Soft Tissue Sarcoma Study: Association of Genetic Alterations in the Apoptosis Pathways with Chemoresistance to Doxorubicin
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Tumor Specimens
2.2. Primary Cancer Cells Cultures and Chemosensitivity Assay
2.3. Cytology for STS Primary Cultures
2.4. Bioinformatic and Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lazar, A.J.; McLellan, M.D.; Bailey, M.H.; Miller, C.A.; Appelbaum, E.L.; Cordes, M.G.; Lichtenberg, T.M. Cancer Genome Atlas Research Network. Electronic address: Elizabeth.demicco@sinaihealthsystem.ca; Cancer Genome Atlas Research Network Comprehensive and Integrated Genomic Characterization of Adult Soft Tissue Sarcomas. Cell 2017, 171, 950–965. [Google Scholar] [CrossRef]
- Gronchi, A.; Miah, A.B.; Dei Tos, A.P.; Abecassis, N.; Bajpai, J.; Bauer, S.; Biagini, R.; Bielack, S.; Blay, J.Y.; Bolle, S.; et al. ESMO Guidelines Committee, EURACAN and GENTURIS. Electronic address: Clinicalguidelines@esmo.org. Soft tissue and visceral sarcomas: ESMO-EURACAN-GENTURIS Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2021, 32, 1348–1365. [Google Scholar] [CrossRef] [PubMed]
- Judson, I.; Verweij, J.; Gelderblom, H.; Hartmann, J.T.; Schöffski, P.; Blay, J.-Y.; Kerst, J.M.; Sufliarsky, J.; Whelan, J.; Hohenberger, P.; et al. Doxorubicin alone versus intensified doxorubicin plus ifosfamide for first-line treatment of advanced or metastatic soft-tissue sarcoma: A randomised controlled phase 3 trial. Lancet Oncol. 2014, 15, 415–423. [Google Scholar] [CrossRef]
- The WHO Classification of Tumours Editorial Board. Soft Tissue and Bone Tumours WHO Classification of Tumours, 5th ed.; IARC Press: Lyon, France, 2020. [Google Scholar]
- Thorn, C.F.; Oshiro, C.; Marsh, S.; Hernandez-Boussard, T.; McLeod, H.; Klein, T.E.; Altman, R.B. Doxorubicin pathways: Pharmacodynamics and adverse effects. Pharm. Genom. 2011, 21, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Tacar, O.; Sriamornsak, P.; Dass, C.R. Doxorubicin: An update on anticancer molecular action, toxicity and novel drug delivery systems. J. Pharm. Pharmacol. 2013, 65, 157–170. [Google Scholar] [CrossRef]
- Gangireddy, M.; Nookala, V. Ifosfamide; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Heinemann, V.; Xu, Y.Z.; Chubb, S.; Sen, A.; Hertel, L.W.; Grindey, G.B.; Plunkett, W. Cellular elimination of 2′,2′-difluorodeoxycytidine 5′-triphosphate: A mechanism of self-potentiation. Cancer Res. 1992, 52, 533–539. [Google Scholar]
- Cunha, K.S. Taxanes: The genetic toxicity of paclitaxel and docetaxel in somatic cells of Drosophila melanogaster. Mutagenesis 2001, 16, 79–84. [Google Scholar] [CrossRef][Green Version]
- Williams, A.B.; Schumacher, B. p53 in the DNA-Damage-Repair Process. Cold Spring Harb. Perspect. Med. 2016, 6, a026070. [Google Scholar] [CrossRef]
- Thoenen, E.; Curl, A.; Iwakuma, T. TP53 in bone and soft tissue sarcomas. Pharmacol. Ther. 2019, 202, 149–164. [Google Scholar] [CrossRef]
- Cote, G.M.; He, J.; Choy, E. Next-Generation Sequencing for Patients with Sarcoma: A Single Center Experience. Oncologist 2018, 23, 234–242. [Google Scholar] [CrossRef]
- Zhan, M.; Yu, D.; Lang, A.; Li, L.; Pollock, R.E. Wild type p53 sensitizes soft tissue sarcoma cells to doxorubicin by down-regulating multidrug resistance-1 expression. Cancer 2001, 92, 1556–1566. [Google Scholar] [CrossRef]
- Lee, Y.S.; Choi, J.-Y.; Lee, J.; Shim, D.M.; Kim, J.; Park, W.-Y.; Nam, D.-H.; Seo, S.W. TP53-dependence on the effect of doxorubicin and Src inhibitor combination therapy. Tumor Biol. 2018, 40, 101042831879421. [Google Scholar] [CrossRef] [PubMed]
- Kurbacher, C.M.; Cree, I.A.; Brenne, U.; Bruckner, H.W.; Kurbacher, J.A.; Mallmann, P.; Andreotti, P.E.; Krebs, D. Heterogeneity of in vitro chemosensitivity in perioperative breast cancer cells to mitoxantrone versus doxorubicin evaluated by a microplate ATP bioluminescence assay. Breast Cancer Res. Treat. 1996, 41, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Morand du Puch, C.B.; Vanderstraete, M.; Giraud, S.; Lautrette, C.; Christou, N.; Mathonnet, M. Benefits of functional assays in personalized cancer medicine: More than just a proof-of-concept. Theranostics 2021, 11, 9538–9556. [Google Scholar] [CrossRef] [PubMed]
- Xia, F.; Ma, S.; Bian, Y.; Yu, D.; Ma, W.; Miao, M.; Huang, C.; Miao, L. A retrospective study of the correlation of in vitro chemosensitivity using ATP-TCA with patient clinical outcomes in acute myeloid leukemia. Cancer Chemother. Pharmacol. 2020, 85, 509–515. [Google Scholar] [CrossRef]
- Zhang, J.; Li, H. Heterogeneity of tumor chemosensitivity in ovarian epithelial cancer revealed using the adenosine triphosphate-tumor chemosensitivity assay. Oncol. Lett. 2015, 9, 2374–2380. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, S.; Ma, S.; Li, C.; Xu, C.; Shen, Y.; Zhao, J.; Miao, L. Evaluation of the in vitro Chemosensitivity and Correlation with Clinical Outcomes in Lung Cancer using the ATP-TCA. Anticancer Agents Med. Chem. 2018, 18, 139–145. [Google Scholar] [CrossRef]
- Sargent, J.M.; Taylor, C.G. Appraisal of the MTT assay as a rapid test of chemosensitivity in acute myeloid leukaemia. Br. J. Cancer 1989, 60, 206–210. [Google Scholar] [CrossRef]
- Bosserman, L.; Prendergast, F.; Herbst, R.; Fleisher, M.; Salom, E.; Strickland, S.; Raptis, A.; Hallquist, A.; Perree, M.; Rajurkar, S.; et al. The microculture-kinetic (MiCK) assay: The role of a drug-induced apoptosis assay in drug development and clinical care. Cancer Res. 2012, 72, 3901–3905. [Google Scholar] [CrossRef][Green Version]
- Tapias, L.F.; Gilpin, S.E.; Ren, X.; Wei, L.; Fuchs, B.C.; Tanabe, K.K.; Lanuti, M.; Ott, H.C. Assessment of Proliferation and Cytotoxicity in a Biomimetic Three-Dimensional Model of Lung Cancer. Ann. Thorac. Surg. 2015, 100, 414–421. [Google Scholar] [CrossRef]
- Andreotti, P.E.; Cree, I.A.; Kurbacher, C.M.; Hartmann, D.M.; Linder, D.; Harel, G.; Gleiberman, I.; Caruso, P.A.; Ricks, S.H.; Untch, M. Chemosensitivity testing of human tumors using a microplate adenosine triphosphate luminescence assay: Clinical correlation for cisplatin resistance of ovarian carcinoma. Cancer Res. 1995, 55, 5276–5282. [Google Scholar] [PubMed]
- Fischer, A.H.; Zhao, C.; Li, Q.K.; Gustafson, K.S.; Eltoum, I.E.; Tambouret, R.; Benstein, B.; Savaloja, L.C.; Kulesza, P. The cytologic criteria of malignancy. J. Cell Biochem. 2010, 110, 795–811. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118, iso-2; iso-3. Fly (Austin) 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Geoffroy, V.; Herenger, Y.; Kress, A.; Stoetzel, C.; Piton, A.; Dollfus, H.; Muller, J. AnnotSV: An integrated tool for structural variations annotation. Bioinformatics 2018, 34, 3572–3574. [Google Scholar] [CrossRef]
- Dees, N.D.; Zhang, Q.; Kandoth, C.; Wendl, M.C.; Schierding, W.; Koboldt, D.C.; Mooney, T.B.; Callaway, M.B.; Dooling, D.; Mardis, E.R.; et al. MuSiC: Identifying mutational significance in cancer genomes. Genome Res. 2012, 22, 1589–1598. [Google Scholar] [CrossRef]
- Mayakonda, A.; Lin, D.-C.; Assenov, Y.; Plass, C.; Koeffler, H.P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018, 28, 1747–1756. [Google Scholar] [CrossRef]
- Miller, M.; Shirole, N.; Tian, R.; Pal, D.; Sordella, R. The Evolution of TP53 Mutations: From Loss-of-Function to Separation-of-Function Mutants. J. Cancer Biol. Res. 2016, 4. PMID: 28191499. [Google Scholar]
- Trostel, S.Y.; Sackett, D.L.; Fojo, T. Oligomerization of p53 precedes its association with dynein and nuclear accumulation. Cell Cycle 2006, 5, 2253–2259. [Google Scholar] [CrossRef] [PubMed]
- Clarke, S.L.; Thompson, L.R.; Dandekar, E.; Srinivasan, A.; Montgomery, M.R. Distinct TP53 Mutation Subtypes Differentially Influence Cellular Iron Metabolism. Nutrients 2019, 11, 2144. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, T.; Hermann, E.; Lin, D.; Chowanadisai, W.; Hull, E.; Montgomery, M. HDAC inhibition induces EMT and alterations in cellular iron homeostasis to augment ferroptosis sensitivity in SW13 cells. Redox Biol. 2021, 47, 102149. [Google Scholar] [CrossRef] [PubMed]
- Kamada, R.; Nomura, T.; Anderson, C.W.; Sakaguchi, K. Cancer-associated p53 tetramerization domain mutants: Quantitative analysis reveals a low threshold for tumor suppressor inactivation. J. Biol. Chem. 2011, 286, 252–258. [Google Scholar] [CrossRef]
- Banin, S.; Moyal, L.; Shieh, S.; Taya, Y.; Anderson, C.W.; Chessa, L.; Smorodinsky, N.I.; Prives, C.; Reiss, Y.; Shiloh, Y.; et al. Enhanced phosphorylation of p53 by ATM in response to DNA damage. Science 1998, 281, 1674–1677. [Google Scholar] [CrossRef]
- Guarini, A.; Marinelli, M.; Tavolaro, S.; Bellacchio, E.; Magliozzi, M.; Chiaretti, S.; De Propris, M.S.; Peragine, N.; Santangelo, S.; Paoloni, F.; et al. ATM gene alterations in chronic lymphocytic leukemia patients induce a distinct gene expression profile and predict disease progression. Haematologica 2012, 97, 47–55. [Google Scholar] [CrossRef]
- Zhou, X.; Hao, Q.; Lu, H. Mutant p53 in cancer therapy-the barrier or the path. J. Mol. Cell Biol. 2019, 11, 293–305. [Google Scholar] [CrossRef]
- Duffy, M.J.; Synnott, N.C.; O’Grady, S.; Crown, J. Targeting p53 for the treatment of cancer. Semin. Cancer Biol. 2020, 79, 58–67. [Google Scholar] [CrossRef]
- Bykov, V.J.N.; Issaeva, N.; Shilov, A.; Hultcrantz, M.; Pugacheva, E.; Chumakov, P.; Bergman, J.; Wiman, K.G.; Selivanova, G. Restoration of the tumor suppressor function to mutant p53 by a low-molecular-weight compound. Nat. Med. 2002, 8, 282–288. [Google Scholar] [CrossRef]

| Drug/Combination | 100% TDC (mg/mL) |
|---|---|
| Doxorubicin | 1.0 |
| Ifosfamide (4-hydroxy-ifosfamide) | 3.0 |
| Doxorubicin + Ifosfamide | 1.0 + 3.0 |
| Docetaxel | 11.3 |
| Gemcitabine | 25.0 |
| Docetaxel + Gemcitabine | 11.3 + 25.0 |
| Characteristics | Percentage, % |
|---|---|
| Histological subtype SS UPS | 12 (32%) 25 (68%) |
| Age <40 >40 | 12 (32%) 25 (68%) |
| Sex Male Female | 19 (51%) 18 (49%) |
| Tumor grade G1–G2 G3 | 2 (6%) 35 (94%) |
| Stage I–II III–IV | 8 (22%) 29 (78%) |
| Newly diagnosed Recurrent | 23 (62%) 14 (38%) |
| NeoCT NeoRT Without neoadjuvant therapy | 20 (54%) 3 (8%) 14 (38%) |
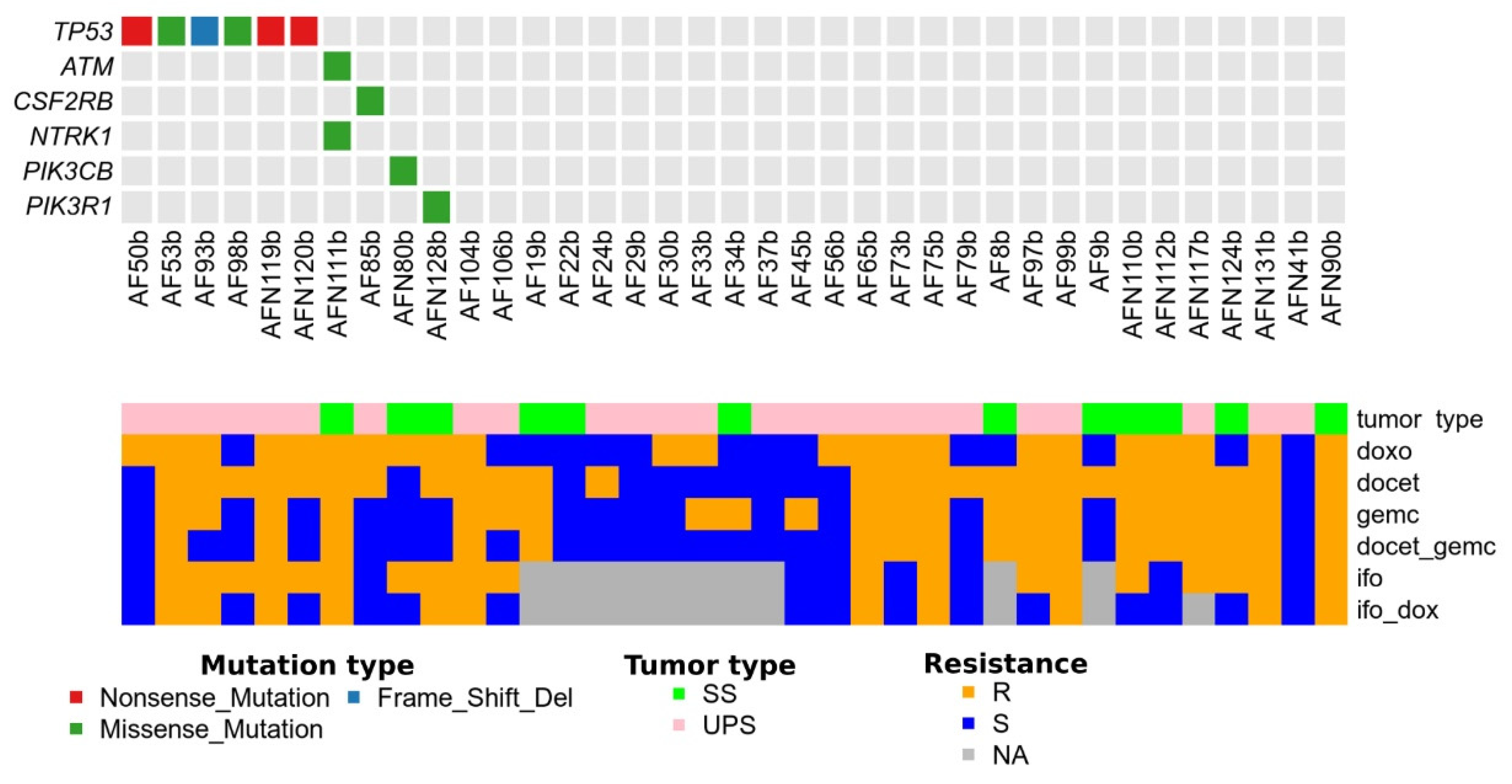
| No of Specimen | Gene | Specimen Code in Sequencing | Gene Mutation | Protein Mutation | Cosmic Database ID |
|---|---|---|---|---|---|
| AF50b | TP53 | tumor_50.GRCh38DH.exome | g.chr17:7675234G>C | p.Y126 * | COSM10862 |
| AF53b | TP53 | tumor_53.GRCh38DH.exome | g.chr17:7674954G>A | p.H193Y | COSM10672 |
| AF93b | TP53 | tumor_93.GRCh38DH.exome | g.chr17:7674191delC | p.E258fs | COSM7340859 |
| AF98b | TP53 | tumor_98.GRCh38DH.exome | g.chr17:7670678A>C | p.L344R | COSM46303 |
| AFN119b | TP53 | tumor_N119.GRCh38DH.exome | g.chr17:7670685G>A | p.R342 * | COSM11073 |
| AFN120b | TP53 | tumor_N120.GRCh38DH.exome | g.chr17:7675218T>A | p.K132 * | COSM44641 |
| AF85b | CSF2RB | tumor_85.GRCh38DH.exome | g.chr22:36936630A>G | p.S516G | |
| AFN111b | ATM | tumor_N111.GRCh38DH.exome | g.chr11:108335029C>T | p.R2691C | COSM922745 |
| NTRK1 | g.chr1:156879231G>T | p.V639L | |||
| AFN128b | PIK3R1 | tumor_N128.GRCh38DH.exome | g.chr5:68294573A>G | p.N125S | Analogue of COSM6960758 |
| AFN80b | PIK3CB | tumor_N80.GRCh38DH.exome | g.chr3:138714521T>C | p.T417A | Analogue of COSM419799 |
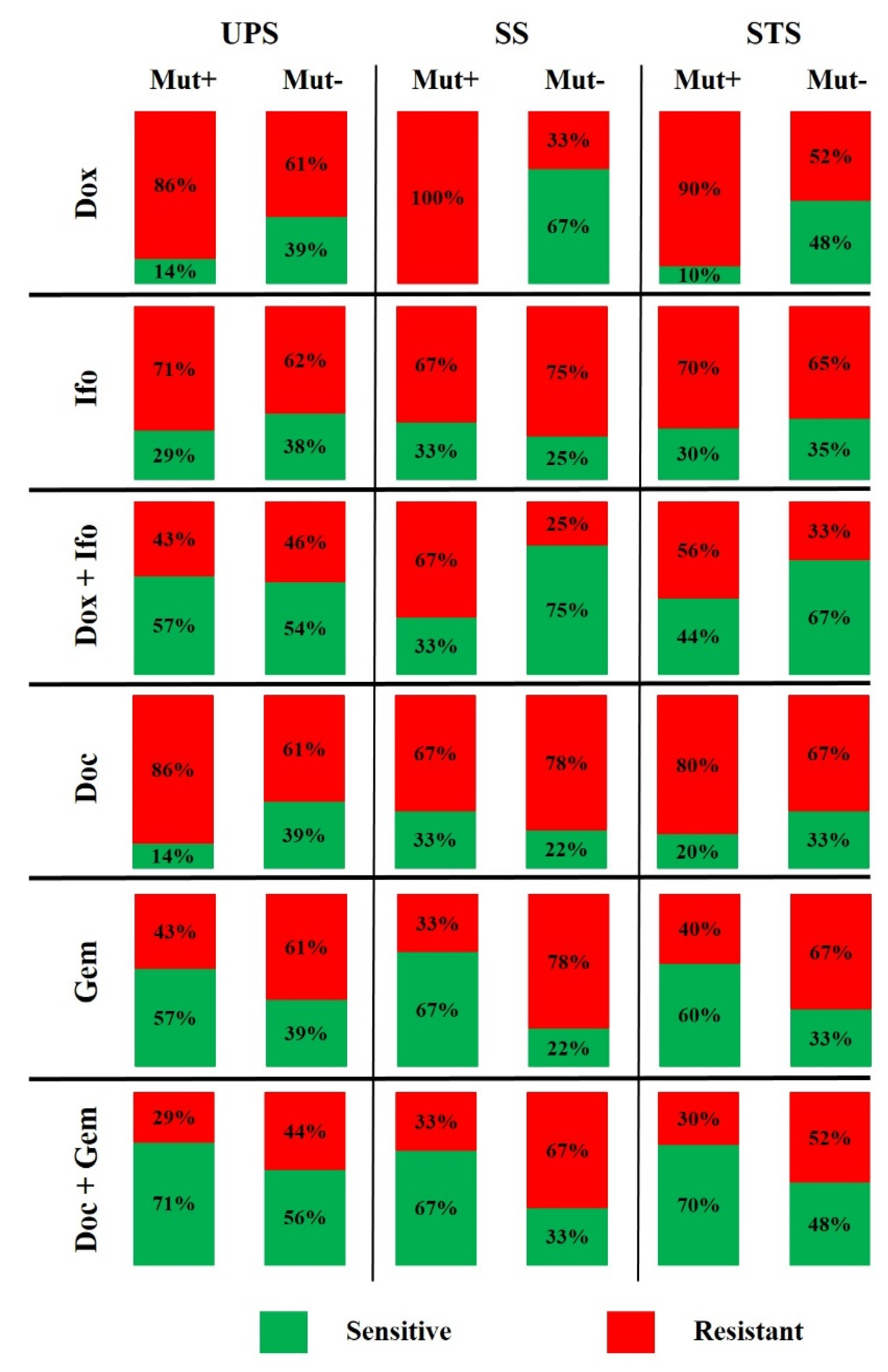
| Drug | Characteristics | UPS | SS | STS | |||
|---|---|---|---|---|---|---|---|
| 25 | 12 | 37 | |||||
| Mut+ | Mut− | Mut+ | Mut− | Mut+ | Mut− | ||
| 7 | 18 | 3 | 9 | 10 | 27 | ||
| Dox | Sensitive (n) | 1 | 7 | 0 | 6 | 1 | 13 |
| Resistant (n) | 6 | 11 | 3 | 3 | 9 | 14 | |
| p-value | 0.25 | 0.38 | 0.036 * | ||||
| Ifo | Sensitive (n) | 2 | 5 | 1 | 1 | 3 | 6 |
| Resistant (n) | 5 | 8 | 2 | 3 | 7 | 11 | |
| p-value | 0.52 | 0.71 | 0.56 | ||||
| Dox + Ifo | Sensitive (n) | 4 | 7 | 1 | 3 | 4 | 12 |
| Resistant (n) | 3 | 6 | 2 | 1 | 5 | 6 | |
| p-value | 0.63 | 0.37 | 0.24 | ||||
| Doc | Sensitive (n) | 1 | 7 | 1 | 2 | 2 | 9 |
| Resistant (n) | 6 | 11 | 2 | 7 | 8 | 18 | |
| p-value | 0.25 | 0.62 | 0.36 | ||||
| Gem | Sensitive (n) | 4 | 7 | 2 | 2 | 6 | 9 |
| Resistant (n) | 3 | 11 | 1 | 7 | 4 | 18 | |
| p-value | 0.35 | 0.24 | 0.14 | ||||
| Doc + Gem | Sensitive (n) | 5 | 10 | 2 | 3 | 7 | 13 |
| Resistant (n) | 2 | 8 | 1 | 6 | 3 | 14 | |
| p-value | 0.40 | 0.36 | 0.21 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kirilin, E.M.; Fetisov, T.I.; Moiseeva, N.I.; Lesovaya, E.A.; Laletina, L.A.; Makhmudova, L.F.; Manikaylo, A.E.; Fomina, L.Y.; Burov, D.A.; Bokhyan, B.Y.; et al. Soft Tissue Sarcoma Study: Association of Genetic Alterations in the Apoptosis Pathways with Chemoresistance to Doxorubicin. Cancers 2022, 14, 1796. https://doi.org/10.3390/cancers14071796
Kirilin EM, Fetisov TI, Moiseeva NI, Lesovaya EA, Laletina LA, Makhmudova LF, Manikaylo AE, Fomina LY, Burov DA, Bokhyan BY, et al. Soft Tissue Sarcoma Study: Association of Genetic Alterations in the Apoptosis Pathways with Chemoresistance to Doxorubicin. Cancers. 2022; 14(7):1796. https://doi.org/10.3390/cancers14071796
Chicago/Turabian StyleKirilin, Evgeny M., Timur I. Fetisov, Natalia I. Moiseeva, Ekaterina A. Lesovaya, Lidia A. Laletina, Leyla F. Makhmudova, Angelika E. Manikaylo, Liliya Y. Fomina, Denis A. Burov, Beniamin Yu. Bokhyan, and et al. 2022. "Soft Tissue Sarcoma Study: Association of Genetic Alterations in the Apoptosis Pathways with Chemoresistance to Doxorubicin" Cancers 14, no. 7: 1796. https://doi.org/10.3390/cancers14071796
APA StyleKirilin, E. M., Fetisov, T. I., Moiseeva, N. I., Lesovaya, E. A., Laletina, L. A., Makhmudova, L. F., Manikaylo, A. E., Fomina, L. Y., Burov, D. A., Bokhyan, B. Y., Zinovieva, V. Y., Vilkova, A. S., Mekheda, L. V., Kozlov, N. A., Scherbakov, A. M., Belitsky, G. A., Švedas, V., Kirsanov, K. I., & Yakubovskaya, M. G. (2022). Soft Tissue Sarcoma Study: Association of Genetic Alterations in the Apoptosis Pathways with Chemoresistance to Doxorubicin. Cancers, 14(7), 1796. https://doi.org/10.3390/cancers14071796







