Review on Techniques for Plant Leaf Classification and Recognition
Abstract
1. Introduction
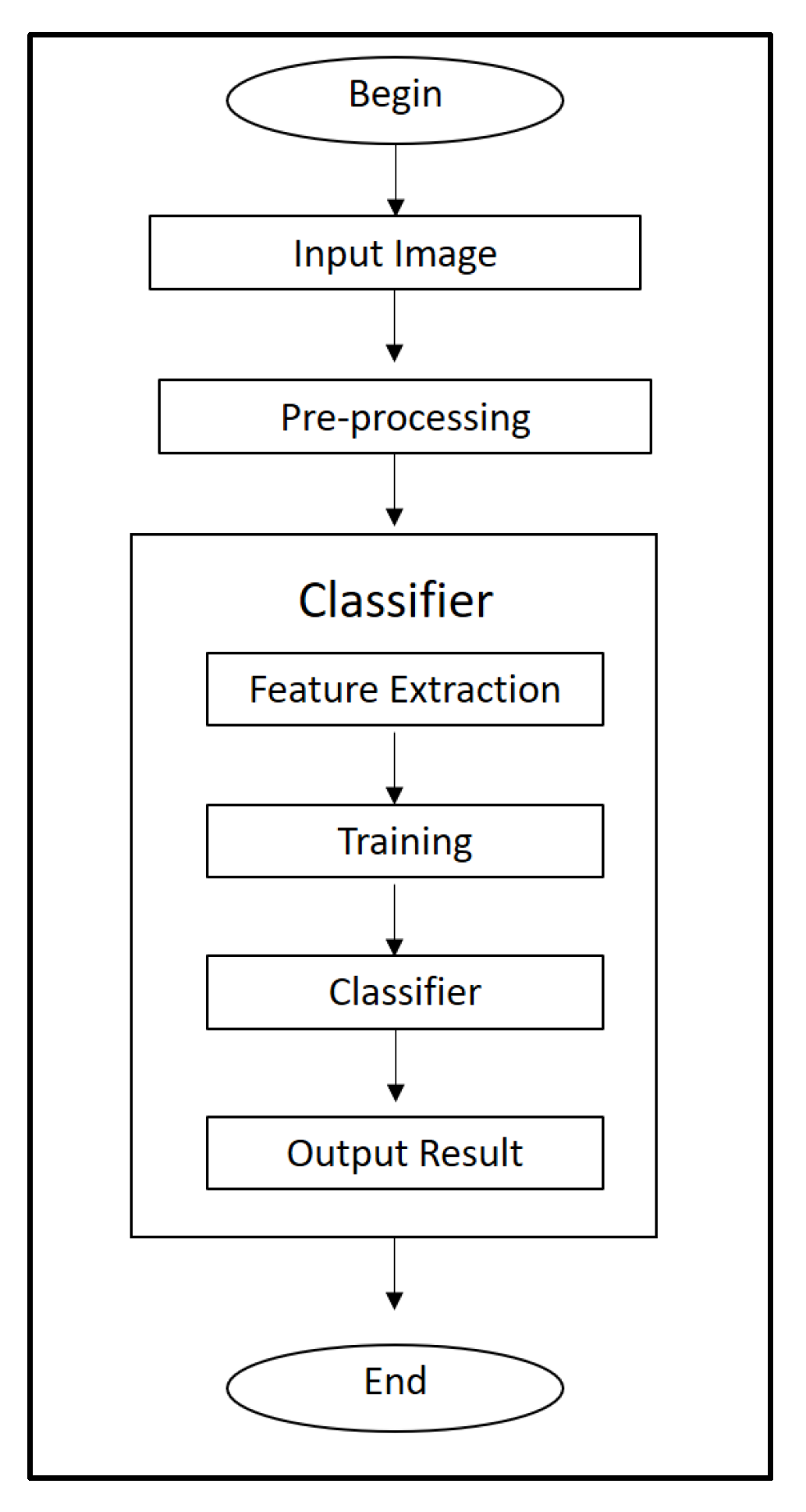
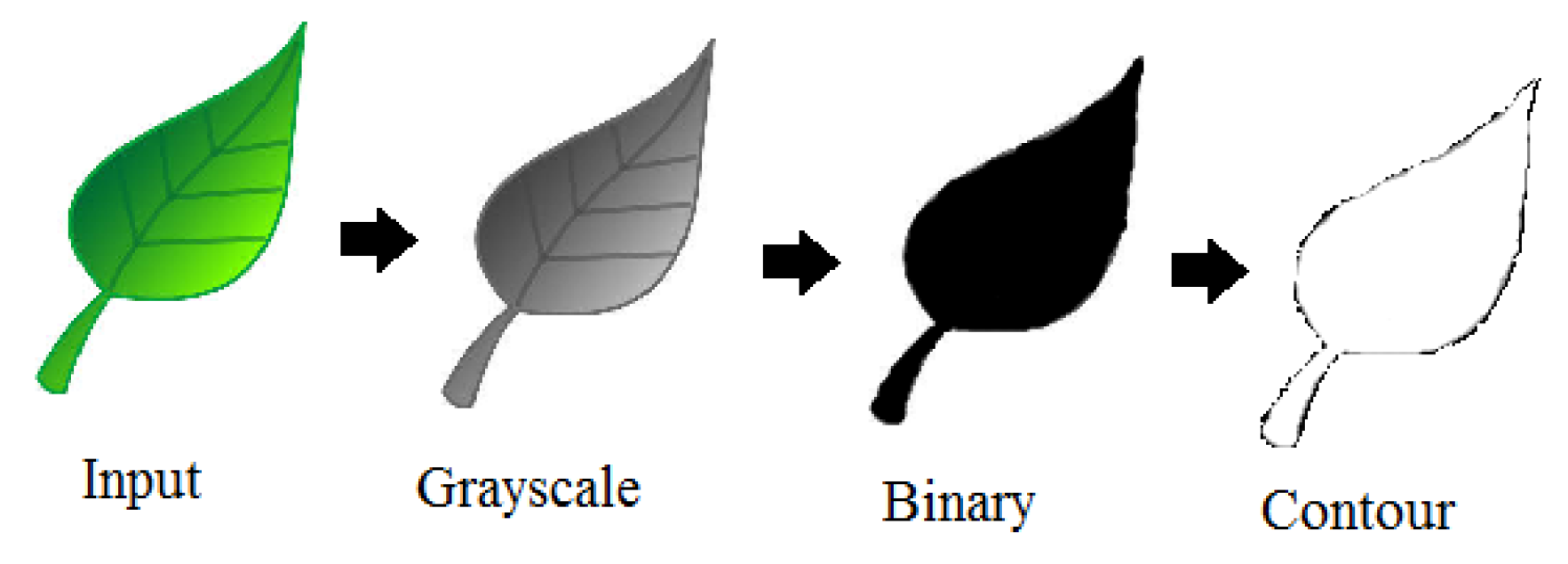
2. Image Processing in Leaf Pattern Recognition
3. Leaf Feature Extraction
4. Mathematical Classifiers
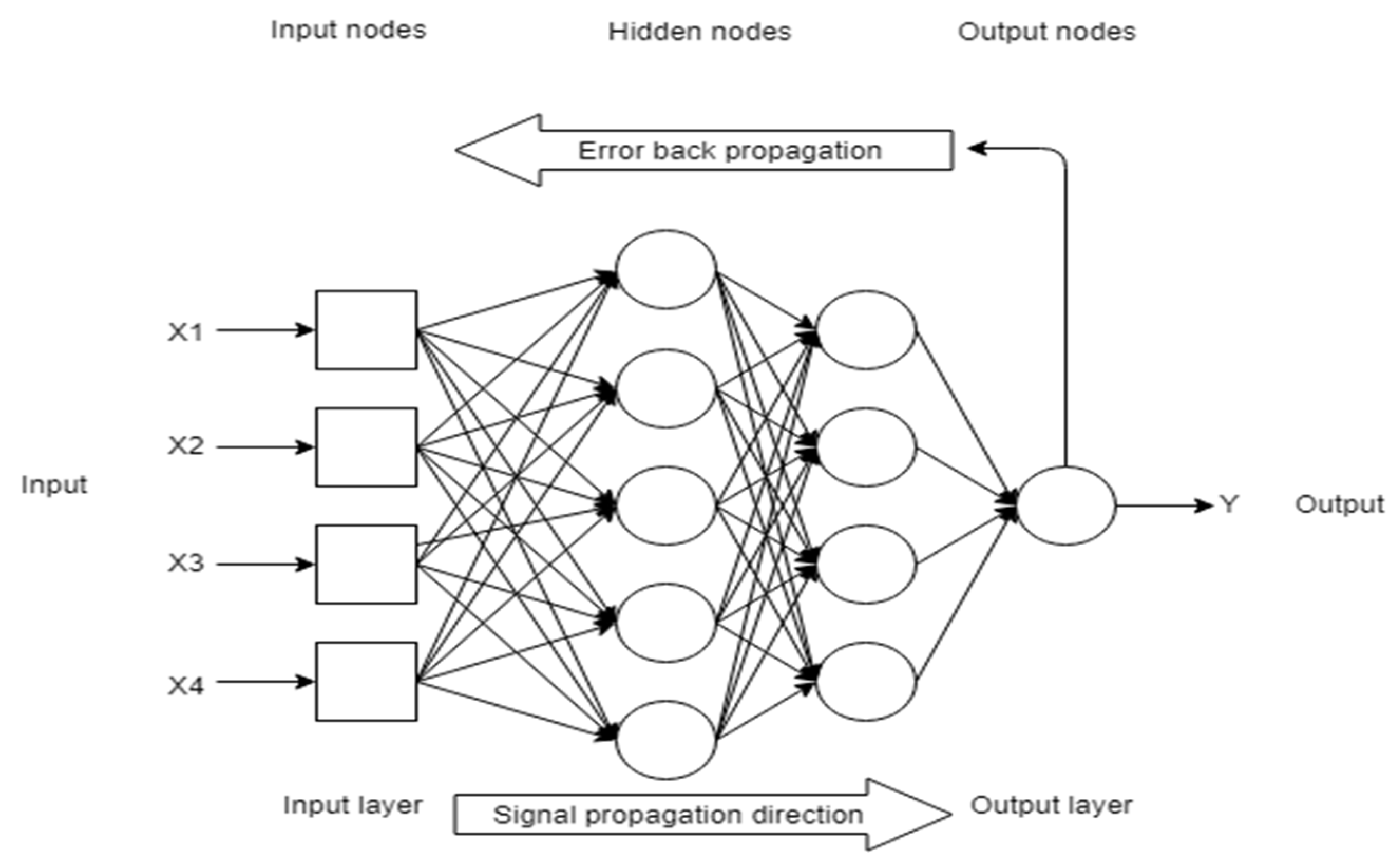
4.1. Artificial Neural Network (ANN)
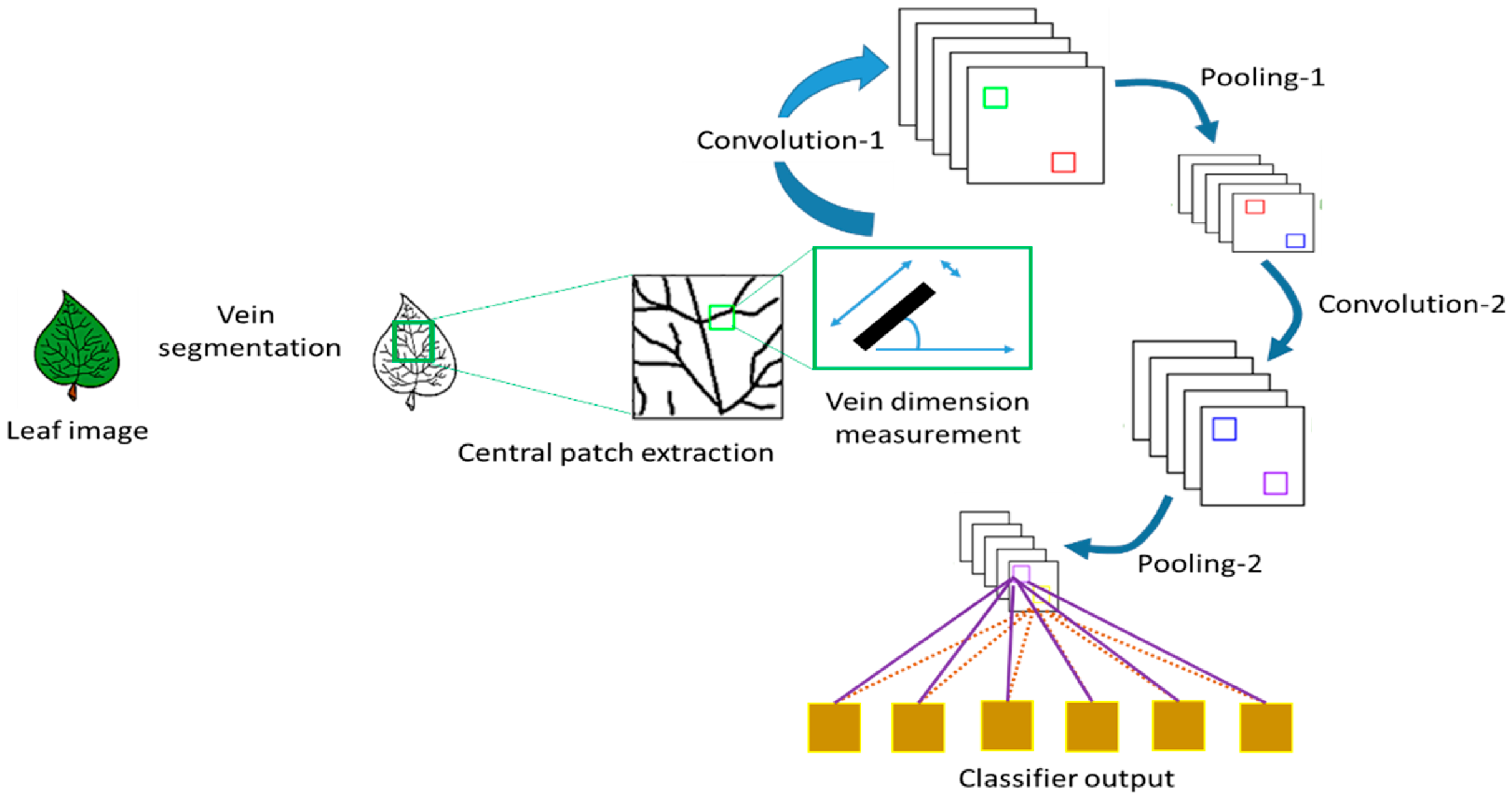
4.2. Convolutional Neural Network (CNN)
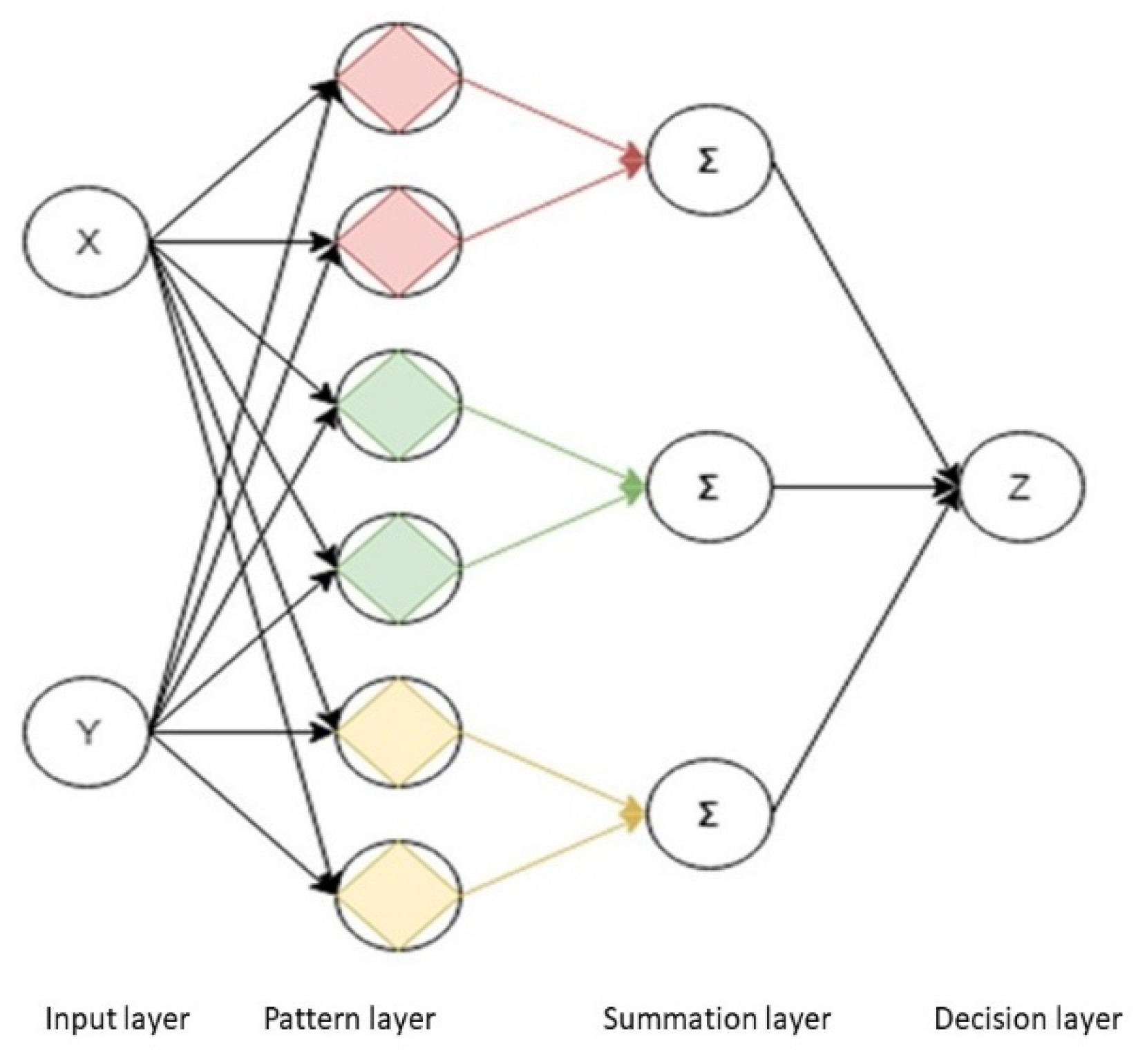
4.3. Probabilistic Neural Network (PNN)
4.4. Support Vector Machine (SVM)
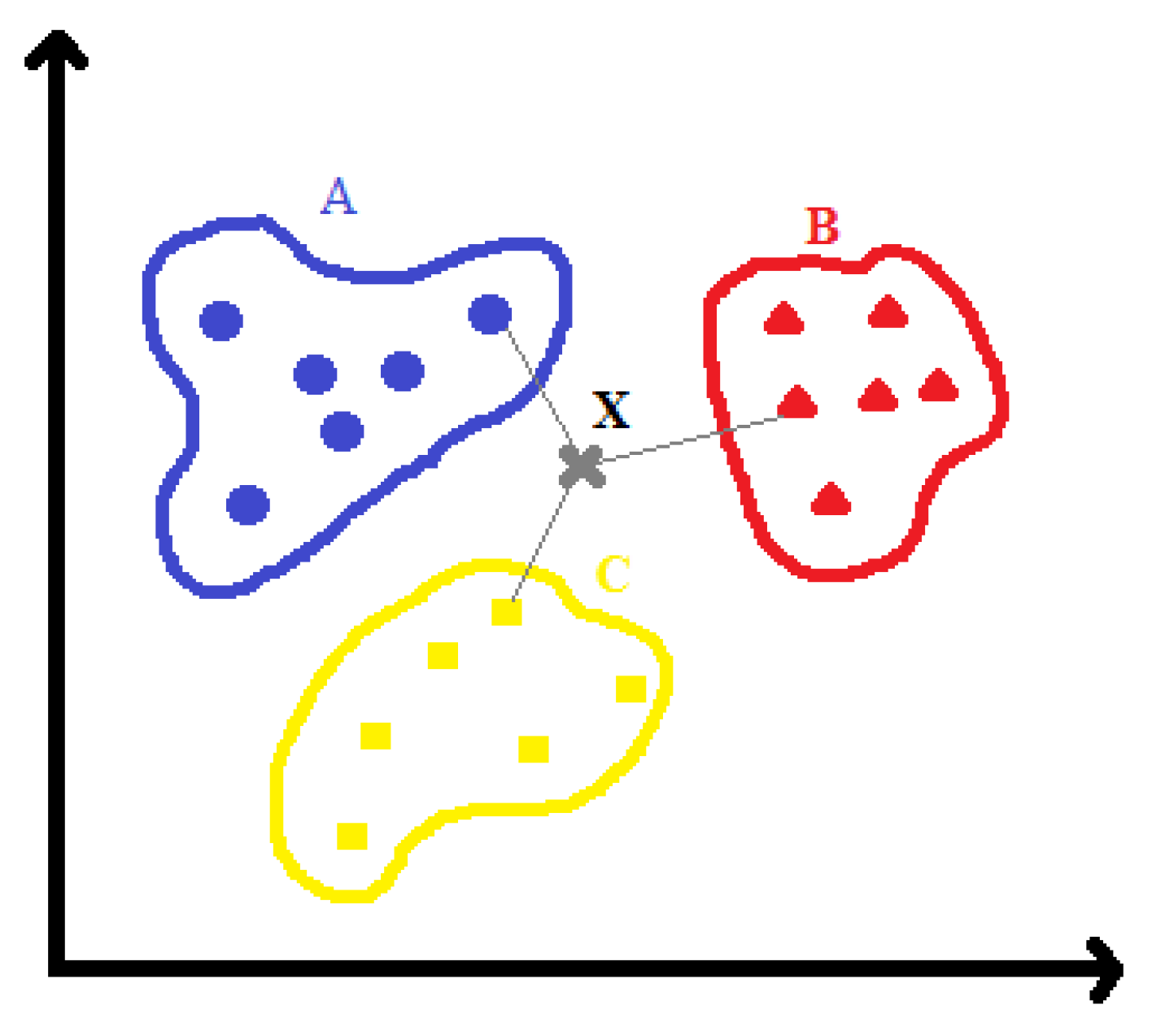
4.5. K-Nearest Neighbor (KNN)
5. Pattern Recognition Method
6. Conclusions and Recommendation
Author Contributions
Funding
Conflicts of Interest
Appendix A
| No. | Goals | Method | Results | References |
|---|---|---|---|---|
| Artificial Neural Network (ANN) | ||||
| 1 | Identification Of Selected Medicinal Plant Leaves Using Image Features And ANN | Method utilizes feature extraction based on shape, color, and texture of leaf images and training via ANN classifier for leaf classification of the system. The image processing and neural network toolboxes were obtained from MATLAB. | Implementation of 8 input features into the system were found to be optimal in terms of the complexity as it demands minimum input as well as less computational time required. The accuracy of the system was 94.4% against 63 leaf images in the dataset. | [30] |
| 2 | Neural Network Based Leaf Recognition | Implementation of Pulse Coupled Neural Network into the feature extraction of leaf images to obtain the entropy sequence, aspect ratio, Zernike moments, Hu’s invariants, form factor, rectangularity, circularity, and area. The classifier used is Artificial Neural Network. | The accuracy of the system proposed was better compared to the other methods in the study, where the entropy was taken as the key feature in the leaf classification, yielding an accuracy of 91.98%. | [63] |
| 3 | Combined Thresholding nnd Neural Network Approach for Vein Pattern Extraction From Leaf Images | A proposed model that extracts leaf vein patterns via combining thresholding methods and an ANN classifier. | The experimental results of the method proposed have shown to possess the capability of extracting a more accurate venation pattern of the leaf samples for pattern recognition with 97.3% accuracy while reducing computing time as well compared to direct neural network approach which achieved an accuracy of 84.4%. | [31] |
| 4 | Classification of Plant Leaves using Morphological Features and Zernike Moments | The proposed model utilizes leaf morphological features and Zernike moments which are independent of leaf growth and image translation, rotation and scaling and then classified with several different classifiers to achieve optimal result. | The dataset tested were Flavia dataset of 32 species (1907 images, 50-60 leaves each species) and a medicinal plants dataset of 6 species (30 images each). SVM and PNN have comparably higher accuracy than k-NN and naïve Bayes classifiers due to the latter being lazy learners algorithms. | [65] |
| Convolutional Neural Network (CNN) | ||||
| 5 | Leaf Classification based on Marginalized Shape Context and Shape-Texture Dual-Path Deep Convolutional Neural Network | Utilizing a dual-path deep CNN to learn leaf image features and optimize them for classification compared to vanilla convolutional network systems. | Dual-path CNN method outperforms various other CNN methods, being: uni-patch CNN, texture-patch CNN, marginalized shape context with SVM classifier, Multiscale Distance Matrix with SVM classifier, curvature histogram, giving near-perfect-top-1-match results on Flavia dataset and top-3-match results on all other datasets. | [60] |
| Probabilistic Neural Network (PNN) | ||||
| 6 | Mobile Application for Indonesian Medicinal Plants Identification using Fuzzy Local Binary Pattern and Fuzzy Color Histogram | Utilizing the combined methods of Fuzzy Local Binary Pattern (FLBP) and the Fuzzy Color Histogram (FCH) in order to identify the medicinal plants found in Indonesia. The combination was done via Product Decision Rule (PDR) method, and is capable of extracting leaf image texture and color. The classifier utilized was PNN classifier. | Fusion of methods FLBP and FCH has shown to be capable of increasing the overall accuracy of plant identification methods and yielded an accuracy of 74.51%. The accuracy of the system without the fusion features were 59.61% for FLBP features and 50.78% for FCH feature respectively. | [42] |
| 7 | Plant Leaf Classification Using Centroid Distance and Axis of Least Inertia Method | The utilization of centroid distance and axis of least inertia for plant classification. Canary operator was implemented for the binary images converted from RGB (Red, Green, Blue) to detect and thin out the edges of the leaf images before the shape is traced. The centroid distances of these points as well as the distance of sampling points from the axis of least inertial lines were computed. The classifier used was the PNN classifier for two public leaf databases, Flavia and the Swedish leaf dataset. | Combination of both these methods, which are invariant to translation, rotation and scale into the extraction of feature vectors for leaf image classification has been proven to be able to improve the accuracy. Experimentation results have shown that the accuracy of the system from the Flavia dataset was 82.05% and Swedish leaf dataset was 80.10%, an improvement from previous methods proposed. | [44] |
| Support Vector Machine (SVM) | ||||
| 8 | Combined Classifier for Plant Classification and Identification from Leaf Image based on Visual Attribute | The identification of the plant images based on varying features such as shape, color, texture, and vein pattern with combined classifier using majority voting technique has been proposed for recognizing leaf’s category along with SVM, Naïve Bay’s and Decision tree as the combined classifiers | Utilization of the combined classifiers with several features extraction of the leaf images have been shown to yield maximum accuracy output as compared to them individually and all existing methodologies with accuracy of 93.11% while with separately, SVM is 91.8576, Naïve Bayes is 88.2736%, and Decision tree is 85.6678% respectively. | [52] |
| 9 | Comparative Analysis of Leaf Classification and Recognition by different SVM Classifiers | A new approach proposed by extracting 14 features of leaf by utilizing shape detector and SVM as the classifier of the system | 16 different plants species from the Flavia database comprising of 480 images were used as the training dataset with 14 features extracted for optimal performance. Quadratic SVM yielded the highest accuracy of 90.9% as compared to Medium Gaussian SVM and Cubic SVM gave 89.4% and 89.8% test accuracy separately. | [48] |
| 10 | Multiple Classifier System for Plant Leaf Recognition | The methodology presents a multiple classifier system (MCS) of plant identification based on texture and shape features of the leaf images. SVM and Neural Network classifiers are trained on four different feature sets, namely, Local Binary Pattern (LBP), Histogram of Gradients (HOG), Speed of Robust Features (SURF) and Zernike Moments (ZM). Then, a static classifiers selection method is used to search for the ensembles that maximize the average classification score. | ImageCLEF 2011 and ImageCLEF 2012 datasets were used for the experiments with the proposed MCS methods showing an improvement of 11.76% and 4.23% in the scan category and 4.06% and 5.87% for the scan-like category in the average classification score relative to the best results reported in the literature for ImageCLEF 2011 and ImageCLEF 2012 datasets respectively. MCS approach also overcomes the performances of monolithic methodologies compared in the study. | [49] |
| 11 | Identifying leaf in a natural image using morphological characters | Proposed methodology of plant detection in their natural habitat, designed in an environment with heavy interference and overlapping. The images are captured and segmented using marker-controlled watershed segmentation which is generated automatically using morphological operations and the shapes obtained are converted into features via Hu moments. The classifier utilized is SVM classifier. | 3 species with 300 different leaf image samples were used for the experimentation of the system and captured in real time with 86.7% identification accuracy. The accuracy of the system can be further improved by the addition of more features extracted via the system’s operation as well as increasing the dataset used for the experiment. | [54] |
| k-Nearest Neighbor (KNN) | ||||
| 12 | Recognition of Whole and Deformed Plant Leaves using Statistical Shape Features and Neuro-Fuzzy Classifier | A method-logiical recognition of 32 pre-defined classes of plants using a Neuro-fuzzy classifier and comparing it to other classifiers such as k-Nearest Neighbor and Neural Network classifiers. | 640 leaf sample images of varying sizes, shapes, and orientation were tested in the experiment and have been proven to show improvements compared to k-NNC and NNC with an accuracy of 97.5% and 79.7%.A limitation of the proposed method is that it works reliably only when deformations do not alter the major and minor axis lengths. | [62] |
| 13 | Leaf Classification based on Shape and Edge feature with k-NN Classifier | A proposed approach via the utilization of edge based and shape based features extraction for the classification of leaf images. | The method was tested on Flavia dataset of 32 plant species and yielded an average classification accuracy rate of 94.4%, an improvement compared to existing methods, with size and rotation invariant being the novelty of the algorithm. | [59] |
| No Classifier | ||||
| 14 | Leaf Recognition and Classification using GLCM and Hierarchical Centroid Based Technique | Utilization of various techniques for preprocessing, feature extraction, and classification of leaf based on shape and texture, via Gray Level Co-occurrence Matrix and hierarchical centroid based technique. | Gray Level Co-occurrence Matrix, Discrete Wavelet Transform, and hierarchical centroid have been proven to be able to obtain the special information of leaves accurately and improve on the accuracy. 300 leaf samples of 30 different species from Flavia dataset were used for the experiment and the model achieved an accuracy of 96.66%. | [61] |
| 15 | Plant Leaf Recognition Using a Layered Approach | A proposed methodology using layered architecture with each layer handling a specific type of visual characteristics using a set of features to form a customized data model. The different layers are then fed into an array of custom classifiers for robust recognition. Color based modeling is used for non-green leaves while shape based modeling is used for green simple and compound leaves, using a layered approach. | The database of 600 leaf images from 30 different classes of leaves were used to test the layered model and yielded an accuracy of over 90% for all the different approaches. The utilization of different visuals, segmentation to reduce computational load, different customization in the classifiers, and addition of different layers make the model more robust than all the other models proposed in previous studies. | [64] |
| 16 | Leaf Plant Identification System based on Hidden naïve Bays Classifier | A proposed computational model in plant identification system of digital images of plants utilizing biometric features such as shape and vein patterns via hidden naïve Bays classifier. | 1907 leaf image samples from 32 different species of plants taken from Flavia dataset were used in the study in determining the accuracy of the system and the model has proven to have an average identification accuracy of 97% based on 10 different biometric information extracted. | [53] |
| 17 | Leaf species classification based on a botanical shape sub-classifier strategy | The implementation of fusion strategy and its corresponding random-forest-based sub-classifiers as part of a leaf recognition system. | The fusion technique provides a significant accuracy enhancement compared to other proposed methods, while providing necessary information for educational purposes. | [43] |
References
- Hoffman, H.J.; Cruickshanks, K.J.; Davis, B. Perspectives on population-based epidemiological studies of olfactory and taste impairment. Ann. N. Y. Acad. Sci. 2009, 1170, 514. [Google Scholar] [CrossRef] [PubMed]
- Shivling, V.D.; Singla, A.; Ghanshyam, C.; Kapur, P.; Gupta, S. Plant leaf imaging technique for agronomy. In Proceedings of the 2011 International Conference on Image Information Processing, Shimla, India, 3–5 November 2011; pp. 1–5. [Google Scholar]
- Hossain, J.; Amin, M.A. Leaf shape identification based plant biometrics. In Proceedings of the 2010 13th International Conference on Computer and Information Technology (ICCIT), Dhaka, Bangladesh, 23–25 December 2010; pp. 458–463. [Google Scholar]
- Khmag, A.; Al-Haddad, S.A.R.; Kamarudin, N. Recognition system for leaf images based on its leaf contour and centroid. In Proceedings of the 2017 IEEE 15th Student Conference on Research and Development (SCOReD), Putrajaya, Malaysia, 13–14 December 2017; pp. 467–472. [Google Scholar]
- Wäldchen, J.; Rzanny, M.; Seeland, M.; Mäder, P. Automated plant species identification-trends and future directions. PLoS Comput. Biol. 2018, 14, e1005993. [Google Scholar]
- Sabu, A.; Sreekumar, K. Literature review of image features and classifiers used in leaf based plant recognition through image analysis approach. In Proceedings of the 2017 International Conference on Inventive Communication and Computational Technologies (ICICCT), Coimbatore, India, 10–11 March 2017; pp. 145–149. [Google Scholar]
- Wu, S.G.; Bao, F.S.; Xu, E.Y.; Wang, Y.; Chang, Y.; Xiang, Q. A Leaf Recognition Algorithm for Plant Classification Using Probabilistic Neural Network. In Proceedings of the 2007 IEEE International Symposium on Signal Processing and Information Technology, Giza, Egypt, 15–18 December 2007; pp. 11–16. [Google Scholar]
- Singh, V.; Misra, A.K. Detection of plant leaf diseases using image segmentation and soft computing techniques. Inf. Process. Agric. 2017, 4, 41–49. [Google Scholar] [CrossRef]
- Chaki, J.; Parekh, R.; Bhattacharya, S. Plant leaf classification using multiple descriptors: A hierarchical approach. J. King Saud Univ. Comp. Inf. Sci. 2018, in press. [Google Scholar] [CrossRef]
- Munisami, T.; Ramsurn, M.; Kishnah, S.; Pudaruth, S. Plant Leaf Recognition Using Shape Features and Colour Histogram with K-nearest Neighbour Classifiers. Procedia Comput. Sci. 2015, 58, 740–747. [Google Scholar] [CrossRef]
- Turkoglu, M.; Hanbay, D. Recognition of plant leaves: An approach with hybrid features produced by dividing leaf images into two and four parts. Appl. Math. Comput. 2019, 352, 1–14. [Google Scholar] [CrossRef]
- Ma, L.; Fang, J.; Chen, Y.; Gong, S. Color analysis of leaf images of deficiencies and excess nitrogen content in soybean leaves. In Proceedings of the 2010 International Conference on E-Product E-Service and E-Entertainment, Henan, China, 7–9 November 2010; Volume 11541023, pp. 1–3. [Google Scholar]
- Gonzalez, R.C.; Woods, R.E. Digital Image Processing, 3rd ed.; Pearson Prentice Hall: Upper Saddle River, NJ, USA, 2002. [Google Scholar]
- Kittler, J.; Illingworth, J. Minimum error thresholding thresholding minimum error decision rule Classification error Dynamic clustering. Pattern Recognit. 1986, 19, 41–47. [Google Scholar] [CrossRef]
- ASM International. Practical Guide to Image Analysis; ASM International: Novelty, OH, USA, 2000. [Google Scholar]
- Bo, Z.; Hua, W.H.; Jun, L.S.; Hua, M.W.; Chao, Z.X. Research on weed recognition method based on invariant moments. In Proceeding of the 11th World Congress on Intelligent Control and Automation, Shenyang, China, 29 June–4 July 2014; pp. 2167–2169. [Google Scholar]
- Achanta, R.; Shaji, A.; Smith, K.; Lucchi, A.; Fua, P.; Süsstrunk, S. SLIC superpixels compared to state-of-the-art superpixel methods. IEEE Trans. Pattern Anal. Mach. Intell. 2012, 34, 2274–2281. [Google Scholar] [CrossRef]
- Cerutti, G.; Tougne, L.; Mille, J.; Vacavant, A.; Coquin, D. Understanding leaves in natural images—A model-based approach for tree species identification. Comput. Vis. Image Underst. 2013, 117, 1482–1501. [Google Scholar] [CrossRef]
- Chan, T.F.; Vese, L.A. Donald Middleton, L.D.S. Br. Dent. J. 1977, 10, 266–277. [Google Scholar]
- Kurtz, C.; Passat, N.; Gançarski, P.; Puissant, A. Extraction of complex patterns from multiresolution remote sensing images: A hierarchical top-down methodology. Pattern Recognit. 2012, 45, 685–706. [Google Scholar] [CrossRef]
- Grand-Brochier, M.; Vacavant, A.; Cerutti, G.; Kurtz, C.; Weber, J.; Tougne, L. Tree leaves extraction in natural images: Comparative study of preprocessing tools and segmentation methods. IEEE Trans. Image Process. 2015, 24, 1549–1560. [Google Scholar] [CrossRef] [PubMed]
- Couprie, C.; Grady, L.J.; Najman, L.; Talbot, H. Power watersheds: A new image segmentation framework extending graph cuts, random walker and optimal spanning forest. In Proceedings of the 2009 IEEE 12th International Conference on Computer Vision, Kyoto, Japan, 29 September–2 October 2009; Volume 9, pp. 731–738. [Google Scholar]
- Lü, C.; Ren, H.; Zhang, Y.; Shen, Y. Leaf area measurement based on image processing. In Proceedings of the 2010 International Conference on Measuring Technology and Mechatronics Automation, Changsha, China, 13–14 March 2010; Volume 2, pp. 580–582. [Google Scholar]
- Gopal, A.; Prudhveeswar Reddy, S.; Gayatri, V. Classification of selected medicinal plants leaf using image processing. In Proceedings of the 2012 International Conference on Machine Vision and Image Processing (MVIP), Taipei, Taiwan, 14–15 December 2012; pp. 5–8. [Google Scholar]
- Bama, B.S.; Valli, S.M.; Raju, S.; Kumar, V.A. Content Based Image Retrieval Using Advanced Color and Texture Features. Int. J. Comput. Appl. 2011, 2, 10–14. [Google Scholar]
- Ni, F.; Wang, B. Integral contour angle: An invariant shape descriptor for classification and retrieval of leaf images. In Proceedings of the 2018 25th IEEE International Conference on Image Processing (ICIP), Athens, Greece, 7–10 October 2018; pp. 1223–1227. [Google Scholar]
- Kpalma, K.; Yang, M.; Ronsin, J. Planar shapes descriptors based on the turning angle scalogram. Int. Conf. Image Anal. Recognit. 2008, 5112, 547–556. [Google Scholar]
- How Artificial Intelligence Has Switched-Up the SEO Game—More than you Imagined. 2016. Available online: http://backlinksforseo.com/artificial-intelligence-switched-up-seo/ (accessed on 31 January 2018).
- Wu, Q.; Zhou, C.; Wang, C. Feature Extraction and Automatic Recognition of Plant Leaf Using Artificial Neural Network. Adv. Artif. Intell. 2006, 3, 5–12. [Google Scholar]
- Janani, R.; Gopal, A. Identification of selected medicinal plant leaves using image features and ANN. In Proceedings of the 2013 International Conference on Advanced Electronic Systems (ICAES), Pilani, India, 21–23 September 2013; pp. 238–242. [Google Scholar]
- Fu, H.; Chi, Z. Combined thresholding and neural network approach for vein pattern extraction from leaf images. IEE Proc. Vis. Image Signal Process. 2006, 153, 881–892. [Google Scholar] [CrossRef]
- Grinblat, G.L.; Uzal, L.C.; Larese, M.G.; Granitto, P.M. Deep learning for plant identification using vein morphological patterns. Comput. Electron. Agric. 2016, 127, 418–424. [Google Scholar] [CrossRef]
- Vaidya, A.; Pujari, D.; Desai, H.; Borse, K. Leaf Recognition—A Technical Review. Int. J. Res. Emerg. Sci. Technol. 2015, 2, 46–51. [Google Scholar]
- Hang, J.; Zhang, D.; Chen, P.; Zhang, J.; Wang, B. Classification of Plant Leaf Diseases Based on Improved Convolutional Neural Network. Sensors 2019, 19, 4161. [Google Scholar] [CrossRef]
- Sibiya, M.; Sumbwanyambe, M. A Computational procedure for the recognition and classification of maize leaf diseases out of healthy leaves using convolutional neural networks. AgriEngineering 2019, 1, 119–131. [Google Scholar] [CrossRef]
- Toda, Y.; Okura, F. How convolutional neural networks diagnose plant disease. Plant Phenomics 2019, 2019, 9237136. [Google Scholar] [CrossRef]
- Jeon, W.S.; Rhee, S.Y. Plant Leaf Recognition Using a Convolution Neural Network. Int. J. Fuzzy Log. Intell. Syst. 2017, 17, 26–34. [Google Scholar] [CrossRef]
- Zeiler, M.D.; Fergus, R. Stochastic Pooling for Regularization of Deep Convolutional Neural Networks. arXiv 2013, arXiv:1301.3557. [Google Scholar]
- Lin, H.; Peng, H. Machine recognition for broad-leaved trees based on synthetic features of leaves using probabilistic neural network. In Proceedings of the 2008 International Conference on Computer Science and Software Engineering, Hubei, China, 12–14 Decemner 2008; Volume 4, pp. 871–877. [Google Scholar]
- McKenzie, J.S.; Jurado, J.M.; de Pablos, F. Characterisation of tea leaves according to their total mineral content by means of probabilistic neural networks. Food Chem. 2010, 123, 859–864. [Google Scholar] [CrossRef]
- Sawant, S.S.; Topannavar, P.S. Introduction to Probabilistic Neural Network—Used for Image Classifications. Int. J. Adv. Res. Comput. Sci. Softw. Eng. 2015, 5, 279–283. [Google Scholar]
- Herdiyeni, Y.; Kadek, N.; Wahyuni, S. Mobile Application for Indonesian Medicinal Plants Identification using Fuzzy Local Binary Pattern and Fuzzy Color Histogram. In Proceedings of the 2012 International Conference on Advanced Computer Science and Information Systems (ICACSIS), Depok, Indonesia, 1–2 December 2012; pp. 978–979. [Google Scholar]
- Liu, H.; Coquin, D.; Valet, L.; Cerutti, G. Leaf species classification based on a botanical shape sub-classifier strategy. In Proceedings of the 2014 22nd International Conference on Pattern Recognition, Stockholm, Sweden, 24–28 August 2014; Volume 5, pp. 1496–1501. [Google Scholar]
- Mahdikhanlou, K.; Ebrahimnezhad, H. Plant leaf classification using centroid distance and axis of least inertia method. In Proceedings of the 2014 22nd Iranian Conference on Electrical Engineering (ICEE), Tehran, Iran, 20–22 May 2014; pp. 1690–1694. [Google Scholar]
- Lukic, M.; Tuba, E.; Tuba, M. Leaf recognition algorithm using support vector machine with Hu moments and local binary patterns. In Proceedings of the 2017 IEEE 15th International Symposium on Applied Machine Intelligence and Informatics, Herl′any, Slovakia, 26–28 January 2017; pp. 000485–000490. [Google Scholar]
- Zhang, H.; Yanne, P.; Liang, S. Plant species classification using leaf shape and texture. In Proceedings of the 2012 International Conference on Industrial Control and Electronics Engineering, Xi′an, China, 23–25 August 2012; pp. 2025–2028. [Google Scholar]
- Salman, A.; Semwal, A.; Bhatt, U.; Thakkar, V.M. Leaf classification and identification using Canny Edge Detector and SVM classifier. In Proceedings of the 2017 International Conference on Inventive Systems and Control (ICISC), Coimbatore, India, 19–20 January 2017; pp. 1–4. [Google Scholar]
- Srivastava, V.; Khunteta, A. Comparative Analysis of Leaf Classification and Recognition by Different SVM Classifiers. In Proceedings of the 2018 International Conference on Inventive Research in Computing Applications (ICIRCA), Coimbatore, India, 11–12 July 2018; pp. 626–631. [Google Scholar]
- Aráujo, V.; Britto, A.S.; Brun, A.L.; Koerich, A.L.; Falate, R. Multiple classifier system for plant leaf recognition. In Proceedings of the 2017 IEEE International Conference on Systems, Man, and Cybernetics (SMC), Banff, AB, Canada, 5–8 October 2017; pp. 1880–1885. [Google Scholar]
- Goeau, H.; Bonnet, P.; Joly, A.; Nozha, B.; Daniel, B.; Jean-Francois, M.; Philippe, B.; Elise, M.; Marie, P. The Image CLEF 2011 plant images classification task. In Proceedings of the CLEF’2011: Conference and Labs of the Evaluation Forum, Amsterdam, The Netherlands, 19–22 September 2011; pp. 1–23. [Google Scholar]
- Goeau, H.; Bonnet, P.; Joly, A.; Yahiaoui, I.; Barthelemy, D.; Boujemaa, N.; Molino, J.F. The Image CLEF 2012 plant identification task. In Proceedings of the CLEF’2012: Conference and Labs of the Evaluation Forum, Rome, Italy, 17–20 September 2012. [Google Scholar]
- Mittal, P.; Kansal, M.; Jhajj, H.K. Combined classifier for plant classification and identification from leaf image based on visual attributes. In Proceedings of the 2018 International Conference on Intelligent Circuits and Systems (ICICS), Phagwara, India, 19–20 April 201; pp. 188–194.
- Eid, H.F.; Hassanien, A.E.; Kim, T.H. Leaf Plant Identification System Based on Hidden Naïve Bays Classifier. In Proceedings of the 2015 4th International Conference on Advanced Information Technology and Sensor Application (AITS), Harbin, China, 21–23 August 2015; pp. 76–79. [Google Scholar]
- James Nesaratnam, R.; Balamurugan, C. Identifying leaf in a natural image using morphological characters. In Proceedings of the 2015 International Conference on Innovations in Information, Embedded and Communication Systems (ICIIECS), Coimbatore, India, 19–20 March 2015. [Google Scholar]
- Ehsanirad, A. Plant Classification Based on Leaf Recognition. Int. J. Comput. Sci. Inf. Secur. 2010, 8, 78–81. [Google Scholar]
- Zhang, S.; Chau, K. Dimension Reduction Using Semi-Supervised Locally Linear Embedding for Plant Leaf Classification. Int. Conf. Intell. Comput. 2009, 5754, 948–955. [Google Scholar]
- Kherkhah, F.M.; Asghari, H. Plant leaf classification using GIST texture features. IET Comput. Vis. 2019, 13, 369. [Google Scholar] [CrossRef]
- Rahmani, M.E.; Amine, A.; Hamou, M.R. Plant Leaves Classification. ALLDATA 2015, 82, 75–80. [Google Scholar]
- Kumar, P.S.; Rao, K.N.V.; Raju, A.S.N.; Kumar, D.J.N. Leaf classification based on shape and edge feature with k-NN classifier. In Proceedings of the 2016 2nd International Conference on Contemporary Computing and Informatics (IC3I), Noida, India, 14–17 December 2016; pp. 548–552. [Google Scholar]
- Shah, M.P.; Singha, S.; Awate, S.P. Leaf classification using marginalized shape context and shape+ texture dual-path deep convolutional neural network. In Proceedings of the 2017 IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 860–864. [Google Scholar]
- Pankaja, K.; Suma, V. Leaf Recognition and Classification Using GLCM and Hierarchical Centroid Based Technique. In Proceedings of the 2018 International Conference on Inventive Research in Computing Applications (ICIRCA), Coimbatore, India, 11–12 July 2018; pp. 1190–1194. [Google Scholar]
- Chaki, J.; Parekh, R.; Bhattacharya, S. Plant leaf recognition using texture and shape features with neural classifiers. Pattern Recognit. Lett. 2015, 58, 61–68. [Google Scholar] [CrossRef]
- Wable, P.B.; Chilveri, P.G. Neural network based leaf recognition. In Proceedings of the 2016 International Conference on Automatic Control and Dynamic Optimization Techniques (ICACDOT), Pune, India, 9–10 September 2016; pp. 645–648. [Google Scholar]
- Chaki, J.; Parekh, R.; Bhattacharya, S. Plant leaf recognition using a layered approach. In Proceedings of the 2016 International Conference on Microelectronics, Computing and Communications (MicroCom), Durgapur, India, 23–25 January 2016; pp. 6–11. [Google Scholar]
- Harish, B.S.; Hedge, A.; Venkatesh, O.; Spoorthy, D.G.; Sushma, D. Classification of plant leaves using Morphological features and Zernike moments. In Proceedings of the 2013 International Conference on Advances in Computing, Communications and Informatics (ICACCI), Mysore, India, 22–25 August 2013; pp. 1827–1831. [Google Scholar]
| Classifiers. | Advantages | Disadvantages |
|---|---|---|
| Artificial Neural Network (ANN) | 1. Capable in distinguishing complex nonlinear relationship between independent and dependent variables. 2. Simplistic statistical training. | 1. Great tendency of data overfitting. 2. Bigger computational load. |
| Convolutional Neural Network (CNN) | 1. Multiple features can be extracted simultaneously. 2. Robust to noise. | 1. High computation level. 2. No capable for generalization. |
| Probabilistic Neural Network (PNN) | 1. High distortion resistance. 2. Flexible to changing data. 3. Specimen can be classified into multiple output. | 1. Long training duration. 2. Complicated network layout. 3. Have tendency for overfitting with too many traits. |
| Support Vector Machine (SVM) | 1. Great generalization potential. 2. Exceptionally robust. | 1. Speed and size constraint for both training and testing. 2. Complex algorithm structure. 3. Slow training. |
| K-Nearest Neighbor (KNN) | 1. No training needed. 2. Robust in term of research space. 3. Simplest classifier. | 1. Susceptible to noise. 2. Lazy learning. 3. Costly testing for each instance. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Azlah, M.A.F.; Chua, L.S.; Rahmad, F.R.; Abdullah, F.I.; Wan Alwi, S.R. Review on Techniques for Plant Leaf Classification and Recognition. Computers 2019, 8, 77. https://doi.org/10.3390/computers8040077
Azlah MAF, Chua LS, Rahmad FR, Abdullah FI, Wan Alwi SR. Review on Techniques for Plant Leaf Classification and Recognition. Computers. 2019; 8(4):77. https://doi.org/10.3390/computers8040077
Chicago/Turabian StyleAzlah, Muhammad Azfar Firdaus, Lee Suan Chua, Fakhrul Razan Rahmad, Farah Izana Abdullah, and Sharifah Rafidah Wan Alwi. 2019. "Review on Techniques for Plant Leaf Classification and Recognition" Computers 8, no. 4: 77. https://doi.org/10.3390/computers8040077
APA StyleAzlah, M. A. F., Chua, L. S., Rahmad, F. R., Abdullah, F. I., & Wan Alwi, S. R. (2019). Review on Techniques for Plant Leaf Classification and Recognition. Computers, 8(4), 77. https://doi.org/10.3390/computers8040077






