1. Introduction
New functional products are increasingly demanded by the food, cosmetic, and pharmaceutical industries. Enzyme degrading processes of vegetable biomasses are a ‘white biotechnology’ for sustainable production of these new functional products [
1]. Degradation of the cell membrane before the extraction of biocomponents facilitates the recovery of cytoplasmic products [
2]. Since the great part of cyanobacteria biocomponents are inside the cell [
3,
4], their extraction can be improved via destructive pre-treatment of the cellular and subcellular structures [
2,
5]. To recover the biomass constituents, enzymatic tools provide energy savings compared to mechanical treatment processes or chemical catalytic hydrolysis at high temperatures. They also compete well in the selectivity of extracted biocomponents. Depending on the type of enzyme, the cellular membrane and cell components may be degraded in different ways. Consequently, the potential of this technology is high, considering the great variety of degradation products that can be obtained through the action of different types of selective catalysts. This fact opens many opportunities for enzyme technology. Examples of advantageous enzymatic processes able to extract high value products from vegetable and animal biomasses are: (i) degradation of cellulose to glucose using cellulases, and (ii) membrane phospholipid hydrolysis by lipases for production of essential fatty acids.
The food chain is based on algae and microalgae products. Microalgae are good protein and metabolite sources. Dietary supplements are obtained from them [
6].
Cyanobacteria (also named blue algae) are prokaryotic, photosynthetic, unicellular microorganisms. To this group belong
Arthrospira platensis and
Arthrospira maxima (commonly named
Spirulina platensis and
Spirulina maxima).
Spirulina platensis grows in Africa, Asia, and South America, with a characteristic helicoidal morphology [
7].
Spirulina biomass (spirulina) has great potential for its composition and nutritional properties [
8]. More than two thousand years ago, it was already being consumed by Aztecs. Spirulina properties are very much appreciated by modern society, being widely consumed for different types of healthy diets. Spirulina is a good source of essential amino acids, vitamins, carbohydrates, macro- and trace minerals, and other nutrients. The composition percent dry weight of spirulina is 64–73% protein, 12–17% carbohydrate, 5–7% lipids, 0.9% P, and 10.3–11.6% N. [
9]. Blue pigments (phycocianins) of spirulina contribute to increasing the protein and iron availability [
10]. Since 2011, spirulina is considered a safe ingredient in class A diet supplements [
11]. Spirulina improves the immune system [
12], and it has therapeutic effects against cancer and different virus, microbial, and inflammatory processes [
1]. Spirulina provides high amounts of a unique antioxidant amino acid: L-ergothioneine (EGT; 2-mercaptohistidine trimethylbetaine) [
13]. Spirulina supplementation provides vegetable proteins to the organism. To increase bioassimilation of its proteic material, extracts of its degraded proteic biocomponents must be obtained [
14]. Additionally, the efficient extraction of intracellular components is limited by the cell membrane.
The cell membrane of
Spirulina is analogous to that of Gram-negative bacteria [
15]; they have two lipid membrane layers (cellular and cytoplasmic), separated by the murein layer. Murein is a rigid macromolecular structure formed by complex polymers of peptidoglycans and lipopolysaccharides. Peptidoglycans are covalently linked disaccharides and tetrapeptides. They are placed between cellular and cytoplasmic membranes, and linked to the external membrane layer by lipoproteins. Lipopolysaccharides are formed by a lipid and a complex polysaccharide chains. The cellular membrane has proteins non-covalently linked to lipids, whereas the cytoplasmic membrane is formed by lipoproteins (proteins covalently linked to lipids). The sugar complexes of cell membranes function as energy reserves (e.g., glucogen) [
12].
Methods for cell membrane degradation may be physical (ultrasounds, microwaves [
16,
17], osmotic sock, pulse electric field, heat treatment [
18], etc.), but may also be chemical or enzymatic [
19,
20]. The recovery of biocomponents can be achieved by different methods, such as those based on phase separation with solvents [
21,
22], supercritical fluids, pervaporation, etc. Safi et al. compared different fragile-cell-walled microalgae using several physical [
23] and chemical [
24] cell disruption methods. They were attacked according to the following order:
Haematococcus pluvialis <
Nannochloropsis oculata <
Chlorella vulgaris <
Porphyridium cruentum ≤
Arthrospira platensis. These authors determined that among the physical methods employed for protein extraction, high-pressure cell disruption was the most efficient one, although it was not enough to recover more than 50% of the proteins from these green microalgae, indicating that more passes are required to completely disrupt their macrostructure, and thus more energy input is necessary [
23]. Using several cycles, mechanical treatments released more proteins from all the microalgae compared to chemical treatments. Percentages of protein extracted from
Spirulina biomass using an alkaline pre-treatment or high-pressure homogenization method were 68% and 75%, respectively [
24]. On the other hand, some reports described different enzyme treatments of protein extracts from spirulina, obtaining several bioactive products. In all the cases, the protein extracts were obtained by mechanical methods, prior to being submitted to a given enzyme degradation. More precisely, the iron-chelating peptide Thr-Asp-Pro-Ile(Leu)-Ala-Ala-Cys-Ile(Leu), with a molecular weight of 802 Da, was obtained through the combined action of two proteases [
25]. The protein extract was first obtained by homogenization of the cell suspension, centrifugation, and further precipitation of the supernatant with ammonium sulfate. The extracted material was then consecutively submitted to two proteolytic steps by two different proteases. In other study, the antihypertensive peptide Ile-Gln-Pro was prepared via Alcalase
® digestion of previously extracted proteins by freeze–thawing and sonication procedures [
26]. Similarly, two potential anti-inflammatory peptides (LDAVNR and MMLDF) were obtained by subsequent proteolysis with trypsin, chymotrypsin, and pepsine of the proteins that were previously extracted from spirulina by freeze–thawing and sonication procedures [
27]. All these studies were focused on purification and characterization of particular bioactive peptides [
25,
26,
27]. In these cases, only the fractions of proteins previously extracted by physical or mechanical methods were treated with enzymes, and in none of the cases did the authors take into consideration the advantage that the direct action of the enzymes on the spirulina cells provides: higher recovery of intracellular components (e.g., proteins). Hence, more studies based on direct enzyme extraction protocols are required. In order to take full advantage of the enzyme-assisted extractions of proteins, direct studies of the digestion of
Spirulina cells by enzymes should be carried out. In that respect, appropriate use of the enzyme technology requires determining for each biocatalyst the influence of different operation parameters on the enzyme degrading activity for each specific biomass.
In this work, the application of different enzymes for obtaining easily adsorbed proteic extracts was studied. Different selective enzyme degradation processes of Spirulina biomass were investigated for the extraction of polar spirulina biocomponents. The biomass was enzymatically degraded using four different cheap and easily accessible commercial enzyme preparations. These processes at their corresponding best operation conditions were compared with the extraction done without any prior enzyme treatment of the biomass. In particular, two different enzyme treatments, based on the degradation of membrane proteins, lipoproteins, and peptidoglucan by two proteases (Alcalase® and Flavourzyme®), and two other biomass treatments using endo- and exo-glucanases (Ultraflo® and Vinoflow®) to breakdown the sugar polymer structure, were comparatively studied. The best values of the most important parameters of all the enzymatic pre-treatments for biomass degradation were determined. The corresponding extractive yields in dry weight of aqueous extracts were determined for the four enzyme extraction processes at their respective best conditions, and they were compared with those of the control extract (without enzyme assistance). Changes occurring at the cellular level after the different extraction processes were comparatively analyzed by transmission electron microscopy (TEM). Total amino acid contents of all hydrophilic extracts were compared.
2. Results and Discussion
A commercial dry biomass of the cyanobacteria Arthrospira platensis (Spirulina) was submitted to different enzyme degradations. S. platensis extractions after four different enzyme pre-treatments were independently studied. These processes were carried out in their respective best operation conditions and compared with the control (no enzyme added) extraction. Two distinct proteases and two distinct glucanases were used to favor biocomponent recovery via membrane enzymatic degradation. The four different enzyme extractions were analyzed in terms of both the weight yields of the dry aqueous extracts and their respective amino acids contents.
2.1. Enzyme-Assisted Extraction
The most important parameters of the enzyme-assisted degradation of spirulina were studied. The best operation conditions for each enzyme extraction were determined by following changes in each peak area value of the high performance liquid chromatography with a evaporative light scattering detector (HPLC-ELSD) chromatograms of the aqueous phase. In all cases, the area changes in all the significant peaks were inspected (see below). All significant peaks of the HPLC chromatogram exhibited the same dependence of the studied parameters, that is, the same variation (increase or decrease) of their area values with the studied parameter value, and the same optimal value. Hence, for the sake of clarity, the influences of pH, time, temperature, and enzyme loading are depicted with respect to the total area values of all significant peaks. Nevertheless, a couple of examples of the parameter (pH and time) effect in one peak area are also given.
2.1.1. Influence of pH
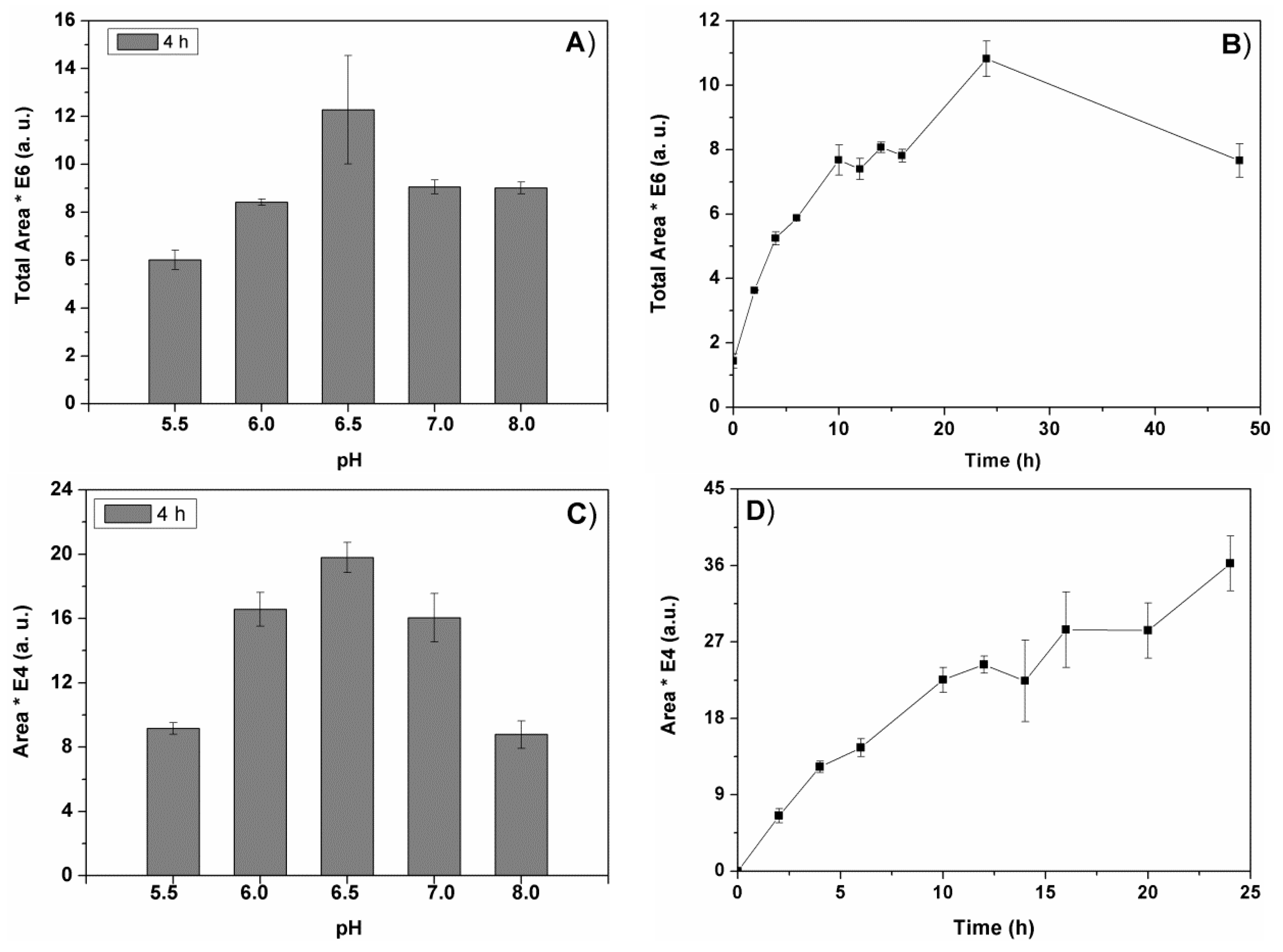
Figure 1 represents the change of the total peak areas in the chromatogram (
Figure 1A), and the area of a representative individual peak (
Figure 1C) obtained after 4h of biomass pre-treatment with Alcalase
®. The maximal initial rate (4h) of the biocomponent extraction with Alcalase
® was obtained at pH 6.5. The enzyme degradation process was faster at this pH value, giving rise to the maximal total peak area value obtained for a short time pre-treatment (4h,
Figure 1A). For the extractions with Flavourzyme
®, Ultraflo
®, and Vinoflow
®, the corresponding best pH values determined were 6.0, 7.0, and 6.5, respectively (not shown).
Lu et al. used a different pH value for the extraction of an anti-hypertensive peptide from spirulina using Alcalase
® (pH 8.5 at 50 °C for 10h) after three cycles of freeze–thawing the biomass [
26]. Kim et al. also used different operation conditions to obtain the iron-chelating peptide with a multi-enzymatic biomass degradation (Alcalase
® pH 8.0 at 50 °C for 1 h + Flavourzyme
® pH 7.0 at 50 °C for 8 h) [
25]. The obtained products of their peptide degradation were also different from those of this study (work in progress, personal communication).
The dependence of the extraction yield on the pH parameter is a consequence of the effect of pH on the process of biomass degradation by the enzyme. Variations in the total charge of proteins with pH should determine their mutual interaction and interaction with the biocatalyst employed. For the extraction of anti-cancer biocomponents from spirulina, papaine presented an optimal pH of 6.5, while other enzymes exhibited very different optimal pH values (pH 2–8.5) [
28]. Optimal values in the basic range of pH were earlier described for the extraction of spirulina antioxidants with different enzymes, although other group of enzymes exhibited low activities under these conditions [
29]. Different proteases from those used here were earlier studied to extract spirulina oil, reporting an optimal pH range of 7.5–10 for the best biocatalyst [
30].
The time course of the enzyme-assisted extraction at the best pH value was studied for the first 48 h. The results obtained with Alcalase
® for all the chromatogram peaks are represented in
Figure 1B, and for a representative peak in
Figure 1D. Similarly to Alcalase
® extraction, the other three types of enzyme extractions required 24 h of enzyme pre-treatment of spirulina to reach maximal recovery of biocomponents (not shown). These results (
Figure 1B,D) were obtained via analyses of aliquots taken from the reaction mixtures at the indicated times, while those of
Figure 1A,B correspond to analyses of lyophilized extracts. The two studies correspond to samples of different extract concentrations, so the area values of these two figures cannot be compared (see Materials and Methods
Section 3.3.1).
2.1.2. Effect of Temperature
The effect of the temperature of the biomass pre-treatment on the extraction of polar biocomponents was carried out at the respective optimal pH values of each enzyme (Alcalase
® at pH 6.5; Flavourzyme
® at pH 6.0; Ultraflo
® at pH 7.0; Vinoflow
® at pH 6.5) and an enzyme loading of 1% (
v/
w) in the range 30–50 °C after 24 h of biomass treatment (
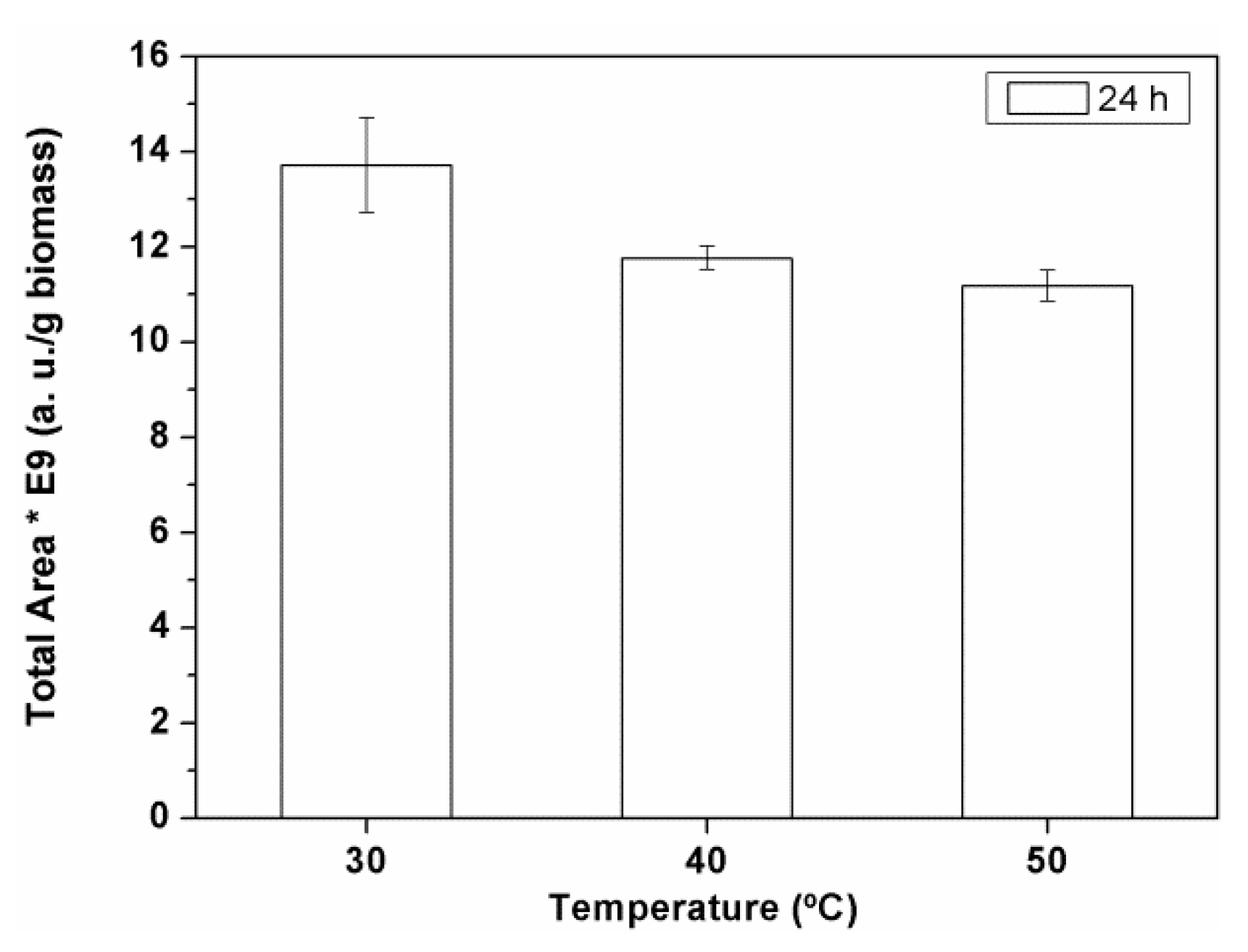
Figure 2). The total area values of peak products of the corresponding aqueous extracts were calculated from the obtained HPLC-ELSD chromatograms. The area value corresponding to 1 g of spirulina was calculated and represented against temperature.
Figure 2 depicts the total peaks area values obtained for Alcalase
®.
Considering the experimental error, 30 °C was the best temperature determined for the extractions with Alcalase
®. Similar results were obtained with the other three enzymes (not shown). From these results, 30 °C was determined to be the best temperature for the two proteases (Alcalase
® and Flavourzyme
®) and Ultraflo
®, the latter one having β-glucanase and xylanase activities as well as several side activities (Cellulase, Hemicellulase, and Pentosanase). The optimal temperature value determined for Vinoflow
® was 40 °C (not shown). In all cases, the best temperature to achieve the maximum recovery of biocomponents in the aqueous extract was relatively low. Vitamins, antioxidants, and other thermo-labile components of the extracts were relatively well preserved in this temperature range. At mild temperatures the energy expenses and the product lability were reduced, while the operational stability of the biocatalysts employed was increased. In fact, the decrease observed in the extraction of biocomponents at the higher temperatures (
Figure 2), might be explained by a decay of the operational stability of the enzyme.
In the case of proteases, Zhang & Zhang reported higher optimal temperatures for enzyme-assisted extraction of anti-tumor polypeptides from spirulina with trypsin, pepsine, and papain, namely 42 °C, 37 °C, and 55 °C, respectively, than the ones herein determined [
28].
2.1.3. Effect of the Enzyme Loading
Catalyst charge affects the speed of biomass biodegradation prior to solvent extraction, determining the extraction yield value obtained at a given time.
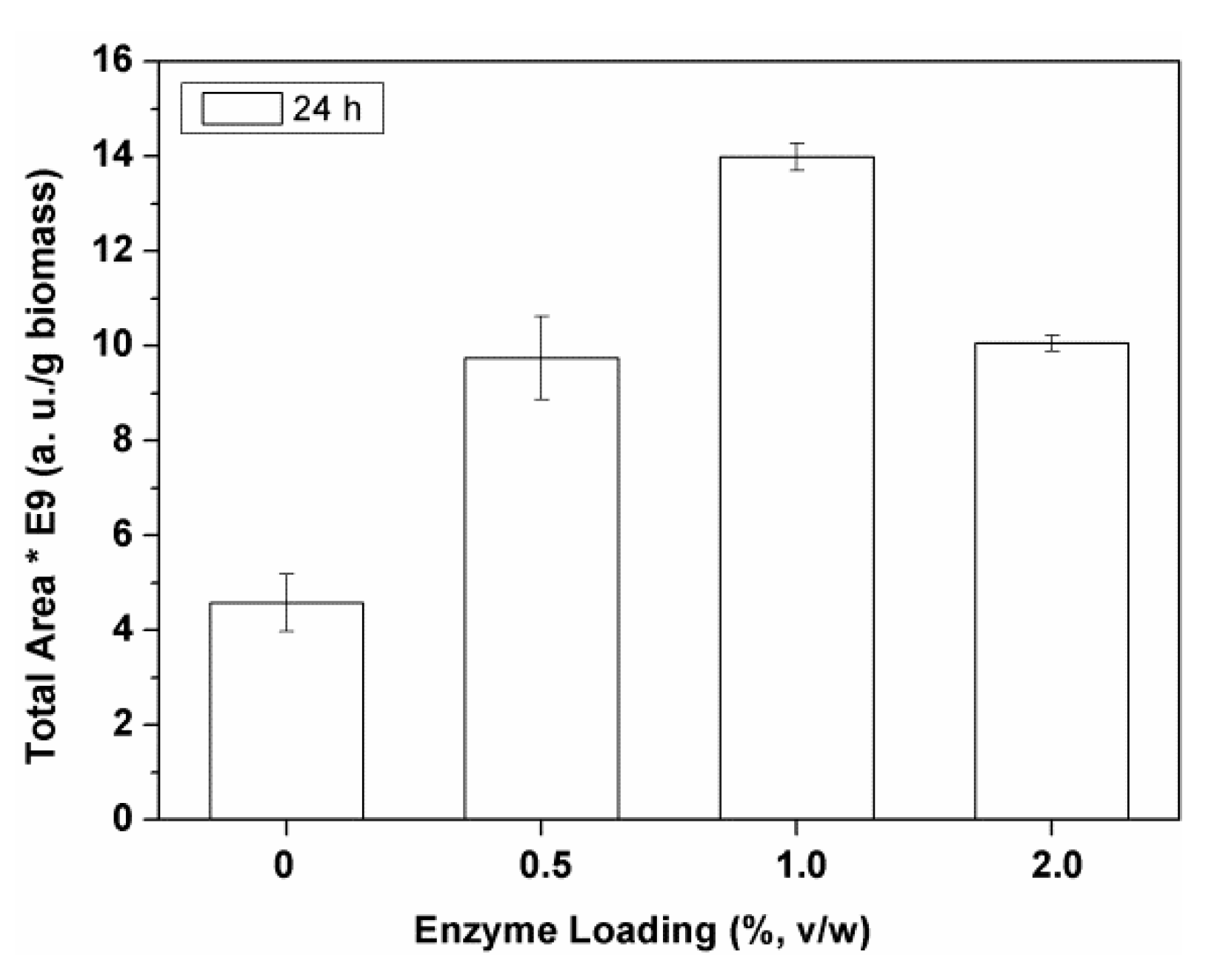
The results obtained with different Alcalase
® loadings (0–2%
v/
w enzyme) are represented in the
Figure 3. These results were compared with those of a control extraction assay, where, instead of the enzyme solution, an equivalent volume of water solution was used (0% enzyme loading in
Figure 3).
The extraction of polar biocomponents increased with the increase of the enzyme loading from 0.5% to 1% (
v/
w) Alcalase
®, but it dramatically decreased with an higher enzyme amount (i.e., 2%
v/
w;
Figure 3). The same best value was found for Flavourzyme
® and Ultraflo
®. The best enzyme loading of Vinoflow
® was 2% (
v/
w) (not shown). These values correspond to 1.2–2.4% (
w/
w), considering the corresponding density values.
Our results are in good agreement with those of the literature, where the dependence of the extraction yield exhibits a maximum value, with a value of biocatalyst loading above which the process becomes less efficient [
31]. The decrease of the product recovery when using high enzyme charges suggests that high enzyme concentrations might favor an excessive biodegradation of the biomass. Different types of biomasses require different amounts of biocatalysts. All these reports and the results of this work indicate that the optimal charge of enzyme requested depends on the type of biomass, type of the biocatalyst, and the biocomponents to be extracted.
The duration of the enzyme pre-treatment is an important parameter, involving important considerations in the process scale up. In this work, 24 h seemed to be necessary to get maximum extraction of intra- and extracellular biocomponents from spirulina. Liang et al. [
5] studied an enzyme-assisted extraction of microalgae oil, where optimal yield extraction was found at 12 h of biomass biodegradation. These results, and those of the literature, suggest that the necessary time to degrade the biomass depends on both the enzyme and the biomass studied. It also substantially varies with the method used to degrade the biomass and the product to be extracted. Extractions based on the use of ultrasounds and microwave irradiation for biomass degradation use relatively short times (20 min–4 h) for antioxidant and pigment extractions from microalgae [
16].
2.2. Scale Up and Extraction Yields
The four different enzyme-assisted extraction processes were carried out at their corresponding optimal operation conditions at a scale increased by a factor of 2.5. Ten grams of Spirulina biomass were used for each experiment. All the extracts were prepared in their respective optimal conditions (pH 6.5, 1% v/w and 30 °C for Alcalase®; pH 6.0, 1% v/w and 30 °C for Flavourzyme®; pH 7.0, 1% v/w and 30 °C for Ultraflo®; pH 6.5, 2% v/w and 40 °C for Vinoflow®). The control extract was obtained in this scale without any enzyme assistance, using milli-Q water instead of the enzyme solution, at 30 °C and 24h. The aqueous extracts obtained with the different enzyme-assisted methods were liophylyzed and kept at –70 °C.
Table 1 summarizes the recovery yields of aqueous extracts obtained. They are expressed in dry weight percent (calculated with respect to the starting
Spirulina biomass).
All the enzyme-assisted extraction processes enabled greater yields of aqueous extracts than the control extraction, being higher the yields obtained with the two proteases (Alcalase
® and Flavourzyme
®). The highest extraction yield was obtained with Alcalase
®. This biocatalyst enables an increase by 1.9 times of the weight of extract obtained, compared to the one obtained in the control extraction process without any enzyme assistance (
Table 1).
2.3. Compositional Analysis of Polar Extracts
All dry extracts (obtained in their respective best conditions) were analyzed to determine their amino acid and peptide compositions. The extracts were rich in free amino acids and short peptides. The total values obtained for both hydrolyzed and free amino acids are given in
Table 1.
Alcalase
® extract had a significantly higher content of amino acids (45%
w/
w dry extract) than the extract obtained without any enzyme assistance (34%
w/
w), and also higher than the other three enzymatic extracts (13–17%
w/
w). Alcalase
® extract has higher nutritional interest, and some of the peptides could exhibit bioactivities of therapeutic interest. This value corresponded to most amino acids of the spirulina (50–65%
w/
w,
Table 2).
Total (hydrolyzed + free) amino acids content values were also calculated per gram of dry spirulina. The ratio between amounts of extracted amino acids for each particular method and the quantity obtained with the control extraction was greater than 1 only in the case of Alcalase®, indicating the superiority of Alcalase® method not only with respect to the control extraction, but also with respect to the other enzymatic methods (p ≤ 0.05). Remarkably, among the two proteolytic enzymes and the two studied glucanases, only Alcalase® significantly increased the recovery percentage of amino acids with respect to the control assay (853 µmol/g spirulina more). These results suggest that amino acid extraction is not necessarily improved by any type of protease-assisted extraction. However, only the process with Alcalase® yielded a weight of amino acids 2.5 times higher than the control extraction, and 3.1 times higher than the extraction with the other protease (Flavourzyme®). From 1 g biomass, the Alcalase® extraction method obtained 1426 µmol amino acids, a quantity 2.5–6.1 times higher than the total amount obtained with the other methods (control without enzyme and enzyme-assisted methods). The control extraction obtained only 573 µmol amino acids/g biomass.
Liquid chromatography coupled to an electrospray ionization mass spectrometer in positive ionization mode (LC ESI-MS/MS) analyses revealed that the most abundant peptides in the spirulina extracts obtained with four enzymes and without an enzyme pre-treatment (control) were all different from one extract to another. More specifically, the eight peptides present in Alcalase® extract were: MKKIEAIIRPF, LPPL, ALAVGIGSIGPGLGQGQ, TTAASVIAAAL, DFPGDDIPIVS, LELL, WKLLP, and CHLLLSM (438–1493 Da). By contrast, the control extract contained 12 peptides, namely: NGDPFVGHL, VFETGIKVVDL, DFFVDKL, SGPPLDIKL, DVNETVLDNLPKTRTQI, DVNETVLDNLP, DSLISGAAQAVY, GIGNDPLEIQF, GLILLPHLATL, GLILLPHLA, AVLGAGALFHTF, and DVNETVLDNLP (851–1955 Da).
2.4. Biomass Analyses by TEM
Structural changes in the spirulina were investigated by TEM analysis (
Figure 4 and
Figure 5).
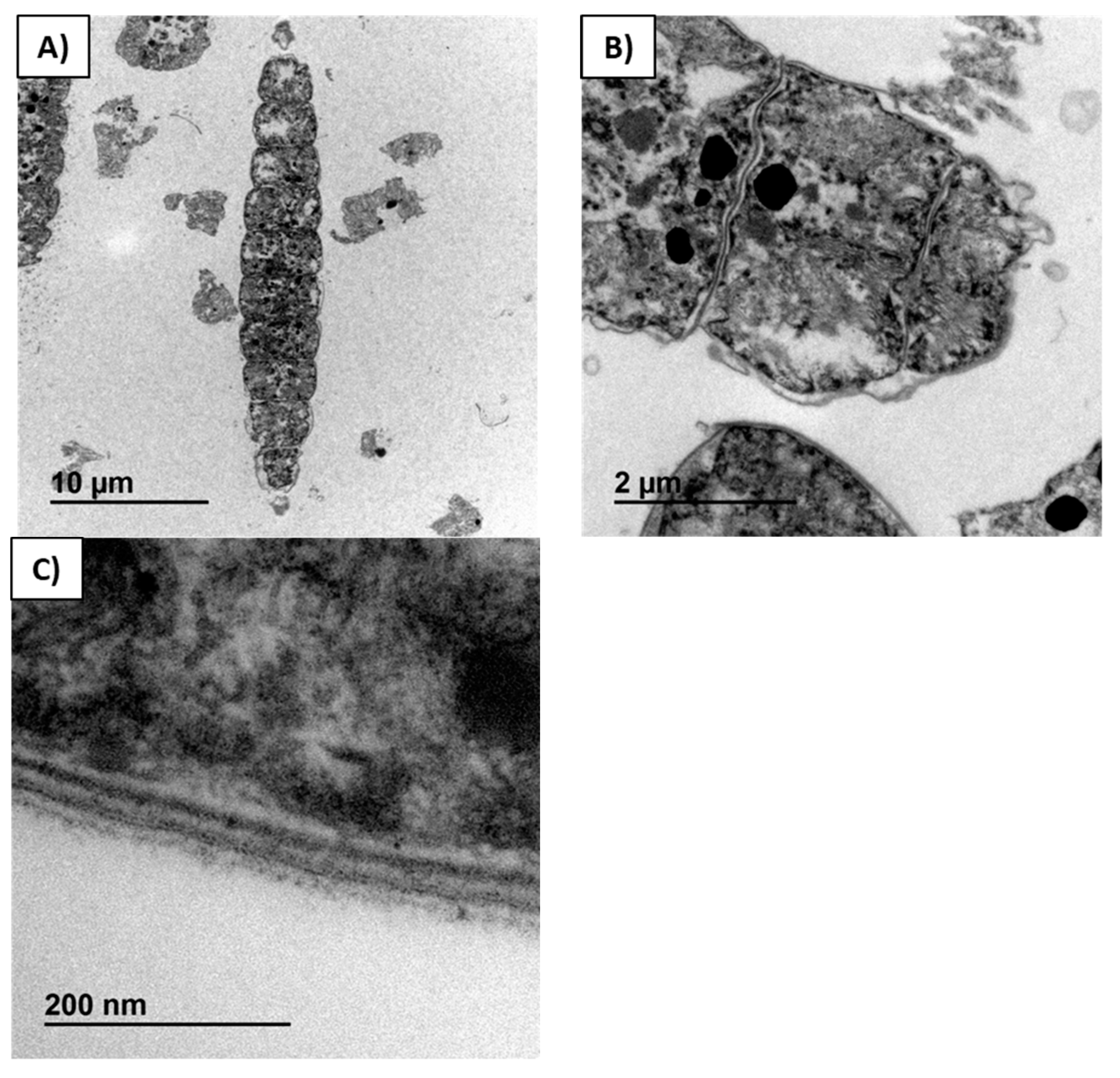
Figure 4A is a micrograph of commercial dry biomass not treated with enzymes nor extracted with any solvent, where a longitudinal section through a trichome with twelve cells is visualized. Complete integrity of the cellular material, including the membrane, is clearly visualized in
Figure 4B,C. The four layers of cell membrane were identified (
Figure 4C). In the starting spirulina, the cytoplasm and thylakoid system were compressed against the internal cell membrane.
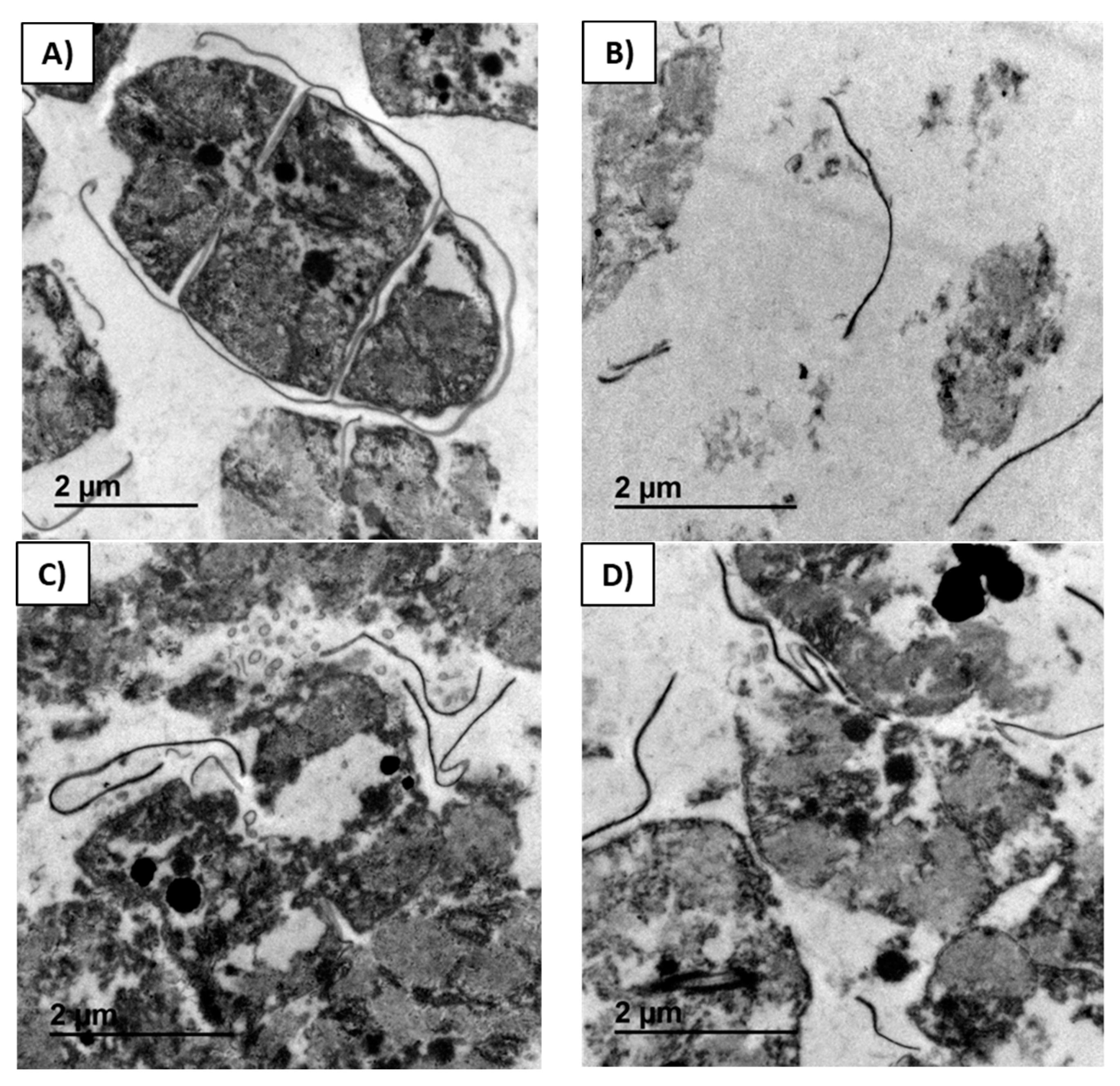
In
Figure 5, the micrographs of residual biomass obtained after the control extraction (
Figure 5A), and residual biomasses obtained after extraction with Alcalase
® (
Figure 5B), Ultraflo
® (
Figure 5C), and Vinoflow
® (
Figure 5D) assistance are compared at the same scale. Trichomes were still observed after the control extraction of the biomass, although their thylakoid system was not any more compressed against the internal cell membrane (
Figure 5A). Cells after the control extraction exhibited a swelling phenomenon, and a few were detached from the trichome. By contrast, after all the enzyme extractions studied, the number of trichomes observed was minimal and cellular degradation was evident (
Figure 5B–D). A dispersion of remaining (non-extracted) intracellular material and membrane rests was obtained after biomass treatment with Alcalase
® (
Figure 5B). After extraction with Alcalase
® assistance, most of the cellular material disappeared. Consequently, the residual biomass is more transparent (
Figure 5B) than the other ones (
Figure 5A,C,D). This fact is due to the higher recovery of biocomponents with Alcalase
® than with the control and other enzyme extractions (
Figure 5A,B). These observations agree with previous studies, where cyanobacteria cells modified their shape and size, swelling after a Lysozyme treatment [
32]. Less cellular material remained in the residual biomasses extracted with Ultraflo
® and Vinoflow
® (
Figure 5C,D) than in the control experiment (lower observed electronic density). However, in all these cases the amount of cell residues was significantly greater than in the case of the Alcalase
® extraction (
Figure 3B). These findings are in good correspondence with the values of extraction yields obtained (
Table 1), where the Alcalase
® extraction process was the most effective one for biocomponent recovery of spirulina.
The micrographs of this study show that Alcalase
® pre-treatment of the biomass is the most efficient, resulting in most of trichomes and spheroplasts degraded. Vladimirescu revealed the existence of differences in enzymatic sensitivity of cells [
33]. Compared with other bacteria,
Spirulina sp. has a thicker cell membrane that makes more difficult to detach the cells or spheroplasts from the trichome. Because of that, cell disintegration is not efficiently achieved with lysozyme treatment [
34].
The comparative inspection of micrographs of residual biomasses obtained with the different enzyme-assisted extraction processes (
Figure 5) reveals that the enzyme pre-treatment of the biomass that allowed the highest biocomponent recovery (Alcalase
®) produces the greatest cellular degradation. Compared with the other enzyme-assisted extraction types, the biomass extracted after Alcalase
® treatment appeared nearly completely disintegrated and with very low electronic density, as a result of the greater extraction of its biocomponents (
Figure 5B–D).
The results here described support the implementation of enzyme technology to replace conventional extraction processes. The Alcalase
® extraction herein reported significantly increased the yield and quality of aqueous extracts from
A. (Spirulina) platensis biomass. The enzyme technology developed uses food grade enzymes and hexane, which are accepted by regulatory agencies for food and drug safety. This enzyme technology looks promising for a more efficient, safe, and environmentally clean industrial production of cyanobacteria extracts with high value in the nutrition, cosmetic, and pharmaceutical industries [
14]. The potential of this extraction method will be further clarified once the extracted biocomponents are molecularly and functionally characterized. A complete characterization of the different aqueous extracts obtained with all the enzymes in their respective optimal operation conditions will be reported.