The Prognostic Value of Haplotypes in the Vascular Endothelial Growth Factor A Gene in Colorectal Cancer
Abstract
:1. Introduction
2. Results and Discussion
2.1. Patient Characteristics
| Test cohort | Validation cohort | |
|---|---|---|
| Number (%) | Number (%) | |
| Sum of the percentages do not always equal 100 % due to rounding of data. Not all patients had complete patient characteristics available. | ||
| Sex | ||
| Male | 98 (51) | 159 (54) |
| Female | 93 (49) | 136 (46) |
| Age (years) | ||
| Mean (SD) | 70.1 (11.6) | 70.5 (11.2) |
| Range | 31–97 | 33–92 |
| pT category | ||
| 1–3 | 159 (83) | 248 (84) |
| 4 | 32 (17) | 47 (16) |
| pN category | ||
| 0 | 99 (52) | 163 (55) |
| 1–2 | 92 (48) | 132 (45) |
| Stage | ||
| II | 98 (51) | 163 (55) |
| III | 93 (49) | 132 (45) |
| Localization | ||
| Colon | 121 (63) | 197 (67) |
| Rectum | 70 (37) | 98 (33) |
| Vascular invasion | ||
| Yes | 11 (9) | 35 (12) |
| No | 111 (91) | 249 (88) |
| Neuronal invasion | ||
| Yes | 11 (9) | 31 (12) |
| No | 109 (91) | 236 (88) |
| Peritoneal perforation | ||
| Yes | 26 (14) | 38 (13) |
| No | 160 (86) | 253 (87) |
| Adjuvant chemotherapy | ||
| No | 157 (82) | 232 (81) |
| Yes | 34 (18) | 53 (19) |
| 5-FU/Leucovorin or capecitabine | 34 (100) | 28 (53) |
| XELOX | 0 (0) | 25 (47) |
| −2578 C/A SNP | ||
| CC | 47 (25) | 80 (27) |
| CA | 85 (45) | 147 (50) |
| AA | 59 (31) | 68 (23) |
| −460 C/T SNP | ||
| CC | 59 (31) | 69 (23) |
| CT | 85 (45) | 146 (49) |
| TT | 47 (25) | 80 (27) |
| 405 G/C SNP | ||
| GG | 95 (50) | 128 (43) |
| GC | 74 (39) | 129 (44) |
| CC | 22 (12) | 38 (13) |
2.2. The Prognostic Value of VEGF-A SNPs
| Disease free survival (DFS) | |||||||||
| Test cohort | Validation cohort | ||||||||
| VEGF-A SNP and genotype comparison* | Events | HR (95% CI) | p-value | VEGF-A SNP and genotype comparison* | Events | HR (95% CI) | p-value | ||
| −2578 C/A | −2578 C/A | ||||||||
| CC (NR) v CA (3.3) | 19 v 55 | 0.47 (0.30–0.75) | <0.01 | CC (5.6) v CA (5.8) | 17 v 49 | 0.62 (0.37–1.03) | 0.09 | ||
| CC (NR) v AA (8.5) | 19 v 29 | 0.83 (0.47–1.46) | 0.52 | CC (5.6) v AA (NR) | 17 v 15 | 1.13 (0.57–2.26) | 0.73 | ||
| CA (3.3) v AA (8.5) | 55 v 29 | 1.78 (1.16–2.74) | 0.01 | CA (5.8) v AA (NR) | 49 v 15 | 1.75 (1.05–2.93) | >0.05 | ||
| CA v CC + AA | 1.93 (1.29–2.89) | <0.01 | CA v CC + AA | 1.68 (1.09–2.60) | 0.02 | ||||
| −460 C/T | −460 C/T | ||||||||
| CC (8.5) v CT (3.3) | 29 v 55 | 0.56 (0.37–0.86) | 0.01 | CC (NR) v CT (5.8) | 15 v 50 | 0.55 (0.33–0.91) | 0.04 | ||
| CC (8.5) v TT (NR) | 29 v 19 | 1.21 (0.68–2.14) | 0.52 | CC (NR) v TT (5.6) | 15 v 16 | 0.91 (0.45–1.85) | 0.80 | ||
| CT (3.3) v TT (NR) | 55 v 19 | 2.11 (1.33–3.35) | <0.01 | CT (5.8) v TT (5.6) | 50 v 16 | 1.73 (1.05–2.87) | >0.05 | ||
| CT v CC + TT | 1.93 (1.29–2.89) | <0.01 | CT v CC + TT | 1.78 (1.15–2.75) | 0.01 | ||||
| 405 G/C | 405 G/C | ||||||||
| GG (7.5) v GC (4.3) | 50 v 43 | 0.73 (0.48–1.11) | 0.13 | GG (NR) v GC (5.0) | 24 v 48 | 0.41 (0.26–0.65) | <0.01 | ||
| GG (7.5) v CC (NR) | 50 v 10 | 1.16 (0.61–2.22) | 0.67 | GG (NR) v CC (NR) | 24 v 9 | 0.69 (0.30–1.60) | 0.34 | ||
| GC (4.3) v CC (NR) | 43 v 10 | 1.55 (0.84–2.85) | 0.21 | GC (5.0) v CC (NR) | 48 v 9 | 1.66 (0.90–3.06) | 0.16 | ||
| GC v GG + CC | 1.40 (0.93–2.12) | 0.09 | GC v GG + CC | 2.22 (1.42–3.46) | <0.01 | ||||
| Overall survival (OS) | |||||||||
| Test cohort | Validation cohort | ||||||||
| VEGF-A SNP and genotype comparison* | Events | HR (95% CI) | p-value | VEGF-A SNP and genotype comparison* | Events | HR (95% CI) | p-value | ||
| −2578 C/A | −2578 C/A | ||||||||
| CC (NR) v CA (4.7) | 18 v 55 | 0.46 (0.29–0.74) | <0.01 | CC (6.1) v CA (NR) | 12 v 40 | 0.55 (0.31–0.97) | 0.06 | ||
| CC (NR) v AA (8.8) | 18 v 29 | 0.79 (0.44–1.40) | 0.42 | CC (6.1) v AA (7.0) | 12 v 15 | 0.84 (0.39–1.79) | 0.65 | ||
| CA (4.7) v AA (8.8) | 55 v 29 | 1.71 (1.12–2.63) | 0.02 | CA (NR) v AA (7.0) | 40 v 15 | 1.38 (0.79–2.41) | 0.28 | ||
| CA v CC + AA | 1.89 (1.27–2.82) | <0.01 | CA v CC v AA | 1.57 (0.97–2.54) | 0.07 | ||||
| −460 C/T | −460 C/T | ||||||||
| CC (8.8) v CT (4.7) | 29 v 55 | 0.58 (0.38–0.90) | 0.02 | CC (7.0) v CT (NR) | 15 v 41 | 0.69 (0.40–1.20) | 0.22 | ||
| CC (8.8) v TT (NR) | 29 v 18 | 1.27 (0.72–2.26) | 0.42 | CC (7.0) v TT (NR) | 15 v 11 | 1.26 (0.58–2.72) | 0.56 | ||
| CT (4.7) v TT (NR) | 55 v 18 | 2.15 (1.35–3.43) | <0.01 | CT (NR) v TT (NR) | 41 v 11 | 1.99 (1.13–3.53) | 0.04 | ||
| CT v CC + TT | 1.89 (1.27–2.82) | <0.01 | CT v CC + TT | 1.66 (1.03–2.68) | 0.04 | ||||
| 405 G/C | 405 G/C | ||||||||
| GG (8.3) v GC (4.8) | 50 v 43 | 0.72 (0.47–1.10) | 0.11 | GG (NR) v GC (5.8) | 23 v 39 | 0.51 (0.31–0.84) | 0.01 | ||
| GG (8.3) v CC (NR) | 50 v 9 | 1.33 (0.70–2.54) | 0.43 | GG (NR) v CC (NR) | 23 v 5 | 1.21 (0.49–3.02) | 0.70 | ||
| GC (4.8) v CC (NR) | 43 v 9 | 1.82 (0.99–3.35) | 0.10 | GC (5.8) v CC (NR) | 39 v 5 | 2.25 (1.11–4.58) | 0.08 | ||
| GC v GG + CC | 1.46 (0.97–2.20) | 0.06 | GC v GG + CC | 2.00 (1.23–3.25) | <0.01 | ||||
| Genotypes | Disease free survival (DFS) | Overall survival (OS) | ||
|---|---|---|---|---|
| HR (95% CI) | p-value | HR (95% CI) | p-value | |
| −2578 C/A | ||||
| CC | 1 | 1 | ||
| CA | 1.98 (1.10–3.56) | 0.02 | 1.92 (0.98–3.76) | 0.06 |
| AA | 1.10 (0.53–2.28) | 0.80 | 1.32 (0.60–2.92) | 0.49 |
| −460 C/T | ||||
| CC | 1 | 1 | ||
| CT | 1.81 (0.99–3.31) | 0.06 | 1.46 (0.79–2.72) | 0.23 |
| TT | 0.89 (0.42–1.87) | 0.76 | 0.73 (0.32–1.64) | 0.44 |
| 405 G/C | ||||
| GG | 1 | 1 | ||
| GC | 1.79 (1.06–3.01) | 0.03 | 1.52 (0.88–2.63) | 0.13 |
| CC | 0.98 (0.43–2.26) | 0.97 | 0.73 (0.27–1.98) | 0.53 |
2.3. Haplotype Analysis
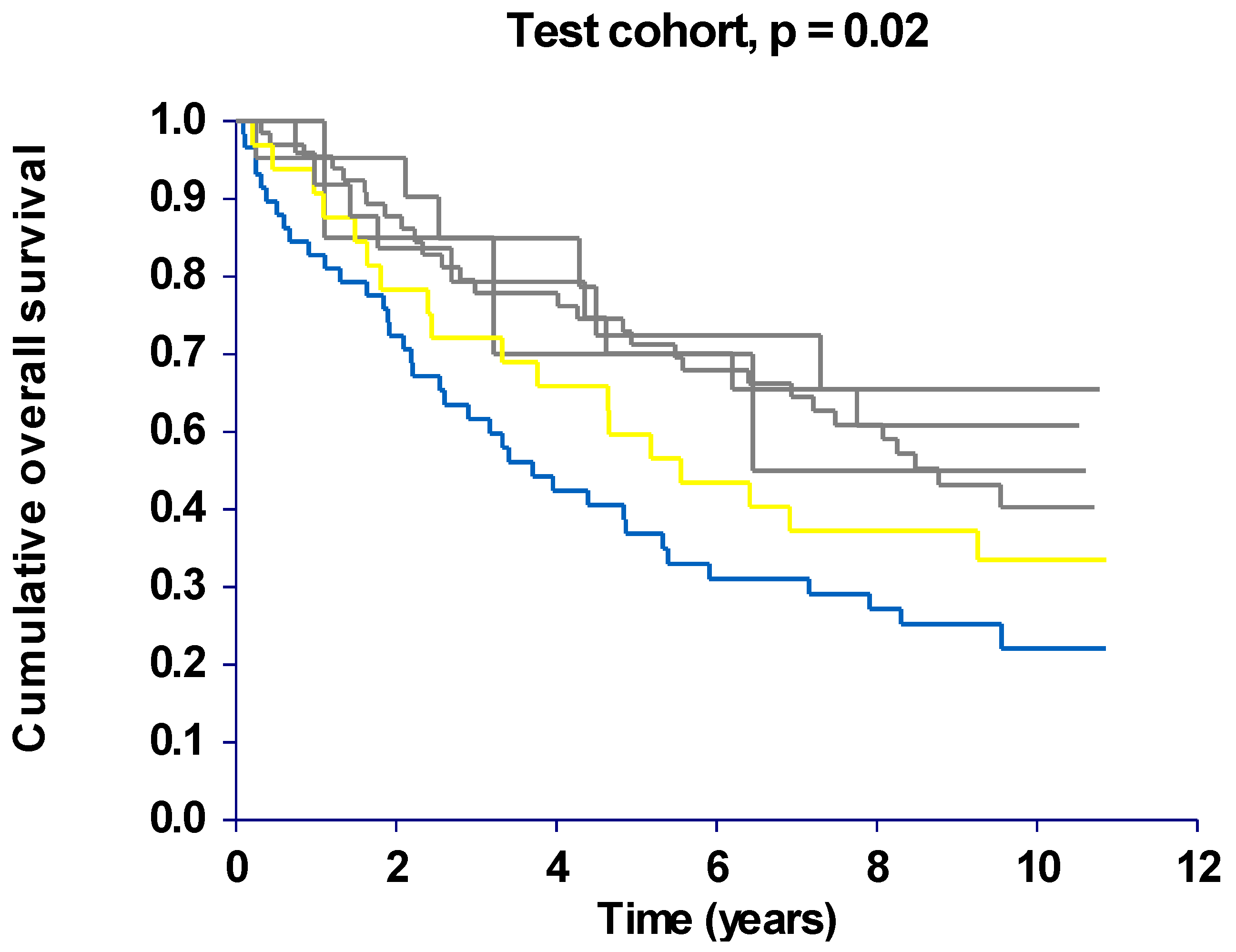
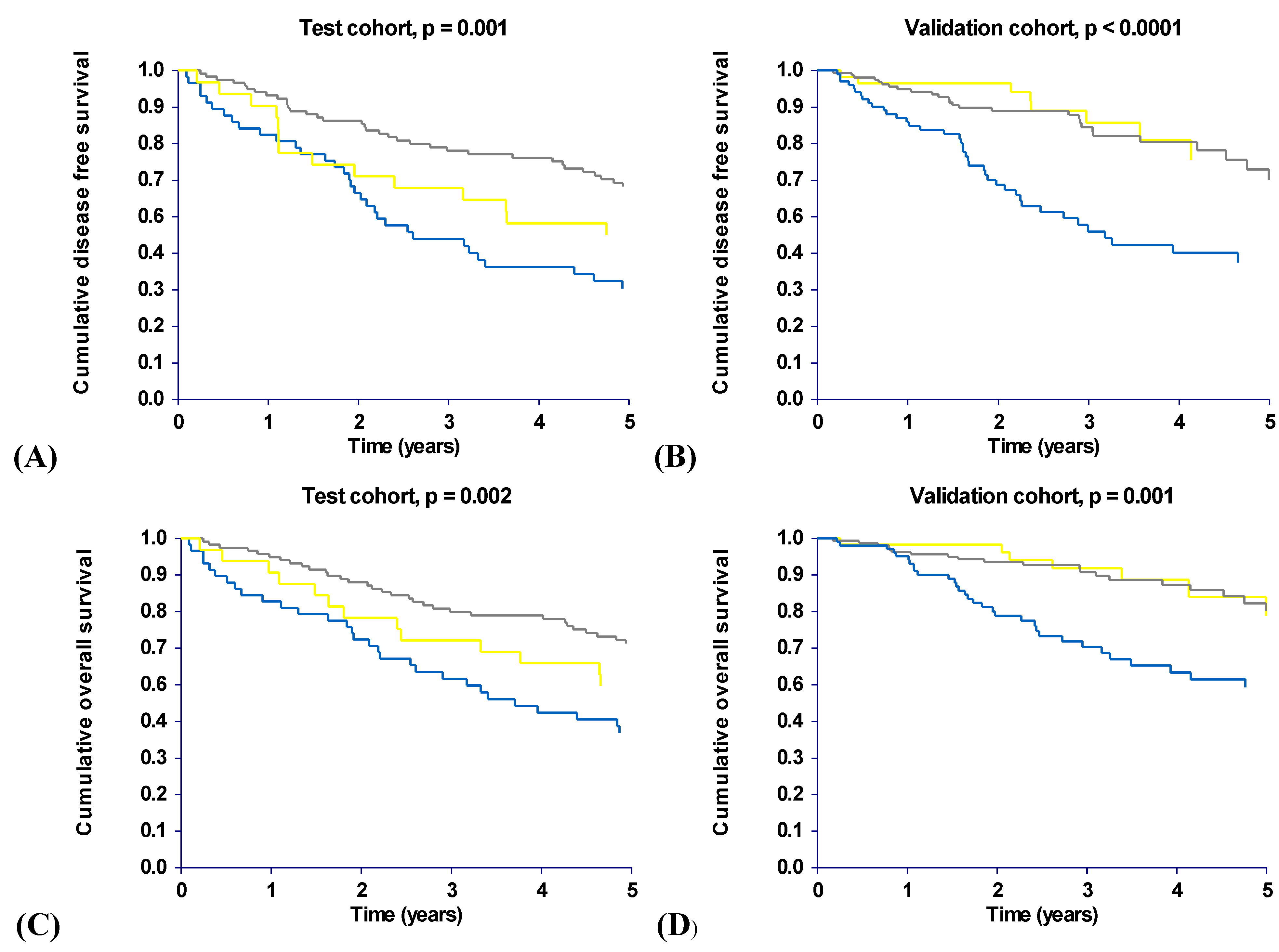
| Disease free survival (DFS) | ||||||
| Test cohort | Validation cohort | |||||
| HR (95% CI) | p-value | HR (95% CI) | p-value | |||
| Remaining combinations | 1 | Remaining combinations | 1 | |||
| (CTG, ACG) | 1.85 (0.95–3.62) | 0.07 | (CTG, ACG) | 0.98 (0.44–2.18) | 0.97 | |
| (CTC, ACG) | 2.04 (1.14–3.67) | 0.02 | (CTC, ACG) | 2.46 (1.49–4.06) | <0.01 | |
| Overall survival (OS) | ||||||
| Test cohort | Validation cohort | |||||
| HR (95% CI) | p-value | HR (95% CI) | p-value | |||
| Remaining combinations | 1 | Remaining combinations | 1 | |||
| (CTG, ACG) | 1.72 (0.89–3.36) | 0.11 | (CTG, ACG) | 0.95 (0.40–2.21) | 0.90 | |
| (CTC, ACG) | 2.15 (1.21–3.82) | 0.01 | (CTC, ACG) | 2.07 (1.20–3.57) | 0.01 | |
2.4. Discussion
3. Experimental Section
3.1. Study Population
3.2. Analysis of Single Nucleotide Polymorphism
3.3. Statistical Analysis
4. Conclusions
Acknowledgements
References
- Andre, T.; Boni, C.; Navarro, M.; Tabernero, J.; Hickish, T.; Topham, C.; Bonetti, A.; Clingan, P.; Bridgewater, J.; Rivera, F.De.G.A. Improved overall survival with oxaliplatin, fluorouracil, and leucovorin as adjuvant treatment in stage II or III colon cancer in the MOSAIC trial. J. Clin. Oncol. 2009, 27, 3109–3116. [Google Scholar] [CrossRef]
- Benson, A.B.I.; Schrag, D.; Somerfield, M.R.; Cohen, A.M.; Figueredo, A.T.; Flynn, P.J.; Krzyzanowska, M.K.; Maroun, J.; McAllister, P.; van Cutsem, E.; Brouwers, M.; Charette, M.; Haller, D.G. American Society of Clinical Oncology recommendations on adjuvant chemotherapy for stage II colon cancer. J. Clin. Oncol. 2004, 22, 3408–3419. [Google Scholar]
- Moertel, C.G.; Fleming, T.R.; Macdonald, J.S.; Haller, D.G.; Laurie, J.A.; Tangen, C.M.; Ungerleider, J.S.; Emerson, W.A.; Tormey, D.C.; Glick, J.H.; Veeder, M.H.; Mailliard, J.A. Fluorouracil plus levamisole as effective adjuvant therapy after resection of stage III colon carcinoma: a final report. Ann. Intern. Med. 1995, 122, 321–326. [Google Scholar] [CrossRef]
- Van Cutsem, E. Colon cancer: ESMO clinical recommendations for diagnosis, adjuvant treatment and follow-up. Ann. Oncol. 2007, 18 (Suppl. 2), 21–22. [Google Scholar]
- Glimelius, B. Rectal cancer: ESMO clinical recommendations for diagnosis, treatment and follow-up. Ann. Oncol. 2007, 18 (Suppl. 2), 23–24. [Google Scholar]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Ferrara, N.; Gerber, H.P.; LeCouter, J. The biology of VEGF and its receptors. Nat. Med. 2003, 9, 669–676. [Google Scholar] [CrossRef]
- Kim, K.J.; Li, B.; Winer, J.; Armanini, M.; Gillett, N.; Phillips, H.S.; Ferrara, N. Inhibition of vascular endothelial growth factor-induced angiogenesis suppresses tumor growth in vivo. Nature 1993, 362, 841–844. [Google Scholar] [CrossRef]
- Takahashi, H.; Shibuya, M. The vascular endothelial growth factor (VEGF)/VEGF receptor system and its role under physiological and pathological conditions. Clin. Sci. (Lond.) 2005, 109, 227–241. [Google Scholar] [CrossRef]
- Stevens, A.; Soden, J.; Brenchley, P.E.; Ralph, S.; Ray, D.W. Haplotype analysis of the polymorphic human vascular endothelial growth factor gene promoter. Cancer Res. 2003, 63, 812–816. [Google Scholar]
- Watson, C.J.; Webb, N.J.; Bottomley, M.J.; Brenchley, P.E. Identification of polymorphisms within the vascular endothelial growth factor (VEGF) gene: correlation with variation in VEGF protein production. Cytokine 2000, 12, 1232–1235. [Google Scholar] [CrossRef]
- Lu, H.; Shu, X.O.; Cui, Y.; Kataoka, N.; Wen, W.; Cai, Q.; Ruan, Z.X.; Gao, Y.T.; Zheng, W. Association of genetic polymorphisms in the VEGF gene with breast cancer survival. Cancer Res. 2005, 65, 5015–5019. [Google Scholar] [CrossRef]
- Kawai, Y.; Sakano, S.; Korenaga, Y.; Eguchi, S.; Naito, K. Associations of single nucleotide polymorphisms in the vascular endothelial growth factor gene with the characteristics and prognosis of renal cell carcinomas. Eur. Urol. 2007, 52, 1147–1155. [Google Scholar] [CrossRef]
- Kim, J.G.; Sohn, S.K.; Chae, Y.S.; Cho, Y.Y.; Bae, H.I.; Yan, G.; Park, J.Y.; Lee, M.H.; Chung, H.Y.; Yu, W. Vascular endothelial growth factor gene polymorphisms associated with prognosis for patients with gastric cancer. Ann. Oncol. 2007, 18, 1030–1036. [Google Scholar] [CrossRef]
- Kim, J.G.; Chae, Y.S.; Sohn, S.K.; Cho, Y.Y.; Moon, J.H.; Park, J.Y.; Jeon, S.W.; Lee, I.T.; Choi, G.S.; Jun, S.H. Vascular endothelial growth factor gene polymorphisms associated with prognosis for patients with colorectal cancer. Clin. Cancer Res. 2008, 14, 62–66. [Google Scholar] [CrossRef]
- Lurje, G.; Zhang, W.; Schultheis, A.M.; Yang, D.; Groshen, S.; Hendifar, A.E.; Husain, H.; Gordon, M.A.; Nagashima, F.; Chang, H.M.; Lenz, H.J. Polymorphisms in VEGF and IL-8 predict tumor recurrence in stage III colon cancer. Ann. Oncol. 2008, 19, 1734–1741. [Google Scholar] [CrossRef]
- Dassoulas, K.; Gazouli, M.; Rizos, S.; Theodoropoulos, G.; Christoni, Z.; Nikiteas, N.; Karakitsos, P. Common polymorphisms in the vascular endothelial growth factor gene and colorectal cancer development, prognosis, and survival. Mol. Carcinog. 2009, 48, 563–569. [Google Scholar] [CrossRef]
- Smerdel, M.P.; Waldstrom, M.; Brandslund, I.; Steffensen, K.D.; Andersen, R.F.; Jakobsen, A. Prognostic importance of vascular endothelial growth factor-A expression and vascular endothelial growth factor polymorphisms in epithelial ovarian cancer. Int. J. Gynecol. Cancer 2009, 19, 578–584. [Google Scholar] [CrossRef]
- Schneider, B.P.; Radovich, M.; Miller, K.D. The role of vascular endothelial growth factor genetic variability in cancer. Clin. Cancer Res. 2009, 15, 5297–5302. [Google Scholar] [CrossRef]
- Cacev, T.; Loncar, B.; Seiwerth, S.; Spaventi, S.; Kapitanovic, S. Vascular endothelial growth factor polymorphisms −1154 G/A and −460 C/T are not associated with VEGF mRNA expression and susceptibility to sporadic colon cancer. DNA Cell Biol. 2008, 27, 569–574. [Google Scholar] [CrossRef]
- Hofmann, G.; Langsenlehner, U.; Renner, W.; Langsenlehner, T.; Yazdani-Biuki, B.; Clar, H.; Gerger, A.; Wehrschuetz, M.; Samonigg, H.; Krippl, P. Common single nucleotide polymorphisms in the vascular endothelial growth factor gene and colorectal cancer risk. J. Cancer Res. Clin. Oncol. 2008, 134, 591–595. [Google Scholar] [CrossRef]
- Maltese, P.; Canestrari, E.; Ruzzo, A.; Graziano, F.; Falcone, A.; Loupakis, F.; Tonini, G.; Santini, D.; Magnani, M. VEGF gene polymorphisms and susceptibility to colorectal cancer disease in Italian population. Int. J. Colorectal Dis. 2008, 24, 165–170. [Google Scholar]
- Chae, Y.S.; Kim, J.G.; Sohn, S.K.; Cho, Y.Y.; Ahn, B.M.; Moon, J.H.; Jeon, S.W.; Park, J.Y.; Lee, I.T.; Choi, G.S.; Jun, S.H. Association of vascular endothelial growth factor gene polymorphisms with susceptibility and clinicopathologic characteristics of colorectal cancer. J. Korean Med. Sci. 2008, 23, 421–427. [Google Scholar] [CrossRef]
- Stephens, M.; Smith, N.J.; Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 2001, 68, 978–989. [Google Scholar] [CrossRef]
- Stephens, M.; Scheet, P. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am. J. Hum. Genet. 2005, 76, 449–462. [Google Scholar] [CrossRef]
© 2010 by the authors; licensee MDPI, Basel, Switzerland. This article is an Open Access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hansen, T.F.; Spindler, K.-L.G.; Andersen, R.F.; Lindebjerg, J.; Kølvraa, S.; Brandslund, I.; Jakobsen, A. The Prognostic Value of Haplotypes in the Vascular Endothelial Growth Factor A Gene in Colorectal Cancer. Cancers 2010, 2, 1405-1418. https://doi.org/10.3390/cancers2031405
Hansen TF, Spindler K-LG, Andersen RF, Lindebjerg J, Kølvraa S, Brandslund I, Jakobsen A. The Prognostic Value of Haplotypes in the Vascular Endothelial Growth Factor A Gene in Colorectal Cancer. Cancers. 2010; 2(3):1405-1418. https://doi.org/10.3390/cancers2031405
Chicago/Turabian StyleHansen, Torben F., Karen-Lise G. Spindler, Rikke F. Andersen, Jan Lindebjerg, Steen Kølvraa, Ivan Brandslund, and Anders Jakobsen. 2010. "The Prognostic Value of Haplotypes in the Vascular Endothelial Growth Factor A Gene in Colorectal Cancer" Cancers 2, no. 3: 1405-1418. https://doi.org/10.3390/cancers2031405
APA StyleHansen, T. F., Spindler, K.-L. G., Andersen, R. F., Lindebjerg, J., Kølvraa, S., Brandslund, I., & Jakobsen, A. (2010). The Prognostic Value of Haplotypes in the Vascular Endothelial Growth Factor A Gene in Colorectal Cancer. Cancers, 2(3), 1405-1418. https://doi.org/10.3390/cancers2031405




