Comprehensive Detection of Chromosomal and Genomic Abnormalities via Next-Generation Sequencing-Based Genomic Proximity Mapping Improves Diagnostic Classification of Hematologic Neoplasms
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Case Selection
2.2. Genomic Proximity Mapping
3. Results
3.1. Patient Characteristics (Table 1)
| Number (%) | |
|---|---|
| Patients No. | 18 |
| Age, median (range) | 57 (16–82) |
| Gender | |
| Male | 9 (50%) |
| Female | 9 (50%) |
| Tissue type | |
| Peripheral blood | 6 (33.3%) |
| Bone marrow | 7 (38.9%) |
| Lymph node | 5 (27.8%) |
| Pathologic diagnosis | |
| Acute lymphoblastic leukemia | 4 (22.2%) |
| Acute myeloid leukemia | 3 (16.7%) |
| Diffuse large B-cell lymphoma | 3 (16.7%) |
| High-grade B-cell lymphoma | 2 (11.1%) |
| Mantle cell lymphoma | 1 (5.6%) |
| T-prolymphocytic leukemia | 1 (5.6%) |
| Primary myelofibrosis | 1 (5.6%) |
| Myelodysplastic neoplasm | 1 (5.6%) |
| Chronic myeloid leukemia in B-lymphoid blast crisis | 1 (5.6%) |
| Rosai–Dorfman disease | 1 (5.6%) |
| Abnormal cells by flow cytometry, median (range) | 0.75–86% |
| <20% | 7.9% (0.75–15.8%) |
| 20–50% | 38.0% (29.0–49.3%) |
| >50% | 75.0% (60.2–86.0%) |
3.2. Comparison of Genomic Aberrations Detected by GPM and Other Cytogenomic Methods, Including Novel Findings by GPM (Table 2)
| Case | Original Pathologic Diagnosis | Diagnostic Impact of GPM | GPM (Relevant Aberrations) | Karyotype/FISH |
|---|---|---|---|---|
| 1 | Mantle cell lymphoma | Comprehensive genomic analysis | t(11;14) IGH::CCND1, TP53 deletion, other CNAs | nuc ish(CCND1,IGH)x3(IGH con CCND1)x2[188/200] |
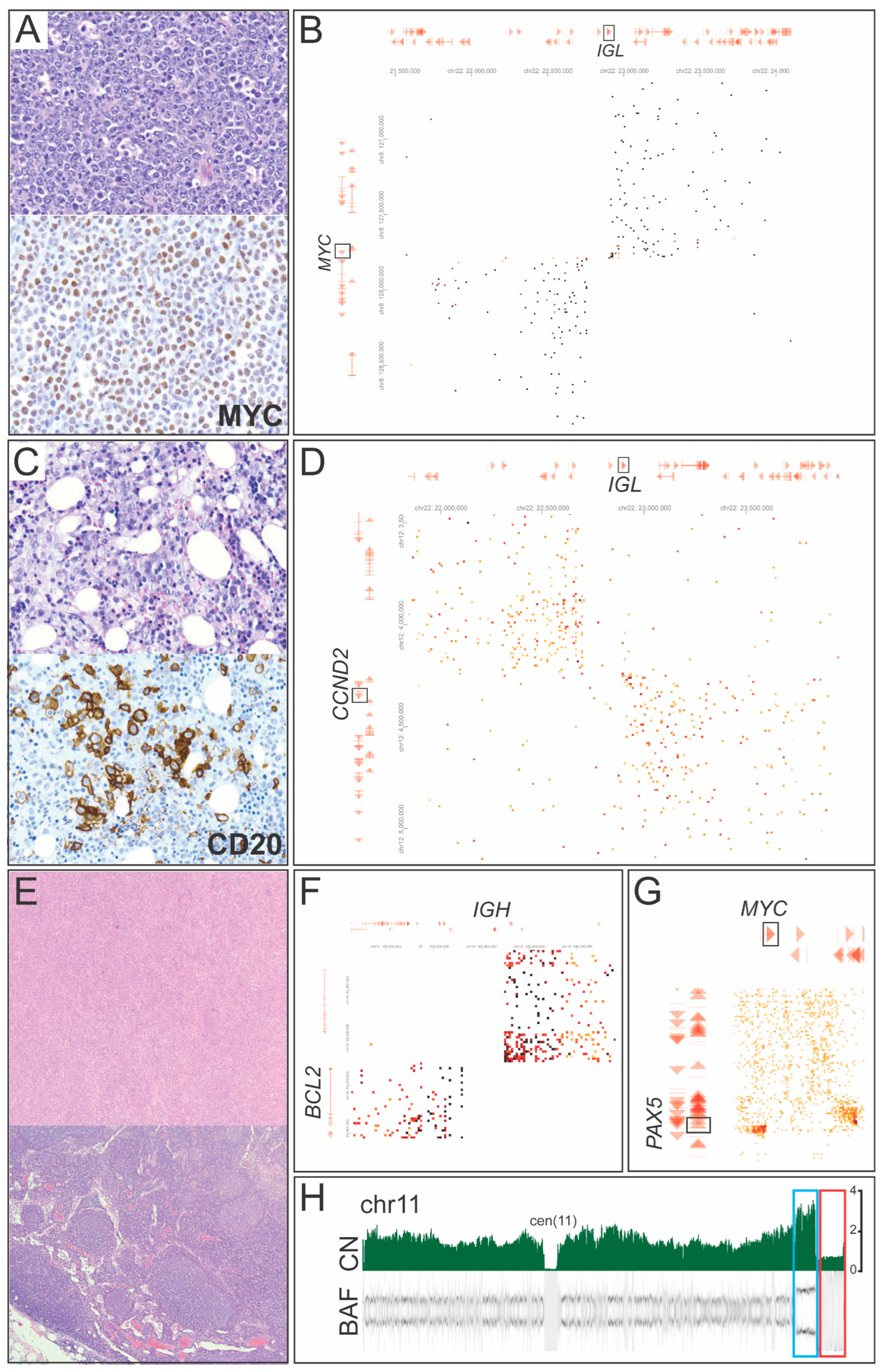
| 2 | High-grade B-cell lymphoma | Diagnostic support for Burkitt lymphoma | t(8;22) IGL::MYC | nuc ish(BCL6)x2[200],(MYC)x2(5′MYC sep 3′MYC)x1[200],(MYCx3,IGHx2)[200],(IGH,BCL2)x2[200] |
| 3 | High-grade B-cell lymphoma | Comprehensive genomic analysis | 6q deletion, 9p cnLOH, copy gains of segments of 2p, 11p, and 12q, +13, +17, +22 | nuc ish(MYC)x2[200] |
| 4 | Diffuse large B-cell lymphoma | Accurate genetic characterization for diagnostic clarity | t(12;22) IGL::CCND2, inv(3) RPN1::EVI1, t(5;15), t(10;18) | 46,XY,add(3)(q27)[7]/46,XY,t(5;15)(q33;q15),t(10;18)(p10;p10),t(12;22)(p13;q11.2)[13]. nuc ish(CCND1,IGH)x2[200],(TP53,CEP17)x2[200] |
| 5 | Diffuse large B-cell lymphoma transformed from follicular lymphoma | Diagnostic support for DLBCL/high-grade B-cell lymphoma with MYC and BCL2 rearrangements | t(8;9) PAX5::MYC, t(14;18) IGH::BCL2, 11q aberration, TP53 deletion, other CNAs | nuc ish(5′MYCx2~3,3′MYCx1~2)(5′MYC con 3′MYC)x1~2[152/300],(IGH,BCL2)x2~3(IGH con BCL2)x1~2[171/200] PAX5::MYC (Exon2::Exon2) by RNA sequencing |
| 6 | Diffuse large B-cell lymphoma | Comprehensive genomic analysis | Multiple CNAs | nuc ish(MYC)x2[200] |
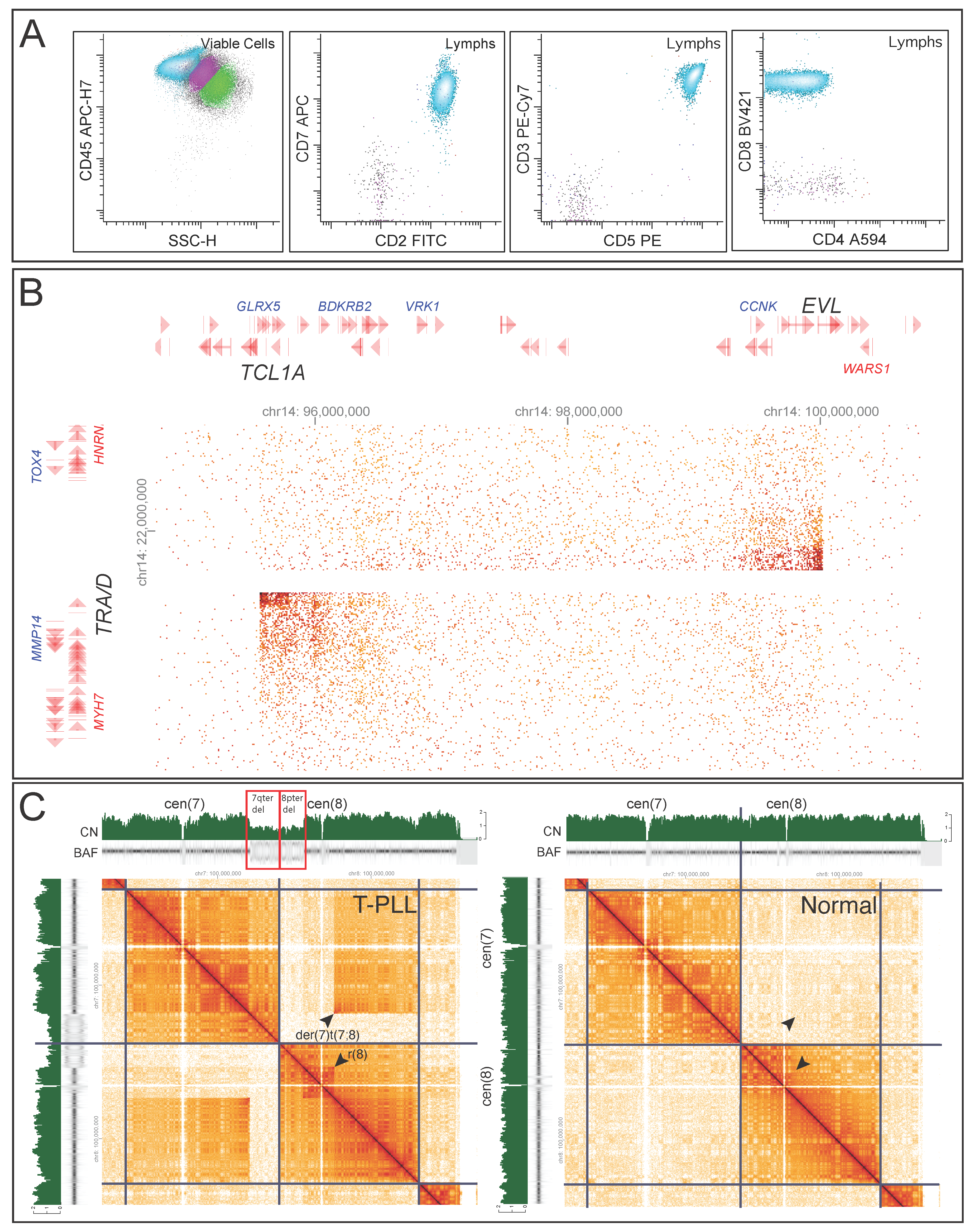
| 7 | Peripheral T-cell lymphoma | Diagnostic support for T-prolymphocytic leukemia | ins(14;14)(q11.2;q32.1q32.2) TRA/D::TCL1A | nuc ish(TCL1Ax3,TCL1Bx2)(TCL1A con TCL1B)x2[138/200] |
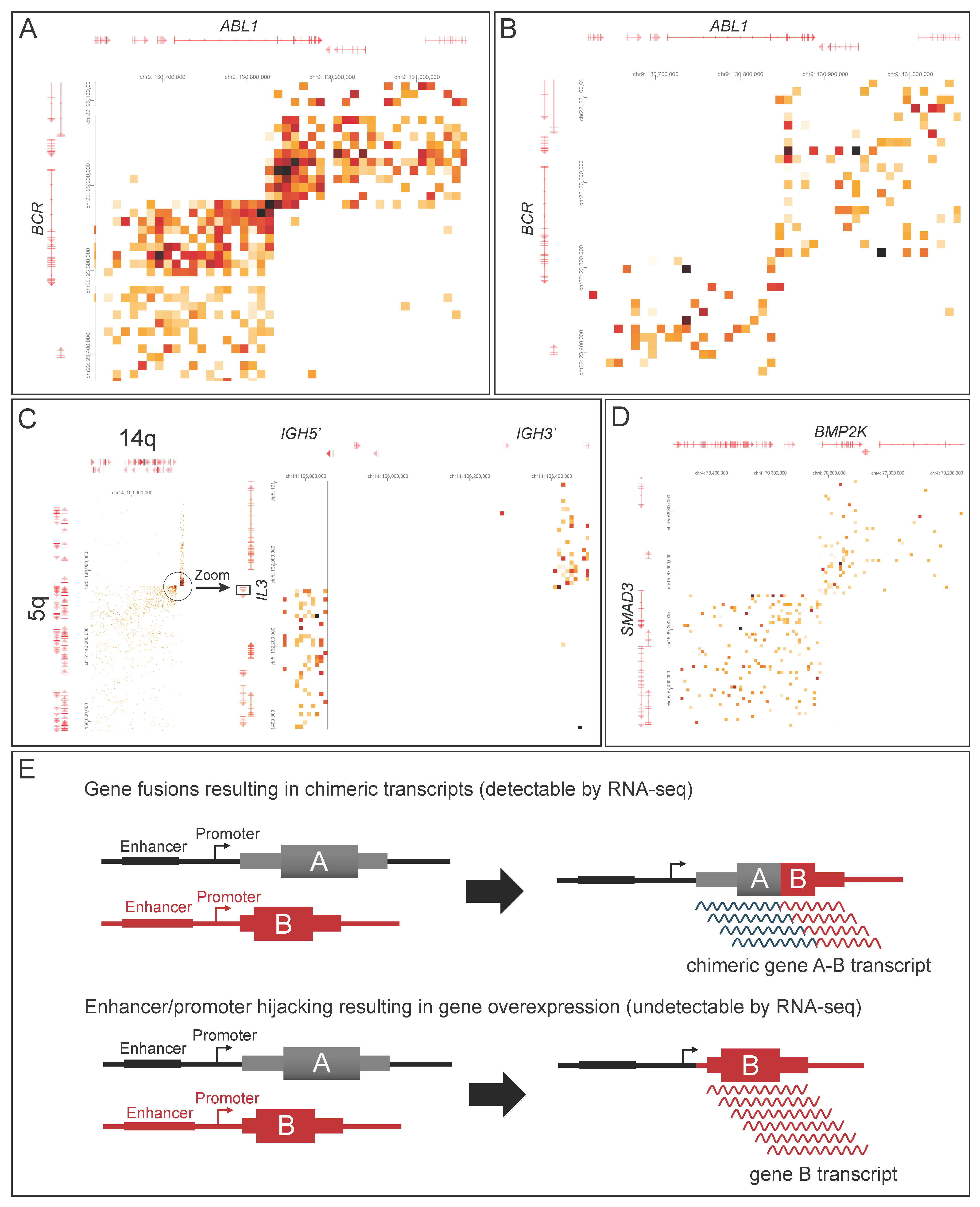
| 8 | B-lymphoblastic leukemia/lymphoma | Comprehensive genomic analysis | t(9;22) BCR::ABL1, 7p deletion (IKZF1), 17q deletion | 46,XX,t(9;22)(q34;q11.2)[13]/46,sl,der(16)t(1;16)(q12;q11.2)[2]/47,sl,-16,+2r [5]. nuc ish(ABL1x3,ASS1x2,BCRx3)(ABL1 con BCR)x2[177/200] BCR::ABL1 p190 by RT-PCR |
| 9 | B-lymphoblastic leukemia/lymphoma | Comprehensive genomic analysis | t(4;11) AFF1::KMT2A, trisomy 6, 7q gain | 46,XY[20]. nuc ish(KMT2A)x2(5′KMT2A sep 3′KMT2A)x1[181/200] |
| 10 | B-lymphoblastic leukemia/lymphoma | Accurate genetic characterization for diagnostic clarity | t(5;14) IGH::IL3, t(4;15) BMP2K::SMAD3 | t(5;14) (per outside report) |
| 11 | T-lymphoblastic leukemia/lymphoma | Accurate genetic characterization for diagnostic clarity | t(1;17), der(12)t(12;17), t(2;9;4), t(14;16) with BCL11B rearrangement, biallelic CDKN2A deletion, +18 | 47,XY,t(1;17)(q21;p13),t(2;9;4)(p23;p23;q25),del(4)(q27q31),der(12)t(12;17)(p13;q11.2),+18[33]/47,XY,idem,add(4)(q35)[5]/47,XY,idem,del(4)(q11.2q25)[2]. nuc ish (CDKN2Ax0,CEP9x2)[194/200] |
| 12 | Acute myeloid leukemia | Comprehensive genomic analysis | inv(16) MYH11::CBFB, t(1;12), t(6;12) | 46,XY,inv(16)(p13.1q22)[13]/46,idem,inv(7)(p13p22)[5]/46,idem,t(1;12)(q32;p13)[2] |
| 13 | Acute myeloid leukemia | Comprehensive genomic analysis | Deletions of 3p and 5q, and other CNAs | 52,XX,del(3)(p13),del(5)(q13q33),+del(5)(q13q33),add(9)(q34),+11,+13,+14,+21,+22[8]/52,sl,add(14)(q32)[cp6]/45~48,del(3)(p13),t(4;11)(q25;q25),del(5) (q13q33),add(9)(q34),add(12)(p13)[cp3]/46,XX,-3,del(5)(q13q33),der(12)t(3;12)(q21;p13),-18,add(21)(q22),+2mar [cp3]. nuc ish(D5S23x2,EGR1x1)[50/200]/(D5S23x3,EGR1x1)[68/200] |
| 14 | Acute myeloid leukemia | Concordant | Normal | 46,XX[20] |
| 15 | Chronic myeloid leukemia in B-lymphoid blast crisis | Concordant | t(9;22) BCR::ABL1 | t(9;22) by FISH (per outside report) Negative for BCR::ABL1 by RT-PCR BCR::ABL1 (Exon 13::Exon 3) by RNA sequencing |
| 16 | Primary myelofibrosis | Concordant | 9p cnLOH | 46,XX[20] |
| 17 | Myelodysplastic neoplasm | Comprehensive genomic analysis | Deletions of 5q, 16q, 17p, and 17q, -18, der(20)t(5;20), and other CNAs | 43,X,-Y,-5,-17,-18,add(20)(p13),+r[6]/44,idem,+r[10]/43,idem,-r,+mar[2]/46,XY[2]. nuc ish(EGR1x1,D5S23x2)[184/200] |
| 18 | Rosai–Dorfman disease | Concordant | Normal | Failed |
3.2.1. Mature Lymphomas
3.2.2. Acute Lymphoblastic Leukemia/Lymphoma
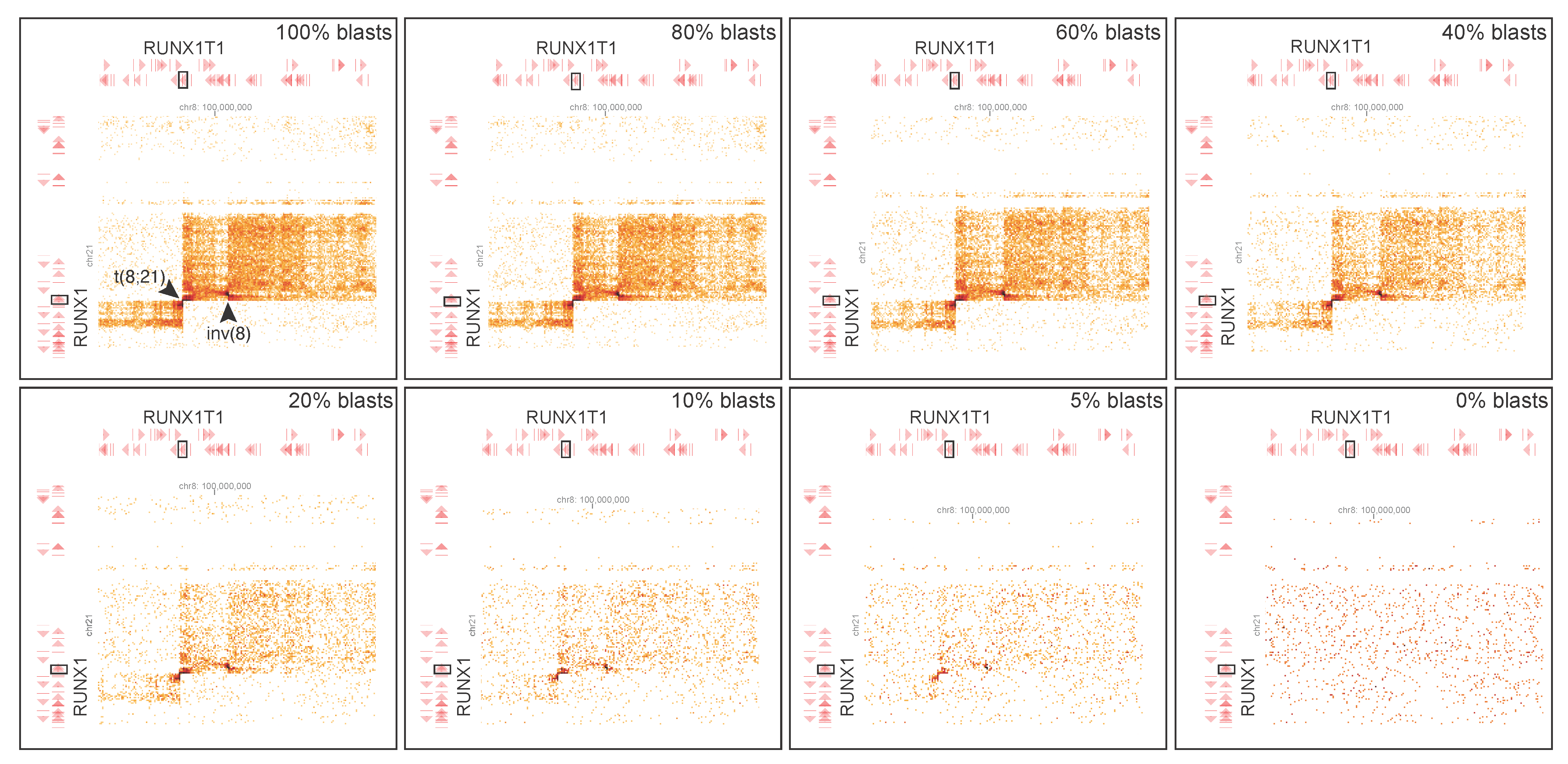
3.2.3. Acute Myeloid Leukemia
3.2.4. Other Myeloid Neoplasms
3.3. GPM Performance and Concordance with Other Methods (Table 3)
| (A) | |||||
| Total | Translocation | Inversion | CBD | ||
| Total variants by GPM | 91 | 60 | 14 | 17 | |
| Variants by other tests * | 31 | 21 | 3 | 7 | |
| Additional variants by GPM | 62 | 40 | 12 | 10 | |
| Concordant variants by GPM | 29 | 20 | 2 | 7 | |
| Concordance rate (%) | 93.5 | 95.2 | |||
| (B) | |||||
| Total | Deletion | Duplication | Aneuploidy | cnLOH | |
| Variants by other tests * | 83 | 32 | 18 | 11 | 22 |
| Concordant variants by GPM | 74 | 31 | 18 | 10 | 15 |
| Concordance rate (%) | 89.2 | 96.9 | 100 | 90.9 | 68.2 |
4. Discussion
4.1. Performance of GPM
4.2. GPM Improved Diagnostic Accuracy in Mature B-Cell and T-Cell Lymphomas
4.3. Diagnostic Relevance of GPM in B- and T-Lymphoblastic Leukemia/Lymphoma
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AML | Acute myeloid leukemia |
| B-LL | B-lymphoblastic leukemia/lymphoma |
| CBD | Contact by discontinuity |
| CLL/SLL | Chronic lymphocytic leukemia/small lymphocytic lymphoma |
| CMA | Chromosomal microarray analysis |
| CML | Chronic myeloid leukemia |
| CNA | Copy number alteration |
| cnLOH | Copy-neutral loss of heterozygosity |
| DLBCL | Diffuse large B-cell lymphoma |
| FFPE | Formalin-fixed, paraffin-embedded |
| FISH | Fluorescence in situ hybridization |
| FL | Follicular lymphoma |
| GPM | Genomic proximity mapping |
| Hi-C | High-throughput chromosome conformation capture |
| ICC | International Consensus Classification |
| IG | Immunoglobulin |
| LRS | Long-red sequencing |
| MDS | Myelodysplastic neoplasm |
| MPAL | Mixed phenotype acute leukemia |
| NGS | Next-generation sequencing |
| OGM | Optical genome mapping |
| PMF | Primary myelofibrosis |
| SV | Structural variant |
| TCR | T-cell receptor |
| T-LL | T-lymphoblastic leukemia/lymphoma |
| T-PLL | T-prolymphocytic leukemia |
| CNN | Convolutional neural network |
| MAF | Minor allele frequency |
References
- Arber, D.A.; Orazi, A.; Hasserjian, R.P.; Borowitz, M.J.; Calvo, K.R.; Kvasnicka, H.M.; Wang, S.A.; Bagg, A.; Barbui, T.; Branford, S.; et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: Integrating morphologic, clinical, and genomic data. Blood 2022, 140, 1200–1228. [Google Scholar] [CrossRef] [PubMed]
- Khoury, J.D.; Solary, E.; Abla, O.; Akkari, Y.; Alaggio, R.; Apperley, J.F.; Bejar, R.; Berti, E.; Busque, L.; Chan, J.K.C.; et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia 2022, 36, 1703–1719. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Fei, Z.; Xiao, P. Methods to improve the accuracy of next-generation sequencing. Front. Bioeng. Biotechnol. 2023, 11, 982111. [Google Scholar] [CrossRef]
- Satam, H.; Joshi, K.; Mangrolia, U.; Waghoo, S.; Zaidi, G.; Rawool, S.; Thakare, R.P.; Banday, S.; Mishra, A.K.; Das, G.; et al. Next-Generation Sequencing Technology: Current Trends and Advancements. Biology 2023, 12, 997. [Google Scholar] [CrossRef]
- Amarasinghe, S.L.; Su, S.; Dong, X.; Zappia, L.; Ritchie, M.E.; Gouil, Q. Opportunities and challenges in long-read sequencing data analysis. Genome Biol. 2020, 21, 30. [Google Scholar] [CrossRef]
- De Coster, W.; Weissensteiner, M.H.; Sedlazeck, F.J. Towards population-scale long-read sequencing. Nat. Rev. Genet. 2021, 22, 572–587. [Google Scholar] [CrossRef]
- Jiang, T.; Liu, S.; Cao, S.; Liu, Y.; Cui, Z.; Wang, Y.; Guo, H. Long-read sequencing settings for efficient structural variation detection based on comprehensive evaluation. BMC Bioinform. 2021, 22, 552. [Google Scholar] [CrossRef]
- Schmitt, A.D.; Sikkink, K.; Ahmed, A.A.; Melnyk, S.; Reid, D.; Van Meter, L.; Guest, E.M.; Lansdon, L.A.; Pastinen, T.; Pushel, I.; et al. Evaluation of Hi-C Sequencing for Detection of Gene Fusions in Hematologic and Solid Tumor Pediatric Cancer Samples. Cancers 2024, 16, 2936. [Google Scholar] [CrossRef]
- Harewood, L.; Kishore, K.; Eldridge, M.D.; Wingett, S.; Pearson, D.; Schoenfelder, S.; Collins, V.P.; Fraser, P. Hi-C as a tool for precise detection and characterisation of chromosomal rearrangements and copy number variation in human tumours. Genome Biol. 2017, 18, 125. [Google Scholar] [CrossRef]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Burton, J.N.; Adey, A.; Patwardhan, R.P.; Qiu, R.; Kitzman, J.O.; Shendure, J. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 2013, 31, 1119–1125. [Google Scholar] [CrossRef]
- Rickman, D.S.; Soong, T.D.; Moss, B.; Mosquera, J.M.; Dlabal, J.; Terry, S.; MacDonald, T.Y.; Tripodi, J.; Bunting, K.; Najfeld, V.; et al. Oncogene-mediated alterations in chromatin conformation. Proc. Natl. Acad. Sci. USA 2012, 109, 9083–9088. [Google Scholar] [CrossRef] [PubMed]
- Song, F.; Xu, J.; Dixon, J.; Yue, F. Analysis of Hi-C Data for Discovery of Structural Variations in Cancer. Methods Mol. Biol. 2022, 2301, 143–161. [Google Scholar] [CrossRef] [PubMed]
- Fang, H.; Eacker, S.M.; Wu, Y.; Neufeld-Kaiser, W.; Laurino, M.; Keel, S.; Horwitz, M.S.; Liu, Y.J. Genetic and functional characterization of inherited complex chromosomal rearrangements in a family with multisystem anomalies. Genet. Med. Open 2025, 3, 103423. [Google Scholar] [CrossRef]
- Fang, H.; Eacker, S.M.; Wu, Y.; Paschal, C.; Wood, M.; Nelson, B.; Muratov, A.; Liu, Y. Evaluation of Genomic Proximity Mapping for Detecting Genomic and Chromosomal Structural Variants in Constitutional Disorders. J. Mol. Diagn. 2025, 27, 1054–1069. [Google Scholar] [CrossRef] [PubMed]
- Zook, J.M.; Catoe, D.; McDaniel, J.; Vang, L.; Spies, N.; Sidow, A.; Weng, Z.; Liu, Y.; Mason, C.E.; Alexander, N.; et al. Extensive sequencing of seven human genomes to characterize benchmark reference materials. Sci. Data 2016, 3, 160025. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Lee, S.; Bakker, C.R.; Vitzthum, C.; Alver, B.H.; Park, P.J. Pairs and Pairix: A file format and a tool for efficient storage and retrieval for Hi-C read pairs. Bioinformatics 2022, 38, 1729–1731. [Google Scholar] [CrossRef]
- Abdennur, N.; Mirny, L.A. Cooler: Scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics 2020, 36, 311–316. [Google Scholar] [CrossRef]
- Gonzales, P.R. Integration of Newer Genomic Technologies into Clinical Cytogenetics Laboratories. Genes 2025, 16, 688. [Google Scholar] [CrossRef]
- Hans, C.P.; Weisenburger, D.D.; Greiner, T.C.; Gascoyne, R.D.; Delabie, J.; Ott, G.; Muller-Hermelink, H.K.; Campo, E.; Braziel, R.M.; Jaffe, E.S.; et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 2004, 103, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Javed, S.; Hassan, J.; Naz, M.; Shan, S.; Abid, M.; Shamsi, T.S. Cytogenetic culture failure and its causes in hematological disorders; a single centre perspective. Mol. Cytogenet. 2023, 16, 4. [Google Scholar] [CrossRef] [PubMed]
- Di Giacomo, D.; La Starza, R.; Gorello, P.; Pellanera, F.; Kalender Atak, Z.; De Keersmaecker, K.; Pierini, V.; Harrison, C.J.; Arniani, S.; Moretti, M.; et al. 14q32 rearrangements deregulating BCL11B mark a distinct subgroup of T-lymphoid and myeloid immature acute leukemia. Blood 2021, 138, 773–784. [Google Scholar] [CrossRef] [PubMed]
- Somasundaram, R.; Jensen, C.T.; Tingvall-Gustafsson, J.; Ahsberg, J.; Okuyama, K.; Prasad, M.; Hagman, J.R.; Wang, X.; Soneji, S.; Strid, T.; et al. EBF1 and PAX5 control pro-B cell expansion via opposing regulation of the Myc gene. Blood 2021, 137, 3037–3049. [Google Scholar] [CrossRef]
- Aukema, S.M.; Kreuz, M.; Kohler, C.W.; Rosolowski, M.; Hasenclever, D.; Hummel, M.; Kuppers, R.; Lenze, D.; Ott, G.; Pott, C.; et al. Biological characterization of adult MYC-translocation-positive mature B-cell lymphomas other than molecular Burkitt lymphoma. Haematologica 2014, 99, 726–735. [Google Scholar] [CrossRef]
- Aukema, S.M.; van Pel, R.; Nagel, I.; Bens, S.; Siebert, R.; Rosati, S.; van den Berg, E.; Bosga-Bouwer, A.G.; Kibbelaar, R.E.; Hoogendoorn, M.; et al. MYC expression and translocation analyses in low-grade and transformed follicular lymphoma. Histopathology 2017, 71, 960–971. [Google Scholar] [CrossRef]
- Bertrand, P.; Bastard, C.; Maingonnat, C.; Jardin, F.; Maisonneuve, C.; Courel, M.N.; Ruminy, P.; Picquenot, J.M.; Tilly, H. Mapping of MYC breakpoints in 8q24 rearrangements involving non-immunoglobulin partners in B-cell lymphomas. Leukemia 2007, 21, 515–523. [Google Scholar] [CrossRef]
- Martin-Garcia, D.; Navarro, A.; Valdes-Mas, R.; Clot, G.; Gutierrez-Abril, J.; Prieto, M.; Ribera-Cortada, I.; Woroniecka, R.; Rymkiewicz, G.; Bens, S.; et al. CCND2 and CCND3 hijack immunoglobulin light-chain enhancers in cyclin D1− mantle cell lymphoma. Blood 2019, 133, 940–951. [Google Scholar] [CrossRef]
- Grimaldi, J.C.; Meeker, T.C. The t(5;14) chromosomal translocation in a case of acute lymphocytic leukemia joins the interleukin-3 gene to the immunoglobulin heavy chain gene. Blood 1989, 73, 2081–2085. [Google Scholar] [CrossRef]
- Meeker, T.C.; Hardy, D.; Willman, C.; Hogan, T.; Abrams, J. Activation of the interleukin-3 gene by chromosome translocation in acute lymphocytic leukemia with eosinophilia. Blood 1990, 76, 285–289. [Google Scholar] [CrossRef]
- Swerdlow, S.H.; Campo, E.; Harris, N.L.; Jaffe, E.S.; Pileri, S.A.; Stein, H.; Thiele, J.; Arber, D.A.; Hasserjian, R.P.; Le Beau, M.M.; et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th ed.; International Agency for Research on Cancer: Lyon, France, 2017. [Google Scholar]
- Fournier, B.; Balducci, E.; Duployez, N.; Clappier, E.; Cuccuini, W.; Arfeuille, C.; Caye-Eude, A.; Delabesse, E.; Bottollier-Lemallaz Colomb, E.; Nebral, K.; et al. B-ALL with t(5;14)(q31;q32); IGH-IL3 Rearrangement and Eosinophilia: A Comprehensive Analysis of a Peculiar IGH-Rearranged B-ALL. Front. Oncol. 2019, 9, 1374. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Mizuta, S.; Yamane, N.; Ueno, H.; Yoshida, K.; Kato, I.; Umeda, K.; Hiramatsu, H.; Suehiro, M.; Maihara, T.; et al. Paraneoplastic hypereosinophilic syndrome associated with IL3-IgH positive acute lymphoblastic leukemia. Pediatr. Blood Cancer 2019, 66, e27449. [Google Scholar] [CrossRef] [PubMed]
- Montefiori, L.E.; Bendig, S.; Gu, Z.; Chen, X.; Polonen, P.; Ma, X.; Murison, A.; Zeng, A.; Garcia-Prat, L.; Dickerson, K.; et al. Enhancer Hijacking Drives Oncogenic BCL11B Expression in Lineage-Ambiguous Stem Cell Leukemia. Cancer Discov. 2021, 11, 2846–2867. [Google Scholar] [CrossRef] [PubMed]
- Gianni, F.; Belver, L.; Ferrando, A. The Genetics and Mechanisms of T-Cell Acute Lymphoblastic Leukemia. Cold Spring Harb. Perspect. Med. 2020, 10, a035246. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, X.; Fang, H.; Wu, Y.; Meshinchi, S.; Naresh, K.N.; Reister, E.; Langford, K.; Eacker, S.M.; Liu, Y.J. Comprehensive Detection of Chromosomal and Genomic Abnormalities via Next-Generation Sequencing-Based Genomic Proximity Mapping Improves Diagnostic Classification of Hematologic Neoplasms. Cancers 2025, 17, 3775. https://doi.org/10.3390/cancers17233775
Chen X, Fang H, Wu Y, Meshinchi S, Naresh KN, Reister E, Langford K, Eacker SM, Liu YJ. Comprehensive Detection of Chromosomal and Genomic Abnormalities via Next-Generation Sequencing-Based Genomic Proximity Mapping Improves Diagnostic Classification of Hematologic Neoplasms. Cancers. 2025; 17(23):3775. https://doi.org/10.3390/cancers17233775
Chicago/Turabian StyleChen, Xueyan, He Fang, Yu Wu, Soheil Meshinchi, Kikkeri N. Naresh, Emily Reister, Kyle Langford, Stephen M. Eacker, and Yajuan J. Liu. 2025. "Comprehensive Detection of Chromosomal and Genomic Abnormalities via Next-Generation Sequencing-Based Genomic Proximity Mapping Improves Diagnostic Classification of Hematologic Neoplasms" Cancers 17, no. 23: 3775. https://doi.org/10.3390/cancers17233775
APA StyleChen, X., Fang, H., Wu, Y., Meshinchi, S., Naresh, K. N., Reister, E., Langford, K., Eacker, S. M., & Liu, Y. J. (2025). Comprehensive Detection of Chromosomal and Genomic Abnormalities via Next-Generation Sequencing-Based Genomic Proximity Mapping Improves Diagnostic Classification of Hematologic Neoplasms. Cancers, 17(23), 3775. https://doi.org/10.3390/cancers17233775






