ITGA2 Mediates the Resistance of Hepatocellular Carcinoma to Lenvatinib by Activating the AKT/FOXO3A Signaling Pathway
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Bioinformatics Analysis
2.2. Cell Culture and Transfection
2.3. Protein Extraction and WB
2.4. CCK-8 Assay
2.5. Colony Formation Assay
2.6. Flow Cytometry for Detecting Apoptosis and the Cell Cycle
2.7. Animal Experiments
2.8. Immunohistochemistry and Scoring
2.9. Transcriptome Sequencing
2.10. Statistical Analysis
2.11. Visualization
3. Results
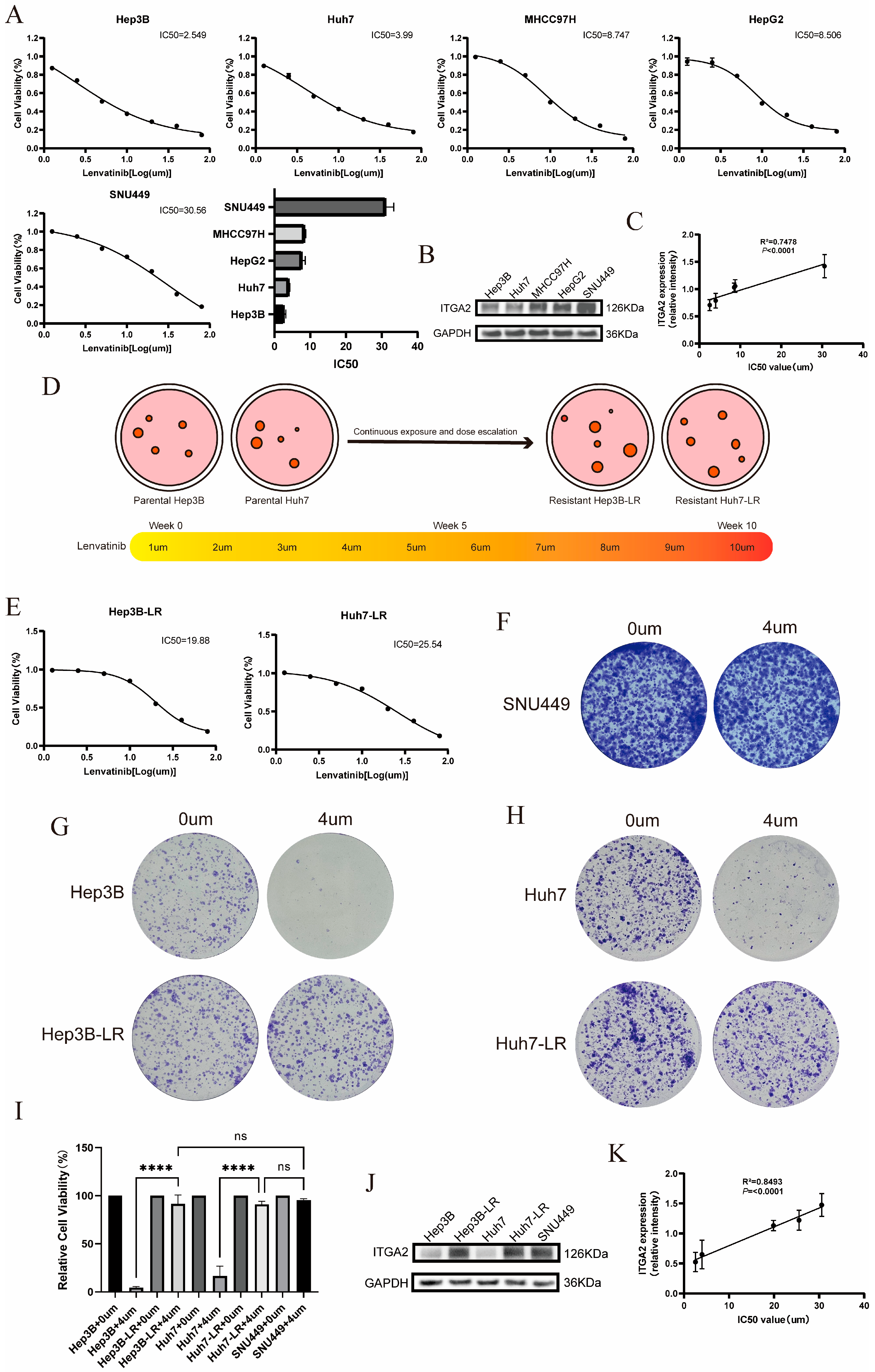
3.1. The Expression Level of ITGA2 Is Negatively Correlated with the Prognosis of Liver Cancer Patients and the Sensitivity to Lenvatinib
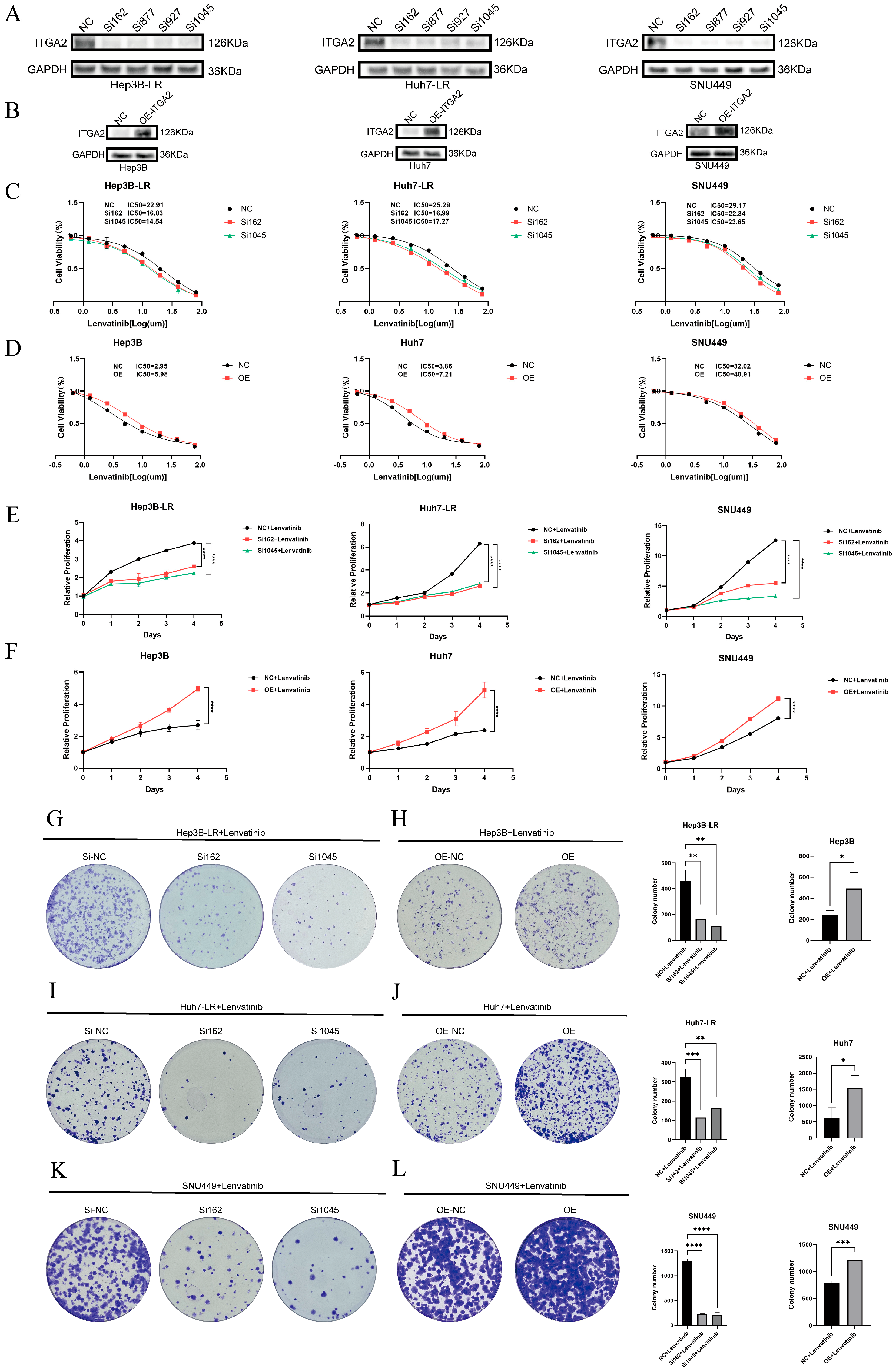
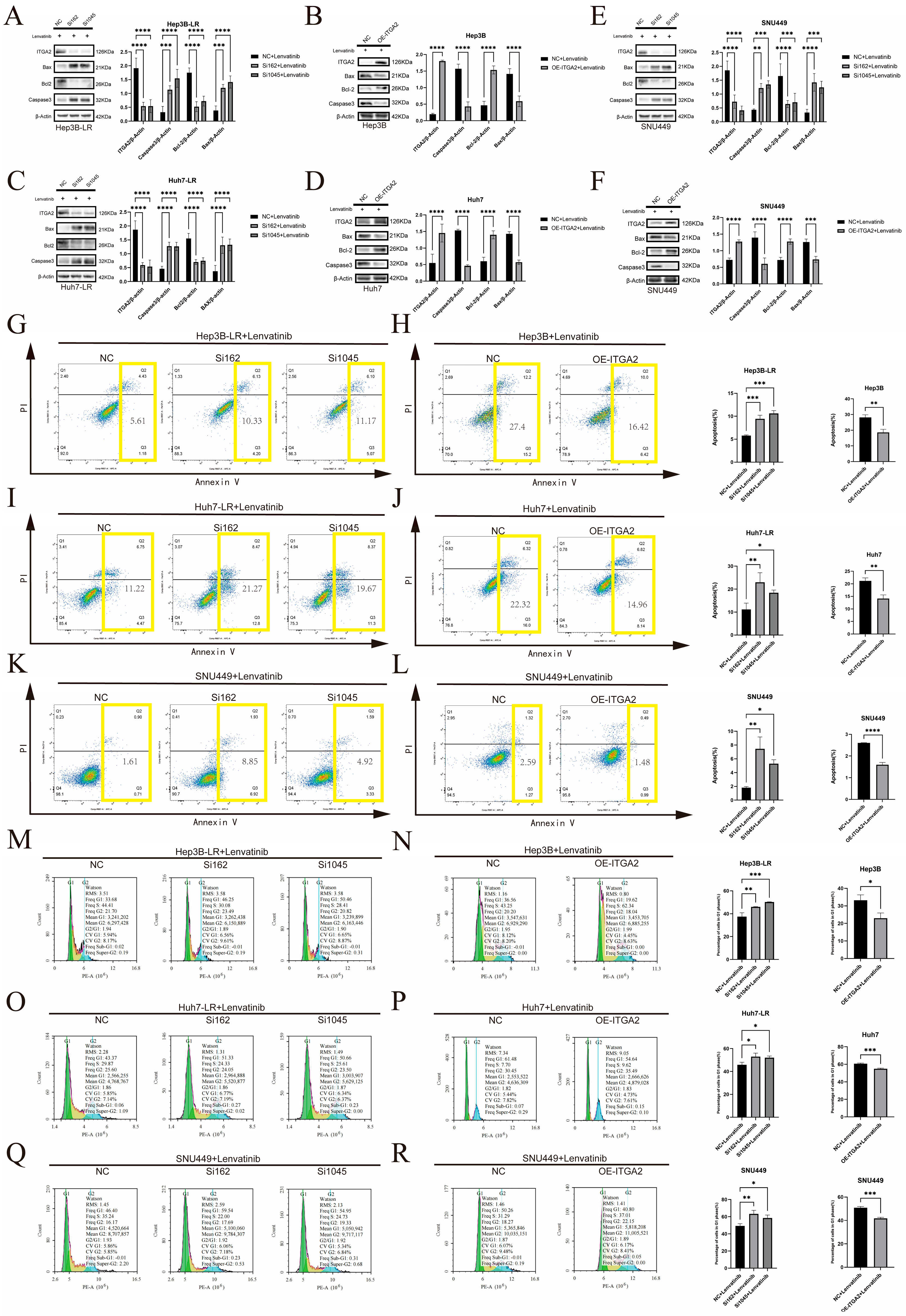
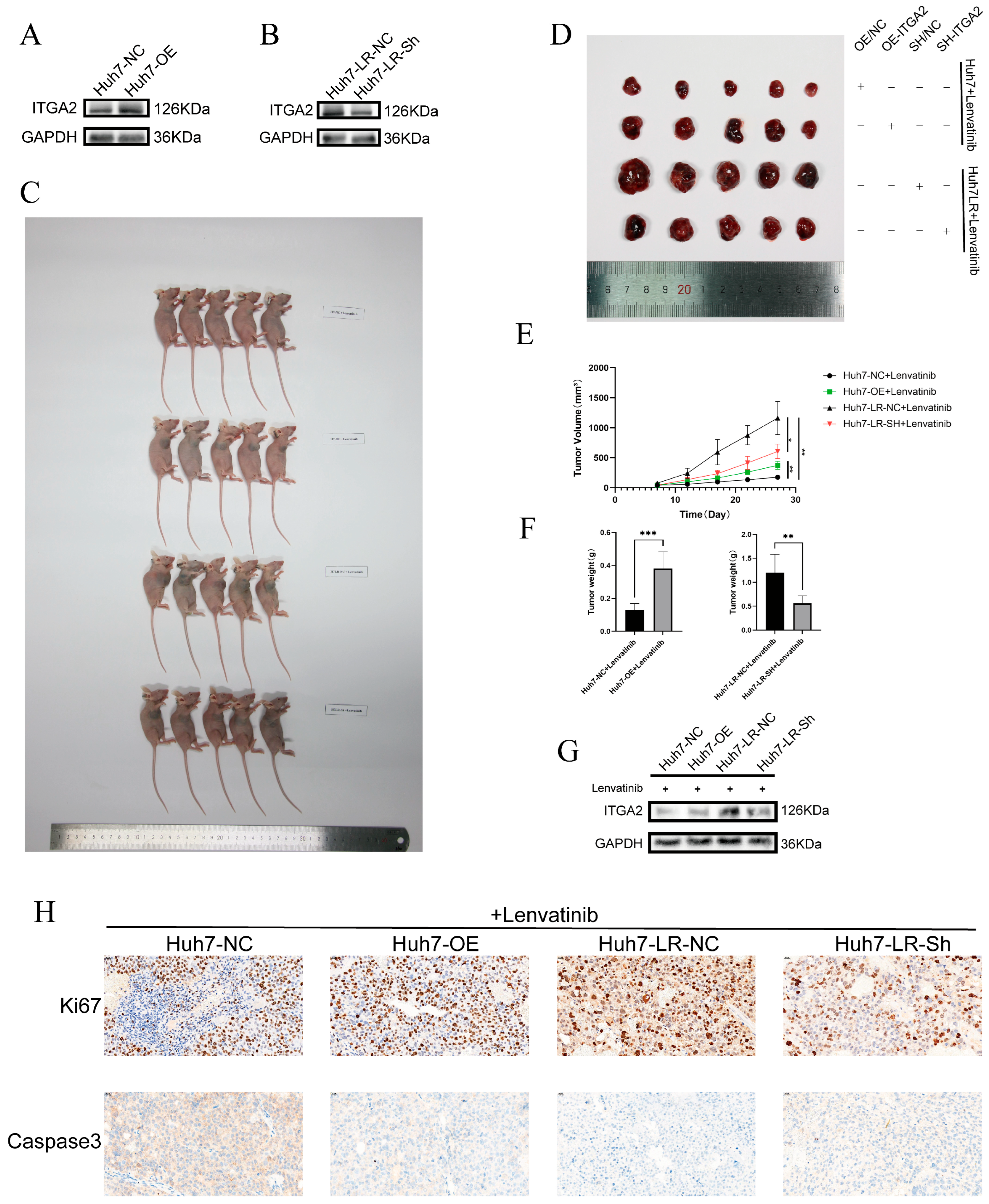
3.2. ITGA2 Promotes the Resistance of Hepatocellular Carcinoma to Lenvatinib
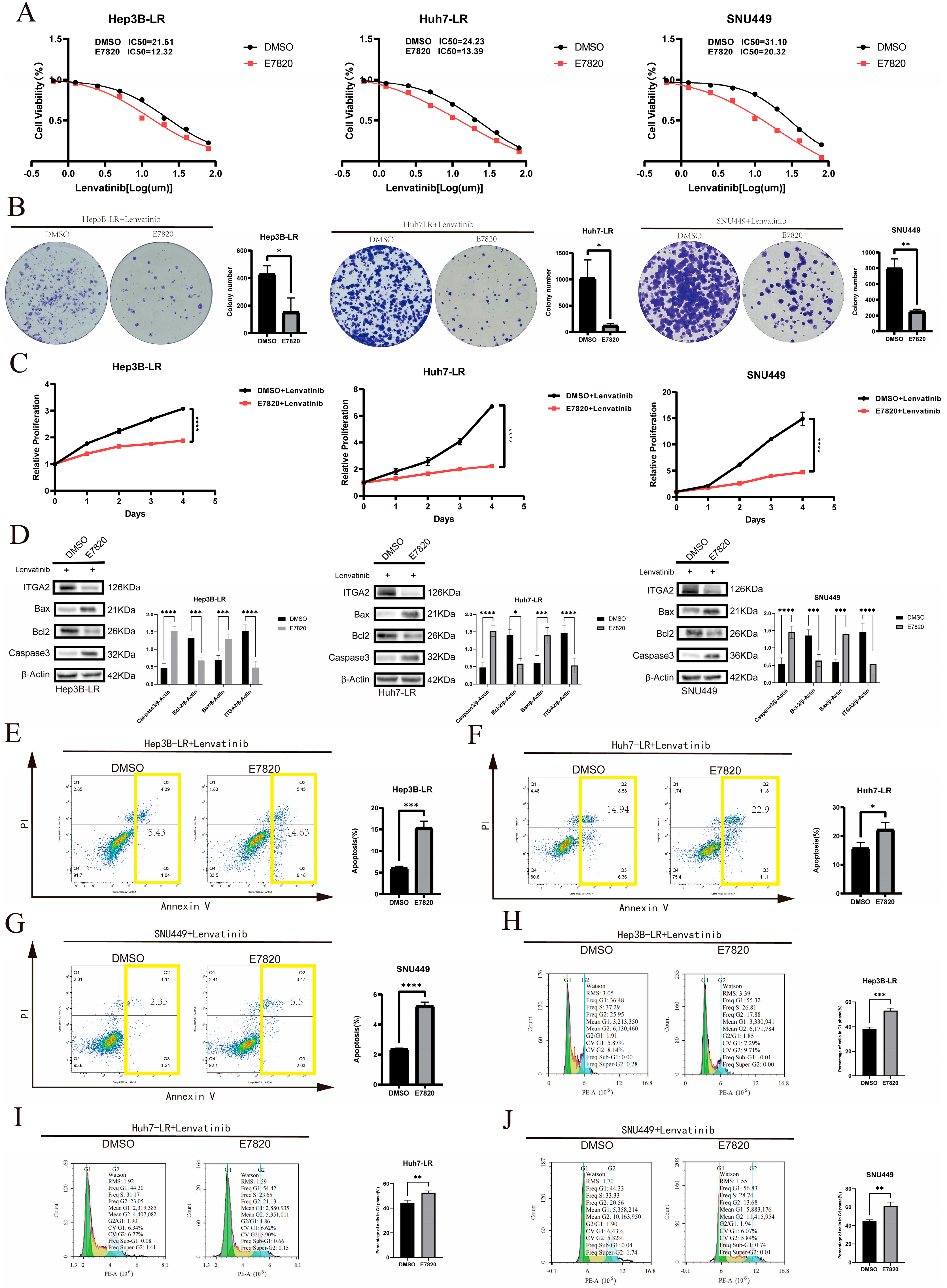
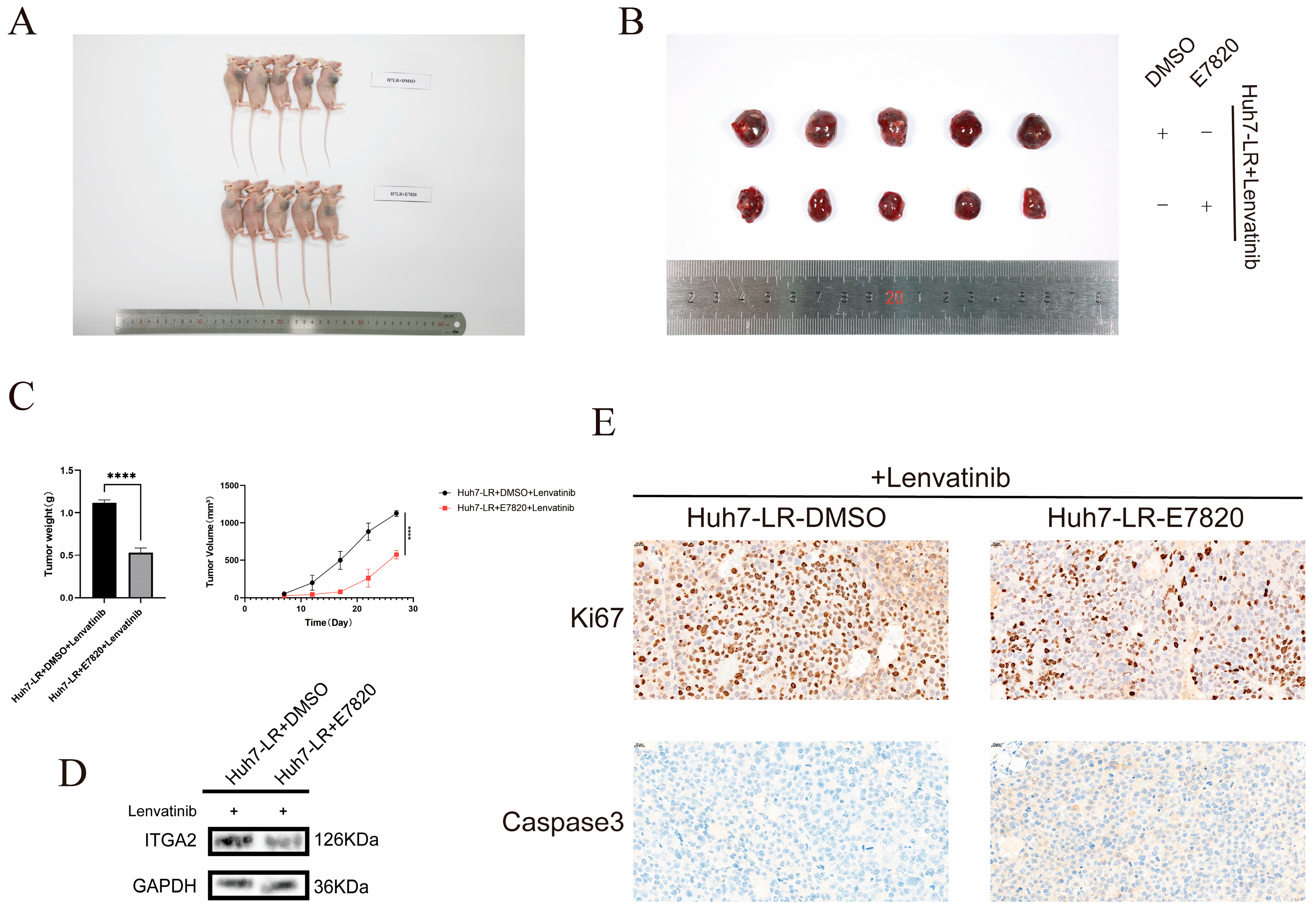
3.3. The ITGA2 Inhibitor E7820 Enhances the Sensitivity of Hepatocellular Carcinoma to Lenvatinib
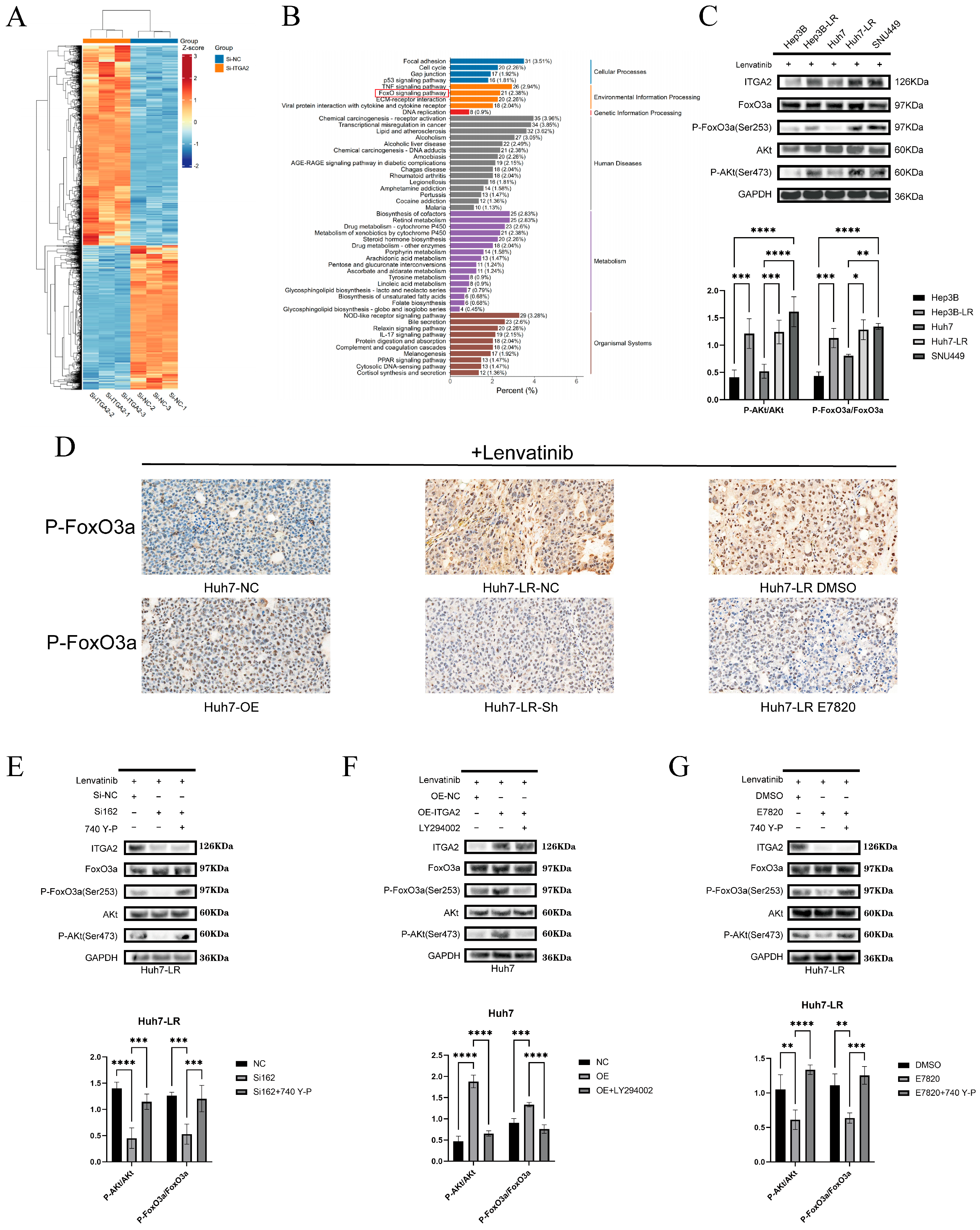
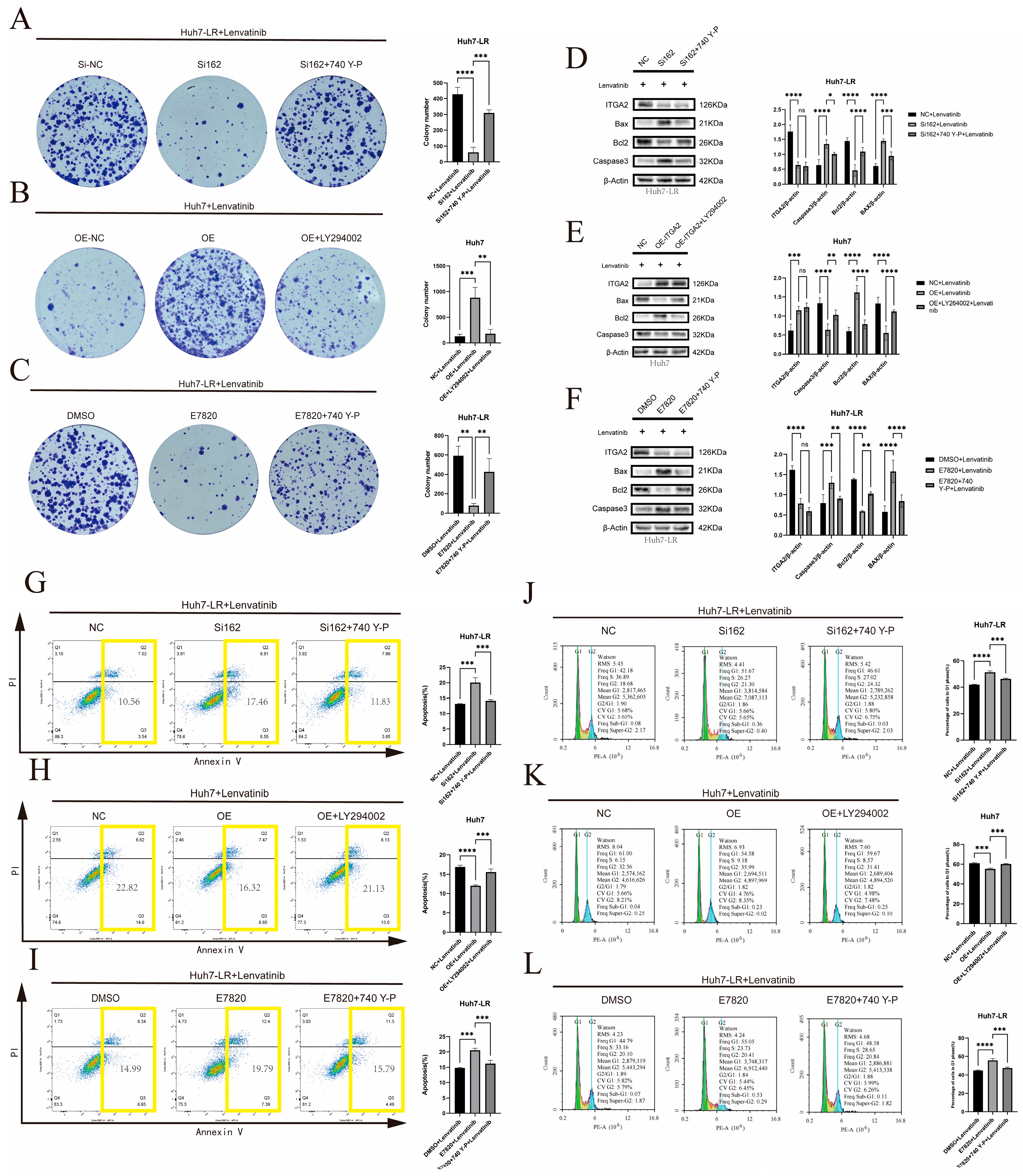
3.4. The ITGA2/AKT/FOXO3A Signaling Axis Promotes Resistance to Lenvatinib in Hepatocellular Carcinoma
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| HCC | Hepatocellular Carcinoma |
| ICC | Intrahepatic Cholangiocarcinoma |
| cHCC-CCA | Combined Hepatocellular-Cholangiocarcinoma |
| TACE | Transarterial Chemoembolization |
| TKI | Tyrosine Kinase Inhibitor |
| ECM | Extracellular Matrix |
| GSCA | Gene Set Cancer Analysis |
| CTRP | The Cancer Therapeutics Response Portal |
| WB | Western Blotting |
| CCK-8 | Cell Counting Kit-8 |
| PVDF | Polyvinylidene Fluoride |
| FBS | Fetal Bovine Serum |
| SPF | Specific Pathogen-Free |
| IACUC | Institutional Animal Care and Use Committee |
| OS | Overall Survival |
| PFS | Progression-Free Survival |
| IC50 | Half-Maximal Inhibitory Concentration |
| CI | Combination Index |
| EMT | Epithelial-Mesenchymal Transition |
| FAK | Focal Adhesion Kinase |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| siRNA | Small Interfering RNA |
| shRNA | Short Hairpin RNA |
| KO | Knockout |
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Zheng, R.; Zhang, S.; Zeng, H.; Wang, S.; Sun, K.; Chen, N.R.; Li, L.; Wei, W.; He, J. Cancer incidence and mortality in China 2016. J. Natl. Cancer Cent. 2022, 2, 1–9. [Google Scholar] [CrossRef]
- Mcglynn, K.A.; Petrick, J.L.; El-Serag, H.B. Epidemiology of Hepatocellular Carcinoma. Hepatology 2021, 73 (Suppl. 1), 4–13. [Google Scholar] [CrossRef] [PubMed]
- Forner, A.; Reig, M.; Bruix, J. Hepatocellular carcinoma. Lancet 2018, 391, 1301–1314. [Google Scholar] [CrossRef]
- Llovet, J.M.; Ricci, S.; Mazzaferro, V.; Hilgard, P.; Gane, E.; Blanc, J.-F.; de Olieira, A.C.; Santoro, A.; Raoul, J.-L.; Forner, A.; et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 2008, 359, 378–390. [Google Scholar] [CrossRef]
- Kudo, M.; Finn, R.S.; Qin, S.; Han, K.-H.; Ikeda, K.; Piscaglia, F.; Baron, A.; Park, J.-W.; Han, G.; Jassem, J.; et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: A randomised phase 3 non-inferiority trial. Lancet 2018, 391, 1163–1173. [Google Scholar] [CrossRef]
- Ladd, A.D.; Duarte, S.; Sahin, I.; Zarrinpar, A. Mechanisms of drug resistance in HCC. Hepatology 2024, 79, 926–940. [Google Scholar] [CrossRef]
- Kanchanawong, P.; Calderwood, D.A. Organization, dynamics and mechanoregulation of integrin-mediated cell-ECM adhesions. Nat. Rev. Mol. Cell Biol. 2023, 24, 142–161. [Google Scholar] [CrossRef]
- Li, M.; Wang, Y.; Li, M.; Wu, X.; Setrerrahmane, S.; Xu, H. Integrins as attractive targets for cancer therapeutics. Acta Pharm. Sin. B 2021, 11, 2726–2737. [Google Scholar] [CrossRef]
- Javadi, S.; Zhiani, M.; Mousavi, M.A.; Fathi, M. Crosstalk between Epidermal Growth Factor Receptors (EGFR) and integrins in resistance to EGFR tyrosine kinase inhibitors (TKIs) in solid tumors. Eur. J. Cell Biol. 2020, 99, 151083. [Google Scholar] [CrossRef] [PubMed]
- Takada, Y.; Ye, X.; Simon, S. The integrins. Genome Biol. 2007, 8, 215. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, Y.; Che, X.; Hou, K.; Wu, J.; Zheng, C.; Cheng, Y.; Liu, Y.; Hu, X.; Zhang, J. Integrin α5 promotes migration and invasion through the FAK/STAT3/AKT signaling pathway in icotinib-resistant non-small cell lung cancer cells. Oncol. Lett. 2021, 22, 556. [Google Scholar] [CrossRef]
- Gregori, A.; Bergonzini, C.; Capula, M.; Mantini, G.; Khojasteh-Leylakoohi, F.; Comandatore, A.; Khalili-Tanha, G.; Khooei, A.; Morelli, L.; Avan, A.; et al. Prognostic Significance of Integrin Subunit Alpha 2 (ITGA2) and Role of Mechanical Cues in Resistance to Gemcitabine in Pancreatic Ductal Adenocarcinoma (PDAC). Cancers 2023, 15, 628. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Bridgeman, B.; Malnassy, G.; Ding, X.; Cotler, S.J.; Dhanarajan, A.; Qiu, W. Integrin subunit beta 8 contributes to lenvatinib resistance in HCC. Hepatol. Commun. 2022, 6, 1786–1802. [Google Scholar] [CrossRef] [PubMed]
- Gang, E.J.; Kim, H.N.; Hsieh, Y.-T.; Ruan, Y.; Ogana, H.A.; Lee, S.; Pham, J.; Geng, H.; Park, E.; Klemm, L.; et al. Integrin α6 mediates the drug resistance of acute lymphoblastic B-cell leukemia. Blood 2020, 136, 210–223. [Google Scholar] [CrossRef]
- Shi, Y.; Shang, J.; Li, Y.; Zhong, D.; Zhang, Z.; Yang, Q.; Lai, C.; Feng, T.; Yao, Y.; Huang, X. ITGA5 and ITGB1 contribute to Sorafenib resistance by promoting vasculogenic mimicry formation in hepatocellular carcinoma. Cancer Med. 2023, 12, 3786–3796. [Google Scholar] [CrossRef]
- Slack, R.J.; Macdonald, S.J.F.; Roper, J.A.; Jenkins, R.G.; Hatley, R.J.D. Emerging therapeutic opportunities for integrin inhibitors. Nat. Rev. Drug Discov. 2022, 21, 60–78. [Google Scholar] [CrossRef]
- Su, C.-Y.; Li, J.-Q.; Zhang, L.-L.; Wang, H.; Wang, F.-H.; Tao, Y.-W.; Wang, Y.-Q.; Guo, Q.-R.; Li, J.-J.; Liu, Y.; et al. The Biological Functions and Clinical Applications of Integrins in Cancers. Front. Pharmacol. 2020, 11, 579068. [Google Scholar] [CrossRef]
- Pang, X.; He, X.; Qiu, Z.; Zhang, H.; Xie, R.; Liu, Z.; Gu, Y.; Zhao, N.; Xiang, Q.; Cui, Y. Targeting integrin pathways: Mechanisms and advances in therapy. Signal Transduct. Target. Ther. 2023, 8, 1–42. [Google Scholar] [CrossRef] [PubMed]
- Kariya, Y.; Nishita, M. Integrins in Cancer Drug Resistance: Molecular Mechanisms and Clinical Implications. Int. J. Mol. Sci. 2025, 26, 3143. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Shi, Y.; Lv, Y.; Yuan, S.; Ramirez, C.F.A.; Lieftink, C.; Wang, L.; Wang, S.; Wang, C.; Dias, M.H.; et al. EGFR activation limits the response of liver cancer to lenvatinib. Nature 2021, 595, 730–734. [Google Scholar] [CrossRef]
- Qin, J.; Hu, S.; Lou, J.; Xu, M.; Gao, R.; Xiao, Q.; Chen, Y.; Ding, M.; Pan, Y.; Wang, S. Selumetinib overcomes ITGA2-induced 5-fluorouracil resistance in colorectal cancer. Int. Immunopharmacol. 2024, 137, 112487. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Cao, T.; Guo, K.; Zhou, Y.; Liu, H.; Pan, Y.; Hou, Q.; Nie, Y.; Fan, D.; Lu, Y.; et al. Regulation of Integrin Subunit Alpha 2 by miR-135b-5p Modulates Chemoresistance in Gastric Cancer. Front. Oncol. 2020, 10, 308. [Google Scholar] [CrossRef]
- Zhang, L.; Huang, Y.; Ling, J.; Zhuo, W.; Yu, Z.; Luo, Y.; Zhu, Y. Is Integrin Subunit Alpha 2 Expression a Prognostic Factor for Liver Carcinoma? A Validation Experiment Based on Bioinformatics Analysis. Pathol. Oncol. Res. 2019, 25, 1545–1552. [Google Scholar] [CrossRef]
- Rattanasinchai, C.; Navasumrit, P.; Ruchirawat, M. Elevated ITGA2 expression promotes collagen type I-induced clonogenic growth of intrahepatic cholangiocarcinoma. Sci. Rep. 2022, 12, 22429. [Google Scholar] [CrossRef]
- Huang, W.; Zhu, J.; Shi, H.; Wu, Q.; Zhang, C. ITGA2 Overexpression Promotes Esophageal Squamous Cell Carcinoma Aggression via FAK/AKT Signaling Pathway. OncoTargets Ther. 2021, 14, 3583–3596. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Duan, Y.; Li, X.; Li, Z.; Hu, J.; Shi, H.; Su, Z.; Li, Z.; Du, B.; Chen, Y.; et al. High expression ITGA2 affects the expression of MET, PD-L1, CD4 and CD8 with the immune microenvironment in pancreatic cancer patients. Front. Immunol. 2023, 14, 1209367. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Zhao, J.; Sun, Y.; Li, D.; Meng, Z.; Wang, B.; Fan, P.; Liu, Z.; Jin, X.; Wu, H. Overexpressed ITGA2 promotes malignant tumor aggression by up-regulating PD-L1 expression through the activation of the STAT3 signaling pathway. J. Exp. Clin. Cancer Res. 2019, 38, 485. [Google Scholar] [CrossRef]
- Hornsveld, M.; Dansen, T.B.; Derksen, P.W.; Burgering, B.M.T. Re-evaluating the role of FOXOs in cancer. Semin. Cancer Biol. 2018, 50, 90–100. [Google Scholar] [CrossRef]
- Farhan, M.; Wang, H.; Gaur, U.; Little, P.J.; Xu, J.; Zheng, W. FOXO Signaling Pathways as Therapeutic Targets in Cancer. Int. J. Biol. Sci. 2017, 13, 815–827. [Google Scholar] [CrossRef]
- Sun, B.; Chen, H.; Wang, X.; Chen, T. Regorafenib induces Bim-mediated intrinsic apoptosis by blocking AKT-mediated FOXO3a nuclear export. Cell Death Discov. 2023, 9, 37. [Google Scholar] [CrossRef]
- Fan, Z.; Lin, W.-H.; Liang, C.; Li, Y.; Peng, C.-J.; Luo, J.-S.; Tang, W.-Y.; Zheng, L.-M.; Huang, D.-P.; Ke, Z.-Y.; et al. MG132 inhibits proliferation and induces apoptosis of acute lymphoblastic leukemia via Akt/FOXO3a/Bim pathway. Hum. Exp. Toxicol. 2024, 43, 9603271241303030. [Google Scholar] [CrossRef]
- Tan, J.; Xiang, Y.; Xiong, Y.; Zhang, Y.; Qiao, B.; Zhang, H. Crebanine induces ROS-dependent apoptosis in human hepatocellular carcinoma cells via the AKT/FoxO3a signaling pathway. Front. Pharmacol. 2023, 14, 1069093. [Google Scholar] [CrossRef]
- Guan, L.; Zhang, L.; Gong, Z.; Hou, X.; Xu, Y.; Feng, X.; Wang, H.; You, H. FoxO3 inactivation promotes human cholangiocarcinoma tumorigenesis and chemoresistance through Keap1-Nrf2 signaling. Hepatology 2016, 63, 1914–1927. [Google Scholar] [CrossRef]
- Liu, M.; Xu, C.; Qin, X.; Liu, W.; Li, D.; Jia, H.; Gao, X.; Wu, Y.; Wu, Q.; Xu, X.; et al. DHW-221, a Dual PI3K/mTOR Inhibitor, Overcomes Multidrug Resistance by Targeting P-Glycoprotein (P-gp/ABCB1) and Akt-Mediated FOXO3a Nuclear Translocation in Non-small Cell Lung Cancer. Front. Oncol. 2022, 12, 873649. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Cao, Q.; Lin, S.; Chen, Y.; Liu, X.; Hong, Q. Knockdown of CALM2 increases the sensitivity to afatinib in HER2-amplified gastric cancer cells by regulating the Akt/FoxO3a/Puma axis. Toxicol. Vitr. 2023, 87, 105531. [Google Scholar] [CrossRef]
- Chen, Y.; Jin, L.; Ma, Y.; Liu, Y.; Zhu, Q.; Huang, Y.; Feng, W. BACH1 promotes lung adenocarcinoma cell metastasis through transcriptional activation of ITGA2. Cancer Sci. 2023, 114, 3568–3582. [Google Scholar] [CrossRef]
- Qi, X.; Zhou, J.; Wang, P.; Li, Y.; Li, H.; Miao, Y.; Ma, X.; Luo, X.; Zhang, Z.; He, Y.; et al. KLF7-regulated ITGA2 as a therapeutic target for inhibiting oral cancer stem cells. Cell Death Dis. 2025, 16, 354. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Gao, Y.; Zhao, X.; Guo, C.; Wang, X.; Yang, Y.; Han, C.; Zhao, L.; Qin, Y.; Liu, L.; et al. HOXD3 was negatively regulated by YY1 recruiting HDAC1 to suppress progression of hepatocellular carcinoma cells via ITGA2 pathway. Cell Prolif. 2020, 53, e12835. [Google Scholar] [CrossRef]
- Qu, H.; Sun, L.; Li, X.; Liu, F.; Sun, H.; He, X.; Gan, D.; Yin, Y.; Tian, B.; Chen, F.; et al. Long non-coding RNA AC018926.2 regulates palmitic acid exposure-compromised osteogenic potential of periodontal ligament stem cells via the ITGA2/FAK/AKT pathway. Cell Prolif. 2023, 56, e13411. [Google Scholar] [CrossRef]
- Yu, J.; Zhang, C.; Yu, Q.; Yu, H.; Zhang, B. ADAR1 p110 Enhances Adhesion of Tumor Cells to Extracellular Matrix in Hepatocellular Carcinoma via Up-Regulating ITGA2 Expression. Med. Sci. Monit. 2019, 25, 1469–1479. [Google Scholar] [CrossRef]
- Dong, J.; Wang, R.; Ren, G.; Li, X.; Wang, J.; Sun, Y.; Liang, J.; Nie, Y.; Wu, K.; Feng, B.; et al. HMGA2-FOXL2 Axis Regulates Metastases and Epithelial-to-Mesenchymal Transition of Chemoresistant Gastric Cancer. Clin. Cancer Res. 2017, 23, 3461–3473. [Google Scholar] [CrossRef]
- Cai, H.; Guo, F.; Wen, S.; Jin, X.; Wu, H.; Ren, D. Overexpressed integrin alpha 2 inhibits the activation of the transforming growth factor β pathway in pancreatic cancer via the TFCP2-SMAD2 axis. J. Exp. Clin. Cancer Res. 2022, 41, 73. [Google Scholar] [CrossRef]
- Zheng, J.; Wang, S.; Xia, L.; Sun, Z.; Chan, K.M.; Bernards, R.; Qin, W.; Chen, J.; Xia, Q.; Jin, H. Hepatocellular carcinoma: Signaling pathways and therapeutic advances. Signal Transduct. Target. Ther. 2025, 10, 35. [Google Scholar] [CrossRef]
- Mita, M.; Kelly, K.R.; Mita, A.; Ricart, A.D.; Romero, O.; Tolcher, A.; Hook, L.; Okereke, C.; Krivelevich, I.; Rossignol, D.P.; et al. Phase I study of E7820, an oral inhibitor of integrin alpha-2 expression with antiangiogenic properties, in patients with advanced malignancies. Clin. Cancer Res. 2011, 17, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Kerklaan, B.M.; Slater, S.; Flynn, M.; Greystoke, A.; Witteveen, P.O.; Megui-Roelvink, M.; de Vos, F.; Dean, E.; Reyderman, L.; Ottesen, L.; et al. A phase I, dose escalation, pharmacodynamic, pharmacokinetic, and food-effect study of α2 integrin inhibitor E7820 in patients with advanced solid tumors. Investig. New Drugs 2016, 34, 329–337. [Google Scholar] [CrossRef] [PubMed]
- Ito, K.; Semba, T.; Uenaka, T.; Wakabayashi, T.; Asada, M.; Funahashi, Y. Enhanced anti-angiogenic effect of E7820 in combination with erlotinib in epidermal growth factor receptor-tyrosine kinase inhibitor-resistant non-small-cell lung cancer xenograft models. Cancer Sci. 2014, 105, 1023–1031. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zheng, L.; Chen, W.; Zhang, X.; Lv, D.; Zeng, C.; Zhang, C.; Zhang, R.; Kang, T.; Zhong, L. Targeting RBM39 suppresses tumor growth and sensitizes osteosarcoma cells to cisplatin. Oncogene 2025, 44, 575–586. [Google Scholar] [CrossRef]
- Wu, X.; Cai, J.; Zuo, Z.; Li, J. Collagen facilitates the colorectal cancer stemness and metastasis through an integrin/PI3K/AKT/Snail signaling pathway. Biomed. Pharmacother. 2019, 114, 108708. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, K.; Xu, B.; Gong, L.; Zhong, M.; Han, C.; Liu, Y.; Yin, Z.; Liang, X.; Wang, Q.; Ye, G.; et al. ITGA2 Mediates the Resistance of Hepatocellular Carcinoma to Lenvatinib by Activating the AKT/FOXO3A Signaling Pathway. Cancers 2025, 17, 2846. https://doi.org/10.3390/cancers17172846
Gong K, Xu B, Gong L, Zhong M, Han C, Liu Y, Yin Z, Liang X, Wang Q, Ye G, et al. ITGA2 Mediates the Resistance of Hepatocellular Carcinoma to Lenvatinib by Activating the AKT/FOXO3A Signaling Pathway. Cancers. 2025; 17(17):2846. https://doi.org/10.3390/cancers17172846
Chicago/Turabian StyleGong, Kai, Bin Xu, Lian Gong, Ming Zhong, Chun Han, Yuechuan Liu, Zeli Yin, Xiangnan Liang, Qiuxiang Wang, Genhua Ye, and et al. 2025. "ITGA2 Mediates the Resistance of Hepatocellular Carcinoma to Lenvatinib by Activating the AKT/FOXO3A Signaling Pathway" Cancers 17, no. 17: 2846. https://doi.org/10.3390/cancers17172846
APA StyleGong, K., Xu, B., Gong, L., Zhong, M., Han, C., Liu, Y., Yin, Z., Liang, X., Wang, Q., Ye, G., Sun, B., & Wang, L. (2025). ITGA2 Mediates the Resistance of Hepatocellular Carcinoma to Lenvatinib by Activating the AKT/FOXO3A Signaling Pathway. Cancers, 17(17), 2846. https://doi.org/10.3390/cancers17172846




