperatures. Biochemistry 2004, 43, 3537–3554. [Google Scholar] [CrossRef]
- Kreysing, M.; Keil, L.; Lanzmich, S.; Braun, D. Heat flux across an open pore enables the continuous replication and selection of oligonucleotides towards increasing length. Nat. Chem. 2015, 7, 203–208. [Google Scholar] [CrossRef]
- Mullineaux, L.S.; Metaxas, A.; Beaulieu, S.E.; Bright, M.; Gollner, S.; Grupe, B.M.; Herrera, S.; Kellner, J.B.; Levin, L.A.; Mitarai, S.; et al. Exploring the Ecology of Deep-Sea Hydrothermal Vents in a Metacommunity Framework. Front. Mar. Sci. 2018, 5, 1158. [Google Scholar] [CrossRef]
- Fu, J.; Schlenoff, J.B. Driving Forces for Oppositely Charged Polyion Association in Aqueous Solutions: Enthalpic, Entropic, but Not Electrostatic. J. Am. Chem. Soc. 2016, 138, 980–990. [Google Scholar] [CrossRef] [PubMed]
- Kilchherr, F.; Wachauf, C.; Pelz, B.; Rief, M.; Zacharias, M.; Dietz, H. Single-molecule dissection of stacking forces in DNA. Science 2016, 353. [Google Scholar] [CrossRef] [PubMed]
- SantaLucia, J., Jr.; Hicks, D. The thermodynamics of DNA structural motifs. Annu. Rev. Biophys. Biomol. Struct. 2004, 33, 415–440. [Google Scholar] [CrossRef]
- Elder, R.M.; Emrick, T.; Jayaraman, A. Understanding the effect of polylysine architecture on DNA binding using molecular dynamics simulations. Biomacromolecules 2011, 12, 3870–3879. [Google Scholar] [CrossRef]
- Vieregg, J.R.; Lueckheide, M.; Marciel, A.B.; Leon, L.; Bologna, A.J.; Rivera, J.R.; Tirrell, M.V. Oligonucleotide-Peptide Complexes: Phase Control by Hybridization. J. Am. Chem. Soc. 2018, 140, 1632–1638. [Google Scholar] [CrossRef]
- Zouhir, S.; Perchat, S.; Nicaise, M.; Perez, J.; Guimaraes, B.; Lereclus, D.; Nessler, S. Peptide-binding dependent conformational changes regulate the transcriptional activity of the quorum-sensor NprR. Nucleic Acids Res. 2013, 41, 7920–7933. [Google Scholar] [CrossRef]
- Upadhyay, S.K. Binding and thermodynamics of REV peptide-ctDNA interaction. Biopolymers 2017, 108, e22902. [Google Scholar] [CrossRef]
- Solovyev, A.Y.; Tarnovskaya, S.I.; Chernova, I.A.; Shataeva, L.K.; Skorik, Y.A. The interaction of amino acids, peptides, and proteins with DNA. Int. J. Biol. Macromol. 2015, 78, 39–45. [Google Scholar] [CrossRef]
- Ross, D.S.; Deamer, D. Dry/Wet Cycling and the Thermodynamics and Kinetics of Prebiotic Polymer Synthesis. Life 2016, 6, 28. [Google Scholar] [CrossRef]
- Campbell, T.D.; Febrian, R.; McCarthy, J.T.; Kleinschmidt, H.E.; Forsythe, J.G.; Bracher, P.J. Prebiotic condensation through wet-dry cycling regulated by deliquescence. Nat. Commun. 2019, 10, 4508. [Google Scholar] [CrossRef]
- Chandru, K.; Guttenberg, N.; Giri, C.; Hongo, Y.; Butch, C.; Mamajanov, I.; James Cleaves, H. Simple prebiotic synthesis of high diversity dynamic combinatorial polyester libraries. Commun. Chem. 2018, 1, 30. [Google Scholar] [CrossRef]
- Mamajanov, I.; MacDonald, P.J.; Ying, J.; Duncanson, D.M.; Dowdy, G.R.; Walker, C.A.; Engelhart, A.E.; Fernández, F.M.; Grover, M.A.; Hud, N.V.; et al. Ester Formation and Hydrolysis during Wet–Dry Cycles: Generation of Far-from-Equilibrium Polymers in a Model Prebiotic Reaction. Macromolecules 2014, 47, 1334–1343. [Google Scholar] [CrossRef]
- Becker, S.; Schneider, C.; Okamura, H.; Crisp, A.; Amatov, T.; Dejmek, M.; Carell, T. Wet-dry cycles enable the parallel origin of canonical and non-canonical nucleosides by continuous synthesis. Nat. Commun. 2018, 9, 163. [Google Scholar] [CrossRef]
- Hud, N.V. Searching for lost nucleotides of the pre-RNA World with a self-refining model of early Earth. Nat. Commun. 2018, 9, 5171. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, S. Seawater Quality Detection. In Water Quality Monitoring and Management; Elsevier: Amsterdam, The Netherlands, 2019; pp. 233–249. ISBN 9780128113301. [Google Scholar]
- Garcia, A.K.; Schopf, J.W.; Yokobori, S.-I.; Akanuma, S.; Yamagishi, A. Reconstructed ancestral enzymes suggest long-term cooling of Earth’s photic zone since the Archean. Proc. Natl. Acad. Sci. USA 2017, 114, 4619–4624. [Google Scholar] [CrossRef]
- Krissansen-Totton, J.; Arney, G.N.; Catling, D.C. Constraining the climate and ocean pH of the early Earth with a geological carbon cycle model. Proc. Natl. Acad. Sci. USA 2018, 115, 4105–4110. [Google Scholar] [CrossRef] [PubMed]
- Knauth, L.P. Temperature and salinity history of the Precambrian ocean: Implications for the course of microbial evolution. In Geobiology: Objectives, Concepts, Perspectives; Elsevier: Amsterdam, The Netherlands, 2005; pp. 53–69. ISBN 9780444520197. [Google Scholar]
- Marty, B.; Avice, G.; Bekaert, D.V.; Broadley, M.W. Salinity of the Archaean oceans from analysis of fluid inclusions in quartz. C. R. Geosci. 2018, 350, 154–163. [Google Scholar] [CrossRef]
- Stephens, D.W. Changes in lake levels, salinity and the biological community of Great Salt Lake (Utah, USA), 1847–1987. In Saline Lakes; Comín, F.A., Northcote, T.G., Eds.; Utah Geological and Mineral Survey Bull; Springer: Dordrecht, The Netherlands, 1990; Volume 913, pp. 139–146. ISBN 9789401067591. [Google Scholar]
- Baxter, B.K. Great Salt Lake microbiology: A historical perspective. Int. Microbiol. 2018, 21, 79–95. [Google Scholar] [CrossRef]
- Lensky, N.G.; Dvorkin, Y.; Lyakhovsky, V.; Gertman, I.; Gavrieli, I. Water, salt, and energy balances of the Dead Sea: MASS AND ENERGY BALANCES OF THE DEAD SEA. Water Resour. Res. 2005, 41, 513. [Google Scholar] [CrossRef]
- Obolewski, K.; Glińska-Lewczuk, K.; Szymańska, M.; Mrozińska, N.; Bąkowska, M.; Astel, A.; Lew, S.; Paturej, E. Patterns of salinity regime in coastal lakes based on structure of benthic invertebrates. PLoS ONE 2018, 13, e0207825. [Google Scholar] [CrossRef] [PubMed]
- Antunes, A.; Ngugi, D.K.; Stingl, U. Microbiology of the Red Sea (and other) deep-sea anoxic brine lakes. Environ. Microbiol. Rep. 2011, 3, 416–433. [Google Scholar] [CrossRef] [PubMed]
- Blokhuis, A.; Lacoste, D.; Nghe, P.; Peliti, L. Selection Dynamics in Transient Compartmentalization. Phys. Rev. Lett. 2018, 120, 158101. [Google Scholar] [CrossRef] [PubMed]
- Szostak, J.W. The eightfold path to non-enzymatic RNA replication. J. Syst. Chem. 2012, 3, 2. [Google Scholar] [CrossRef]
- He, C.; Gállego, I.; Laughlin, B.; Grover, M.A.; Hud, N.V. A viscous solvent enables information transfer from gene-length nucleic acids in a model prebiotic replication cycle. Nat. Chem. 2017, 9, 318–324. [Google Scholar] [CrossRef]
- Mariani, A.; Bonfio, C.; Johnson, C.M.; Sutherland, J.D. pH-Driven RNA Strand Separation under Prebiotically Plausible Conditions. Biochemistry 2018, 57, 6382–6386. [Google Scholar] [CrossRef]
- Singleton, M.R.; Dillingham, M.S.; Wigley, D.B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 2007, 76, 23–50. [Google Scholar] [CrossRef]
- Krishnamurthy, R. Giving Rise to Life: Transition from Prebiotic Chemistry to Protobiology. Acc. Chem. Res. 2017, 50, 455–459. [Google Scholar] [CrossRef]
- Lubock, N.B.; Zhang, D.; Sidore, A.M.; Church, G.M.; Kosuri, S. A systematic comparison of error correction enzymes by next-generation sequencing. Nucleic Acids Res. 2017, 45, 9206–9217. [Google Scholar] [CrossRef]
- Prévôt, M.E.; Ustunel, S.; Hegmann, E. Liquid Crystal Elastomers-A Path to Biocompatible and Biodegradable 3D-LCE Scaffolds for Tissue Regeneration. Materials 2018, 11, 377. [Google Scholar] [CrossRef]
- Frenkel-Pinter, M.; Samanta, M.; Ashkenasy, G.; Leman, L.J. Prebiotic Peptides: Molecular Hubs in the Origin of Life. Chem. Rev. 2020, 120, 4707–4765. [Google Scholar] [CrossRef] [PubMed]
- Kitadai, N.; Maruyama, S. Origins of building blocks of life: A review. Geosci. Front. 2018, 9, 1117–1153. [Google Scholar] [CrossRef]
- Naskar, S.; Saurabh, S.; Jang, Y.H.; Lansac, Y.; Maiti, P.K. Liquid crystal ordering of nucleic acids. Soft Matter 2020, 16, 634–641. [Google Scholar] [CrossRef]
- Huang, M.; Song, J.; Huang, P.; Chen, X.; Wang, W.; Zhu, Z.; Song, Y.; Yang, C. Molecular Crowding Evolution for Enabling Discovery of Enthalpy-Driven Aptamers for Robust Biomedical Applications. Anal. Chem. 2019, 91, 10879–10886. [Google Scholar] [CrossRef] [PubMed]
- Saad, N.Y. A ribonucleopeptide world at the origin of life: Co-evolution of RNA, amino acids and peptide. J. Syst. Evol. 2018, 56, 1–13. [Google Scholar] [CrossRef]
- Piette, B.M.A.G.; Heddle, J.G. A Peptide-Nucleic Acid Replicator Origin for Life. Trends Ecol. Evol. 2020, 35, 397–406. [Google Scholar] [CrossRef]
- Poudyal, R.R.; Pir Cakmak, F.; Keating, C.D.; Bevilacqua, P.C. Physical Principles and Extant Biology Reveal Roles for RNA-Containing Membraneless Compartments in Origins of Life Chemistry. Biochemistry 2018, 57, 2509–2519. [Google Scholar] [CrossRef]
- Damer, B.; Deamer, D. The Hot Spring Hypothesis for an Origin of Life. Astrobiology 2019, 20, 429–452. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, T.Z.; Fraccia, T.P. Liquid Crystal Peptide/DNA Coacervates in the Context of Prebiotic Molecular Evolution. Crystals 2020, 10, 964. https://doi.org/10.3390/cryst10110964
Jia TZ, Fraccia TP. Liquid Crystal Peptide/DNA Coacervates in the Context of Prebiotic Molecular Evolution. Crystals. 2020; 10(11):964. https://doi.org/10.3390/cryst10110964
Chicago/Turabian StyleJia, Tony Z., and Tommaso P. Fraccia. 2020. "Liquid Crystal Peptide/DNA Coacervates in the Context of Prebiotic Molecular Evolution" Crystals 10, no. 11: 964. https://doi.org/10.3390/cryst10110964
APA StyleJia, T. Z., & Fraccia, T. P. (2020). Liquid Crystal Peptide/DNA Coacervates in the Context of Prebiotic Molecular Evolution. Crystals, 10(11), 964. https://doi.org/10.3390/cryst10110964





