1. Introduction
Neuroblastoma is the most common solid tumour in early childhood [
1]. Almost 50% of children with neuroblastoma present with clinically advanced disease and, despite multimodal therapy, survival is poor with treatment options being highly toxic. The v-myc avian myelocytomatosis viral oncogene neuroblastoma derived homolog (MYCN) is a common cancer driver gene in the most aggressive form of neuroblastoma [
2] and other cancers, such as medulloblastoma [
3], ovarian [
4], and prostate cancer [
5]. Aberrant high MYCN expression and amplification is a major clinical determinant of patient relapse [
1].
The development of direct inhibitors of MYCN proteins, thus far, has been difficult. The MYC protein family, of which MYCN is a member, are intrinsically disordered proteins with no deep pockets for drug design. A variety of MYCN protein functions are dependent on protein-protein interactions [
6]. Extensive evidence from us and other researchers has shown that some MYCN binding proteins markedly increase MYCN protein stability to drive tumour formation [
7,
8,
9]. Inhibition of MYCN protein binding partners can markedly reduce MYCN stability and is an emerging treatment strategy for MYCN-driven cancers [
10,
11].
We have previously shown that proliferation-associated protein 2G4 (PA2G4) is a MYCN binding protein that stabilised MYCN in a positive feed-forward expression loop [
11]. WS6, a chemical inhibitor of PA2G4, disrupted the PA2G4-MYCN binding in neuroblastoma cells with significant antitumour effects in vitro and in vivo [
11]. By disrupting the PA2G4-MYCN interaction, treatment with WS6 increased MYCN degradation, in turn decreasing overall levels of MYCN and PA2G4 protein in neuroblastoma cells. However, WS6 exhibited a narrow therapeutic window, with significant toxicity for normal fibroblasts in vitro and in perinatal mice. Furthermore, WS6′s physicochemical properties, in particular its high molecular weight (656.74 Da) and clogP (7.31), suggest it may have significant off-target effects, which preclude clinical development [
12].
Small molecule inhibition of protein-protein interfaces is a challenge due to the undruggable nature of a protein-protein interface [
13]. However, the advantages of small molecules have led to research into the ideal physicochemical properties of successful protein-protein interaction inhibitors [
14,
15,
16]. Our recent analysis showed that, although protein-protein modulators on the market tend to have more variability in their physicochemical properties, those in clinical trials more frequently demonstrate traditional drug-like parameters [
14]. This study indicated that protein-protein inhibitor hit compounds should aim for a cLogP < 3.5, molecular weight < 500, total polar surface area < 100, hydrogen bond donors < 3, and rotatable bonds < 6.
In order to increases WS6 drug-like properties, we studied the cytotoxic effects of different WS6 analogues, with improved physicochemical properties. Two of our hit compounds, #5333 (also called SVI-5333) and #5338 (also called SVI-5338), had a significantly increased therapeutic window compared to WS6 and inhibited the malignant phenotype of neuroblastoma cells in vitro, which partly depended on PA2G4 and MYCN expression. Quantitative mass spectrometry revealed numerous differentially regulated protein targets of #5333 and #5338, providing insights into functional pathways and off target activities, as well as potential biomarkers of the drug response.
2. Materials and Methods
2.1. 2D Analogue Search
A virtual screen of our in-house library of ~16.5 million commercially available compounds was conducted with UNITY (Sybylx2.1), using a backbone of the previously identified compounds and a minimum 80% Tanimoto similarity index [
17]. All compounds were purchased from Chemdiv (
https://www.chemdiv.com/).
2.2. Molecular Docking
The human PA2G4 crystal structure (PDB: 2Q8K) was retrieved from the PDB [
18]. All waters and crystallisation ligands (glycerol and sulfate ions) were removed, and hydrogen atoms were added in Sybylx2.1 (Sybylx2.1, Certara L.P, St. Louis, MO, USA). The Geom-Dock (Sybylx2.1) software function was used to create a protomer (docking receptor) with a bloat protomer level of 2 Å (default 0) and threshold reduced to 0.43 (default 0.50). Three-dimensional (3D) conformers of compounds were created in GeomX-Dock (Sybylx2.1), with parameters set to allow flexibility of rings and movement of heavy atoms and hydrogens. The starting conformations were increased to 5 (default 0) and the number of poses to optimise was set at 20 (default 10). All other parameters were left at the default setting. The resultant dock was analysed visually, and clusters were identified for each ligand.
A second program, Fast-Rigid Exhaustive Docking (FRED) (OpenEye Scientific Software, Santa Fe, NM, USA), was used to confirm the binding mode. A docking receptor was created using the PA2G4 structure (PDB: 2Q8K) and the Make Receptor program (OpenEye Scientific Software, Santa Fe, NM, USA) [
19]. OMEGA v2.4.3 was then used to create a multi-conformer database of WS6 and the analogues. FRED was used to perform a rigid binding pose exploration for PA2G4 protein and ligand complexes. Poses were ranked using the Chemgauss4 scoring function as an indication of pose and docking quality [
19,
20]. FRED docking results were then visualised using VIDA (OpenEye Software, Santa Fe, NM, USA) to identify conformational clusters for each ligand.
2.3. Cell Lines
The MYCN amplified neuroblastoma cell line SK-N-BE(2)-C was kindly provided by Professor June Biedler (Memorial Sloan Kettering Cancer Centre, New York, NY, USA), and Kelly was obtained from European Collection of Cell Cultures. The two normal lung fibroblast cell lines (MRC-5 and WI-38) were purchased from ATCC. SK-N-BE(2)-C shMYCN#2 doxycycline-inducible cell and Kelly shMYCN#2 doxycycline-inducible cell were custom-made by GenScript (Piscataway, NJ, USA). SK-N-BE(2)-C, SK-N-BE(2)-C shMYCN#2, and SHEP MYCN3 doxycycline-inducible cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) (Invitrogen, Life Technologies, Waltham, MA, USA) with 10% foetal calf serum (FCS). Kelly and Kelly shMYCN#2 doxycycline-inducible cell were cultured in Roswell Park Memorial Institute (RPMI) 1640 (Invitrogen, Life Technologies) with 10% FCS. MRC-5 and WI-38 were cultured in Alpha-MEM media (Invitrogen, Life Technologies) with 10% FCS. All cells were not cultured for more than 2 months. All used cell lines were authenticated by Cell Bank Australia (Westmead, NSW, Australia) as being free from mycoplasma, and cultured at 37 °C and 5% CO2 in a humidifier incubator.
2.4. Protein Production
PA2G4 protein construct was ordered from GenScript (Piscataway, NJ, USA) and cloned into a pET22b vector. The construct was expressed in BL21 (DE3) E. coli, which was transformed by adding 2 μL of DNA, as per the Addgene Bacterial Transformation Protocol (Addgene, Cambridge, MA, USA). A total of 250 μL of transformed cells were grown in Luria-Bertani (LB) agar, containing 100 μg/mL of ampicillin, overnight at 37 °C. Starting cultures were created by inoculating 100 mL of LB with several bacterial culture colonies, followed by an overnight shaking incubation at 125 rpm and 37 °C. The overnight culture was added (10 mL/L) to 2 L flasks of LB and incubated at 125 rpm and 37 °C until log phase growth was reached (OD600 = 0.6). Following incubation, the cell line was centrifuged at 4200 rpm for 30 min in 1 L bottles. The cell pellet was then resuspended in 25 mL of phosphate-buffered saline (PBS) and centrifuged again at 4200 rpm for 30 min.
Cell pellets were resuspended in 25 mL of Ni-IMAC buffer A (100 mM Tris (pH 8.0), 0.5 M NaCl, 1 mM TCEP), filtered through 22–25 μm Miracloth, and placed on ice. Filtered cells were lysed using the EmulsiFlex-C5 High-Pressure homogenizer (Avestin, Ottawa, ON, Canada), followed by high-speed centrifugation at 18,000 rpm for 20 min. The supernatant, containing the soluble cell lysate, was loaded onto a 5 mL His-Trap column (GE Healthcare Life Sciences, Buckinghamshire, UK) equilibrated in Ni-IMAC buffer A. An incremental gradient of Ni-IMAC buffer B (100 mM Tris pH 8.0, 0.5 M NaCl, 1 mM TCEP, and 1 M imidazole) was introduced. Eluted fractions were analysed by SDS-PAGE and pooled accordingly. Each protein sample was then loaded onto a HiPrep 26/10 Desalting column (GE Healthcare Life Sciences, Buckinghamshire, UK) and equilibrated into anion exchange buffer A (100 mM Tris pH 8.0, 50 mM NaCl, 1 mM TCEP). Samples were loaded onto the column and eluted fractions were analysed by SDS-PAGE and pooled accordingly.
Protein was loaded onto a 5 mL HiTrap Q (Sephrose-Q) anion exchange column (GE Healthcare Life Sciences, Buckinghamshire, UK) and equilibrated in anion exchange buffer A. Samples were loaded, eluted in the flow-through, and analysed using SDS-PAGE. Samples were pooled and concentrated using 10 kDa MWCO Ultra-4 centrifugal filters (Merck, Macquarie Park, NSW, Australia) and intermittent centrifugation at 4000 rpm. Protein samples were then purified using Size Exclusion Chromatography (SEC). Each sample was concentrated to 10 mL using MWCO Ultra-4 centrifugal filters, then loaded onto a Superdex 75 16/60 SEC column (GE Healthcare Life Sciences, Buckinghamshire, UK) and equilibrated in SEC buffer (25 mM Hepes pH 7.7, 100 mM NaCl, 1 mM TCEP). Samples were eluted, analysed by SDS-PAGE, and concentrated to 10 mg/mL using 10 kDa MWCO Ultra-4 centrifugal filters, with intermitted centrifugation at 4000 rpm.
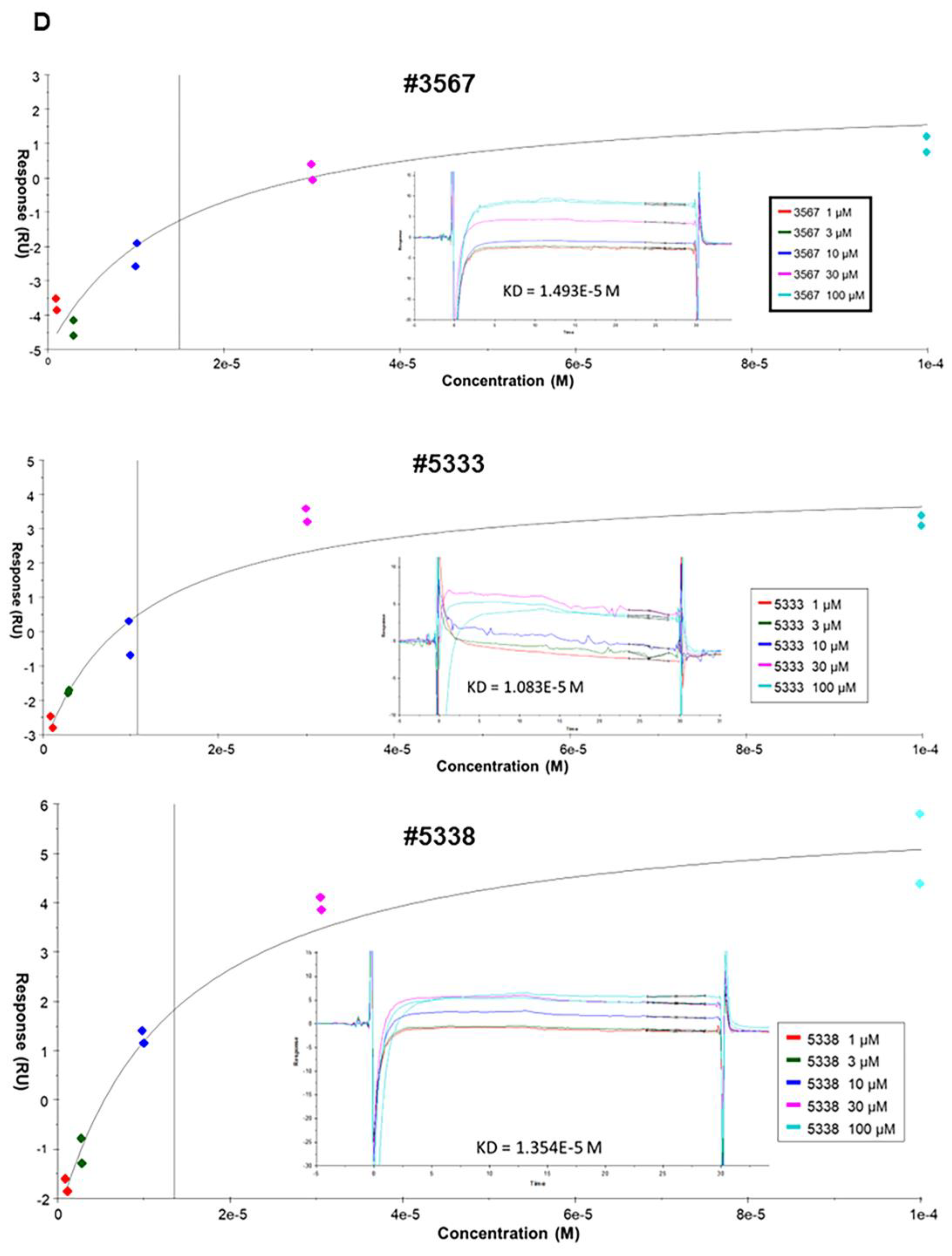
2.5. Surface Plasmon Resonance (SPR)
Recombinant PA2G4 was buffer exchanged into PBS buffer using an overnight dialysis at 4 °C. Protein was concentrated to 0.5 mg/mL and incubated for 3 h on ice with a 20:1 molar excess of EZ-Link Sulfo-NHS-LC-Biotin (Thermo Scientific, Melbourne, VIC, Australia). Biotinylated protein samples were isolated from excess biotin using a Superdex 200 10/300 SEC column (GE Healthcare Life Sciences, Buckinghamshire, UK). All SPR experiments were performed on a Biacore T200 instrument (GE Healthcare Life Sciences, Uppsala, Sweden), at 25 °C, in the presence of HEPES Buffered Saline (HBS) (20 mM phosphate (pH 7.5) 137 mM NaCl, 2.7 mM KCl, and 0.05% Tween 20). Streptavidin was coupled to a CM5 chip (GE Healthcare Life Sciences, Buckinghamshire, UK) using amine coupling. Biotinylated protein samples were then capture-coupled to the streptavidin, with flow cell 1 blocked as a reference.
Compounds were dissolved in 100% DMSO and diluted to 10 mM. Prior to conducting compound SPR, a DMSO solvent correction was performed following the Biacore Laboratory guideline 29-0057-18 (GE Healthcare Life Sciences, Buckinghamshire, UK). Compounds were injected at 30 μL/min for 60 s across the chip in a 3-fold dose-response manner. SPR results were analysed using the Biacore T200 Evaluation Software v.1 [
21].
2.6. Colony Formation Assays
A total of 250 SK-N-BE(2)-C and 500 Kelly cells were seeded in 6-well plates and treated with #5333 and #5338 24 h after seeding. Once colonies had formed (14 days for Kelly, 10 days for SK-N-BE(2)-C), cells were fixed and stained with 0.5% crystal violet in 50% methanol and washed twice to remove the unincorporated stain. The photos of colonies were taken by ChemiDoc MP Imaging System (Bio-Rad, South Granville, NSW, Australia). Stained colonies were counted, and colony size was measured using Image J software.
2.7. Cell Proliferation Assays
Cells were seeded in 96-well plates the day before and then treated with #5333 and #5338 for 72 h. Cell proliferation was measured using the BrdU ELISA kit (Roche, Millers Point, Australia), according to the manufacturer’s instructions. 10 μM BrdU was added to media and incubated at 37 °C in 5% CO2 for 2 h. Cells were fixed with supplied fixing solution for 30 min and then blocked with 10% FCS in PBS for 10 min. Peroxidase-conjugated anti-BrdU (1:100) was added and incubated for 90 min at room temperature. After washing with PBS for three times, peroxidase substrate was added to the cells. Changes in cell proliferation were calculated from the absorbance readings at 370 nm (495 nm reference wavelength) on the Benchmark Plus microplate reader (Bio-Rad).
2.8. Cell Viability Assays
Cell viability was determined by the Alamar Blue assay (75 mg resazurin, 12.5 mg methylene blue, 164.5 mg potassium hexacyanoferrate (III), 211 mg potassium hexacyanoferrate (II) trihydrate in 500 mL of phosphate buffered saline). After 6 h incubation with Alamar Blue, the change of fluorescence was measured by Victor 3 multilabel Plate Reader (Perkin Elmer Australia, Melburne, Australia) at an excitation wavelength of 560 nm and an emission wavelength of 590 nm.
2.9. Western Blots
Protein was extracted from cell pellets using RIPA buffer (Sigma) with 10% protease inhibitor (Sigma, Sydney, Australia). Quantification of proteins was conducted using Pierce BCA protein assay kit (Thermo Scientific), according to manufacturer’s instructions. A total of 30–50 μg protein samples were loaded onto Criterion TGX 10% precast gel (Bio-Rad) or NuPAGE™ 4–12% Mini Protein Gel (Invitrogen, Life technologies) and transferred to a nitrocellulose membrane (Bio-Rad). The membrane was blocked in 5% skim milk in Tris-buffered saline with Tween-20 (20 mM Tris-HCl (pH 7.6), 137 mM NaCl, 0.1% Tween-20) for an hour before incubation at 4 °C overnight with the following primary antibodies: mouse anti-MYCN(1:2000, Santa-Cruz, Cat# sc-53993), rabbit anti-PA2G4 (1:2000, Atlas Antibodies, Cat# HPA016484), rabbit anti-PARP (1:1000, Cell Signalling Technologies, Danvers, MA, USA, Cat# 9542), rabbit anti-Caspase-3 (1:500, Cell Signalling Technologies, Cat# 9662), mouse anti-Bcl-2 (1:1000, Cell Signalling Technologies, Cat# 15071), rabbit anti-Bax (1:1000, Cell Signalling Technologies, Cat# 2772), mouse anti-Vinculin (1:2000, Sigma, Cat# V9131), mouse anti-GAPDH (1:2000, Santa-cruz, Cat# sc-365062), rabbit anti-β-actin (1:5000, Sigma, Cat# SAB2100037), rabbit anti-GATA2 (1:1000, Cell Signalling Technologies, Cat# 4595), mouse anti-Flag/DYKDDDDK tag (1:1000, Cell Signalling Technologies, Cat# 8146). Anti-mouse or anti-rabbit horseradish peroxidase secondary antibodies (1:5000, Life technologies, Cat# 31430, 31460) were added and incubated for 2 h at room temperature. Immunoblots were visualised by Clarity ECL reagent (Bio-Rad) and ChemiDoc MP Imaging System (Bio-Rad). Quantification of protein expression was measured by Image Lab software (Bio-Rad) and normalised to loading control.
2.10. siRNA and Plasmid DNA Transfections
For siRNA-mediated knockdown, 40 nM of Ambion Silencer Select GATA2 siRNAs (Thermo Fisher Scientific) (GATA2 siRNA#1: 5′-GGCUCGUUCCUGUUCAGAATT-3′, GATA2 siRNA #2: 5′-GGUACAGCUGUAUAUAAACTT-3′) (Cat# AM16704) were transfected with lipofectamine 2000 (Life Technologies), according to manufacturer’s instruction. Dharmacon on-target plus control siRNA (Cat# D-001810-10-20) was used as siControl. Cells were transfected between 24, 48, and 96 h, depending on the experimental requirements. For overexpression, 1 µg/mL of pCMV6-GATA2-Myc/DDK (Origene, Rockville, MD, USA) was transfected with pCMV6-Empty (Origene) as control. Lipofectamine 2000 (Life Technologies) was used, following manufacturer’s instruction.
2.11. Flow Cytometry
Neuroblastoma cell lines were seeded 24 h before treating with #5333 or #5338 for 48 h. The apoptotic and necrotic effects of the #5333 and #5338 treatment were determined by staining the cells with Annexin V PE/7-AAD using the Annexin V: PE Apoptosis Detection Kit (BD Biosciences, North Ryde, NSW, Australia), according to the manufacturer’s instruction. Stained cells were measured by flow cytometry using the BD LSRFortessa (BD Biosciences, NSW, Australia) and the data was analysed using the FlowJo software.
2.12. Methods for LFQ LC/MS
SK-N-BE(2)-C cells were incubated with #5333 or #5338 at IC50 concentration of the compound for 72 h. Cells were harvested and the pellets were lysed with RIPA buffer containing protease inhibitors (Sigma-Aldrich, St. Louis, MO, USA). Cell lysates were sent for LFQ/LC/MS analysis at the Bioanalytical Mass Spectrometry Facility, UNSW Sydney. Differentially expressed proteins where identified as those that had a
p-value < 0.05 and a log2 fold-change (log2FC) of >1.5 for upregulated and <−1.5 for downregulated proteins. Differential protein expression analysis was performed using the DEP (v1.14.0) [
22] package in R(v4.1.1). Variance stabilising transformation (vsn) method was used to normalise the protein expression data. Threshold of
p-value < 0.05 and log2 fold-change value > |0.6| was used as a cut off to identify differentially expressed proteins. Functional enrichment analysis was performed using DAVID (v 6.8) [
23].
2.13. Co-Immunoprecipitation Assay
SK-N-BE(2)-C cells were incubated with #5333 (53 µM) or #5338 (47 µM) at 2 × IC50 concentration of the compound for 1 h. The protein was harvested and the pellet was lysed in ice-cold BC100 buffer (20 mm Tris-HCl, pH 7.9, 100 mm NaCl, 10% glycerol, 0.2 mm EDTA, 0.2% Triton X-100, and freshly supplemented protease inhibitor) and incubated on ice for 30 min. After centrifugation at 12,000× g for 20 min at 4 °C, the supernatant was collected. For immunoblotting, 20–40 µg (calculated using the ThermoFischer Scientific BCA Assay kit) of whole protein lysates were resolved on either 10.5% or 10–14% Tris-HCl Criterion gels (Bio-Rad, South Granville, NSW, Australia). MYCN and GAPDH antibodies were purchased from Santa Cruz Biotechnology. A total of 750 µg of cell lysate was incubated with either MYCN or control mouse IgG antibody, then captured by Gammabind G-Sepharose beads (GE Healthcare). Bound proteins were resolved by SDS-PAGE. All primary antibodies were probed overnight at 4 °C and secondary antibody incubation was carried out at room temperature for 2–4 h. Immunoblots were visualised after incubation with Clarity™ Western ECL substrate and imaged with the ChemiDoc-Touch Imaging System (Bio-Rad, NSW, Australia).
2.14. Statistical Analysis
All statistical analysis was conducted by GraphPad Prism 9 software and differences were analysed using unpaired two-sided t-tests. All data were expressed as mean ± standard error of combined data from at least three independent biological replicates.
4. Discussion
We have previously reported that WS6 inhibits the MYCN-PA2G4 protein-protein interface and markedly reduces MYCN protein stability and neuroblastoma tumorigenesis. Here, we identified two analogues of WS6 with 80% structural similarity and a 70-fold lower toxicity for normal human myofibroblasts than WS6. WS6 analogues also inhibited the malignant neuroblastoma cell phenotype in vitro; their cytotoxicity was partially dependent on PA2G4 and MYCN protein expression. We also found that a transcription factor and MYC target, GATA2, is a downstream target for the anti-cancer effects of the WS6 analogues. Our data provides a basis for the future design of competitive inhibitors of the PA2G4-MYCN protein interface.
In recent years, many groups attempted to target MYCN oncogenic signalling at various stages of its formation and function. Inhibitors of MYCN downstream oncogenic signalling pathways have also been developed [
28]. However, inhibition of MYCN protein binding partners (MYCN-BPs) is emerging as an efficient strategy for targeting MYCN oncogenic signalling in the treatment of MYCN-driven cancers [
6,
7,
10,
11]. A variety of functions of the MYCN gene product are attributable to protein-protein interactions (PPIs). Extensive evidence, from our lab and others, have shown that MYCN-BPs can markedly increase MYCN protein stability to drive tumour formation [
9,
11,
29,
30]. In neuroblastoma cells, PA2G4 is a transactivation target for MYCN. PA2G4 directly binds to and stabilises MYCN by preventing its ubiquitin-mediated proteolysis [
11]. Inhibition of PA2G4 and MYCN binding by WS6 can reduce levels of both proteins and neuroblastoma tumorigenicity in vitro and in vivo. In this study, the two WS6 analogues displayed a much larger therapeutic window compared to WS6. Nonetheless, potential off-target effects of the WS6 analogues need to be investigated. Additionally, the half maximal inhibitory concentration (IC50) of two WS6 analogous in treated neuroblastoma cells was relatively high; therefore, chemical modification to synthesise more potent WS6 analogues is required to improve the effectiveness on neuroblastoma cells in future study. As the 2,3 dimethyl phenoxy ring at R
1 position was important for cytotoxicity (
Figure 1B), we plan to keep this group constant at R
1 position and explore the R
2 position to enhance activity. First, we would explore the effect of bioisosteric equivalents of the trifluoromethyl (CF
3) group, such as iodide (I), isopropyl (iPr), and tert-butyl (tBu), at the meta position of the phenyl ring in R
2 position. Second, we would explore the effect of the carbon chain length between amino (NH) group and the phenyl ring by a systematic increase of carbon linkage from 0–5 carbons for compound #5338. Third, we would explore the bioisosteric replacements of phenyl ring with thiophene, pyridyl rings. Fourth, we would synthesise pure enantiomers of #5338 to study the effect of spatial orientation of the methyl group on activity. In this study, SAR studies revealed that 2,3 dimethyl substitution on the phenoxy ring at R1 position was important for cytotoxicity of #3567 against neuroblastoma cell lines. The compound without this dimethyl substitution, #3561, was not cytotoxic towards neuroblastoma cell lines, SK-N-BE(2)C, and Kelly cells (IC50 > 100 µM). Furthermore, investigating the effect of trifluoromethyl (CF3) group at different positions of phenyl ring at the R2 position revealed that the CF3 group is required at meta position to the amino-methyl group, as #3567 displayed a 2.7-fold therapeutic window against normal cells. Moving the CF3 to the para-position of amino-methyl group decreased the therapeutic window to 1.4-fold and changing the CF3 to ortho-position of amino-methyl group resulted in complete loss of cytotoxicity (#3566, IC50 > 100 µM)). Replacing the CF3 group of #3567 with a bromine atom resulted in an increase in the therapeutic window to 3.6-fold. The series of compounds reduced the cytotoxic effect compared to the parent compound WS6, although there was an increase in the therapeutic window from 2.1- to 3.8-fold.
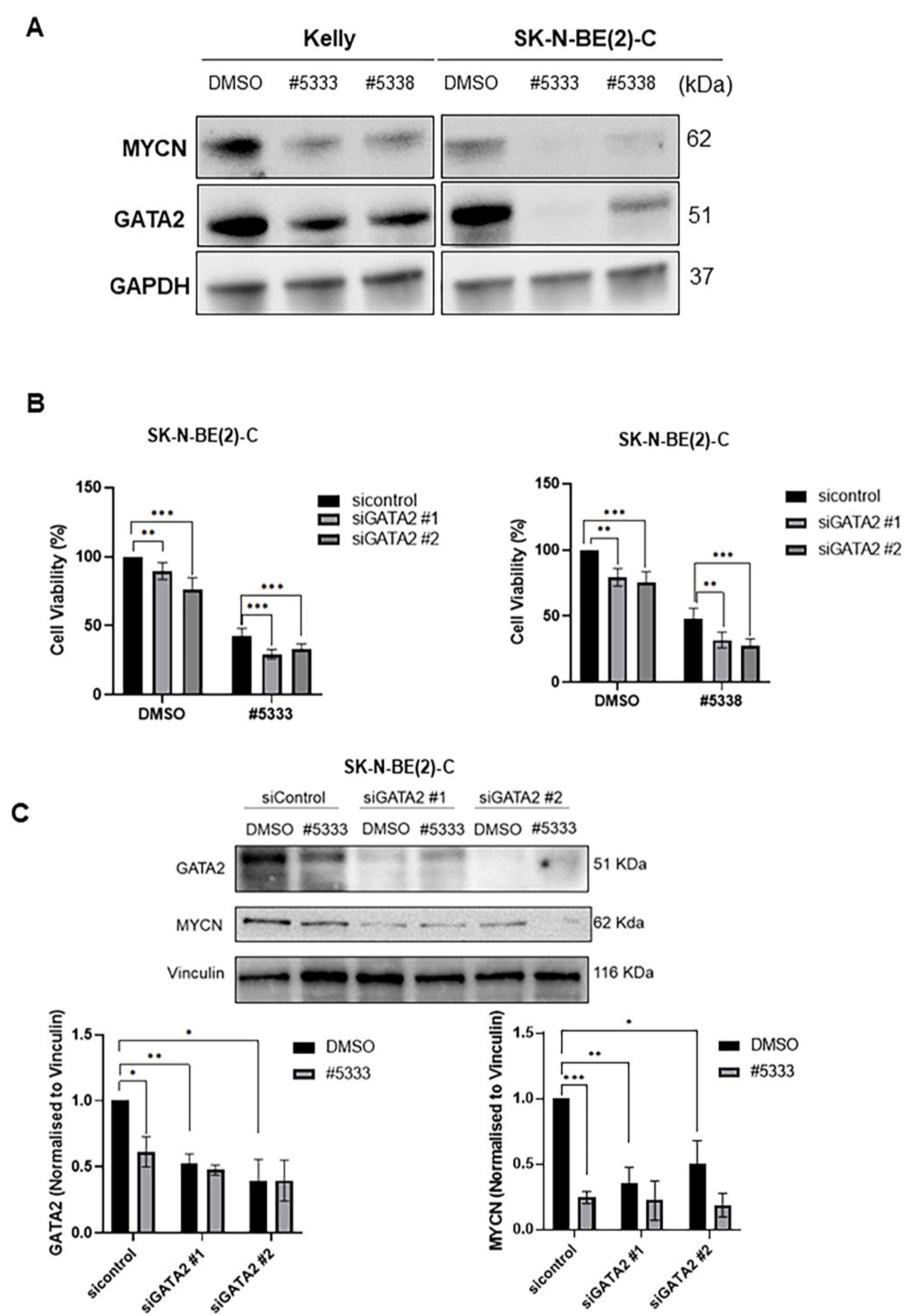
GATA2 is a transcription factor that plays an essential role in gene regulation during vascular development and hematopoietic differentiation. A study found that haploinsufficiency of GATA2 underlies primary lymphedema and predisposes to acute myeloid leukaemia [
31]. GATA2 mutation is also one of the causes of B and NK lymphoid deficiency, which could contribute to leukemic transformation [
32]. Importantly, GATA2 regulates the metastatic potential of prostate cancer cells in the early stages of disease. GATA2 has also been shown to increase the metastatic potential of androgen responsive LNCaP cells. Thus, targeting GATA2 is an attractive therapeutic strategy that might improve the clinical outcome of patients with prostate cancer [
27,
33]. A study in neuroblastoma suggests that GATA2 plays a role in ATRA-induced neuronal differentiation in neuroblastoma SH-SY5Y cells [
34]. GATA2 may also play critical roles in maintaining adrenergic features in poorly differentiated tumours [
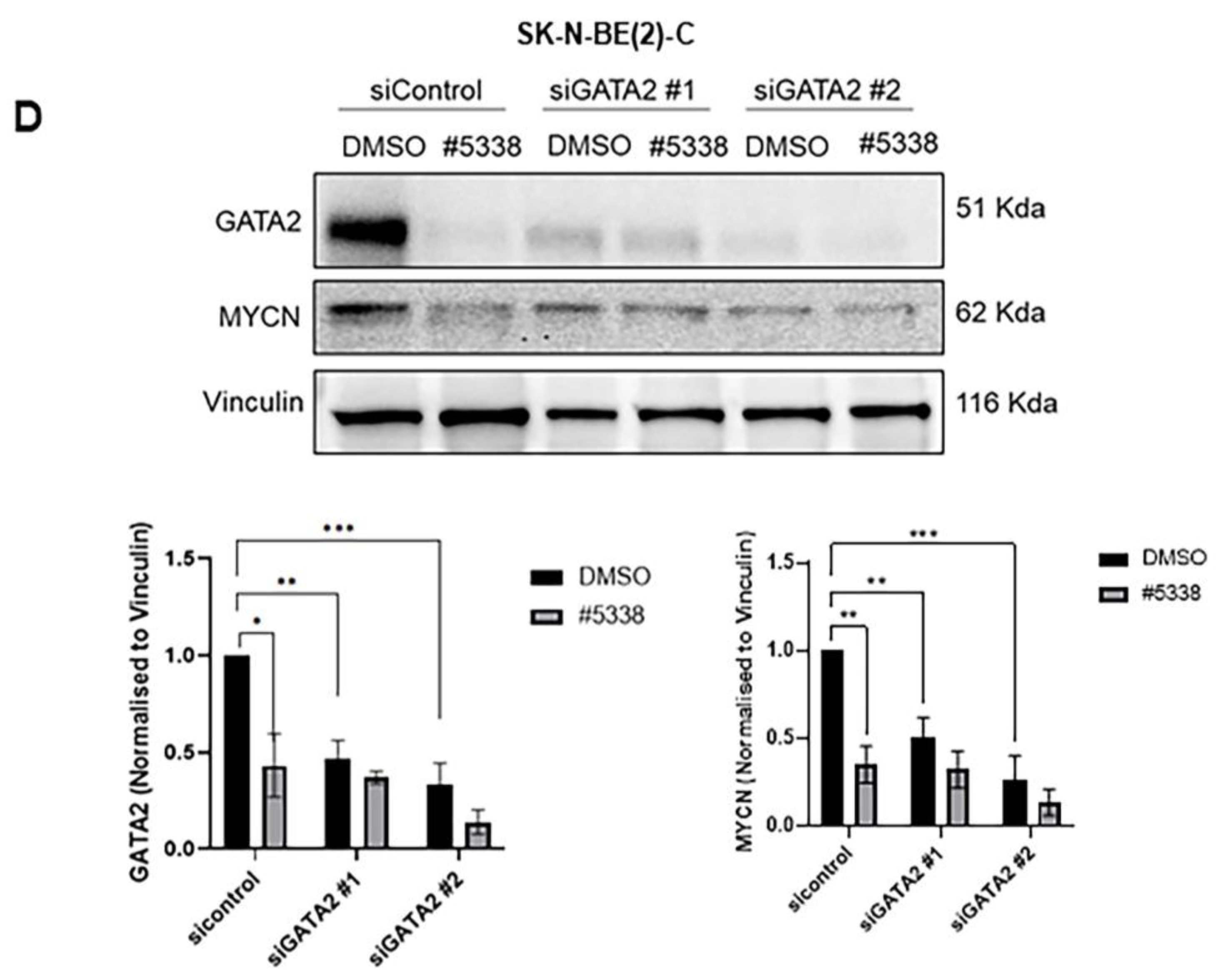
35]. However, the functional role of GATA2 in neuroblastoma has not been fully studied. In this study, we found GATA2 was the most differentially downregulated protein after the treatment of #5333 and #5338, identified by a label-free LC/MS study, and the downregulation of GATA2 protein was confirmed by immunoblotting. GATA2 repression by siRNA increased the sensitivity of neuroblastoma cells to the two compounds, suggesting that GATA2 could be the downstream target in mediating the anti-cancer effects of #5333 and #5338. However, further studies are needed to determine the molecular mechanism for how GATA2 mediates the anti-cancer effects of these compounds.