Surface Display—An Alternative to Classic Enzyme Immobilization
Abstract
1. Introduction
2. Immobilization Strategies
2.1. Protein Surface Display in Saccharomyces cerevisiae
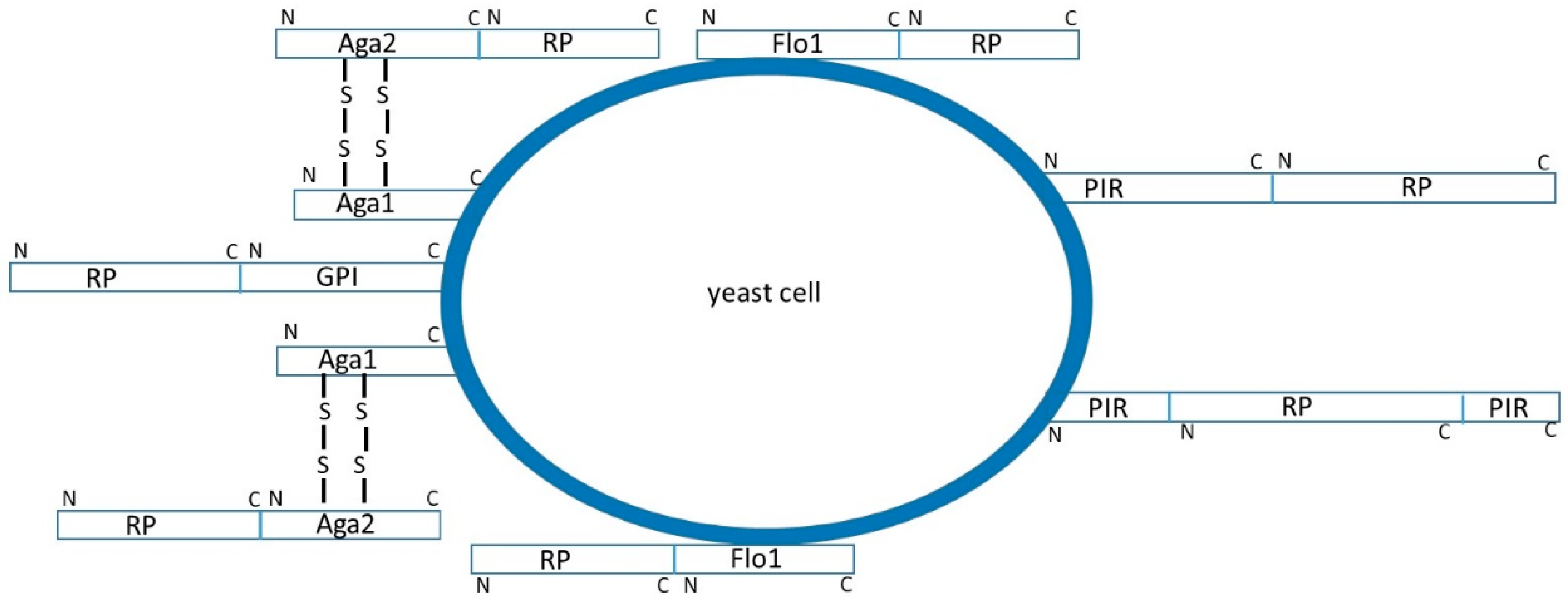
2.1.1. Recombinant Protein Fusion to α-agglutinin
2.1.2. Recombinant Protein Fusion to Other GPI-Anchored Proteins
2.1.3. Recombinant Protein Fusion to PIR Proteins
2.1.4. Approaches for Increasing Efficiency of Recombinant Protein Surface Display in S. cerevisiae
Engineering of Cell Wall Surface and Secretory Mechanisms
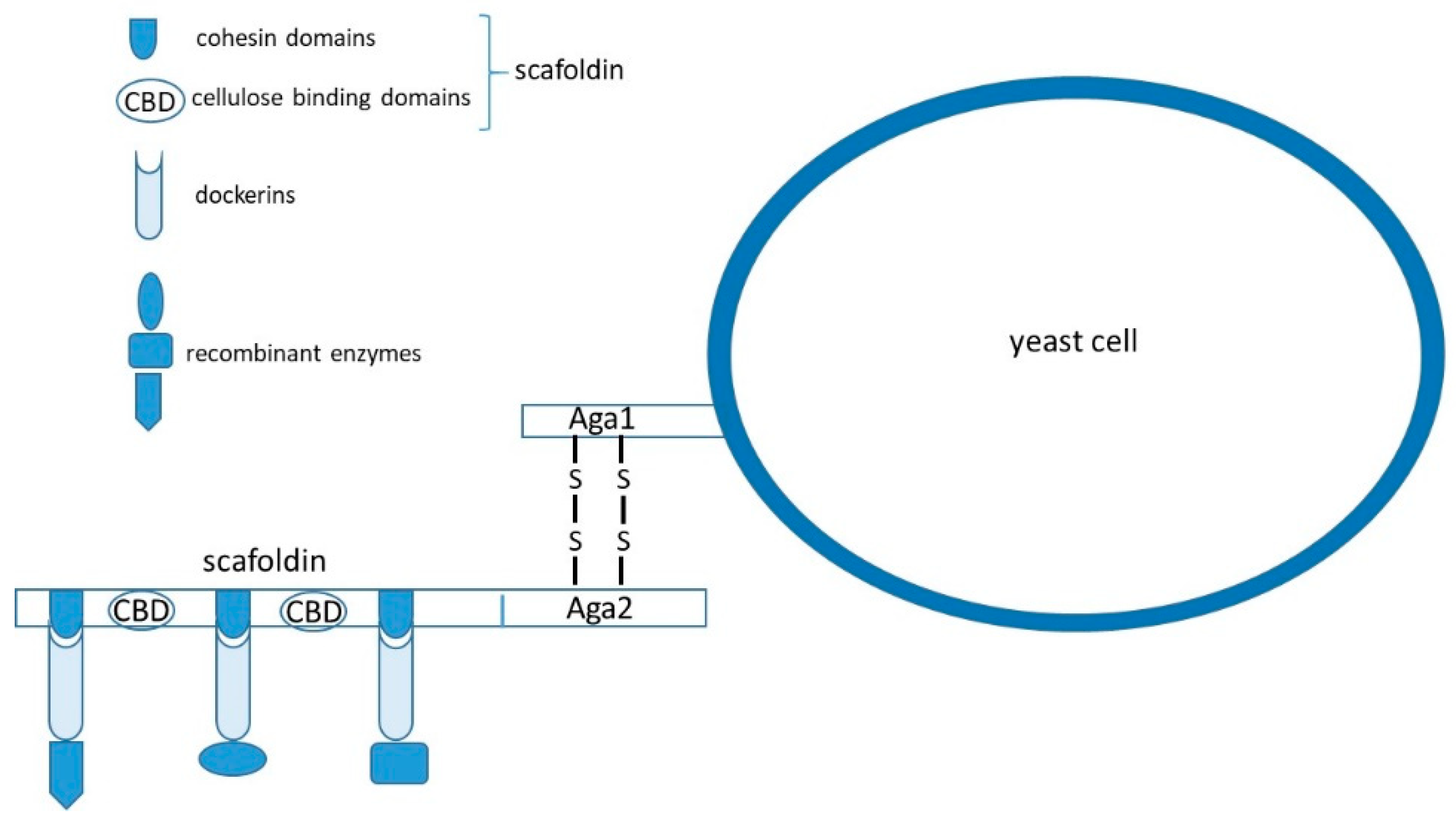
Construction of Synthetic Yeast Cellulosomes
2.2. Protein Surface Display in Pichia pastoris
2.3. Protein Surface Display in Yarrowia lipolytica
2.4. Protein Surface Display in Blastobotrys adeninivorans
2.5. Protein Surface Display in Schizosaccharomyces pombe
2.6. Protein Surface Display in Debaryomyces hansenii
2.7. Protein Surface Display in Kluyveromyces marxianus
3. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Abe, H.; Ohba, M.; Shimma, Y.; Jigami, Y. Yeast cells harboring human β-1,3-fucosyltransferase at the cell surface engineered using Pir, a cell wall-anchored protein. FEMS Yeast Res. 2004, 4, 417–425. [Google Scholar] [CrossRef]
- Matsumoto, T.; Fukudu, H.; Ueda, M.; Tanaka, A.; Kondo, A. Construction of yeast strains with high cell surface lipase activity by using novel display systems based on the Flo1p flocculation functional domain. Appl. Environ. Microbiol. 2002, 68, 4517–4522. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Shibasaki, S.; Ueda, M.; Tanaka, A.; Fukuda, H.; Kondo, A. Development of novel whole-cell immunoadsorbents by yeast surface display of the IgG-binding domain. Appl. Microbiol. Biotechnol. 2001, 57, 500–505. [Google Scholar] [CrossRef] [PubMed]
- van der Vaart, J.M.; Biesbeke, R.; Chapman, J.W.; Toschka, H.Y.; Klis, F.M.; Verrips, T. Comparison of cell wall proteins of Saccharomyces cerevisiae as anchors for cell expression of heterologous proteins. Appl. Environ. Microbiol. 1997, 63, 615–620. [Google Scholar] [PubMed]
- Ito, J.; Kosugi, A.; Tanaka, T.; Kuroda, K.; Shibasaki, S.; Ogino, C.; Ueda, M.; Fukuda, H.; Doi, R.H.; Kondo, A. Regulation of the display ratio of enzymes on the Saccharomyces cerevisiae cell surface by the immunoglobulin G and cellulosomal enzyme binding domains. Appl. Environ. Microbiol. 2009, 75, 4149–4154. [Google Scholar] [CrossRef] [PubMed]
- Tsai, S.L.; Oh, J.; Singh, S.; Chen, R.; Chen, W. Functional assembly of minicellulosomes on the Saccharomyces cerevisiae cell surface for cellulose hydrolysis and ethanol production. Appl. Environ. Microbiol. 2009, 75, 6087–6093. [Google Scholar] [CrossRef] [PubMed]
- Wen, F.; Sun, J.; Zhao, H. Yeast surface display of trifunctional minicellulosomes for simultaneous saccharification and fermentation of cellulose to ethanol. Appl. Environ. Microbiol. 2010, 76, 1251–1260. [Google Scholar] [CrossRef] [PubMed]
- Kondo, A.; Ueda, M. Yeast cell-surface display—applications of molecular display. Appl. Microbiol. Biotechnol. 2004, 64, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Jose, J.; Maas, R.M.; Teese, M.G. Autodisplay of enzymes—Molecular basis and perspectives. J. Biotechnol. 2012, 161, 92–103. [Google Scholar] [CrossRef] [PubMed]
- Detzel, C.; Maas, R.; Jose, J. Autodisplay of Nitrilase from Alcaligenes faecalis in E. coli Yields a Whole Cell Biocatalyst for the Synthesis of Enantiomerically Pure (R)-Mandelic Acid. Chem. Cat. Chem. 2011, 3, 719–725. [Google Scholar] [CrossRef]
- Watari, J.; Takata, Y.; Ogawa, M.; Sahara, H.; Koshino, S.; Onnela, M.L.; Airaksinen, U.; Jaatinen, R.; Penttilä, M.; Keränen, S. Molecular cloning and analysis of the yeast flocculation gene FLO1. Yeast 1994, 10, 211–225. [Google Scholar] [CrossRef] [PubMed]
- Breinig, F.; Diehl, B.; Rau, S.; Zimmer, C.; Schwab, H.; Schmitt, M.J. Cell surface expression of bacterial esterase a by Saccharomyces cerevisiae and its enhancement by constitutive activation of the cellular unfolded protein response. Appl. Environ. Microbiol. 2006, 72, 7140–7147. [Google Scholar] [CrossRef] [PubMed]
- Shigechi, H.; Koh, J.; Fujita, Y.; Matsumoto, T.; Bito, Y.; Ueda, M.; Satoh, E.; Fukuda, H.; Kondo, A. Direct production of ethanol from raw corn starch via fermentation by use of a novel surface-engineered yeast strain codisplaying glucoamylase and α-amylase. Appl. Environ. Microbiol. 2004, 70, 5037–5040. [Google Scholar] [CrossRef] [PubMed]
- Sato, N.; Matsumoto, T.; Ueda, M.; Tanaka, A.; Fukuda, H.; Kondo, A. Long anchor using Flo1p protein enhances reactivity of cell surface-displayed glucoamylase to polymer substrates. Appl. Microbiol. Biotechnol. 2002, 60, 549–747. [Google Scholar]
- Nakamura, Y.; Matsumoto, T.; Nomoto, F.; Ueda, M.; Fukuda, H.; Kondo, A. Enhancement of activity of lipase-displaying yeast cells and their application to optical resolution of (R, S)-1-benzyloxy-3-chloro-2-propyl monosuccinate. Biotechnol. Progr. 2006, 22, 998–1002. [Google Scholar] [CrossRef] [PubMed]
- Tanino, T.; Ohno, T.; Aoki, T.; Fukuda, H.; Kondo, A. Development of yeast cells displaying Candida antarctica lipase B and their application to ester synthesis reaction. Appl. Microbiol. Biotechnol. 2007, 75, 1319–1325. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Gao, B.; Ren, R.; Tao, X.; Ma, Y.; Wei, D. Efficient display of active lipase LipB52 with a Pichia pastoris cell surface display system and comparison with the LipB52 displayed on Saccharomyces cerevisiae cell surface. BMC Biotechnol. 2008, 8, 4–10. [Google Scholar] [CrossRef]
- Schreuder, M.P.; Brekelmans, S.; van den Ende, H.; Klis, F.M. Targeting of heterologous proteins on the surface of yeast cells. Yeast 1993, 9, 399–409. [Google Scholar] [CrossRef]
- Murai, T.; Ueda, M.; Yamamura, M.; Atomi, H.; Shibasaki, Y.; Kamasawa, N.; Osumi, M.; Amachi, T.; Tanaka, A. Construction of a starch-utilizing yeast by cell surface engineering. Appl. Environ. Microbiol. 1997, 63, 1362–1366. [Google Scholar]
- Washida, M.; Takahashi, S.; Ueda, M.; Tanaka, A. Spacer mediated display of active lipase on the yeast cell surface. Appl. Microbiol. Biotechnol. 2001, 56, 681–686. [Google Scholar] [CrossRef]
- Tokuhiro, K.; Ishida, N.; Kondo, A.; Takahashi, H. Lactic fermentation of cellobiose by a yeast strain displaying β-glucosidase on the cell surface. Appl. Microbiol. Biotechnol. 2008, 79, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, A.; Bae, J.G.; Fukai, K.; Tokumoto, N.; Kuroda, K.; Ogawa, J.; Nakatani, M.; Shimizu, S.; Ueda, M. Effect of pretreatment of hydrothermally processed rice straw with laccase-displaying yeast on ethanol fermentation. Appl. Microbiol. Biotechnol. 2012, 94, 939–948. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.G.; Han, S.Y.; Wei, D.Z.; Lin, Y.; Wang, X.N. Functional display of Rhizomucor miehei lipase on surface of Saccharomyces cerevisiae with higher activity and its practical properties. J. Chem. Technol. Biotechnol. 2008, 83, 329–335. [Google Scholar] [CrossRef]
- Inaba, C.; Maekawa, K.; Morisaka, H.; Kuroda, K.; Ueda, M. Efficient synthesis of enantiomeric ethyl lactate by Candida antarctica lipase B (CALB)-displaying yeasts. Appl. Microbiol. Biotechnol. 2009, 83, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Ryckaert, S.; Martens, V.; Vusser, K.D.; Contreras, R. Development of a S. cerevisiae whole cell biocatalyst for in vitro sialylation of oligosaccharides. J. Biotechnol. 2005, 119, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Jia, B.; Zhao, H.; Xu, L.; Yan, Y. Preparation of a whole-cell biocatalyst of Aspergillus niger lipase and its practical properties. J. Agric. Food Chem. 2010, 58, 10426–10430. [Google Scholar] [CrossRef] [PubMed]
- Parthasarathy, R.; Subramanian, S.; Boder, E.T.; Discher, D.E. Post-translational regulation of expression and conformation of an immunoglobulin domain in yeast surface display. Biotechnol. Bioeng. 2006, 93, 159–168. [Google Scholar] [CrossRef]
- Van den Beucken, T.; Pieters, H.; Steukers, M.; van der Vaart, M.; Ladner, R.C.; Hoogenboom, H.R.; Hufton, S.E. Affinity maturation of Fab antibody fragments by fluorescent-activated cell sorting of yeast-displayed libraries. FEBS Lett. 2003, 546, 288–294. [Google Scholar] [CrossRef]
- Lin, Y.; Tsumuraya, T.; Wakabayashi, T.; Shiraga, S.; Fujii, I.; Kondo, A.; Ueda, M. Display of a functional hetero-oligomeric catalytic antibody on the yeast cell surface. Appl. Microbiol. Biotechnol. 2003, 62, 226–232. [Google Scholar] [CrossRef]
- Boder, E.T.; Bill, J.R.; Nields, A.W.; Marrack, P.C.; Kappler, J.W. Yeast surface display of a noncovalent MHC class II heterodimer complexed with antigenic peptide. Biotechnol. Bioeng. 2005, 92, 485–491. [Google Scholar] [CrossRef]
- Liu, W.; Zhao, H.; Jia, B.; Xu, L.; Yan, Y. Surface display of active lipase in Saccharomyces cerevisiae using Cwp2 as an anchor protein. Biotechnol. Lett. 2010, 32, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Ram, A.F.; van den Ende, H.; Klis, F.M. Green fluorescent protein-cell wall fusion proteins are covalently incorporated into the cell wall of Saccharomyces cerevisiae. FEMS Microbiol. Lett. 1998, 162, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Abe, H.; Shimma, Y.; Jigami, Y. In vitro oligosaccharide synthesis using intact yeast cells that display glycosyltransferases at the cell surface through cell-wall anchored protein Pir. Glycobiology 2003, 13, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Sumita, T.; Yoko-o, T.; Shimma, Y.; Jigami, Y. Comparison of cell wall localization among Pir family proteins and functional dissection of the region required for cell wall binding and bud scar recruitment of Pir1p. Eukaryot Cell 2005, 4, 1872–1881. [Google Scholar] [CrossRef] [PubMed]
- Andres, I.; Gallardo, O.; Parascandola, P.; Pastor, F.I.J.; Zueco, J. Use of the cell wall protein Pir4 as a fusion partner for the expression of Bacillus sp. BP-7 Xylanase A in Saccharomyces cerevisiae. Biotechnol. Bioeng. 2005, 89, 690–697. [Google Scholar] [CrossRef]
- Andres, I.; Rodriguez-Diaz, J.; Buesa, J.; Zueco, J. Yeast expression of the VP8* fragment of the rotavirus spike protein and its use as immunogen in mice. Biotechnol. Bioeng. 2006, 93, 89–98. [Google Scholar] [CrossRef]
- Shimma, Y.I.; Saito, F.; Oosawa, F.; Jigami, Y. Construction of a library of human glycosyltransferases immobilized in the cell wall of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2006, 72, 7003–7012. [Google Scholar] [CrossRef]
- Feldhaus, M.J.; Siegel, R.W.; Opresko, L.K.; Coleman, J.R.; Feldhous, J.M.; Yeung, Y.A.; Cochran, J.R.; Heinzelman, P.; Colby, D.; Swers, J.; et al. Flow-cytometric isolation of human antibodies from a nonimmune Saccharomyces cerevisiae surface display library. Nat. Biotechnol. 2003, 21, 163–170. [Google Scholar] [CrossRef]
- Idiris, A.; Tohda, H.; Kumagai, H.; Takegawa, K. Engineering of protein secretion in yeast: Strategies and impact on protein production. Appl. Microbiol. Biotechnol. 2010, 86, 403–417. [Google Scholar] [CrossRef]
- Murai, T.; Ueda, M.; Atomi, H.; Shibasaki, Y.; Kamasawa, N.; Osumi, M.; Kawaguchi, T.; Arai, M.; Tanaka, A. Genetic imobilization of cellulase on the cell surface of Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 1997, 48, 499–503. [Google Scholar] [CrossRef]
- Ota, M.; Sakuragi, H.; Morisaka, H.; Kuroda, K.; Miyake, H.; Tamaru, Y.; Ueda, M. Display of Clostridium cellulovorans xylose isomerase on the cell surface of Saccharomyces cerevisiae and its direct application to xylose fermentation. Biotechnol. Prog. 2013, 29, 346–351. [Google Scholar] [CrossRef] [PubMed]
- Kaya, M.; Ito, J.; Kotaka, A.; Matsumura, K.; Bando, H.; Sahara, H.; Ogino, C.; Shibasaki, S.; Kuroda, K.; Ueda, M.; et al. Isoflavone aglycones production from isoflavone glycosides by display of β-glucosidase from Aspergillus oryzae on yeast cell surface. Appl. Microbiol. Biotechnol. 2008, 79, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Izumi, T.; Piskula, M.K.; Osawa, S.; Obata, A.; Tobe, K.; Saito, M.; Kataoka, S.; Kubota, Y.; Kikuchi, M. Soy Isoflavone Aglycones Are Absorbed Faster and in Higher Amounts than Their Glucosides in Humans. J. Nutr. 2000, 130, 1695–1699. [Google Scholar] [CrossRef] [PubMed]
- Arai, Y.; Watanabe, S.; Kimira, M.; Shimoi, K.; Mochizuki, R.; Kinae, N. Dietary intakes of flavonols, flavones and isoflavones by Japanese women and the inverse correlation between quercetin intake and plasma LDL cholesterol concentration. J. Nutr. 2000, 130, 2243–2250. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, K.; Ueda, M.; Shibasaki, S.; Tanaka, A. Cell surface-engineered yeast with ability to bind, and self-aggregate in response to, copper ion. Appl. Microbiol. Biotechnol. 2002, 59, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, T.; Shimada, M.; Kuroda, K.; Ueda, M. Molecular design of yeast cell surface for adsorption and recovery of molybdenum, one of rare metals. Appl. Microbiol Biotechnol. 2010, 86, 641–648. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, K.; Nishitani, T.; Ueda, M. Specific adsorption of tungstate by cell surface display of the newly designed ModE mutant. Appl. Microbiol. Biotechnol. 2012, 96, 153–159. [Google Scholar] [CrossRef]
- Krauland, E.M.; Peelle, B.R.; Wittrup, K.D.; Belcher, A.M. Peptide tags for enhanced cellular and protein adhesion to single-crystalline sapphire. Biotechnol. Bioeng. 2007, 97, 1009–1020. [Google Scholar] [CrossRef]
- Boder, E.T.; Raeeszadeh-Sarmazdeh, M.; Price, J.V. Engineering antibodies by yeast display. Arch. Biochem. Biophys. 2012, 526, 99–106. [Google Scholar] [CrossRef]
- Tafakori, V.; Torktaz, I.; Doostmohammadi, M.; Ahmadian, G. Microbial cell surface display; its medical and environmental applications. Iran. J. Biotech. 2012, 10, 231–239. [Google Scholar]
- Yang, X.; Tang, H.; Song, M.; Shen, Y.; Hou, J.; Bao, X. Development of novel surface display platforms for anchoring heterologous proteins in Saccharomyces cerevisiae. Microb Cell Fact. 2019, 18, 85. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Wang, J.; Wang, S.; Shen, Y.; Petranovic, D.; Hou, J.; Bao, X. Efficient yeast surface-display of novel complex synthetic cellulosomes. Microb. Cell Fact. 2018, 17, 122. [Google Scholar] [CrossRef] [PubMed]
- Inokuma, K.; Hasunuma, T.; Kondo, A. Efficient yeast cell-surface display of exo- and endo-cellulase using the SED1 anchoring region and its original promoter. Biotechnol. Biofuels 2014, 7, 965–969. [Google Scholar] [CrossRef] [PubMed]
- Inokuma, K.; Bamba, T.; Ishii, J.; Ito, Y.; Hasunuma, T.; Kondo, A. Enhanced cell-surface display and secretory production of cellulolytic enzymes with Saccharomyces cerevisiae Sed1 signal peptide. Biotechnol. Bioeng. 2016, 113, 2358–2366. [Google Scholar] [CrossRef] [PubMed]
- Andreu, C.; del Olmo, M. Development of a new yeast surface display system based on Spi1 as an anchor protein. Appl. Microbiol. Biotechnol. 2017, 101, 287–299. [Google Scholar] [CrossRef] [PubMed]
- Puig, S.; Pérez-Ortín, J.E. Stress response and expression patterns in wine fermentations of yeast genes induced at the diauxic shift. Yeast 2000, 16, 139–148. [Google Scholar] [CrossRef]
- Gasch, A.P.; Spellman, P.T.; Kao, C.M.; Carmel-Harel, O.; Eisen, M.B.; Storz, G.; Botstein, D.; Brown, P.O. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 2000, 11, 4241–4257. [Google Scholar] [CrossRef]
- Aranda, A.; del Olmo, M. Exposure of Saccharomyces cerevisiae to acetaldehyde induces sulfur amino acid metabolism and polyamine transporter genes, which depend on Met4p and Haa1p transcription factors, respectively. Appl. Environ. Microbiol. 2004, 70, 1913–1922. [Google Scholar] [CrossRef]
- Cardona, F.; Del Olmo, M.; Aranda, A. Phylogenetic origin and transcriptional regulation at the post-diauxic phase of SPI1, in Saccharomyces cerevisiae. Cell Mol. Biol. Lett. 2012, 17, 393–407. [Google Scholar] [CrossRef]
- Teparić, R.; Didak, B.; Ščulac, E.; Mrša, V. Genetic immobilization of RNase Rny1p at the Saccharomyces cerevisiae cell surface. J. Gen. Appl. Microbiol. 2013, 59, 75–82. [Google Scholar] [CrossRef]
- Hossain, A.S.; Teparic, R.; Mrsa, V. Comparison of two models of surface display of xylose reductase in the Saccharomyces cerevisiae cell wall. Enzyme Microb. Technol. 2019, 123, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Salo, H.; Sievi, E.; Suntio, T.; Mecklin, M.; Mattila, P.; Renkonen, R.; Makarow, M. Co-expression of two mammalian glycosyltransferases in the yeast cell wall allows synthesis of sLex. FEMS Yeast Res. 2005, 5, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.S.; Walter, P. A novel mechanism for regulating activity of a transcription factor that controls the unfolded protein response. Cell 1996, 87, 391–404. [Google Scholar] [CrossRef]
- Teunissen, A.W.; Holub, E.; van der Hucht, J.; van den Berg, J.A.; Steensma, H.Y. Sequence of the open reading frame of the FLO1 gene from Saccharomyces cerevisiae. Yeast 1993, 9, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Breinig, F.; Schmitt, M.J. Spacer-elongated cell wall fusion proteins improve cell surface expression in the yeast Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 2002, 58, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, K.; Matsui, K.; Higuchi, S.; Kotaka, A.; Sahara, H.; Hata, Y.; Ueda, M. Enhancement of display efficiency in yeast display system by vector engineering and gene disruption. Appl. Microbiol. Biotechnol. 2009, 82, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Macreadie, I.G.; Horaitis, O.; Verkuylen, A.J.; Savin, K.W. Improved shuttle vectors for cloning and high-level Cu2+-mediated expression of foreign genes in yeast. Gene 1991, 104, 107–111. [Google Scholar] [CrossRef]
- Shimoi, H.; Kitagaki, H.; Ohmori, H.; Iimura, Y.; Ito, K. Sed1p Is a Major Cell Wall Protein of Saccharomyces cerevisiae in the Stationary Phase and Is Involved in Lytic Enzyme Resistance. J. Bacteriol. 1998, 180, 3381–3387. [Google Scholar]
- Wentz, A.E.; Shusta, E.V. A novel high-throughput screen reveals yeast genes that increase secretion of heterologous proteins. Appl. Environ. Microbiol. 2007, 73, 1189–1198. [Google Scholar] [CrossRef]
- Shusta, E.V.; Holler, P.D.; Kieke, M.C.; Kranz, D.M.; Wittrup, K.D. Directed evolution of a stable scaffold for T-cell receptor engineering. Nat. Biotechnol. 2000, 18, 754–759. [Google Scholar] [CrossRef]
- Tu, B.P.; Ho-Schleyer, S.C.; Travers, K.J.; Weissman, J.S. Biochemical basis of oxidative protein folding in the endoplasmic reticulum. Science 2000, 290, 1571–1574. [Google Scholar] [CrossRef] [PubMed]
- Frand, A.R.; Kaiser, C.A. The ERO1 gene of yeast is required for oxidation of protein dithiols in the endoplasmic reticulum. Mol. Cell 1998, 1, 161–170. [Google Scholar] [CrossRef]
- Mitsui, K.; Tsurugi, K. cDNA and deduced amino acid sequence of 38 kDa-type acidic ribosomal protein A0 from Saccharomyces cerevisiae. Nucleic Acids Res. 1988, 16, 3573. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, H.; Hashimoto, K.; Saijo, A.; Takada, Y.; Kondo, A.; Ueda, M.; Ooshima, H.; Tachibana, T.; Azuma, M. Cell wall structure suitable for surface display of proteins in Saccharomyces cerevisiae. Yeast 2014, 31, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Song, M.; He, Y.; Wang, J.; Wang, S.; Shen, Y.; Hou, J.; Bao, X. Engineering vesicle trafficking improves the extracellular activity and surface display efficiency of cellulases in Saccharomyces cerevisiae. Biotechnol. Biofuels 2017, 10, 53. [Google Scholar] [CrossRef]
- Demain, A.L.; Newcomb, M.; Wu, J.H.D. Cellulase, clostridia, and ethanol. Microbiol Mol. Biol. Rev. 2005, 69, 124–154. [Google Scholar] [CrossRef] [PubMed]
- Adams, J.J.; Pal, G.; Jia, Z.; Smith, S.P. Mechanism of bacterial cell-surface attachment revealed by the structure of cellulosomal type II cohesin–dockerin complex. Proc. Natl. Acad. Sci. USA 2006, 103, 305–310. [Google Scholar] [CrossRef]
- Goyal, G.; Tsai, S.L.; Madan, B.; Dasilva, N.A.; Chen, W. Simultaneous cell growth and ethanol production from cellulose by an engineered yeast consortium displaying a functional mini-cellulosome. Microb. Cell Fact. 2011, 10, 1–8. [Google Scholar] [CrossRef]
- Fan, L.H.; Zhang, Z.J.; Yu, X.Y.; Xue, Y.X.; Tan, T.W. Self-surface assembly of cellulosomes with two miniscaffoldins on Saccharomyces cerevisiae for cellulosic ethanol production. Proc. Natl. Acad. Sci. USA 2012, 109, 13260–13265. [Google Scholar] [CrossRef]
- Liang, Y.Y.; Si, T.; EeLui, A.; Zhao, H.M. Engineered pentafunctional minicellulosome for simultaneous saccharification and ethanol fermentation in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2014, 80, 6677–6684. [Google Scholar] [CrossRef]
- Mogelsvang, S.; Gomez-Ospina, N.; Soderholm, J.; Glick, B.S.; Staehelin, L.A. Tomographic Evidence for Continuous Turnover of Golgi Cisternae in Pichia pastoris. Mol. Biol. Cell 2003, 14, 2277–2291. [Google Scholar] [CrossRef] [PubMed]
- Cereghino, J.L.; Cregg, J.M. Heterologous protein expression in the methylotrophic yeast Pichia pastoris. FEMS Microbiol. Rev. 2000, 24, 45–66. [Google Scholar] [CrossRef] [PubMed]
- Macauley-Patrick, S.; Fazenda, M.L.; McNeil, B.; Harvey, L.M. Heterologous protein production using the Pichia pastoris expression system. Yeast 2005, 22, 249–270. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, L.; Chen, M.; Qi, Q.; Wang, P.G. Construction of a novel system for cell surface display of heterologous proteins on Pichia pastoris. Biotechnol. Lett. 2007, 29, 1561–1566. [Google Scholar] [CrossRef] [PubMed]
- Su, G.D.; Huang, D.F.; Han, S.Y.; Zheng, S.P.; Lin, Y. Display of Candida antarctica lipase B on Pichia pastoris and its application to flavor ester synthesis. Appl. Microbiol. Biotechnol. 2010, 86, 1493–1501. [Google Scholar] [CrossRef] [PubMed]
- Khasa, Y.P.; Conrad, S.; Sengul, M.; Plautz, S.; Meagher, M.M.; Inan, M. Isolation of Pichia pastoris PIR genes and their utilization for cell surface display and recombinant protein secretion. Yeast 2011, 28, 213–226. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Shi, H.; Ding, H.; Wang, L.; Zhang, Y.; Li, X.; Wang, F. Cell surface display and characterization of Rhizopus oryzae lipase in Pichia pastoris using Sed1p as an anchor protein. Curr. Microbiol. 2015, 71, 150–155. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liang, S.; Zhou, X.; Jin, Z.; Jiang, F.; Han, S.; Zhen, S.; Lin, Y. Screening for glycosylphosphatidylinositol-modified cell wall proteins in Pichia pastoris and their recombinant expression on the cell surface. Appl. Environ. Microbiol. 2013, 79, 5519–5526. [Google Scholar] [CrossRef]
- Moura, M.V.; da Silva, G.P.; Machado, A.C.; Torres, F.A.; Freire, D.M.; Almeida, R.V. Displaying lipase B from Candida antarctica in Pichia pastoris using the yeast surface display approach: Prospection of a new anchor and characterization of the whole cell biocatalyst. PLoS ONE 2015, 10, e0141454. [Google Scholar] [CrossRef]
- Mergler, M.; Wolf, K.; Zimmermann, M. Development of a bisphenol A-adsorbing yeast by surface display of the Kluyveromyces yellow enzyme on Pichia pastoris. Appl. Microbiol. Biotechnol. 2004, 63, 418–421. [Google Scholar] [CrossRef]
- Jacobs, P.P.; Ryckaert, S.; Geysens, S.; De Vusser, K.; Callewaert, N.; Contreras, R. Pichia surface display: Display of proteins on the surface of glycoengineered Pichia pastoris strains. Biotechnol. Lett. 2008, 30, 2173–2181. [Google Scholar] [CrossRef] [PubMed]
- Shaheen, H.H.; Prinz, B.; Chen, M.; Pavoor, T.; Lin, S.; Houston-Cummings, N.R.; Moore, R.; Stadheim, T.A.; Zha, D. A Dual-Mode Surface Display System for the Maturation and Production of Monoclonal Antibodies in Glyco-Engineered Pichia pastoris. PLoS ONE 2013, 8, e70190. [Google Scholar] [CrossRef] [PubMed]
- Wasilenko, J.L.; Sarmento, L.; Spatz, S.; Pantin-Jackwood, M. Cell Surface Display of Highly Pathogenic Avian Influenza Virus Hemagglutinin on the Surface of Pichia pastoris Cells Using a-Agglutinin for Production of Oral Vaccines. Biotechnol. Prog. 2010, 26, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.F.; Lin, Y.; Zhang, J.H.; Zheng, S.P.; Ye, Y.R.; Li, X.X.; Han, S.Y. Double Candida antarctica lipase B co-display on Pichia pastoris cell surface based on a self-processing foot-and-mouth disease virus 2A peptide. Appl. Microbiol. Biotechnol. 2012, 96, 1539–1555. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Houston-Cummings, N.R.; Prinz, B.; Moore, R.; Bobrowicz, B.; Davidson, R.C.; Wildt, S.; Stadheim, T.A.; Zha, D. A novel fragment of antigen binding (Fab) surface display platform using glycoengineered Pichia pastoris. J. Immunol. Methods 2012, 375, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Lv, X.; Wang, X.; Wang, J.; Wang, R.; Wang, T. Cell-Surface Displayed Expression of Trehalose Synthase from Pseudomonas putida ATCC 47054 in Pichia Pastoris Using Pir1p as an Anchor Protein. Front. Microbiol. 2017, 8, 2583. [Google Scholar] [CrossRef] [PubMed]
- Jo, J.H.; Im, E.M.; Kim, S.H.; Lee, H.H. Surface display of human lactoferrin using a glycosylphosphatidylinositol-anchored protein of Saccharomyces cerevisiae in Pichia pastoris. Biotechnol. Lett. 2011, 33, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Herrera, J.; Sentandreu, R. Different effectors of dimorphism in Yarrowia lipolytica. Arch. Microbiol. 2002, 178, 477–483. [Google Scholar] [CrossRef]
- Fickers, P.; Benetti, P.H.; Waché, Y.; Marty, A.; Mauersberger, S.; Smit, M.S.; Nicaud, J.M. Hydrophobic substrate utilisation by the yeast Yarrowia lipolytica, and its potential applications. FEMS Yeast Res. 2005, 5, 527–543. [Google Scholar] [CrossRef]
- Rodrigues, G.; Pais, C. The influence of acetic and other weak carboxylic acids on growth and cellular death of the yeast Yarrowia lipolytica. Food Technol. Biotechnol. 1997, 38, 27–32. [Google Scholar]
- Barth, G.; Gaillardin, C. Physiology and genetics of the dimorphic fungus Yarrowia lipolytica. FEMS Microbiol. Rev. 1997, 19, 219–237. [Google Scholar] [CrossRef] [PubMed]
- Madzak, C.; Gaillardin, C.; Beckerich, J.M. Heterologous protein expression and secretion in the non-conventional yeast Yarrowia lipolytica: A review. J. Biotechnol. 2004, 109, 63–81. [Google Scholar] [CrossRef] [PubMed]
- Dominguez, A.; Ferminan, E.; Sanchez, M.; Gonzalez, F.J.; Perez-Campo, F.M.; Garcia, S.; Herrero, A.B.; San Vicente, A.; Cabello, J.; Prado, M.; et al. Non-conventional yeasts as hosts for heterologous protein production. Int. Microbiol. 1998, 1, 131–142. [Google Scholar] [PubMed]
- Yue, L.; Chi, Z.; Wang, L.; Liu, J.; Madzak, C.; Li, J.; Wang, X. Construction of a new plasmid for surface display on cells of Yarrowia lipolytica. J. Microbiol. Methods 2008, 72, 116–123. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.Y.; Chi, Z.; Liu, G.L.; Wang, F.; Madzak, C.; Chi, Z.M. Inulin hydrolysis and citric acid production from inulin using the surface-engineered Yarrowia lipolytica displaying inulinase. Metab. Eng. 2010, 12, 469–476. [Google Scholar] [CrossRef] [PubMed]
- Ni, X.; Yue, L.; Chi, Z.; Li, J.; Wang, X.; Madzak, C. Alkaline protease gene cloning from the marine yeast Aureobasidium pullulans HN2-3 and the protease surface display on Yarrowia lipolytica for bioactive peptide production. Mar. Biotechnol. 2009, 11, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.J.; Madzak, C.; Li, H.J.; Chi, Z.M.; Li, J. Surface display of acid protease on the cells of Yarrowia lipolytica for milk clotting. Appl. Microbiol. Biotechnol. 2010, 87, 669–677. [Google Scholar] [CrossRef]
- Zheng, Y.; Wang, Z.; Ji, X.; Sheng, J. Display of a sucrose isomerase on the cell surface of Yarrowia lipolytica for synthesis of isomaltulose from sugar cane by-products. 3 Biotech 2019, 9, 179. [Google Scholar] [CrossRef]
- Liu, G.; Yue, L.; Chi, Z.; Yu, W.; Chi, Z.; Madzak, C. The surface display of the alginate lyase on the cells of Yarrowia lipolytica for hydrolysis of alginate. Mar. Biotechnol. 2009, 11, 619–626. [Google Scholar] [CrossRef]
- Yuzbasheva, E.Y.; Yuzbashev, T.V.; Konstantinova, T.K.; Laptev, I.A.; Sineoky, S.P. Capacity of the N- and C-domains of the cell wall protein Flo1p homologue in yeast Yarrowia lipolytica to display lipase Lip2 on the cell surface. Biotechnologiya 2010, 6, 23–36. [Google Scholar]
- Yang, X.S.; Jiang, Z.B.; Song, H.T.; Jiang, S.J.; Madzak, C.; Ma, L.X. Cell-surface display of the active mannanase in Yarrowia lipolytica with a novel surface-display system. Biotechnol. Appl. Biochem. 2009, 54, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Yuzbasheva, E.Y.; Yuzbashev, T.V.; Laptev, I.A.; Konstantinova, T.K.; Sineoky, S.P. Efficient cell surface display of Lip2 lipase using C-domains of glycosylphosphatidylinositol-anchored cell wall proteins of Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2011, 91, 645–654. [Google Scholar] [CrossRef] [PubMed]
- Tanino, T.; Aoki, T.; Chung, W.Y.; Watanabe, Y.; Ogino, C.; Fukuda, H.; Kondo, A. Improvement of a Candida antarctica lipase B-displaying yeast whole-cell biocatalyst and its application to the polyester synthesis reaction. Appl. Microbiol. Biotechnol. 2009, 82, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.S.; Pan, X.X.; Jia, B.; Zhao, H.Y.; Xu, L.; Liu, Y.; Yan, Y.J. Surface display of active lipases Lip7 and Lip8 from Yarrowia lipolytica on Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 2010, 88, 885–891. [Google Scholar] [CrossRef] [PubMed]
- Su, G.D.; Zhang, X.; Lin, Y. Surface display of active lipase in Pichia pastoris using Sed1 as an anchor protein. Biotechnol. Lett. 2010, 32, 1131–1136. [Google Scholar] [CrossRef] [PubMed]
- Alloue, W.A.; Destain, J.; El Medjoub, T.; Ghalfi, H.; Kabran, P.; Thonart, P. Comparison of Yarrowia lipolytica lipase immobilization yield of entrapment, adsorption, and covalent bond techniques. Appl. Biochem. Biotechnol. 2008, 150, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Yuzbasheva, E.Y.; Yuzbashev, T.V.; Perkovskaya, N.I.; Mostova, E.B.; Vybornaya, T.V.; Sukhozhenko, A.V.; Toropygin, I.Y.; Sineoky, S.P. Cell surface display of Yarrowia lipolytica lipase Lip2p using the cell wall protein YlPir1p, its characterization, and application as a whole-cell biocatalyst. Appl. Biochem. Biotechnol. 2015, 175, 3888–3900. [Google Scholar] [CrossRef] [PubMed]
- Duquesne, S.; Bozonnet, S.; Bordes, F.; Dumon, C.; Nicaud, J.M.; Marty, A. Construction of a highly active xylanase displaying oleaginous yeast: Comparison of anchoring systems. PLoS ONE 2014, 9, e95128. [Google Scholar] [CrossRef] [PubMed]
- Middelhoven, W.J.; Hoogkamer-Te Niet, M.V.; Kreger-Van Rij, N.J.W. Trichosporon adeninovorans sp. nov., a yeast species utilizing adenine, xanthine, uric acid, putrescine and primary n-alkylamines as the sole source of carbon, nitrogen and energy. Antonie Van Leeuwenhoek 1984, 50, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Middelhoven, W.J.; de Jong, I.M.; de Winter, M. Arxula adeninivorans, a yeast assimilating many nitrogenous and aromatic compounds. Antonie Van Leeuwenhoek 1991, 59, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Kurtzman, C.; Fell, J.W. The Yeasts–A Taxonomic Study, 4th ed.; Elsevier Science: Amsterdam, The Netherlands, 1998. [Google Scholar] [CrossRef]
- Yang, X.; Wartmann, T.; Stoltenburg, R.; Kunze, G. Halotolerance of the yeast Arxula adeninivorans LS3. Antonie Van Leeuwenhoek 2000, 77, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Wartmann, T.; Erdman, J.; Kunze, I.; Kunze, K. Morphology-related effects on gene expression and protein accumulation of the yeast Arxula adeninivorans LS3. Arch. Microbiol. 2000, 173, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Böer, E.; Steinborn, G.; Matros, A.; Mock, H.P.; Gellissen, G.; Kunze, G. Production of interleukin-6 in Arxula adeninivorans, Hansenula polymorpha and Saccharomyces cerevisiae by applying the wide-range yeast vector (CoMedTM) system to simultaneous comparative assessment. FEMS Yeast Res. 2007, 7, 1181–1187. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wartmann, T.; Stephan, U.W.; Bube, I.; Böer, E.; Melzer, M.; Manteuffel, R.; Stoltenburg, R.; Guengerich, L.; Gellissen, G.; Kunze, G. Post-translational modifications of the AFET3 gene product—A component of the iron transport system in budding cells and mycelia of the yeast Arxula adeninivorans. Yeast 2002, 19, 849–862. [Google Scholar] [CrossRef] [PubMed]
- Kunze, G.; Gaillardin, C.; Czernicka, M.; Durrens, P.; Martin, T.; Erik, B.; Toni, G.; Cruz, J.A.; Talla, E.; Marck, C.; et al. The complete genome of Blastobotrys (Arxula) adeninivorans LS3: A yeast of biotechnological interest. Biotechnol. Biofuels 2014, 7, 66. [Google Scholar] [CrossRef] [PubMed]
- Terentiev, Y.; Pico, A.H.; Böer, E.; Wartmann, T.; Klabunde, J.; Breuer, U.; Babel, W.; Suckow, M.; Gellissen, G.; Kunze, K. A wide range integrative yeast expression vector system based on Arxula adeninivorans derived elements. J. Ind. Microbiol. Biotechnol. 2004, 31, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Böer, E.; Bode, R.; Mock, H.P.; Piontek, M.; Kunze, G. Atan1p: An extracellular tannase from the dimorphic yeast Arxula adeninivorans: Molecular cloning of the ATAN1 gene and characterization of the recombinant enzyme. Yeast 2009, 26, 323–337. [Google Scholar] [CrossRef]
- Böer, E.; Piontek, M.; Kunze, G. Xplor® 2—An optimized transformation/expression system for recombinant protein production in the yeast Arxula adeninivorans. Appl. Microbiol. Biotechnol. 2009, 84, 583–594. [Google Scholar] [CrossRef]
- Wartmann, T.; Bellebna, C.; Böer, E.; Bartelsen, O.; Gellissen, G.; Kunze, G. The constitutive AHSB4 promoter: A novel component of the Arxula adeninivorans-based expression platform. Appl. Microb. Biotechnol. 2003, 62, 528–535. [Google Scholar] [CrossRef]
- Wartmann, T.; Stoltenburg, R.; Böer, E.; Sieber, H.; Bartelsen, O.; Gellissen, G.; Kunze, G. The ALEU2 gene: A new component for an Arxula adeninivorans-based expression platform. FEMS Yeast Res. 2003, 3, 223–232. [Google Scholar] [CrossRef]
- Giga-Hama, Y. Fission Yeast Schizosaccharomyces pombe: An Attractive Host for Heterologous Protein Production. In Foreign Gene Expression in Fission Yeast: Schizosaccharomyces pombe; Giga-Hama, Y., Kumagai, H., Eds.; Biotechnology Intelligence Unit. Springer: Berlin/Heidelberg, Germany, 1997. [Google Scholar] [CrossRef]
- Takegawa, K.; Tohda, H.; Sasaki, M.; Idiris, A.; Ohashi, T.; Mukaiyama, H.; Giga-Hama, Y.; Kumagai, H. Production of heterologous proteins using the fission-yeast (Schizosaccharomyces pombe) expression system. Biotechnol. Appl. Biochem. 2009, 53, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Lieberman, H.B. Schizosaccharomyces pombe: A model for molecular studies of eukaryotic genes. DNA Cell Biol. 1995, 14, 359–371. [Google Scholar] [CrossRef] [PubMed]
- Sander, P.; Grunewald, S.; Reilander, H.; Michel, H. Expression of the human D2S dopamine receptor in the yeasts Saccharomyces cerevisiae and Schizosaccharomyces pombe: A comparative study. FEBS Lett. 1994, 344, 41–46. [Google Scholar] [CrossRef]
- Idiris, A.; Tohda, H.; Bi, K.W.; Isoai, A.; Kumagai, H.; Giga-Hama, Y. Enhanced productivity of protease-sensitive heterologous proteins by disruption of multiple protease genes in the fission yeast Schizosaccharomyces pombe. Appl. Microbiol. Biotechnol. 2006, 73, 404–420. [Google Scholar] [CrossRef] [PubMed]
- Mukaiyama, H.; Tohda, H.; Takegawa, K. Overexpression of protein disulfide isomerases enhances secretion of recombinant human transferrin in Schizosaccharomyces pombe. Appl. Microbiol. Biotechnol. 2010, 86, 1135–1143. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Matsumoto, S.; Yamada, M.; Yamada, R.; Matsuda, F.; Kondo, A. Display of active beta-glucosidase on the surface of Schizosaccharomyces pombe cells using novel anchor proteins. Appl. Microbiol. Biotechnol. 2013, 97, 4343–4352. [Google Scholar] [CrossRef]
- Barnett, J.A.; Payne, R.W.; Yarrow, D. Yeasts: Characteristics and Identification, 3rd ed.; Cambridge University Press: Cambridge, UK, 2000. [Google Scholar] [CrossRef]
- Cerniglia, C.E.; Crow, S.A. Metabolism of aromatic hydrocarbons by yeasts. Arch. Microbiol. 1981, 129, 9–13. [Google Scholar] [CrossRef]
- Yadav, J.S.; Loper, J.C. Multiple P450alk (cytochrome P450 alkane hydroxylase) genes from the halotolerant yeast Debaryomyces hansenii. Gene 1999, 226, 139–146. [Google Scholar] [CrossRef]
- Hooley, P.; Fincham, D.A.; Whithead, M.P.; Clipson, N.J.W. Fungal osmotolerance. Adv. Appl. Microbiol. 2003, 53, 177–211. [Google Scholar]
- Singh, D.; Nigam, P.; Banat, I.M.; Marchant, R.; McHale, A.P. Ethanol production at elevated temperatures and alcohol concentrations. Part II. Use of Kluyveromyces marxianus IMB3. World J. Microbiol. Biotechnol. 1998, 14, 823–834. [Google Scholar] [CrossRef]
- Yanase, S.; Hasunuma, T.; Yamada, R.; Tanaka, T.; Ogino, C.; Fukuda, H.; Kondo, A. Direct ethanol production from cellulosic materials at high temperature using the thermotolerant yeast Kluyveromyces marxianus displaying cellulolytic enzymes. Appl. Microbiol. Biotechnol. 2010, 88, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Galan, C.; Berenguer-Murcia, A.; Fernandez-Lafuente, R.; Rodrigues, R.C. Potential of different enzyme immobilization strategies to improve enzyme performance. Adv. Synth. Catal. 2011, 353, 2885–2904. [Google Scholar] [CrossRef]
- Sheldon, R.A.; Van Pelt, S. Enzyme immobilisation in biocatalysis: Why, what and how. Chem. Soc. Rev. 2013, 42, 6223–6235. [Google Scholar] [CrossRef] [PubMed]
- Zdarta, J.; Meyer, A.S.; Jesionowski, T.; Pinelo, M. A general overview of support materials for enzyme immobilization: Characteristics, properties, practical utility. Catalysts 2018, 8, 92. [Google Scholar] [CrossRef]
- Andler, S.M.; Goddard, J.M. Transforming food waste: How immobilized enzymes can valorize waste streams into revenue streams. NPJ Sci. Food 2018, 2, 19. [Google Scholar] [CrossRef] [PubMed]
- Velasco-Lozano, S.; López-Gallego, F.; Mateos-Díaz, J.C.; Favela-Torres, E. Cross-linked enzyme aggregates (CLEA) in enzyme improvement—A review. Biocatalysis 2015, 1, 166–177. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lozančić, M.; Sk. Hossain, A.; Mrša, V.; Teparić, R. Surface Display—An Alternative to Classic Enzyme Immobilization. Catalysts 2019, 9, 728. https://doi.org/10.3390/catal9090728
Lozančić M, Sk. Hossain A, Mrša V, Teparić R. Surface Display—An Alternative to Classic Enzyme Immobilization. Catalysts. 2019; 9(9):728. https://doi.org/10.3390/catal9090728
Chicago/Turabian StyleLozančić, Mateja, Amir Sk. Hossain, Vladimir Mrša, and Renata Teparić. 2019. "Surface Display—An Alternative to Classic Enzyme Immobilization" Catalysts 9, no. 9: 728. https://doi.org/10.3390/catal9090728
APA StyleLozančić, M., Sk. Hossain, A., Mrša, V., & Teparić, R. (2019). Surface Display—An Alternative to Classic Enzyme Immobilization. Catalysts, 9(9), 728. https://doi.org/10.3390/catal9090728




