Krüppel-like Factor-9 and Krüppel-like Factor-13: Highly Related, Multi-Functional, Transcriptional Repressors and Activators of Oncogenesis
Abstract
Simple Summary
Abstract
1. Introduction
2. General Aspects of KLF9 and KLF13
2.1. Selected Highlights in the KLF9/KLF13 Field
2.1.1. KLF9
2.1.2. KLF13
2.2. Oncogenic Features of KLF9 and KLF13
2.3. Hormonal and Other Regulators of KLF9 and KLF13 Genes
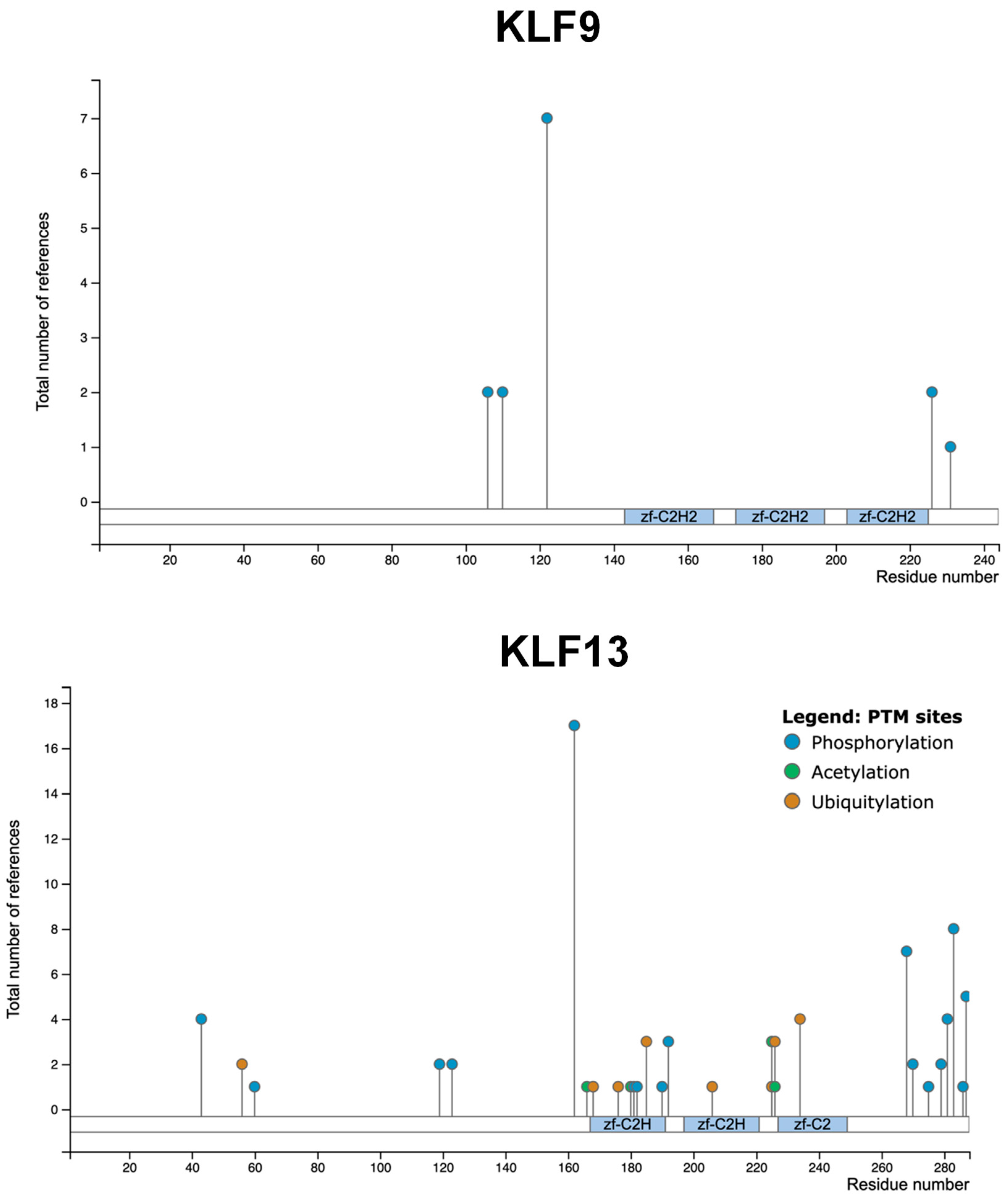
2.4. KLF9 and KLF13 Post-Translational Modifications and Interactomes
2.5. Epigenomics of KLF9 and KLF13
2.6. Redundancy, Overlapping Actions, and Functional Networks of KLF9 and KLF13
2.7. Known Pathways Subserving KLF9 and/or KLF13 Actions
2.8. KLF9 and KLF13 in Oxidative Stress (OS)
2.9. KLF9, KLF13 and Tissue Fibrosis
2.10. KLF9 and Cancer Stem Cells
3. Cancer-Specific Aspects of KLF9 and KLF13
3.1. Bladder Cancer
3.2. Breast Cancer
3.3. Cervical Cancer
3.4. Colorectal Cancer
3.5. Cutaneous Squamous Cell Carcinoma
3.6. Endometrial Cancer
3.7. Esophageal Squamous Cell Carcinoma (ESCC)
3.8. Gastric Cancer (GC)
3.9. Gliomas and Glioblastoma (GBM)
3.10. Hepatocellular Carcinoma (HCC)
3.11. Kidney Cancer (Renal Cell Carcinoma; RCC)
3.12. Melanoma
3.13. Multiple Myeloma (MM)
3.14. Nasopharyngeal Carcinoma (NPC)
3.15. Neuroblastoma (NB)
3.16. Non-Small Cell Lung Cancer (NSCLC)
3.17. Oral Squamous Cell Carcinoma (OSCC)
3.18. Osteosarcoma (OS)
3.19. Ovarian Cancer
3.20. Pancreatic Cancer
3.21. Papillary Thyroid Cancer (PTC)
3.22. Pedatric Acute Lymphoblastic Leukemia (ALL)
3.23. Prostate Cancer
3.24. Testicular Seminoma
4. Gaps in Knowledge and Future Directions
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Kadonaga, J.T.; Carner, K.R.; Masiarz, F.R.; Tjian, R. Isolation of cDNA Encoding Transcription Factor Sp1 and Functional Analysis of the DNA Binding Domain. Cell 1987, 51, 1079–1090. [Google Scholar] [CrossRef] [PubMed]
- Pei, J.; Grishin, N.V. A New Family of Predicted Krüppel-like Factor Genes and Pseudogenes in Placental Mammals. PLoS ONE 2013, 8, e81109. [Google Scholar] [CrossRef] [PubMed]
- Simmen, R.C.M.; Heard, M.E.; Simmen, A.M.; Montales, M.T.M.; Marji, M.; Scanlon, S.; Pabona, J.M.P. The Krüppel-like Factors in Female Reproductive System Pathologies. J. Mol. Endocrinol. 2015, 54, R89–R101. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.-K.; He, P.; Bialkowska, A.B.; Yang, V.W. SP and KLF Transcription Factors in Digestive Physiology and Diseases. Gastroenterology 2017, 152, 1845–1875. [Google Scholar] [CrossRef] [PubMed]
- Presnell, J.S.; Schnitzler, C.E.; Browne, W.E. KLF/SP Transcription Factor Family Evolution: Expansion, Diversification, and Innovation in Eukaryotes. Genome Biol. Evol. 2015, 7, 2289–2309. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Pearce, M.; Tivey, A.R.N.; Basutkar, P.; Lee, J.; Edbali, O.; Madhusoodanan, N.; Kolesnikov, A.; Lopez, R. Search and Sequence Analysis Tools Services from EMBL-EBI in 2022. Nucleic Acids Res. 2022, 50, W276–W279. [Google Scholar] [CrossRef] [PubMed]
- Oishi, Y.; Manabe, I. Krüppel-Like Factors in Metabolic Homeostasis and Cardiometabolic Disease. Front. Cardiovasc. Med. 2018, 5, 69. [Google Scholar] [CrossRef]
- Hsieh, P.N.; Fan, L.; Sweet, D.R.; Jain, M.K. The Krüppel-Like Factors and Control of Energy Homeostasis. Endocr. Rev. 2019, 40, 137–152. [Google Scholar] [CrossRef]
- Zakeri, S.; Aminian, H.; Sadeghi, S.; Esmaeilzadeh-Gharehdaghi, E.; Razmara, E. Krüppel-like Factors in Bone Biology. Cell Signal 2022, 93, 110308. [Google Scholar] [CrossRef]
- Abe, M.; Saeki, N.; Ikeda, Y.; Ohba, S. Kruppel-like Factors in Skeletal Physiology and Pathologies. Int. J. Mol. Sci. 2022, 23, 15174. [Google Scholar] [CrossRef]
- García-Niño, W.R.; Zazueta, C. New Insights of Krüppel-like Transcription Factors in Adipogenesis and the Role of Their Regulatory Neighbors. Life Sci. 2021, 265, 118763. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, G.; Feng, L.; Lu, H.; Wang, X. Krüppel-like Factors in Breast Cancer: Function, Regulation and Clinical Relevance. Biomed. Pharmacother. 2020, 123, 109778. [Google Scholar] [CrossRef] [PubMed]
- Orzechowska-Licari, E.J.; LaComb, J.F.; Mojumdar, A.; Bialkowska, A.B. SP and KLF Transcription Factors in Cancer Metabolism. Int. J. Mol. Sci. 2022, 23, 9956. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yao, C.; Ju, Z.; Jiao, D.; Hu, D.; Qi, L.; Liu, S.; Wu, X.; Zhao, C. Krüppel-like Factors in Tumors: Key Regulators and Therapeutic Avenues. Front. Oncol. 2023, 13, 1080720. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Li, L.; Zheng, X.-L.; Yin, W.-D.; Tang, C.-K. The Role of Krüppel-like Factor 14 in the Pathogenesis of Atherosclerosis. Atherosclerosis 2017, 263, 352–360. [Google Scholar] [CrossRef]
- Park, C.S.; Lewis, A.; Chen, T.; Lacorazza, D. Concise Review: Regulation of Self-Renewal in Normal and Malignant Hematopoietic Stem Cells by Krüppel-Like Factor 4. Stem Cells Transl. Med. 2019, 8, 568–574. [Google Scholar] [CrossRef]
- Taracha-Wisniewska, A.; Kotarba, G.; Dworkin, S.; Wilanowski, T. Recent Discoveries on the Involvement of Krüppel-Like Factor 4 in the Most Common Cancer Types. Int. J. Mol. Sci. 2020, 21, 8843. [Google Scholar] [CrossRef]
- Imataka, H.; Sogawa, K.; Yasumoto, K.; Kikuchi, Y.; Sasano, K.; Kobayashi, A.; Hayami, M.; Fujii-Kuriyama, Y. Two Regulatory Proteins That Bind to the Basic Transcription Element (BTE), a GC Box Sequence in the Promoter Region of the Rat P-4501A1 Gene. EMBO J. 1992, 11, 3663–3671. [Google Scholar] [CrossRef]
- Ohe, N.; Yamasaki, Y.; Sogawa, K.; Inazawa, J.; Ariyama, T.; Oshimura, M.; Fujii-Kuriyama, Y. Chromosomal Localization and cDNA Sequence of Human BTEB, a GC Box Binding Protein. Somat. Cell Mol. Genet. 1993, 19, 499–503. [Google Scholar] [CrossRef]
- Imataka, H.; Mizuno, A.; Fujii-Kuriyama, Y.; Hayami, M. Activation of the Human Immunodeficiency Virus Type 1 Long Terminal Repeat by BTEB, a GC Box-Binding Transcription Factor. AIDS Res. Hum. Retroviruses 1993, 9, 825–831. [Google Scholar] [CrossRef]
- Kobayashi, A.; Sogawa, K.; Imataka, H.; Fujii-Kuriyama, Y. Analysis of Functional Domains of a GC Box-Binding Protein, BTEB. J. Biochem. 1995, 117, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Michel, F.J.; Wing, A.; Simmen, F.A.; Simmen, R.C. Cell-Type Expression, Immunolocalization, and Deoxyribonucleic Acid-Binding Activity of Basic Transcription Element Binding Transcription Factor, an Sp-Related Family Member, in Porcine Endometrium of Pregnancy. Biol. Reprod. 1997, 57, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Simmen, R.C.; Chung, T.E.; Imataka, H.; Michel, F.J.; Badinga, L.; Simmen, F.A. Trans-Activation Functions of the Sp-Related Nuclear Factor, Basic Transcription Element-Binding Protein, and Progesterone Receptor in Endometrial Epithelial Cells. Endocrinology 1999, 140, 2517–2525. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Denver, R.J.; Ouellet, L.; Furling, D.; Kobayashi, A.; Fujii-Kuriyama, Y.; Puymirat, J. Basic Transcription Element-Binding Protein (BTEB) Is a Thyroid Hormone-Regulated Gene in the Developing Central Nervous System. Evidence for a Role in Neurite Outgrowth. J. Biol. Chem. 1999, 274, 23128–23134. [Google Scholar] [CrossRef] [PubMed]
- Morita, M.; Kobayashi, A.; Yamashita, T.; Shimanuki, T.; Nakajima, O.; Takahashi, S.; Ikegami, S.; Inokuchi, K.; Yamashita, K.; Yamamoto, M.; et al. Functional Analysis of Basic Transcription Element Binding Protein by Gene Targeting Technology. Mol. Cell. Biol. 2003, 23, 2489–2500. [Google Scholar] [CrossRef] [PubMed]
- Velarde, M.C.; Geng, Y.; Eason, R.R.; Simmen, F.A.; Simmen, R.C.M. Null Mutation of Kruppel-like Factor9/Basic Transcription Element Binding Protein-1 Alters Peri-Implantation Uterine Development in Mice. Biol. Reprod. 2005, 73, 472–481. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Scobie, K.N.; Hall, B.J.; Wilke, S.A.; Klemenhagen, K.C.; Fujii-Kuriyama, Y.; Ghosh, A.; Hen, R.; Sahay, A. Krüppel-like Factor 9 Is Necessary for Late-Phase Neuronal Maturation in the Developing Dentate Gyrus and during Adult Hippocampal Neurogenesis. J. Neurosci. 2009, 29, 9875–9887. [Google Scholar] [CrossRef] [PubMed]
- Besnard, A.; Langberg, T.; Levinson, S.; Chu, D.; Vicidomini, C.; Scobie, K.N.; Dwork, A.J.; Arango, V.; Rosoklija, G.B.; Mann, J.J.; et al. Targeting Kruppel-like Factor 9 in Excitatory Neurons Protects against Chronic Stress-Induced Impairments in Dendritic Spines and Fear Responses. Cell Rep. 2018, 23, 3183–3196. [Google Scholar] [CrossRef]
- Guo, N.; McDermott, K.D.; Shih, Y.-T.; Zanga, H.; Ghosh, D.; Herber, C.; Meara, W.R.; Coleman, J.; Zagouras, A.; Wong, L.P.; et al. Transcriptional Regulation of Neural Stem Cell Expansion in the Adult Hippocampus. eLife 2022, 11, e72195. [Google Scholar] [CrossRef]
- Zucker, S.N.; Fink, E.E.; Bagati, A.; Mannava, S.; Bianchi-Smiraglia, A.; Bogner, P.N.; Wawrzyniak, J.A.; Foley, C.; Leonova, K.I.; Grimm, M.J.; et al. Nrf2 Amplifies Oxidative Stress via Induction of Klf9. Mol. Cell 2014, 53, 916–928. [Google Scholar] [CrossRef]
- Simmen, F.A.; Su, Y.; Xiao, R.; Zeng, Z.; Simmen, R.C.M. The Krüppel-like Factor 9 (KLF9) Network in HEC-1-A Endometrial Carcinoma Cells Suggests the Carcinogenic Potential of Dys-Regulated KLF9 Expression. Reprod. Biol. Endocrinol. 2008, 6, 41. [Google Scholar] [CrossRef]
- Simmons, C.D.; Pabona, J.M.P.; Heard, M.E.; Friedman, T.M.; Spataro, M.T.; Godley, A.L.; Simmen, F.A.; Burnett, A.F.; Simmen, R.C.M. Krüppel-like Factor 9 Loss-of-Expression in Human Endometrial Carcinoma Links Altered Expression of Growth-Regulatory Genes with Aberrant Proliferative Response to Estrogen. Biol. Reprod. 2011, 85, 378–385. [Google Scholar] [CrossRef] [PubMed]
- Song, A.; Chen, Y.F.; Thamatrakoln, K.; Storm, T.A.; Krensky, A.M. RFLAT-1: A New Zinc Finger Transcription Factor That Activates RANTES Gene Expression in T Lymphocytes. Immunity 1999, 10, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Martin, K.M.; Cooper, W.N.; Metcalfe, J.C.; Kemp, P.R. Mouse BTEB3, a New Member of the Basic Transcription Element Binding Protein (BTEB) Family, Activates Expression from GC-Rich Minimal Promoter Regions. Biochem. J. 2000, 345 Pt 3, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Scohy, S.; Gabant, P.; Van Reeth, T.; Hertveldt, V.; Drèze, P.L.; Van Vooren, P.; Rivière, M.; Szpirer, J.; Szpirer, C. Identification of KLF13 and KLF14 (SP6), Novel Members of the SP/XKLF Transcription Factor Family. Genomics 2000, 70, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Song, A.; Patel, A.; Thamatrakoln, K.; Liu, C.; Feng, D.; Clayberger, C.; Krensky, A.M. Functional Domains and DNA-Binding Sequences of RFLAT-1/KLF13, a Krüppel-like Transcription Factor of Activated T Lymphocytes. J. Biol. Chem. 2002, 277, 30055–30065. [Google Scholar] [CrossRef]
- Martin, K.M.; Ellis, P.D.; Metcalfe, J.C.; Kemp, P.R. Selective Modulation of the SM22alpha Promoter by the Binding of BTEB3 (Basal Transcription Element-Binding Protein 3) to TGGG Repeats. Biochem. J. 2003, 375 Pt 2, 457–463. [Google Scholar] [CrossRef]
- Mitsuma, A.; Asano, H.; Kinoshita, T.; Murate, T.; Saito, H.; Stamatoyannopoulos, G.; Naoe, T. Transcriptional Regulation of FKLF-2 (KLF13) Gene in Erythroid Cells. Biochim. Biophys. Acta 2005, 1727, 125–133. [Google Scholar] [CrossRef]
- Zhang, P.; Basu, P.; Redmond, L.C.; Morris, P.E.; Rupon, J.W.; Ginder, G.D.; Lloyd, J.A. A Functional Screen for Krüppel-like Factors That Regulate the Human Gamma-Globin Gene through the CACCC Promoter Element. Blood Cells Mol. Dis. 2005, 35, 227–235. [Google Scholar] [CrossRef]
- Lavallée, G.; Andelfinger, G.; Nadeau, M.; Lefebvre, C.; Nemer, G.; Horb, M.E.; Nemer, M. The Kruppel-like Transcription Factor KLF13 Is a Novel Regulator of Heart Development. EMBO J. 2006, 25, 5201–5213. [Google Scholar] [CrossRef]
- Nemer, M.; Horb, M.E. The KLF Family of Transcriptional Regulators in Cardiomyocyte Proliferation and Differentiation. Cell Cycle 2007, 6, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; McPherson, L.; Feng, D.; Song, A.; Dong, C.; Lyu, S.-C.; Zhou, L.; Shi, X.; Ahn, Y.-T.; Wang, D.; et al. Kruppel-like Transcription Factor 13 Regulates T Lymphocyte Survival in Vivo. J. Immunol. 2007, 178, 5496–5504. [Google Scholar] [CrossRef] [PubMed]
- Outram, S.V.; Gordon, A.R.; Hager-Theodorides, A.L.; Metcalfe, J.; Crompton, T.; Kemp, P. KLF13 Influences Multiple Stages of Both B and T Cell Development. Cell Cycle 2008, 7, 2047–2055. [Google Scholar] [CrossRef] [PubMed]
- Gordon, A.R.; Outram, S.V.; Keramatipour, M.; Goddard, C.A.; Colledge, W.H.; Metcalfe, J.C.; Hager-Theodorides, A.L.; Crompton, T.; Kemp, P.R. Splenomegaly and Modified Erythropoiesis in KLF13-/- Mice. J. Biol. Chem. 2008, 283, 11897–11904. [Google Scholar] [CrossRef] [PubMed]
- Lai, D.; Zhu, J.; Wang, T.; Hu-Li, J.; Terabe, M.; Berzofsky, J.A.; Clayberger, C.; Krensky, A.M. KLF13 Sustains Thymic Memory-like CD8+ T Cells in BALB/c Mice by Regulating IL-4-Generating Invariant Natural Killer T Cells. J. Exp. Med. 2011, 208, 1093–1103. [Google Scholar] [CrossRef]
- Kwon, S.J.; Crespo-Barreto, J.; Zhang, W.; Wang, T.; Kim, D.S.; Krensky, A.; Clayberger, C. KLF13 Cooperates with C-Maf to Regulate IL-4 Expression in CD4+ T Cells. J. Immunol. 2014, 192, 5703–5709. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.-H.; Wang, J.; Guo, X.-J.; Gao, R.-F.; Yang, C.-X.; Li, L.; Sun, Y.-M.; Qiu, X.-B.; Xu, Y.-J.; Yang, Y.-Q. KLF13 Loss-of-Function Mutations Underlying Familial Dilated Cardiomyopathy. J. Am. Heart Assoc. 2022, 11, e027578. [Google Scholar] [CrossRef] [PubMed]
- Abhinav, P.; Zhang, G.-F.; Zhao, C.-M.; Xu, Y.-J.; Wang, J.; Yang, Y.-Q. A Novel KLF13 Mutation Underlying Congenital Patent Ductus Arteriosus and Ventricular Septal Defect, as Well as Bicuspid Aortic Valve. Exp. Ther. Med. 2022, 23, 311. [Google Scholar] [CrossRef]
- Henson, B.J.; Gollin, S.M. Overexpression of KLF13 and FGFR3 in Oral Cancer Cells. Cytogenet. Genome Res. 2010, 128, 192–198. [Google Scholar] [CrossRef]
- Martin, K.M.; Metcalfe, J.C.; Kemp, P.R. Expression of Klf9 and Klf13 in Mouse Development. Mech. Dev. 2001, 103, 149–151. [Google Scholar] [CrossRef]
- Zhang, Y.; Xue, Y.; Cao, C.; Huang, J.; Hong, Q.; Hai, T.; Jia, Q.; Wang, X.; Qin, G.; Yao, J.; et al. Thyroid Hormone Regulates Hematopoiesis via the TR-KLF9 Axis. Blood 2017, 130, 2161–2170. [Google Scholar] [CrossRef] [PubMed]
- Good, K.L.; Tangye, S.G. Decreased Expression of Kruppel-like Factors in Memory B Cells Induces the Rapid Response Typical of Secondary Antibody Responses. Proc. Natl. Acad. Sci. USA 2007, 104, 13420–13425. [Google Scholar] [CrossRef] [PubMed]
- Savignac, M.; Mellström, B.; Bébin, A.-G.; Oliveros, J.C.; Delpy, L.; Pinaud, E.; Naranjo, J.R. Increased B Cell Proliferation and Reduced Ig Production in DREAM Transgenic Mice. J. Immunol. 2010, 185, 7527–7536. [Google Scholar] [CrossRef] [PubMed]
- Cai, W.; Li, Y.; Cao, W. Prognostic Value and Immunological Role of Kruppel-like Transcription Factor 9 Gene in Pan-Carcinoma. Medicine 2022, 101, e32027. [Google Scholar] [CrossRef] [PubMed]
- Simmen, F.A.; Xiao, R.; Velarde, M.C.; Nicholson, R.D.; Bowman, M.T.; Fujii-Kuriyama, Y.; Oh, S.P.; Simmen, R.C.M. Dysregulation of Intestinal Crypt Cell Proliferation and Villus Cell Migration in Mice Lacking Kruppel-like Factor 9. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 292, G1757–G1769. [Google Scholar] [CrossRef] [PubMed]
- Li, S.S.-L.; Yu, S.-L.; Kao, L.-P.; Tsai, Z.Y.; Singh, S.; Chen, B.Z.; Ho, B.-C.; Liu, Y.-H.; Yang, P.-C. Target Identification of microRNAs Expressed Highly in Human Embryonic Stem Cells. J. Cell. Biochem. 2009, 106, 1020–1030. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, K.; Takehara, A.; Chiba, N.; Matsui, Y. Identification of KLF9 and BCL3 as Transcription Factors That Enhance Reprogramming of Primordial Germ Cells. PLoS ONE 2018, 13, e0205004. [Google Scholar] [CrossRef]
- Fernandez-Zapico, M.E.; Lomberk, G.A.; Tsuji, S.; DeMars, C.J.; Bardsley, M.R.; Lin, Y.-H.; Almada, L.L.; Han, J.-J.; Mukhopadhyay, D.; Ordog, T.; et al. A Functional Family-Wide Screening of SP/KLF Proteins Identifies a Subset of Suppressors of KRAS-Mediated Cell Growth. Biochem. J. 2011, 435, 529–537. [Google Scholar] [CrossRef]
- Mitchell, D.L.; DiMario, J.X. Bimodal, Reciprocal Regulation of Fibroblast Growth Factor Receptor 1 Promoter Activity by BTEB1/KLF9 during Myogenesis. Mol. Biol. Cell 2010, 21, 2780–2787. [Google Scholar] [CrossRef][Green Version]
- Lambert, S.A.; Yang, A.W.H.; Sasse, A.; Cowley, G.; Albu, M.; Caddick, M.X.; Morris, Q.D.; Weirauch, M.T.; Hughes, T.R. Similarity Regression Predicts Evolution of Transcription Factor Sequence Specificity. Nat. Genet. 2019, 51, 981–989. [Google Scholar] [CrossRef]
- Kaczynski, J.; Cook, T.; Urrutia, R. Sp1- and Krüppel-like Transcription Factors. Genome Biol. 2003, 4, 206. [Google Scholar] [CrossRef] [PubMed]
- Imataka, H.; Nakayama, K.; Yasumoto, K.; Mizuno, A.; Fujii-Kuriyama, Y.; Hayami, M. Cell-Specific Translational Control of Transcription Factor BTEB Expression. The Role of an Upstream AUG in the 5′-Untranslated Region. J. Biol. Chem. 1994, 269, 20668–20673. [Google Scholar] [CrossRef] [PubMed]
- Nikolcheva, T.; Pyronnet, S.; Chou, S.; Sonenberg, N.; Song, A.; Clayberger, C.; Krensky, A.M. A Translational Rheostat for RFLAT-1 Regulates RANTES Expression in T Lymphocytes. J. Clin. Investig. 2002, 110, 119–126. [Google Scholar] [CrossRef]
- Martel, J.; Cayrou, C.; Puymirat, J. Identification of New Thyroid Hormone-Regulated Genes in Rat Brain Neuronal Cultures. Neuroreport 2002, 13, 1849–1851. [Google Scholar] [CrossRef] [PubMed]
- Dugas, J.C.; Ibrahim, A.; Barres, B.A. The T3-Induced Gene KLF9 Regulates Oligodendrocyte Differentiation and Myelin Regeneration. Mol. Cell. Neurosci. 2012, 50, 45–57. [Google Scholar] [CrossRef] [PubMed]
- Avci, H.X.; Lebrun, C.; Wehrlé, R.; Doulazmi, M.; Chatonnet, F.; Morel, M.-P.; Ema, M.; Vodjdani, G.; Sotelo, C.; Flamant, F.; et al. Thyroid Hormone Triggers the Developmental Loss of Axonal Regenerative Capacity via Thyroid Hormone Receptor A1 and Krüppel-like Factor 9 in Purkinje Cells. Proc. Natl. Acad. Sci. USA 2012, 109, 14206–14211. [Google Scholar] [CrossRef] [PubMed]
- Gil-Ibáñez, P.; Bernal, J.; Morte, B. Thyroid Hormone Regulation of Gene Expression in Primary Cerebrocortical Cells: Role of Thyroid Hormone Receptor Subtypes and Interactions with Retinoic Acid and Glucocorticoids. PLoS ONE 2014, 9, e91692. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Abe, K.; Milanesi, A.; Liu, Y.-Y.; Brent, G.A. Thyroid Hormone Protects Primary Cortical Neurons Exposed to Hypoxia by Reducing DNA Methylation and Apoptosis. Endocrinology 2019, 160, 2243–2256. [Google Scholar] [CrossRef]
- Bonett, R.M.; Hu, F.; Bagamasbad, P.; Denver, R.J. Stressor and Glucocorticoid-Dependent Induction of the Immediate Early Gene Kruppel-like Factor 9: Implications for Neural Development and Plasticity. Endocrinology 2009, 150, 1757–1765. [Google Scholar] [CrossRef]
- Bagamasbad, P.; Ziera, T.; Borden, S.A.; Bonett, R.M.; Rozeboom, A.M.; Seasholtz, A.; Denver, R.J. Molecular Basis for Glucocorticoid Induction of the Kruppel-like Factor 9 Gene in Hippocampal Neurons. Endocrinology 2012, 153, 5334–5345. [Google Scholar] [CrossRef]
- Kennedy, C.L.M.; Price, E.M.; Mifsud, K.R.; Salatino, S.; Sharma, E.; Engledow, S.; Broxholme, J.; Goss, H.M.; Reul, J.M.H.M. Genomic Regulation of Krüppel-like-Factor Family Members by Corticosteroid Receptors in the Rat Brain. Neurobiol. Stress 2023, 23, 100532. [Google Scholar] [CrossRef] [PubMed]
- Shewade, L.H.; Schneider, K.A.; Brown, A.C.; Buchholz, D.R. In-Vivo Regulation of Krüppel-like Factor 9 by Corticosteroids and Their Receptors across Tissues in Tadpoles of Xenopus Tropicalis. Gen. Comp. Endocrinol. 2017, 248, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Mostafa, M.M.; Rider, C.F.; Shah, S.; Traves, S.L.; Gordon, P.M.K.; Miller-Larsson, A.; Leigh, R.; Newton, R. Glucocorticoid-Driven Transcriptomes in Human Airway Epithelial Cells: Commonalities, Differences and Functional Insight from Cell Lines and Primary Cells. BMC Med. Genom. 2019, 12, 29. [Google Scholar] [CrossRef] [PubMed]
- Mostafa, M.M.; Bansal, A.; Michi, A.N.; Sasse, S.K.; Proud, D.; Gerber, A.N.; Newton, R. Genomic Determinants Implicated in the Glucocorticoid-Mediated Induction of KLF9 in Pulmonary Epithelial Cells. J. Biol. Chem. 2021, 296, 100065. [Google Scholar] [CrossRef] [PubMed]
- Cui, A.; Fan, H.; Zhang, Y.; Zhang, Y.; Niu, D.; Liu, S.; Liu, Q.; Ma, W.; Shen, Z.; Shen, L.; et al. Dexamethasone-Induced Krüppel-like Factor 9 Expression Promotes Hepatic Gluconeogenesis and Hyperglycemia. J. Clin. Investig. 2019, 129, 2266–2278. [Google Scholar] [CrossRef]
- Chinenov, Y.; Coppo, M.; Gupte, R.; Sacta, M.A.; Rogatsky, I. Glucocorticoid Receptor Coordinates Transcription Factor-Dominated Regulatory Network in Macrophages. BMC Genom. 2014, 15, 656. [Google Scholar] [CrossRef] [PubMed]
- Ai, F.; Zhao, G.; Lv, W.; Liu, B.; Lin, J. Dexamethasone Induces Aberrant Macrophage Immune Function and Apoptosis. Oncol. Rep. 2020, 43, 427–436. [Google Scholar] [CrossRef]
- Spörl, F.; Korge, S.; Jürchott, K.; Wunderskirchner, M.; Schellenberg, K.; Heins, S.; Specht, A.; Stoll, C.; Klemz, R.; Maier, B.; et al. Krüppel-like Factor 9 Is a Circadian Transcription Factor in Human Epidermis That Controls Proliferation of Keratinocytes. Proc. Natl. Acad. Sci. USA 2012, 109, 10903–10908. [Google Scholar] [CrossRef]
- Lili, L.N.; Klopot, A.; Readhead, B.; Baida, G.; Dudley, J.T.; Budunova, I. Transcriptomic Network Interactions in Human Skin Treated with Topical Glucocorticoid Clobetasol Propionate. J. Investig. Dermatol. 2019, 139, 2281–2291. [Google Scholar] [CrossRef]
- Gans, I.; Hartig, E.I.; Zhu, S.; Tilden, A.R.; Hutchins, L.N.; Maki, N.J.; Graber, J.H.; Coffman, J.A. Klf9 Is a Key Feedforward Regulator of the Transcriptomic Response to Glucocorticoid Receptor Activity. Sci. Rep. 2020, 10, 11415. [Google Scholar] [CrossRef]
- Jing, D.; Bhadri, V.A.; Beck, D.; Thoms, J.A.I.; Yakob, N.A.; Wong, J.W.H.; Knezevic, K.; Pimanda, J.E.; Lock, R.B. Opposing Regulation of BIM and BCL2 Controls Glucocorticoid-Induced Apoptosis of Pediatric Acute Lymphoblastic Leukemia Cells. Blood 2015, 125, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Topete, D.; He, B.; Xu, X.; Cidlowski, J.A. Krüppel-like Factor 13 Is a Major Mediator of Glucocorticoid Receptor Signaling in Cardiomyocytes and Protects These Cells from DNA Damage and Death. J. Biol. Chem. 2016, 291, 19374–19386. [Google Scholar] [CrossRef] [PubMed]
- Velarde, M.C.; Iruthayanathan, M.; Eason, R.R.; Zhang, D.; Simmen, F.A.; Simmen, R.C.M. Progesterone Receptor Transactivation of the Secretory Leukocyte Protease Inhibitor Gene in Ishikawa Endometrial Epithelial Cells Involves Recruitment of Krüppel-like Factor 9/Basic Transcription Element Binding Protein-1. Endocrinology 2006, 147, 1969–1978. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Zhang, H.; Ke, J.; Zhang, Y.; Dai, C.; Zhu, M.; Jiang, F.; Zhu, H.; Zhang, L.; Zuo, X.; et al. Progesterone Receptor Inhibits the Proliferation and Invasion of Endometrial Cancer Cells by up Regulating Krüppel-like Factor 9. Transl. Cancer Res. 2020, 9, 2220–2230. [Google Scholar] [CrossRef]
- Du, H.; Sarno, J.; Taylor, H.S. HOXA10 Inhibits Kruppel-like Factor 9 Expression in the Human Endometrial Epithelium. Biol. Reprod. 2010, 83, 205–211. [Google Scholar] [CrossRef][Green Version]
- Shen, P.; Cao, X.; Sun, L.; Qian, Y.; Wu, B.; Wang, X.; Shi, G.; Wang, D. KLF9 Suppresses Cell Growth and Induces Apoptosis via the AR Pathway in Androgen-Dependent Prostate Cancer Cells. Biochem. Biophys. Rep. 2021, 28, 101151. [Google Scholar] [CrossRef]
- Song, C.-Z.; Keller, K.; Chen, Y.; Stamatoyannopoulos, G. Functional Interplay between CBP and PCAF in Acetylation and Regulation of Transcription Factor KLF13 Activity. J. Mol. Biol. 2003, 329, 207–215. [Google Scholar] [CrossRef]
- Apara, A.; Galvao, J.; Wang, Y.; Blackmore, M.; Trillo, A.; Iwao, K.; Brown, D.P.; Fernandes, K.A.; Huang, A.; Nguyen, T.; et al. KLF9 and JNK3 Interact to Suppress Axon Regeneration in the Adult CNS. J. Neurosci. 2017, 37, 9632–9644. [Google Scholar] [CrossRef]
- Bernhardt, C.; Sock, E.; Fröb, F.; Hillgärtner, S.; Nemer, M.; Wegner, M. KLF9 and KLF13 Transcription Factors Boost Myelin Gene Expression in Oligodendrocytes as Partners of SOX10 and MYRF. Nucleic Acids Res. 2022, 50, 11509–11528. [Google Scholar] [CrossRef]
- Pang, Y.-P.; Kumar, G.A.; Zhang, J.-S.; Urrutia, R. Differential Binding of Sin3 Interacting Repressor Domains to the PAH2 Domain of Sin3A. FEBS Lett. 2003, 548, 108–112. [Google Scholar] [CrossRef]
- Kaczynski, J.; Zhang, J.S.; Ellenrieder, V.; Conley, A.; Duenes, T.; Kester, H.; van Der Burg, B.; Urrutia, R. The Sp1-like Protein BTEB3 Inhibits Transcription via the Basic Transcription Element Box by Interacting with mSin3A and HDAC-1 Co-Repressors and Competing with Sp1. J. Biol. Chem. 2001, 276, 36749–36756. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Xiang, J.; Ma, C.; Yang, G.; Wang, X.; Liu, H.; Fan, G.; Kang, L.; Liang, Z. Sp1-like Protein KLF13 Acts as a Negative Feedback Regulator of TGF-β Signaling and Fibrosis. Cell Rep. 2023, 42, 112367. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.-Y.; Li, S.; Wang, M.; Li, X.; Yang, Y.; Xu, Z.; Li, B.; Li, Y.; Xia, K.; Chen, H.; et al. Krüppel-like Factor 9 down-Regulates Matrix Metalloproteinase 9 Transcription and Suppresses Human Breast Cancer Invasion. Cancer Lett. 2018, 412, 224–235. [Google Scholar] [CrossRef]
- Zhang, D.; Zhang, X.-L.; Michel, F.J.; Blum, J.L.; Simmen, F.A.; Simmen, R.C.M. Direct Interaction of the Krüppel-like Family (KLF) Member, BTEB1, and PR Mediates Progesterone-Responsive Gene Expression in Endometrial Epithelial Cells. Endocrinology 2002, 143, 62–73. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Tencer, A.; Xuan, H.; Kutateladze, T.G.; Shi, X. ZZEF1 Is a Histone Reader and Transcriptional Coregulator of Krüppel-Like Factors. J. Mol. Biol. 2021, 433, 166722. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Ahn, Y.-T.; McPherson, L.; Clayberger, C.; Krensky, A.M. Interaction of PRP4 with Kruppel-like Factor 13 Regulates CCL5 Transcription. J. Immunol. 2007, 178, 7081–7087. [Google Scholar] [CrossRef] [PubMed]
- Ahn, Y.-T.; Huang, B.; McPherson, L.; Clayberger, C.; Krensky, A.M. Dynamic Interplay of Transcriptional Machinery and Chromatin Regulates “Late” Expression of the Chemokine RANTES in T Lymphocytes. Mol. Cell. Biol. 2007, 27, 253–266. [Google Scholar] [CrossRef]
- Krensky, A.M.; Ahn, Y.-T. Mechanisms of Disease: Regulation of RANTES (CCL5) in Renal Disease. Nat. Clin. Pract. Nephrol. 2007, 3, 164–170. [Google Scholar] [CrossRef]
- Zhang, J.S.; Moncrieffe, M.C.; Kaczynski, J.; Ellenrieder, V.; Prendergast, F.G.; Urrutia, R. A Conserved Alpha-Helical Motif Mediates the Interaction of Sp1-like Transcriptional Repressors with the Corepressor mSin3A. Mol. Cell. Biol. 2001, 21, 5041–5049. [Google Scholar] [CrossRef]
- Ellenrieder, V.; Zhang, J.-S.; Kaczynski, J.; Urrutia, R. Signaling Disrupts mSin3A Binding to the Mad1-like Sin3-Interacting Domain of TIEG2, an Sp1-like Repressor. EMBO J. 2002, 21, 2451–2460. [Google Scholar] [CrossRef]
- Fernandez-Zapico, M.E.; Mladek, A.; Ellenrieder, V.; Folch-Puy, E.; Miller, L.; Urrutia, R. An mSin3A interaction domain links the transcriptional activity of KLF11 with its role in growth regulation. EMBO J. 2003, 22, 4748–4758. [Google Scholar] [CrossRef] [PubMed]
- Gowri, P.M.; Yu, J.H.; Shaufl, A.; Sperling, M.A.; Menon, R.K. Recruitment of a Repressosome Complex at the Growth Hormone Receptor Promoter and Its Potential Role in Diabetic Nephropathy. Mol. Cell. Biol. 2003, 23, 815–825. [Google Scholar] [CrossRef] [PubMed]
- Darwich, R.; Li, W.; Yamak, A.; Komati, H.; Andelfinger, G.; Sun, K.; Nemer, M. KLF13 Is a Genetic Modifier of the Holt-Oram Syndrome Gene TBX5. Hum. Mol. Genet. 2017, 26, 942–954. [Google Scholar] [CrossRef] [PubMed]
- Göös, H.; Kinnunen, M.; Salokas, K.; Tan, Z.; Liu, X.; Yadav, L.; Zhang, Q.; Wei, G.-H.; Varjosalo, M. Human Transcription Factor Protein Interaction Networks. Nat. Commun. 2022, 13, 766. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.-Y.; Hsu, S.-D.; Huang, H.-Y.; Sun, Y.-M.; Chou, C.-H.; Weng, S.-L.; Huang, H.-D. MethHC: A Database of DNA Methylation and Gene Expression in Human Cancer. Nucleic Acids Res. 2015, 43, D856–D861. [Google Scholar] [CrossRef]
- Huang, H.-Y.; Li, J.; Tang, Y.; Huang, Y.-X.; Chen, Y.-G.; Xie, Y.-Y.; Zhou, Z.-Y.; Chen, X.-Y.; Ding, S.-Y.; Luo, M.-F.; et al. MethHC 2.0: Information Repository of DNA Methylation and Gene Expression in Human Cancer. Nucleic Acids Res. 2021, 49, D1268–D1275. [Google Scholar] [CrossRef]
- Wang, L.; Mao, Q.; Zhou, S.; Ji, X. Hypermethylated KLF9 Is an Independent Prognostic Factor for Favorable Outcome in Breast Cancer. Onco Targets Ther. 2019, 12, 9915–9926. [Google Scholar] [CrossRef]
- Lafontaine, N.; Campbell, P.J.; Castillo-Fernandez, J.E.; Mullin, S.; Lim, E.M.; Kendrew, P.; Lewer, M.; Brown, S.J.; Huang, R.-C.; Melton, P.E.; et al. Epigenome-Wide Association Study of Thyroid Function Traits Identifies Novel Associations of fT3 With KLF9 and DOT1L. J. Clin. Endocrinol. Metab. 2021, 106, e2191–e2202. [Google Scholar] [CrossRef]
- Weihs, A.; Chaker, L.; Martin, T.C.; Braun, K.V.E.; Campbell, P.J.; Cox, S.R.; Fornage, M.; Gieger, C.; Grabe, H.J.; Grallert, H.; et al. Epigenome-Wide Association Study Reveals CpG Sites Associated with Thyroid Function and Regulatory Effects on KLF9. Thyroid 2023, 33, 301–311. [Google Scholar] [CrossRef]
- Alfano, R.; Zugna, D.; Barros, H.; Bustamante, M.; Chatzi, L.; Ghantous, A.; Herceg, Z.; Keski-Rahkonen, P.; de Kok, T.M.; Nawrot, T.S.; et al. Cord Blood Epigenome-Wide Meta-Analysis in Six European-Based Child Cohorts Identifies Signatures Linked to Rapid Weight Growth. BMC Med. 2023, 21, 17. [Google Scholar] [CrossRef]
- Wolf, I.; Bose, S.; Desmond, J.C.; Lin, B.T.; Williamson, E.A.; Karlan, B.Y.; Koeffler, H.P. Unmasking of Epigenetically Silenced Genes Reveals DNA Promoter Methylation and Reduced Expression of PTCH in Breast Cancer. Breast Cancer Res. Treat. 2007, 105, 139–155. [Google Scholar] [CrossRef] [PubMed]
- Khamas, A.; Ishikawa, T.; Shimokawa, K.; Mogushi, K.; Iida, S.; Ishiguro, M.; Mizushima, H.; Tanaka, H.; Uetake, H.; Sugihara, K. Screening for Epigenetically Masked Genes in Colorectal Cancer Using 5-Aza-2′-Deoxycytidine, Microarray and Gene Expression Profile. Cancer Genom. Proteom. 2012, 9, 67–75. [Google Scholar]
- Koh, I.-U.; Lee, H.-J.; Hwang, J.-Y.; Choi, N.-H.; Lee, S. Obesity-Related CpG Methylation (Cg07814318) of Kruppel-like Factor-13 (KLF13) Gene with Childhood Obesity and Its Cis-Methylation Quantitative Loci. Sci. Rep. 2017, 7, 45368. [Google Scholar] [CrossRef] [PubMed]
- Wiemerslage, L.; Islam, R.; van der Kamp, C.; Cao, H.; Olivo, G.; Ence-Eriksson, F.; Castillo, S.; Larsen, A.L.; Bandstein, M.; Dahlberg, L.S.; et al. A DNA Methylation Site within the KLF13 Gene Is Associated with Orexigenic Processes Based on Neural Responses and Ghrelin Levels. Int. J. Obes. 2017, 41, 990–994. [Google Scholar] [CrossRef]
- Wu, R.; Yun, Q.; Zhang, J.; Bao, J. Downregulation of KLF13 through DNMT1-Mediated Hypermethylation Promotes Glioma Cell Proliferation and Invasion. Onco Targets Ther. 2019, 12, 1509–1520. [Google Scholar] [CrossRef]
- Simmen, R.C.M.; Eason, R.R.; McQuown, J.R.; Linz, A.L.; Kang, T.-J.; Chatman, L.; Till, S.R.; Fujii-Kuriyama, Y.; Simmen, F.A.; Oh, S.P. Subfertility, Uterine Hypoplasia, and Partial Progesterone Resistance in Mice Lacking the Kruppel-like Factor 9/Basic Transcription Element-Binding Protein-1 (Bteb1) Gene. J. Biol. Chem. 2004, 279, 29286–29294. [Google Scholar] [CrossRef]
- Natesampillai, S.; Kerkvliet, J.; Leung, P.C.K.; Veldhuis, J.D. Regulation of Kruppel-like Factor 4, 9, and 13 Genes and the Steroidogenic Genes LDLR, StAR, and CYP11A in Ovarian Granulosa Cells. Am. J. Physiol. Endocrinol. Metab. 2008, 294, E385–E391. [Google Scholar] [CrossRef]
- Pabona, J.M.P.; Zeng, Z.; Simmen, F.A.; Simmen, R.C.M. Functional Differentiation of Uterine Stromal Cells Involves Cross-Regulation between Bone Morphogenetic Protein 2 and Kruppel-like Factor (KLF) Family Members KLF9 and KLF13. Endocrinology 2010, 151, 3396–3406. [Google Scholar] [CrossRef]
- Heard, M.E.; Pabona, J.M.P.; Clayberger, C.; Krensky, A.M.; Simmen, F.A.; Simmen, R.C.M. The Reproductive Phenotype of Mice Null for Transcription Factor Krüppel-like Factor 13 Suggests Compensatory Function of Family Member Krüppel-like Factor 9 in the Peri-Implantation Uterus. Biol. Reprod. 2012, 87, 115. [Google Scholar] [CrossRef]
- Heard, M.E.; Velarde, M.C.; Giudice, L.C.; Simmen, F.A.; Simmen, R.C.M. Krüppel-Like Factor 13 Deficiency in Uterine Endometrial Cells Contributes to Defective Steroid Hormone Receptor Signaling but Not Lesion Establishment in a Mouse Model of Endometriosis. Biol. Reprod. 2015, 92, 140. [Google Scholar] [CrossRef]
- Knoedler, J.R.; Subramani, A.; Denver, R.J. The Krüppel-like Factor 9 Cistrome in Mouse Hippocampal Neurons Reveals Predominant Transcriptional Repression via Proximal Promoter Binding. BMC Genom. 2017, 18, 299. [Google Scholar] [CrossRef] [PubMed]
- Knoedler, J.R.; Ávila-Mendoza, J.; Subramani, A.; Denver, R.J. The Paralogous Krüppel-like Factors 9 and 13 Regulate the Mammalian Cellular Circadian Clock Output Gene Dbp. J. Biol. Rhythms 2020, 35, 257–274. [Google Scholar] [CrossRef] [PubMed]
- Ávila-Mendoza, J.; Subramani, A.; Denver, R.J. Krüppel-Like Factors 9 and 13 Block Axon Growth by Transcriptional Repression of Key Components of the cAMP Signaling Pathway. Front. Mol. Neurosci. 2020, 13, 602638. [Google Scholar] [CrossRef]
- Ávila-Mendoza, J.; Subramani, A.; Sifuentes, C.J.; Denver, R.J. Molecular Mechanisms for Krüppel-Like Factor 13 Actions in Hippocampal Neurons. Mol. Neurobiol. 2020, 57, 3785–3802. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Kubo, M.; Ohmiya, H.; Takahashi, A.; Kumasaka, N.; Hosono, N.; Maeda, S.; Wen, W.; Dorajoo, R.; Go, M.J.; et al. Common Variants at CDKAL1 and KLF9 Are Associated with Body Mass Index in East Asian Populations. Nat. Genet. 2012, 44, 302–306. [Google Scholar] [CrossRef]
- Pei, H.; Yao, Y.; Yang, Y.; Liao, K.; Wu, J.-R. Krüppel-like Factor KLF9 Regulates PPARγ Transactivation at the Middle Stage of Adipogenesis. Cell Death Differ. 2011, 18, 315–327. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Wei, H.; Song, T.; Yang, Y.; Zhang, F.; Zhou, Y.; Peng, J.; Jiang, S. KLF13 Promotes Porcine Adipocyte Differentiation through PPARγ Activation. Cell Biosci. 2015, 5, 28. [Google Scholar] [CrossRef]
- Zhang, X.L.; Simmen, F.A.; Michel, F.J.; Simmen, R.C. Increased Expression of the Zn-Finger Transcription Factor BTEB1 in Human Endometrial Cells Is Correlated with Distinct Cell Phenotype, Gene Expression Patterns, and Proliferative Responsiveness to Serum and TGF-Beta1. Mol. Cell. Endocrinol. 2001, 181, 81–96. [Google Scholar] [CrossRef]
- Simmen, R.C.M.; Zhang, X.-L.; Michel, F.J.; Min, S.H.; Zhao, G.; Simmen, F.A. Molecular Markers of Endometrial Epithelial Cell Mitogenesis Mediated by the Sp/Krüppel-like Factor BTEB1. DNA Cell Biol. 2002, 21, 115–128. [Google Scholar] [CrossRef]
- Mazzoccoli, G.; Colangelo, T.; Panza, A.; Rubino, R.; Tiberio, C.; Palumbo, O.; Carella, M.; Trombetta, D.; Gentile, A.; Tavano, F.; et al. Analysis of Clock Gene-miRNA Correlation Networks Reveals Candidate Drivers in Colorectal Cancer. Oncotarget 2016, 7, 45444–45461. [Google Scholar] [CrossRef]
- Drepanos, L.; Gans, I.M.; Grendler, J.; Guitar, S.; Fuqua, J.H.; Maki, N.J.; Tilden, A.R.; Graber, J.H.; Coffman, J.A. Loss of Krüppel-like Factor 9 Deregulates Both Physiological Gene Expression and Development. Sci. Rep. 2023, 13, 12239. [Google Scholar] [CrossRef] [PubMed]
- Rezaeian, A.-H.; Dang, F.; Wei, W. The Circadian Clock, Aging and Its Implications in Cancer. Neoplasia 2023, 41, 100904. [Google Scholar] [CrossRef] [PubMed]
- Ybañez, W.S.; Bagamasbad, P.D. Krüppel-like Factor 9 (KLF9) Links Hormone Dysregulation and Circadian Disruption to Breast Cancer Pathogenesis. Cancer Cell Int. 2023, 23, 33. [Google Scholar] [CrossRef] [PubMed]
- Yoshitane, H.; Ozaki, H.; Terajima, H.; Du, N.-H.; Suzuki, Y.; Fujimori, T.; Kosaka, N.; Shimba, S.; Sugano, S.; Takagi, T.; et al. CLOCK-Controlled Polyphonic Regulation of Circadian Rhythms through Canonical and Noncanonical E-Boxes. Mol. Cell. Biol. 2014, 34, 1776–1787. [Google Scholar] [CrossRef] [PubMed]
- Welcker, M.; Clurman, B.E. FBW7 Ubiquitin Ligase: A Tumour Suppressor at the Crossroads of Cell Division, Growth and Differentiation. Nat. Rev. Cancer 2008, 8, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Hong, S.; Maniar, K.P.; Cheng, S.; Jie, C.; Rademaker, A.W.; Krensky, A.M.; Clayberger, C. KLF13 Regulates the Differentiation-Dependent Human Papillomavirus Life Cycle in Keratinocytes through STAT5 and IL-8. Oncogene 2016, 35, 5565–5575. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Wu, Y.-B.; Wang, L.-H.; Yin, J.-N. Involvement of Kruppel-like Factor 9 in Bleomycin-Induced Pulmonary Toxicity. Mol. Med. Rep. 2015, 12, 5262–5266. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Lv, Z.; Zhang, H.; Liu, B.; Jiang, H.; Tan, X.; Lu, J.; Baiyun, R.; Zhang, Z. Activation of the Nrf2 Signaling Pathway Involving KLF9 Plays a Critical Role in Allicin Resisting Against Arsenic Trioxide-Induced Hepatotoxicity in Rats. Biol. Trace Elem. Res. 2017, 176, 192–200. [Google Scholar] [CrossRef]
- Bagheri-Yarmand, R.; Sinha, K.M.; Li, L.; Lu, Y.; Cote, G.J.; Sherman, S.I.; Gagel, R.F. Combinations of Tyrosine Kinase Inhibitor and ERAD Inhibitor Promote Oxidative Stress-Induced Apoptosis through ATF4 and KLF9 in Medullary Thyroid Cancer. Mol. Cancer Res. 2019, 17, 751–760. [Google Scholar] [CrossRef]
- Parga, J.A.; Rodriguez-Perez, A.I.; Garcia-Garrote, M.; Rodriguez-Pallares, J.; Labandeira-Garcia, J.L. Angiotensin II Induces Oxidative Stress and Upregulates Neuroprotective Signaling from the NRF2 and KLF9 Pathway in Dopaminergic Cells. Free. Radic. Biol. Med. 2018, 129, 394–406. [Google Scholar] [CrossRef]
- Yan, Q.; He, B.; Hao, G.; Liu, Z.; Tang, J.; Fu, Q.; Jiang, C.X. KLF9 Aggravates Ischemic Injury in Cardiomyocytes through Augmenting Oxidative Stress. Life Sci. 2019, 233, 116641. [Google Scholar] [CrossRef] [PubMed]
- Chhunchha, B.; Kubo, E.; Singh, D.P. Sulforaphane-Induced Klf9/Prdx6 Axis Acts as a Molecular Switch to Control Redox Signaling and Determines Fate of Cells. Cells 2019, 8, 1159. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Weng, Y.; Lai, L.; Lei, H.; Xu, S.; Zhang, Y.; Li, L. KLF9 Regulates PRDX6 Expression in Hyperglycemia-Aggravated Bupivacaine Neurotoxicity. Mol. Cell Biochem. 2021, 476, 2125–2134. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xing, R.; Ou, Z.; Zhao, J.; Hong, G.; Zhao, T.-J.; Han, Y.; Chen, Y. G-Protein-Coupled Receptor GPR17 Inhibits Glioma Development by Increasing Polycomb Repressive Complex 1-Mediated ROS Production. Cell Death Dis. 2021, 12, 610. [Google Scholar] [CrossRef]
- Taqi, M.O.; Saeed-Zidane, M.; Gebremedhn, S.; Salilew-Wondim, D.; Tholen, E.; Neuhoff, C.; Hoelker, M.; Schellander, K.; Tesfaye, D. NRF2-Mediated Signaling Is a Master Regulator of Transcription Factors in Bovine Granulosa Cells under Oxidative Stress Condition. Cell Tissue Res. 2021, 385, 769–783. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Peng, J.; Feng, H.; Yang, Y.; Gao, J.; Liu, C.; Xu, J.; Zhao, Y.; Pan, S.; Wang, Y.; et al. KLF9 Aggravates Streptozotocin-Induced Diabetic Cardiomyopathy by Inhibiting PPARγ/NRF2 Signalling. Cells 2022, 11, 3393. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.R.; Alhallak, I.; Simmen, R.C.M.; Melnyk, S.B.; Heard-Lipsmeyer, M.E.; Montales, M.T.E.; Habenicht, D.; Van, T.T.; Simmen, F.A. Krüppel-like Factor 9 (KLF9) Suppresses Hepatocellular Carcinoma (HCC)-Promoting Oxidative Stress and Inflammation in Mice Fed High-Fat Diet. Cancers 2022, 14, 1737. [Google Scholar] [CrossRef]
- Heard, M.E.; Melnyk, S.B.; Simmen, F.A.; Yang, Y.; Pabona, J.M.P.; Simmen, R.C.M. High-Fat Diet Promotion of Endometriosis in an Immunocompetent Mouse Model Is Associated with Altered Peripheral and Ectopic Lesion Redox and Inflammatory Status. Endocrinology 2016, 157, 2870–2882. [Google Scholar] [CrossRef]
- Heard-Lipsmeyer, M.E.; Alhallak, I.; Simmen, F.A.; Melnyk, S.B.; Simmen, R.C.M. Lesion Genotype Modifies High-Fat Diet Effects on Endometriosis Development in Mice. Front. Physiol. 2021, 12, 702674. [Google Scholar] [CrossRef]
- Mannava, S.; Zhuang, D.; Nair, J.R.; Bansal, R.; Wawrzyniak, J.A.; Zucker, S.N.; Fink, E.E.; Moparthy, K.C.; Hu, Q.; Liu, S.; et al. KLF9 Is a Novel Transcriptional Regulator of Bortezomib- and LBH589-Induced Apoptosis in Multiple Myeloma Cells. Blood 2012, 119, 1450–1458. [Google Scholar] [CrossRef]
- Fink, E.E.; Moparthy, S.; Bagati, A.; Bianchi-Smiraglia, A.; Lipchick, B.C.; Wolff, D.W.; Roll, M.V.; Wang, J.; Liu, S.; Bakin, A.V.; et al. XBP1-KLF9 Axis Acts as a Molecular Rheostat to Control the Transition from Adaptive to Cytotoxic Unfolded Protein Response. Cell Rep. 2018, 25, 212–223.e4. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.; Davis, B.H. The DNA Binding Protein BTEB Mediates Acetaldehyde-Induced, Jun N-Terminal Kinase-Dependent alphaI (I) Collagen Gene Expression in Rat Hepatic Stellate Cells. Mol. Cell. Biol. 2000, 20, 2818–2826. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Ma, C.; Dai, W.; Fang, E.; Li, W.; Yang, F. Ghrelin Attenuates Transforming Growth Factor-Β1-Induced Pulmonary Fibrosis via the miR-125a-5p/Kruppel-like Factor 13 Axis. Arch. Biochem. Biophys. 2022, 715, 109082. [Google Scholar] [CrossRef] [PubMed]
- Ying, M.; Sang, Y.; Li, Y.; Guerrero-Cazares, H.; Quinones-Hinojosa, A.; Vescovi, A.L.; Eberhart, C.G.; Xia, S.; Laterra, J. Krüppel-like Family of Transcription Factor 9, a Differentiation-Associated Transcription Factor, Suppresses Notch1 Signaling and Inhibits Glioblastoma-Initiating Stem Cells. Stem Cells 2011, 29, 20–31. [Google Scholar] [CrossRef]
- Tung, B.; Ma, D.; Wang, S.; Oyinlade, O.; Laterra, J.; Ying, M.; Lv, S.-Q.; Wei, S.; Xia, S. Krüppel-like Factor 9 and Histone Deacetylase Inhibitors Synergistically Induce Cell Death in Glioblastoma Stem-like Cells. BMC Cancer 2018, 18, 1025. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Zhang, M.; Tian, N.; Li, D.; Wu, F.; Hu, P.; Wang, Z.; Wang, L.; Hao, W.; Kang, J.; et al. The Antibiotic Clofoctol Suppresses Glioma Stem Cell Proliferation by Activating KLF13. J. Clin. Investig. 2019, 129, 3072–3085. [Google Scholar] [CrossRef]
- Wang, K.; Liu, S.; Dou, Z.; Zhang, S.; Yang, X. Loss of Krüppel-like Factor 9 Facilitates Stemness in Ovarian Cancer Ascites-Derived Multicellular Spheroids via Notch1/Slug Signaling. Cancer Sci. 2021, 112, 4220–4233. [Google Scholar] [CrossRef]
- Shan, L.; Song, P.; Zhao, Y.; An, N.; Xia, Y.; Qi, Y.; Zhao, H.; Ge, J. miR-600 Promotes Ovarian Cancer Cells Stemness, Proliferation and Metastasis via Targeting KLF9. J. Ovarian Res. 2022, 15, 52. [Google Scholar] [CrossRef]
- He, Q.; Huang, L.; Yan, D.; Bi, J.; Yang, M.; Huang, J.; Lin, T. CircPTPRA Acts as a Tumor Suppressor in Bladder Cancer by Sponging miR-636 and Upregulating KLF9. Aging 2019, 11, 11314–11328. [Google Scholar] [CrossRef]
- Limame, R.; de Beeck, K.O.; Van Laere, S.; Croes, L.; De Wilde, A.; Dirix, L.; Van Camp, G.; Peeters, M.; De Wever, O.; Lardon, F.; et al. Expression Profiling of Migrated and Invaded Breast Cancer Cells Predicts Early Metastatic Relapse and Reveals Krüppel-like Factor 9 as a Potential Suppressor of Invasive Growth in Breast Cancer. Oncoscience 2014, 1, 69–81. [Google Scholar] [CrossRef]
- Jiang, Z.; Xu, Z.; Hu, T.; Song, B.; Li, F.; Wang, K. Expression of Krüppel-like Factor 9 in Breast Cancer Patients and Its Effect on Prognosis. Oncol. Lett. 2020, 20, 1311–1317. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Jiang, X.; Liu, Y.; Wang, Y.; Jiang, X.; Song, G.; Qiu, H.; Zhang, Q. Krüppel-like Factor 9 Upregulates E-Cadherin Transcription and Represses Breast Cancer Invasion and Metastasis. Am. J. Cancer Res. 2021, 11, 3660–3673. [Google Scholar] [PubMed]
- Kadamb, R.; Leibovitch, B.A.; Farias, E.F.; Dahiya, N.; Suryawanshi, H.; Bansal, N.; Waxman, S. Invasive Phenotype in Triple Negative Breast Cancer Is Inhibited by Blocking SIN3A-PF1 Interaction through KLF9 Mediated Repression of ITGA6 and ITGB1. Transl. Oncol. 2022, 16, 101320. [Google Scholar] [CrossRef] [PubMed]
- Safi, S.; Badshah, Y.; Shabbir, M.; Zahra, K.; Khan, K.; Dilshad, E.; Afsar, T.; Almajwal, A.; Alruwaili, N.W.; Al-Disi, D.; et al. Predicting 3D Structure, Cross Talks, and Prognostic Significance of KLF9 in Cervical Cancer. Front. Oncol. 2021, 11, 797007. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Xia, L.; Jiang, H.; Hu, Y.; Li, L.; Xu, L.; Xia, R. Long Non-coding RNA DANCR Represses the Viability, Migration and Invasion of Multiple Myeloma Cells by Sponging miR-135b-5p to Target KLF9. Mol. Med. Rep. 2021, 24, 649. [Google Scholar] [CrossRef]
- Fernandes, L.M.; Al-Dwairi, A.; Simmen, R.C.M.; Marji, M.; Brown, D.M.; Jewell, S.W.; Simmen, F.A. Malic Enzyme 1 (ME1) Is pro-Oncogenic in ApcMin/+ Mice. Sci. Rep. 2018, 8, 14268. [Google Scholar] [CrossRef]
- Kang, L.; Lü, B.; Xu, J.; Hu, H.; Lai, M. Downregulation of Krüppel-like Factor 9 in Human Colorectal Cancer. Pathol. Int. 2008, 58, 334–338. [Google Scholar] [CrossRef]
- Brown, A.R.; Simmen, R.C.M.; Raj, V.R.; Van, T.T.; MacLeod, S.L.; Simmen, F.A. Krüppel-like Factor 9 (KLF9) Prevents Colorectal Cancer through Inhibition of Interferon-Related Signaling. Carcinogenesis 2015, 36, 946–955. [Google Scholar] [CrossRef]
- Pan, F.; Chen, T.; Sun, X.; Li, K.; Jiang, X.; Försti, A.; Zhu, Y.; Lai, M. Prognosis Prediction of Colorectal Cancer Using Gene Expression Profiles. Front. Oncol. 2019, 9, 252. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Z.; Yi, Y.; Wang, Y.; Fu, J. CircNOL10 Acts as a Sponge of miR-135a/b-5p in Suppressing Colorectal Cancer Progression via Regulating KLF9. Onco Targets Ther. 2020, 13, 5165–5176. [Google Scholar] [CrossRef]
- Yu, Y.; Li, C.; Wang, Y.; Wang, Q.; Wang, S.; Wei, S.; Yang, M.; Qin, Q. Molecular Cloning and Characterization of Grouper Krϋppel-like Factor 9 Gene: Involvement in the Fish Immune Response to Viral Infection. Fish Shellfish Immunol. 2019, 89, 677–686. [Google Scholar] [CrossRef]
- Chen, Z.Y.; Shie, J.; Tseng, C. Up-Regulation of Gut-Enriched Krüppel-like Factor by Interferon-Gamma in Human Colon Carcinoma Cells. FEBS Lett. 2000, 477, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Peng, Z.; Tang, H.; Wei, P.; Kong, X.; Yan, D.; Huang, F.; Li, Q.; Le, X.; Li, Q.; et al. KLF4-Mediated Negative Regulation of IFITM3 Expression Plays a Critical Role in Colon Cancer Pathogenesis. Clin. Cancer Res. 2011, 17, 3558–3568. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-F.; Chen, L.; Luo, J.; He, H.-X. KLF5 Is Involved in Regulation of IFITM1, 2, and 3 Genes during H5N1 Virus Infection in A549 Cells. Cell. Mol. Biol. 2016, 62, 65–70. [Google Scholar] [CrossRef][Green Version]
- Lee, K.S.; Kim, B.H.; Oh, H.-K.; Kim, D.-W.; Kang, S.-B.; Kim, H.; Shin, E. Programmed Cell Death Ligand-1 Protein Expression and CD274/PD-L1 Gene Amplification in Colorectal Cancer: Implications for Prognosis. Cancer Sci. 2018, 109, 2957–2969. [Google Scholar] [CrossRef] [PubMed]
- Gao, P.; He, M.; Zhang, C.; Geng, C. Integrated Analysis of Gene Expression Signatures Associated with Colon Cancer from Three Datasets. Gene 2018, 654, 95–102. [Google Scholar] [CrossRef]
- Yao, W.; Jiao, Y.; Zhou, Y.; Luo, X. KLF13 Suppresses the Proliferation and Growth of Colorectal Cancer Cells through Transcriptionally Inhibiting HMGCS1-Mediated Cholesterol Biosynthesis. Cell Biosci. 2020, 10, 76. [Google Scholar] [CrossRef]
- Xing, J.; Jia, Z.; Xu, Y.; Chen, M.; Yang, Z.; Chen, Y.; Han, Y. KLF9 (Kruppel Like Factor 9) Induced PFKFB3 (6-Phosphofructo-2-Kinase/Fructose-2,6-Biphosphatase 3) Downregulation Inhibits the Proliferation, Metastasis and Aerobic Glycolysis of Cutaneous Squamous Cell Carcinoma Cells. Bioengineered 2021, 12, 7563–7576. [Google Scholar] [CrossRef]
- Bulun, S.E.; Wan, Y.; Matei, D. Epithelial Mutations in Endometriosis: Link to Ovarian Cancer. Endocrinology 2019, 160, 626–638. [Google Scholar] [CrossRef]
- Ye, J.; Peng, H.; Huang, X.; Qi, X. The Association between Endometriosis and Risk of Endometrial Cancer and Breast Cancer: A Meta-Analysis. BMC Women’s Health 2022, 22, 455. [Google Scholar] [CrossRef]
- Pabona, J.M.P.; Simmen, F.A.; Nikiforov, M.A.; Zhuang, D.; Shankar, K.; Velarde, M.C.; Zelenko, Z.; Giudice, L.C.; Simmen, R.C.M. Krüppel-like Factor 9 and Progesterone Receptor Coregulation of Decidualizing Endometrial Stromal Cells: Implications for the Pathogenesis of Endometriosis. J. Clin. Endocrinol. Metab. 2012, 97, E376–E392. [Google Scholar] [CrossRef]
- Heard, M.E.; Simmons, C.D.; Simmen, F.A.; Simmen, R.C.M. Krüppel-like Factor 9 Deficiency in Uterine Endometrial Cells Promotes Ectopic Lesion Establishment Associated with Activated Notch and Hedgehog Signaling in a Mouse Model of Endometriosis. Endocrinology 2014, 155, 1532–1546. [Google Scholar] [CrossRef]
- Brown, D.M.; Lee, H.-C.; Liu, S.; Quick, C.M.; Fernandes, L.M.; Simmen, F.A.; Tsai, S.-J.; Simmen, R.C.M. Notch-1 Signaling Activation and Progesterone Receptor Expression in Ectopic Lesions of Women with Endometriosis. J. Endocr. Soc. 2018, 2, 765–778. [Google Scholar] [CrossRef] [PubMed]
- Korani, M.; Fallah, S.; Tehranian, A.; Nourbakhsh, M.; Samadikuchaksaraei, A.; Pour, M.S.; Maleki, J. The Evaluation of the FOXO1, KLF9 and YT521 Genes Expression in Human Endometrial Cancer. Clin. Lab. 2013, 59, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Velarde, M.C.; Zeng, Z.; McQuown, J.R.; Simmen, F.A.; Simmen, R.C.M. Kruppel-like Factor 9 Is a Negative Regulator of Ligand-Dependent Estrogen Receptor Alpha Signaling in Ishikawa Endometrial Adenocarcinoma Cells. Mol. Endocrinol. 2007, 21, 2988–3001. [Google Scholar] [CrossRef]
- Simmons, C.D.; Pabona, J.M.P.; Zeng, Z.; Velarde, M.C.; Gaddy, D.; Simmen, F.A.; Simmen, R.C.M. Response of Adult Mouse Uterus to Early Disruption of Estrogen Receptor-Alpha Signaling Is Influenced by Krüppel-like Factor 9. J. Endocrinol. 2010, 205, 147–157. [Google Scholar] [CrossRef][Green Version]
- Qiao, F.; Yao, F.; Chen, L.; Lu, C.; Ni, Y.; Fang, W.; Jin, H. Krüppel-like Factor 9 Was Down-Regulated in Esophageal Squamous Cell Carcinoma and Negatively Regulated Beta-Catenin/TCF Signaling. Mol. Carcinog. 2016, 55, 280–291. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Huang, Y.; Wang, Y.; Wu, Q.; Hong, S.; Huang, Z. THBS4 Predicts Poor Outcomes and Promotes Proliferation and Metastasis in Gastric Cancer. J. Physiol. Biochem. 2019, 75, 117–123. [Google Scholar] [CrossRef]
- Li, Y.; Sun, Q.; Jiang, M.; Li, S.; Zhang, J.; Xu, Z.; Guo, D.; Gu, T.; Wang, B.; Xiao, L.; et al. KLF9 Suppresses Gastric Cancer Cell Invasion and Metastasis through Transcriptional Inhibition of MMP28. FASEB J. 2019, 33, 7915–7928. [Google Scholar] [CrossRef]
- Ding, Y.; Xu, Y.; Fu, Y.; Zhang, H.; Zhao, L.; Fan, X. Kruppel-like Factor 13 Inhibits Cell Proliferation of Gastric Cancer by Inducing Autophagic Degradation of β-Catenin. Discov. Oncol. 2022, 13, 121. [Google Scholar] [CrossRef]
- Huang, S.; Wang, C.; Yi, Y.; Sun, X.; Luo, M.; Zhou, Z.; Li, J.; Cai, Y.; Jiang, X.; Ke, Y. Krüppel-like Factor 9 Inhibits Glioma Cell Proliferation and Tumorigenicity via Downregulation of miR-21. Cancer Lett. 2015, 356 Pt B, 547–555. [Google Scholar] [CrossRef]
- Zhang, D.; Hao, P.; Jin, L.; Wang, Y.; Yan, Z.; Wu, S. MicroRNA-940 Promotes Cell Proliferation and Invasion of Glioma by Directly Targeting Kruppel-like Factor 9. Mol. Med. Rep. 2019, 19, 734–742. [Google Scholar] [CrossRef] [PubMed]
- Ying, M.; Tilghman, J.; Wei, Y.; Guerrero-Cazares, H.; Quinones-Hinojosa, A.; Ji, H.; Laterra, J. Kruppel-like Factor-9 (KLF9) Inhibits Glioblastoma Stemness through Global Transcription Repression and Integrin A6 Inhibition. J. Biol. Chem. 2014, 289, 32742–32756. [Google Scholar] [CrossRef] [PubMed]
- Cvoro, A.; Devito, L.; Milton, F.A.; Noli, L.; Zhang, A.; Filippi, C.; Sakai, K.; Suh, J.H.; Sieglaff, D.H.; Dhawan, A.; et al. A Thyroid Hormone Receptor/KLF9 Axis in Human Hepatocytes and Pluripotent Stem Cells. Stem Cells 2015, 33, 416–428. [Google Scholar] [CrossRef] [PubMed]
- Escalona-Nandez, I.; Guerrero-Escalera, D.; Estanes-Hernández, A.; Ortíz-Ortega, V.; Tovar, A.R.; Pérez-Monter, C. The Activation of Peroxisome Proliferator-Activated Receptor γ Is Regulated by Krüppel-like Transcription Factors 6 & 9 under Steatotic Conditions. Biochem. Biophys. Res. Commun. 2015, 458, 751–756. [Google Scholar] [CrossRef]
- Zhou, S.-S.; Zhang, Y.-L.; Chang, Y.-S. KLF9 Regulates Hepatic Lipid Metabolism via Inducing CD36 Expression. Acta Physiol. Sin. 2021, 73, 772–780. [Google Scholar]
- Liu, S.; Liu, M.; Zhang, M.-L.; Wang, C.-Z.; Zhang, Y.-L.; Zhang, Y.-J.; Du, C.-Y.; Sheng, S.-F.; Wang, W.; Fan, Y.-T.; et al. Transcription Factor Klf9 Controls Bile Acid Reabsorption and Enterohepatic Circulation in Mice via Promoting Intestinal Asbt Expression. Acta Pharmacol. Sin. 2022, 43, 2362–2372. [Google Scholar] [CrossRef]
- Yang, H.; Arif, M.; Yuan, M.; Li, X.; Shong, K.; Türkez, H.; Nielsen, J.; Uhlén, M.; Borén, J.; Zhang, C.; et al. A Network-Based Approach Reveals the Dysregulated Transcriptional Regulation in Non-Alcoholic Fatty Liver Disease. iScience 2021, 24, 103222. [Google Scholar] [CrossRef]
- Fu, D.-Z.; Cheng, Y.; He, H.; Liu, H.-Y.; Liu, Y.-F. The Fate of Krüppel-like Factor 9-Positive Hepatic Carcinoma Cells May Be Determined by the Programmed Cell Death Protein 5. Int. J. Oncol. 2014, 44, 153–160. [Google Scholar] [CrossRef]
- Sun, J.; Wang, B.; Liu, Y.; Zhang, L.; Ma, A.; Yang, Z.; Ji, Y.; Liu, Y. Transcription Factor KLF9 Suppresses the Growth of Hepatocellular Carcinoma Cells in Vivo and Positively Regulates P53 Expression. Cancer Lett. 2014, 355, 25–33. [Google Scholar] [CrossRef]
- Wang, T.; Feng, L.; Shi, Z.; Yang, L.; Yu, X.; Wu, J.; Sun, J.; Zhang, J.; Feng, Y.; Wang, W. A Negative Feedback Loop between KLF9 and the EMT Program Dictates Metastasis of Hepatocellular Carcinoma. J. Cell Mol. Med. 2023, 27, 2372–2384. [Google Scholar] [CrossRef] [PubMed]
- Kowalik, M.A.; Puliga, E.; Cabras, L.; Sulas, P.; Petrelli, A.; Perra, A.; Ledda-Columbano, G.M.; Morandi, A.; Merlin, S.; Orrù, C.; et al. Thyroid Hormone Inhibits Hepatocellular Carcinoma Progression via Induction of Differentiation and Metabolic Reprogramming. J. Hepatol. 2020, 72, 1159–1169. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-C.; Xie, X.-M.; Zhao, X.-K.; Zuo, S.; Li, H.-Y. Krüppel-like Factor 13 Promotes HCC Progression by Transcriptional Regulation of HMGCS1-Mediated Cholesterol Synthesis. J. Clin. Transl. Hepatol. 2022, 10, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Chen, C.; Feng, S.; Zuo, S.; Zhao, X.; Li, H. Acyl-CoA Thioesterase 7 Is Transcriptionally Activated by Krüppel-Like Factor 13 and Promotes the Progression of Hepatocellular Carcinoma. J. Hepatocell. Carcinoma 2021, 8, 1623–1641. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Huang, X.; Wang, W.; Xie, H.; Li, J.; Hu, Z.; Zheng, Z.; Li, H.; Teng, L. LncRNA CDKN2BAS Predicts Poor Prognosis in Patients with Hepatocellular Carcinoma and Promotes Metastasis via the miR-153-5p/ARHGAP18 Signaling Axis. Aging 2018, 10, 3371–3381. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Li, J.; Zhang, X.; Xiong, T.; Ye, J.; Yu, J.; Gui, Y. The miR-140-5p/KLF9/KCNQ1 Axis Promotes the Progression of Renal Cell Carcinoma. FASEB J. 2020, 34, 10623–10639. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wang, X.; Wang, Q.; Ding, K.; Chen, X.; Zhao, Y.; Gao, Y.; Wang, Y. Effects and Prognostic Values of Circadian Genes CSNK1E/GNA11/KLF9/THRAP3 in Kidney Renal Clear Cell Carcinoma via a Comprehensive Analysis. Bioengineering 2022, 9, 306. [Google Scholar] [CrossRef]
- Yang, L.; Yin, H.; Chen, Y.; Pan, C.; Hang, H.; Lu, Y.; Ma, W.; Li, X.; Gan, W.; Guo, H.; et al. Low Expression of PEBP1P2 Promotes Metastasis of Clear Cell Renal Cell Carcinoma by Post-Transcriptional Regulation of PEBP1 and KLF13 mRNA. Exp. Hematol. Oncol. 2022, 11, 87. [Google Scholar] [CrossRef]
- Bagati, A.; Moparthy, S.; Fink, E.E.; Bianchi-Smiraglia, A.; Yun, D.H.; Kolesnikova, M.; Udartseva, O.O.; Wolff, D.W.; Roll, M.V.; Lipchick, B.C.; et al. KLF9-Dependent ROS Regulate Melanoma Progression in Stage-Specific Manner. Oncogene 2019, 38, 3585–3597. [Google Scholar] [CrossRef]
- Yi, T.-W.; Lv, X.-X.; Fan, H.; Zan, N.; Su, X.-D. LncRNA SNHG15 Promotes the Proliferation of Nasopharyngeal Carcinoma via Sponging miR-141-3p to Upregulate KLF9. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 6744–6751. [Google Scholar] [CrossRef]
- Chen, S.; Gu, S.; Xu, M.; Mei, D.; Xiao, Y.; Chen, K.; Yan, Z. Krüppel-like Factor 9 Promotes Neuroblastoma Differentiation via Targeting the Sonic Hedgehog Signaling Pathway. Pediatr. Blood Cancer 2020, 67, e28108. [Google Scholar] [CrossRef] [PubMed]
- Tong, X.-D.; Liu, T.-Q.; Wang, G.-B.; Zhang, C.-L.; Liu, H.-X. MicroRNA-570 Promotes Lung Carcinoma Proliferation through Targeting Tumor Suppressor KLF9. Int. J. Clin. Exp. Pathol. 2015, 8, 2829–2834. [Google Scholar] [PubMed]
- Kong, Y.-J.; Tan, X.-X.; Zhang, Y.; He, Q.-J.; Zhao, L.; Meng, Q. MiR-141 Promotes Cell Proliferation and Invasion in Non-Small Cell Lung Cancer by Targeting KLF9. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 10370–10378. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Tang, Y.; Dai, Y.; Hu, S.; Zhou, J.; Liu, X.; Zhu, J.; Wu, Y. MiR-889 Promotes Cell Growth in Human Non-Small Cell Lung Cancer by Regulating KLF9. Gene 2019, 699, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Huang, Y.; Lin, J.; Sun, S.; Che, K.; Shen, J.; Liao, J.; Chen, Y.; Chen, K.; Lin, Z.; et al. Mxi1 Participates in the Progression of Lung Cancer via the microRNA-300/KLF9/GADD34 Axis. Cell Death Dis. 2022, 13, 425. [Google Scholar] [CrossRef]
- Boyero, L.; Noguera-Uclés, J.F.; Castillo-Peña, A.; Salinas, A.; Sánchez-Gastaldo, A.; Alonso, M.; Benedetti, J.C.; Bernabé-Caro, R.; Paz-Ares, L.; Molina-Pinelo, S. Aberrant Methylation of the Imprinted C19MC and MIR371-3 Clusters in Patients with Non-Small Cell Lung Cancer. Cancers 2023, 15, 1466. [Google Scholar] [CrossRef]
- Wang, C.; Wang, Q.; Weng, Z. LINC00664/miR-411-5p/KLF9 Feedback Loop Contributes to the Human Oral Squamous Cell Carcinoma Progression. Oral Dis. 2023, 29, 672–685. [Google Scholar] [CrossRef]
- Peng, N.; Miao, Z.; Wang, L.; Liu, B.; Wang, G.; Guo, X. MiR-378 Promotes the Cell Proliferation of Osteosarcoma through Down-Regulating the Expression of Kruppel-like Factor 9. Biochem. Cell Biol. 2018, 96, 515–521. [Google Scholar] [CrossRef]
- Jin, Y.; Yang, L.; Li, X. MicroRNA-652 Promotes Cell Proliferation and Osteosarcoma Invasion by Directly Targeting KLF9. Exp. Ther. Med. 2020, 20, 2953–2960. [Google Scholar] [CrossRef]
- Chen, Q.; Zhou, H.; Rong, W. Circular RNA_0078767 Upregulates Kruppel-like Factor 9 Expression by Targeting microRNA-889, Thereby Inhibiting the Progression of Osteosarcoma. Bioengineered 2022, 13, 14313–14328. [Google Scholar] [CrossRef]
- Liu, Y.; Han, K.; Cao, Y.; Hu, Y.; Shao, Z.; Tong, W.; Han, Y.; Liu, Y. KLF9 Regulates miR-338-3p/NRCAM Axis to Block the Progression of Osteosarcoma Cells. J. Cancer 2022, 13, 2029–2039. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Dou, H.; Tang, Y.; Su, S.; Liu, P. Lentivirus-Mediated Knockdown of Krüppel-like Factor 9 Inhibits the Growth of Ovarian Cancer. Arch Gynecol. Obstet. 2015, 291, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Mao, Z.; Fan, X.; Zhang, J.; Wang, X.; Ma, X.; Michalski, C.W.; Zhang, Y. KLF9 Is a Prognostic Indicator in Human Pancreatic Ductal Adenocarcinoma. Anticancer. Res. 2017, 37, 3795–3799. [Google Scholar] [CrossRef] [PubMed]
- Ji, P.; Fan, X.; Ma, X.; Wang, X.; Zhang, J.; Mao, Z. Krüppel-like Factor 9 Suppressed Tumorigenicity of the Pancreatic Ductal Adenocarcinoma by Negatively Regulating Frizzled-5. Biochem. Biophys. Res. Commun. 2018, 499, 815–821. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Zhou, F.; Wang, D.; Wu, M.; Zhou, W.; Zou, Y.; Li, J.; Wu, L.; Yin, X. Expression of KLF9 in Pancreatic Cancer and Its Effects on the Invasion, Migration, Apoptosis, Cell Cycle Distribution, and Proliferation of Pancreatic Cancer Cell Lines. Oncol. Rep. 2018, 40, 3852–3860. [Google Scholar] [CrossRef]
- Li, B.; Pang, S.; Dou, J.; Zhou, C.; Shen, B.; Zhou, Y. The Inhibitory Effect of LINC00261 Upregulation on the Pancreatic Cancer EMT Process Is Mediated by KLF13 via the mTOR Signaling Pathway. Clin. Transl. Oncol. 2022, 24, 1059–1072. [Google Scholar] [CrossRef]
- Wang, Z.; Wu, P.; Shi, J.; Ji, X.; He, L.; Dong, W.; Wang, Z.; Zhang, H.; Sun, W. A Novel Necroptosis-Related Gene Signature Associated with Immune Landscape for Predicting the Prognosis of Papillary Thyroid Cancer. Front. Genet. 2022, 13, 947216. [Google Scholar] [CrossRef]
- Shen, P.; Sun, J.; Xu, G.; Zhang, L.; Yang, Z.; Xia, S.; Wang, Y.; Liu, Y.; Shi, G. KLF9, a Transcription Factor Induced in Flutamide-Caused Cell Apoptosis, Inhibits AKT Activation and Suppresses Tumor Growth of Prostate Cancer Cells. Prostate 2014, 74, 946–958. [Google Scholar] [CrossRef]
- Li, J.-Z.; Li, J.; Wang, H.-Q.; Li, X.; Wen, B.; Wang, Y.-J. MiR-141-3p Promotes Prostate Cancer Cell Proliferation through Inhibiting Kruppel-like Factor-9 Expression. Biochem. Biophys. Res. Commun. 2017, 482, 1381–1386. [Google Scholar] [CrossRef]
- Wang, Q.; Peng, R.; Wang, B.; Wang, J.; Yu, W.; Liu, Y.; Shi, G. Transcription Factor KLF13 Inhibits AKT Activation and Suppresses the Growth of Prostate Carcinoma Cells. Cancer Biomark. 2018, 22, 533–541. [Google Scholar] [CrossRef]
- Zhang, L.; Ruan, Y.; Qin, Z.; Gao, X.; Xu, K.; Shi, X.; Gao, S.; Liu, S.; Zhu, K.; Wang, W.; et al. miR-483-3p, Mediated by KLF9, Functions as Tumor Suppressor in Testicular Seminoma via Targeting MMP9. Front. Oncol. 2020, 10, 596574. [Google Scholar] [CrossRef] [PubMed]
- Pelka, K.; Hofree, M.; Chen, J.H.; Sarkizova, S.; Pirl, J.D.; Jorgji, V.; Bejnood, A.; Dionne, D.; Ge, W.H.; Xu, K.H.; et al. Spatially Organized Multicellular Immune Hubs in Human Colorectal Cancer. Cell 2021, 184, 4734–4752.e20. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.-F.; Wu, L.; Liu, S.-P.; Jiang, M.-M.; Hu, B.; Zhou, K.-Q.; Guo, W.; Xu, Y.; Zhong, Y.; Zhou, X.-R.; et al. Dissecting Spatial Heterogeneity and the Immune-Evasion Mechanism of CTCs by Single-Cell RNA-Seq in Hepatocellular Carcinoma. Nat. Commun. 2021, 12, 4091. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wu, L.; Sun, Y.; Li, J.; Mao, N.; Yang, Y.; Zhao, M.; Ren, S. CCL5 Might Be a Prognostic Biomarker and Associated with Immuno-Therapeutic Efficacy in Cancers: A Pan-Cancer Analysis. Heliyon 2023, 9, e18215. [Google Scholar] [CrossRef] [PubMed]
- Bang, S.; Li, J.; Zhang, M.; Cui, R.; Wu, X.; Xin, Z.; Ma, D.; Zhang, J.; Zhang, H. The Clinical Relevance and Function of Krüppel-Like Factor 16 in Breast Cancer. Cancer Manag. Res. 2020, 12, 6373–6383. [Google Scholar] [CrossRef]
- Zhang, J.; Yu, W.; Wang, X.; Hu, B.; Wu, D.; Shi, G. KLF16 Affects the MYC Signature and Tumor Growth in Prostate Cancer. Onco Targets Ther. 2020, 13, 1303–1310. [Google Scholar] [CrossRef]
- Jiao, X.; Gao, W.; Ren, H.; Wu, Y.; Li, T.; Li, S.; Yan, H. Kruppel like Factor 16 Promotes Lung Adenocarcinoma Progression by Upregulating Lamin B2. Bioengineered 2022, 13, 9482–9494. [Google Scholar] [CrossRef]
- Ma, X.-D.; Xu, S.-D.; Hao, S.-H.; Han, K.; Chen, J.-W.; Ling, H.; Chen, R.-X.; Jin, X.-H.; Cao, J.-H.; Lin, J.-L.; et al. KLF16 Enhances Stress Tolerance of Colorectal Carcinomas by Modulating Nucleolar Homeostasis and Translational Reprogramming. Mol. Ther. 2022, 30, 2828–2843. [Google Scholar] [CrossRef]



| KLF9 b | N c | N c | Cancer d | Normal d | p Value |
|---|---|---|---|---|---|
| Colon Cancer | 309 | 38 | 0.0568 | 0.0504 | <0.05 |
| Esophageal Cancer | 186 | 16 | 0.0545 | 0.0444 | <0.05 |
| Kidney Clear Cell Carcinoma | 323 | 160 | 0.0557 | 0.0510 | <0.05 |
| Kidney Papillary Cell Carcinoma | 276 | 45 | 0.0503 | 0.0449 | <0.01 |
| Hepatocellular Carcinoma | 380 | 50 | 0.0557 | 0.0492 | <0.01 |
| Lung Squamous Cell Carcinoma | 370 | 42 | 0.0569 | 0.0465 | <0.01 |
| Thyroid Cancer e | 515 | 56 | 0.0490 | 0.0548 | <0.01 |
| KLF13 b | N c | N c | Cancer d | Normal d | p value |
| Breast Cancer e | 794 | 96 | 0.1675 | 0.1752 | <0.01 |
| Cholangiocarcinoma e | 36 | 9 | 0.1466 | 0.1682 | <0.05 |
| Kidney Clear Cell | 323 | 160 | 0.1422 | 0.1102 | <0.01 |
| Kidney Papillary | 276 | 45 | 0.1313 | 0.1153 | <0.01 |
| Hepatocellular Carcinoma e | 380 | 50 | 0.1431 | 0.1666 | <0.01 |
| Pheochromocytoma & Paraganglioma e | 184 | 3 | 0.1236 | 0.1718 | <0.01 |
| Endometrial Cancer e | 436 | 46 | 0.1216 | 0.1711 | <0.01 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Simmen, F.A.; Alhallak, I.; Simmen, R.C.M. Krüppel-like Factor-9 and Krüppel-like Factor-13: Highly Related, Multi-Functional, Transcriptional Repressors and Activators of Oncogenesis. Cancers 2023, 15, 5667. https://doi.org/10.3390/cancers15235667
Simmen FA, Alhallak I, Simmen RCM. Krüppel-like Factor-9 and Krüppel-like Factor-13: Highly Related, Multi-Functional, Transcriptional Repressors and Activators of Oncogenesis. Cancers. 2023; 15(23):5667. https://doi.org/10.3390/cancers15235667
Chicago/Turabian StyleSimmen, Frank A., Iad Alhallak, and Rosalia C. M. Simmen. 2023. "Krüppel-like Factor-9 and Krüppel-like Factor-13: Highly Related, Multi-Functional, Transcriptional Repressors and Activators of Oncogenesis" Cancers 15, no. 23: 5667. https://doi.org/10.3390/cancers15235667
APA StyleSimmen, F. A., Alhallak, I., & Simmen, R. C. M. (2023). Krüppel-like Factor-9 and Krüppel-like Factor-13: Highly Related, Multi-Functional, Transcriptional Repressors and Activators of Oncogenesis. Cancers, 15(23), 5667. https://doi.org/10.3390/cancers15235667








