Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers
Simple Summary
Abstract
1. Introduction
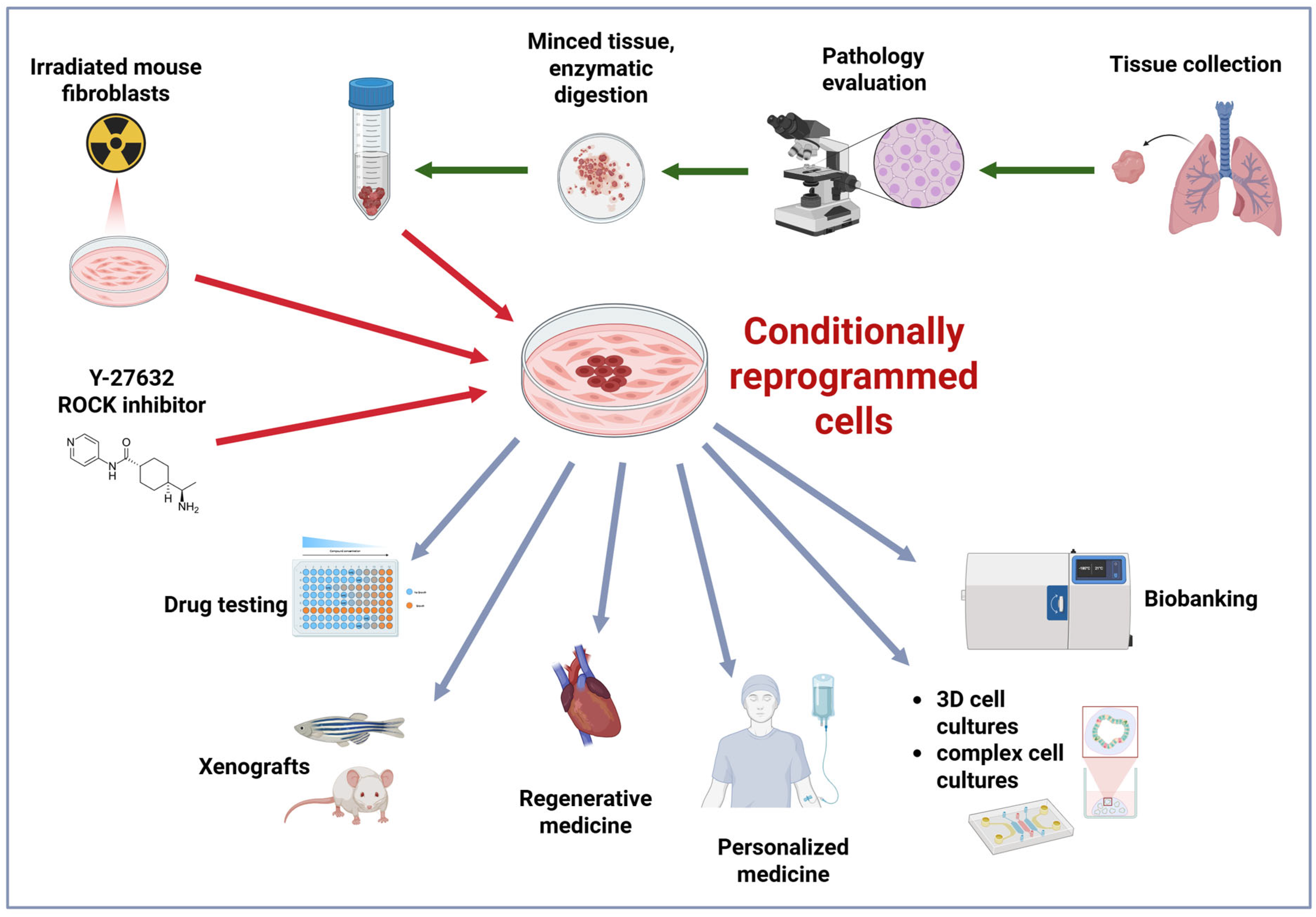
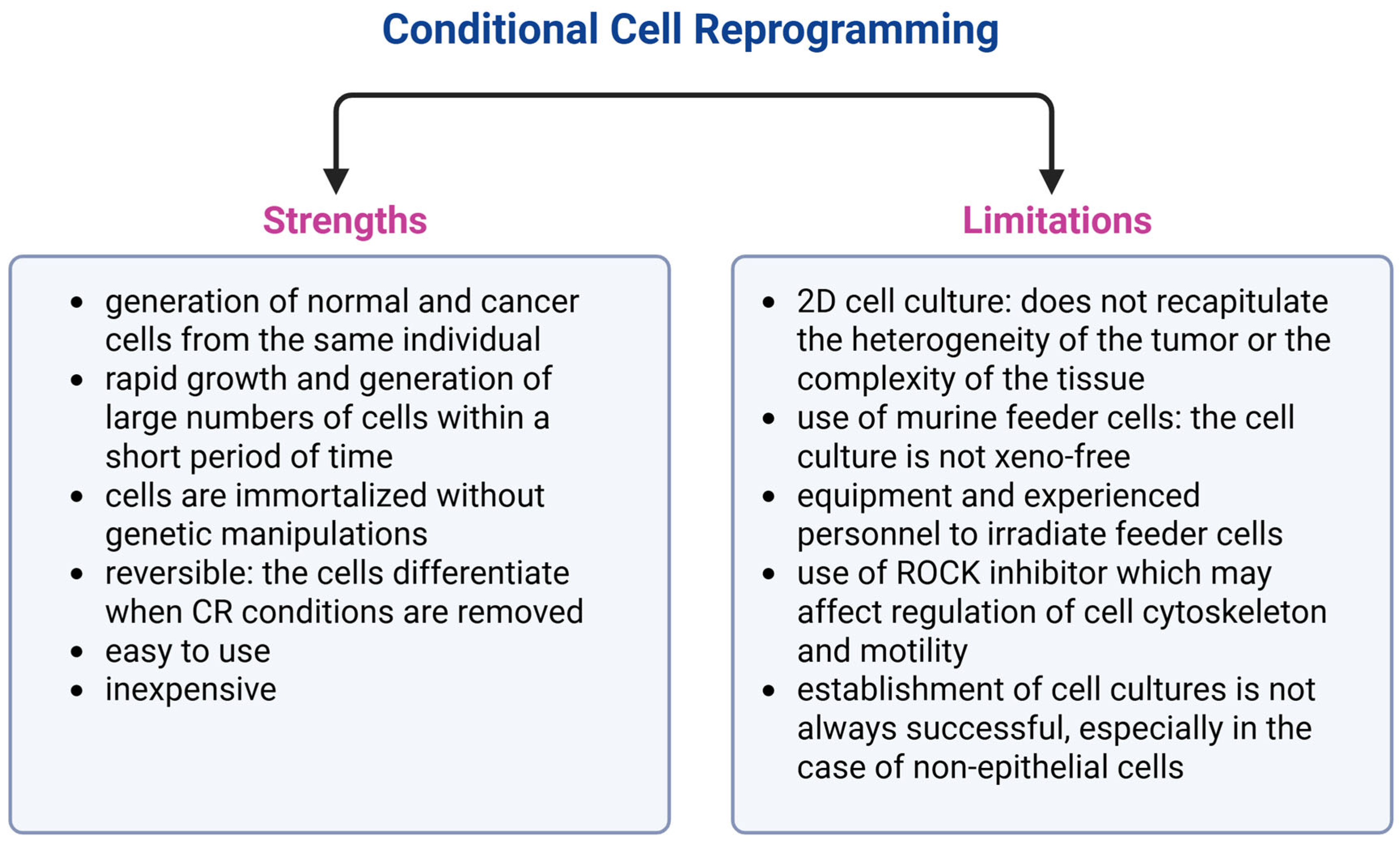
2. Conditional Cell Reprogramming
3. Conditional Cell Reprogramming Applications in Cancer Research
4. Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers
4.1. Neuroblastoma
4.2. Neuroendocrine Cervical Carcinoma
4.3. Neuroendocrine Prostate Cancer
4.4. GIST
4.5. Gliomas and Other Glioneural Tumors
4.5.1. Ependymoma
4.5.2. Pilocytic Astrocytoma
4.5.3. Other Gliomas
4.6. Ameloblastoma
4.7. Laryngeal and Hypopharyngeal Carcinoma
4.8. Adenoid Cystic Carcinoma
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Krattiger, L.A.; Guex, A.G. Complex in Vitro Model Systems to Understand the Biointerfaces of Dental Implants. Dent. Mater. 2025, 41, 810–826. [Google Scholar] [CrossRef] [PubMed]
- Abbot, A. Cell Culture: Biology’s New Dimension. Nature 2003, 424, 870–872. [Google Scholar] [CrossRef] [PubMed]
- Aljitawi, O.S.; Li, D.; Xiao, Y.; Zhang, D.; Ramachandran, K.; Stehno-Bittel, L.; Van Veldhuizen, P.; Lin, T.L.; Kambhampati, S.; Garimella, R. A Novel Three-Dimensional Stromal-Based Model for in Vitro Chemotherapy Sensitivity Testing of Leukemia Cells. Leuk. Lymphoma 2014, 55, 378–391. [Google Scholar] [CrossRef]
- Senft, D.; Leiserson, M.D.M.; Ruppin, E.; Ronai, Z.A. Precision Oncology: The Road Ahead. Trends Mol. Med. 2017, 23, 874–898. [Google Scholar] [CrossRef]
- Novelli, G.; Spitalieri, P.; Murdocca, M.; Centanini, E.; Sangiuolo, F. Organoid Factory: The Recent Role of the Human Induced Pluripotent Stem Cells (HiPSCs) in Precision Medicine. Front. Cell Dev. Biol. 2023, 10, 1059579. [Google Scholar] [CrossRef]
- He, Z.; Wilson, A.; Rich, F.; Kenwright, D.; Stevens, A.; Low, Y.S.; Thunders, M. Chromosomal Instability and Its Effect on Cell Lines. Cancer Rep. 2023, 6, e1822. [Google Scholar] [CrossRef]
- Krüger, M.; Kopp, S. Tumor Models and Drug Targeting In Vitro—Where Are We Today? Where Do We Go from Here? Cancers 2023, 15, 1768. [Google Scholar] [CrossRef]
- Krendl, F.J.; Primavesi, F.; Oberhuber, R.; Neureiter, D.; Ocker, M.; Bekric, D.; Kiesslich, T.; Mayr, C. The Importance of Preclinical Models for Cholangiocarcinoma Drug Discovery. Expert. Opin. Drug Discov. 2025, 20, 205–216. [Google Scholar] [CrossRef]
- Movia, D.; Prina-Mello, A. (Eds.) Cancer Cell Culture Methods and Protocols Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2023. [Google Scholar]
- Festing, S.; Wilkinson, R. The Ethics of Animal Research: Talking Point on the Use of Animals in Scientific Research. EMBO Rep. 2007, 8, 526–530. [Google Scholar] [CrossRef]
- Charitos, I.A.; Ballini, A.; Cantore, S.; Boccellino, M.; Di Domenico, M.; Borsani, E.; Nocini, R.; Di Cosola, M.; Santacroce, L.; Bottalico, L. Stem Cells: A Historical Review about Biological, Religious, and Ethical Issues. Stem Cells Int. 2021, 2021, 9978837. [Google Scholar] [CrossRef] [PubMed]
- Boehm, J.S.; Golub, T.R. An Ecosystem of Cancer Cell Line Factories to Support a Cancer Dependency Map. Nat. Rev. Genet. 2015, 16, 373–374. [Google Scholar] [CrossRef]
- Barretina, J.; Caponigro, G.; Stransky, N.; Venkatesan, K.; Margolin, A.A.; Kim, S.; Wilson, C.J.; Lehár, J.; Kryukov, G.V.; Sonkin, D.; et al. The Cancer Cell Line Encyclopedia Enables Predictive Modelling of Anticancer Drug Sensitivity. Nature 2012, 483, 603–607. [Google Scholar] [CrossRef]
- Voskoglou-Nomikos, T.; Pater, J.L.; Seymour, L. Clinical Predictive Value of the in Vitro Cell Line, Human Xenograft, and Mouse Allograft Preclinical Cancer Models. Clin. Cancer Res. 2003, 9, 4227–4239. [Google Scholar] [PubMed]
- Palechor-Ceron, N.; Krawczyk, E.; Dakic, A.; Simic, V.; Yuan, H.; Blancato, J.; Wang, W.; Hubbard, F.; Zheng, Y.L.; Dan, H.; et al. Conditional Reprogramming for Patient-Derived Cancer Models and next-Generation Living Biobanks. Cells 2019, 8, 1327. [Google Scholar] [CrossRef] [PubMed]
- Coppola, L.; Cianflone, A.; Grimaldi, A.M.; Incoronato, M.; Bevilacqua, P.; Messina, F.; Baselice, S.; Soricelli, A.; Mirabelli, P.; Salvatore, M. Biobanking in Health Care: Evolution and Future Directions. J. Transl. Med. 2019, 17, 172. [Google Scholar] [CrossRef]
- Augustine, R.; Kalva, S.N.; Ahmad, R.; Zahid, A.A.; Hasan, S.; Nayeem, A.; McClements, L.; Hasan, A. 3D Bioprinted Cancer Models: Revolutionizing Personalized Cancer Therapy. Transl. Oncol. 2021, 14, 101015. [Google Scholar] [CrossRef] [PubMed]
- Grainger, D.W. Cell-Based Drug Testing; This World is Not Flat. Adv. Drug Deliv. Rev. 2014, 69–70, vii–xi. [Google Scholar] [CrossRef]
- Bhuker, S.; Sinha, A.K.; Arora, A.; Tuli, H.S.; Datta, S.; Saini, A.K.; Saini, R.V.; Ramniwas, S. Genes and Proteins Expression Profile of 2D vs 3D Cancer Models: A Comparative Analysis for Better Tumor Insights. Cytotechnology 2025, 77, 51. [Google Scholar] [CrossRef]
- Bittman-Soto, X.S.; Thomas, E.S.; Ganshert, M.E.; Mendez-Santacruz, L.L.; Harrell, J.C. The Transformative Role of 3D Culture Models in Triple-Negative Breast Cancer Research. Cancers 2024, 16, 1859. [Google Scholar] [CrossRef]
- Hickman, J.A.; Graeser, R.; de Hoogt, R.; Vidic, S.; Brito, C.; Gutekunst, M.; van der Kuip, H.; Imi Predect consortium. Three-Dimensional Models of Cancer for Pharmacology and Cancer Cell Biology: Capturing Tumor Complexity in Vitro/Ex Vivo. Biotechnol. J. 2014, 9, 1115–1128. [Google Scholar] [CrossRef]
- Kamb, A. What’s Wrong with Our Cancer Models? Nat. Rev. Drug Discov. 2005, 4, 161–165. [Google Scholar] [CrossRef]
- Ben-David, U.; Siranosian, B.; Ha, G.; Tang, H.; Oren, Y.; Hinohara, K.; Strathdee, C.A.; Dempster, J.; Lyons, N.J.; Burns, R.; et al. Genetic and Transcriptional Evolution Alters Cancer Cell Line Drug Response. Nature 2018, 560, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Santillo, S. Changes in Biophysical Properties of Undifferentiated SH-SY5Y Cells During Long-Term Cultures. Neuroscience 2022, 482, 143–158. [Google Scholar] [CrossRef] [PubMed]
- Nestor, C.E.; Ottaviano, R.; Reinhardt, D.; Cruickshanks, H.A.; Mjoseng, H.K.; McPherson, R.C.; Lentini, A.; Thomson, J.P.; Dunican, D.S.; Pennings, S.; et al. Rapid Reprogramming of Epigenetic and Transcriptional Profiles in Mammalian Culture Systems. Genome Biol. 2015, 16, 11. [Google Scholar] [CrossRef] [PubMed]
- Shay, J.; Wright, W. Hayflick, His Limit, and Cellular Ageing. Nat. Rev. Mol. Cell Biol. 2000, 1, 72–76. [Google Scholar] [CrossRef]
- Drexler, H.G.; Dirks, W.G.; Matsuo, Y.; MacLeod, R.A.F. False Leukemia-Lymphoma Cell Lines: An Update on over 500 Cell Lines. Leukemia 2003, 17, 416–426. [Google Scholar] [CrossRef]
- Lacroix, M. Persistent Use of “False” Cell Lines. Int. J. Cancer 2008, 122, 1–4. [Google Scholar] [CrossRef]
- Ye, F.; Chen, C.; Qin, J.; Liu, J.; Zheng, A.C. Genetic Profiling Reveals an Alarming Rate of Cross-Contamination among Human Cell Lines Used in China. FASEB J. 2015, 29, 4268–4272. [Google Scholar] [CrossRef]
- Weiskirchen, S.; Schröder, S.K.; Buhl, E.M.; Weiskirchen, R. A Beginner’s Guide to Cell Culture: Practical Advice for Preventing Needless Problems. Cells 2023, 12, 682. [Google Scholar] [CrossRef]
- Hughes, P.; Marshall, D.; Reid, Y.; Parkes, H.; Gelber, C. The Costs of Using Unauthenticated, over-Passaged Cell Lines: How Much More Data Do We Need? Biotechniques 2007, 43, 575–586. [Google Scholar] [CrossRef]
- Alston-Roberts, C.; Barallon, R.; Bauer, S.R.; Butler, J.; Capes-Davis, A.; Dirks, W.G.; Elmore, E.; Furtado, M.; Kerrigan, L.; Kline, M.C.; et al. Cell Line Misidentification: The Beginning of the End. Nat. Rev. Cancer 2010, 10, 441–448. [Google Scholar] [CrossRef]
- Boyd, N.; Dancey, J.E.; Gilks, B.; Huntsman, D.G. Rare Cancers: A Sea of Opportunity. Lancet Oncol. 2016, 17, e52–e61. [Google Scholar] [CrossRef]
- Beltinger, C.; Debatin, K.M. Murine Models for Experimental Therapy of Pediatric Solid Tumors with Poor Prognosis. Int. J. Cancer 2001, 92, 313–318. [Google Scholar] [CrossRef]
- Kawai, A.; Higashi, T.; Shibata, T.; Yoshida, A.; Katoh, Y.; Fujiwara, Y.; Nishida, T. Rare Cancers in Japan: Definition, Clinical Features and Future Perspectives. Jpn. J. Clin. Oncol. 2020, 50, 970–975. [Google Scholar] [CrossRef] [PubMed]
- Kondo, T. Current Status and Future Outlook for Patient-Derived Cancer Models from a Rare Cancer Research Perspective. Cancer Sci. 2021, 112, 953–961. [Google Scholar] [CrossRef] [PubMed]
- Sharifnia, T.; Hong, A.L.; Painter, C.A.; Boehm, J.S. Emerging Opportunities for Target Discovery in Rare Cancers. Cell Chem. Biol. 2017, 24, 1075–1091. [Google Scholar] [CrossRef]
- Liu, X.; Ory, V.; Chapman, S.; Yuan, H.; Albanese, C.; Kallakury, B.; Timofeeva, O.A.; Nealon, C.; Dakic, A.; Simic, V.; et al. ROCK Inhibitor and Feeder Cells Induce the Conditional Reprogramming of Epithelial Cells. Am. J. Pathol. 2012, 180, 599–607. [Google Scholar] [CrossRef]
- Liu, X.; Krawczyk, E.; Suprynowicz, F.A.; Palechor-Ceron, N.; Yuan, H.; Dakic, A.; Simic, V.; Zheng, Y.L.; Sripadhan, P.; Chen, C.; et al. Conditional Reprogramming and Long-Term Expansion of Normal and Tumor Cells from Human Biospecimens. Nat. Protoc. 2017, 12, 439–451. [Google Scholar] [CrossRef]
- Friedman, A.A.; Letai, A.; Fisher, D.E.; Flaherty, K.T. Precision Medicine for Cancer with Next-Generation Functional Diagnostics. Nat. Rev. Cancer 2015, 15, 747–756. [Google Scholar] [CrossRef]
- Lisanti, M.P.; Tanowitz, H.B. Translational Discoveries, Personalized Medicine, and Living Biobanks of the Future. Am. J. Pathol. 2012, 180, 1334–1336. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wu, X.; Wang, S.; Li, M.; Li, J.; Shen, J.; Zhao, Y.; Pang, J.; Wen, Q.; Chen, M.; Wei, B.; et al. Conditional Reprogramming: Next Generation Cell Culture. Acta Pharm. Sin. B 2020, 10, 1360–1381. [Google Scholar] [CrossRef]
- Yuan, H.; Myers, S.; Wang, J.; Zhou, D.; Woo, J.A.; Kallakury, B.; Ju, A.; Bazylewicz, M.; Carter, Y.M.; Albanese, C.; et al. Use of Reprogrammed Cells to Identify Therapy for Respiratory Papillomatosis. N. Engl. J. Med. 2012, 367, 1220–1227. [Google Scholar] [CrossRef]
- Dakic, A.; Divito, K.; Fang, S.; Suprynowicz, F.; Gaur, A.; Li, X.; Palechor-Ceron, N.; Simic, V.; Choudhury, S.; Yu, S.; et al. ROCK Inhibitor Reduces Myc-Induced Apoptosis and Mediates Immortalization of Human Keratinocytes. Oncotarget 2016, 7, 66740–66753. [Google Scholar] [CrossRef]
- Chapman, S.; Liu, X.; Meyers, C.; Schlegel, R.; McBride, A.A. Human Keratinocytes Are Efficiently Immortalized by a Rho Kinase Inhibitor. J. Clin. Investig. 2010, 120, 2619–2626. [Google Scholar] [CrossRef] [PubMed]
- Piltti, J.; Varjosalo, M.; Qu, C.; Häyrinen, J.; Lammi, M.J. Rho-Kinase Inhibitor Y-27632 Increases Cellular Proliferation and Migration in Human Foreskin Fibroblast Cells. Proteomics 2015, 15, 2953–2965. [Google Scholar] [CrossRef]
- Rheinwald, J.G.; Green, H. Serial Cultivation of Strains of Human Epidermal Keratinocytes: The Formation of Keratinizing Colonies from Single Cells. Cell 1975, 6, 331–343. [Google Scholar] [CrossRef] [PubMed]
- Suprynowicz, F.A.; Upadhyay, G.; Krawczyk, E.; Kramer, S.C.; Hebert, J.D.; Liu, X.; Yuan, H.; Cheluvaraju, C.; Clapp, P.W.; Boucher, R.C.; et al. Conditionally Reprogrammed Cells Represent a Stem-like State of Adult Epithelial Cells. Proc. Natl. Acad. Sci. USA 2012, 109, 20035–20040. [Google Scholar] [CrossRef] [PubMed]
- Timofeeva, O.A.; Palechor-Ceron, N.; Li, G.; Yuan, H.; Krawczyk, E.; Zhong, X.; Liu, G.; Upadhyay, G.; Dakic, A.; Yu, S.; et al. Conditionally Reprogrammed Normal and Primary Tumor Prostate Epithelial Cells: A Novel Patient-Derived Cell Model for Studies of Human Prostate Cancer. Oncotarget 2017, 8, 22741–22758. [Google Scholar] [CrossRef]
- Alamri, A.M.; Liu, X.; Blancato, J.K.; Haddad, B.R.; Wang, W.; Zhong, X.; Choudhary, S.; Krawczyk, E.; Kallakury, B.V.; Davidson, B.J.; et al. Expanding Primary Cells from Mucoepidermoid and Other Salivary Gland Neoplasms for Genetic and Chemosensitivity Testing. DMM Dis. Models Mech. 2018, 11, dmm031716. [Google Scholar] [CrossRef]
- Saenz, F.R.; Ory, V.; AlOtaiby, M.; Rosenfield, S.; Furlong, M.; Cavalli, L.R.; Johnson, M.D.; Liu, X.; Schlegel, R.; Wellstein, A.; et al. Conditionally Reprogrammed Normal and Transformed Mouse Mammary Epithelial Cells Display a Progenitor-Cell-like Phenotype. PLoS ONE 2014, 9, e97666. [Google Scholar] [CrossRef]
- Suprynowicz, F.A.; Kamonjoh, C.M.; Krawczyk, E.; Agarwal, S.; Wellstein, A.; Agboke, F.A.; Choudhury, S.; Liu, X.; Schlegel, R. Conditional Cell Reprogramming Involves Noncanonical β-Catenin Activation and MTOR-Mediated Inactivation of Akt. PLoS ONE 2017, 12, e0180897. [Google Scholar] [CrossRef]
- Chalak, M.; Hesaraki, M.; Mirbahari, S.N.; Yeganeh, M.; Abdi, S.; Rajabi, S.; Hemmatzadeh, F. Cell Immortality: In Vitro Effective Techniques to Achieve and Investigate Its Applications and Challenges. Life 2024, 14, 417. [Google Scholar] [CrossRef] [PubMed]
- Cerneckis, J.; Cai, H.; Shi, Y. Induced Pluripotent Stem Cells (IPSCs): Molecular Mechanisms of Induction and Applications. Signal Transduct. Target Ther. 2024, 9, 112. [Google Scholar] [CrossRef]
- He, Z.; Cole, K.D.; He, H.J. A Novel Immortalization Method for Immortalizing Human Primary CD8+ T Cells by Inserting a Single Copy of Human Telomerase Reverse Transcriptase via CRISPR/Cas9. Tissue Cell 2025, 95, 102908. [Google Scholar] [CrossRef] [PubMed]
- Gao, B.; Huang, C.; Kernstine, K.; Pelekanou, V.; Kluger, Y.; Jiang, T.; Peters-Hall, J.R.; Coquelin, M.; Girard, L.; Zhang, W.; et al. Non-Malignant Respiratory Epithelial Cells Preferentially Proliferate from Resected Non-Small Cell Lung Cancer Specimens Cultured under Conditionally Reprogrammed Conditions. Oncotarget 2017, 8, 1114. [Google Scholar] [CrossRef]
- Yu, F.; Lu, Y.; Tao, L.; Jiang, Y.Y.; Lin, D.C.; Wang, L.; Petersson, F.; Yoshiyama, H.; Koeffler, P.H.; Goh, B.C.; et al. Non-Malignant Epithelial Cells Preferentially Proliferate from Nasopharyngeal Carcinoma Biopsy Cultured under Conditionally Reprogrammed Conditions. Sci. Rep. 2017, 7, 17359. [Google Scholar] [CrossRef]
- Tricoli, L.; Naeem, A.; Parasido, E.; Mikhaiel, J.P.; Choudhry, M.U.; Berry, D.L.; Abdelgawad, I.A.; Lee, R.J.; Feldman, A.S.; Ihemelandu, C.; et al. Characterization of the Effects of Defined, Multidimensional Culture Conditions on Conditionally Reprogrammed Primary Human Prostate Cells. Oncotarget 2018, 9, 2193. [Google Scholar] [CrossRef]
- Krawczyk, E.; Hong, S.H.; Galli, S.; Trinh, E.; Wietlisbach, L.; Misiukiewicz, S.F.; Tilan, J.U.; Chen, Y.S.; Schlegel, R.; Kitlinska, J. Murine Neuroblastoma Cell Lines Developed by Conditional Reprogramming Preserve Heterogeneous Phenotypes Observed in Vivo. Lab. Investig. 2020, 100, 38–51. [Google Scholar] [CrossRef]
- Yuan, H.; Krawczyk, E.; Blancato, J.; Albanese, C.; Zhou, D.; Wang, N.; Paul, S.; Alkhilaiwi, F.; Palechor-Ceron, N.; Dakic, A.; et al. HPV Positive Neuroendocrine Cervical Cancer Cells Are Dependent on Myc but Not E6/E7 Viral Oncogenes. Sci. Rep. 2017, 7, 45617. [Google Scholar] [CrossRef]
- Alothman, S.J.; Kang, K.; Liu, X.; Krawczyk, E.; Azhar, R.I.; Hu, R.; Goerlitz, D.; Kallakury, B.V.; Furth, P.A. Characterization of Transcriptome Diversity and in Vitro Behavior of Primary Human High-Risk Breast Cells. Sci. Rep. 2022, 12, 6159. [Google Scholar] [CrossRef] [PubMed]
- Mondal, A.M.; Ma, A.H.; Li, G.; Krawczyk, E.; Yuan, R.; Lu, J.; Schlegel, R.; Stamatakis, L.; Kowalczyk, K.J.; Philips, G.K.; et al. Fidelity of a PDX-CR Model for Bladder Cancer. Biochem. Biophys. Res. Commun. 2019, 517, 49–56. [Google Scholar] [CrossRef]
- Borodovsky, A.; McQuiston, T.J.; Stetson, D.; Ahmed, A.; Whitston, D.; Zhang, J.; Grondine, M.; Lawson, D.; Challberg, S.S.; Zinda, M.; et al. Generation of Stable PDX Derived Cell Lines Using Conditional Reprogramming. Mol. Cancer 2017, 16, 177. [Google Scholar] [CrossRef]
- Daneshdoust, D.; Yin, M.; Luo, M.; Sundi, D.; Dang, Y.; Lee, C.; Li, J.; Liu, X. Conditional Reprogramming Modeling of Bladder Cancer for Clinical Translation. Cells 2023, 12, 1714. [Google Scholar] [CrossRef]
- Mahajan, A.S.; Sugita, B.M.; Duttargi, A.N.; Saenz, F.; Krawczyk, E.; McCutcheon, J.N.; Fonseca, A.S.; Kallakury, B.; Pohlmann, P.; Gusev, Y.; et al. Genomic Comparison of Early-Passage Conditionally Reprogrammed Breast Cancer Cells to Their Corresponding Primary Tumors. PLoS ONE 2017, 12, e0186190. [Google Scholar] [CrossRef] [PubMed]
- Zhong, M.; Fu, L. Culture and Application of Conditionally Reprogrammed Primary Tumor Cells. Gastroenterol. Rep. (Oxf.) 2020, 8, 224–233. [Google Scholar] [CrossRef]
- Zhou, X.; Kang, Y.; Chang, Y.; Xia, S.; Wu, M.; Liu, J.; Dong, D.; Zhang, W.; Chen, H.; Li, H. CRC Therapy Identifies Indian Hedgehog Signaling in Mouse Endometrial Epithelial Cells and Inhibition of Ihh-KLF9 as a Novel Strategy for Treating IUA. Cells 2022, 11, 4053. [Google Scholar] [CrossRef]
- Alkhilaiwi, F.; Wang, L.; Zhou, D.; Raudsepp, T.; Ghosh, S.; Paul, S.; Palechor-Ceron, N.; Brandt, S.; Luff, J.; Liu, X.; et al. Long-Term Expansion of Primary Equine Keratinocytes That Maintain the Ability to Differentiate into Stratified Epidermis. Stem Cell Res. Ther. 2018, 9, 181. [Google Scholar] [CrossRef]
- Zhou, D.; Wang, A.; Maxwell, S.; Schlegel, R.; Yuan, H. Long-Term Culture of Canine Ocular Cells That Maintain Canine Papillomaviruses. Viruses 2022, 14, 2675. [Google Scholar] [CrossRef] [PubMed]
- Narumiya, S.; Ishizaki, T.; Uehata, M. Use and Properties of ROCK-Specific Inhibitor Y-27632. Methods Enzymol. 2000, 325, 273–284. [Google Scholar] [PubMed]
- Ligaba, S.B.; Khurana, A.; Graham, G.; Krawczyk, E.; Jablonski, S.; Petricoin, E.F.; Glazer, R.I.; Upadhyay, G. Multifactorial Analysis of Conditional Reprogramming of Human Keratinocytes. PLoS ONE 2015, 10, e0116755. [Google Scholar] [CrossRef] [PubMed]
- Mondal, A.M.; Zhou, H.; Horikawa, I.; Suprynowicz, F.A.; Li, G.; Dakic, A.; Rosenthal, B.; Ye, L.; Harris, C.C.; Schlegel, R.; et al. Δ133p53α, a Natural P53 Isoform, Contributes to Conditional Reprogramming and Long-Term Proliferation of Primary Epithelial Cells. Cell Death Dis. 2018, 9, 750. [Google Scholar] [CrossRef]
- Ji, H.; Tang, H.; Lin, H.; Mao, J.; Gao, L.; Liu, J.; Wu, T. Rho/Rock Cross-Talks with Transforming Growth Factor-β/Smad Pathway Participates in Lung Fibroblast-Myofibroblast Differentiation. Biomed. Rep. 2014, 2, 787–792. [Google Scholar] [CrossRef] [PubMed]
- Hynds, R.E.; Gowers, K.H.C.; Nigro, E.; Butler, C.R.; Bonfanti, P.; Giangreco, A.; Prêle, C.M.; Janes, S.M. Cross-Talk between Human Airway Epithelial Cells and 3T3-J2 Feeder Cells Involves Partial Activation of Human MET by Murine HGF. PLoS ONE 2018, 13, e0197129. [Google Scholar] [CrossRef]
- Calautti, E.; Li, J.; Saoncella, S.; Brissette, J.L.; Goetinck, P.F. Phosphoinositide 3-Kinase Signaling to Akt Promotes Keratinocyte Differentiation versus Death. J. Biol. Chem. 2005, 280, 32856–32865. [Google Scholar] [CrossRef]
- Alitalo, K.; Kuismanen, E.; Myllyl, R.; Kiistala, U.; Asko-Seljavaara, S.; Vaheri, A. Extracellular Matrix Proteins of Human Epidermal Keratinocytes and Feeder 3T3 Cells. J. Cell Biol. 1982, 94, 497–505. [Google Scholar] [CrossRef]
- Palechor-Ceron, N.; Suprynowicz, F.A.; Upadhyay, G.; Dakic, A.; Minas, T.; Simic, V.; Johnson, M.; Albanese, C.; Schlegel, R.; Liu, X. Radiation Induces Diffusible Feeder Cell Factor(s) That Cooperate with ROCK Inhibitor to Conditionally Reprogram and Immortalize Epithelial Cells. Am. J. Pathol. 2013, 183, 1862–1870. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.J.; Hong, Y.; Yoon, Y.-J.; Sim, N.S.; Hong, S.-M.; Lim, J.-Y. Chemical Reprogramming Culture for the Expansion of Salivary Gland Epithelial Basal Progenitor Cells. Stem Cell Res. Ther. 2025, 16, 187. [Google Scholar] [CrossRef]
- Witkowski, T.A.; Li, B.; Andersen, J.G.; Kumar, B.; Mroz, E.A.; Rocco, J.W. Y-27632 Acts beyond ROCK Inhibition to Maintain Epidermal Stem-like-Cells in Culture. J. Cell Sci. 2023, 136, jcs260990. [Google Scholar] [CrossRef]
- Yugawa, T.; Nishino, K.; Ohno, S.; Nakahara, T.; Fujita, M.; Goshima, N.; Umezawa, A.; Kiyono, T. Noncanonical NOTCH Signaling Limits Self-Renewal of Human Epithelial and Induced Pluripotent Stem Cells through ROCK Activation. Mol. Cell Biol. 2013, 33, 4434–4447. [Google Scholar] [CrossRef] [PubMed]
- Correa, B.R.S.; Hu, J.; Penalva, L.O.F.; Schlegel, R.; Rimm, D.L.; Galante, P.A.F.; Agarwal, S. Patient-Derived Conditionally Reprogrammed Cells Maintain Intra-Tumor Genetic Heterogeneity. Sci. Rep. 2018, 8, 1327. [Google Scholar] [CrossRef]
- Krawczyk, E.; Kitlinska, J. Preclinical Models of Neuroblastoma—Current Status and Perspectives. Cancers 2023, 15, 3314. [Google Scholar] [CrossRef]
- Abualsaud, N.; Caprio, L.; Galli, S.; Krawczyk, E.; Alamri, L.; Zhu, S.; Gallicano, G.I.; Kitlinska, J. Neuropeptide Y/Y5 Receptor Pathway Stimulates Neuroblastoma Cell Motility Through RhoA Activation. Front. Cell Dev. Biol. 2021, 8, 627090. [Google Scholar] [CrossRef]
- Chen, C.; Choudhury, S.; Wangsa, D.; Lescott, C.J.; Wilkins, D.J.; Sripadhan, P.; Liu, X.; Wangsa, D.; Ried, T.; Moskaluk, C.; et al. A Multiplex Preclinical Model for Adenoid Cystic Carcinoma of the Salivary Gland Identifies Regorafenib as a Potential Therapeutic Drug. Sci. Rep. 2017, 7, 11410. [Google Scholar] [CrossRef]
- Daneshdoust, D.; Luo, M.; Li, Z.; Mo, X.; Alothman, S.; Kallakury, B.; Schlegel, R.; Zhang, J.; Guo, D.; Furth, P.A.; et al. Unlocking Translational Potential: Conditionally Reprogrammed Cells in Advancing Breast Cancer Research. Cells 2023, 12, 2388. [Google Scholar] [CrossRef]
- Vaclova, T.; Maguire, S.; Pugh, M.; Barry, P.; Orr, N. Abstract 816: Molecular and Genomic Characterization of a Newly Established Male Breast Cancer Cell Line. Cancer Res. 2017, 77, 816. [Google Scholar] [CrossRef]
- Jeong, Y.J.; Park, S.H.; Jeon, C.H. Detection of Circulating Tumor Cells in Patients with Breast Cancer Using the Conditionally Reprogrammed Cell Culture Method and Reverse Transcription-PCR of Htert and Mage A1-6. Oncol. Lett. 2020, 20, 78. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.D.; Dabbs, D.J.; Lee, A.V.; McGuire, K.P.; Ahrendt, G.M.; Bhargava, R.; Davidson, N.E.; Brufsky, A.M.; Johnson, R.R.; Oesterreich, S.; et al. Developing in Vitro Models of Human Ductal Carcinoma in Situ from Primary Tissue Explants. Breast Cancer Res. Treat. 2015, 153, 311–321. [Google Scholar] [CrossRef]
- Alamri, A.M.; Kang, K.; Groeneveld, S.; Wang, W.; Zhong, X.; Kallakury, B.; Hennighausen, L.; Liu, X.; Furth, P.A. Primary Cancer Cell Culture: Mammary-Optimized vs Conditional Reprogramming. Endocr. Relat. Cancer 2016, 23, 535–554. [Google Scholar] [CrossRef]
- Jung, M.; Kowalczyk, K.; Hankins, R.; Bandi, G.; Kallakury, B.; Carrasquilla, M.A.; Banerjee, P.P.; Grindrod, S.; Dritschilo, A. Novel Paired Normal Prostate and Prostate Cancer Model Cell Systems Derived from African American Patients. Cancer Res. Commun. 2022, 2, 1617–1625. [Google Scholar] [CrossRef] [PubMed]
- Naeem, A.; Dakshanamurthy, S.; Walthieu, H.; Parasido, E.; Avantaggiati, M.; Tricoli, L.; Kumar, D.; Lee, R.J.; Feldman, A.; Noon, M.S.; et al. Predicting New Drug Indications for Prostate Cancer: The Integration of an in Silico Proteochemometric Network Pharmacology Platform with Patient-Derived Primary Prostate Cells. Prostate 2020, 80, 1233–1243. [Google Scholar] [CrossRef]
- Elbialy, A.; Kappala, D.; Desai, D.; Wang, P.; Fadiel, A.; Wang, S.J.; Makary, M.S.; Lenobel, S.; Sood, A.; Gong, M.; et al. Patient-Derived Conditionally Reprogrammed Cells in Prostate Cancer Research. Cells 2024, 13, 1005. [Google Scholar] [CrossRef]
- Stover, E.H.; Oh, C.; Keskula, P.; Choudhury, A.D.; Tseng, Y.Y.; Adalsteinsson, V.A.; Lohr, J.G.; Thorner, A.R.; Ducar, M.; Kryukov, G.V.; et al. Implementation of a Prostate Cancer-Specific Targeted Sequencing Panel for Credentialing of Patient-Derived Cell Lines and Genomic Characterization of Patient Samples. Prostate 2022, 82, 584–597. [Google Scholar] [CrossRef]
- Cao, J.; Chan, W.C.; Chow, M.S.S. Use of Conditional Reprogramming Cell, Patient Derived Xenograft and Organoid for Drug Screening for Individualized Prostate Cancer Therapy: Current and Future Perspectives (Review). Int. J. Oncol. 2022, 60, 52. [Google Scholar] [CrossRef]
- Saeed, K.; Rahkama, V.; Eldfors, S.; Bychkov, D.; Mpindi, J.P.; Yadav, B.; Paavolainen, L.; Aittokallio, T.; Heckman, C.; Wennerberg, K.; et al. Comprehensive Drug Testing of Patient-Derived Conditionally Reprogrammed Cells from Castration-Resistant Prostate Cancer. Eur. Urol. 2017, 71, 319–327. [Google Scholar] [CrossRef]
- Choudhary, S.; Ramasundaram, P.; Dziopa, E.; Mannion, C.; Kissin, Y.; Tricoli, L.; Albanese, C.; Lee, W.; Zilberberg, J. Human Ex Vivo 3D Bone Model Recapitulates Osteocyte Response to Metastatic Prostate Cancer. Sci. Rep. 2018, 8, 17975. [Google Scholar] [CrossRef]
- Ringer, L.; Sirajuddin, P.; Tricoli, L.; Waye, S.; Choudhry, M.U.; Parasido, E.; Sivakumar, A.; Heckler, M.; Naeem, A.; Abdelgawad, I.; et al. The Induction of the P53 Tumor Suppressor Protein Bridges the Apoptotic and Autophagic Signaling Pathways to Regulate Cell Death in Prostate Cancer Cells. Oncotarget 2014, 5, 10678–10691. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ow, T.J.; Mehta, V.; Li, D.; Thomas, C.; Shrivastava, N.; Kawachi, N.; Gersten, A.J.; Zhu, J.; Schiff, B.A.; Smith, R.V.; et al. Characterization of a Diverse Set of Conditionally Reprogrammed Head and Neck Cancer Cell Cultures. Laryngoscope 2024, 134, 2748–2756. [Google Scholar] [CrossRef]
- Li, D.; Thomas, C.; Shrivastava, N.; Gersten, A.; Gadsden, N.; Schlecht, N.; Kawachi, N.; Schiff, B.A.; Smith, R.V.; Rosenblatt, G.; et al. Establishment of a Diverse Head and Neck Squamous Cancer Cell Bank Using Conditional Reprogramming Culture Methods. J. Med. Virol. 2023, 95, e28388. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, H. Conditional Reprogramming Technology: A New Tool for Personalized Medicine in Bladder Cancer? Transl. Cancer Res. 2019, 8, S636–S638. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Shan, D.; Wang, G.; Mao, X.; You, W.; Wang, X.; Wang, Z. Elafibranor Emerged as a Potential Chemotherapeutic Drug for Non-Muscle Invasive Bladder Cancer. Cell Insight 2024, 3, 100149. [Google Scholar] [CrossRef]
- Kettunen, K.; Boström, P.J.; Lamminen, T.; Heinosalo, T.; West, G.; Saarinen, I.; Kaipio, K.; Rantala, J.; Albanese, C.; Poutanen, M.; et al. Personalized Drug Sensitivity Screening for Bladder Cancer Using Conditionally Reprogrammed Patient-Derived Cells. Eur. Urol. 2019, 76, 430–434. [Google Scholar] [CrossRef] [PubMed]
- Ge, Q.; Wang, P.; Wang, S.; Sood, A.; Meng, L.; Lee, C.; Parwani, A.V.; Li, J.; Liu, X. Urine PD-L1 as a Non-Invasive Biomarker for Immune Checkpoint Inhibitor (ICI) Therapy in Bladder Cancer. Adv. Biomark. Sci. Technol. 2025, 7, 172–179. [Google Scholar] [CrossRef]
- Liu, W.; Ju, L.; Cheng, S.; Wang, G.; Qian, K.; Liu, X.; Xiao, Y.; Wang, X. Conditional Reprogramming: Modeling Urological Cancer and Translation to Clinics. Clin. Transl. Med. 2020, 10, e95. [Google Scholar] [CrossRef]
- Simbulan-Rosenthal, C.M.; Dougherty, R.; Vakili, S.; Ferraro, A.M.; Kuo, L.W.; Alobaidi, R.; Aljehane, L.; Gaur, A.; Sykora, P.; Glasgow, E.; et al. CRISPR-Cas9 Knockdown and Induced Expression of CD133 Reveal Essential Roles in Melanoma Invasion and Metastasis. Cancers 2019, 11, 1490. [Google Scholar] [CrossRef]
- Xing, B.; Pu, C.; Chen, Y.; Sheng, Y.; Zhang, B.; Cui, J.; Wu, G.; Zhao, Y. Insights into the Characteristics of Primary Radioresistant Cervical Cancer Using Single-Cell Transcriptomics. Hum. Cell 2023, 36, 1135–1146. [Google Scholar] [CrossRef]
- Hynds, R.E.; Ben Aissa, A.; Gowers, K.H.C.; Watkins, T.B.K.; Bosshard-Carter, L.; Rowan, A.J.; Veeriah, S.; Wilson, G.A.; Quezada, S.A.; Swanton, C.; et al. Expansion of Airway Basal Epithelial Cells from Primary Human Non-Small Cell Lung Cancer Tumors. Int. J. Cancer 2018, 143, 160–166. [Google Scholar] [CrossRef]
- Wu, G.D.; Xiao, Y.G.; Fang, F.Y.; Yao, D.; Liu, J.; Cao, Y.H.; Mao, Y.; Yu, B.; Yao, T.R.; Wu, Y.M.; et al. Monitoring of Lung Malignant Epithelial Cells by Gene Methylation Analysis in the Conditionally Reprogrammed Cell Cultures. Neoplasma 2020, 67, 692–699. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Hong, G.; Chen, Y.; Ye, L.; Zhang, K.; Cai, K.; Yang, H.; Long, X.; Gao, W.; Li, H. Personalized Drug Testing in a Patient with Non-Small-Cell Lung Cancer Using Cultured Cancer Cells from Pleural Effusion. J. Int. Med. Res. 2020, 48, 300060520955058. [Google Scholar] [CrossRef] [PubMed]
- Kodack, D.P.; Farago, A.F.; Dastur, A.; Held, M.A.; Dardaei, L.; Friboulet, L.; von Flotow, F.; Damon, L.J.; Lee, D.; Parks, M.; et al. Primary Patient-Derived Cancer Cells and Their Potential for Personalized Cancer Patient Care. Cell Rep. 2017, 21, 3298–3309. [Google Scholar] [CrossRef]
- Zhao, Z.; Feng, X.; Wu, H.; Chen, S.; Ma, C.; Guan, Z.; Lei, L.; Tang, K.; Chen, X.; Dong, Y.; et al. Construction of a Lung Cancer 3D Culture Model Based on Alginate/Gelatin Micro-Beads for Drug Evaluation. Transl. Lung Cancer Res. 2024, 13, 2698–2712. [Google Scholar] [CrossRef] [PubMed]
- Sette, G.; Salvati, V.; Giordani, I.; Pilozzi, E.; Quacquarini, D.; Duranti, E.; De Nicola, F.; Pallocca, M.; Fanciulli, M.; Falchi, M.; et al. Conditionally Reprogrammed Cells (CRC) Methodology Does Not Allow the in Vitro Expansion of Patient-Derived Primary and Metastatic Lung Cancer Cells. Int. J. Cancer 2018, 143, 88–99. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Li, R.; An, T.; Liu, X. Conditional Cell Reprogramming in Modeling Digestive System Diseases. Front. Cell Dev. Biol. 2021, 9, 669756. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.S.; Kim, I.H.; Chae, H.D.; Kim, D.D.; Jeon, C.H. Detection of Circulating Gastrointestinal Cancer Cells in Conditionally Reprogrammed Cell Culture. In Vivo 2021, 35, 1515–1520. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Guo, D.; Zhang, Y.; Wang, L.; Sun, T.; Li, Z.; Zhang, X.; Wang, S.; Chen, Y.; Wu, A. Rapid Screening for Individualized Chemotherapy Optimization of Colorectal Cancer: A Novel Conditional Reprogramming Technology-Based Functional Diagnostic Assay. Transl. Oncol. 2021, 14, 100935. [Google Scholar] [CrossRef]
- Wang, Y.; Liao, H.; Zheng, T.; Wang, J.; Guo, D.; Lu, Z.; Li, Z.; Chen, Y.; Shen, L.; Zhang, Y.; et al. Conditionally Reprogrammed Colorectal Cancer Cells Combined with Mouse Avatars Identify Synergy between EGFR and MEK or CDK4/6 Inhibitors. Am. J. Cancer Res. 2020, 10, 249–262. [Google Scholar]
- Kim, B.K.; Nam, S.W.; Min, B.S.; Ban, H.S.; Paik, S.; Lee, K.; Im, J.Y.; Lee, Y.; Park, J.T.; Kim, S.Y.; et al. Bcl-2-Dependent Synthetic Lethal Interaction of the IDF-11774 with the V0 Subunit C of Vacuolar ATPase (ATP6V0C) in Colorectal Cancer. Br. J. Cancer 2018, 119, 1347–1357. [Google Scholar] [CrossRef]
- Parasido, E.; Avetian, G.S.; Naeem, A.; Graham, G.; Pishvaian, M.; Glasgow, E.; Mudambi, S.; Lee, Y.; Ihemelandu, C.; Choudhry, M.; et al. The Sustained Induction of C-MYC Drives Nab-Paclitaxel Resistance in Primary Pancreatic Ductal Carcinoma Cells. Mol. Cancer Res. 2019, 17, 1815–1827. [Google Scholar] [CrossRef]
- Beglyarova, N.; Banina, E.; Zhou, Y.; Mukhamadeeva, R.; Andrianov, G.; Bobrov, E.; Lysenko, E.; Skobeleva, N.; Gabitova, L.; Restifo, D.; et al. Screening of Conditionally Reprogrammed Patient-Derived Carcinoma Cells Identifies ERCC3-MYC Interactions as a Target in Pancreatic Cancer. Clin. Cancer Res. 2016, 22, 6153–6163. [Google Scholar] [CrossRef]
- Hollevoet, K.; Mason-Osann, E.; Liu, X.F.; Imhof-Jung, S.; Niederfellner, G.; Pastan, I. In Vitro and in Vivo Activity of the Low-Immunogenic Antimesothelin Immunotoxin RG7787 in Pancreatic Cancer. Mol. Cancer Ther. 2014, 13, 2040–2049. [Google Scholar] [CrossRef]
- Su, S.; Di Poto, C.; Roy, R.; Liu, X.; Cui, W.; Kroemer, A.; Ressom, H.W. Long-Term Culture and Characterization of Patient-Derived Primary Hepatocytes Using Conditional Reprogramming. Exp. Biol. Med. 2019, 244, 857–864. [Google Scholar] [CrossRef]
- Su, S.; Di Poto, C.; Kroemer, A.H.; Cui, W.; Roy, R.; Liu, X.; Ressom, H.W. Establishment of Ornithine Transcarbamylase Deficiency-Derived Primary Human Hepatocyte with Hepatic Functions. Exp. Cell Res. 2019, 384, 111621. [Google Scholar] [CrossRef]
- Wang, Z.; Bi, B.; Song, H.; Liu, L.; Zheng, H.; Wang, S.; Shen, Z. Proliferation of Human Hepatocellular Carcinoma Cells from Surgically Resected Specimens under Conditionally Reprogrammed Culture. Mol. Med. Rep. 2019, 19, 4623–4630. [Google Scholar] [CrossRef]
- Liu, X.; Mondal, A.M. Conditional Cell Reprogramming for Modeling Host-Virus Interactions and Human Viral Diseases. J. Med. Virol. 2020, 92, 2440–2452. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wu, Y.; Rong, L. Conditionally Reprogrammed Human Normal Airway Epithelial Cells at ALI: A Physiological Model for Emerging Viruses. Virol. Sin. 2020, 35, 280–289. [Google Scholar] [CrossRef]
- Matsumoto, N.; Ueha, S.; Ueha, R.; Koyama, M.; Yamakawa, K.; Sato, T.; Goto, T.; Kono, T.; Shichino, S.; Matsushima, K.; et al. Induced Gene Expression Signature in Recurrent Respiratory Papillomatosis. Laryngoscope 2025. [Google Scholar] [CrossRef]
- Rani, A.Q.; Nurmemet, D.; Liffick, J.; Khan, A.; Mitchell, D.; Li, J.; Zhao, B.; Liu, X. Conditional Cell Reprogramming and Air–Liquid Interface Modeling Life Cycle of Oncogenic Viruses (HPV and EBV) in Epithelial Cells and Virus-Associated Human Carcinomas. Viruses 2023, 15, 1388. [Google Scholar] [CrossRef] [PubMed]
- Roberts, N.; Al Mubarak, R.; Francisco, D.; Kraft, M.; Chu, H.W. Comparison of Paired Human Nasal and Bronchial Airway Epithelial Cell Responses to Rhinovirus Infection and IL-13 Treatment. Clin. Transl. Med. 2018, 7, 13. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Guthjahr, L.; Sauter, A.; Zech, F.; Nchioua, R.; Stenger, S.; Frick, M.; Kirchhoff, F.; Dietl, P.; Wittekindt, O.H. Serially Passaged, Conditionally Reprogrammed Nasal Epithelial Cells as a Model to Study Epithelial Functions and SARS-CoV-2 Infection. Am. J. Physiol. Cell Physiol. 2022, 322, C591–C604. [Google Scholar] [CrossRef]
- Xiao, Y.; Wang, L.; Li, S.; Fang, S.; Luo, F.; Chen, S.; Zou, X.; Ye, L.; Hou, W. Conditional Reprogrammed Human Limbal Epithelial Cell Model for Anti-SARS-CoV-2 Drug Screening. Heliyon 2024, 10, e30044. [Google Scholar] [CrossRef]
- Bukowy-Bieryłło, Z. Long-Term Differentiating Primary Human Airway Epithelial Cell Cultures: How Far Are We? Cell Communication and Signaling. Cell Commun. Signal. 2021, 19, 63. [Google Scholar] [CrossRef]
- Kaneko, Y.; Konno, T.; Kohno, T.; Kakuki, T.; Miyata, R.; Ohkuni, T.; Kakiuchi, A.; Yajima, R.; Ohwada, K.; Kurose, M.; et al. Induction of Airway Progenitor Cells via P63 and KLF11 by Rho-Kinase Inhibitor Y27632 in HTERT-Human Nasal Epithelial Cells. Am. J. Transl. Res. 2019, 11, 599–611. [Google Scholar]
- Kelly, N.A.; Shontz, K.M.; Bergman, M.; Manning, A.M.; Reynolds, S.D.; Chiang, T. Biobanked Tracheal Basal Cells Retain the Capacity to Differentiate. Laryngoscope Investig. Otolaryngol. 2022, 7, 2119–2125. [Google Scholar] [CrossRef]
- Kurokawa, A.; Kondo, M.; Honda, N.; Orimo, M.; Miyoshi, A.; Kobayashi, F.; Abe, K.; Akaba, T.; Tsuji, M.; Arimura, K.; et al. Analysis of the Diagnosis of Japanese Patients with Primary Ciliary Dyskinesia Using a Conditional Reprogramming Culture. Respir. Investig. 2022, 60, 407–417. [Google Scholar] [CrossRef]
- Peters-Hall, J.R.; Min, J.; Tedone, E.; Sho, S.; Siteni, S.; Mender, I.; Shay, J.W. Proliferation of Adult Human Bronchial Epithelial Cells without a Telomere Maintenance Mechanism for over 200 Population Doublings. FASEB J. 2020, 34, 386–398. [Google Scholar] [CrossRef]
- Peters-Hall, J.R.; Coquelin, M.L.; Torres, M.J.; Laranger, R.; Alabi, B.R.; Sho, S.; Calva-Moreno, J.F.; Thomas, P.J.; Shay, J.W. Long-Term Culture and Cloning of Primary Human Bronchial Basal Cells That Maintain Multipotent Differentiation Capacity and CFTR Channel Function. Am. J. Physiol. Lung Cell Mol. Physiol. 2018, 315, L313–L327. [Google Scholar] [CrossRef]
- Zhang, Z.; Bai, Q.; Chen, Y.; Ye, L.; Wu, X.; Long, X.; Ye, L.; Liu, J.; Li, H. Conditionally Reprogrammed Human Normal Bronchial Epithelial Cells Express Comparable Levels of Cytochromes P450 and Are Sensitive to BaP Induction. Biochem. Biophys. Res. Commun. 2018, 503, 2132–2138. [Google Scholar] [CrossRef] [PubMed]
- Veerati, P.C.; Nichol, K.S.; Read, J.M.; Bartlett, N.W.; Wark, P.A.B.; Knight, D.A.; Grainge, C.L.; Reid, A.T. Conditionally Reprogrammed Asthmatic Bronchial Epithelial Cells Express Lower FOXJ1 at Terminal Differentiation and Lower IFNs Following RV-A1 Infection. Am. J. Physiol. Lung Cell Mol. Physiol. 2022, 323, L495–L502. [Google Scholar] [CrossRef] [PubMed]
- Gentzsch, M.; Boyles, S.E.; Cheluvaraju, C.; Chaudhry, I.G.; Quinney, N.L.; Cho, C.; Dang, H.; Liu, X.; Schlegel, R.; Randell, S.H. Pharmacological Rescue of Conditionally Reprogrammed Cystic Fibrosis Bronchial Epithelial Cells. Am. J. Respir. Cell Mol. Biol. 2017, 56, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Martinovich, K.M.; Iosifidis, T.; Buckley, A.G.; Looi, K.; Ling, K.M.; Sutanto, E.N.; Kicic-Starcevich, E.; Garratt, L.W.; Shaw, N.C.; Montgomery, S.; et al. Conditionally Reprogrammed Primary Airway Epithelial Cells Maintain Morphology, Lineage and Disease Specific Functional Characteristics. Sci. Rep. 2017, 7, 17971. [Google Scholar] [CrossRef]
- Kabbesh, H.; Riaz, M.A.; Jensen, A.D.; Scheiner-Bobis, G.; Konrad, L. Long-Term Maintenance of Viable Adult Rat Sertoli Cells Able to Establish Testis Barrier Components and Function in Response to Androgens. Cells 2021, 10, 2405. [Google Scholar] [CrossRef]
- Park, D.; Reddy, A.P.; Wilmarth, P.A.; Jensen, J.T.; Han, L. Mucus Secretions from a Conditionally Reprogrammed Primary Endocervical Cell Culture. F S Sci. 2022, 3, 159–165. [Google Scholar] [CrossRef]
- Wu, M.; Zhang, X.; Kang, Y.; Zhu, Y.; Su, Z.; Liu, J.; Zhang, W.; Chen, H.; Li, H. The First Human Vulvar Intraepithelial Neoplasia Cell Line with Naturally Infected Episomal HPV18 Genome. Viruses 2022, 14, 2054. [Google Scholar] [CrossRef]
- Looney, R.J.; Roberts, M.; Markovetz, M.; Godiah, R.; Yao, S.; Golgotiu, K.; Wei, S.; Cellucci, C.; Han, L. In Vitro Inhibition of the CFTR Ion Channel in the Macaca mulatta Cervix Thickens Cervical Mucus. Biol. Reprod. 2025, ioaf103. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Wu, M.; Chen, S.; Zhang, T.; Ye, L.; Liu, J.; Li, H. Long Term Culture of Human Kidney Proximal Tubule Epithelial Cells Maintains Lineage Functions and Serves as an Ex Vivo Model for Coronavirus Associated Kidney Injury. Virol. Sin. 2020, 35, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Jensen, T.J.; Foster, C.; Sayej, W.; Finck, C.M. Conditional Reprogramming of Pediatric Human Esophageal Epithelial Cells for Use in Tissue Engineering and Disease Investigation. J. Vis. Exp. 2017, 121, 55243. [Google Scholar] [CrossRef]
- Liu, L.; Mondal, A.M.; Liu, X. Crosstalk of Moderate ROS and PARP-1 Contributes to Sustainable Proliferation of Conditionally Reprogrammed Keratinocytes. J. Biochem. Mol. Toxicol. 2023, 37, e23262. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Qu, Y.; Gomez, L.J.; Chung, S.; Han, B.; Gao, B.; Yue, Y.; Gong, Y.; Liu, X.; Amersi, F.; et al. Characterization of Primary Human Mammary Epithelial Cells Isolated and Propagated by Conditional Reprogrammed Cell Culture. Oncotarget 2018, 9, 11503–11514. [Google Scholar] [CrossRef]
- Sayej, W.N.; Foster, C.; Jensen, T.; Chatfield, S.; Finck, C. Expanding and Characterizing Esophageal Epithelial Cells Obtained from Children with Eosinophilic Esophagitis. Pediatr. Res. 2018, 84, 306–313. [Google Scholar] [CrossRef]
- Daneshdoust, D.; He, K.; Wang, Q.E.; Li, J.; Liu, X. Modeling Respiratory Tract Diseases for Clinical Translation Employing Conditionally Reprogrammed Cells. Cell Insight 2024, 3, 100201. [Google Scholar] [CrossRef]
- Katta, S.S.; Nagati, V.; Paturi, A.S.V.; Murakonda, S.P.; Murakonda, A.B.; Pandey, M.K.; Gupta, S.C.; Pasupulati, A.K.; Challagundla, K.B. Neuroblastoma: Emerging Trends in Pathogenesis, Diagnosis, and Therapeutic Targets. J. Control. Release 2023, 357, 444–459. [Google Scholar] [CrossRef]
- Ornell, K.J.; Coburn, J.M. Developing Preclinical Models of Neuroblastoma: Driving Therapeutic Testing. BMC Biomed. Eng. 2019, 1, 33. [Google Scholar] [CrossRef] [PubMed]
- Kolb, E.A.; Houghton, P.J.; Kurmasheva, R.T.; Mosse, Y.P.; Maris, J.M.; Erickson, S.W.; Guo, Y.; Teicher, B.A.; Smith, M.A.; Gorlick, R. Preclinical Evaluation of the Combination of AZD1775 and Irinotecan against Selected Pediatric Solid Tumors: A Pediatric Preclinical Testing Consortium Report. Pediatr. Blood Cancer 2020, 67, e28098. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; Zhou, D.; Chen, L.; Tian, Y.; Zhong, B.; Cao, Y.; Dong, Q.; Zhou, M.; Yan, J.; Wang, Y.; et al. Polo-like Kinase 4 Mediates Epithelial-Mesenchymal Transition in Neuroblastoma via PI3K/Akt Signaling Pathway Article. Cell Death Dis. 2018, 9, 54. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Krawczyk, E.; Lourdusamy, A.; Storer, L.C.; Xian, L.; Cohen, K.J.; Schlegel, R.; Grundy, R.; Resar, L. Abstract LB-224: A. Novel Model of Pediatric Spinal Ependymoma Using Conditionally Reprogrammed Cells from a Primary Tumor Demonstrates Aberrant Expression of HMGA, HOX, MYC and Other Wnt Target Genes. Cancer Res. 2017, 77 (Suppl. 13), LB-224. [Google Scholar] [CrossRef]
- Yuan, M.; White, D.; Resar, L.; Bar, E.; Groves, M.; Cohen, A.; Jackson, E.; Bynum, J.; Rubens, J.; Mumm, J.; et al. Conditional Reprogramming Culture Conditions Facilitate Growth of Lower-Grade Glioma Models. Neuro Oncol. 2021, 23, 770–782. [Google Scholar] [CrossRef]
- Nguyen, J.; Saffari, P.S.; Pollack, A.S.; Vennam, S.; Gong, X.; West, R.B.; Pollack, J.R. New Ameloblastoma Cell Lines Enable Preclinical Study of Targeted Therapies. J. Dent. Res. 2022, 101, 1517–1525. [Google Scholar] [CrossRef]
- Saffari, P.S.; Vapniarsky, N.; Pollack, A.S.; Gong, X.; Vennam, S.; Pollack, A.J.; Verstraete, F.J.M.; West, R.B.; Arzi, B.; Pollack, J.R. Most Canine Ameloblastomas Harbor HRAS Mutations, Providing a Novel Large-Animal Model of RAS-Driven Cancer. Oncogenesis 2019, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Panaccione, A.; Chang, M.T.; Carbone, B.E.; Guo, Y.; Moskaluk, C.A.; Virk, R.K.; Chiriboga, L.; Prasad, M.L.; Judson, B.; Mehra, S.; et al. NOTCH1 and SOX10 Are Essential for Proliferation and Radiation Resistance of Cancer Stem-like Cells in Adenoid Cystic Carcinoma. Clin. Cancer Res. 2016, 22, 2083–2095. [Google Scholar] [CrossRef]
- Ci, X.; Hao, J.; Dong, X.; Xue, H.; Wu, R.; Choi, S.Y.C.; Haegert, A.M.; Collins, C.C.; Liu, X.; Lin, D.; et al. Conditionally Reprogrammed Cells from Patient-Derived Xenograft to Model Neuroendocrine Prostate Cancer Development. Cells 2020, 9, 1398. [Google Scholar] [CrossRef]
- Dong, Y.; Wang, J.; Ji, W.; Zheng, M.; Wang, P.; Liu, L.; Li, S. Preclinical Application of Conditional Reprogramming Culture System for Laryngeal and Hypopharyngeal Carcinoma. Front. Cell Dev. Biol. 2021, 9, 744969. [Google Scholar] [CrossRef] [PubMed]
- Grossmann, L.D.; Chen, C.-H.; Uzun, Y.; Thadi, A.; Wolpaw, A.J.; Louault, K.; Goldstein, Y.; Surrey, L.F.; Martinez, D.; Calafatti, M.; et al. Identification and Characterization of Chemotherapy Resistant High-Risk Neuroblastoma Persister Cells. Cancer Discov. 2024, 12, 2387–2406. [Google Scholar] [CrossRef] [PubMed]
- Wolf, A.B.; Reynolds, C.P.; Barbieri, E. Characterization of Persister Cells Provides Insights into Mechanisms of Therapy Resistance in Neuroblastoma. Cancer Discov. 2024, 14, 2308–2311. [Google Scholar] [CrossRef]
- Polychronopoulos, P.A.; Bedoya-Reina, O.C.; Johnsen, J.I. The Neuroblastoma Microenvironment, Heterogeneity and Immunotherapeutic Approaches. Cancers 2024, 16, 1863. [Google Scholar] [CrossRef]
- Yu, W.; Biyik-Sit, R.; Uzun, Y.; Chen, C.-H.; Thadi, A.; Sussman, J.H.; Pang, M.; Wu, C.-Y.; Grossmann, L.D.; Gao, P.; et al. Longitudinal Single-Cell Multiomic Atlas of High-Risk Neuroblastoma Reveals Chemotherapy-Induced Tumor Microenvironment Rewiring. Nat. Genet. 2025, 1–13. [Google Scholar] [CrossRef]
- Amoli, M.S.; Rezapourdamanab, S.; Jin, L.; Cadena, M.A.; Kaw, K.; Sridhar, V.; Meselhe, M.; Krikor, S.; Mahmoudi, M.; Ning, L.; et al. Protocol for 3D Bioprinting of a 3D in Vitro Model of Neuroblastoma. STAR Protoc. 2025, 6, 103725. [Google Scholar] [CrossRef]
- Yamada, Y.; Beltran, H. Clinical and Biological Features of Neuroendocrine Prostate Cancer. Curr. Oncol. Rep. 2021, 23, 15. [Google Scholar] [CrossRef]
- Shaheen, S.; Guddati, A.K. Gastrointestinal Stromal Tumor: A Rare Abdominal Tumor. Case Rep. Oncol. 2013, 6, 148–153. [Google Scholar] [CrossRef]
- Rheinwald, J.G.; Green, H. Formation of a Keratinizing Epithelium in Culture by a Cloned Cell Line Derived from a Teratoma. Cell 1975, 6, 317–330. [Google Scholar] [CrossRef] [PubMed]
- Kresbach, C.; Neyazi, S.; Schüller, U. Updates in the Classification of Ependymal Neoplasms: The 2021 WHO Classification and Beyond. Brain Pathol. 2022, 32, e13068. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Ge, P. Long Non-Coding RNA HCG18 Facilitates the Progression of Laryngeal and Hypopharyngeal Squamous Cell Carcinoma by Upregulating FGFR1 via MiR-133b. Mol. Med. Rep. 2022, 25, 46. [Google Scholar] [CrossRef] [PubMed]

| Transformed Cell Lines | iPSc * | CR Cells ** | |
|---|---|---|---|
| Success rate | Medium | Medium | High |
| Lifespan | Long | Long | Long |
| Timing | 1–2 months | 2–10 weeks | 1–10 days |
| Expansion | High | Medium | High |
| Genetic stability | Low | Medium | High |
| Tissue specificity | Low | Low | High |
| Heterogeneity | No | Medium | Medium |
| Differentiation | Low | Medium | High |
| High-throughput drug screening applicability | High | Low | High |
| Low-throughput drug screening applicability | High | High | High |
| Cost | Low | Medium | Low |
| Tumor | Standard CR Technology | CM Medium | ACC ** Medium | 2% O2 | Collagen Coating | Number of Generated CR Cell Lines | Refs | |||
|---|---|---|---|---|---|---|---|---|---|---|
| CR Medium | Feeder Cells | Y-27632 | ||||||||
| Neuroblastoma (murine) | + | + | + | 21 | [59,82,83] | |||||
| Neuroendocrine cervical cancer | + | + | + | 1 | [60] | |||||
| Ependymoma | + | + | + | 1 | [155] | |||||
| + | + | + | 2 | [156] | ||||||
| Pilocytic astrocytoma | + | + | + | 3 | [156] | |||||
| Pleomorphic xanthoastrocytoma | + | + | + | 1 | [156] | |||||
| Other lower-grade gliomas | + | + | + | 8 (total) | [156] | |||||
| GIST | + | + | + | 1 | [39] | |||||
| Ameloblastoma | human * | + | + | + | 6 | [157] | ||||
| canine | + | + | + | 4 | [158] | |||||
| ACC | [159] | + | + | + | 6 (total) | [159] | ||||
| [84] | + | + | + | + | 2 | [84] | ||||
| Neuroendocrine prostate cancer | + | + | + | 1 | [160] | |||||
| Laryngeal and hypopharyngeal carcinoma | + | + | + | 28 | [161] | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Krawczyk, E. Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers. Cancers 2025, 17, 2834. https://doi.org/10.3390/cancers17172834
Krawczyk E. Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers. Cancers. 2025; 17(17):2834. https://doi.org/10.3390/cancers17172834
Chicago/Turabian StyleKrawczyk, Ewa. 2025. "Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers" Cancers 17, no. 17: 2834. https://doi.org/10.3390/cancers17172834
APA StyleKrawczyk, E. (2025). Conditionally Reprogrammed Cells as Preclinical Model for Rare Cancers. Cancers, 17(17), 2834. https://doi.org/10.3390/cancers17172834






