KRAS Mutations in Colorectal Adenocarcinoma: Incidence and Association with Histological Features with Particular Reference to Gly12Asp in a Multicenter GIPAD Real-World Study
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinicopathologic Data
2.2. Pathologic Features
2.3. MMR/MSI Status
2.4. KRAS Mutational Analysis
- Center 1: Myriapod NGS Cancer Panel (Diatech Pharmacogenetics, Jesi, Italy), sequencing on Illumina platform.
- Center 2: EasyPGX ready KRAS (Diatech Pharmacogenetics).
- Center 3: Oncomine DX Target Test (Thermo Fisher Scientific, Waltham, MA, USA), sequencing on Ion Torrent platform.
- Center 4: Oncomine Focus Assay (Thermo Fisher Scientific), sequencing on Ion Torrent platform; Myriapod NGS Cancer Panel (Diatech Pharmacogenetics).
- Center 5: Myriapod Colon Panel (Diatech Pharmacogenetics), sequencing on Ion Torrent platform; Myriapod Colon Status Kit (Diatech Pharmacogenetics) on the MassARRAY system (Sequenom, San Diego, CA, USA).
- Center 6, Center 7: Myriapod NGS Cancer Panel (Diatech Pharmacogenetics), sequencing on Illumina; EasyPGX ready KRAS (Diatech Pharmacogenetics).
- Center 8: Oncomine Focus Assay (Thermo Fisher Scientific), sequencing on Ion Torrent platform; EasyPGX ready KRAS (Diatech Pharmacogenetics).
- Center 9: RAS Mutation Screening Panel (EntroGen, Woodland Hills, CA, USA); EasyPGX ready KRAS (Diatech Pharmacogenetics).
- Centers 10 and 11: Myriapod NGS Cancer Panel DNA (Diatech Pharmacogenetics), sequencing on Illumina platform (San Diego, CA, USA); Idylla KRAS Mutation Assay (Biocartis, Mechelen, Belgium).
- Center 12: Idylla KRAS Mutation Test (Biocartis).
2.5. Statistical Analysis
3. Results
3.1. Patient Cohort
3.2. Clinicopathological Features of KRAS-Mutated CRC
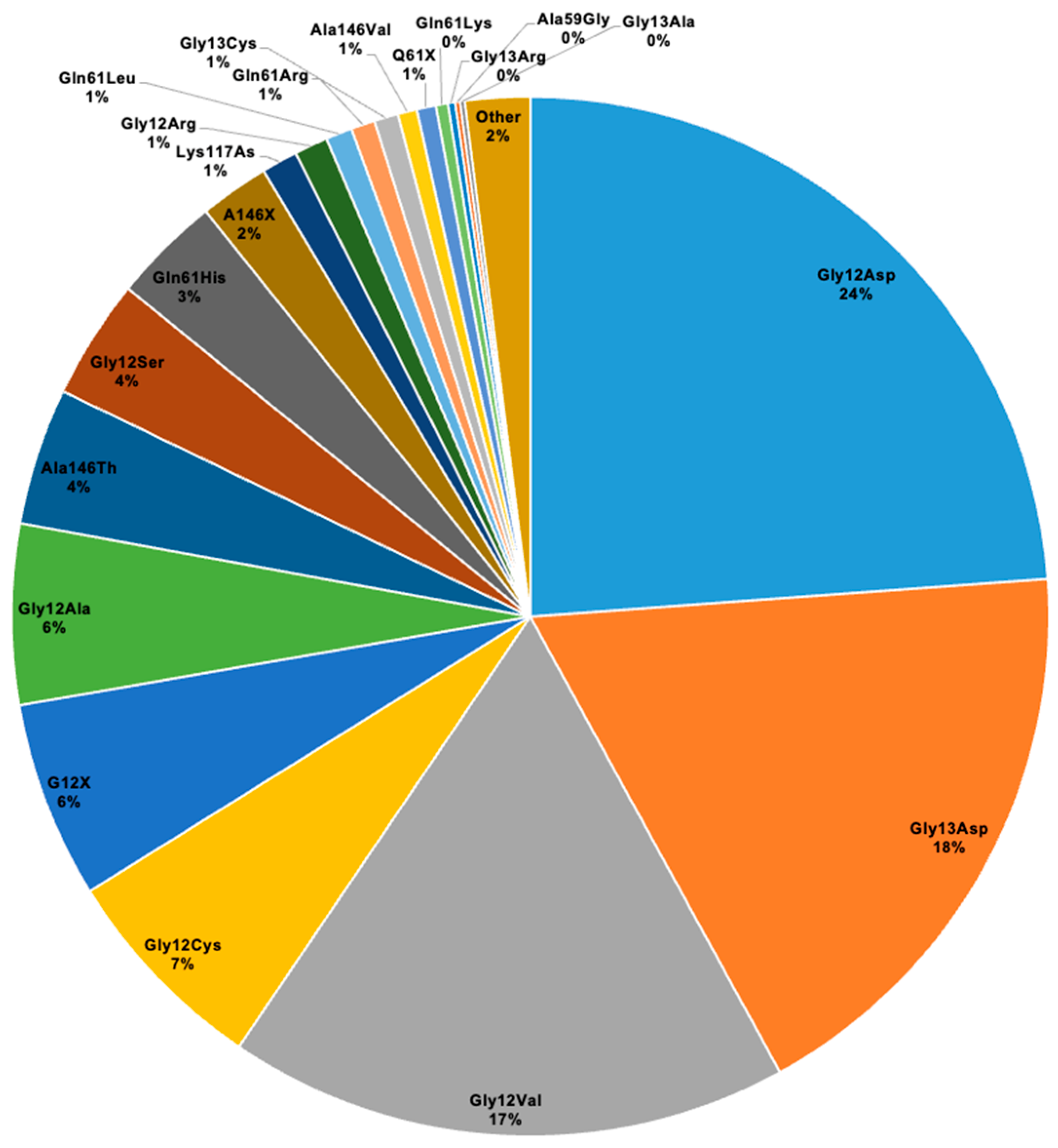
3.3. KRAS Mutational Landscape
3.4. Clinicopathologic Features of Gly12Asp-Mutated CRC
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CRC | Colorectal cancer |
| iMMR | Indeterminate MMR |
| mKRAS | Mutated KRAS |
| MMR | Mismatch repair |
| MMRd | Mismatch repair deficiency |
| MMRp | Mismatch repair proficiency |
| MS | Microsatellite assay |
| MSI | Microsatellite instability |
| MSS | Microsatellite stability |
| NGS | Next-generation sequencing |
| NOS | Not otherwise specified |
| OS | Overall survival |
| PCR | Polymerase reaction chain |
| wtKRAS | Wild-type KRAS |
References
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. [Google Scholar] [CrossRef]
- Siegel, R.L.; Wagle, N.S.; Cercek, A.; Smith, R.A.; Jemal, A. Colorectal cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 233–254. [Google Scholar] [CrossRef]
- Takeda, M.; Yoshida, S.; Inoue, T.; Sekido, Y.; Hata, T.; Hamabe, A.; Ogino, T.; Miyoshi, N.; Uemura, M.; Yamamoto, H.; et al. The Role of KRAS Mutations in Colorectal Cancer: Biological Insights, Clinical Implications, and Future Therapeutic Perspectives. Cancers 2025, 17, 428. [Google Scholar] [CrossRef]
- Eng, C.; Yoshino, T.; Ruíz-García, E.; Mostafa, N.; Cann, C.G.; O’Brian, B.; Benny, A.; Perez, R.O.; Cremolini, C. Colorectal cancer. Lancet 2024, 404, 294–310. [Google Scholar] [CrossRef] [PubMed]
- Nagtegaal, I.D.; Odze, R.D.; Klimstra, D.; Paradis, V.; Rugge, M.; Schirmacher, P.; Washington, K.M.; Carneiro, F.; Cree, I.A. The 2019 WHO classification of tumours of the digestive system. Histopathology 2020, 76, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Remo, A.; Fassan, M.; Vanoli, A.; Bonetti, L.R.; Barresi, V.; Tatangelo, F.; Gafà, R.; Giordano, G.; Pancione, M.; Grillo, F.; et al. Morphology and Molecular Features of Rare Colorectal Carcinoma Histotypes. Cancers 2019, 11, 1036. [Google Scholar] [CrossRef]
- Adorisio, R.; Ciardiello, D.; Rappa, A.; Gervaso, L.; Pelizzari, G.; Marinucci, L.; Fusco, N.; Zampino, M.G.; Fazio, N.; Venetis, K.; et al. Investigating the Pathogenicity of Uncommon KRAS Mutations and Their Association with Clinicopathologic Characteristics in Patients with Colorectal Cancer. J. Mol. Diagn. 2025, 27, 130–138. [Google Scholar] [CrossRef]
- Malapelle, U.; Angerilli, V.; Pepe, F.; Fontanini, G.; Lonardi, S.; Scartozzi, M.; Memeo, L.; Pruneri, G.; Marchetti, A.; Perrone, G.; et al. The ideal reporting of RAS testing in colorectal adenocarcinoma: A pathologists’ perspective. Pathologica 2023, 115, 137–147. [Google Scholar] [CrossRef]
- Malapelle, U.; Passiglia, F.; Cremolini, C.; Reale, M.L.; Pepe, F.; Pisapia, P.; Avallone, A.; Cortinovis, D.; De Stefano, A.; Fassan, M.; et al. RAS as a positive predictive biomarker: Focus on lung and colorectal cancer patients. Eur. J. Cancer 2021, 146, 74–83. [Google Scholar] [CrossRef]
- Lu, X.; Li, Y.; Zhang, X.; Shi, J.; Feng, H.; Yu, Z.; Gao, Y. Prognostic and predictive biomarkers for anti-EGFR monoclonal antibody therapy in RAS wild-type metastatic colorectal cancer: A systematic review and meta-analysis. BMC Cancer 2023, 23, 1117. [Google Scholar] [CrossRef] [PubMed]
- Schirripa, M.; Nappo, F.; Cremolini, C.; Salvatore, L.; Rossini, D.; Bensi, M.; Businello, G.; Pietrantonio, F.; Randon, G.; Fucà, G.; et al. KRAS G12C Metastatic Colorectal Cancer: Specific Features of a New Emerging Target Population. Clin. Colorectal. Cancer 2020, 19, 219–225. [Google Scholar] [CrossRef]
- Morris, V.K.; Kennedy, E.B.; Baxter, N.N.; Benson, A.B., 3rd; Cercek, A.; Cho, M.; Ciombor, K.K.; Cremolini, C.; Davis, A.; Deming, D.A.; et al. Treatment of Metastatic Colorectal Cancer: ASCO Guideline. J. Clin. Oncol. 2023, 41, 678–700. [Google Scholar] [CrossRef]
- Van Cutsem, E.; Cervantes, A.; Adam, R.; Sobrero, A.; Van Krieken, J.H.; Aderka, D.; Aranda Aguilar, E.; Bardelli, A.; Benson, A.; Bodoky, G.; et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann. Oncol. 2016, 27, 1386–1422. [Google Scholar] [CrossRef]
- Andre, T.; Shiu, K.K.; Kim, T.W.; Jensen, B.V.; Jensen, L.H.; Punt, C.; Smith, D.; Garcia-Carbonero, R.; Benavides, M.; Gibbs, P.; et al. Pembrolizumab in Microsatellite-Instability-High Advanced Colorectal Cancer. N. Engl. J. Med. 2020, 383, 2207–2218. [Google Scholar] [CrossRef]
- Ciardiello, D.; Maiorano, B.A.; Martinelli, E. Targeting KRAS (G12C) in colorectal cancer: The beginning of a new era. ESMO Open 2023, 8, 100745. [Google Scholar] [CrossRef]
- Di Nicolantonio, F.; Bardelli, A. Precision oncology for KRAS (G12C)-mutant colorectal cancer. Nat. Rev. Clin. Oncol. 2023, 20, 355–356. [Google Scholar] [CrossRef] [PubMed]
- Basso, M.; Signorelli, C.; Calegari, M.A.; Lucchetti, J.; Zurlo, I.V.; Dell’Aquila, E.; Arrivi, G.; Zoratto, F.; Santamaria, F.; Saltarelli, R.; et al. Efficacy of Regorafenib and Trifluridine/Tipiracil According to Extended RAS Evaluation in Advanced Metastatic Colorectal Cancer Patients: A Multicenter Retrospective Analysis. Target. Oncol. 2024, 19, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Lugli, A.; Kirsch, R.; Ajioka, Y.; Bosman, F.; Cathomas, G.; Dawson, H.; El Zimaity, H.; Fléjou, J.F.; Hansen, T.P.; Hartmann, A.; et al. Recommendations for reporting tumor budding in colorectal cancer based on the International Tumor Budding Consensus Conference (ITBCC) 2016. Mod. Pathol. 2017, 30, 1299–1311. [Google Scholar] [CrossRef]
- Loughrey, M.B.; Webster, F.; Arends, M.J.; Brown, I.; Burgart, L.J.; Cunningham, C.; Flejou, J.F.; Kakar, S.; Kirsch, R.; Kojima, M.; et al. Dataset for Pathology Reporting of Colorectal Cancer: Recommendations From the International Collaboration on Cancer Reporting (ICCR). Ann. Surg. 2022, 275, e549–e561. [Google Scholar] [CrossRef] [PubMed]
- Parente, P.; Grillo, F.; Vanoli, A.; Macciomei, M.C.; Ambrosio, M.R.; Scibetta, N.; Filippi, E.; Cataldo, I.; Baron, L.; Ingravallo, G.; et al. The Day-To-Day Practice of MMR and MSI Assessment in Colorectal Adenocarcinoma: What We Know and What We Still Need to Explore. Dig. Dis. 2023, 41, 746–756. [Google Scholar] [CrossRef]
- Dienstmann, R.; Connor, K.; Byrne, A.T.; Consortium, C. Precision Therapy in RAS Mutant Colorectal Cancer. Gastroenterology 2020, 158, 806–811. [Google Scholar] [CrossRef]
- Rebersek, M.; Boc, M.; Cerkovnik, P.; Benedik, J.; Hlebanja, Z.; Volk, N.; Novakovic, S.; Ocvirk, J. Efficacy of first-line systemic treatment in correlation with BRAF V600E and different KRAS mutations in metastatic colorectal cancer—A single institution retrospective analysis. Radiol. Oncol. 2011, 45, 285–291. [Google Scholar] [CrossRef]
- Hayama, T.; Hashiguchi, Y.; Okamoto, K.; Okada, Y.; Ono, K.; Shimada, R.; Ozawa, T.; Toyoda, T.; Tsuchiya, T.; Iinuma, H.; et al. G12V and G12C mutations in the gene KRAS are associated with a poorer prognosis in primary colorectal cancer. Int. J. Colorectal Dis. 2019, 34, 1491–1496. [Google Scholar] [CrossRef] [PubMed]
- Koulouridi, A.; Karagianni, M.; Messaritakis, I.; Sfakianaki, M.; Voutsina, A.; Trypaki, M.; Bachlitzanaki, M.; Koustas, E.; Karamouzis, M.V.; Ntavatzikos, A.; et al. Prognostic Value of KRAS Mutations in Colorectal Cancer Patients. Cancers 2022, 14, 3320. [Google Scholar] [CrossRef]
- Strickler, J.H.; Yoshino, T.; Stevinson, K.; Eichinger, C.S.; Giannopoulou, C.; Rehn, M.; Modest, D.P. Prevalence of KRAS G12C Mutation and Co-mutations and Associated Clinical Outcomes in Patients with Colorectal Cancer: A Systematic Literature Review. Oncologist 2023, 28, e981–e994. [Google Scholar] [CrossRef] [PubMed]
- Rosty, C.; Young, J.P.; Walsh, M.D.; Clendenning, M.; Walters, R.J.; Pearson, S.; Pavluk, E.; Nagler, B.; Pakenas, D.; Jass, J.R.; et al. Colorectal carcinomas with KRAS mutation are associated with distinctive morphological and molecular features. Mod. Pathol. 2013, 26, 825–834. [Google Scholar] [CrossRef] [PubMed]
- Zeissig, M.N.; Ashwood, L.M.; Kondrashova, O.; Sutherland, K.D. Next batter up! Targeting cancers with KRAS-G12D mutations. Trends Cancer 2023, 9, 955–967. [Google Scholar] [CrossRef]
- Fassan, M.; Scarpa, A.; Remo, A.; De Maglio, G.; Troncone, G.; Marchetti, A.; Doglioni, C.; Ingravallo, G.; Perrone, G.; Parente, P.; et al. Current prognostic and predictive biomarkers for gastrointestinal tumors in clinical practice. Pathologica 2020, 112, 248–259. [Google Scholar] [CrossRef]
- Fusco, N.; Pruneri, G.; Pagni, F.; Malapelle, U. Molecular Testing in Solid Tumors: Best Practices from the Molecular Pathology and Precision Medicine Study Group of the Italian Society of Pathology (PMMP/SIAPeC): Shaping Excellence in Molecular Diagnostics. Pathologica 2025. Available online: https://www.pathologica.it/article/view/1214 (accessed on 6 August 2025).
- Dawson, H.; Kirsch, R.; Messenger, D.; Driman, D. A Review of Current Challenges in Colorectal Cancer Reporting. Arch. Pathol. Lab. Med. 2019, 143, 869–882. [Google Scholar] [CrossRef]
| KRAS mut (n = 1334; 47.4%) | KRAS wt (n = 1482; 52.6%) | P (Chi-Square) | ||||
|---|---|---|---|---|---|---|
| Center | Center 1 | 201 | 15.1% | 356 | 24.0% | <0.001 |
| Center 2 | 175 | 13.1% | 187 | 12.6% | ||
| Center 3 | 145 | 10.9% | 163 | 11.0% | ||
| Center 4 | 121 | 9.1% | 170 | 11.5% | ||
| Center 5 | 122 | 9.1% | 108 | 7.3% | ||
| Center 6 | 105 | 7.9% | 93 | 6.3% | ||
| Center 7 | 106 | 7.9% | 84 | 5.7% | ||
| Center 8 | 98 | 7.3% | 90 | 6.1% | ||
| Center 9 | 85 | 6.4% | 100 | 6.7% | ||
| Center 10 | 67 | 5.0% | 56 | 3.8% | ||
| Center 11 | 67 | 5.0% | 41 | 2.8% | ||
| Center 12 | 42 | 3.1% | 34 | 2.3% | ||
| Assay | NGS-based assay | 783 | 58.7% | 939 | 63.3% | 0.011 |
| PCR-based assay/MALDI-TOF | 551 | 41.3% | 543 | 36.7% | ||
| Age | >70 years | 669 | 50.1% | 803 | 54.2% | 0.031 |
| ≤70 years | 665 | 49.9% | 678 | 45.8% | ||
| Sex | M | 781 | 58.5% | 846 | 57.2% | 0.458 |
| F | 553 | 41.5% | 634 | 42.8% | ||
| Specimen | Biopsy | 363 | 27.2% | 329 | 22.2% | 0.002 |
| Surgical | 971 | 72.8% | 1153 | 77.8% | ||
| Site | Proximal colon | 522 | 41.0% | 489 | 34.3% | 0.002 |
| Distal colon | 215 | 16.8% | 268 | 18.8% | ||
| Rectosigmoid | 539 | 42.2% | 669 | 46.9% | ||
| Histotype | NOS | 1096 | 87.7% | 1226 | 86.8% | 0.02 |
| Mucinous | 125 | 10.0% | 122 | 8.6% | ||
| Signet Ring cell | 9 | 0.7% | 19 | 1.3% | ||
| Medullary | 3 | 0.2% | 17 | 1.2% | ||
| Micropapillary | 6 | 0.5% | 14 | 1.0% | ||
| Adenosquamous | 8 | 0.6% | 4 | 0.3% | ||
| Undifferentiated | 0 | 0.0% | 8 | 0.6% | ||
| Serrated | 3 | 0.2% | 2 | 0.1% | ||
| Grading | High | 352 | 31.1% | 514 | 40.6% | <0.001 |
| Low | 781 | 68.9% | 752 | 59.4% | ||
| Budding | Bd1 | 203 | 33.3% | 325 | 41.6% | <0.001 |
| Bd2 | 168 | 27.6% | 230 | 29.4% | ||
| Bd3 | 238 | 39.1% | 226 | 28.9% | ||
| Lymphovascular invasion | No | 293 | 31.3% | 349 | 30.8% | 0.830 |
| Yes | 644 | 68.7% | 783 | 69.2% | ||
| Perineural invasion | No | 502 | 55.1% | 564 | 51.2% | 0.079 |
| Yes | 409 | 44.9% | 538 | 48.8% | ||
| pT | pT1 | 6 | 0.8% | 10 | 1.2% | 0.388 |
| pT2 | 38 | 5.0% | 54 | 6.7% | ||
| pT3 | 361 | 47.4% | 382 | 47.3% | ||
| pT4 | 356 | 46.8% | 362 | 44.8% | ||
| pN | pN0 | 213 | 28.2% | 233 | 29.1% | 0.013 |
| pN1 | 321 | 42.6% | 288 | 35.9% | ||
| pN2 | 220 | 29.2% | 281 | 35.0% | ||
| MMR/MS status | MMRd/MSI | 93 | 7.7% | 343 | 26.2% | <0.001 |
| MMRp/MSS | 1116 | 92.3% | 966 | 73.8% | ||
| Gly12Asp | Gly13Asp | Gly12Val | Gly12Cys | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gly12Asp N = 319 (25.5%) | KRAS non-Gly12Asp mut * N = 934 (74.5%) | Gly13Asp N = 243 (18.2%) | KRAS non-Gly13Asp mut N = 1091 (81.8%) | Gly12Val N = 234 (17.5%) | KRAS non- Gly12Val mut * N = 1019 (82.5%) | Gly12Cys N = 89 (6.7%) | KRAS non-Gly12Cys mut N = 1245 (93.3%) | ||||||||||
| A Center | Center 1 | 59 | 18.5% | 142 | 15.2% | 45 | 18.5% | 156 | 14.3% | 30 | 12.8% | 171 | 16.8% | 14 | 15.7% | 187 | 15.0% |
| Center 2 | 29 | 9.1% | 116 | 12.4% | 33 | 13.6% | 112 | 10.3% | 23 | 9.8% | 122 | 12.0% | 9 | 10.1% | 136 | 10.9% | |
| Center 3 | 50 | 15.7% | 123 | 13.2% | 27 | 11.1% | 148 | 13.6% | 42 | 17.9% | 131 | 12.9% | 16 | 18.0% | 159 | 12.8% | |
| Center 4 | 35 | 11.0% | 86 | 9.2% | 20 | 8.2% | 101 | 9.3% | 28 | 12.0% | 93 | 9.1% | 9 | 10.1% | 112 | 9.0% | |
| Center 5 | 30 | 9.4% | 92 | 9.9% | 16 | 6.6% | 106 | 9.7% | 25 | 10.7% | 97 | 9.5% | 6 | 6.7% | 116 | 9.3% | |
| Center 6 | 27 | 8.5% | 78 | 8.4% | 15 | 6.2% | 90 | 8.2% | 19 | 8.1% | 86 | 8.4% | 6 | 6.7% | 99 | 8.0% | |
| Center 7 | 11 | 3.4% | 51 | 5.5% | 20 | 8.2% | 86 | 7.9% | 4 | 1.7% | 58 | 5.7% | 5 | 5.6% | 101 | 8.1% | |
| Center 8 | 9 | 2.8% | 54 | 5.8% | 18 | 7.4% | 80 | 7.3% | 5 | 2.1% | 58 | 5.7% | 5 | 5.6% | 93 | 7.5% | |
| Center 9 | 25 | 7.8% | 60 | 6.4% | 22 | 9.1% | 63 | 5.8% | 13 | 5.6% | 72 | 7.1% | 3 | 3.4% | 82 | 6.6% | |
| Center 10 | 12 | 3.8% | 55 | 5.9% | 10 | 4.1% | 57 | 5.2% | 17 | 7.3% | 50 | 4.9% | 10 | 11.2% | 57 | 4.6% | |
| Center 11 | 21 | 6.6% | 46 | 4.9% | 9 | 3.7% | 58 | 5.3% | 16 | 6.8% | 51 | 5.0% | 3 | 3.4% | 64 | 5.1% | |
| Center 12 | 11 | 3.4% | 31 | 3.3% | 8 | 3.3% | 34 | 3.1% | 12 | 5.1% | 30 | 2.9% | 3 | 3.4% | 39 | 3.1% | |
| p-value | 0.148 | 0.253 | 0.008 | 0.299 | |||||||||||||
| B Assay | NGS-based assay | 197 | 61.8% | 584 | 62.5% | 145 | 59.7% | 638 | 58.5% | 144 | 61.5% | 636 | 62.4% | 55 | 61.8% | 728 | 58.5% |
| PCR-based assay/MALDI-TOF | 122 | 38.2% | 350 | 37.5% | 98 | 40.3% | 453 | 41.5% | 90 | 38.5% | 383 | 37.6% | 34 | 38.2% | 517 | 41.5% | |
| p-value | 0.806 | 0.732 | 0.803 | 0.538 | |||||||||||||
| C Site | Distal Colon | 129 | 41.9% | 393 | 40.4% | 112 | 48.5% | 410 | 39.0% | 74 | 32.7% | 448 | 42.5% | 33 | 37.5% | 489 | 41.0% |
| Proximal Colon | 35 | 11.4% | 187 | 19.2% | 34 | 14.7% | 188 | 17.9% | 47 | 20.8% | 175 | 16.6% | 18 | 20.5% | 204 | 17.1% | |
| Rectosigmoid Colon | 144 | 46.8% | 393 | 40.4% | 85 | 36.8% | 452 | 43.0% | 105 | 46.5% | 432 | 40.9% | 37 | 42.0% | 500 | 41.9% | |
| p-value | 0.005 | 0.03 | 0.023 | 0.678 | |||||||||||||
| D Histotype | NOS | 264 | 87.1% | 773 | 88.3% | 201 | 87.0% | 895 | 87.8% | 196 | 90.3% | 841 | 87.5% | 72 | 87.8% | 1024 | 87.7% |
| Mucinous | 34 | 11.2% | 80 | 9.1% | 22 | 9.5% | 103 | 10.1% | 19 | 8.8% | 95 | 9.9% | 7 | 8.5% | 118 | 10.1% | |
| Signet Ring cell | 2 | 0.7% | 6 | 0.7% | 0 | 0.0% | 9 | 0.9% | 1 | 0.5% | 7 | 0.7% | 3 | 3.7% | 6 | 0.5% | |
| Medullary | 0 | 0.0% | 2 | 0.2% | 0 | 0.0% | 3 | 0.3% | 0 | 0.0% | 2 | 0.2% | 0 | 0.0% | 3 | 0.3% | |
| Micropapillary | 1 | 0.3% | 5 | 0.6% | 2 | 0.9% | 4 | 0.4% | 0 | 0.0% | 6 | 0.6% | 0 | 0.0% | 6 | 0.5% | |
| Adenosquamous | 0 | 0.0% | 8 | 0.9% | 5 | 2.2% | 3 | 0.3% | 1 | 0.5% | 7 | 0.7% | 0 | 0.0% | 8 | 0.7% | |
| Undifferentiated | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | |
| Serrated | 2 | 0.7% | 1 | 0.1% | 1 | 0.4% | 2 | 0.2% | 0 | 0.0% | 3 | 0.3% | 0 | 0.0% | 3 | 0.3% | |
| p-value | 0.294 | 0.037 | 0.965 | 0.205 | |||||||||||||
| E Grading | High | 70 | 26.1% | 235 | 29.8% | 77 | 37.2% | 275 | 29.7% | 46 | 24.2% | 259 | 29.9% | 18 | 25.4% | 334 | 31.5% |
| Low | 198 | 73.9% | 553 | 70.2% | 130 | 62.8% | 651 | 70.3% | 144 | 75.8% | 607 | 70.1% | 53 | 74.6% | 728 | 68.5% | |
| p-value | 0.248 | 0.035 | 0.117 | 0.282 | |||||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parente, P.; Angerilli, V.; Grillo, F.; Ambrosio, M.R.; Petrelli, F.; Gasparello, J.; Antoci, F.; Pilozzi, E.; Scarpino, S.; Adotti, F.; et al. KRAS Mutations in Colorectal Adenocarcinoma: Incidence and Association with Histological Features with Particular Reference to Gly12Asp in a Multicenter GIPAD Real-World Study. Cancers 2025, 17, 2721. https://doi.org/10.3390/cancers17172721
Parente P, Angerilli V, Grillo F, Ambrosio MR, Petrelli F, Gasparello J, Antoci F, Pilozzi E, Scarpino S, Adotti F, et al. KRAS Mutations in Colorectal Adenocarcinoma: Incidence and Association with Histological Features with Particular Reference to Gly12Asp in a Multicenter GIPAD Real-World Study. Cancers. 2025; 17(17):2721. https://doi.org/10.3390/cancers17172721
Chicago/Turabian StyleParente, Paola, Valentina Angerilli, Federica Grillo, Maria Raffaella Ambrosio, Federica Petrelli, Jessica Gasparello, Francesca Antoci, Emanuela Pilozzi, Stefania Scarpino, Flavia Adotti, and et al. 2025. "KRAS Mutations in Colorectal Adenocarcinoma: Incidence and Association with Histological Features with Particular Reference to Gly12Asp in a Multicenter GIPAD Real-World Study" Cancers 17, no. 17: 2721. https://doi.org/10.3390/cancers17172721
APA StyleParente, P., Angerilli, V., Grillo, F., Ambrosio, M. R., Petrelli, F., Gasparello, J., Antoci, F., Pilozzi, E., Scarpino, S., Adotti, F., Ascione, A., Veccia, N., Caputo, A., Giobbe, M., Gafà, R., Melocchi, L., Gandolfi, L., Parrella, P., Pasculli, B., ... Fassan, M. (2025). KRAS Mutations in Colorectal Adenocarcinoma: Incidence and Association with Histological Features with Particular Reference to Gly12Asp in a Multicenter GIPAD Real-World Study. Cancers, 17(17), 2721. https://doi.org/10.3390/cancers17172721









