Biodegradation of Both Ethanol and Acetaldehyde by Acetobacter ghanensis JN01
Abstract
1. Introduction
2. Results and Discussion
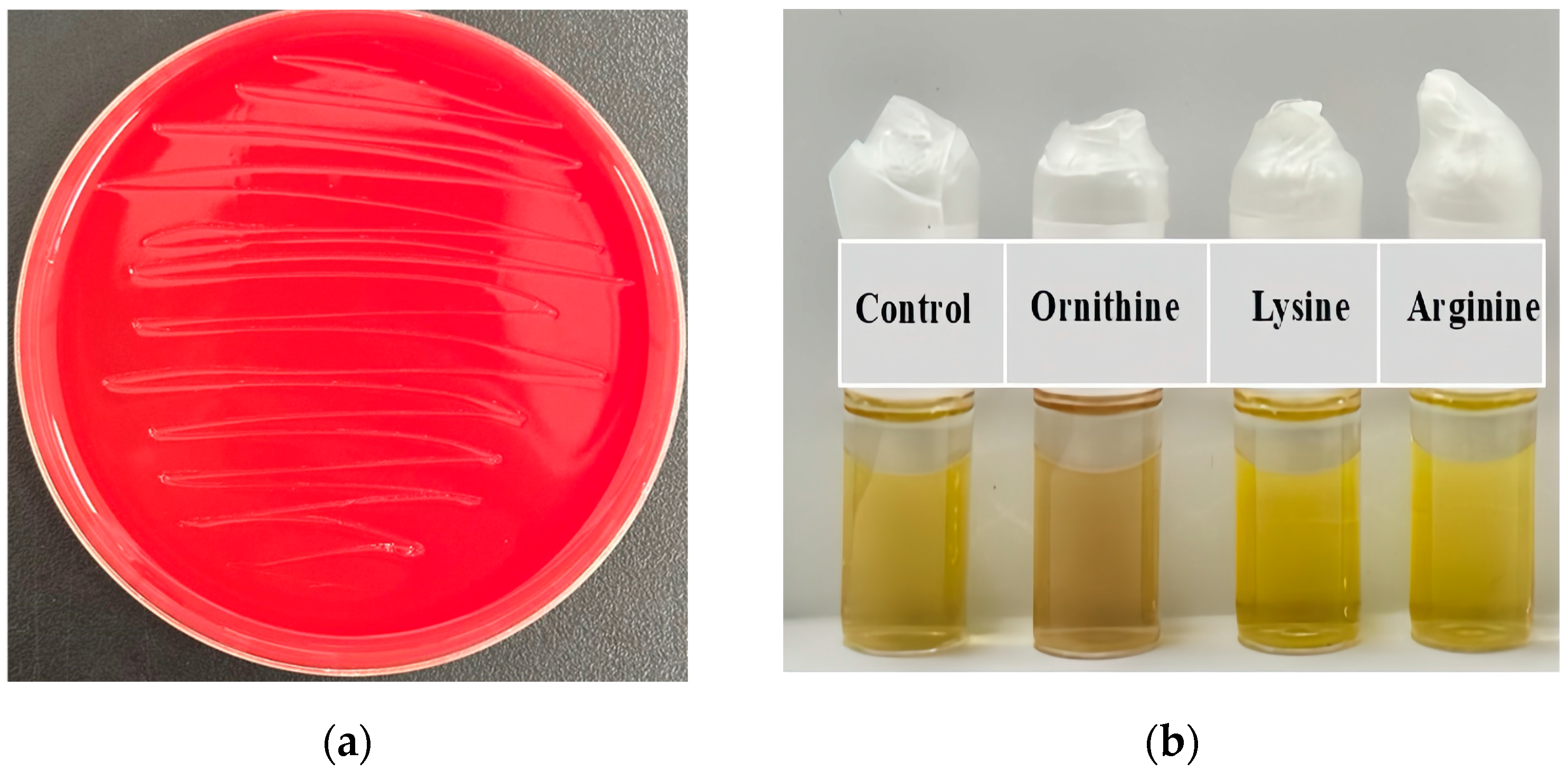
2.1. Isolation and Identification of a Bacterial Strain for Biodegrading Both Ethanol and Acetaldehyde
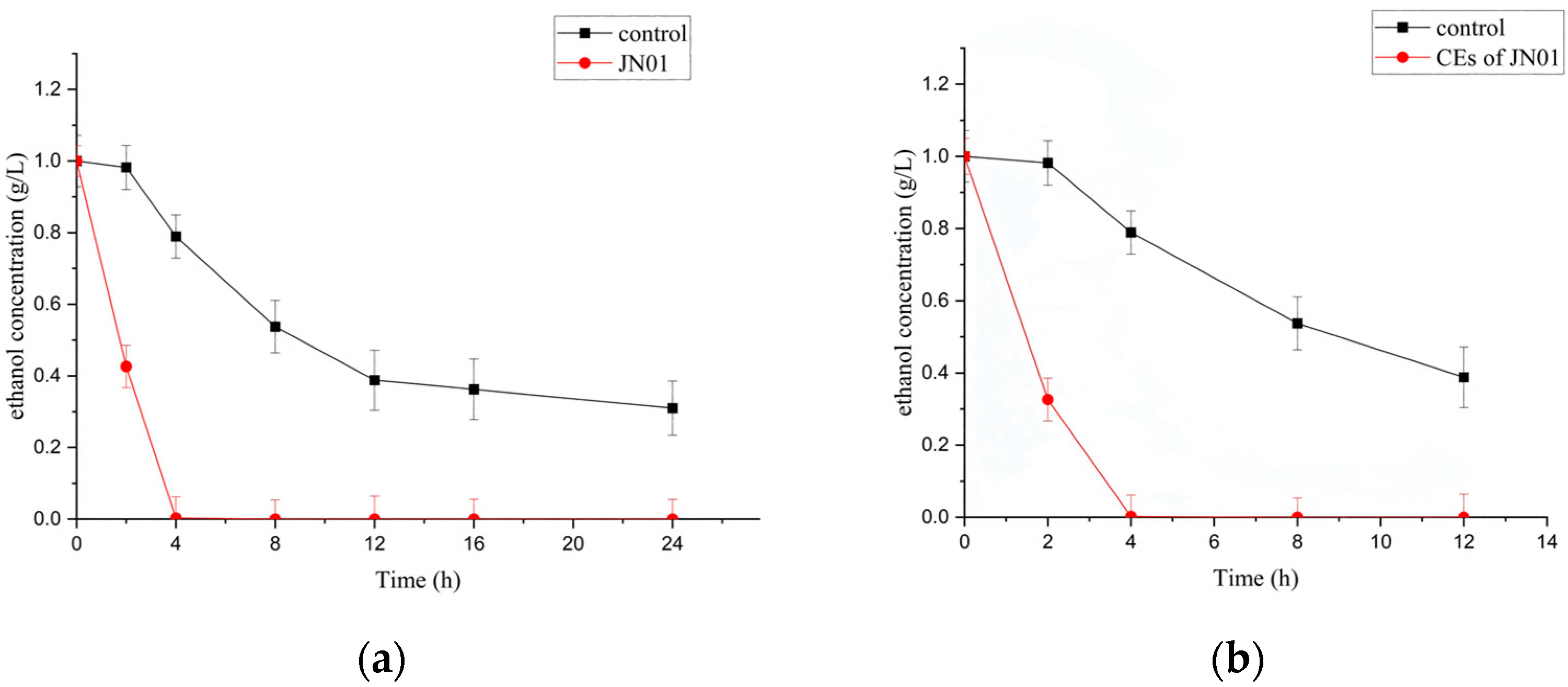
2.2. Biodegradation of Ethanol by Whole Cells and Cell-Free Extracts (CEs) of JN01
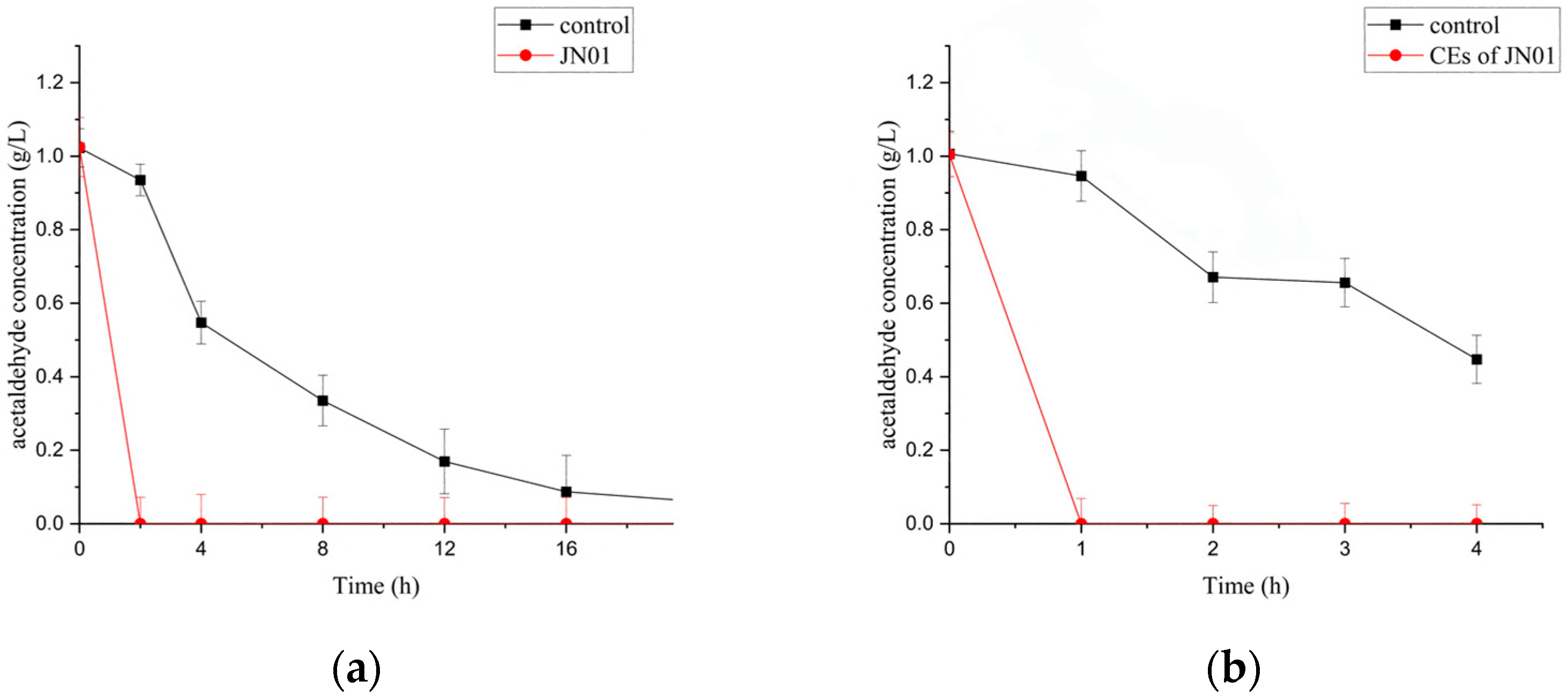
2.3. Biodegradation of Acetaldehyde by Whole Cells and CEs of JN01
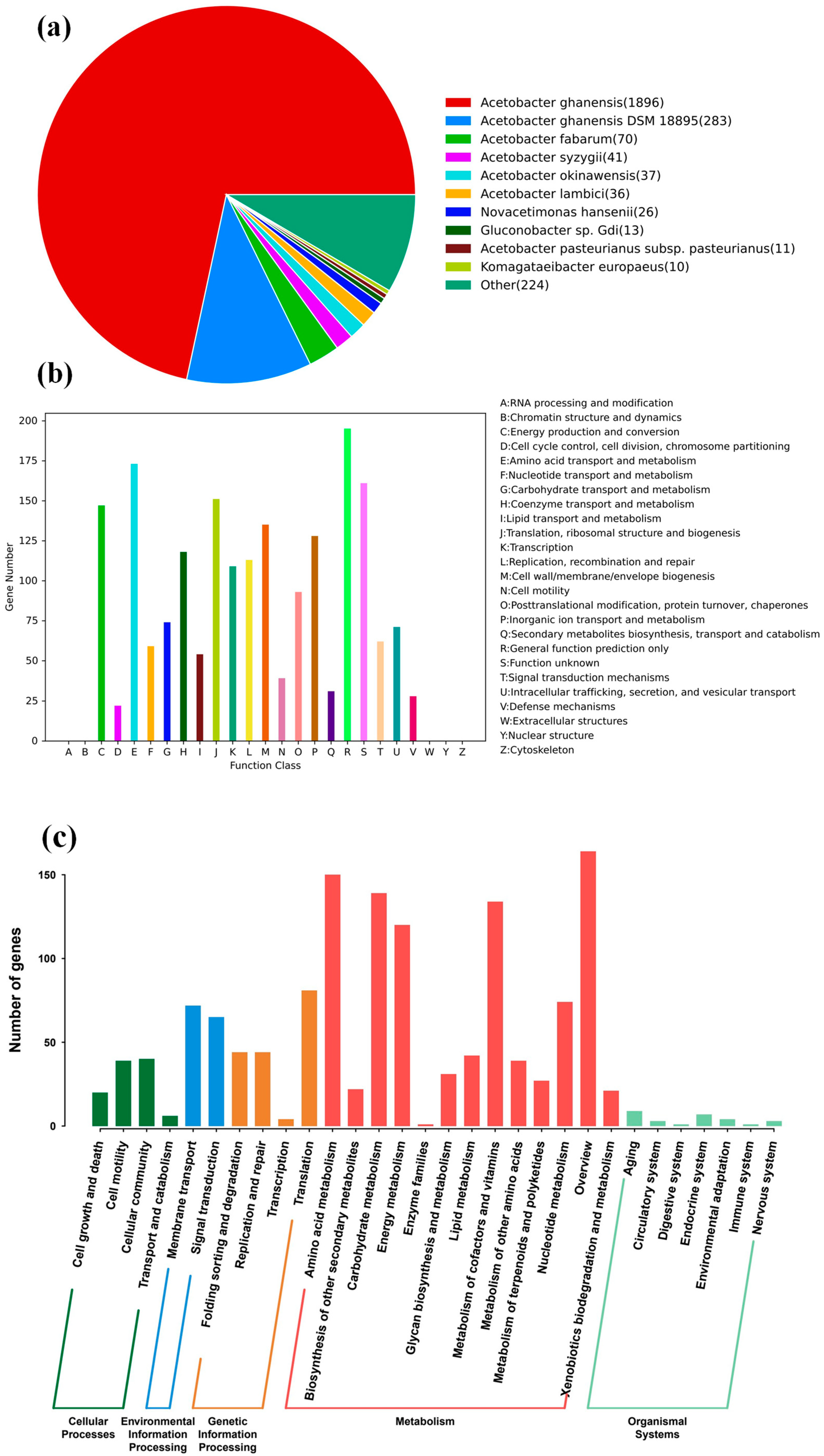
2.4. Overview of JN01 Genome Analysis
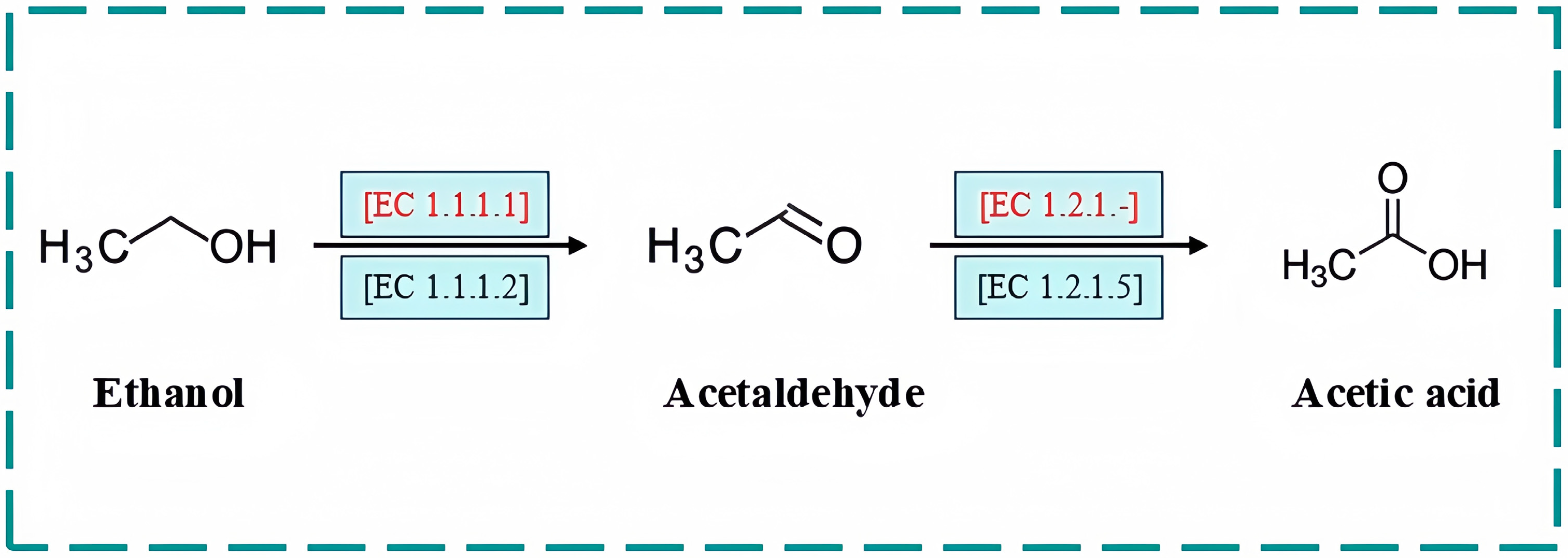
2.5. Pathway for Biodegrading Both Ethanol and Acetaldehyde by JN01
2.6. Genetic Features of Probiotic Properties of A. ghanensis JN01
2.7. Antibiotic Resistance and Virulence Factor Evaluation
2.8. In Vitro Safety Assessment of A. ghanensis JN01
3. Materials and Methods
3.1. Samples and Media
3.2. Isolation of Bacterial Strain for Biodegrading Both Ethanol and Acetaldehyde
3.3. Determination Biodegradation Ability of Both Ethanol and Acetaldehyde by JN01
3.4. Analytical Methods
3.5. Whole-Genome Sequencing and Genome Annotation
3.6. Identification of JN01
3.7. Safety Assessment
3.8. Statistical Analysis
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Macfarlane, V.F.; Prentice, D.A.; Walsh, M.S. The Auckland alcohol detoxification outcome study: Measuring changes in quality of life in individuals completing a medicated withdrawal from alcohol in a detoxification unit. Drug Alcohol Depend. 2019, 202, 156–161. [Google Scholar] [CrossRef]
- Song, S.; Lei, L.; Liu, H.; Yang, F.; Li, N.; Chen, W.; Ren, J. Impact of changing the prevalence of smoking, alcohol consumption and overweight/obesity on cancer incidence in China from 2021 to 2050: A simulation modelling study. EClinicalMedicine 2023, 63, 102215. [Google Scholar] [CrossRef]
- Liu, F.; Guo, H.; Ren, H.; Wang, C.; Zhang, Y.; Zhou, J. Pathogenic Mechanisms and Regulatory Factors Involved in Alcoholic Liver Disease. J. Transl. Med. 2023, 21, 369. [Google Scholar] [CrossRef]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Arsenic, metals, fibres, and dusts. IARC Monogr. Eval. Carcinog. Risks Hum. 2012, 100C, 11–465. [Google Scholar]
- Shortall, K.; Djeghader, A.; Magner, E.; Soulimane, T. Insights into aldehyde dehydrogenase enzymes: A structural perspective. Front. Mol. Biosci. 2021, 8, 659550. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Fu, L. The role of ALDH2 in tumorigenesis and tumor progression: Targeting ALDH2 as a potential cancer treatment. Acta Pharm. Sin. B 2021, 11, 1400–1411. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.C.; Guerrero-Morán, J.D.; González-Espinosa, C. Alcohol: Immunomodulatory Effects and Cancer. Rev. Investig. Clin. 2023, 75, 129–142. [Google Scholar] [CrossRef]
- Väkeväinen, S.; Tillonen, J.; Agarwal, D.P.; Srivastava, N.; Salaspuro, M. High salivary acetaldehyde after a moderate dose of alcohol in ALDH2-deficient subjects: Strong evidence for the local carcinogenic action of acetaldehyde. Alcohol. Clin. Exp. Res. 2000, 24, 873–877. [Google Scholar] [CrossRef]
- Li, L.B.; Chen, Y.; Lu, K.K. Advancements in nursing research regarding acute alcohol poisoning. Asian J. Surg. 2024, 47, 5205–5206. [Google Scholar] [CrossRef]
- Hyun, J.; Han, J.; Lee, C.; Yoon, M.; Jung, Y. Pathophysiological Aspects of Alcohol Metabolism in the Liver. Int. J. Mol. Sci. 2021, 22, 5717. [Google Scholar] [CrossRef]
- Mackowiak, B.; Fu, Y.; Maccioni, L.; Gao, B. Alcohol-associated liver disease. J. Clin. Investig. 2024, 134, e176345. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, L.; Qian, S.; Chen, Y.; Ya, R.; Ma, N.; Hao, Y.; Ge, S.; Zhang, X.; Yang, L.; et al. Hepatic microRNA-320 restrains ferroptosis to mitigate acute-on-chronic alcohol-induced liver injury. Biochim. Biophys. Acta Mol. Basis Dis. 2025, 1871, 167748. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Deng, S.; Wu, Z.; Cui, Z.; Mei, H.; Wang, J.; Zhang, Y. Angelica sinensis polysaccharide could alleviate the gastrointestinal damage in alcoholic fatty liver disease mice: Regulation of alcohol metabolism and enhancement of short-chain fatty acids utilization. J. Ethnopharmacol. 2025, 338, 119117. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Liu, W.; Ba, T.; Luo, Z.; Ma, Y.; Tang, G.; Kang, Y.J. Zinc-glutathione in Chinese Baijiu prevents alcohol-associated liver injury. Heliyon 2023, 9, e13722. [Google Scholar] [CrossRef]
- Mirijello, A.; Addolorato, G. Treatment of acute alcohol intoxication: The role of metadoxine. Eur. J. Intern. Med. 2023, 110, 128. [Google Scholar] [CrossRef]
- Higuera-de la Tijera, F.; Servín-Caamaño, A.I.; Cruz-Herrera, J.; Serralde-Zúñiga, A.E.; Abdo-Francis, J.M.; Gutiérrez-Reyes, G.; Pérez-Hernández, J.L. Treatment with metadoxine and its impact on early mortality in patients with severe alcoholic hepatitis. Ann. Hepatol. 2014, 13, 343–352. [Google Scholar] [CrossRef]
- Díaz Martínez, M.C.; Díaz Martínez, A.; Villamil Salcedo, V.; Cruz Fuentes, C. Efficacy of metadoxine in the management of acute alcohol intoxication. J. Int. Med. Res. 2002, 30, 44–51. [Google Scholar] [CrossRef]
- Silberman, Y.; Hajnal, A. Diet, diet access, and metabolic physiology as critically understudied factors in rodent models of alcohol intake: A commentary on Emous et al. (2025). Alcohol Clin. Exp. Res. 2025, 49, 1111–1112. [Google Scholar] [CrossRef]
- Hamurcu Varol, P.; Kaya, E.; Alphan, E.; Yilmaz, Y. Role of intensive dietary and lifestyle interventions in the treatment of lean nonalcoholic fatty liver disease patients. Eur. J. Gastroenterol. Hepatol. 2020, 32, 1352–1357. [Google Scholar] [CrossRef]
- Jiang, X.; Yan, C.; Zhang, H.; Chen, L.; Jiang, R.; Zheng, K.; Jin, W.; Ma, H.; Liu, X.; Dong, M. Oral Probiotic Expressing Human Ethanol Dehydrogenase Attenuates Damage Caused by Acute Alcohol Consumption in Mice. Microbiol. Spectr. 2023, 11, e0429422. [Google Scholar] [CrossRef]
- Lu, J.; Zhao, Y.; Cheng, Y.; Hu, R.; Fang, Y.; Lyu, M.; Wang, S.; Lu, Z. Optimal Secretory Expression of Acetaldehyde Dehydrogenase from Issatchenkia terricola in Bacillus subtilis through a Combined Strategy. Molecules 2022, 27, 747. [Google Scholar] [CrossRef]
- Kim, J.H.; Kim, H.J.; Son, J.H.; Chun, H.N.; Yang, J.O.; Choi, S.J.; Kim, S.K. Effect of Lactobacillus fermentum MG590 on alcohol metabolism and liver function in rats. J. Microbiol. Biotechnol. 2003, 13, 919–925. [Google Scholar]
- Jung, S.J.; Hwang, J.H.; Park, E.O.; Lee, S.O.; Chung, Y.J.; Chung, M.J.; Chae, S.W. Regulation of alcohol and acetaldehyde metabolism by a mixture of Lactobacillus and Bifidobacterium species in human. Nutrients 2021, 13, 1875. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Z.; Liu, C.; Song, M.; Xu, Q.; Liu, Y.; Yan, H. Genome analysis of a newly isolated Bacillus velezensis-YW01 for biodegrading acetaldehyde. Biodegradation 2024, 35, 539–549. [Google Scholar] [CrossRef]
- Wang, Z.; He, J.J.; Liu, X.X.; Shi, H.L.; Lu, Y.F.; Shi, J.Y.; Tang, C.D. High-level expression of an acetaldehyde dehydrogenase from Lactiplantibacillus plantarum and preliminary evaluation of its potential as a functional food additive. Electron. J. Biotechnol. 2023, 63, 1–9. [Google Scholar] [CrossRef]
- Brodmann, T.; Endo, A.; Gueimonde, M.; Vinderola, G.; Kneifel, W.; de Vos, W.M.; Gómez-Gallego, C. Safety of novel microbes for human consumption: Practical examples of assessment in the European Union. Front. Microbiol. 2017, 8, 1725. [Google Scholar] [CrossRef]
- Qiu, X.M.; Zhang, Y.; Hong, H.S. Classification of acetic acid bacteria and their acid resistant mechanism. AMB Express 2021, 11, 29. [Google Scholar] [CrossRef] [PubMed]
- Cleenwerck, I.; Camu, N.; Engelbeen, K.; De Winter, T.; Vandemeulebroecke, K.; De Vos, P.; De Vuyst, L. Acetobacter ghanensis sp. nov., a novel acetic acid bacterium isolated from traditional heap fermentations of Ghanaian cocoa beans. Int. J. Syst. Evol. Microbiol. 2007, 57, 1647–1652. [Google Scholar] [CrossRef]
- Sengun, I.Y.; Kilic, G.; Charoenyingcharoen, P.; Yukphan, P.; Yamada, Y. Investigation of the microbiota associated with traditionally produced fruit vinegars with focus on acetic acid bacteria and lactic acid bacteria. Food Biosci. 2022, 47, 101636. [Google Scholar] [CrossRef]
- Okoń, A.; Łepecka, A.; Szymański, P.; Neffe-Skocińska, K. The Effect of the Use of the Beneficial Acetic Acid Bacteria Starter Cultures on the Microbiological and Physicochemical Quality of Raw Ripening Sausages. Appl. Sci. 2024, 15, 263. [Google Scholar] [CrossRef]
- Li, W.Y.; Zhou, X.G.; Peng, S.P.; Li, Y.; Liu, Q.; Liu, Q.Y.; Liu, G.H. Study on the identification of dominant strains and influence on fermentation quality of Pennisetum giganteum silage. Southwest China J. Agric. Sci. 2025, 1, 1–9. [Google Scholar]
- Doguer, C.; Akalan, H.; Tokatlı Demirok, N.; Erdal, B.; Mete, R.; Bilgen, T. Protective effects of Acetobacter ghanensis against gliadin toxicity in intestinal epithelial cells with immunoregulatory and gluten-digestive properties. Eur. J. Nutr. 2023, 62, 605–614. [Google Scholar] [CrossRef] [PubMed]
- Wen, X.; Yang, H.; Li, Z.; Chu, W. Alcohol degradation, learning, and memory-enhancing effect of Acetobacter pasteurianus BP2201 in Caenorhabditis elegans model. J. Appl. Microbiol. 2023, 134, lxad253. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Yu, P.; Lin, X.M.; Chen, X.J.; Wang, T.T.; Peng, Y.Z. Screening, identification, and anti-alcoholic and anti-drunken efficacy evaluation of ethanol-degrading lactic acid bacteria. China Brew. 2024, 43, 106–112. [Google Scholar]
- Fu, Q.; Wang, Q.; Zhou, Q.L.; Jin, Y.; Hu, M.Q.; Zhao, F.L. Screening of ethanol-degrading strains from Siraitia grosvenorii endophytes and optimization of fermentation conditions. China Brew. 2021, 40, 187–191. [Google Scholar]
- Lu, J.; Zhu, X.; Zhang, C.; Lu, F.; Lu, Z.; Lu, Y. Co-expression of alcohol dehydrogenase and aldehyde dehydrogenase in Bacillus subtilis for alcohol detoxification. Food Chem. Toxicol. 2020, 135, 110890. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, Y.; Liu, S.; Song, J.; Suo, H. Degradation effects and mechanisms of Limosilactobacillus fermentum on ethanol. Food Funct. 2024, 15, 10283–10299. [Google Scholar] [CrossRef]
- Sobh, A.; Loguinov, A.; Stornetta, A.; Balbo, S.; Tagmount, A.; Zhang, L.; Vulpe, C.D. Genome-Wide CRISPR Screening Identifies the Tumor Suppressor Candidate OVCA2 As a Determinant of Tolerance to Acetaldehyde. Toxicol. Sci. 2019, 169, 235–245. [Google Scholar] [CrossRef]
- Yao, L.Y.; Yan, R.Y.; Zheng, B.Y.; Yang, K.Y.; Guo, Z.Y.; Liu, J.; Zhao, M.L.; Wu, L.T. Optimization of Fermentation Conditions of an Acetaldehyde-Degrading Bacterium Bacillus velezensis LT-2. Microbiol. China 2023, 50, 3345–3356. [Google Scholar] [CrossRef]
- Peng, Z.; Wang, R.; Xia, X.; Zhang, J. Engineered acetaldehyde dehydrogenase for the efficient degradation of acetaldehyde. J. Environ. Manag. 2023, 331, 117258. [Google Scholar] [CrossRef]
- Corcoran, B.M.; Ross, R.P.; Fitzgerald, G.F.; Stanton, C. Comparative survival of probiotic lactobacilli spray-dried in the presence of prebiotic substances. J. Appl. Microbiol. 2004, 96, 1024–1039. [Google Scholar] [CrossRef] [PubMed]
- Lebeer, S.; Vanderleyden, J.; De Keersmaecker, S.C. Genes and molecules of lactobacilli supporting probiotic action. Microbiol. Mol. Biol. Rev. 2008, 72, 728–764. [Google Scholar] [CrossRef] [PubMed]
- Papadimitriou, K.; Alegría, Á.; Bron, P.A.; De Angelis, M.; Gobbetti, M.; Kleerebezem, M.; Lemos, J.A.; Linares, D.M.; Ross, P.; Stanton, C.; et al. Stress physiology of lactic acid bacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 837–890. [Google Scholar] [CrossRef] [PubMed]
- Parvez, S.; Malik, K.A.; Kang, S.A.; Kim, H.Y. Probiotics and their fermented food products are beneficial for health. J. Appl. Microbiol. 2006, 100, 1171–1185. [Google Scholar] [CrossRef]
- Zhao, J.Y.; Meng, Z.; Ma, X.L.; Zhao, S.M.; An, Y.; Xiao, Z.J. Characterization and regulation of the acetolactate synthase genes involved in acetoin biosynthesis in Acetobacter pasteurianus. Foods 2021, 10, 1013. [Google Scholar] [CrossRef]
- Douillard, F.P.; de Vos, W.M. Functional genomics of lactic acid bacteria: From food to health. Microb. Cell Fact. 2014, 13 (Suppl. 1), S8. [Google Scholar] [CrossRef]
- Jankovic, I.; Sybesma, W.; Phothirath, P.; Ananta, E.; Mercenier, A. Application of probiotics in food products—Challenges and new approaches. Curr. Opin. Biotechnol. 2010, 21, 175–181. [Google Scholar] [CrossRef]
- Liang, L.; Fu, Y.; Deng, S.; Wu, Y.; Gao, M. Genomic, Antimicrobial, and Aphicidal Traits of Bacillus velezensis ATR2, and Its Biocontrol Potential against Ginger Rhizome Rot Disease Caused by Bacillus pumilus. Microorganisms 2022, 10, 63. [Google Scholar] [CrossRef]
- Matsutani, M.; Matsumoto, N.; Hirakawa, H.; Shiwa, Y.; Yoshikawa, H.; Okamoto-Kainuma, A.; Ishikawa, M.; Kataoka, N.; Yakushi, T.; Matsushita, K. Comparative Genomic Analysis of Closely Related Acetobacter pasteurianus Strains Provides Evidence of Horizontal Gene Transfer and Reveals Factors Necessary for Thermotolerance. J. Bacteriol. 2020, 202, e00553-19. [Google Scholar] [CrossRef]
- Elsaadany, K.; El-Sayed, A.I.; Awad, S. Identification, Safety Assessment, and Antimicrobial Characteristics of Cocci Lactic Acid Bacteria Isolated from Traditional Egyptian Dairy Products. Foods 2024, 13, 3059. [Google Scholar] [CrossRef]
- Zhang, Q.; Wang, M.; Ma, X.; Li, Z.; Jiang, C.; Pan, Y.; Zeng, Q. In vitro investigation on lactic acid bacteria isolated from Yak faeces for potential probiotics. Front. Cell. Infect. Microbiol. 2022, 12, 984537. [Google Scholar] [CrossRef]
- Linares, D.M.; del Río, B.; Ladero, V.; Martínez, N.; Fernández, M.; Martín, M.C.; Álvarez, M.A. Factors influencing biogenic amines accumulation in dairy products. Front. Microbiol. 2012, 3, 180. [Google Scholar] [CrossRef] [PubMed]
- Yazgan, H.; Kuley, E.; Gökmen, T.G.; Regenstein, J.M.; Özogul, F. The antimicrobial properties and biogenic amine production of lactic acid bacteria isolated from various fermented food products. J. Food Process. Preserv. 2021, 45, e15085. [Google Scholar] [CrossRef]
- Tsanasidou, C.; Asimakoula, S.; Sameli, N.; Fanitsios, C.; Vandera, E.; Bosnea, L.; Koukkou, A.-I.; Samelis, J. Safety Evaluation, Biogenic Amine Formation, and Enzymatic Activity Profiles of Autochthonous Enterocin-Producing Greek Cheese Isolates of the Enterococcus faecium/durans Group. Microorganisms 2021, 9, 777. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Park, S.; Sathiyaseelan, A.; Wang, M.-H. Isolation and Characterization of Enterococcus faecium from Fermented Korean Soybean Paste with Antibacterial Effects. Fermentation 2023, 9, 760. [Google Scholar] [CrossRef]


| Gene ID | Gene | Product |
|---|---|---|
| ctg00001_00033 | frmA | S-(hydroxymethyl)glutathione dehydrogenase |
| ctg00005_01632 | adh | Alcohol dehydrogenase |
| ctg00009_02255 | ald1 | Long-chain aldehyde dehydrogenase |
| ctg00001_00447 | adhA1 | Putative alcohol dehydrogenase AdhA |
| ctg00013_02492 | yqhD | Alcohol dehydrogenase YqhD |
| ctg00002_00717 | fdhC1 | Fructose dehydrogenase cytochrome subunit |
| ctg00002_00718 | iorB | Isoquinoline 1-oxidoreductase subunit beta |
| ctg00002_00719 | iorA2 | Isoquinoline 1-oxidoreductase subunit alpha |
| ctg00001_00039 | pta | Phosphate acetyltransferase |
| ctg00001_00040 | pduW | Putative propionate kinase |
| Function | Gene | Product |
|---|---|---|
| Acid stress | aspS | Aspartate-tRNA (Asp/Asn) ligase |
| groS | 10 kDa chaperonin | |
| recA | Protein RecA | |
| aspA | Aspartate ammonia-lyase | |
| Acid stress/bile resistance | dnaA | Chromosomal replication initiator protein DnaA |
| dnaG | DNA primase | |
| atpF_1 | ATP synthase subunit b | |
| atpF_2 | ATP synthase subunit b | |
| atpB | ATP synthase subunit a | |
| atpC | ATP synthase epsilon chain | |
| atpD | ATP synthase subunit beta | |
| atpA atpG | ATP synthase subunit alpha | |
| ATP synthase gamma chain | ||
| Bile resistance | pyrG | CTP synthase |
| satP | Succinate-acetate/proton symporter SatP | |
| argS | Arginine-tRNA ligase | |
| Antimicrobial properties | tyrS | Tyrosine-tRNA ligase |
| alsD | Alpha-acetolactate decarboxylase |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, H.; Wang, J.; Xu, Q.; Cao, X.; Du, X.; Lin, K.; Yan, H. Biodegradation of Both Ethanol and Acetaldehyde by Acetobacter ghanensis JN01. Catalysts 2025, 15, 756. https://doi.org/10.3390/catal15080756
Liu H, Wang J, Xu Q, Cao X, Du X, Lin K, Yan H. Biodegradation of Both Ethanol and Acetaldehyde by Acetobacter ghanensis JN01. Catalysts. 2025; 15(8):756. https://doi.org/10.3390/catal15080756
Chicago/Turabian StyleLiu, Hongyan, Jingjing Wang, Qianqian Xu, Xiaoyu Cao, Xinyue Du, Kun Lin, and Hai Yan. 2025. "Biodegradation of Both Ethanol and Acetaldehyde by Acetobacter ghanensis JN01" Catalysts 15, no. 8: 756. https://doi.org/10.3390/catal15080756
APA StyleLiu, H., Wang, J., Xu, Q., Cao, X., Du, X., Lin, K., & Yan, H. (2025). Biodegradation of Both Ethanol and Acetaldehyde by Acetobacter ghanensis JN01. Catalysts, 15(8), 756. https://doi.org/10.3390/catal15080756






