Microbial Reduction of Cholesterol to Coprostanol: An Old Concept and New Insights
Abstract
:1. Introduction
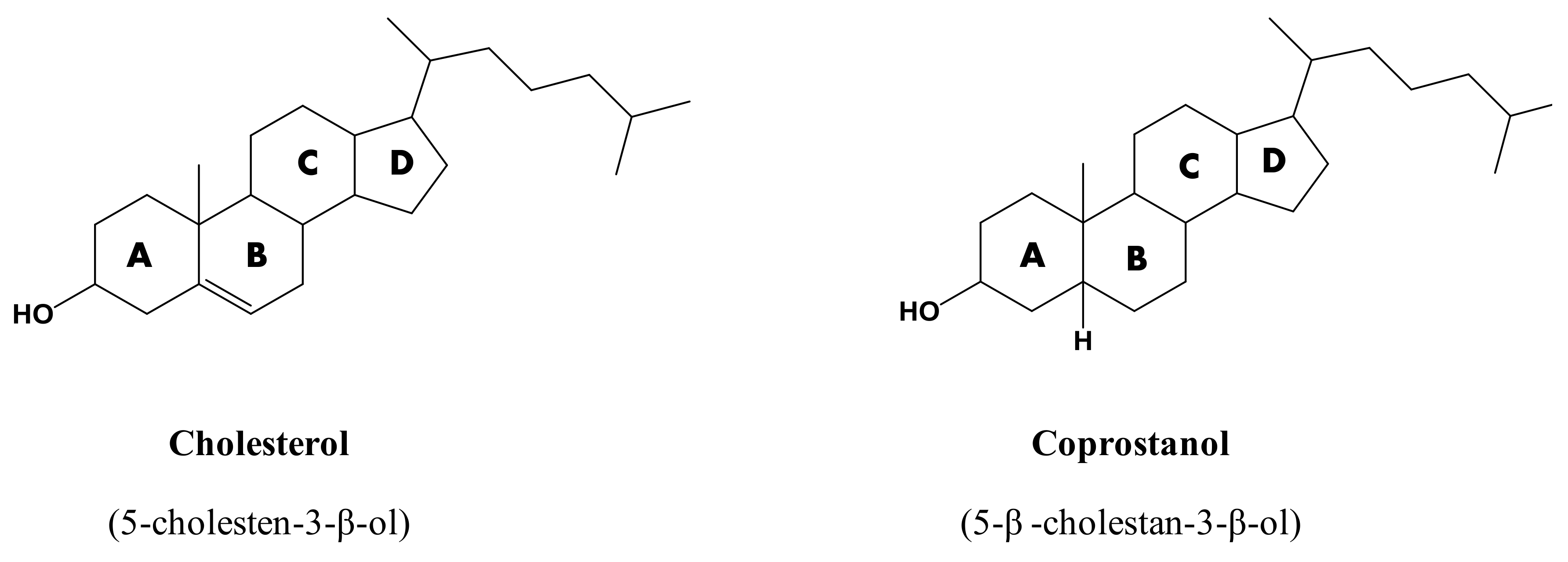
2. Cholesterol versus Coprostanol
2.1. Chemical Behaviors
2.2. Intestinal Uptake
3. Microbial Conversion of Cholesterol to Coprostanol
3.1. Coprostanoligenic Bacteria
3.2. Two Patterns Cepending on Gut Microbiota
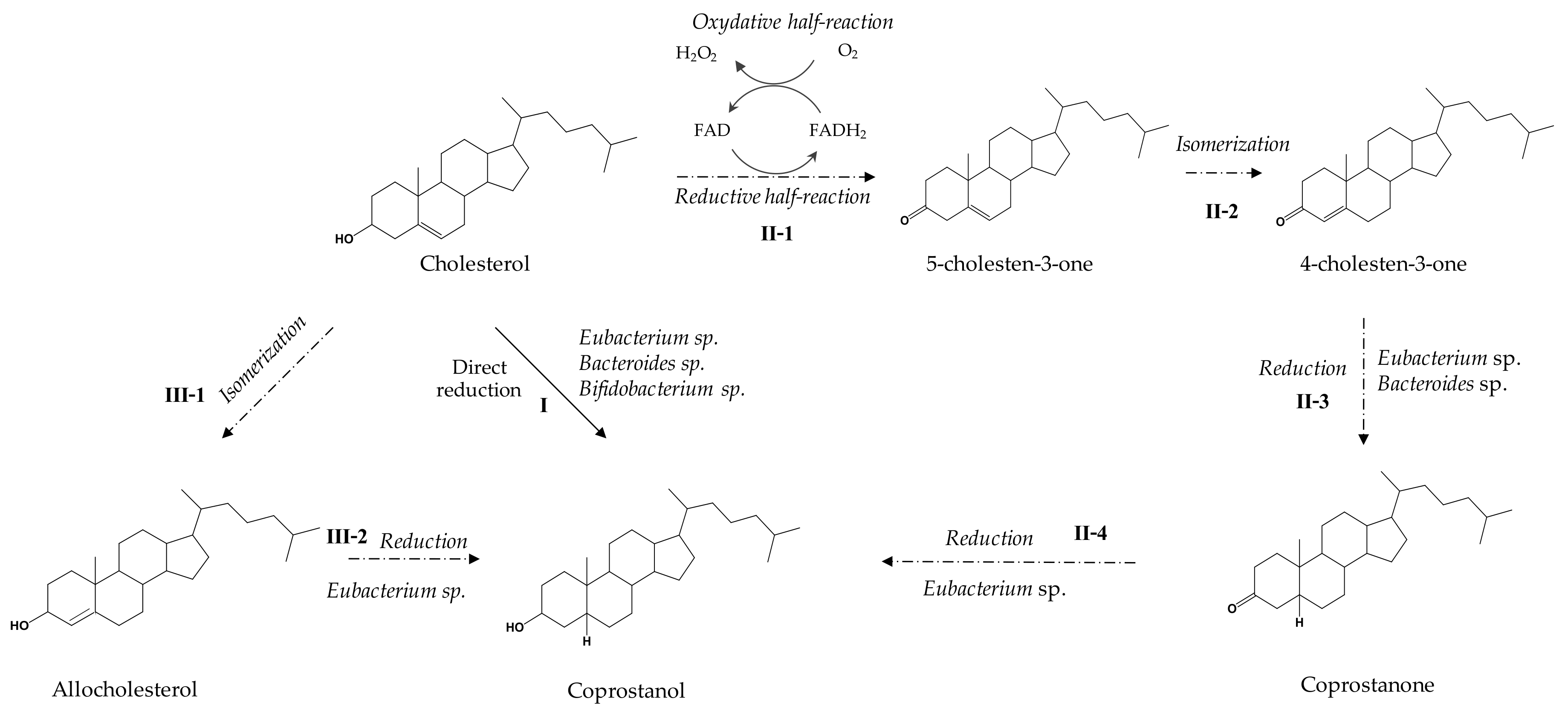
3.3. Metabolic Pathways
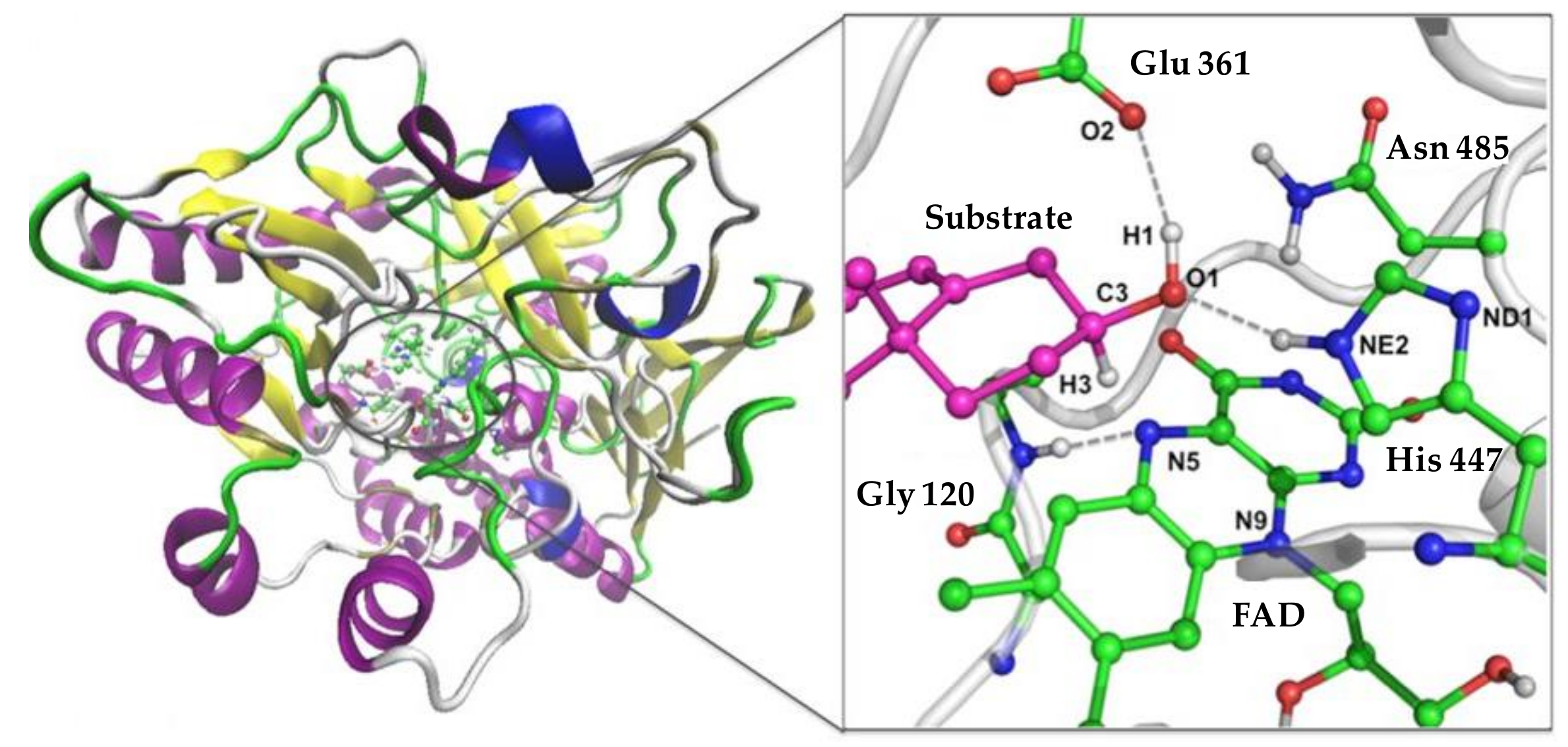
4. Towards a Better Understanding of the Coprostanol Production Pathway
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Rosamond, W.D.; Sampson, K.A.; Satou, G.M.; Shah, S.H.; Spartano, N.L.; et al. Heart Disease and Stroke Statistics—2018 Update: A Report from the American Heart Association. Circulation 2018, 137, e67–e492. [Google Scholar] [CrossRef] [PubMed]
- Krobot, K.J.; Yin, D.D.; Alemao, E.; Steinhagen-Thiessen, E. Realworld effectiveness of lipid-lowering therapy in male and female outpatients with CHD: Relation to pre-treatment LDL-cholesterol, pre-treatment CHD risk and other factors. Eur. J. Cardiovasc. Prev. Rehabil. 2005, 1, 37–450. [Google Scholar] [CrossRef]
- Groen, A.K.; Bloks, V.W.; Verkade, H.; Kuipers, F. Cross-talk between liver and intestine in control of cholesterol and energy homeostasis. Mol. Aspects Med. 2014, 37, 77–88. [Google Scholar] [CrossRef]
- Lee, S.D.; Gershkovich, P.; Darlington, J.W.; Wasan, K.M. Inhibition of cholesterol absorption: Targeting the intestine. Pharm. Res. 2012, 29, 3235–3250. [Google Scholar] [CrossRef] [PubMed]
- Chugh, A.; Ray, A.; Gupta, J.B. Squalene epoxidase as hypocholesterolemic drug target revisited. Prog. Lipid Res. 2003, 42, 37–50. [Google Scholar] [CrossRef]
- Goldstein, J.L.; Brown, M.S. A century of cholesterol and coronaries: From plaques to genes to statins. Cell 2015, 161, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Hou, R.; Goldberg, A.C. Lowering low-density lipoprotein cholesterol: Statins, ezetimibe, bile acid sequestrants, and combinations: Comparative efficacy and safety. Endocrinol. Metab. Clin. N. Am. 2009, 38, 79–97. [Google Scholar] [CrossRef]
- Gérard, P. Metabolism of cholesterol and bile acids by the gut microbiota. Pathogens 2014, 3, 14–24. [Google Scholar] [CrossRef]
- Bhattacharyya, A.K. Differences in uptake and esterification of saturated analogues of cholesterol by rat small intestine. Am. J. Physiol. 1986, 251, 495–500. [Google Scholar] [CrossRef]
- Sekimoto, H.; Shimada, O.; Makanishi, M.; Nakano, T.; Katayama, O. Interrelationship between serum and fecal sterols. Jpn. J. Med. 1983, 22, 14–20. [Google Scholar] [CrossRef]
- Reddy, B.S.; Martin, C.W.; Wynder, E.L. Fecal bile acids and cholesterol metabolites of patients with ulcerative colitis, a high-risk group for development of colon cancer. Cancer Res. 1977, 37, 1697–1701. [Google Scholar] [PubMed]
- Nomura, A.M.; Wilkins, T.D.; Kamiyama, S.; Heilbrun, L.K.; Shimada, A.; Stemmermann, G.N.; Mower, H.F. Fecal neutral steroids in two Japanese populations with different colon cancer risks. Cancer Res. 1983, 43, 1910–1913. [Google Scholar] [PubMed]
- Dam, H. Historical introduction to cholesterol. In Chemistry, Biochemistry and Pathology; Cooked, R.P., Ed.; Academic Press: New York, NY, USA, 1958; pp. 1–14. [Google Scholar]
- Liscum, L.; Underwood, K.W. Intracellular cholesterol transport and compartmentation. J. Biol. Chem. 1995, 270, 15443–15446. [Google Scholar] [CrossRef] [PubMed]
- Simons, K.; Ikonene, E. How cells handle cholesterol. Science 2000, 290, 1721–1725. [Google Scholar] [CrossRef] [PubMed]
- Sheng, R.; Chen, Y.; Yung Gee, H.; Stec, E.; Melowic, H.R.; Blatner, N.R.; Tun, M.P.; Kim, Y.; Källberg, M.; Fujiwara, T.K.; et al. Cholesterol modulates cell signaling and protein networking by specifically interacting with PDZ domain-containing scaffold proteins. Nat. Commun. 2012, 3, 1249. [Google Scholar] [CrossRef] [PubMed]
- Ohvo-Rekilä, H.; Ramstedt, B.; Leppimäki, P.; Slotte, J.P. Cholesterol interactions with phospholipids in membranes. Prog. Lipid Res. 2002, 41, 66–97. [Google Scholar] [CrossRef]
- Róg, T.; Pasenkiewicz-Gierula, M.; Vattulainen, I.; Karttunen, M. Ordering effects of cholesterol and its analogues. Biochim. Biophys. Acta 2009, 1788, 97–121. [Google Scholar] [CrossRef]
- Do, T.Q.; Moshkani, S.; Castillo, P.; Anunta, S.; Pogosyan, A.; Cheung, A.; Marbois, B.; Faull, K.F.; Ernst, W.; Chiang, S.M.; et al. Lipids including cholesteryl linoleate and cholesteryl arachidonate contribute to the inherent antibacterial activity of human nasal fluid. J. Immunol. 2008, 181, 4177–4187. [Google Scholar] [CrossRef]
- Tiangang, L.; Chiang, Y.L. Regulation of bile acid and cholesterol metabolism by PPARs. PPAR Res. 2009, 2009, 501739. [Google Scholar]
- Walker, R.W.; Wun, C.; Litsky, W. Coprostanol as indicator of faecal pollution. Crit. Rev. Env. Control 1982, 12, 91–112. [Google Scholar] [CrossRef]
- Singley, J.E.; Kirchmer, C.J.; Miura, R. Analysis of Coprostanol, an Indicator of Fecal Contamination; Environmental Protection Agency, Office of Research and Development: Washington, DC, USA, 1974; pp. 4–8.
- Christie, W.W. Lipid Analysis: Isolation, Separation, Identification and Structural Analysis of Lipids, 2nd ed.; Pergamon Press: Oxford, UK, 1982. [Google Scholar]
- Saad, H.Y.; Higuchi, W.I. Water solubility of cholesterol. J. Pharm. Sci. 1965, 54, 1205–1206. [Google Scholar] [CrossRef] [PubMed]
- Kanazawa, A.; Teshima, S.I. The occurrence of coprostanol, an indicator of fecal pollution in seawater and sediments. Oceanol. Acta 1978, 1, 39–44. [Google Scholar]
- Freier, T.A. Isolation and Characterization of Unique Cholesterol-Reducing Anaerobes. Ph.D. Thesis, Iowa State University, Ames, IA, USA, 1991. [Google Scholar]
- Li, L.; Buhman, K.K.; Hartman, P.A.; Beitz, D.C. Hypocholesterolemic effect of Eubacterium coprostanoligenes ATCC 51222 in rabbits. Lett. Appl. Microbiol. 1995, 20, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Batt, S.M.; Wannemuehler, M.; Dispirito, A.; Beitz, D.C. Effect of feeding of a cholesterol-reducing bacterium, Eubacterium coprostanoligenes, to germ-free mice. Lab. Anim. Sci. 1998, 48, 253–255. [Google Scholar] [PubMed]
- Iqbal, J.; Hussain, M.M. Intestinal lipid absorption. Am. J. Physiol. Endocrinol. Metab. 2009, 296, E1183–E11894. [Google Scholar] [CrossRef] [PubMed]
- Swell, L.; Troutec, J.; Hopper, J.R.; Fieldh, J.; Treadwell, C.D. Mechanism of cholesterol absorption. II. Changes in free and esterified cholesterol pools of mucosa after feeding cholesterol-4-C14. J. Biol. Chem. 1958, 233, 49–53. [Google Scholar] [PubMed]
- Kellogg, T.F. On the site of the microbiological reduction of cholesterol to coprostanol in the rat. Lipids 1973, 8, 658–659. [Google Scholar] [CrossRef]
- Setty, C.S.; Ivy, A.C. Intestinal absorption of coprostanol (coprosterol) in the rat. Am. J. Physiol. 1960, 199, 1008–1010. [Google Scholar] [CrossRef]
- Wells, W.; Anderson, S.A.; Ma, Q. Lactose diets and cholesterol metabolism. I. Cholesterol absorption, coprostanol formation and bile acid excretion in the rat. J. Nutr. 1960, 71, 405–410. [Google Scholar] [CrossRef]
- Rosenfeld, R.S.; Zumoff, B.; Hellman, L. Metabolism of coprostanol-C14 and cholestanol-4-C14 in man. J. Lipid Res. 1963, 4, 337–340. [Google Scholar]
- Schoenheimer, R. New contributions in sterol metabolism. Science 1931, 74, 579–584. [Google Scholar] [CrossRef]
- Koppel, N.; Maini Rekdal, V.; Balskus, E.P. Chemical transformation of xenobiotics by the human gut microbiota. Science 2017, 356, eaag2770. [Google Scholar] [CrossRef] [PubMed]
- Kellogg, T.F. Steroid balance and tissue cholesterol accumulation in germfree and conventional rats fed diets containing saturated and polyunsaturated fats. J. Lipid Res. 1974, 15, 574–579. [Google Scholar]
- Eyssen, H.J.; Parmentier, G.G.; Compernolle, F.C.; De Pauw, G.; Piessens-Denef, M. Biohydrogenation of sterols by Eubacterium ATCC 21408 nova species. Eur. J. Biochem. 1973, 36, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Brinkley, A.W.; Gottesman, A.R.; Mott, G.E. Growth of cholesterol-reducing Eubacterium on cholesterol-brain agar. Appl. Environ. Microbiol. 1980, 40, 1130–1132. [Google Scholar] [PubMed]
- Brinkley, A.W.; Gottesman, A.R.; Mott, G.E. Isolation and characterization of new strains of cholesterol-reducing bacteria from baboons. Appl. Environ. Microbiol. 1982, 43, 86–89. [Google Scholar] [PubMed]
- Freier, T.A.; Beitz, D.C.; Li, L.; Hartman, P.A. Characterization of Eubacterium coprostanoligenes sp. nov., a cholesterol-reducing anaerobe. Int. J. Syst. Bacteriol. 1994, 44, 137–142. [Google Scholar] [CrossRef]
- Li, L. Characterization and Application of a Novel Cholesterol-Reducing Anaerobe, Eubacterium coprostanoligenes ATCC 51222. Ph.D. Thesis, Iowa State University, Ames, IA, USA, 1995. [Google Scholar]
- Mott, G.E.; Brinkley, A.W. Plasmenylethanolamine: Growth factor for cholesterol reducing Eubacterium. J. Bacteriol. 1979, 139, 755–760. [Google Scholar] [PubMed]
- Sadzikowski, M.R.; Sperry, J.F.; Wilkins, T.D. Cholesterol-reducing bacterium from human feces. Appl. Environ. Microbiol. 1977, 34, 355–362. [Google Scholar]
- Gérard, P.; Lepercq, P.; Leclerc, M.; Gavini, F.; Raibaud, P.; Juste, C. Bacteroides sp. strain D8, the first cholesterol-reducing bacterium isolated from human feces. Appl. Environ. Microbiol. 2007, 73, 5742–5749. [Google Scholar] [CrossRef]
- Snog-Kjaer, A.; Prange, I.; Dam, H. Conversion of cholesterol into coprosterol by bacteria in vitro. J. Gen. Microbiol. 1956, 14, 256–260. [Google Scholar] [CrossRef] [PubMed]
- Crowther, J.S.; Drasar, B.S.; Goddard, P.; Hill, M.J.; Johnson, K. The effect of a chemically defined diet on the faecal flora and faecal steroid concentration. Gut 1973, 14, 790–793. [Google Scholar] [CrossRef] [PubMed]
- Lye, H.S.; Rusul, G.; Liong, M.T. Removal of cholesterol by lactobacilli via incorporation and conversion to coprostanol. J. Dairy Sci. 2010, 93, 1383–1392. [Google Scholar] [CrossRef] [PubMed]
- Antharam, V.C.; McEwen, D.C.; Garrett, T.J.; Dossey, A.T.; Li, E.C.; Kozlov, A.N.; Mesbah, Z.; Wang, G.P. An Integrated metabolomic and microbiome analysis identified specific gut microbiota associated with fecal cholesterol and coprostanol in Clostridium difficile Infection. PLoS ONE 2016, 11, e0148824. [Google Scholar] [CrossRef] [PubMed]
- Midtvedt, A.C.; Midtvedt, T. Conversion of cholesterol to coprostanol by the intestinal microflora during the first two years of human life. J. Pediatr. Gastroenterol. Nutr. 1993, 17, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, T.D.; Hackman, A.S. Two patterns of neutral steroid conversion in the feces of normal North Americans. Cancer Res. 1974, 34, 2250–2254. [Google Scholar] [PubMed]
- Veiga, P.; Juste, C.; Lepercq, P.; Saunier, K.; Beguet, F.; Gérard, P. Correlation between faecal microbial community structure and cholesterol-to-coprostanol conversion in the human gut. FEMS Microbiol. Lett. 2005, 242, 81–86. [Google Scholar] [CrossRef]
- Macdonald, I.A.; Bokkenheuser, V.D.; Winter, J.; McLernon, A.M.; Mosbach, E.H. Degradation of steroids in the human gut. J. Lipid Res. 1983, 24, 675–700. [Google Scholar]
- Gérard, P.; Béguet, F.; Lepercq, P.; Rigottier-Gois, L.; Rochet, V.; Andrieux, C.; Juste, C. Gnotobiotic rats harboring human intestinal microbiota as a model for studying cholesterol-to-coprostanol conversion. FEMS Microbiol. Ecol. 2004, 47, 337–343. [Google Scholar] [CrossRef]
- Rosenfeld, R.S.; Fukushima, D.K.; Hellman, L.; Gallagher, T.F. The transformation of cholesterol to coprostanol. J. Biol. Chem. 1954, 211, 301–311. [Google Scholar]
- Rosenfeld, R.S.; Hellman, L.; Gallagher, T.F. The transformation of cholesterol-3d to coprostanol-d. Location of deuterium in coprostanol. J. Biol. Chem. 1956, 222, 321–323. [Google Scholar]
- Mott, G.E.; Brinkley, A.W.; Mersinger, C.L. Biochemical characterization of cholesterol-reducing Eubacterium. Appl. Environ. Microbiol. 1980, 40, 1017–1022. [Google Scholar] [PubMed]
- Cuevas-Tena, M.; Alegría, A.; Lagarda, M.J. Relationship Between Dietary Sterols and Gut Microbiota: A Review. Eur. J. Lipid Sci. Technol. 2018, 120, 1800054. [Google Scholar] [CrossRef]
- Rosenfeld, R.S.; Gallagher, T.F. Further studies of the biotransformation of cholesterol to coprostanol. Steroids 1964, 4, 515–520. [Google Scholar] [CrossRef]
- Björkhem, I.; Gustafsson, J.A. Mechanism of microbial transformation of cholesterol into coprostanol. Eur. J. Biochem. 1971, 21, 428–432. [Google Scholar] [CrossRef] [PubMed]
- Parmentier, G.; Eyssen, H. Mechanism of biohydrogenation of cholesterol to coprostanol by Eubacterium ATCC 21408. Biochim. Biophys. Acta 1974, 348, 279–284. [Google Scholar] [CrossRef]
- Ren, D.; Li, L.; Schwabacher, A.W.; Young, J.W.; Beitz, D.C. Mechanism of cholesterol reduction to coprostanol by Eubacterium coprostanoligenes ATCC 51222. Steroids 1996, 61, 33–40. [Google Scholar] [CrossRef]
- García, J.L.; Uhía, I.; Galán, B. Catabolism and biotechnological applications of cholesterol degrading bacteria. Microb. Biotechnol. 2012, 5, 679–699. [Google Scholar] [CrossRef]
- Vrielink, A.; Ghisla, S. Cholesterol oxidase: Biochemistry and structural features. FEBS J. 2009, 276, 6826–6843. [Google Scholar] [CrossRef]
- Cheetham, P.S.; Dunnill, P.; Lilly, M.D. The characterization and interconversion of three forms of cholesterol oxidase extracted from Nocardia rhodochrous. Biochem. J. 1982, 201, 515–521. [Google Scholar] [CrossRef]
- Tomioka, H.; Kagawa, M.; Nakamura, S. Some enzymatic properties of 3beta-hydroxysteroid oxidase produced by Streptomyces violascens. J. Biochem. 1976, 79, 903–915. [Google Scholar] [CrossRef] [PubMed]
- Fukuyama, M.; Miyake, Y. Purification and some properties of cholesterol oxidase from Schizophyllum commune with covalently bound flavin. J. Biochem. 1979, 85, 1183–1193. [Google Scholar] [PubMed]
- Sojo, M.; Bru, R.; Lopez-Molina, D.; Garcia-Carmona, F.; Arguelles, J.C. Cell-linked and extracellular cholesterol oxidase activities from Rhodococcus erythropolis isolation and physiological characterization. Appl. Microbiol. Biotechnol. 1997, 47, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Vrielink, A.; Lloyd, L.F.; Blow, D.M. Crystal structure of cholesterol oxidase from Brevibacterium sterolicum refined at 1.8 Å resolution. J. Mol. Biol. 1991, 219, 533–554. [Google Scholar] [CrossRef]
- Sampson, N.S.; Kwak, S. Catalysis at the membrane interface: Cholesterol oxidase as a case study. In Proceedings of the 3rd International Symposium on Experimental Standard Conditions of Enzyme Characterizations (ESCEC), Rudesheim Rhein, Germany, 23–26 September 2008; pp. 13–22. [Google Scholar]
- Joseph, K.; Sampson, N.S. Cholesterol oxidase: Physiological functions. FEBS J. 2009, 276, 6844–6856. [Google Scholar]
- Yu, L.J.; Golden, E.; Chen, N.; Zhao, Y.; Vrielink, A.; Karton, A. Computational insights for the hydride transfer and distinctive roles of key residues in cholesterol oxidase. Sci. Rep. 2017, 7, 17265. [Google Scholar] [CrossRef] [PubMed]
- Björkhem, I.; Gustafsson, J.A.; Wrange, O. Microbial transformation of cholesterol into coprostanol. Properties of a 3-oxo-4-steroid-5 beta-reductase. Eur. J. Biochem. 1973, 37, 143–147. [Google Scholar] [CrossRef] [PubMed]
- Zanotti, I.; Turroni, F.; Piemontese, A.; Mancabelli, L.; Milani, C.; Viappiani, A.; Prevedini, G.; Sanchez, B.; Margolles, A.; Elviri, L.; et al. Evidence for cholesterol-lowering activity by Bifidobacterium bifidum PRL2010 through gut microbiota modulation. Appl. Microbiol. Biotechnol. 2015, 99, 6813–6829. [Google Scholar] [CrossRef]
- Bolotine, A.; (INRA-paris-Saclay University, Jouy-en-Josas, France). Personal communication, 2017.
- Lynch, A.; Crowley, E.; Casey, E.; Cano, R.; Shanahan, R.; McGlacken, G.; Marchesi, J.R.; Clarke, D.J. The Bacteroidales produce an N-acylated derivative of glycine with both cholesterol-solubilising and hemolytic activity. Sci. Rep. 2017, 7, 13270. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kriaa, A.; Bourgin, M.; Mkaouar, H.; Jablaoui, A.; Akermi, N.; Soussou, S.; Maguin, E.; Rhimi, M. Microbial Reduction of Cholesterol to Coprostanol: An Old Concept and New Insights. Catalysts 2019, 9, 167. https://doi.org/10.3390/catal9020167
Kriaa A, Bourgin M, Mkaouar H, Jablaoui A, Akermi N, Soussou S, Maguin E, Rhimi M. Microbial Reduction of Cholesterol to Coprostanol: An Old Concept and New Insights. Catalysts. 2019; 9(2):167. https://doi.org/10.3390/catal9020167
Chicago/Turabian StyleKriaa, Aicha, Mélanie Bourgin, Héla Mkaouar, Amin Jablaoui, Nizar Akermi, Souha Soussou, Emmanuelle Maguin, and Moez Rhimi. 2019. "Microbial Reduction of Cholesterol to Coprostanol: An Old Concept and New Insights" Catalysts 9, no. 2: 167. https://doi.org/10.3390/catal9020167
APA StyleKriaa, A., Bourgin, M., Mkaouar, H., Jablaoui, A., Akermi, N., Soussou, S., Maguin, E., & Rhimi, M. (2019). Microbial Reduction of Cholesterol to Coprostanol: An Old Concept and New Insights. Catalysts, 9(2), 167. https://doi.org/10.3390/catal9020167





