Hybrid Cellular Automata Modeling Reveals the Effects of Glucose Gradients on Tumour Spheroid Growth
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Model Development
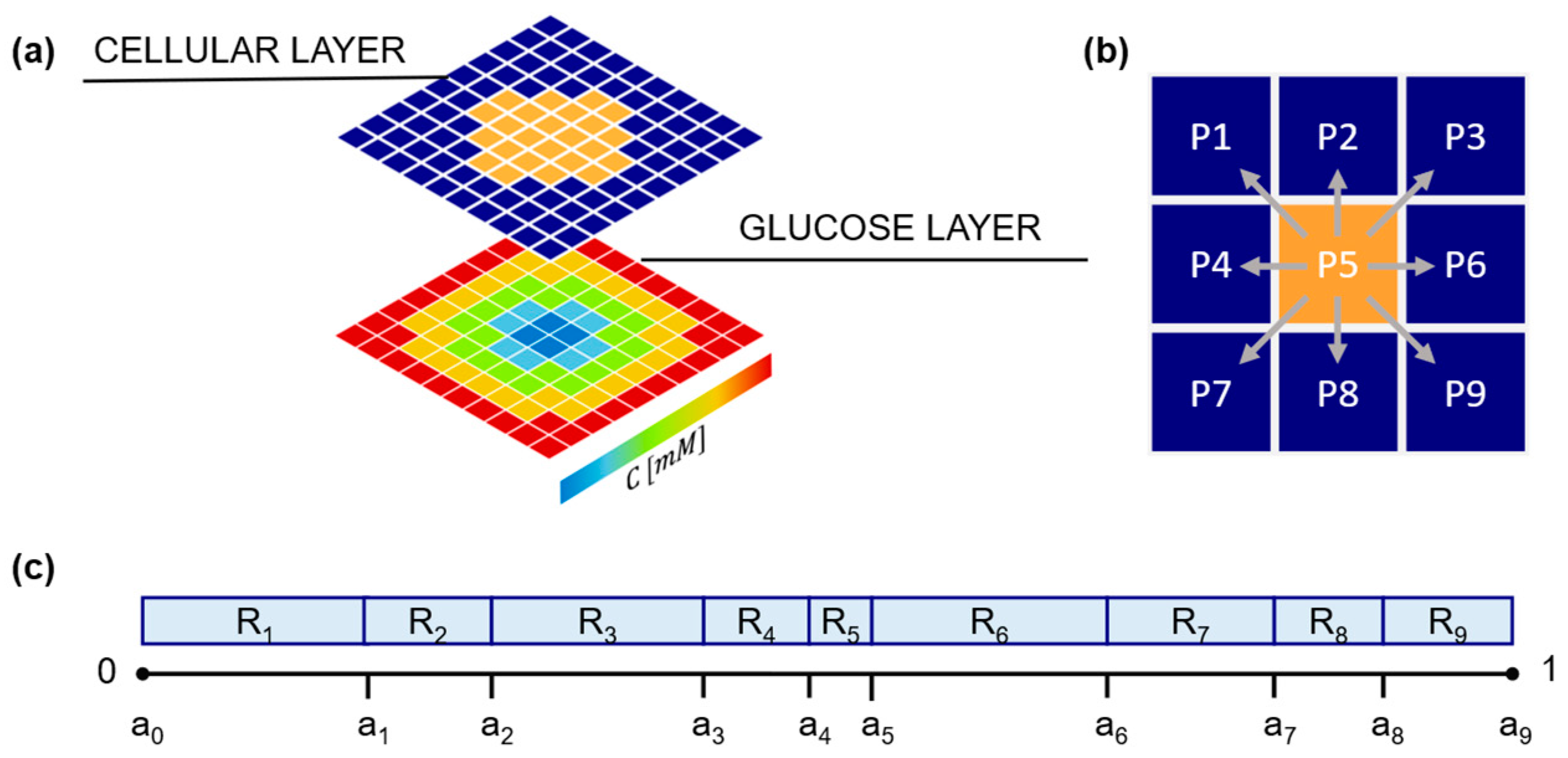
2.1.1. Domain Building
2.1.2. Glucose Layer
2.1.3. Cellular Layer and Cell Phenotypes
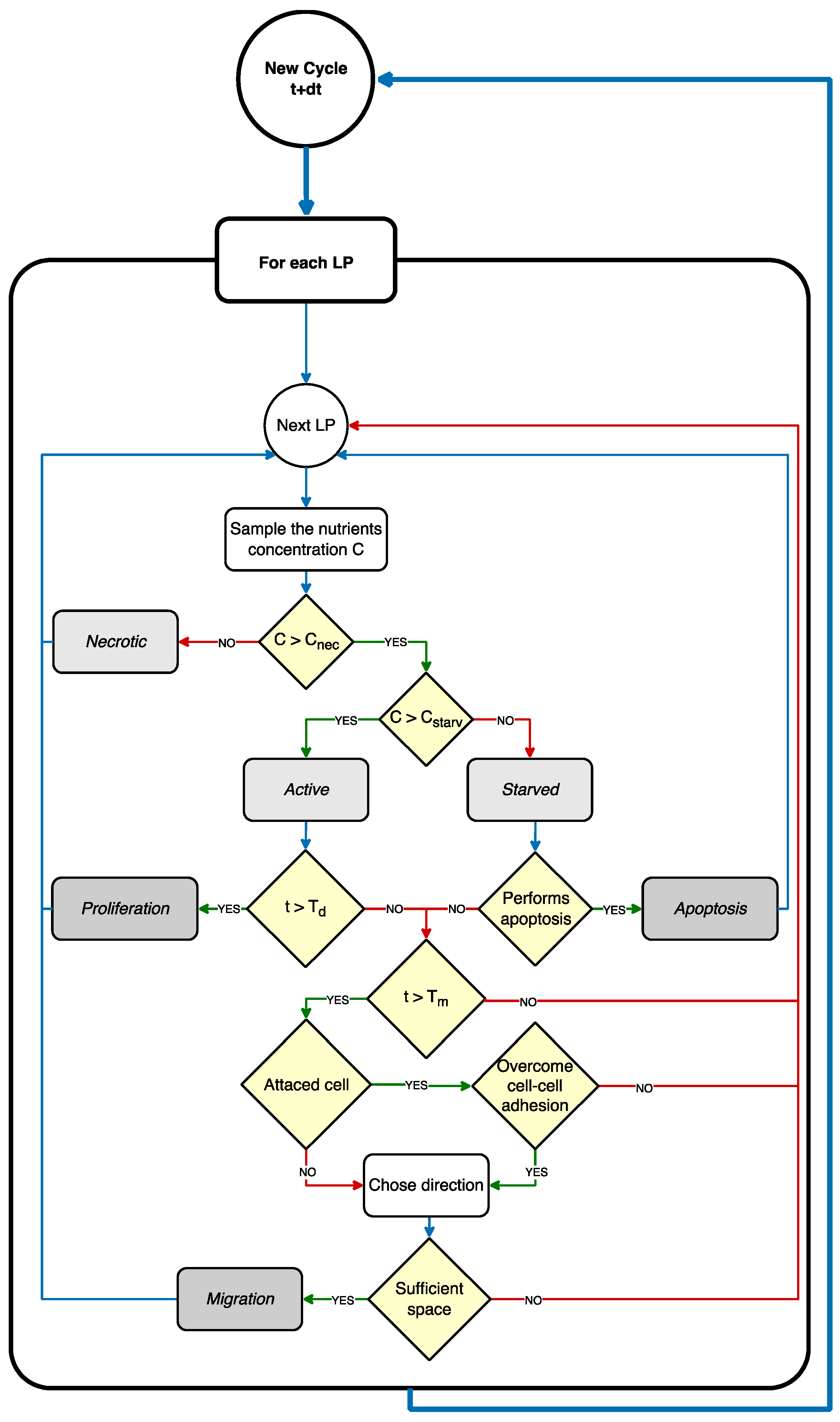
2.1.4. Rules Governing Cellular Dynamics
- (a)
- Cell migration
- (b)
- Cell proliferation
- (c)
- Cell death
2.1.5. Analysis
2.2. Statistical Analysis
2.2.1. Average and Variance Convergence
2.2.2. Global and Sensitivity Analyses
2.2.3. Local Sensitivity Analysis
3. Results
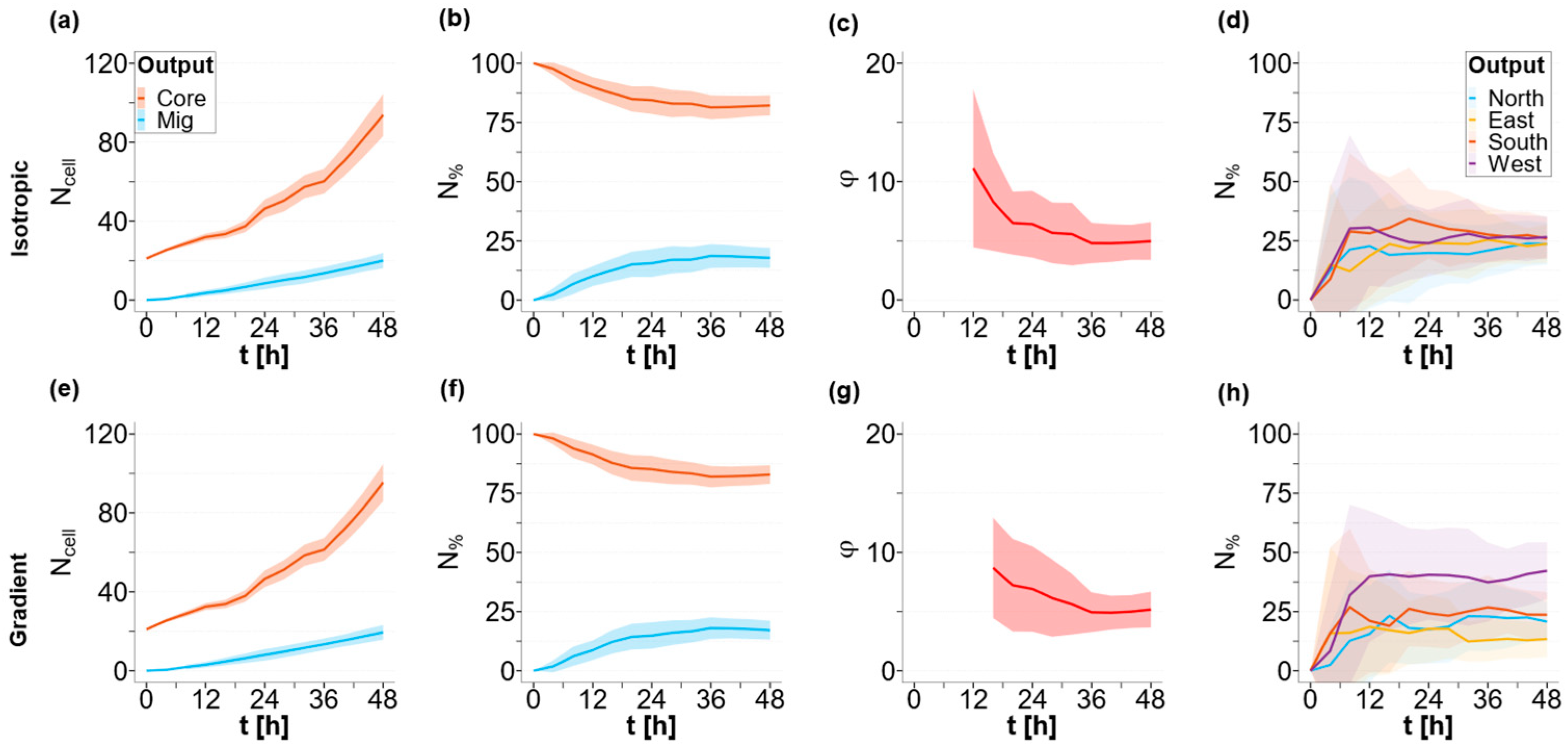
3.1. Baseline Case
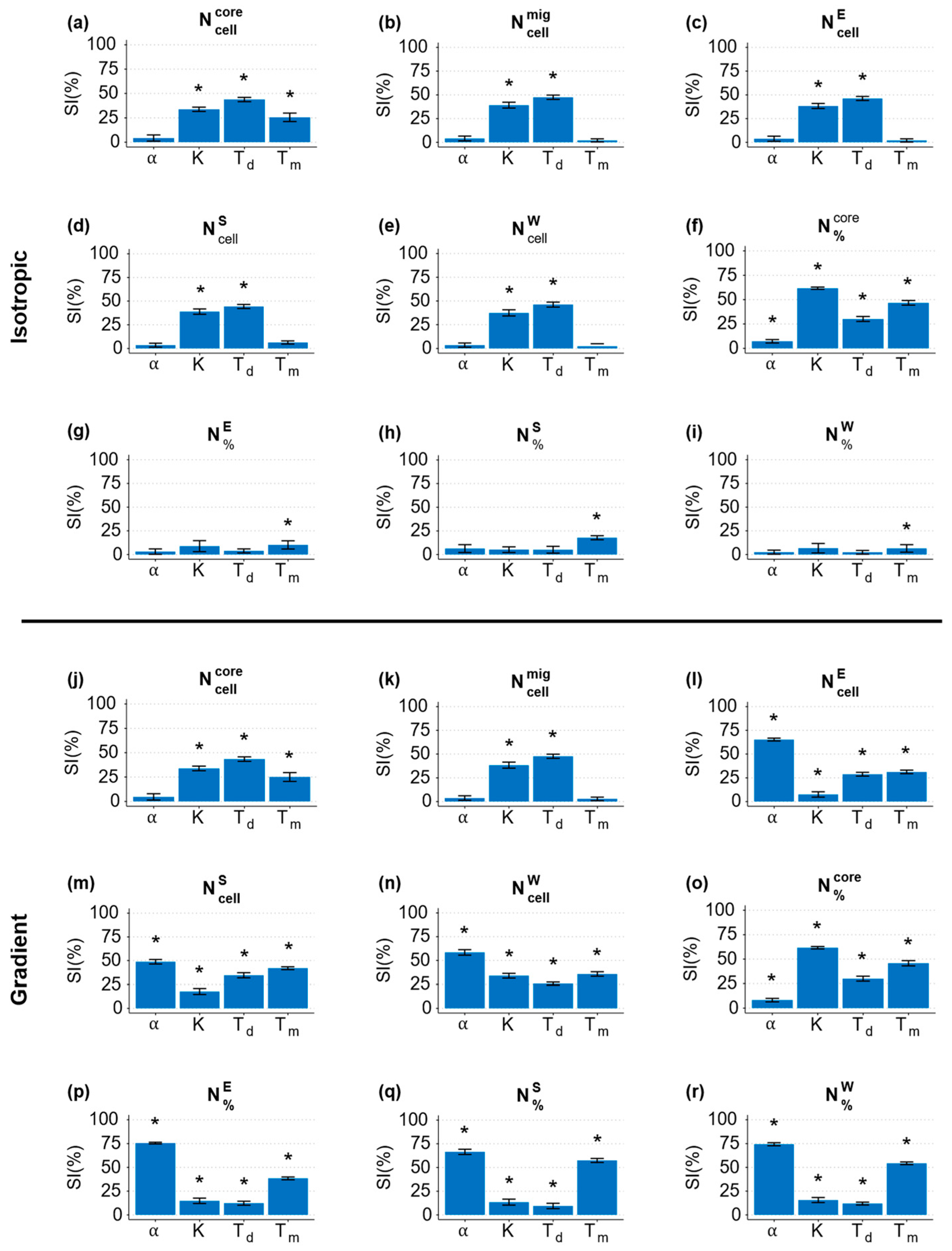
3.2. Global and Sensitivity Analysis (GSA)
3.3. Local Sensitivity Analysis (LSA)
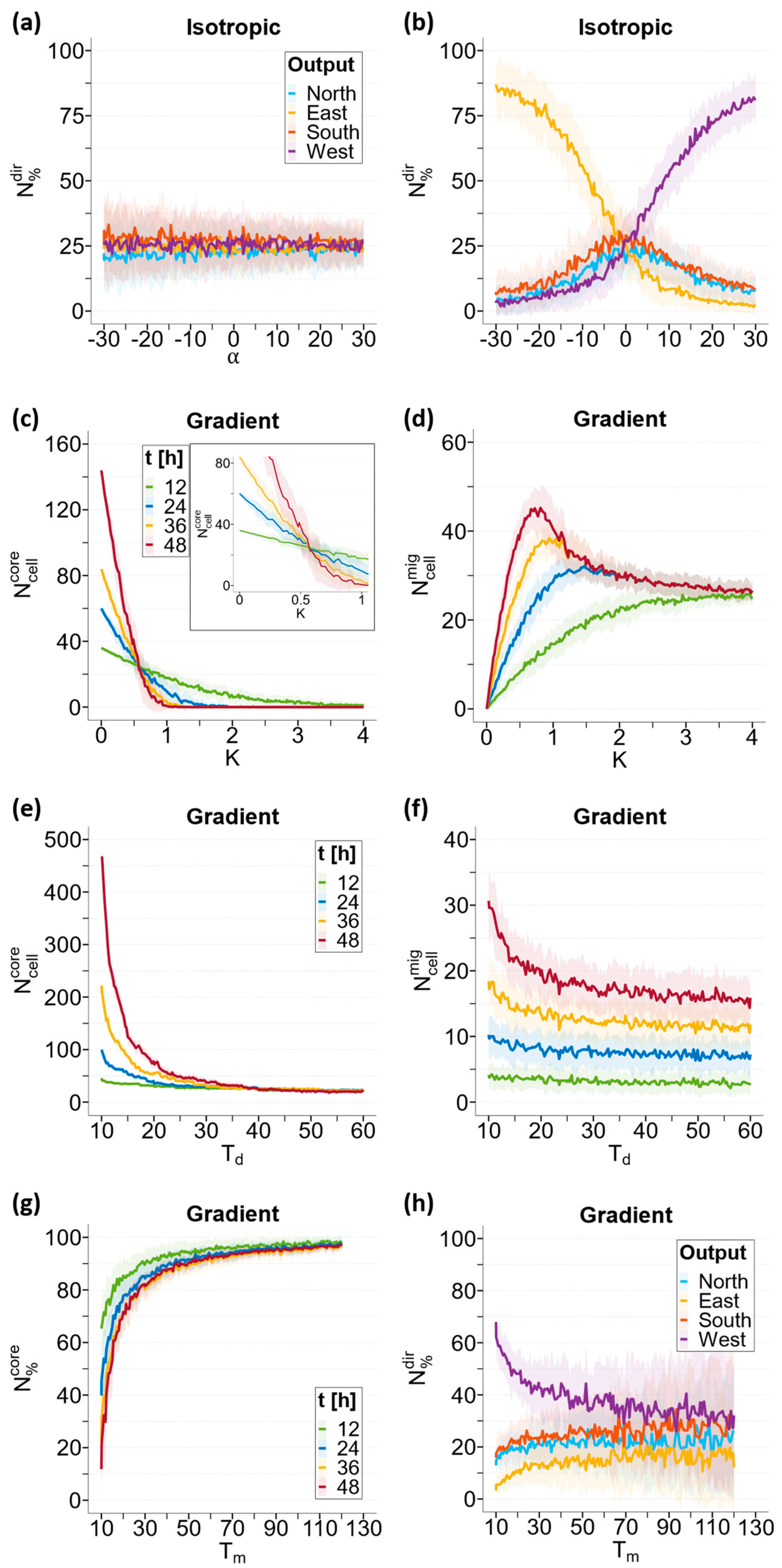
3.3.1. Chemotaxis Sensitivity Index α
3.3.2. Cell–Cell Adhesion Parameter K
3.3.3. Doubling Time Td
3.3.4. Migration Time Tm
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- World Health Organization. Global Health Estimates 2020: Deaths by Cause, Age, Sex, by Country and by Region, 2000–2019; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef]
- Dillekås, H.; Rogers, M.S.; Straume, O. Are 90% of deaths from cancer caused by metastases? Cancer Med. 2019, 8, 5574–5576. [Google Scholar] [CrossRef]
- Fares, J.; Fares, M.Y.; Khachfe, H.H.; Salhab, H.A.; Fares, Y. Molecular principles of metastasis: A hallmark of cancer revisited. Signal Transduct. Target. Ther. 2020, 5, 28. [Google Scholar] [CrossRef]
- Roussos, E.T.; Condeelis, J.S.; Patsialou, A. Chemotaxis in cancer. Nat. Rev. Cancer 2011, 11, 573–587. [Google Scholar] [CrossRef]
- Vasaturo, A.; Caserta, S.; Russo, I.; Preziosi, V.; Ciacci, C.; Guido, S. A novel chemotaxis assay in 3-D collagen gels by time-lapse microscopy. PLoS ONE 2012, 7, e52251. [Google Scholar] [CrossRef]
- Caserta, S.; Campello, S.; Tomaiuolo, G.; Sabetta, L.; Guido, S. A methodology to study chemotaxis in 3-D collagen gels. AIChE J. 2013, 59, 4025–4035. [Google Scholar] [CrossRef]
- Cristini, V.; Frieboes, H.B.; Gatenby, R.; Caserta, S.; Ferrari, M.; Sinek, J. Morphologic instability and cancer invasion. Clin. Cancer Res. 2005, 11, 6772–6779. [Google Scholar] [CrossRef]
- Bearer, E.L.; Lowengrub, J.S.; Frieboes, H.B.; Chuang, Y.-L.; Jin, F.; Wise, S.M.; Ferrari, M.; Agus, D.B.; Cristini, V. Multiparameter computational modeling of tumor invasion. Cancer Res. 2009, 69, 4493–4501. [Google Scholar] [CrossRef]
- Cristini, V.; Lowengrub, J. Multiscale Modeling of Cancer: An Integrated Experimental and Mathematical Modeling Approach; Cambridge University Press: Cambridge, UK, 2010. [Google Scholar]
- Sixt, M.; Lämmermann, T. In vitro analysis of chemotactic leukocyte migration in 3D environments. In Cell Migration; Springer: Berlin/Heidelberg, Germany, 2011; pp. 149–165. [Google Scholar]
- Lämmermann, T.; Bader, B.L.; Monkley, S.J.; Worbs, T.; Wedlich-Söldner, R.; Hirsch, K.; Keller, M.; Förster, R.; Critchley, D.R.; Fässler, R. Rapid leukocyte migration by integrin-independent flowing and squeezing. Nature 2008, 453, 51–55. [Google Scholar] [CrossRef]
- Friedl, P.; Bröcker, E.B. The biology of cell locomotion within three-dimensional extracellular matrix. Cell. Mol. Life Sci. CMLS 2000, 57, 41–64. [Google Scholar] [CrossRef] [PubMed]
- Sant, S.; Johnston, P.A. The production of 3D tumor spheroids for cancer drug discovery. Drug Discov. Today Technol. 2017, 23, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.W.; Leong, D.T.W.; Hutmacher, D.W. The challenge to measure cell proliferation in two and three dimensions. Tissue Eng. 2005, 11, 182–191. [Google Scholar] [CrossRef]
- Rhodes, N.P.; Srivastava, J.K.; Smith, R.F.; Longinotti, C. Metabolic and histological analysis of mesenchymal stem cells grown in 3-D hyaluronan-based scaffolds. J. Mater. Sci. Mater. Med. 2004, 15, 391–395. [Google Scholar] [CrossRef]
- Nyga, A.; Cheema, U.; Loizidou, M. 3D tumour models: Novel in vitro approaches to cancer studies. J. Cell Commun. Signal. 2011, 5, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.M.; Cukierman, E. Modeling tissue morphogenesis and cancer in 3D. Cell 2007, 130, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Hoarau-Véchot, J.; Rafii, A.; Touboul, C.; Pasquier, J. Halfway between 2D and animal models: Are 3D cultures the ideal tool to study cancer-microenvironment interactions? Int. J. Mol. Sci. 2018, 19, 181. [Google Scholar] [CrossRef]
- Li, C.K.N. The glucose distribution in 9L rat brain multicell tumor spheroids and its effect on cell necrosis. Cancer 1982, 50, 2066–2073. [Google Scholar] [CrossRef]
- Kam, Y.; Rejniak, K.A.; Anderson, A.R.A. Cellular modeling of cancer invasion: Integration of in silico and in vitro approaches. J. Cell. Physiol. 2012, 227, 431–438. [Google Scholar] [CrossRef]
- Stillman, N.R.; Kovacevic, M.; Balaz, I.; Hauert, S. In silico modelling of cancer nanomedicine, across scales and transport barriers. NPJ Comput. Mater. 2020, 6, 92. [Google Scholar] [CrossRef]
- Karolak, A.; Rejniak, K.A. Micropharmacology: An in silico approach for assessing drug efficacy within a tumor tissue. Bull. Math. Biol. 2019, 81, 3623–3641. [Google Scholar] [CrossRef]
- Dogra, P.; Butner, J.D.; Chuang, Y.-l.; Caserta, S.; Goel, S.; Brinker, C.J.; Cristini, V.; Wang, Z. Mathematical modeling in cancer nanomedicine: A review. Biomed. Microdevices 2019, 21, 40. [Google Scholar] [CrossRef]
- Burdett, E.; Kasper, F.K.; Mikos, A.G.; Ludwig, J.A. Engineering tumors: A tissue engineering perspective in cancer biology. Tissue Eng. Part B Rev. 2010, 16, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Norton, K.-A.; Gong, C.; Jamalian, S.; Popel, A.S. Multiscale agent-based and hybrid modeling of the tumor immune microenvironment. Processes 2019, 7, 37. [Google Scholar] [CrossRef] [PubMed]
- Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures–a comparison of different types of cancer cell cultures. Arch. Med. Sci. AMS 2018, 14, 910. [Google Scholar] [CrossRef]
- Patel, A.A.; Gawlinski, E.T.; Lemieux, S.K.; Gatenby, R.A. A cellular automaton model of early tumor growth and invasion: The effects of native tissue vascularity and increased anaerobic tumor metabolism. J. Theor. Biol. 2001, 213, 315–331. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, L.; Sagotsky, J.; Deisboeck, T.S. Simulating non-small cell lung cancer with a multiscale agent-based model. Theor. Biol. Med. Model. 2007, 4, 50. [Google Scholar] [CrossRef]
- Costa, E.C.; Moreira, A.F.; de Melo-Diogo, D.; Gaspar, V.M.; Carvalho, M.P.; Correia, I.J. 3D tumor spheroids: An overview on the tools and techniques used for their analysis. Biotechnol. Adv. 2016, 34, 1427–1441. [Google Scholar] [CrossRef]
- Chang, S.L.; Cavnar, S.P.; Takayama, S.; Luker, G.D.; Linderman, J.J. Cell, isoform, and environment factors shape gradients and modulate chemotaxis. PLoS ONE 2015, 10, e0123450. [Google Scholar] [CrossRef]
- Gatenby, R.A.; Gawlinski, E.T. A reaction-diffusion model of cancer invasion. Cancer Res. 1996, 56, 5745–5753. [Google Scholar]
- Masur, K.; Vetter, C.; Hinz, A.; Tomas, N.; Henrich, H.; Niggemann, B.; Zänker, K.S. Diabetogenic glucose and insulin concentrations modulate transcriptom and protein levels involved in tumour cell migration, adhesion and proliferation. Br. J. Cancer 2011, 104, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Lamers, M.L.; Almeida, M.E.S.; Vicente-Manzanares, M.; Horwitz, A.F.; Santos, M.F. High glucose-mediated oxidative stress impairs cell migration. PLoS ONE 2011, 6, e22865. [Google Scholar] [CrossRef] [PubMed]
- Palorini, R.; Votta, G.; Pirola, Y.; De Vitto, H.; De Palma, S.; Airoldi, C.; Vasso, M.; Ricciardiello, F.; Lombardi, P.P.; Cirulli, C. Protein kinase A activation promotes cancer cell resistance to glucose starvation and anoikis. PLoS Genet. 2016, 12, e1005931. [Google Scholar] [CrossRef] [PubMed]
- Leithner, K.; Hrzenjak, A.; Trötzmüller, M.; Moustafa, T.; Köfeler, H.C.; Wohlkoenig, C.; Stacher, E.; Lindenmann, J.; Harris, A.L.; Olschewski, A. PCK2 activation mediates an adaptive response to glucose depletion in lung cancer. Oncogene 2015, 34, 1044–1050. [Google Scholar] [CrossRef] [PubMed]
- Ferraro, R.; Ascione, F.; Dogra, P.; Cristini, V.; Guido, S.; Caserta, S. Diffusion-induced anisotropic cancer invasion: A novel experimental method based on tumor spheroids. AIChE J. 2022, 68, e17678. [Google Scholar] [CrossRef]
- Levenspiel, O. Chemical Reaction Engineering; John Wiley & Sons: Hoboken, NJ, USA, 1998. [Google Scholar]
- Schornack, P.A.; Gillies, R.J. Contributions of cell metabolism and H+ diffusion to the acidic pH of tumors. Neoplasia 2003, 5, 135–145. [Google Scholar] [CrossRef]
- Morana, O.; Wood, W.; Gregory, C.D. The apoptosis paradox in cancer. Int. J. Mol. Sci. 2022, 23, 1328. [Google Scholar] [CrossRef]
- Dogra, P.; Butner, J.D.; Ramírez, J.R.; Chuang, Y.-l.; Noureddine, A.; Brinker, C.J.; Cristini, V.; Wang, Z. A mathematical model to predict nanomedicine pharmacokinetics and tumor delivery. Comput. Struct. Biotechnol. J. 2020, 18, 518–531. [Google Scholar] [CrossRef]
- Saltelli, A. Global sensitivity analysis: An introduction. In Proceedings of the 4th International Conference on Sensitivity Analysis of Model Output (SAMO 2004), Santa Fe, NM, USA, 8–11 March 2004. [Google Scholar]
- Wang, Z.; Deisboeck, T.S.; Cristini, V. Development of a sampling-based global sensitivity analysis workflow for multiscale computational cancer models. IET Syst. Biol. 2014, 8, 191–197. [Google Scholar] [CrossRef]
- Pollock, K.; Stroemer, P.; Patel, S.; Stevanato, L.; Hope, A.; Miljan, E.; Dong, Z.; Hodges, H.; Price, J.; Sinden, J.D. A conditionally immortal clonal stem cell line from human cortical neuroepithelium for the treatment of ischemic stroke. Exp. Neurol. 2006, 199, 143–155. [Google Scholar] [CrossRef]
- Herlyn, M.; Steplewski, Z.; Herlyn, D.; Koprowski, H. Colorectal carcinoma-specific antigen: Detection by means of monoclonal antibodies. Proc. Natl. Acad. Sci. USA 1979, 76, 1438–1442. [Google Scholar] [CrossRef] [PubMed]
- Milo, R.; Phillips, R. Cell Biology by the Numbers; Garland Science: New York, NY, USA, 2015. [Google Scholar]

| Output Variable | Formula | Description |
|---|---|---|
| Number of cells adhered to the spheroid | ||
| Number of cells that migrated away from the spheroid | ||
| Total number of cells | ||
| Percentage of cells adhered to the spheroid | ||
| Percentage of cells that migrated away from the spheroid | ||
| Ratio of adhered cells to migrated cells | ||
| Number of migrated cells located in the North quadrant | ||
| Number of migrated cells located in the South quadrant | ||
| Number of migrated cells located in the East quadrant | ||
| Number of migrated cells located in the West quadrant | ||
| Percentage of migrated cells located in the North quadrant | ||
| Percentage of migrated cells located in the South quadrant | ||
| Percentage of migrated cells located in the East quadrant | ||
| Percentage of migrated cells located in the West quadrant |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Messina, L.; Ferraro, R.; Peláez, M.J.; Wang, Z.; Cristini, V.; Dogra, P.; Caserta, S. Hybrid Cellular Automata Modeling Reveals the Effects of Glucose Gradients on Tumour Spheroid Growth. Cancers 2023, 15, 5660. https://doi.org/10.3390/cancers15235660
Messina L, Ferraro R, Peláez MJ, Wang Z, Cristini V, Dogra P, Caserta S. Hybrid Cellular Automata Modeling Reveals the Effects of Glucose Gradients on Tumour Spheroid Growth. Cancers. 2023; 15(23):5660. https://doi.org/10.3390/cancers15235660
Chicago/Turabian StyleMessina, Luca, Rosalia Ferraro, Maria J. Peláez, Zhihui Wang, Vittorio Cristini, Prashant Dogra, and Sergio Caserta. 2023. "Hybrid Cellular Automata Modeling Reveals the Effects of Glucose Gradients on Tumour Spheroid Growth" Cancers 15, no. 23: 5660. https://doi.org/10.3390/cancers15235660
APA StyleMessina, L., Ferraro, R., Peláez, M. J., Wang, Z., Cristini, V., Dogra, P., & Caserta, S. (2023). Hybrid Cellular Automata Modeling Reveals the Effects of Glucose Gradients on Tumour Spheroid Growth. Cancers, 15(23), 5660. https://doi.org/10.3390/cancers15235660








