Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Patient Information
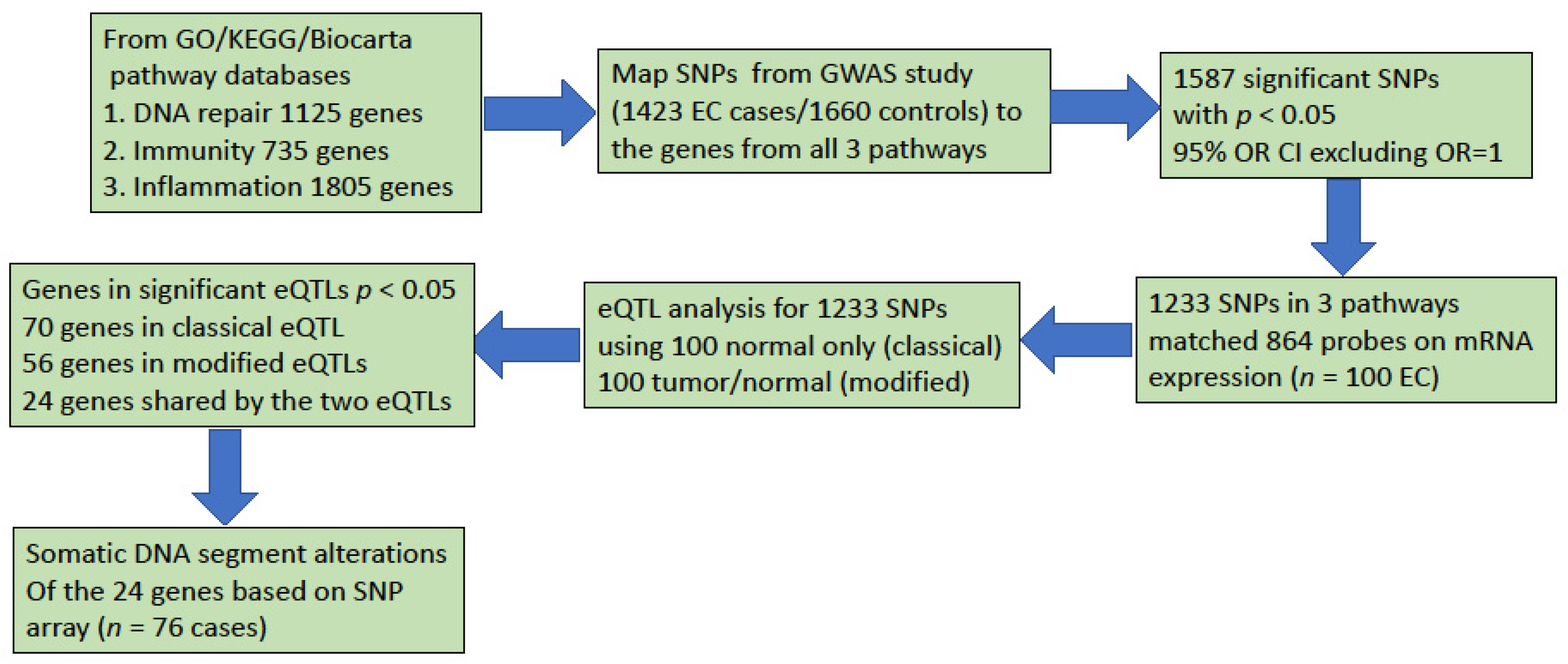
3.2. Pathway-Based Analyses
3.3. Two eQTL Analyses of Gene Expression and SNP Genotype
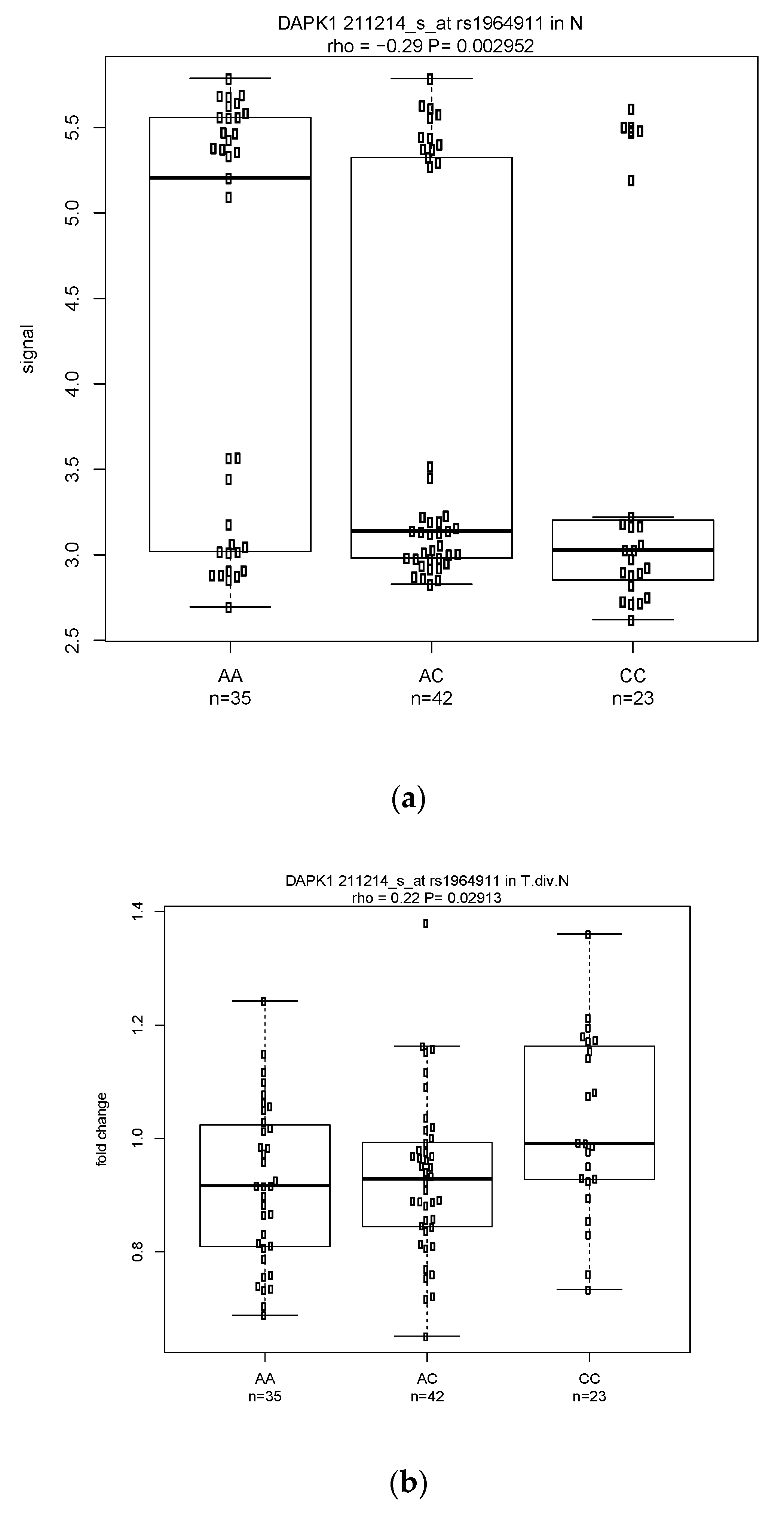
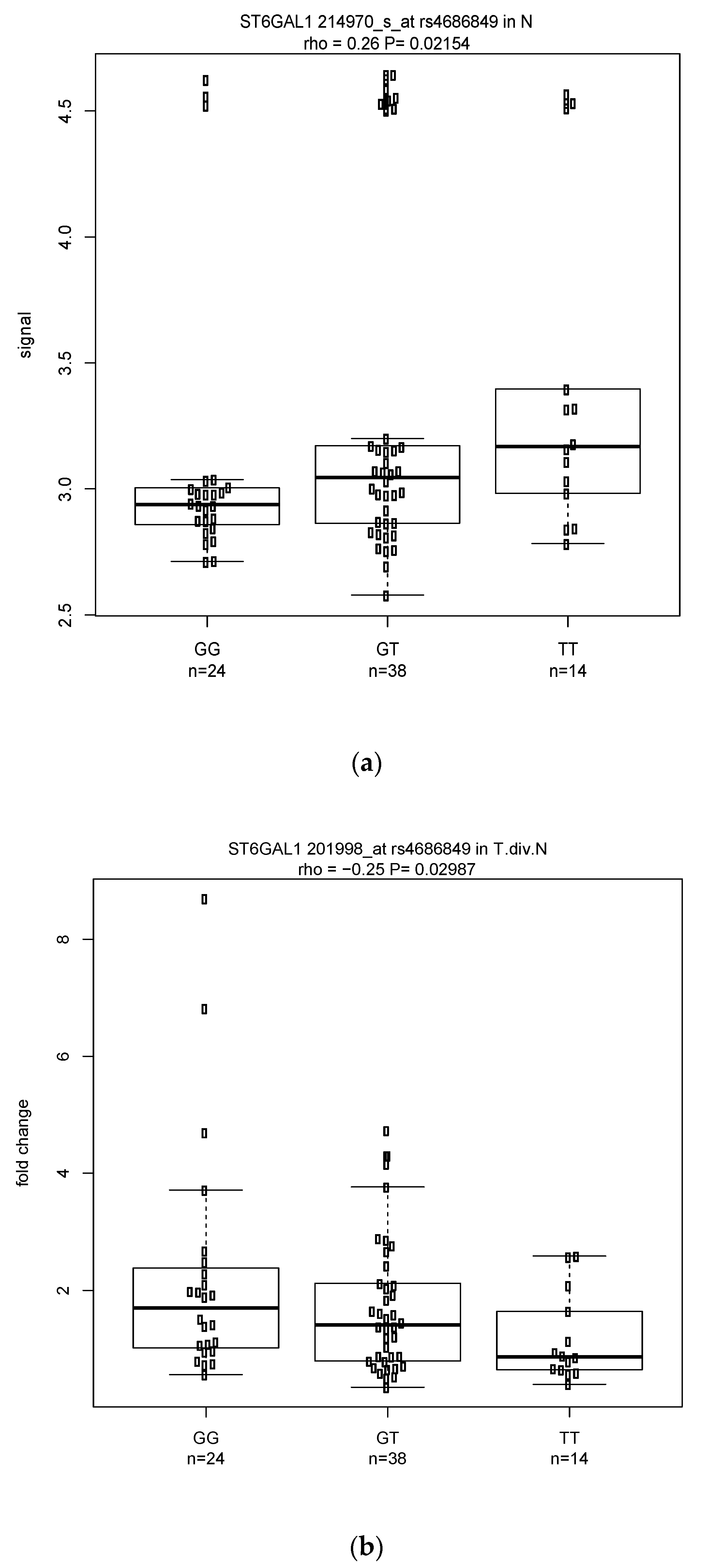
3.3.1. Classical eQTL Analysis (in Normal Esophagus Tissue Only)
3.3.2. Modified eQTL Analysis (Fold Change Based on Tumor vs. Normal Tissue)
3.3.3. Shared Significant eQTLs in the Classical and Modified Approaches
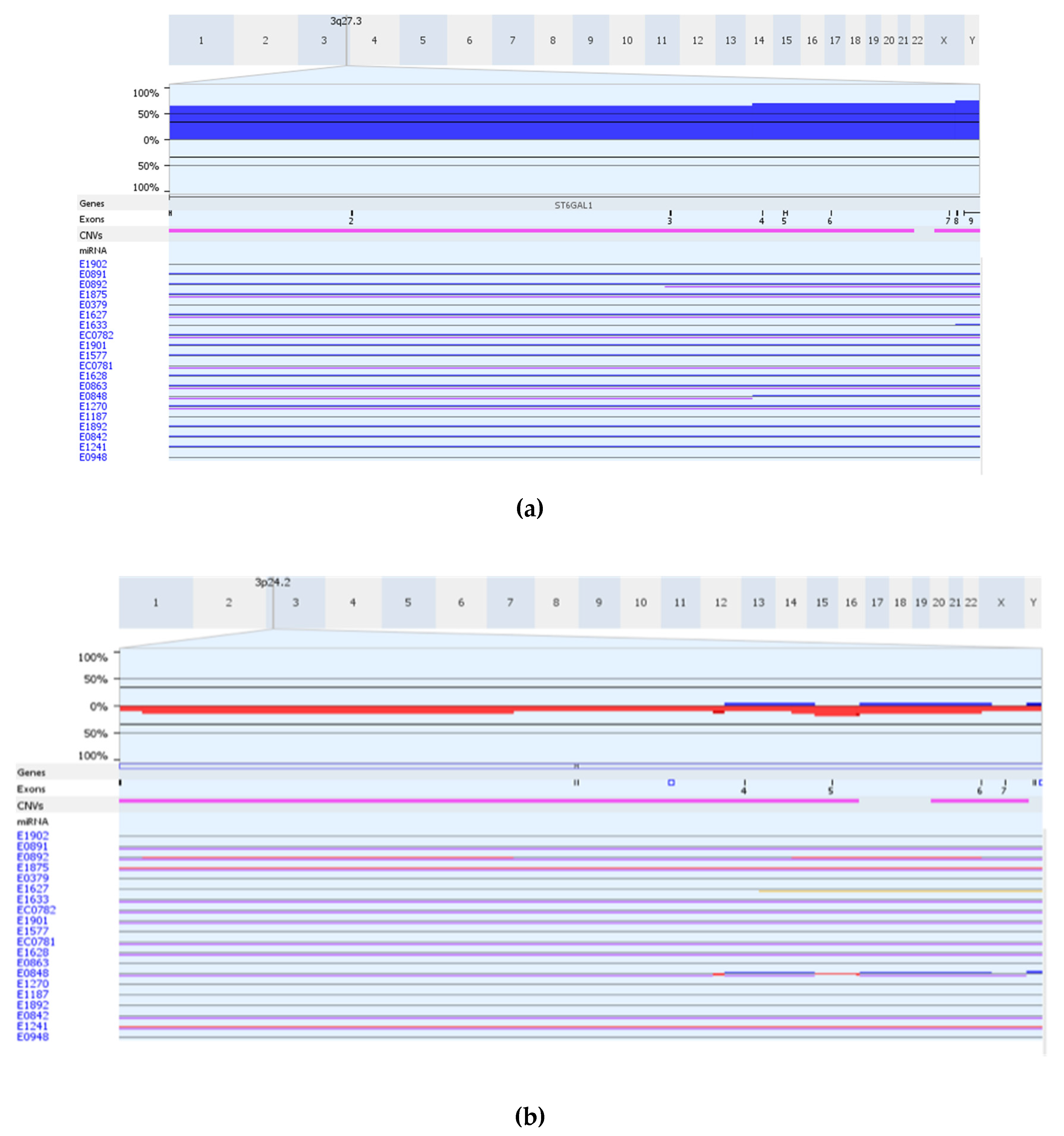
3.4. Somatic DNA Segment Alterations (Copy Number (CN) Alterations, Allele Imbalance (AI) and LOH) in Genes Shared by the Significant eQTLs from the Two Approaches
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix A.1. Biological Specimen Collection and Processing
Appendix A.1.1. U133A Arrays (U133A, U133A v.2.0, and U133 Plus2) and Data Normalization
Appendix A.1.2. GeneChip Human Mapping Arrays (500K, SNP 5.0, and SNP 6.0)
Appendix A.1.3. GeneChip Data Analysis (500K, SNP 5.0, and SNP 6.0 Arrays)
Appendix A.2. Pathway Resources
Appendix A.3. Association and eQTL Analyses
References
- Ferlay, J.; Shin, H.R.; Bray, F.; Forman, D.; Mathers, C.; Parkin, D.M. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int. J. Cancer 2010, 127, 2893–2917. [Google Scholar] [CrossRef]
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global Cancer Statistics, 2012. CA Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, W.; Zheng, R.; Baade, P.D.; Zhang, S.; Zeng, H.; Bray, F.; Jemal, A.; Yu, X.Q.; He, J. Cancer Statistics in China, 2015. CA Cancer J. Clin. 2016, 66, 115–132. [Google Scholar] [CrossRef] [Green Version]
- Li, J.Y. Epidemiology of esophageal cancer in China. Natl. Cancer Inst. Monogr. 1982, 62, 113–120. [Google Scholar]
- Hu, N.; Dawsey, S.M.; Wu, M.; Bonney, G.E.; He, L.J.; Han, X.Y.; Fu, M.; Taylor, P.R. Familial aggregation of esophageal cancer in Yangcheng County, Shanxi Province, China. Int. J. Epidemiol. 1992, 21, 877–882. [Google Scholar] [CrossRef]
- Hu, N.; Dawsey, S.M.; Wu, M.; Taylor, P.T. Family history of esophageal cancer in Shanxi province, China. Eur. J. Cancer 1991, 27, 1336. [Google Scholar] [CrossRef]
- Gao, Y.; Hu, N.; Han, X.Y.; Giffen, C.; Ding, T.; Goldstein, A.M.; Taylor, P.R. Family history of cancer and risk for esophageal and gastric cancer in Shanxi, China. BMC Cancer 2009, 9, 269. [Google Scholar] [CrossRef] [Green Version]
- Hu, N.; Roth, M.J.; Polymeropolous, M.; Tang, Z.Z.; Emmert-Buck, M.R.; Wang, Q.H.; Goldstein, A.M.; Feng, S.S.; Dawsey, S.M.; Ding, T.; et al. Identification of novel regions of allelic loss from a genomewide scan of esophageal squamous-cell carcinoma in a high-risk Chinese population. Genes Chromosomes Cancer 2000, 27, 217–228. [Google Scholar] [CrossRef]
- Hu, N.; Roth, M.J.; Emmert-Buck, M.R.; Tang, Z.Z.; Polymeropolous, M.; Wang, Q.H.; Goldstein, A.M.; Han, X.Y.; Dawsey, S.M.; Ding, T.; et al. Allelic loss in esophageal squamous cell carcinoma patients with and without family history of upper gastrointestinal tract cancer. Clin. Cancer Res. 1999, 5, 3476–3482. [Google Scholar]
- Liberzon, A.; Birger, C.; Thorvaldsdóttir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, N.; Wang, C.; Ng, D.; Clifford, R.; Yang, H.H.; Tang, Z.Z.; Wang, Q.H.; Han, X.Y.; Giffen, C.; Goldstein, A.M.; et al. Genomic characterization of esophageal squamous cell carcinoma from a high-risk population in China. Cancer Res. 2009, 69, 5908–5917. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, N.; Clifford, R.J.; Yang, H.H.; Wang, C.; Goldstein, A.M.; Ding, T.; Taylor, P.R.; Lee, M.P. Genome wide analysis of DNA copy number neutral loss of heterozygosity (CNNLOH) and its relation to gene expression in esophageal squamous cell carcinoma. BMC Genom. 2010, 11, 576. [Google Scholar] [CrossRef] [Green Version]
- Abnet, C.C.; Freedman, N.D.; Hu, N.; Wang, Z.; Yu, K.; Shu, X.O.; Yuan, J.M.; Zheng, W.; Dawsey, S.M.; Dong, L.M.; et al. A shared susceptibility locus in PLCE1 at 10q23 for gastric adenocarcinoma and esophageal squamous cell carcinoma. Nat. Genet. 2010, 42, 764–767. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.-D.; Zhou, F.-Y.; Li, X.-M.; Sun, L.-D.; Song, X.; Jin, Y.; Li, J.-M.; Kong, G.-Q.; Qi, H.; Cui, J.; et al. Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects identifies susceptibility loci at PLCE1 and C20orf54. Nat. Genet. 2010, 42, 759–763. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Hu, Z.B.; He, Z.H.; Jia, W.H.; Wang, F.; Zhou, Y.F.; Liu, Z.H.; Zhan, Q.M.; Liu, Y.; Yu, D.K.; et al. Genome-wide association study identifies three new susceptibility loci for esophageal squamous-cell carcinoma in Chinese populations. Nat. Genet. 2011, 43, 679–684. [Google Scholar] [CrossRef]
- Wu, C.; Wang, Z.; Song, X.; Feng, X.S.; Abnet, C.C.; He, J.; Hu, N.; Zuo, X.B.; Tan, W.; Zhan, Q.; et al. Joint analysis of three genome-wide association studies of esophageal squamous cell carcinoma in Chinese populations. Nat Genet. 2014, 46, 1001–1006. [Google Scholar] [CrossRef] [Green Version]
- Albert, F.W.; Kruglyak, L. The role of regulatory variations in complex traits and disease. Nat. Rev. Genet. 2015, 16, 197–212. [Google Scholar] [CrossRef]
- Coussens, L.M.; Werb, Z. Inflammation and cancer (Review). Nature 2002, 420, 860–867. [Google Scholar] [CrossRef]
- Grivennikov, S.I.; Greten, F.R.; Karin, M. Immunity, inflammation, and cancer (Review). Cell 2010, 140, 883–899. [Google Scholar] [CrossRef] [Green Version]
- Jeggo, P.A.; Pearl, L.H.; Carr, A.M. DNA repair, genome stability and cancer: A historical perspective. Nat. Res Cancer 2016, 16, 35–42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, H.; Su, H.; Hu, N.; Wang, C.; Wang, L.; Giffen, C.; Goldstein, A.M.; Lee, M.P.; Taylor, P.R. Integrated analysis of genome-wide miRNAs and targeted gene expression in esophageal squamous cell carcinoma (ESCC) and relation to prognosis. BMC Cancer 2020, 20, 388. [Google Scholar] [CrossRef] [PubMed]
- Gamazon, E.R.; Stranger, B.E. The impact of human copy number variation on gene expression. Brief. Funct. Genom. 2015, 14, 352–357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shao, X.; Lv, N.; Liao, J.; Long, J.; Xue, R.; Ai, N.; Xu, D.; Fan, X. Copy number variation is highly correlated with differential gene expression: A pan-cancer study. BMC Med. Genet. 2019, 20, 175. [Google Scholar] [CrossRef] [PubMed]
- Stranger, B.E.; Forrest, M.S.; Dunning, M.; Ingle, C.E.; Beazley, C.; Thorne, N.; Redon, R.; Bird, C.P.; de Grassi, A.; Lee, C.; et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 2007, 315, 848–853. [Google Scholar] [CrossRef] [Green Version]
- Garnham, R.; Scott, E.; Livermore, K.E.; Munkley, J. ST6GAL1: A key player in cancer (Review). Oncol. Lett. 2019, 18, 983–989. [Google Scholar] [CrossRef] [Green Version]
- Irizarry, R.A.; Hobbs, B.; Collin, F.; Beazer-Barclay, Y.D.; Antonellis, K.J.; Scherf, U.; Speed, T.P. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003, 4, 249–264. [Google Scholar] [CrossRef] [Green Version]

| (a) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Gene No | Gene Name | Chr | SNP | Probeset | Classical eQTL | p Value | Modified eQTL | p Value | SNP | |
| rho | rho | No | ||||||||
| 1 | CD46 | 1q32 | rs7144 | 208783_s_at | 0.310 | 0.002 | −0.273 | 0.006 | 1 | |
| rs2724391 | 208783_s_at | 0.261 | 0.010 | −0.211 | 0.037 | 2 | ||||
| 2 | CD58 | 1p13 | rs1335532 | 216942_s_at | 0.226 | 0.024 | −0.210 | 0.036 | 3 | |
| 3 | COL11A1 | 1p21 | rs2061705 | 37892_at | −0.223 | 0.028 | 0.242 | 0.017 | 4 | |
| 4 | CYP2C18 | 10q24 | rs1409654 | 215103_at | −0.321 | 0.001 | 0.245 | 0.014 | 5 | |
| rs2296679 | 215103_at | −0.312 | 0.002 | 0.240 | 0.016 | 6 | ||||
| rs1409654 | 208126_s_at | −0.281 | 0.005 | 0.221 | 0.028 | 7 | ||||
| 5 | CYP2C9 | 10q24 | rs4086116 | 214420_s_at | −0.206 | 0.040 | 0.209 | 0.037 | 8 | |
| rs4917639 | 214420_s_at | −0.206 | 0.040 | 0.209 | 0.037 | 9 | ||||
| 6 | DAPK1 | 9q21.33 | rs1964911 | 211214_s_at | −0.294 | 0.003 | 0.218 | 0.029 | 10 | |
| 7 | GLS2 | 12q13 | rs6581096 | 205531_s_at | −0.201 | 0.044 | 0.226 | 0.024 | 11 | |
| 8 | N4BP2L1 | 13q12-q13 | rs1207952 | 211390_at | 0.224 | 0.025 | −0.301 | 0.002 | 12 | |
| 9 | NCAM1 | 11q23.1 | rs2850303 | 212843_at | 0.271 | 0.006 | −0.355 | 0.000 | 13 | |
| rs584427 | 212843_at | 0.251 | 0.013 | −0.314 | 0.002 | 14 | ||||
| rs1821693 | 212843_at | 0.242 | 0.016 | −0.352 | 0.000 | 15 | ||||
| 10 | NCAPD2 | 12p13.3 | rs917634 | 201774_s_at | −0.337 | 0.001 | 0.198 | 0.048 | 16 | |
| 11 | PARD3 | 10p11.22- | rs2496720 | 221280_s_at | −0.262 | 0.009 | 0.247 | 0.014 | 17 | |
| p11.21 | rs2496720 | 210094_s_at | −0.215 | 0.033 | 0.231 | 0.022 | 18 | |||
| 12 | PDCD1LG2 | 9p24.2 | rs1360238 | 220049_s_at | −0.202 | 0.044 | 0.295 | 0.003 | 19 | |
| 13 | PTPRM | 18p11.2 | rs12606738 | 216292_at | −0.202 | 0.046 | 0.222 | 0.028 | 20 | |
| 14 | TCF7L1 | 2p11.2 | rs12714137 | 221016_s_at | −0.211 | 0.037 | 0.213 | 0.035 | 21 | |
| 15 | TCF7L2 | 10q25.3 | rs1028629 | 212761_at | −0.299 | 0.003 | 0.205 | 0.041 | 22 | |
| 16 | ZBTB16 | 11q23.1 | rs2852796 | 205883_at | −0.237 | 0.017 | 0.243 | 0.015 | 23 | |
| 17 | MS4A1 | 11q12 | rs4939363 | 210356_x_at | 0.216 | 0.033 | −0.224 | 0.027 | 24 | |
| rs4939362 | 210356_x_at | 0.199 | 0.047 | −0.202 | 0.044 | 25 | ||||
| rs1941030 | 210356_x_at | 0.199 | 0.047 | −0.202 | 0.044 | 26 | ||||
| 18 | ST6GAL1 | 3q27-q28 | rs12495026 | 214971_s_at | −0.242 | 0.016 | 0.220 | 0.028 | 27 | |
| rs12495023 | 214971_s_at | −0.236 | 0.018 | 0.213 | 0.035 | 28 | ||||
| (b) | ||||||||||
| Gene No | Gene | Cytoband | Classical eQTL SNP | Probeset | rho | pValue | Modified eQTL SNP | Probeset | rho | pValue |
| 1 | CACNA1C | 12p13.3 | rs2239097 | 208020_s_at | 0.229 | 0.0222 | rs2239097 | 211592_s_at | −0.208 | 0.037 |
| rs2283318 | 208020_s_at | 0.219 | 0.0297 | |||||||
| 2 | CACNB2 | 10p12 | rs1034139 | 215365_at | −0.202 | 0.0434 | rs4748472 | 207776_s_at | −0.245 | 0.015 |
| 3 | IGF1R | 15q26.3 | rs2684811 | 203627_at | −0.352 | 0.0003 | rs12908437 | 208441_at | 0.218 | 0.029 |
| 4 | ITPR1 | 3p26-p25 | rs11714599 | 216944_s_at | 0.204 | 0.0431 | rs3805032 | 203710_at | −0.253 | 0.012 |
| rs11714599 | 203710_at | 0.204 | 0.0433 | rs3805032 | 203710_at | −0.253 | 0.012 | |||
| rs304051 | 216944_s_at | 0.250 | 0.012 | |||||||
| rs304053 | 216944_s_at | 0.250 | 0.012 | |||||||
| rs304051 | 222314_x_at | −0.218 | 0.029 | |||||||
| 5 | rs304053 | 222314_x_at | −0.218 | 0.029 | ||||||
| NRP1 | 10p12 | rs4934583 | 210615_at | −0.213 | 0.0335 | rs869636 | 210510_s_at | −0.309 | 0.002 | |
| rs2776928 | 210510_s_at | −0.231 | 0.022 | |||||||
| rs2776928 | 212298_at | −0.215 | 0.032 | |||||||
| 6 | RARB | 3p24 | rs922939 | 208412_s_at | 0.227 | 0.0238 | rs12630664 | 208413_at | −0.293 | 0.003 |
| rs17016781 | 208413_at | −0.220 | 0.0282 | rs12631063 | 208413_at | −0.271 | 0.007 | |||
| rs3773439 | 208412_s_at | 0.271 | 0.007 | |||||||
| rs1730223 | 208412_s_at | 0.269 | 0.007 | |||||||
| rs17016773 | 208412_s_at | 0.266 | 0.008 | |||||||
| rs11707637 | 208412_s_at | 0.265 | 0.008 | |||||||
| rs7610831 | 208412_s_at | 0.253 | 0.011 | |||||||
| rs17029657 | 208412_s_at | 0.239 | 0.017 | |||||||
| rs6800566 | 217020_at | 0.225 | 0.024 | |||||||
| rs17016738 | 208412_s_at | 0.216 | 0.031 | |||||||
| No | Gene | Cytoband | Gene Expression: Frequency | Somatic DNA Segment Alterations: Frequency | |||||
|---|---|---|---|---|---|---|---|---|---|
| Over | Under | Abnormal | AI Only | CN Gain with/without AI | CN Loss or LOH with/without AI | Total with Alterations | |||
| 1 | CACNA1C | 12p13.3 | 0.06 | 0.16 | 0.22 | 0.13 | 0.21 | 0.05 | 0.39 |
| 2 | CACNB2 | 10p12 | 0.09 | 0.36 | 0.45 | 0.22 | 0.05 | 0.16 | 0.43 |
| 3 | CD46 | 1q32 | 0.14 | 0.25 | 0.39 | 0.24 | 0.20 | 0.00 | 0.43 |
| 4 | CD58 | 1p13 | 0.33 | 0.13 | 0.46 | 0.17 | 0.03 | 0.09 | 0.29 |
| 5 | COL11A1 | 1p21 | 0.95 | 0 | 0.95 | 0.16 | 0.03 | 0.12 | 0.3 |
| 6 | CYP2C18 | 10q24 | 0.07 | 0.81 | 0.88 | 0.24 | 0.00 | 0.11 | 0.34 |
| 7 | CYP2C9 | 10q24 | 0.04 | 0.62 | 0.64 | 0.24 | 0.00 | 0.11 | 0.34 |
| 8 | DAPK1 | 9q34.1 | 0.21 | 0.22 | 0.43 | 0.33 | 0.12 | 0.24 | 0.68 |
| 9 | GLS2 | 12q13 | 0.28 | 0.18 | 0.46 | 0.16 | 0.08 | 0.03 | 0.26 |
| 10 | IGF1R | 15q26.3 | 0.54 | 0.03 | 0.57 | 0.17 | 0.18 | 0.08 | 0.43 |
| 11 | ITPR1 | 3p26-p25 | 0.09 | 0.47 | 0.56 | 0.24 | 0.03 | 0.38 | 0.64 |
| 12 | MS4A1 | 11q12 | 0.32 | 0.2 | 0.52 | 0.26 | 0.03 | 0.05 | 0.34 |
| 13 | N4BP2L1 | 13.q13.1 | 0.13 | 0.17 | 0.3 | 0.29 | 0.03 | 0.28 | 0.59 |
| 14 | NCAM1 | 11q23.1 | 0.09 | 0.27 | 0.36 | 0.26 | 0.04 | 0.18 | 0.49 |
| 15 | NCAPD2 | 12p13.3 | 0.81 | 0.05 | 0.86 | 0.18 | 0.14 | 0.03 | 0.36 |
| 16 | NRP1 | 10p12 | 0.52 | 0.08 | 0.6 | 0.22 | 0.07 | 0.09 | 0.38 |
| 17 | PARD3 | 10p11 | 0.03 | 0.47 | 0.5 | 0.17 | 0.07 | 0.16 | 0.39 |
| 18 | PDCD1LG2 | 9p24.2 | 0.13 | 0.08 | 0.21 | 0.43 | 0.05 | 0.29 | 0.78 |
| 19 | PTPRM | 18p11.2 | 0.26 | 0.09 | 0.35 | 0.17 | 0.16 | 0.08 | 0.41 |
| 20 | RARB | 3p24 | 0.04 | 0.15 | 0.19 | 0.30 | 0.00 | 0.36 | 0.66 |
| 21 | ST6GAL1 | 3q27-q28 | 0.33 | 0.07 | 0.4 | 0.04 | 0.68 | 0.01 | 0.74 |
| 22 | TCF7L1 | 2p11.2 | 0.32 | 0.27 | 0.59 | 0.16 | 0.13 | 0.03 | 0.32 |
| 23 | TCF7L2 | 10q25.3 | 0.17 | 0.13 | 0.3 | 0.24 | 0.03 | 0.14 | 0.41 |
| 24 | ZBTB16 | 11q23.1 | 0.05 | 0.74 | 0.79 | 0.22 | 0.03 | 0.21 | 0.46 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, H.H.; Liu, H.; Hu, N.; Su, H.; Wang, C.; Giffen, C.; Goldstein, A.M.; Taylor, P.R.; Lee, M.P. Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation. Cancers 2022, 14, 1629. https://doi.org/10.3390/cancers14071629
Yang HH, Liu H, Hu N, Su H, Wang C, Giffen C, Goldstein AM, Taylor PR, Lee MP. Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation. Cancers. 2022; 14(7):1629. https://doi.org/10.3390/cancers14071629
Chicago/Turabian StyleYang, Howard H., Huaitian Liu, Nan Hu, Hua Su, Chaoyu Wang, Carol Giffen, Alisa M. Goldstein, Philip R. Taylor, and Maxwell P. Lee. 2022. "Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation" Cancers 14, no. 7: 1629. https://doi.org/10.3390/cancers14071629
APA StyleYang, H. H., Liu, H., Hu, N., Su, H., Wang, C., Giffen, C., Goldstein, A. M., Taylor, P. R., & Lee, M. P. (2022). Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation. Cancers, 14(7), 1629. https://doi.org/10.3390/cancers14071629






