DDX3X and Stress Granules: Emerging Players in Cancer and Drug Resistance
Abstract
Simple Summary
Abstract
1. Introduction
2. Overview of DDX3X
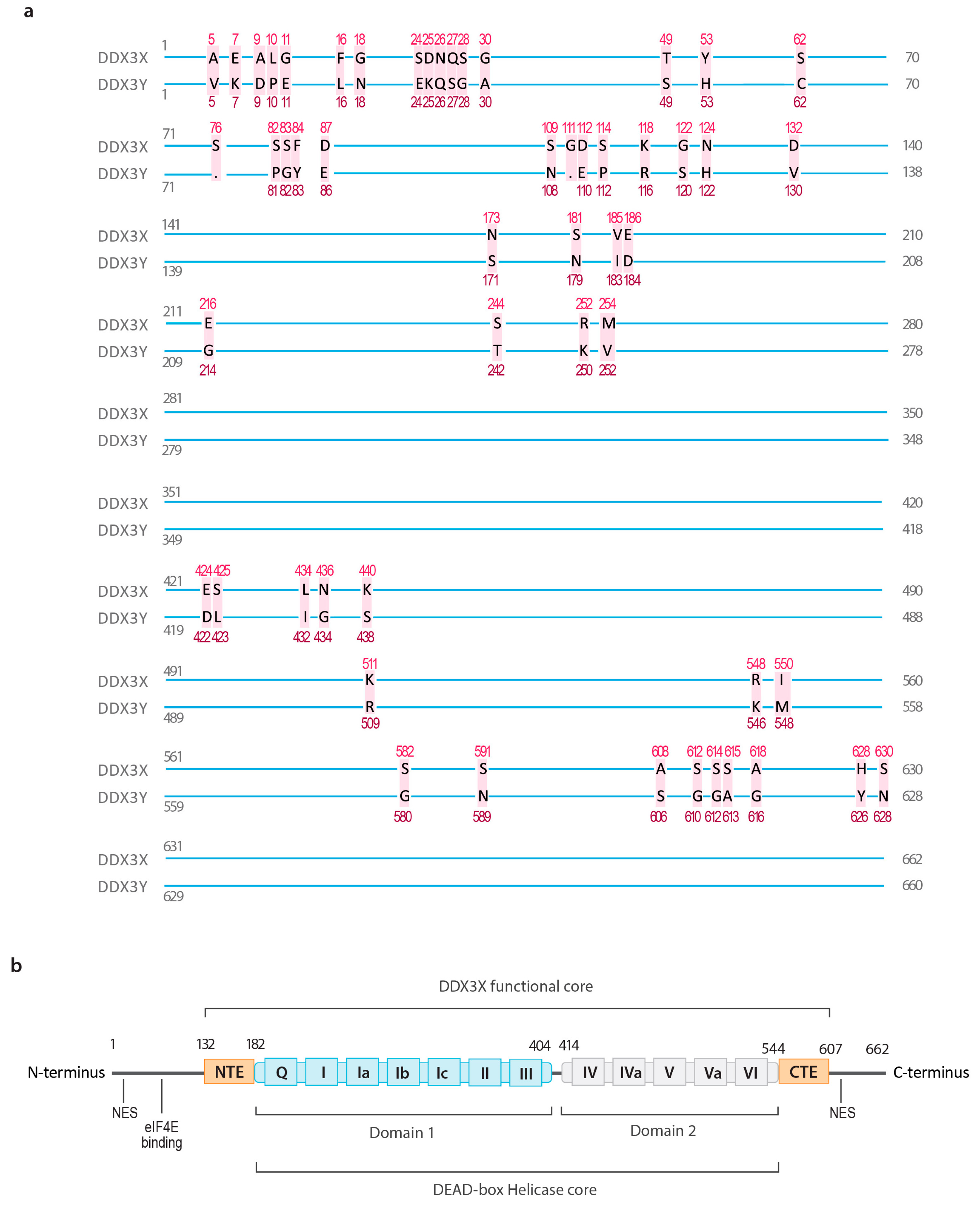
2.1. The Structure of DDX3X
2.2. The Biological Functions of DDX3X in Normal Physiology and Diseases
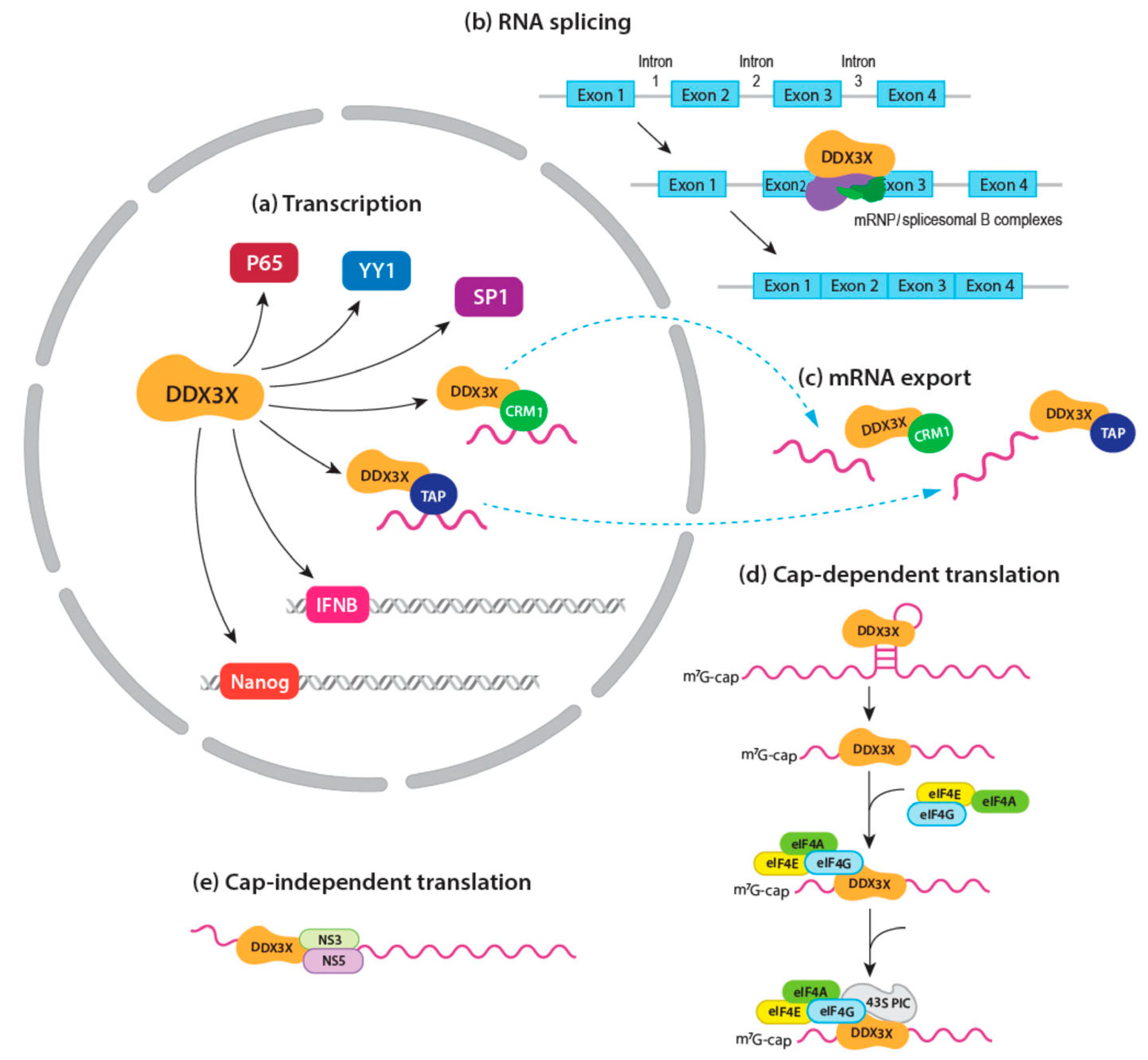
2.2.1. mRNA Transcription
2.2.2. mRNA Splicing
2.2.3. mRNA Export
2.2.4. mRNA Translation
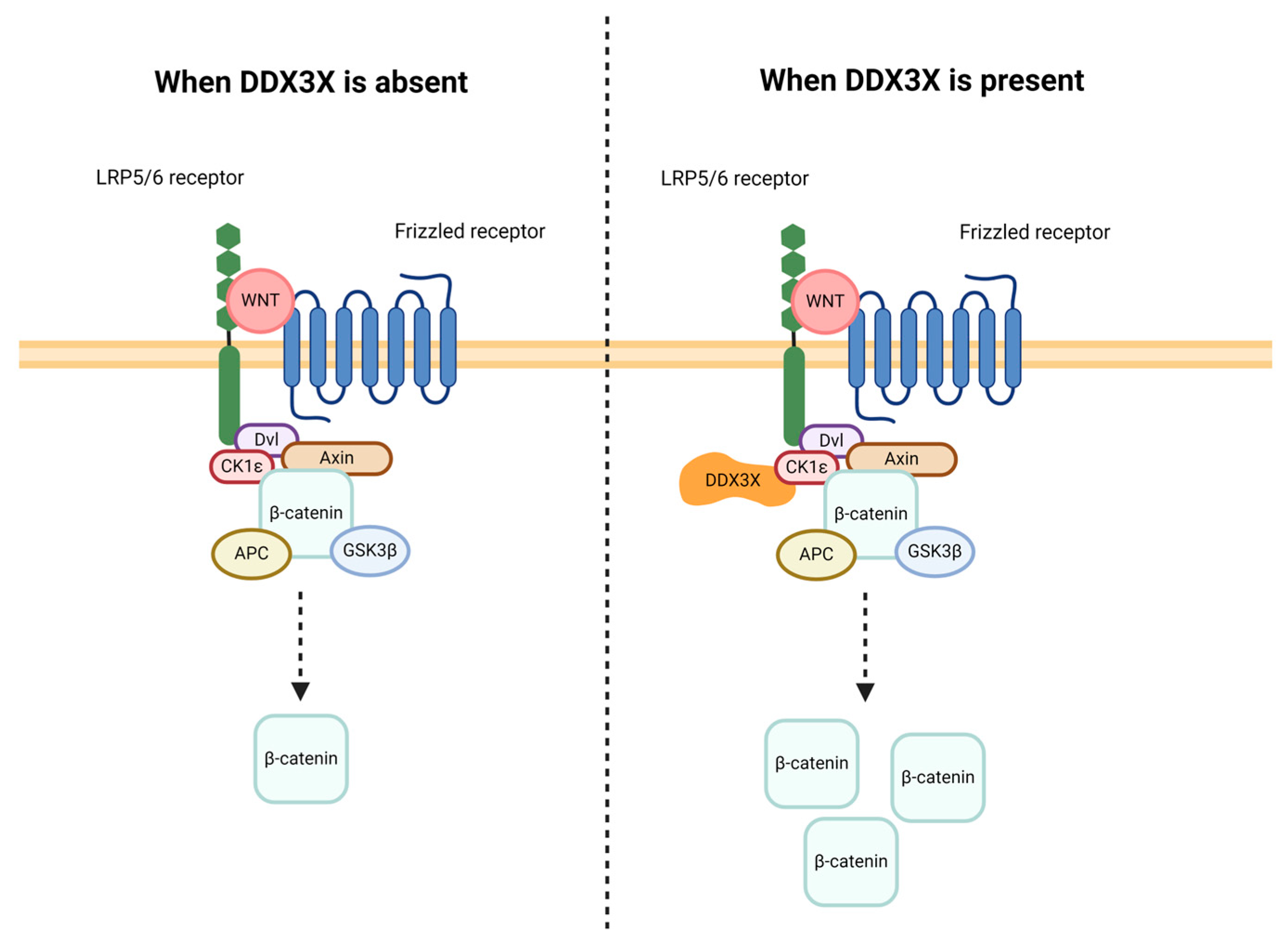
2.2.5. Signal Transduction
2.2.6. Cell Cycle Regulation
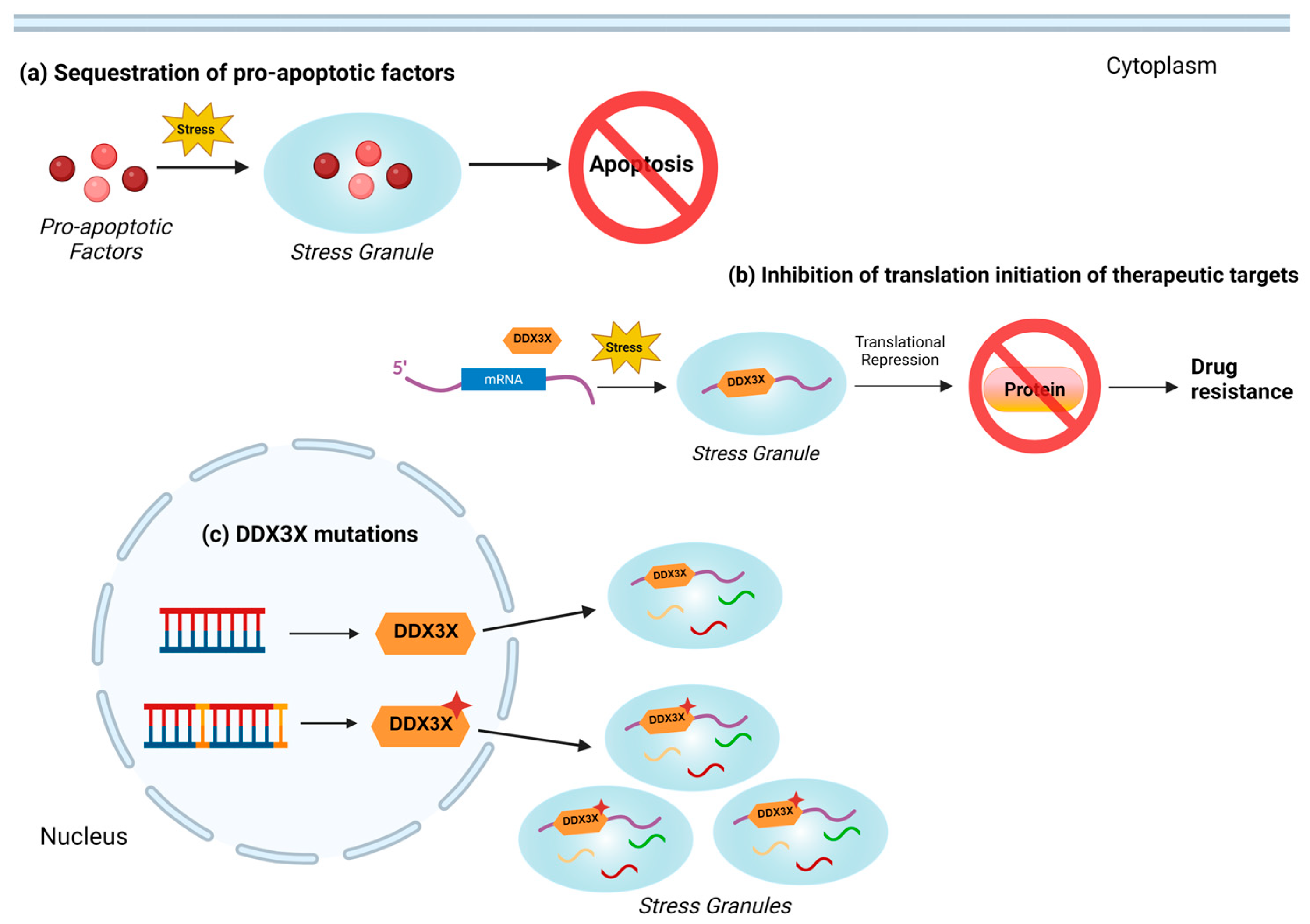
2.2.7. Stress Response
3. DDX3X and Drug Resistance in Cancer
3.1. Overview of Drug Resistance Mechanisms in Cancer
3.2. The Role of DDX3X in Drug Resistance
3.3. DDX3X-Mediated Regulation of Stress Granules in Drug Resistance
3.4. Implications of DDX3X Inhibitors in Cancer Treatment
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Donsbach, P.; Klostermeier, D. Regulation of RNA helicase activity: Principles and examples. Biol. Chem. 2021, 402, 529–559. [Google Scholar] [CrossRef] [PubMed]
- Linder, P.; Jankowsky, E. From unwinding to clamping—The DEAD box RNA helicase family. Nat. Rev. Mol. Cell Biol. 2011, 12, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Bol, G.M.; Xie, M.; Raman, V. DDX3, a potential target for cancer treatment. Mol. Cancer 2015, 14, 188. [Google Scholar] [CrossRef]
- Hoye, M.L.; Calviello, L.; Poff, A.J.; Ejimogu, N.E.; Newman, C.R.; Montgomery, M.D.; Ou, J.; Floor, S.N.; Silver, D.L. Aberrant cortical development is driven by impaired cell cycle and translational control in a DDX3X syndrome model. eLife 2022, 11, e78203. [Google Scholar] [CrossRef] [PubMed]
- Venkataramanan, S.; Gadek, M.; Calviello, L.; Wilkins, K.; Floor, S.N. DDX3X and DDX3Y are redundant in protein synthesis. RNA 2021, 27, 1577–1588. [Google Scholar] [CrossRef]
- Mo, J.; Liang, H.; Su, C.; Li, P.; Chen, J.; Zhang, B. DDX3X: Structure, physiologic functions and cancer. Mol. Cancer 2021, 20, 38. [Google Scholar] [CrossRef]
- Cotton, A.M.; Price, E.M.; Jones, M.J.; Balaton, B.P.; Kobor, M.S.; Brown, C.J. Landscape of DNA methylation on the X chromosome reflects CpG density, functional chromatin state and X-chromosome inactivation. Hum. Mol. Genet. 2015, 24, 1528–1539. [Google Scholar] [CrossRef]
- Calviello, L.; Venkataramanan, S.; Rogowski, K.J.; Wyler, E.; Wilkins, K.; Tejura, M.; Thai, B.; Krol, J.; Filipowicz, W.; Landthaler, M.; et al. DDX3 depletion represses translation of mRNAs with complex 5’ UTRs. Nucleic Acids Res. 2021, 49, 5336–5350. [Google Scholar] [CrossRef]
- Geissler, R.; Golbik, R.P.; Behrens, S.E. The DEAD-box helicase DDX3 supports the assembly of functional 80S ribosomes. Nucleic Acids Res. 2012, 40, 4998–5011. [Google Scholar] [CrossRef]
- Ryan, C.S.; Schroder, M. The human DEAD-box helicase DDX3X as a regulator of mRNA translation. Front. Cell Dev. Biol. 2022, 10, 1033684. [Google Scholar] [CrossRef] [PubMed]
- Vellky, J.E.; McSweeney, S.T.; Ricke, E.A.; Ricke, W.A. RNA-binding protein DDX3 mediates posttranscriptional regulation of androgen receptor: A mechanism of castration resistance. Proc. Natl. Acad. Sci. USA 2020, 117, 28092–28101. [Google Scholar] [CrossRef]
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A.; et al. Proteomics. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef]
- Matsumura, T.; Endo, T.; Isotani, A.; Ogawa, M.; Ikawa, M. An azoospermic factor gene, Ddx3y and its paralog, Ddx3x are dispensable in germ cells for male fertility. J. Reprod. Dev. 2019, 65, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Vakilian, H.; Mirzaei, M.; Sharifi Tabar, M.; Pooyan, P.; Habibi Rezaee, L.; Parker, L.; Haynes, P.A.; Gourabi, H.; Baharvand, H.; Salekdeh, G.H. DDX3Y, a Male-Specific Region of Y Chromosome Gene, May Modulate Neuronal Differentiation. J. Proteome Res. 2015, 14, 3474–3483. [Google Scholar] [CrossRef] [PubMed]
- Marcelo, A.; Koppenol, R.; de Almeida, L.P.; Matos, C.A.; Nobrega, C. Stress granules, RNA-binding proteins and polyglutamine diseases: Too much aggregation? Cell Death Dis. 2021, 12, 592. [Google Scholar] [CrossRef]
- Kedersha, N.; Stoecklin, G.; Ayodele, M.; Yacono, P.; Lykke-Andersen, J.; Fritzler, M.J.; Scheuner, D.; Kaufman, R.J.; Golan, D.E.; Anderson, P. Stress granules and processing bodies are dynamically linked sites of mRNP remodeling. J. Cell Biol. 2005, 169, 871–884. [Google Scholar] [CrossRef]
- Hofmann, S.; Cherkasova, V.; Bankhead, P.; Bukau, B.; Stoecklin, G. Translation suppression promotes stress granule formation and cell survival in response to cold shock. Mol. Biol. Cell 2012, 23, 3786–3800. [Google Scholar] [CrossRef]
- Valentin-Vega, Y.A.; Wang, Y.D.; Parker, M.; Patmore, D.M.; Kanagaraj, A.; Moore, J.; Rusch, M.; Finkelstein, D.; Ellison, D.W.; Gilbertson, R.J.; et al. Cancer-associated DDX3X mutations drive stress granule assembly and impair global translation. Sci. Rep. 2016, 6, 25996. [Google Scholar] [CrossRef]
- Hogbom, M.; Collins, R.; van den Berg, S.; Jenvert, R.M.; Karlberg, T.; Kotenyova, T.; Flores, A.; Karlsson Hedestam, G.B.; Schiavone, L.H. Crystal structure of conserved domains 1 and 2 of the human DEAD-box helicase DDX3X in complex with the mononucleotide AMP. J. Mol. Biol. 2007, 372, 150–159. [Google Scholar] [CrossRef] [PubMed]
- Garbelli, A.; Beermann, S.; Di Cicco, G.; Dietrich, U.; Maga, G. A motif unique to the human DEAD-box protein DDX3 is important for nucleic acid binding, ATP hydrolysis, RNA/DNA unwinding and HIV-1 replication. PLoS ONE 2011, 6, e19810. [Google Scholar] [CrossRef]
- Putnam, A.A.; Jankowsky, E. DEAD-box helicases as integrators of RNA, nucleotide and protein binding. Biochim. Biophys. Acta 2013, 1829, 884–893. [Google Scholar] [CrossRef]
- Soto-Rifo, R.; Ohlmann, T. The role of the DEAD-box RNA helicase DDX3 in mRNA metabolism. Wiley Interdiscip. Rev. RNA 2013, 4, 369–385. [Google Scholar] [CrossRef]
- Floor, S.N.; Condon, K.J.; Sharma, D.; Jankowsky, E.; Doudna, J.A. Autoinhibitory Interdomain Interactions and Subfamily-specific Extensions Redefine the Catalytic Core of the Human DEAD-box Protein DDX3. J. Biol. Chem. 2016, 291, 2412–2421. [Google Scholar] [CrossRef]
- Song, H.; Ji, X. The mechanism of RNA duplex recognition and unwinding by DEAD-box helicase DDX3X. Nat. Commun. 2019, 10, 3085. [Google Scholar] [CrossRef]
- Vellky, J.E.; Ricke, E.A.; Huang, W.; Ricke, W.A. Expression and Localization of DDX3 in Prostate Cancer Progression and Metastasis. Am. J. Pathol. 2019, 189, 1256–1267. [Google Scholar] [CrossRef]
- Shih, J.W.; Wang, W.T.; Tsai, T.Y.; Kuo, C.Y.; Li, H.K.; Wu Lee, Y.H. Critical roles of RNA helicase DDX3 and its interactions with eIF4E/PABP1 in stress granule assembly and stress response. Biochem. J. 2012, 441, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.W.; Lee, M.C.; Wang, J.; Chen, C.Y.; Cheng, Y.W.; Lee, H. DDX3 loss by p53 inactivation promotes tumor malignancy via the MDM2/Slug/E-cadherin pathway and poor patient outcome in non-small-cell lung cancer. Oncogene 2014, 33, 1515–1526. [Google Scholar] [CrossRef] [PubMed]
- Chao, C.H.; Chen, C.M.; Cheng, P.L.; Shih, J.W.; Tsou, A.P.; Lee, Y.H. DDX3, a DEAD box RNA helicase with tumor growth-suppressive property and transcriptional regulation activity of the p21waf1/cip1 promoter, is a candidate tumor suppressor. Cancer Res. 2006, 66, 6579–6588. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.W.; Lin, P.L.; Cheng, Y.W.; Huang, C.C.; Wang, L.; Lee, H. DDX3 enhances oncogenic KRAS-induced tumor invasion in colorectal cancer via the beta-catenin/ZEB1 axis. Oncotarget 2016, 7, 22687–22699. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.W.; Liu, W.S.; Wang, J.; Chen, C.Y.; Cheng, Y.W.; Lee, H. Reduced p21(WAF1/CIP1) via alteration of p53-DDX3 pathway is associated with poor relapse-free survival in early-stage human papillomavirus-associated lung cancer. Clin. Cancer Res. 2011, 17, 1895–1905. [Google Scholar] [CrossRef]
- Yang, F.; Fang, E.; Mei, H.; Chen, Y.; Li, H.; Li, D.; Song, H.; Wang, J.; Hong, M.; Xiao, W.; et al. Cis-Acting circ-CTNNB1 Promotes beta-Catenin Signaling and Cancer Progression via DDX3-Mediated Transactivation of YY1. Cancer Res. 2019, 79, 557–571. [Google Scholar] [CrossRef]
- Xiang, N.; He, M.; Ishaq, M.; Gao, Y.; Song, F.; Guo, L.; Ma, L.; Sun, G.; Liu, D.; Guo, D.; et al. The DEAD-Box RNA Helicase DDX3 Interacts with NF-kappaB Subunit p65 and Suppresses p65-Mediated Transcription. PLoS ONE 2016, 11, e0164471. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Fullam, A.; Brennan, R.; Schroder, M. Human DEAD box helicase 3 couples IkappaB kinase epsilon to interferon regulatory factor 3 activation. Mol. Cell Biol. 2013, 33, 2004–2015. [Google Scholar] [CrossRef] [PubMed]
- Karmakar, S.; Rauth, S.; Nallasamy, P.; Perumal, N.; Nimmakayala, R.K.; Leon, F.; Gupta, R.; Barkeer, S.; Venkata, R.C.; Raman, V.; et al. RNA Polymerase II-Associated Factor 1 Regulates Stem Cell Features of Pancreatic Cancer Cells, Independently of the PAF1 Complex, via Interactions With PHF5A and DDX3. Gastroenterology 2020, 159, 1898–1915.e1896. [Google Scholar] [CrossRef] [PubMed]
- Saikruang, W.; Ang Yan Ping, L.; Abe, H.; Kasumba, D.M.; Kato, H.; Fujita, T. The RNA helicase DDX3 promotes IFNB transcription via enhancing IRF-3/p300 holocomplex binding to the IFNB promoter. Sci. Rep. 2022, 12, 3967. [Google Scholar] [CrossRef] [PubMed]
- Merz, C.; Urlaub, H.; Will, C.L.; Luhrmann, R. Protein composition of human mRNPs spliced in vitro and differential requirements for mRNP protein recruitment. RNA 2007, 13, 116–128. [Google Scholar] [CrossRef] [PubMed]
- Deckert, J.; Hartmuth, K.; Boehringer, D.; Behzadnia, N.; Will, C.L.; Kastner, B.; Stark, H.; Urlaub, H.; Luhrmann, R. Protein composition and electron microscopy structure of affinity-purified human spliceosomal B complexes isolated under physiological conditions. Mol. Cell Biol. 2006, 26, 5528–5543. [Google Scholar] [CrossRef]
- Mahboobi, S.H.; Javanpour, A.A.; Mofrad, M.R. The interaction of RNA helicase DDX3 with HIV-1 Rev-CRM1-RanGTP complex during the HIV replication cycle. PLoS ONE 2015, 10, e0112969. [Google Scholar] [CrossRef]
- Frohlich, A.; Rojas-Araya, B.; Pereira-Montecinos, C.; Dellarossa, A.; Toro-Ascuy, D.; Prades-Perez, Y.; Garcia-de-Gracia, F.; Garces-Alday, A.; Rubilar, P.S.; Valiente-Echeverria, F.; et al. DEAD-box RNA helicase DDX3 connects CRM1-dependent nuclear export and translation of the HIV-1 unspliced mRNA through its N-terminal domain. Biochim. Biophys. Acta 2016, 1859, 719–730. [Google Scholar] [CrossRef]
- Yedavalli, V.S.; Neuveut, C.; Chi, Y.H.; Kleiman, L.; Jeang, K.T. Requirement of DDX3 DEAD box RNA helicase for HIV-1 Rev-RRE export function. Cell 2004, 119, 381–392. [Google Scholar] [CrossRef]
- Hernandez-Diaz, T.; Valiente-Echeverria, F.; Soto-Rifo, R. RNA Helicase DDX3: A Double-Edged Sword for Viral Replication and Immune Signaling. Microorganisms 2021, 9, 1206. [Google Scholar] [CrossRef] [PubMed]
- Ishaq, M.; Hu, J.; Wu, X.; Fu, Q.; Yang, Y.; Liu, Q.; Guo, D. Knockdown of cellular RNA helicase DDX3 by short hairpin RNAs suppresses HIV-1 viral replication without inducing apoptosis. Mol. Biotechnol. 2008, 39, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Culjkovic, B.; Topisirovic, I.; Skrabanek, L.; Ruiz-Gutierrez, M.; Borden, K.L. eIF4E promotes nuclear export of cyclin D1 mRNAs via an element in the 3’UTR. J. Cell Biol. 2005, 169, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.C.; Lee, Y.H.; Tarn, W.Y. The DEAD-box RNA helicase DDX3 associates with export messenger ribonucleoproteins as well as tip-associated protein and participates in translational control. Mol. Biol. Cell 2008, 19, 3847–3858. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.J.; Lee, S.G. The DEAD-box RNA helicase DDX3 interacts with DDX5, co-localizes with it in the cytoplasm during the G2/M phase of the cycle, and affects its shuttling during mRNP export. J. Cell Biochem. 2012, 113, 985–996. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef]
- Chuang, R.Y.; Weaver, P.L.; Liu, Z.; Chang, T.H. Requirement of the DEAD-Box protein ded1p for messenger RNA translation. Science 1997, 275, 1468–1471. [Google Scholar] [CrossRef]
- Marsden, S.; Nardelli, M.; Linder, P.; McCarthy, J.E. Unwinding single RNA molecules using helicases involved in eukaryotic translation initiation. J. Mol. Biol. 2006, 361, 327–335. [Google Scholar] [CrossRef]
- Lee, C.S.; Dias, A.P.; Jedrychowski, M.; Patel, A.H.; Hsu, J.L.; Reed, R. Human DDX3 functions in translation and interacts with the translation initiation factor eIF3. Nucleic Acids Res. 2008, 36, 4708–4718. [Google Scholar] [CrossRef]
- Soto-Rifo, R.; Rubilar, P.S.; Limousin, T.; de Breyne, S.; Decimo, D.; Ohlmann, T. DEAD-box protein DDX3 associates with eIF4F to promote translation of selected mRNAs. EMBO J. 2012, 31, 3745–3756. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, Z. IRES-mediated cap-independent translation, a path leading to hidden proteome. J. Mol. Cell Biol. 2019, 11, 911–919. [Google Scholar] [CrossRef] [PubMed]
- Shatsky, I.N.; Terenin, I.M.; Smirnova, V.V.; Andreev, D.E. Cap-Independent Translation: What’s in a Name? Trends Biochem. Sci. 2018, 43, 882–895. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ge, L.L.; Li, P.P.; Wang, Y.; Dai, J.J.; Sun, M.X.; Huang, L.; Shen, Z.Q.; Hu, X.C.; Ishag, H.; et al. Cellular DDX3 regulates Japanese encephalitis virus replication by interacting with viral un-translated regions. Virology 2014, 449, 70–81. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Putnam, A.A.; Jankowsky, E. Biochemical Differences and Similarities between the DEAD-Box Helicase Orthologs DDX3X and Ded1p. J. Mol. Biol. 2017, 429, 3730–3742. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.; Alam, A.; Pant, R.; Chattopadhyay, S. Wnt Signaling and Its Significance Within the Tumor Microenvironment: Novel Therapeutic Insights. Front. Immunol. 2019, 10, 2872. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.F.; Kaur, P.; Bunnag, N.; Suresh, J.; Sung, I.C.H.; Tan, Q.H.; Gruber, J.; Tolwinski, N.S. WNT Signaling in Disease. Cells 2019, 8, 826. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Mao, Y.; Zhou, J.; Zhao, Y.; Cao, Y.; Chen, X. Multifunctional DDX3: Dual roles in various cancer development and its related signaling pathways. Am. J. Cancer Res. 2016, 6, 387–402. [Google Scholar]
- Cruciat, C.M.; Dolde, C.; de Groot, R.E.; Ohkawara, B.; Reinhard, C.; Korswagen, H.C.; Niehrs, C. RNA helicase DDX3 is a regulatory subunit of casein kinase 1 in Wnt-beta-catenin signaling. Science 2013, 339, 1436–1441. [Google Scholar] [CrossRef]
- Dolde, C.; Bischof, J.; Gruter, S.; Montada, A.; Halekotte, J.; Peifer, C.; Kalbacher, H.; Baumann, U.; Knippschild, U.; Suter, B. A CK1 FRET biosensor reveals that DDX3X is an essential activator of CK1epsilon. J. Cell Sci. 2018, 131, jcs207316. [Google Scholar] [CrossRef]
- Chen, H.H.; Yu, H.I.; Cho, W.C.; Tarn, W.Y. DDX3 modulates cell adhesion and motility and cancer cell metastasis via Rac1-mediated signaling pathway. Oncogene 2015, 34, 2790–2800. [Google Scholar] [CrossRef]
- Gonzalez, D.M.; Medici, D. Signaling mechanisms of the epithelial-mesenchymal transition. Sci. Signal. 2014, 7, re8. [Google Scholar] [CrossRef] [PubMed]
- Botlagunta, M.; Krishnamachary, B.; Vesuna, F.; Winnard, P.T., Jr.; Bol, G.M.; Patel, A.H.; Raman, V. Expression of DDX3 is directly modulated by hypoxia inducible factor-1 alpha in breast epithelial cells. PLoS ONE 2011, 6, e17563. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Song, L.; Zhou, T.; Gillespie, G.Y.; Jope, R.S. The role of DDX3 in regulating Snail. Biochim. Biophys. Acta 2011, 1813, 438–447. [Google Scholar] [CrossRef] [PubMed]
- Su, C.Y.; Lin, T.C.; Lin, Y.F.; Chen, M.H.; Lee, C.H.; Wang, H.Y.; Lee, Y.C.; Liu, Y.P.; Chen, C.L.; Hsiao, M. DDX3 as a strongest prognosis marker and its downregulation promotes metastasis in colorectal cancer. Oncotarget 2015, 6, 18602–18612. [Google Scholar] [CrossRef]
- Wu, D.W.; Lin, P.L.; Wang, L.; Huang, C.C.; Lee, H. The YAP1/SIX2 axis is required for DDX3-mediated tumor aggressiveness and cetuximab resistance in KRAS-wild-type colorectal cancer. Theranostics 2017, 7, 1114–1132. [Google Scholar] [CrossRef]
- Bol, G.M.; Raman, V.; van der Groep, P.; Vermeulen, J.F.; Patel, A.H.; van der Wall, E.; van Diest, P.J. Expression of the RNA helicase DDX3 and the hypoxia response in breast cancer. PLoS ONE 2013, 8, e63548. [Google Scholar] [CrossRef]
- Cannizzaro, E.; Bannister, A.J.; Han, N.; Alendar, A.; Kouzarides, T. DDX3X RNA helicase affects breast cancer cell cycle progression by regulating expression of KLF4. FEBS Lett. 2018, 592, 2308–2322. [Google Scholar] [CrossRef]
- Sergeeva, O.; Zatsepin, T. RNA Helicases as Shadow Modulators of Cell Cycle Progression. Int. J. Mol. Sci. 2021, 22, 2984. [Google Scholar] [CrossRef]
- Ivanov, P.; Kedersha, N.; Anderson, P. Stress Granules and Processing Bodies in Translational Control. Cold Spring Harb. Perspect. Biol. 2019, 11, a032813. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Li, J.; Fan, S.; Jin, Z.; Huang, C. Targeting stress granules: A novel therapeutic strategy for human diseases. Pharmacol. Res. 2020, 161, 105143. [Google Scholar] [CrossRef] [PubMed]
- Khong, A.; Matheny, T.; Jain, S.; Mitchell, S.F.; Wheeler, J.R.; Parker, R. The Stress Granule Transcriptome Reveals Principles of mRNA Accumulation in Stress Granules. Mol. Cell 2017, 68, 808–820.e805. [Google Scholar] [CrossRef]
- Fedorovskiy, A.G.; Burakov, A.V.; Terenin, I.M.; Bykov, D.A.; Lashkevich, K.A.; Popenko, V.I.; Makarova, N.E.; Sorokin, I.I.; Sukhinina, A.P.; Prassolov, V.S.; et al. A Solitary Stalled 80S Ribosome Prevents mRNA Recruitment to Stress Granules. Biochemistry 2023, 88, 1786–1799. [Google Scholar] [CrossRef] [PubMed]
- Drino, A.; Konig, L.; Capitanchik, C.; Sanadgol, N.; Janisiw, E.; Rappol, T.; Vilardo, E.; Schaefer, M.R. Identification of RNA helicases with unwinding activity on angiogenin-processed tRNAs. Nucleic Acids Res. 2023, 51, 1326–1352. [Google Scholar] [CrossRef] [PubMed]
- Cui, B.C.; Sikirzhytski, V.; Aksenova, M.; Lucius, M.D.; Levon, G.H.; Mack, Z.T.; Pollack, C.; Odhiambo, D.; Broude, E.; Lizarraga, S.B.; et al. Pharmacological inhibition of DEAD-Box RNA Helicase 3 attenuates stress granule assembly. Biochem. Pharmacol. 2020, 182, 114280. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Mathieu, C.; Kolaitis, R.M.; Zhang, P.; Messing, J.; Yurtsever, U.; Yang, Z.; Wu, J.; Li, Y.; Pan, Q.; et al. G3BP1 Is a Tunable Switch that Triggers Phase Separation to Assemble Stress Granules. Cell 2020, 181, 325–345.e328. [Google Scholar] [CrossRef] [PubMed]
- Ku, Y.C.; Lai, M.H.; Lo, C.C.; Cheng, Y.C.; Qiu, J.T.; Tarn, W.Y.; Lai, M.C. DDX3 Participates in Translational Control of Inflammation Induced by Infections and Injuries. Mol. Cell Biol. 2019, 39, e00285-18. [Google Scholar] [CrossRef]
- Samir, P.; Kesavardhana, S.; Patmore, D.M.; Gingras, S.; Malireddi, R.K.S.; Karki, R.; Guy, C.S.; Briard, B.; Place, D.E.; Bhattacharya, A.; et al. DDX3X acts as a live-or-die checkpoint in stressed cells by regulating NLRP3 inflammasome. Nature 2019, 573, 590–594. [Google Scholar] [CrossRef] [PubMed]
- Heerma van Voss, M.R.; Brilliant, J.D.; Vesuna, F.; Bol, G.M.; van der Wall, E.; van Diest, P.J.; Raman, V. Combination treatment using DDX3 and PARP inhibitors induces synthetic lethality in BRCA1-proficient breast cancer. Med. Oncol. 2017, 34, 33. [Google Scholar] [CrossRef] [PubMed]
- Cargill, M.J.; Morales, A.; Ravishankar, S.; Warren, E.H. RNA helicase, DDX3X, is actively recruited to sites of DNA damage in live cells. DNA Repair 2021, 103, 103137. [Google Scholar] [CrossRef]
- Li, G.; Manning, A.C.; Bagi, A.; Yang, X.; Gokulnath, P.; Spanos, M.; Howard, J.; Chan, P.P.; Sweeney, T.; Kitchen, R.; et al. Distinct Stress-Dependent Signatures of Cellular and Extracellular tRNA-Derived Small RNAs. Adv. Sci. 2022, 9, e2200829. [Google Scholar] [CrossRef]
- Epling, L.B.; Grace, C.R.; Lowe, B.R.; Partridge, J.F.; Enemark, E.J. Cancer-associated mutants of RNA helicase DDX3X are defective in RNA-stimulated ATP hydrolysis. J. Mol. Biol. 2015, 427, 1779–1796. [Google Scholar] [CrossRef]
- Pavlova, N.N.; Zhu, J.; Thompson, C.B. The hallmarks of cancer metabolism: Still emerging. Cell Metab. 2022, 34, 355–377. [Google Scholar] [CrossRef]
- Bukowski, K.; Kciuk, M.; Kontek, R. Mechanisms of Multidrug Resistance in Cancer Chemotherapy. Int. J. Mol. Sci. 2020, 21, 3233. [Google Scholar] [CrossRef] [PubMed]
- Haider, T.; Pandey, V.; Banjare, N.; Gupta, P.N.; Soni, V. Drug resistance in cancer: Mechanisms and tackling strategies. Pharmacol. Rep. 2020, 72, 1125–1151. [Google Scholar] [CrossRef] [PubMed]
- Pardeshi, J.; McCormack, N.; Gu, L.; Ryan, C.S.; Schröder, M. DDX3X functionally and physically interacts with Estrogen Receptor-alpha. Biochim. Biophys. Acta BBA Gene Regul. Mech. 2022, 1865, 194787. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Chen, G.; Hu, W. lncRNA HLA Complex Group 18 (HCG18) Facilitated Cell Proliferation, Invasion, and Migration of Prostate Cancer Through Modulating miR-370-3p/DDX3X Axis. Reprod. Sci. 2021, 28, 3406–3416. [Google Scholar] [CrossRef]
- Alkallas, R.; Lajoie, M.; Moldoveanu, D.; Hoang, K.V.; Lefrançois, P.; Lingrand, M.; Ahanfeshar-Adams, M.; Watters, K.; Spatz, A.; Zippin, J.H.; et al. Multi-omic analysis reveals significantly mutated genes and DDX3X as a sex-specific tumor suppressor in cutaneous melanoma. Nat. Cancer 2020, 1, 635–652. [Google Scholar] [CrossRef]
- Huang, J.-S.; Chao, C.-C.; Su, T.-L.; Yeh, S.-H.; Chen, D.-S.; Chen, C.-T.; Chen, P.-J.; Jou, Y.-S. Diverse cellular transformation capability of overexpressed genes in human hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 2004, 315, 950–958. [Google Scholar] [CrossRef]
- Botlagunta, M.; Vesuna, F.; Mironchik, Y.; Raman, A.; Lisok, A.; Winnard, P.; Mukadam, S.; Van Diest, P.; Chen, J.H.; Farabaugh, P.; et al. Oncogenic role of DDX3 in breast cancer biogenesis. Oncogene 2008, 27, 3912–3922. [Google Scholar] [CrossRef]
- Nozaki, K.; Kagamu, H.; Shoji, S.; Igarashi, N.; Ohtsubo, A.; Okajima, M.; Miura, S.; Watanabe, S.; Yoshizawa, H.; Narita, I. DDX3X Induces Primary EGFR-TKI Resistance Based on Intratumor Heterogeneity in Lung Cancer Cells Harboring EGFR-Activating Mutations. PLoS ONE 2014, 9, e111019. [Google Scholar] [CrossRef]
- Phung, B.; Cieśla, M.; Sanna, A.; Guzzi, N.; Beneventi, G.; Cao Thi Ngoc, P.; Lauss, M.; Cabrita, R.; Cordero, E.; Bosch, A.; et al. The X-Linked DDX3X RNA Helicase Dictates Translation Reprogramming and Metastasis in Melanoma. Cell Rep. 2019, 27, 3573–3586.e7. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.; Ivanov, P.; Anderson, P. Stress granules and cell signaling: More than just a passing phase? Trends Biochem. Sci. 2013, 38, 494–506. [Google Scholar] [CrossRef] [PubMed]
- Thedieck, K.; Holzwarth, B.; Prentzell, M.T.; Boehlke, C.; Klasener, K.; Ruf, S.; Sonntag, A.G.; Maerz, L.; Grellscheid, S.N.; Kremmer, E.; et al. Inhibition of mTORC1 by astrin and stress granules prevents apoptosis in cancer cells. Cell 2013, 154, 859–874. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.N.Y.; Atkinson, S.C.; Audsley, M.D.; Heaton, S.M.; Jans, D.A.; Borg, N.A. RK-33 Is a Broad-Spectrum Antiviral Agent That Targets DEAD-Box RNA Helicase DDX3X. Cells 2020, 9, 170. [Google Scholar] [CrossRef]
- Xie, M.; Vesuna, F.; Tantravedi, S.; Bol, G.M.; Heerma van Voss, M.R.; Nugent, K.; Malek, R.; Gabrielson, K.; van Diest, P.J.; Tran, P.T.; et al. RK-33 Radiosensitizes Prostate Cancer Cells by Blocking the RNA Helicase DDX3. Cancer Res. 2016, 76, 6340–6350. [Google Scholar] [CrossRef]
- Kondaskar, A.; Kondaskar, S.; Fishbein, J.C.; Carter-Cooper, B.A.; Lapidus, R.G.; Sadowska, M.; Edelman, M.J.; Hosmane, R.S. Structure-based drug design and potent anti-cancer activity of tricyclic 5:7:5-fused diimidazo[4,5-d:4′,5′-f][1,3]diazepines. Bioorg. Med. Chem. 2013, 21, 618–631. [Google Scholar] [CrossRef]
- Samal, S.K.; Routray, S.; Veeramachaneni, G.K.; Dash, R.; Botlagunta, M. Ketorolac salt is a newly discovered DDX3 inhibitor to treat oral cancer. Sci. Rep. 2015, 5, 9982. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Mañán-Mejías, P.M.; Miles, H.N.; Putnam, A.A.; MacGillivray, L.R.; Ricke, W.A. DDX3X and Stress Granules: Emerging Players in Cancer and Drug Resistance. Cancers 2024, 16, 1131. https://doi.org/10.3390/cancers16061131
Zhang H, Mañán-Mejías PM, Miles HN, Putnam AA, MacGillivray LR, Ricke WA. DDX3X and Stress Granules: Emerging Players in Cancer and Drug Resistance. Cancers. 2024; 16(6):1131. https://doi.org/10.3390/cancers16061131
Chicago/Turabian StyleZhang, Han, Paula M. Mañán-Mejías, Hannah N. Miles, Andrea A. Putnam, Leonard R. MacGillivray, and William A. Ricke. 2024. "DDX3X and Stress Granules: Emerging Players in Cancer and Drug Resistance" Cancers 16, no. 6: 1131. https://doi.org/10.3390/cancers16061131
APA StyleZhang, H., Mañán-Mejías, P. M., Miles, H. N., Putnam, A. A., MacGillivray, L. R., & Ricke, W. A. (2024). DDX3X and Stress Granules: Emerging Players in Cancer and Drug Resistance. Cancers, 16(6), 1131. https://doi.org/10.3390/cancers16061131





