Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Samples
2.3. DNA Isolation
2.4. Next Generation Sequencing
2.4.1. Tumor Genotyping
2.4.2. Genotyping for Germline Alterations/Variants
2.5. Multiplex Ligation-Dependent Probe Amplification (MLPA) and Sanger Sequencing
2.6. Statistical Analysis
3. Results
3.1. Frequencies of Germline and Somatic PV/LPV
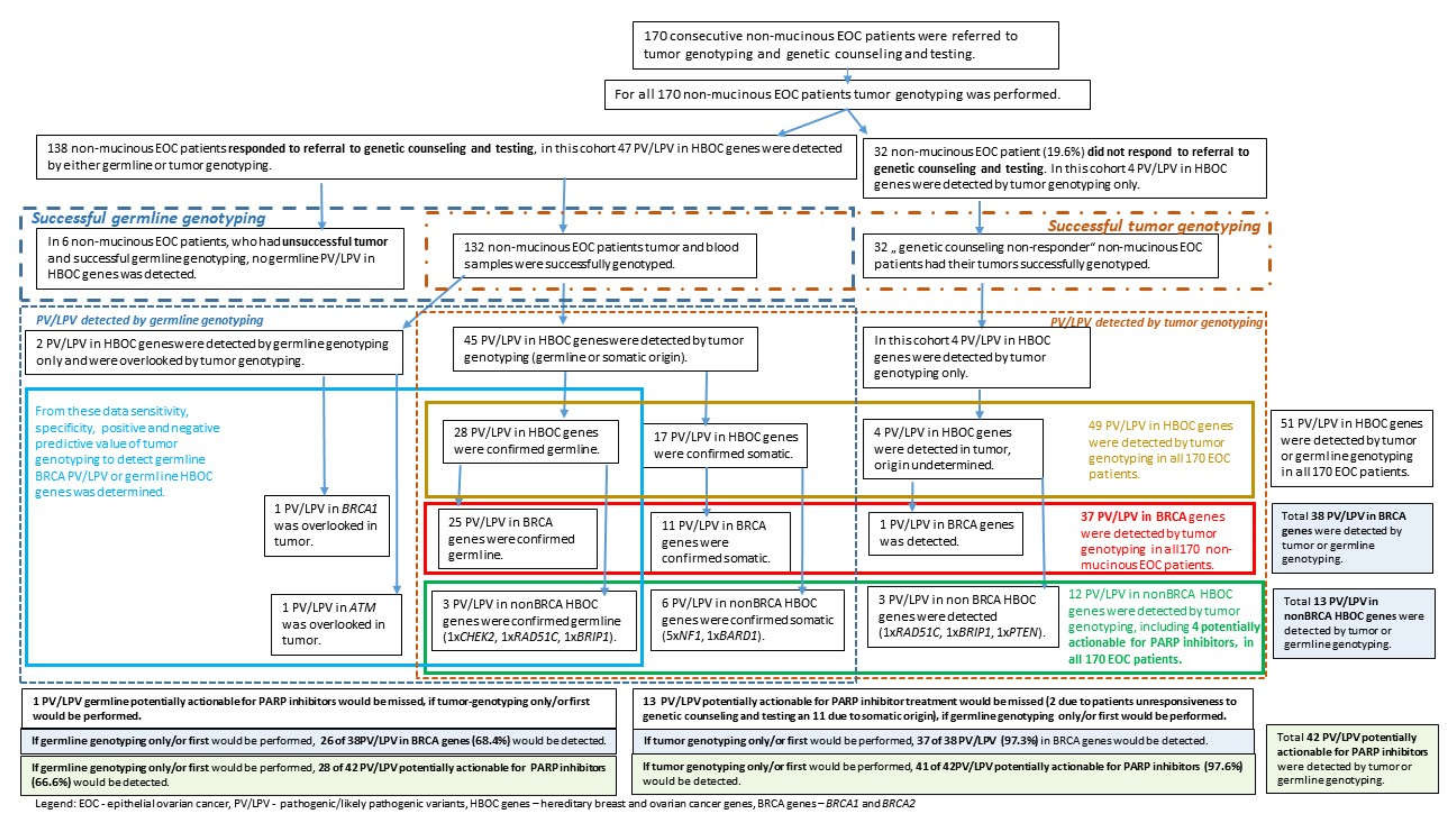
3.2. Proportion of Detected PV/LPV in All 170 Consecutive Advanced Non-Mucinous EOC Patients Depending on the Approach to Testing: Tumor Genotyping Only vs. Germline Genotyping Only
3.3. Frequencies of Germline and Somatic PV/LPV in HBOC Genes among Subgroup of 132 Advanced Non-Mucinous EOC Patients with Matched Successful Tumor and Germline Genotyping
3.4. Efficiency of Tumor Genotyping for Detection of Germline Variants
3.5. Family History of BRCA-Related Cancers among Matched Advanced Non-Mucinous EOC Patients
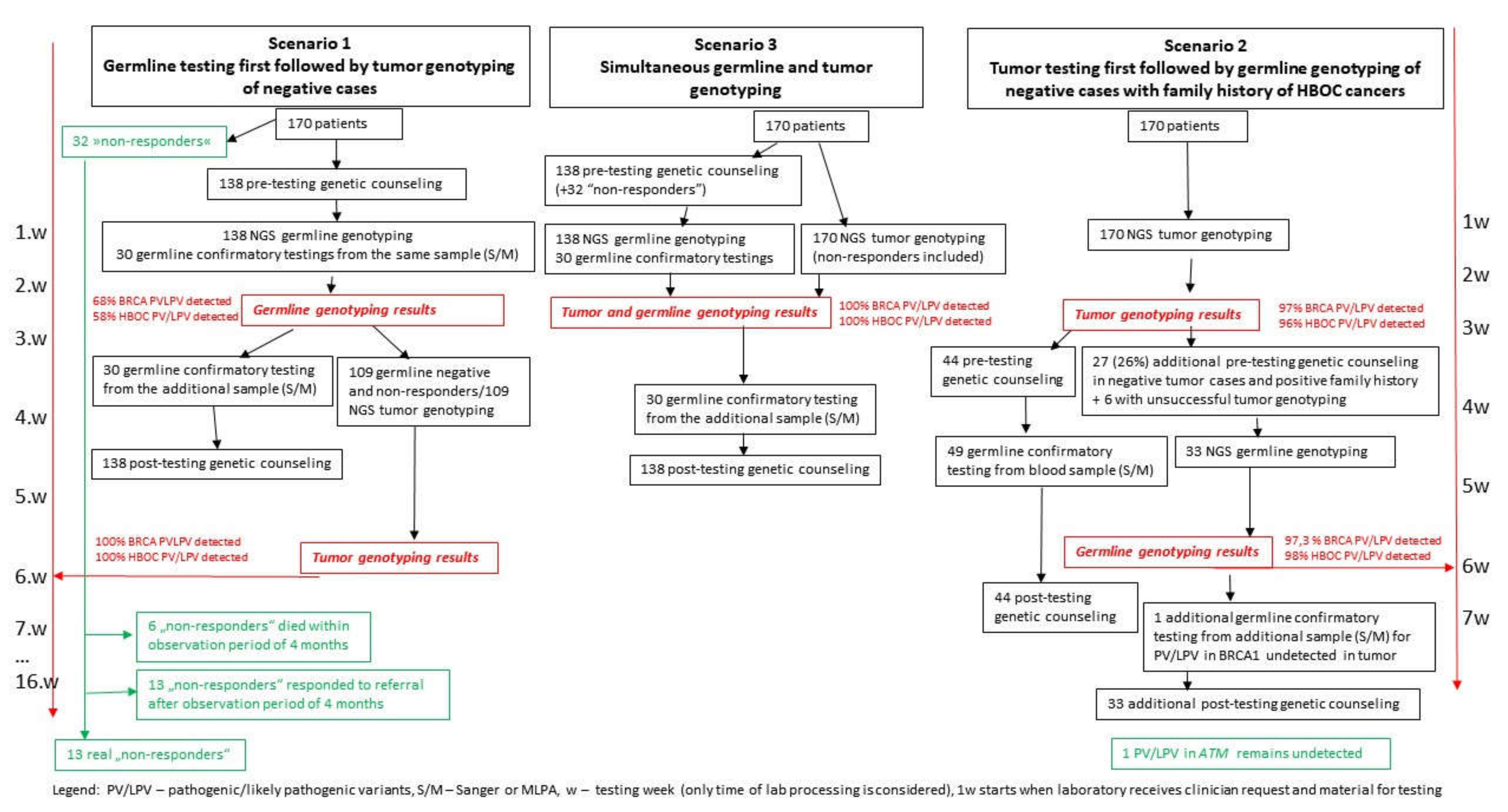
3.6. Analysis of Three Different Approaches to Germline and Tumor Testing
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zadnik, V.; Primic Zakelj, M.; Lokar, K.; Jarm, K.; Ivanus, U.; Zagar, T. Cancer burden in slovenia with the time trends analysis. Radiol. Oncol. 2017, 51, 47–55. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daly, M.B.; Pal, T.; Berry, M.P.; Buys, S.S.; Dickson, P.; Domchek, S.M.; Elkhanany, A.; Friedman, S.; Goggins, M.; Hutton, M.L.; et al. Genetic/Familial High-Risk Assessment: Breast, Ovarian, and Pancreatic, Version 2.2021, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2021, 19, 77–102. [Google Scholar] [CrossRef] [PubMed]
- Prat, J. Ovarian carcinomas: Five distinct diseases with different origins, genetic alterations, and clinicopathological features. Virchows Arch. 2012, 460, 237–249. [Google Scholar] [CrossRef] [PubMed]
- Bell, D.; Berchuck, A.; Birrer, M.; Chien, J.; Cramer, D.W.; Dao, F.; Dhir, R.; DiSaia, P.; Gabra, H.; Glenn, P.; et al. Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef]
- Palacios, J.; de la Hoya, M.; Bellosillo, B.; de Juan, I.; Matías-Guiu, X.; Lázaro, C.; Palanca, S.; Osorio, A.; Rojo, F.; Rosa-Rosa, J.M.; et al. Mutational Screening of BRCA1/2 Genes as a Predictive Factor for Therapeutic Response in Epithelial Ovarian Cancer: A Consensus Guide from the Spanish Society of Pathology (SEAP-IAP) and the Spanish Society of Human Genetics (AEGH). Virchows Arch. 2020, 476, 195–207. [Google Scholar] [CrossRef] [Green Version]
- Ledermann, J.; Harter, P.; Gourley, C.; Friedlander, M.; Vergote, I.; Rustin, G.; Scott, C.L.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: A preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 2014, 15, 852–861. [Google Scholar] [CrossRef]
- Vos, J.R.; Fakkert, I.E.; de Hullu, J.A.; van Altena, A.M.; Sie, A.S.; Ouchene, H.; Willems, R.W.; Nagtegaal, I.D.; Jongmans, M.C.J.; Mensenkamp, A.R.; et al. Universal Tumor DNA BRCA1/2 Testing of Ovarian Cancer: Prescreening PARPi Treatment and Genetic Predisposition. JNCI J. Natl. Cancer Inst. 2020, 112, 161–169. [Google Scholar] [CrossRef]
- Pennington, K.P.; Walsh, T.; Harrell, M.I.; Lee, M.K.; Pennil, C.C.; Rendi, M.H.; Thornton, A.; Norquist, B.M.; Casadei, S.; Nord, A.S.; et al. Germline and Somatic Mutations in Homologous Recombination Genes Predict Platinum Response and Survival in Ovarian, Fallopian Tube, and Peritoneal Carcinomas. Clin. Cancer Res. 2014, 20, 764–775. [Google Scholar] [CrossRef] [Green Version]
- Konstantinopoulos, P.A.; Norquist, B.; Lacchetti, C.; Armstrong, D.; Grisham, R.N.; Goodfellow, P.J.; Kohn, E.C.; Levine, D.A.; Liu, J.F.; Lu, K.H.; et al. Germline and Somatic Tumor Testing in Epithelial Ovarian Cancer: ASCO Guideline. J. Clin. Oncol. 2020, 38, 1222–1245. [Google Scholar] [CrossRef]
- Colombo, N.; Sessa, C.; du Bois, A.; Ledermann, J.; McCluggage, W.G.; McNeish, I.; Morice, P.; Pignata, S.; Ray-Coquard, I.; Vergote, I.; et al. ESMO–ESGO consensus conference recommendations on ovarian cancer: Pathology and molecular biology, early and advanced stages, borderline tumours and recurrent disease. Ann. Oncol. 2019, 30, 672–705. [Google Scholar] [CrossRef] [Green Version]
- Swisher, E.M.; Lin, K.K.; Oza, A.M.; Scott, C.L.; Giordano, H.; Sun, J.; Konecny, G.E.; Coleman, R.L.; Tinker, A.V.; O’Malley, D.M.; et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): An international, multicentre, open-label, phase 2 trial. Lancet Oncol. 2017, 18, 75–87. [Google Scholar] [CrossRef] [Green Version]
- Ledermann, J.A.; Harter, P.; Gourley, C.; Friedlander, M.; Vergote, I.; Rustin, G.; Scott, C.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Overall survival in patients with platinum-sensitive recurrent serous ovarian cancer receiving olaparib maintenance monotherapy: An updated analysis from a randomised, placebo-controlled, double-blind, phase 2 trial. Lancet Oncol. 2016, 17, 1579–1589. [Google Scholar] [CrossRef]
- Capoluongo, E.; Ellison, G.; López-Guerrero, J.A.; Penault-Llorca, F.; Ligtenberg, M.J.L.; Banerjee, S.; Singer, C.; Friedman, E.; Markiefka, B.; Schirmacher, P.; et al. Guidance Statement On BRCA1/2 Tumor Testing in Ovarian Cancer Patients. Semin. Oncol. 2017, 44, 187–197. [Google Scholar] [CrossRef]
- Gornjec, A.; Novakovic, S.; Stegel, V.; Hocevar, M.; Pohar Marinsek, Z.; Gazic, B.; Krajc, M.; Skof, E. Cytology material is equivalent to tumor tissue in determining mutations of BRCA 1/2 genes in patients with tubo-ovarian high grade serous carcinoma. BMC Cancer 2019, 19, 296. [Google Scholar] [CrossRef] [Green Version]
- Klančar, G.; Blatnik, A.; Šetrajčič Dragoš, V.; Vogrič, V.; Stegel, V.; Blatnik, O.; Drev, P.; Gazič, B.; Krajc, M.; Novaković, S. A Novel Germline MLH1 In-Frame Deletion in a Slovenian Lynch Syndrome Family Associated with Uncommon Isolated PMS2 Loss in Tumor Tissue. Genes 2020, 11, 325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.M.; Datto, M.; Duncavage, E.J.; Kulkarni, S.; Lindeman, N.I.; Roy, S.; Tsimberidou, A.M.; Vnencak-Jones, C.L.; Wolff, D.J.; Younes, A.; et al. Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 2017, 19, 4–23. [Google Scholar] [CrossRef] [Green Version]
- Den Dunnen, J.T.; Dalgleish, R.; Maglott, D.R.; Hart, R.K.; Greenblatt, M.S.; McGowan-Jordan, J.; Roux, A.-F.; Smith, T.; Antonarakis, S.E.; Taschner, P.E.M. HGVS Recommendations for the Description of Sequence Variants: 2016 Update. Hum. Mutat. 2016, 37, 564–569. [Google Scholar] [CrossRef] [Green Version]
- Plon, S.E.; Eccles, D.M.; Easton, D.; Foulkes, W.D.; Genuardi, M.; Greenblatt, M.S.; Hogervorst, F.B.L.; Hoogerbrugge, N.; Spurdle, A.B.; Tavtigian, S.V.; et al. Sequence variant classification and reporting: Recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum. Mutat. 2008, 29, 1282–1291. [Google Scholar] [CrossRef] [Green Version]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–423. [Google Scholar] [CrossRef] [Green Version]
- National Comprehensive Cancer Network. Ovarian Cancer Including Fallopian Tube Cancer and Primary Peritoneal Cancer (Version 1.2021). Available online: https://www.nccn.org/professionals/physician_gls/pdf/ovarian.pdf (accessed on 27 April 2021).
- Mirza, M.R.; Coleman, R.L.; González-Martín, A.; Moore, K.N.; Colombo, N.; Ray-Coquard, I.; Pignata, S. The forefront of ovarian cancer therapy: Update on PARP inhibitors. Ann. Oncol. 2020, 31, 1148–1159. [Google Scholar] [CrossRef]
- Miller, R.E.; Leary, A.; Scott, C.L.; Serra, V.; Lord, C.J.; Bowtell, D.; Chang, D.K.; Garsed, D.W.; Jonkers, J.; Ledermann, J.A.; et al. ESMO recommendations on predictive biomarker testing for homologous recombination deficiency and PARP inhibitor benefit in ovarian cancer. Ann. Oncol. 2020, 31, 1606–1622. [Google Scholar] [CrossRef] [PubMed]
- Singer, C.F.; Tan, Y.Y.; Muhr, D.; Rappaport, C.; Gschwantler-Kaulich, D.; Grimm, C.; Polterauer, S.; Pfeiler, G.; Berger, A.; Tea, M.M. Association between family history, mutation locations, and prevalence of BRCA1 or 2 mutations in ovarian cancer patients. Cancer Med. 2019, 8, 1875–1881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rolnick, S.J.; Rahm, A.K.; Jackson, J.M.; Nekhlyudov, L.; Goddard, K.A.B.; Field, T.; McCarty, C.; Nakasato, C.; Roblin, D.; Anderson, C.P.; et al. Barriers in Identification and Referral to Genetic Counseling for Familial Cancer Risk: The Perspective of Genetic Service Providers. J. Genet. Couns. 2011, 20, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, C.; Tomao, F.; Betella, I.; Rappa, A.; Calvello, M.; Bonanni, B.; Bernard, L.; Peccatori, F.; Colombo, N.; Viale, G.; et al. Tumor BRCA Test for Patients with Epithelial Ovarian Cancer: The Role of Molecular Pathology in the Era of PARP Inhibitor Therapy. Cancers 2019, 11, 1641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hennessy, B.T.J.; Timms, K.M.; Carey, M.S.; Gutin, A.; Meyer, L.A.; Flake, D.D.; Abkevich, V.; Potter, J.; Pruss, D.; Glenn, P.; et al. Somatic Mutations in BRCA1 and BRCA2 Could Expand the Number of Patients That Benefit From Poly (ADP Ribose) Polymerase Inhibitors in Ovarian Cancer. J. Clin. Oncol. 2010, 28, 3570–3576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Jonge, M.M.; Ruano, D.; van Eijk, R.; van der Stoep, N.; Nielsen, M.; Wijnen, J.T.; ter Haar, N.T.; Baalbergen, A.; Bos, M.E.M.M.; Kagie, M.J.; et al. Validation and Implementation of BRCA1/2 Variant Screening in Ovarian Tumor Tissue. J. Mol. Diagn. 2018, 20, 600–611. [Google Scholar] [CrossRef]
- Bekos, C.; Grimm, C.; Kranawetter, M.; Polterauer, S.; Oberndorfer, F.; Tan, Y.; Müllauer, L.; Singer, C.F. Reliability of Tumor Testing Compared to Germline Testing for Detecting BRCA1 and BRCA2 Mutations in Patients with Epithelial Ovarian Cancer. J. Pers. Med. 2021, 11, 593. [Google Scholar] [CrossRef]
- Watkins, J.A.; Irshad, S.; Grigoriadis, A.; Tutt, A.N. Genomic scars as biomarkers of homologous recombination deficiency and drug response in breast and ovarian cancers. Breast Cancer Res. 2014, 16, 211. [Google Scholar] [CrossRef] [Green Version]
- Sundar, S.; Manchanda, R.; Gourley, C.; George, A.; Wallace, A.; Balega, J.; Williams, S.; Wallis, Y.; Edmondson, R.; Nicum, S.; et al. British Gynaecological Cancer Society/British Association of Gynaecological Pathology consensus for germline and tumor testing for BRCA1/2 variants in ovarian cancer in the United Kingdom. Int. J. Gynecol. Cancer 2021, 31, 272–278. [Google Scholar] [CrossRef]
| Group/Subgroup No. of Patients | PV/LPV (No.) | Patients with PV/LPV (No., %) | Germline PV/LPV (No.) | Patients with Germline PV/LPV (No., %) | Somatic PV/LPV (No.) | Patients with Somatic PV/LPV (No., %) | PV/LPV with Unknown Origin * (No.) | Patients with PV/LPV with Unknown Origin * (No., %) |
|---|---|---|---|---|---|---|---|---|
| Entire study group 170 | 51 | 46 (27.1%) | 30 *** | 29 (17.1%) | 17 | 15 (8.8%) | 4 | 4 (2.3%) |
| 1. patients with unsuccessful tumor genotyping ** 6 | 0 | 0 | 0 | 0 | NA | NA | NA | NA |
| 2. patient with successful tumor and germline genotyping 132 | 47 | 42 (31.8%) | 30 *** | 29 (21.9%) | 17 | 15 (11.3%) | 0 | 0 |
| 3. non-responders 32 | 4 | 4 | NA | NA | NA | NA | 4 | 4 (12.5%) |
| Group/Subgroup No. of Patients | PV/LPV (No.) | Patients with PV/LPV (No., %) | Germline PV/LPV (No.) | Patients with Germline PV/LPV (No., %) | Somatic PV/LPV (No.) | Patients with Somatic PV/LPV (No.,%) | PV/LPV with Unknown Origin * (No.) | Patients with PV/LPV with Unknown Origin * (No., %) |
|---|---|---|---|---|---|---|---|---|
| Entire study group 170 | 38 | 37 (21.8%) | 26 | 26 (15.3%) *** | 11 | 11 (6.5%) *** | 1 | 1 (0.6%) |
| 1. patients with unsuccessful tumor genotyping ** 6 | 0 | 0 | 0 | 0 | NA | NA | NA | NA |
| 2. patient with successful tumor and germline genotyping 132 | 37 | 36 (27.2) | 26 | 26 (19.6%) *** | 11 | 11 (8.3%) *** | 0 | 0 |
| 3. non-responders 32 | 1 | 1 (3.1%) | NA | NA | NA | NA | 1 | 1 (3.1%) |
| Group/Subgroup No. of Patients | PV/LPV (No.) | Patients with PV/LPV (No., %) | Germline PV/LPV (No.) | Patients with Germline PV/LPV (No., %) | Somatic PV/LPV (No.) | Patients with Somatic PV/LPV (No., %) | PV/LPV with Unknown Origin * (No.) | Patients with PV/LPV with Unknown Origin * (No., %) |
|---|---|---|---|---|---|---|---|---|
| Entire study group. 170 | 13 | 11 (6.4%) | 4 *** | 3 (1.7%) | 6 **** | 5 (2.9%) | 3 ***** | 3 (1.7%) |
| 1. patients with unsuccessful tumor genotyping ** 6 | 0 | 0 | 0 | 0 | NA | NA | NA | NA |
| 2. patient with successful tumor and germline genotyping 132 | 10 | 8 (6.0%) | 4 *** | 3 (2.2%) | 6 **** | 5 (3.8%) | 0 | 0 |
| 3. non-responders 32 | 3 | 3 (9.3%) | NA | NA | NA | NA | 3 ***** | 3 (9.3%) |
| Matched Tumor Samples | Matched Blood Samples | ||
|---|---|---|---|
| No. of patients–carriers of germline PV/LPV in BRCA genes | No. of patients with germline wild type BRCA genes | No. of all patients with genotyped matched blood sample | |
| No. patients with PV/LPV in BRCA genes detected in matched tumor sample | 25 | 10 | 35 |
| No. patients with wild type BRCA genes in matched tumor sample | 1 * | 96 | 97 |
| No. of all patients with genotyped matched tumor samples | 26 | 106 | 132 |
| Matched Tumor Samples | Matched Blood Samples | ||
|---|---|---|---|
| No. of patients–carriers of germline PV/LPV in HBOC genes | No. of patients with germline wild type HBOC genes | No. of all patients with genotyped matched blood samples | |
| No. patients with PV/LPV in HBOC genes detected in matched tumor samples | 27 | 13 ** | 40 |
| No. patients with wild type HBOC genes in matched tumor sample | 2 * | 90 | 92 |
| No. of all patients with genotyped matched tumor samples | 29 | 103 | 132 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stegel, V.; Blatnik, A.; Škof, E.; Dragoš, V.Š.; Krajc, M.; Gregorič, B.; Škerl, P.; Strojnik, K.; Klančar, G.; Banjac, M.; et al. Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing. Cancers 2022, 14, 1434. https://doi.org/10.3390/cancers14061434
Stegel V, Blatnik A, Škof E, Dragoš VŠ, Krajc M, Gregorič B, Škerl P, Strojnik K, Klančar G, Banjac M, et al. Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing. Cancers. 2022; 14(6):1434. https://doi.org/10.3390/cancers14061434
Chicago/Turabian StyleStegel, Vida, Ana Blatnik, Erik Škof, Vita Šetrajčič Dragoš, Mateja Krajc, Brigita Gregorič, Petra Škerl, Ksenija Strojnik, Gašper Klančar, Marta Banjac, and et al. 2022. "Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing" Cancers 14, no. 6: 1434. https://doi.org/10.3390/cancers14061434
APA StyleStegel, V., Blatnik, A., Škof, E., Dragoš, V. Š., Krajc, M., Gregorič, B., Škerl, P., Strojnik, K., Klančar, G., Banjac, M., Žgajnar, J., Ravnik, M., & Novaković, S. (2022). Real-World Data on Detection of Germline and Somatic Pathogenic/Likely Pathogenic Variants in BRCA1/2 and Other Susceptibility Genes in Ovarian Cancer Patients Using Next Generation Sequencing. Cancers, 14(6), 1434. https://doi.org/10.3390/cancers14061434






