Applying Explainable Machine Learning Models for Detection of Breast Cancer Lymph Node Metastasis in Patients Eligible for Neoadjuvant Treatment
Abstract
Simple Summary
Abstract
1. Introduction
- machine learning model training, optimization, and evaluation curated specifically for patients eligible for NST;
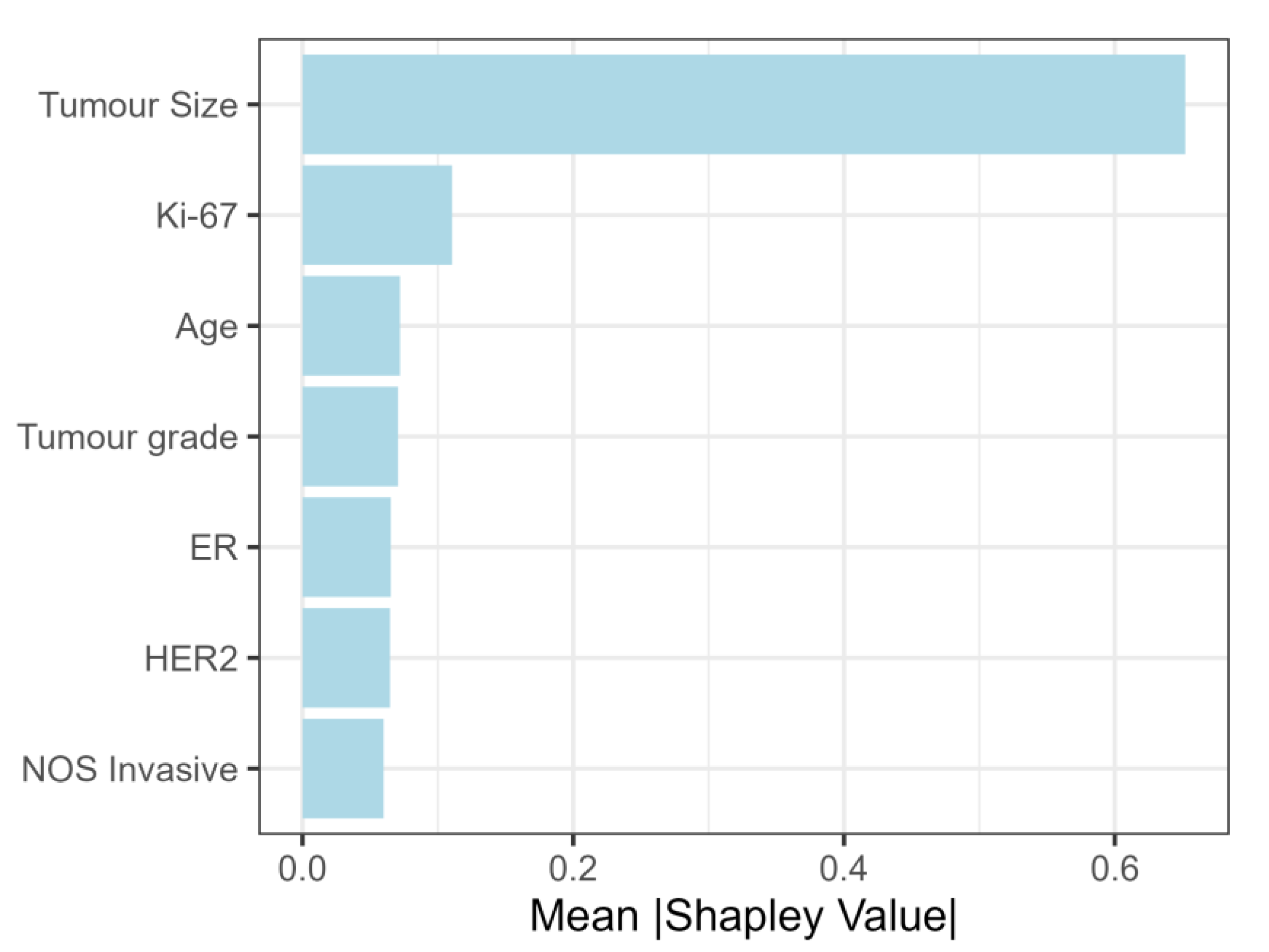
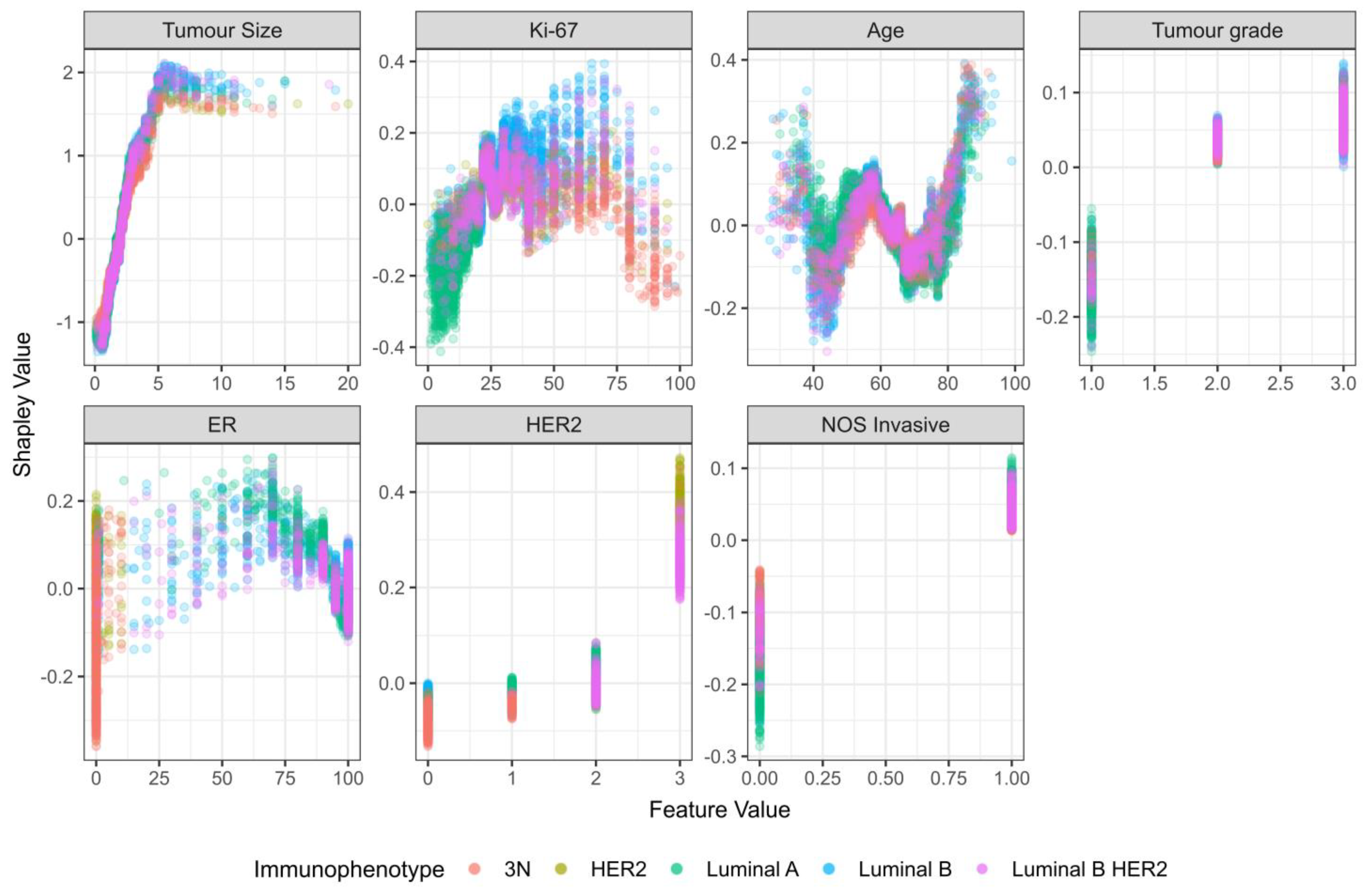
- exhibiting what the model learned and which predictors were the most important in its decision-making process through the use of Shapley values;
- presenting model results for our whole breast cancer population (n = 8381).
2. Materials and Methods
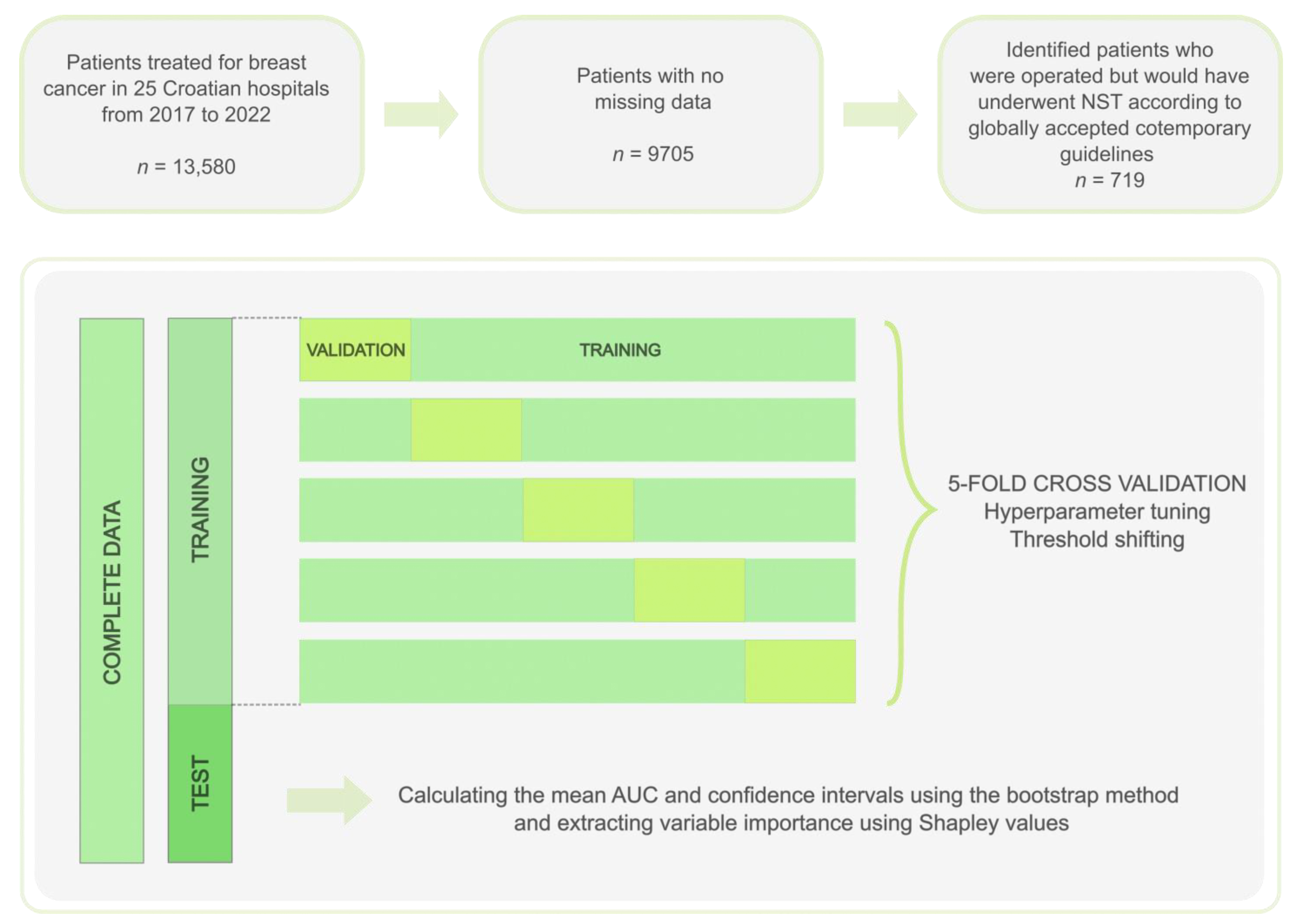
2.1. Data Source and Preparation
- all tumors with size >5 cm (irrespective to subtype),
- tumors with size ≥2 cm of triple-negative or HER-2 positive subtype,
- tumors of inflammatory subtype [30].
2.2. Prediction Model Training, Optimization and Validation
2.3. Model Evaluation
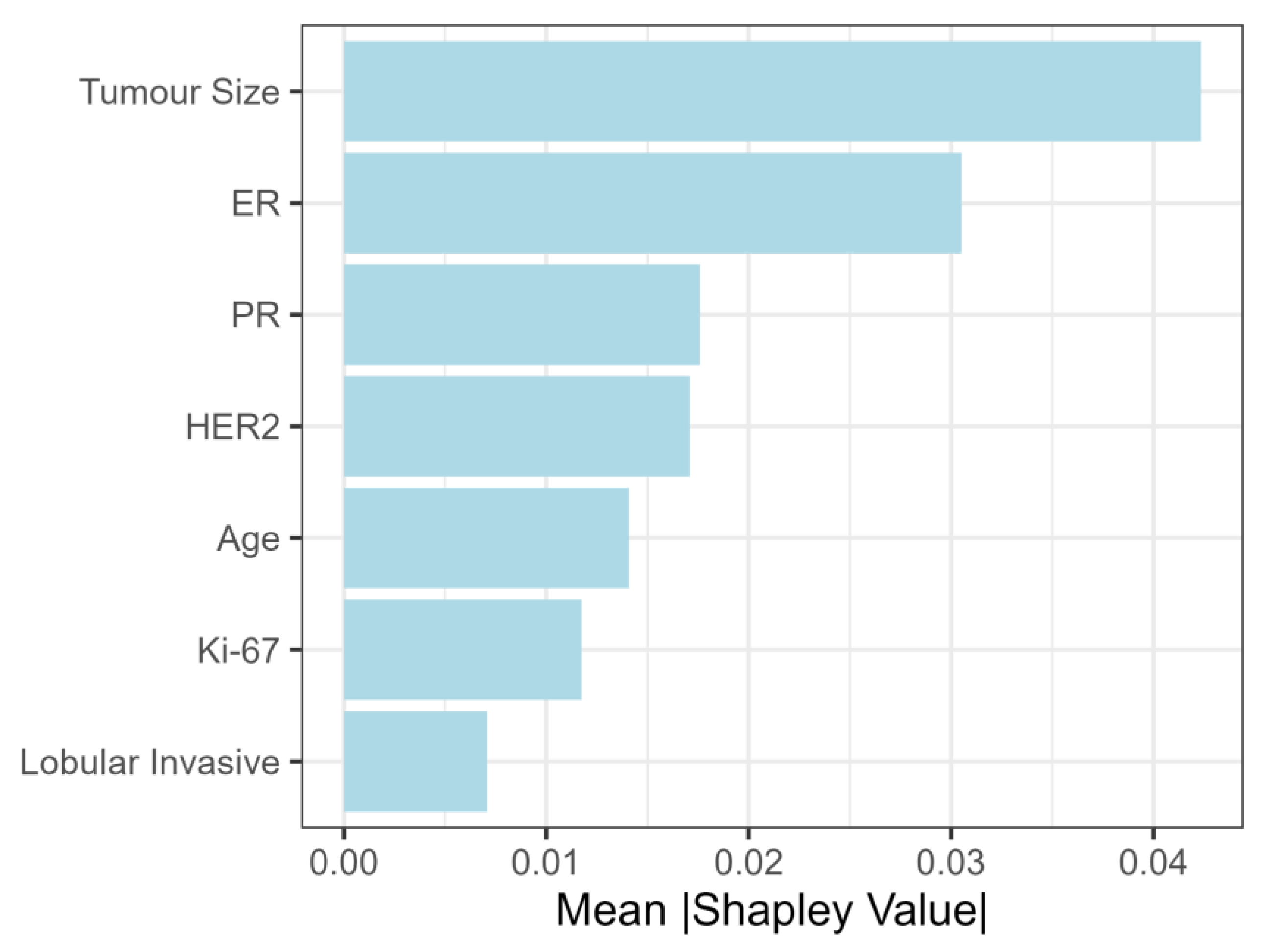
2.4. Feature Importance Analysis and Model Explainability
2.5. Statistical Analysis
3. Results
3.1. Patient Characteristics
3.2. Prediction Model Performance
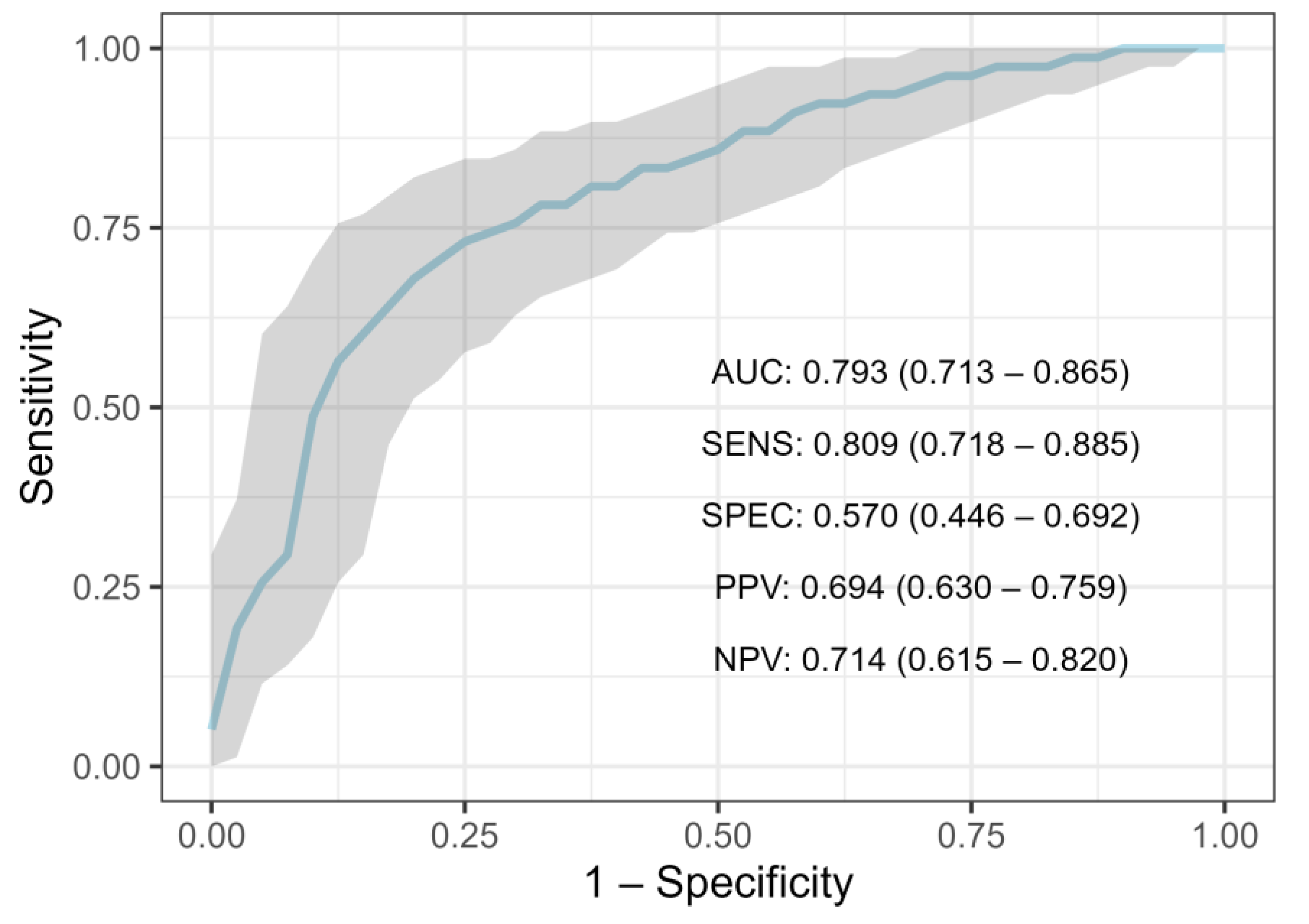
3.2.1. Performance on NST Criteria Group
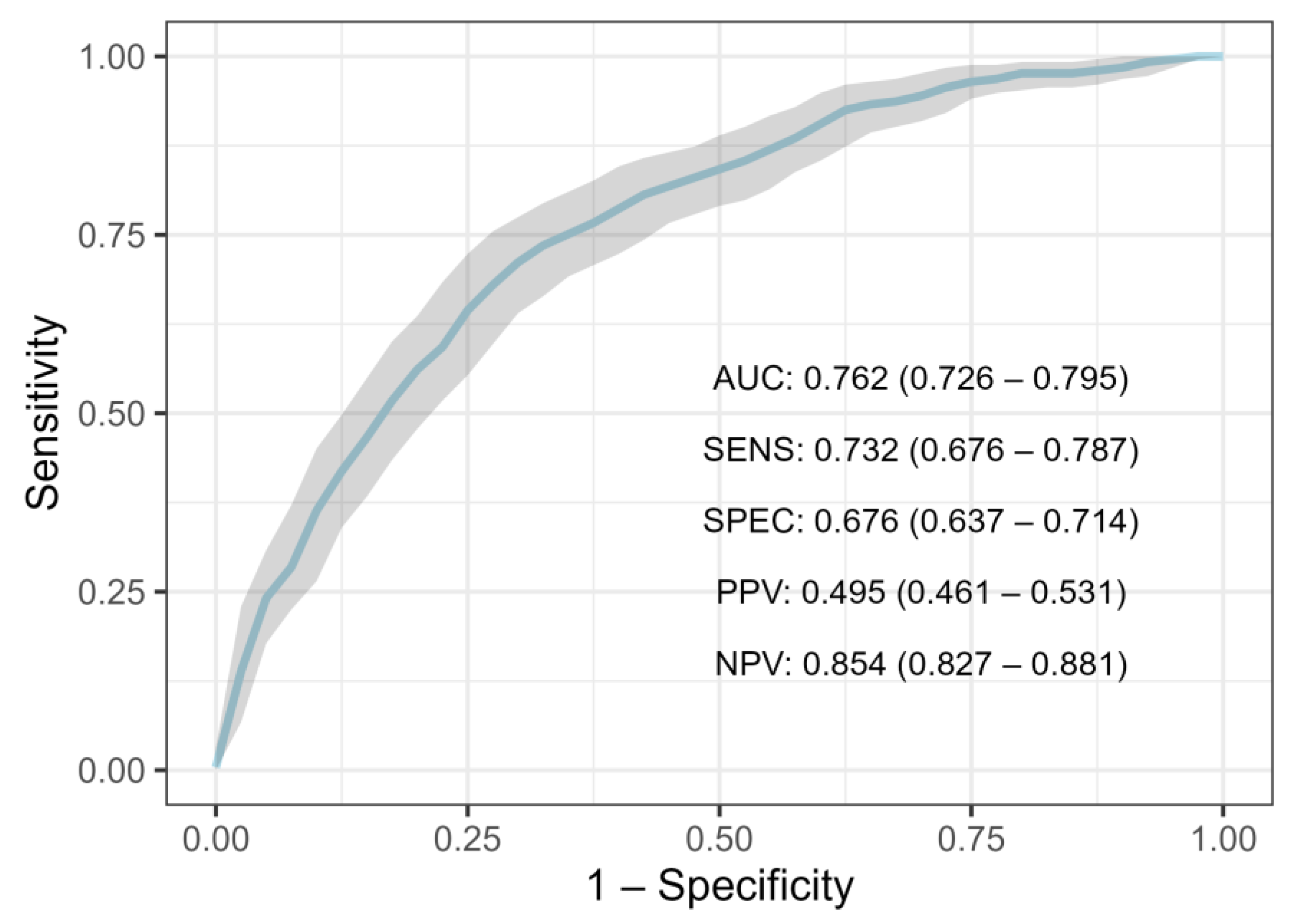
3.2.2. Performance on Entire Population
3.2.3. Feature Importance for Predicting Lymph Node Metastasis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Cancer Today. Available online: https://gco.iarc.fr/today/online-analysis-table?v=2020&mode=cancer&mode_population=continents&population=900&populations=900&key=asr&sex=2&cancer=39&type=1&statistic=5&prevalence=0&population_group=0&ages_group%5B%5D=0&ages_group%5B%5D=17&group_cancer=1&include_nmsc=0&include_nmsc_other=1 (accessed on 10 October 2022.).
- Centers for Disease Control and Prevention. Incidence and Relative Survival by Stage at Diagnosis for Common Cancers; USCS Data Brief, n.A., GA: Centers for Disease Control and Prevention, US Department of Health and Human Services: Washington, DC, USA, 2021. [Google Scholar]
- Nordenskjöld, A.E.; Fohlin, H.; Arnesson, L.G.; Einbeigi, Z.; Holmberg, E.; Albertsson, P.; Karlsson, P. Breast cancer survival trends in different stages and age groups—A population-based study 1989–2013. Acta Oncol. 2019, 58, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Walsh, M.F.; Nathanson, K.L.; Couch, F.J.; Offit, K. Genomic Biomarkers for Breast Cancer Risk. Adv. Exp. Med. Biol. 2016, 882, 1–32. [Google Scholar] [PubMed]
- Kim, J.Y.; Ryu, M.R.; Choi, B.O.; Park, W.C.; Oh, S.J.; Won, J.M.; Chung, S.M. The prognostic significance of the lymph node ratio in axillary lymph node positive breast cancer. J. Breast Cancer 2011, 14, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Gradishar, W.J.; Moran, M.S.; Abraham, J.; Aft, R.; Agnese, D.; Allison, K.H.; Anderson, B.; Burstein, H.J.; Chew, H.; Dang, C.; et al. Breast Cancer, Version 3.2022, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2022, 20, 691–722. [Google Scholar] [CrossRef]
- Cortazar, P.; Zhang, L.; Untch, M.; Mehta, K.; Costantino, J.P.; Wolmark, N.; Bonnefoi, H.; Cameron, D.; Gianni, L.; Valagussa, P.; et al. Pathological complete response and long-term clinical benefit in breast cancer: The CTNeoBC pooled analysis. Lancet 2014, 384, 164–172. [Google Scholar] [CrossRef]
- Symmans, W.F.; Wei, C.; Gould, R.; Yu, X.; Zhang, Y.; Liu, M.; Walls, A.; Bousamra, A.; Ramineni, M.; Sinn, B.; et al. Long-Term Prognostic Risk After Neoadjuvant Chemotherapy Associated With Residual Cancer Burden and Breast Cancer Subtype. J. Clin. Oncol. 2017, 35, 1049–1060. [Google Scholar] [CrossRef]
- Choi, H.Y.; Park, M.; Seo, M.; Song, E.; Shin, S.Y.; Sohn, Y.M. Preoperative Axillary Lymph Node Evaluation in Breast Cancer: Current Issues and Literature Review. Ultrasound Q. 2017, 33, 6–14. [Google Scholar] [CrossRef]
- Piccart, M.; Procter, M.; Fumagalli, D.; Azambuja, E.d.; Clark, E.; Ewer, M.S.; Restuccia, E.; Jerusalem, G.; Dent, S.; Reaby, L.; et al. Adjuvant Pertuzumab and Trastuzumab in Early HER2-Positive Breast Cancer in the APHINITY Trial: 6 Years’ Follow-Up. J. Clin. Oncol. 2021, 39, 1448–1457. [Google Scholar] [CrossRef]
- Kann, B.H.; Hosny, A.; Aerts, H. Artificial intelligence for clinical oncology. Cancer Cell 2021, 39, 916–927. [Google Scholar] [CrossRef]
- Boehm, K.M.; Khosravi, P.; Vanguri, R.; Gao, J.; Shah, S.P. Harnessing multimodal data integration to advance precision oncology. Nat. Rev. Cancer 2022, 22, 114–126. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, M.S.; Ahmad, W.; Alizadehsani, R.; Hussain, S.; Rehman, R. Breast Cancer Dataset, Classification and Detection Using Deep Learning. Healthcare 2022, 10, 2395. [Google Scholar] [CrossRef] [PubMed]
- Sarker, I.H. Machine Learning: Algorithms, Real-World Applications and Research Directions. SN Comput. Sci. 2021, 2, 160. [Google Scholar] [CrossRef] [PubMed]
- Kabiraj, S.; Raihan, M.; Alvi, N.; Afrin, M.; Akter, L.; Sohagi, S.A.; Podder, E. Breast Cancer Risk Prediction using XGBoost and Random Forest Algorithm. In Proceedings of the 2020 11th International Conference on Computing, Communication and Networking Technologies (ICCCNT), Kharagpur, India, 1–3 July 2020; pp. 1–4. [Google Scholar]
- Tseng, Y.J.; Huang, C.E.; Wen, C.N.; Lai, P.Y.; Wu, M.H.; Sun, Y.C.; Wang, H.Y.; Lu, J.J. Predicting breast cancer metastasis by using serum biomarkers and clinicopathological data with machine learning technologies. Int. J. Med. Inform. 2019, 128, 79–86. [Google Scholar] [CrossRef]
- Paik, E.S.; Lee, J.W.; Park, J.Y.; Kim, J.H.; Kim, M.; Kim, T.J.; Choi, C.H.; Kim, B.G.; Bae, D.S.; Seo, S.W. Prediction of survival outcomes in patients with epithelial ovarian cancer using machine learning methods. J. Gynecol. Oncol. 2019, 30, e65. [Google Scholar] [CrossRef]
- Li, Q.; Yang, H.; Wang, P.; Liu, X.; Lv, K.; Ye, M. XGBoost-based and tumor-immune characterized gene signature for the prediction of metastatic status in breast cancer. J. Transl. Med. 2022, 20, 177. [Google Scholar] [CrossRef]
- Altameem, A.; Mahanty, C.; Poonia, R.C.; Saudagar, A.K.J.; Kumar, R. Breast Cancer Detection in Mammography Images Using Deep Convolutional Neural Networks and Fuzzy Ensemble Modeling Techniques. Diagnostics 2022, 12, 1812. [Google Scholar] [CrossRef]
- Muduli, D.; Dash, R.; Majhi, B. Automated diagnosis of breast cancer using multi-modal datasets: A deep convolution neural network based approach. Biomed. Signal Process. Control 2022, 71, 102825. [Google Scholar] [CrossRef]
- Wakili, M.A.; Shehu, H.A.; Sharif, M.H.; Sharif, M.H.U.; Umar, A.; Kusetogullari, H.; Ince, I.F.; Uyaver, S. Classification of Breast Cancer Histopathological Images Using DenseNet and Transfer Learning. Comput. Intell. Neurosci. 2022, 10, 8904768. [Google Scholar] [CrossRef]
- Heenaye-Mamode Khan, M.; Boodoo-Jahangeer, N.; Dullull, W.; Nathire, S.; Gao, X.; Sinha, G.R.; Nagwanshi, K.K. Multi- class classification of breast cancer abnormalities using Deep Convolutional Neural Network (CNN). PLoS ONE 2021, 16, e0256500. [Google Scholar] [CrossRef]
- Hammond, M.E. ASCO-CAP guidelines for breast predictive factor testing: An update. Appl. Immunohistochem. Mol. Morphol. 2011, 19, 499–500. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R., Jr. The International Conference on Harmonization Good Clinical Practice guideline. Qual. Assur. 1998, 6, 65–74. [Google Scholar] [CrossRef]
- World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. JAMA 2013, 310, 2191–2194. [CrossRef] [PubMed]
- Ahn, S.J.; Kim, Y.S.; Kim, E.Y.; Park, H.K.; Cho, E.K.; Kim, Y.K.; Sung, Y.M.; Choi, H.Y. The value of chest CT for prediction of breast tumor size: Comparison with pathology measurement. World J. Surg. Oncol. 2013, 11, 1477–7819. [Google Scholar] [CrossRef] [PubMed]
- Cortadellas, T.; Argacha, P.; Acosta, J.; Rabasa, J.; Peiró, R.; Gomez, M.; Rodellar, L.; Gomez, S.; Navarro-Golobart, A.; Sanchez-Mendez, S.; et al. Estimation of tumor size in breast cancer comparing clinical examination, mammography, ultrasound and MRI—Correlation with the pathological analysis of the surgical specimen. Gland. Surg. 2017, 6, 330–335. [Google Scholar] [CrossRef]
- Hyder, T.; Bhattacharya, S.; Gade, K.; Nasrazadani, A.; Brufsky, A.M. Approaching Neoadjuvant Therapy in the Management of Early-Stage Breast Cancer. Breast Cancer 2021, 13, 199–211. [Google Scholar] [CrossRef]
- Korde, L.A.; Somerfield, M.R.; Carey, L.A.; Crews, J.R.; Denduluri, N.; Hwang, E.S.; Khan, S.A.; Loibl, S.; Morris, E.A.; Perez, A.; et al. Neoadjuvant Chemotherapy, Endocrine Therapy, and Targeted Therapy for Breast Cancer: ASCO Guideline. J. Clin. Oncol. 2021, 39, 1485–1505. [Google Scholar] [CrossRef]
- Grinsztajn, L.; Oyallon, E.; Varoquaux, G. Why do tree-based models still outperform deep learning on tabular data? arXiv 2022, arXiv:2207.08815. [Google Scholar]
- Shwartz-Ziv, R.; Armon, A. Tabular data: Deep learning is not all you need. Inf. Fusion 2022, 81, 84–90. [Google Scholar] [CrossRef]
- Blair, D.C. Information Retrieval, 2nd ed. C.J. Van Rijsbergen. London: Butterworths; 1979: 208 pp. Price: $32.50. J. Am. Soc. Inf. Sci. 1979, 30, 374–375. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.-I. A unified approach to interpreting model predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 4768–4777. [Google Scholar]
- Pesek, S.; Ashikaga, T.; Krag, L.E.; Krag, D. The false-negative rate of sentinel node biopsy in patients with breast cancer: A meta-analysis. World J. Surg. 2012, 36, 2239–2251. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Jun, Z.; Zhi-Cheng, G.; Xiang, Q. Factors that affect the false negative rate of sentinel lymph node mapping with methylene blue dye alone in breast cancer. J. Int. Med. Res. 2019, 47, 4841–4853. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Bentéjac, C.; Csörgő, A.; Martínez-Muñoz, G. A Comparative Analysis of XGBoost. arXiv 2019, arXiv:1911.01914. [Google Scholar]
- Sopik, V.; Narod, S.A. The relationship between tumour size, nodal status and distant metastases: On the origins of breast cancer. Breast Cancer Res. Treat. 2018, 170, 647–656. [Google Scholar] [CrossRef]
- Alsumai, T.S.; Alhazzaa, N.; Alshamrani, A.; Assiri, S.; Alhefdhi, A. Factors Predicting Positive Sentinel Lymph Node Biopsy in Clinically Node-Negative Breast Cancer. Breast Cancer 2022, 14, 323–334. [Google Scholar] [CrossRef]
- Yoshihara, E.; Smeets, A.; Laenen, A.; Reynders, A.; Soens, J.; Van Ongeval, C.; Moerman, P.; Paridaens, R.; Wildiers, H.; Neven, P.; et al. Predictors of axillary lymph node metastases in early breast cancer and their applicability in clinical practice. Breast 2013, 22, 357–361. [Google Scholar] [CrossRef]
- Wildiers, H.; Van Calster, B.; van de Poll-Franse, L.V.; Hendrickx, W.; Røislien, J.; Smeets, A.; Paridaens, R.; Deraedt, K.; Leunen, K.; Weltens, C.; et al. Relationship between age and axillary lymph node involvement in women with breast cancer. J. Clin. Oncol. 2009, 27, 2931–2937. [Google Scholar] [CrossRef]
- Rivadeneira, D.E.; Simmons, R.M.; Christos, P.J.; Hanna, K.; Daly, J.M.; Osborne, M.P. Predictive factors associated with axillary lymph node metastases in T1a and T1b breast carcinomas: Analysis in more than 900 patients. J. Am. Coll. Surg. 2000, 191, 1–6. [Google Scholar] [CrossRef]
- Gajdos, C.; Tartter, P.I.; Bleiweiss, I.J. Lymphatic invasion, tumor size, and age are independent predictors of axillary lymph node metastases in women with T1 breast cancers. Ann. Surg. 1999, 230, 692–696. [Google Scholar] [CrossRef]
- Yin, Y.; Zeng, K.; Wu, M.; Ding, Y.; Zhao, M.; Chen, Q. The levels of Ki-67 positive are positively associated with lymph node metastasis in invasive ductal breast cancer. Cell Biochem. Biophys. 2014, 70, 1145–1151. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Xu, H.; Zhang, H.; Ou, X.; Xu, Z.; Ai, L.; Sun, L.; Liu, C. Nomogram for prediction of level 2 axillary lymph node metastasis in proven level 1 node-positive breast cancer patients. Oncotarget 2017, 8, 72389–72399. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Zheng, T.; Wang, Y.; Li, Z.; Xiao, Q.; He, J.; Tan, J. Development of a prediction model based on LASSO regression to evaluate the risk of non-sentinel lymph node metastasis in Chinese breast cancer patients with 1-2 positive sentinel lymph nodes. Sci. Rep. 2021, 11, 19972. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Tan, Y.; Xie, C.; Hu, Q.; Ouyang, J.; Chen, Y.; Gu, Y.; Li, A.; Lu, N.; He, Z.; et al. Development and Validation of a Preoperative Magnetic Resonance Imaging Radiomics-Based Signature to Predict Axillary Lymph Node Metastasis and Disease-Free Survival in Patients With Early-Stage Breast Cancer. JAMA Netw. Open 2020, 3, 28086. [Google Scholar] [CrossRef]
- Pesapane, F.; Rotili, A.; Agazzi, G.M.; Botta, F.; Raimondi, S.; Penco, S.; Dominelli, V.; Cremonesi, M.; Jereczek-Fossa, B.A.; Carrafiello, G.; et al. Recent Radiomics Advancements in Breast Cancer: Lessons and Pitfalls for the Next Future. Curr. Oncol. 2021, 28, 2351–2372. [Google Scholar] [CrossRef]
- Guo, X.; Liu, Z.; Sun, C.; Zhang, L.; Wang, Y.; Li, Z.; Shi, J.; Wu, T.; Cui, H.; Zhang, J.; et al. Deep learning radiomics of ultrasonography: Identifying the risk of axillary non-sentinel lymph node involvement in primary breast cancer. EBioMedicine 2020, 60, 24. [Google Scholar] [CrossRef]
- Sannasi Chakravarthy, S.R.; Rajaguru, H. Automatic Detection and Classification of Mammograms Using Improved Extreme Learning Machine with Deep Learning. IRBM 2022, 43, 49–61. [Google Scholar] [CrossRef]
- Zheng, X.; Yao, Z.; Huang, Y.; Yu, Y.; Wang, Y.; Liu, Y.; Mao, R.; Li, F.; Xiao, Y.; Hu, Y.; et al. Deep learning radiomics can predict axillary lymph node status in early-stage breast cancer. Nat. Commun. 2020, 11, 020–15027. [Google Scholar] [CrossRef]
- Dihge, L.; Ohlsson, M.; Edén, P.; Bendahl, P.O.; Rydén, L. Artificial neural network models to predict nodal status in clinically node-negative breast cancer. BMC Cancer 2019, 19, 610. [Google Scholar] [CrossRef]
- Harris, G.C.; Denley, H.E.; Pinder, S.E.; Lee, A.H.; Ellis, I.O.; Elston, C.W.; Evans, A. Correlation of histologic prognostic factors in core biopsies and therapeutic excisions of invasive breast carcinoma. Am. J. Surg. Pathol. 2003, 27, 11–15. [Google Scholar] [CrossRef]
- Takada, M.; Sugimoto, M.; Naito, Y.; Moon, H.G.; Han, W.; Noh, D.Y.; Kondo, M.; Kuroi, K.; Sasano, H.; Inamoto, T.; et al. Prediction of axillary lymph node metastasis in primary breast cancer patients using a decision tree-based model. BMC Med. Inform. Decis. Mak. 2012, 12, 1472–6947. [Google Scholar] [CrossRef] [PubMed]

| Variable | Cases, Lymph Node Metastasis Group (n = 2536) | Controls, Non-Lymph Node Metastasis Group (n = 5845) | Total (n = 8381) | p-Value |
|---|---|---|---|---|
| Age (range) | 63.6 (21–92) | 62.3 (25–89) | 62.7 (12.6) | 0.535 * |
| Tumor Size (cm) | 2.7 (1.9) | 1.7 (1.1) | 2.01 (1.5) | <0.001 * |
| Ki-67 | 29.7 (18.7) | 25.1 (18.4) | 26.5 (18.7) | <0.001 * |
| ER | 80.7 (33.7) | 83.1 (32.2) | 82.4 (32.7) | 0.064 * |
| PR | 50.8 (39.5) | 54.3 (39.4) | 53.2 (39.4) | <0.001 * |
| Tumor Grade (%) | <0.001 † | |||
| 1 | 376 (14.8) | 1638 (28) | 2014 (24) | |
| 2 | 1460 (57.6) | 3148 (53.8) | 4608 (54.9) | |
| 3 | 700 (27.6) | 1059 (18.2) | 1759 (20.9) | |
| HER-2 (%) | <0.001 * | |||
| 0 | 1092 (43.1) | 2776 (47.5) | 3868 (46.2) | |
| 1 | 815 (32.1) | 1953 (33.4) | 2768 (33) | |
| 2 | 344 (1.3) | 629 (10.8) | 914 (10.9) | |
| 3 | 285 (11.2) | 487 (8.3) | 831 (9.9) | |
| Histological Type (%) | <0.001 † | |||
| NOS-invasive | 2055 (81) | 4586 (78.5) | 6641 (79.2) | |
| Lobular Invasive | 324 (12.8) | 693 (11.9) | 1017 (12.1) | |
| Ca with Medullary Characteristics | 24 (0.9) | 47 (0.8) | 71 (0.8) | |
| Other (Rare Types) | 133 (5.2) | 519 (8.9) | 652 (7.8) | |
| Immunophenotype (%) | <0.001 † | |||
| Luminal B | 1517 (59.8) | 3154 (53.9) | 4671 (55.7) | |
| Luminal A | 429 (16.9) | 1628 (27.8) | 2057 (24.6) | |
| Luminal B-her2 | 310 (12.3) | 508 (8.8) | 818 (9.8) | |
| Triple Negative | 160 (6.3) | 407 (6.9) | 567 (6.7) | |
| HER2 Positive | 120 (4.7) | 148 (2.6) | 268 (3.2) |
| Variable | Cases, Lymph Node Metastasis Group (n = 392) | Controls, Non-Lymph Node Metastasis Group (n = 327) | Total (n = 719) | p-Value |
|---|---|---|---|---|
| Age (range) | 66.9 (21–92) | 64.2 (25–87) | 65.7 (14.6) | 0.016 * |
| Tumor Size (cm) | 5.7 (3.02) | 3.9 (2.5) | 4.9 (2.9) | <0.001 * |
| Ki-67 | 43.4 (23.3) | 50.6 (25.7) | 46.7 (24.7) | <0.001 * |
| ER | 40.01 (45.9) | 13.7 (32.6) | 28.1 (42.5) | <0.001 * |
| PR | 21.8 (35.04) | 7.51 (22.5) | 15.2 (30.6) | <0.001 * |
| Tumor Grade (%) | 0.027 † | |||
| 1 | 14 (3.6) | 8 (2.4) | 22 (3) | |
| 2 | 133 (33.9) | 97 (29.6) | 230 (32) | |
| 3 | 245 (62.5) | 222 (68) | 467 (65) | |
| HER-2 (%) | 0.060 * | |||
| 0 | 180 (46) | 173 (53) | 353 (49.1) | |
| 1 | 79 (20.1) | 55 (16.8) | 134 (18.7) | |
| 2 | 35 (8.9) | 42 (12.8) | 77 (10.7) | |
| 3 | 98 (25) | 57 (17.4) | 155 (21.5) | |
| Histological Type (%) | <0.001 † | |||
| NOS-invasive | 282 (71.9) | 252 (77.2) | 534 (74.3) | |
| Lobular Invasive | 63 (16.1) | 20 (6.1) | 83 (11.5) | |
|
Ca with Medullary Characteristics | 15 (3.8) | 12 (3.6) | 27 (3.7) | |
| Other (Rare Types) | 32 (8.2) | 43 (13.1) | 75 (10.4%) | |
| Immunophenotype (%) | <0.001 † | |||
| Luminal B | 111 (28.4) | 35 (10.7) | 146 (20.3) | |
| Luminal A | 26 (6.6) | 7 (2.3) | 33 (4.6) | |
| Luminal B-her2 | 35 (8.9) | 8 (2.4) | 43 (5.9) | |
| >Triple Negative | 137 (34.9) | 210 (64.2) | 347 (48.4) | |
| HER-2 Positive | 83 (21.2) | 67 (20.4) | 150 (20.8) |
| Model | Mean AUC (95% CI) |
|---|---|
| Random Forest | 0.793 (0.713–0.865) |
| XGBoost | 0.783 (0.703–0.858) |
| Logistic Regression | 0.763 (0.683–0.838) |
| Univariate Logistic Regression | 0.645 (0.556–0.726) |
| Model | Mean AUC (95% CI) |
|---|---|
| XGBoost | 0.762 (0.726–0.795) |
| Random Forest | 0.760 (0.724–0.794) |
| Logistic Regression | 0.741 (0.706–0.775) |
| Univariate Logistic Regression | 0.713 (0.686–0.739) |
| Study (Algorithm Type) | Total Patients | Mean AUC (95% CI) |
|---|---|---|
| This study (XGBoost) | 8381 | 0.76 (0.73–0.80) |
| Takada et al. [55] (ADTree) | 467 | 0.77 (0.69–0.86) |
| Zheng et al. [52] (without radiomics, neural network) | 1342 | 0.72 (0.63–0.82) |
| Dihge et al. [53] (neural network) | 800 | 0.74 (0.72–0.76) |
| Meng et al. [47] (non-sentinel lymph node prediction, Lasso regression) | 714 | 0.77 (0.69–0.86) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vrdoljak, J.; Boban, Z.; Barić, D.; Šegvić, D.; Kumrić, M.; Avirović, M.; Perić Balja, M.; Periša, M.M.; Tomasović, Č.; Tomić, S.; et al. Applying Explainable Machine Learning Models for Detection of Breast Cancer Lymph Node Metastasis in Patients Eligible for Neoadjuvant Treatment. Cancers 2023, 15, 634. https://doi.org/10.3390/cancers15030634
Vrdoljak J, Boban Z, Barić D, Šegvić D, Kumrić M, Avirović M, Perić Balja M, Periša MM, Tomasović Č, Tomić S, et al. Applying Explainable Machine Learning Models for Detection of Breast Cancer Lymph Node Metastasis in Patients Eligible for Neoadjuvant Treatment. Cancers. 2023; 15(3):634. https://doi.org/10.3390/cancers15030634
Chicago/Turabian StyleVrdoljak, Josip, Zvonimir Boban, Domjan Barić, Darko Šegvić, Marko Kumrić, Manuela Avirović, Melita Perić Balja, Marija Milković Periša, Čedna Tomasović, Snježana Tomić, and et al. 2023. "Applying Explainable Machine Learning Models for Detection of Breast Cancer Lymph Node Metastasis in Patients Eligible for Neoadjuvant Treatment" Cancers 15, no. 3: 634. https://doi.org/10.3390/cancers15030634
APA StyleVrdoljak, J., Boban, Z., Barić, D., Šegvić, D., Kumrić, M., Avirović, M., Perić Balja, M., Periša, M. M., Tomasović, Č., Tomić, S., Vrdoljak, E., & Božić, J. (2023). Applying Explainable Machine Learning Models for Detection of Breast Cancer Lymph Node Metastasis in Patients Eligible for Neoadjuvant Treatment. Cancers, 15(3), 634. https://doi.org/10.3390/cancers15030634








