Simple Summary
The development of new strategies for the management of cutaneous metastases is a major clinical challenge because of the poor prognosis. To advance in this field, a better understanding of the molecular alterations involved in the metastatic process is needed. In the present study, the clinicopathological characteristics of breast cancer that develop cutaneous metastases were analyzed and the molecular differences between primary breast tumors and their corresponding cutaneous metastases were compared. We observed that the surrogate molecular type of breast cancer with an increased risk to metastasize to the skin was triple negative. In total, 48.5% of the cutaneous metastases presented some additional molecular alteration with respect to the primary tumor. However, no characteristic mutational pattern related to skin metastasis development was observed. Identifying the genes involved in the development of cutaneous metastases is important to gain insights into the biology of the disease and to identify possible diagnostic and therapeutic biomarkers.
Abstract
Background: The characterization of molecular alterations of primary breast carcinomas (BC) and their cutaneous metastases (CM) to identify genes involved in the metastatic process have not yet been completely accomplished. Methods: To investigate the molecular alterations of BC and their CM, a total of 66 samples (33 BC and 33 CM) from 33 patients were analyzed by immunohistochemical and massive parallel sequencing analyses. In addition, the clinicopathological characteristics of patients and tumors were analyzed. Results: Triple negative (TN) BCs were overrepresented (36.4%) among tumors that developed CM. A change of tumor surrogate molecular type in metastases was found in 15% of patients and 48.5% of the CM presented some additional molecular alteration with respect to the primary tumor, the most frequent were amplification of MYC and MDM4, and mutations in TP53 and PIK3CA. Survival was related to histological grade, tumor surrogate molecular type and TP53 mutations in the univariate analysis but only the tumor surrogate molecular type remained as a prognostic factor in the multivariate analysis. Conclusions: The TN molecular type has a greater risk of developing skin metastases. There are phenotypic changes and additional molecular alterations in skin metastases compared to the corresponding primary breast tumors in nearly half of the patients. Although these changes do not follow a specific pattern and varied from patient to patient, they could impact on the treatment. More studies with larger patient and sample cohorts are needed.
1. Introduction
Breast cancer (BC) is the most prevalent malignancy in females and is the leading cause of cancer death in women [1]. We can distinguish between different groups of BC according to the molecular profile as those that express Estrogen Receptor (ER) and/or Progesterone Receptor (PR) (75%), those that express Epidermal Growth Factor Receptor 2 (Her2) (15%) [2], and tumors that do not express any of these three markers, the triple-negative (TN) tumors (10–20%) [3]. According to these characteristics, we can apply a surrogate molecular classification that distinguishes four surrogate molecular types of BC: luminal A-like, luminal B-like (HER2− and HER2+), HER2+ (non-luminal), and triple negative (TN) [4].
Metastasis accounts for the majority of deaths from BC [5]. It is a complex process in which the cells of a primary tumor are propagated to distal organs, showing uncontrolled growth in these tissues [6,7]. Normally, BC metastasize to the lung, bone, and brain [8]. Moreover, BC is the tumor most prone to develop skin metastases in females [9].
Skin metastases are the result of lymphatic embolization, hematogenous or contiguous spread [10] and are present in around 24% of patients with metastatic BC [9,11,12,13,14,15,16]. In addition, due to the high incidence of BC, these skin manifestations are the most common metastases among women seen by dermatologists, specifically, 69% of these metastases come from BC [13].
Cutaneous breast metastases tend to develop in the vicinity of the primary tumor in the skin of the breast and chest wall, although they can also develop in the abdomen, extremities, head, or neck [9,12,13]. In addition to being able to develop in different locations, breast cutaneous metastases (CM) can manifest in a wide variety of ways. Nodules are the most common presentation (80%), but there are other patterns [11,13] such as telangiectatic pattern with an incidence of 8–11%, erectile pattern with an incidence of 3–6.3%, carcinoma en cuirasse with an incidence of 3–4% and neoplastic alopecia with an incidence of 2–12% [11,13].
All BC metastases, regardless of subtype, usually occur late in the disease, in the later stages of the disease course. Brownstein et al. [17] observed that skin metastasis was the presenting sign of the disease in only 3% of cases of metastatic BC. Kong et al. [15] observed that 56.8% of the patients had more than one visceral metastasis at the time of diagnosis of CM. Therefore, when diagnosed, the primary tumor is widespread and may not be treatable (palliative care, surgical excision, or complete mastectomy is provided) [18]. All this confers a poor prognosis, with an average survival of 3 to 6 months, with few differences regarding whether the lesions are single or multiple, with mortality exceeding 70% in the first year after diagnosis [19].
Since there are few series evaluating the molecular alterations of CM and most do not include the analysis of both the primary tumor and CM, the objective of this study was to compare the molecular alterations of matched primary BC and their CM in a series of 33 patients to better understand the genes implicated in BC progression and to identify potential therapeutic targets.
2. Materials and Methods
Compliance with Ethical Standards: All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and national research committee and the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The use of patient samples to meet the objectives of this study was approved on 14 May 2021 by the Ethics Committee of the Hospital Ramón y Cajal (ethical approval code: 30-21).
2.1. Histology
Histological sections of all primary tumors and their metastases were reviewed by two experienced pathologists (J.P and B.P.-M.). Histological typing and grading was performed according to WHO recommendations [4]. Lymphovascular invasion (LIV) was also evaluated in primary tumors. Cutaneous lesions were diagnosed as metastases from BC based on a biopsy, excluding cases with direct extension from a subjacent breast lesion.
2.2. Immunohistochemistry
All 33 primary tumors and 33 metastases underwent an immunohistochemical study for the expression of estrogen receptors (ER), progesterone receptors (PR), HER2 and Ki67. Immunostaining was performed using the EnVision detection system (K5007, Agilent Dako, Glostrup, Denmark) using the following antibodies: ER (clone EP1, Agilent Dako Omnis, Glostrup, Denmark), PR (clone PR 1294, Agilent Dako Omnis, Glostrup, Denmark), HER2 (SK001, clone poly, Agilent Dako autostainer, Glostrup, Denmark), and Ki-67 (clone MIB-1, Agilent Dako Omnis, Glostrup, Denmark). Evaluation of ER, PR, and HER2 expression was performed according to American Society of Clinical Oncology and the College of American Pathologists (ASCO-CAP) guidelines) [20]. HER2 equivocal cases (2+) underwent FISH analysis, using the PathVysion HER-2 DNA Probe Kit (PathVysion II kit, Abbot Laboratories, Abbot Park, IL) on complete tumor sections. Results were interpreted according to 2018 ASCO-CAP guidelines [20].
In the 7 invasive lobular carcinomas, the expression of E-cadherin (clone NHC-38, Agilent Dako Omnis, Glostrup, Denmark) was studied to confirm the histological type. Androgen receptor (AR) (SP107 Cell Marque, Ventana Medical Systems, Rocklin, CA, USA) was also determined to evaluate the possible apocrine phenotype in 12 primary TN tumors and their metastases.
Tumors were classified into different surrogate molecular types as Luminal HER2-, Luminal HER2+, HER2 (non-luminal), and TN.
2.3. Massive Parallel Sequencing
Sufficient DNA for sequencing was obtained from the 33 primary breast carcinomas and their corresponding CMs. Areas with >30% of tumor cells were obtained by “punching” paraffin blocks in selected areas previously marked on hematoxylin/eosin (H&E) slides. The QIAamp DNA FFPE Tissue Kit (Qiagen, Valencia, CA, USA) was used to extract DNA from all samples. Quality of DNA samples was measured using TapeStation (Agilent 2200 TapeStation, Santa Clara, CA, USA), whereas quantification was performed by QUBIT 2.0. (Thermo Fisher Scientific Qubit 2.0 Fluorometer, Waltham, MA, USA).
A custom gene panel was designed using the SureDesign platform by Agilent Tech. (Santa Clara, CA, USA) to consistently target 61 genes (AKT1, ARID1A, ARID1B, ARID5B, ATR, BCOR, BRCA1, BRCA2, BRAF, CASP8, CCNE1, CDH1, CDH4, CDH19, COL1A1, CSMD3, CTCF, CTNNB1, EGFR, ERBB2, ESR1, FBXW7, FGFR1, FGFR2, FOXA2, GRB7, GSDMB, MAP2K4, KRAS, MAP3K1, MLL3, MLH1, MKI67, MSH2, MSH6, MYC, NCOR1, NF1, NRAS, PGAP3, PIK3CA3, PIK3R1, PMS2, PNMT, POLE, PPP2R1A, PRPF18, PTEN, KMT2B, RB1, RPL22, SF3B1, SPOP, STARD3, TAF1, TBX3, TCAP, TP53, VGLL1, ZNF217, ZNF703). For library construction, a modified protocol for Agilent SureSelectXT FFPE was selected based on Covaris AFA fragmentation of DNA (Covaris, Woburn, MA, USA) and subsequent probe-mediated hybridization capture. Sequencing of equimolar libraries was performed using the Ion S5™ Torrent (Thermo Fisher Scientific, MA, USA).
Bioinformatics analysis was carried out using a specific pipeline using Novoalign V3 (2021) (http://www.novocraft.com/products/novoalign/ accessed on 4 February 2022) as aligner and VarScan [21] as variant-caller, with no filters. Variant annotation was performed using the VEP from Ensembl version 88 (http://www.ensembl.org/info/docs/tools/vep/index.html accessed on 4 February 2022), which corresponds to the hg38 version of the human reference genome. Variants were latterly filtered using the functional information (taking only deleterious variants), the variant allele frequency (>0.05), and the strand-bias from both the variant and the reference allele. If normal tissue was available, those variants also present in the normal component were excluded. Finally, visual inspection was performed as the final selection criterion using the IGV browser [22].
In addition, 11 mutations were confirmed by Sanger sequencing, 3 in the PIK3CA gene (Pt3, Pt14 and pt36), 6 in the TP53 gene (Pt1, Pt2, Pt4, Pt10 and Pt11), and 2 in the ERBB2 gene in the samples (pt13). (Supplementary Table S1).
2.4. Fluorescent In-Situ Hybridization on Tissue Microarrays
Since our panel was not designed to detect CNVs, a tissue microarray (TMA) was constructed to evaluate gene copy number variations in CCND1, MYC, FGFR1 and MDM4, the genes most frequently amplified in BC, by Fluorescent In-Situ Hybridization (FISH). Only 20 matched primary tumors and metastases (40 samples) were included in the TMA due to sample limitation after the initial immunohistochemical/molecular study. Chromosomal alterations were evaluated by FISH on TMA sections using the following probes: SPEC CCND1/CEN11, MYC/CEN8, FGFR1/CEN8, and MDM4/1p12 dual color Probe Kit (Zytovision GmbH, Bremen, DE). FISH slides were observed with a fluorescence microscope at 100X with immersion oil. A detailed scoring of at least 20 neoplastic cells per sample was performed. Amplification was considered when the tumor cell population had at least twice as many gene signals than centromere signals of the respective chromosome (ratio ≥ 2), and polysomy when the average of centromere signals on tumor cells were >3.
2.5. Statistical Methods
The Kaplan–Meier method was used to calculate overall survival according to clinicopathological characteristics (age, pT, pN, histological type, surrogated molecular type, LIV, histological grade, clinical stage, metastasis location, and neoadjuvant therapy) and mutations in TP53 and PIK3CA genes. Cox proportional hazards models were used to investigate the association between mortality and clinicopathological and molecular features.
3. Results
3.1. Case Selection
A total of 33 patients diagnosed with BC and CM between 2005 and 2020 from the Pathology Department in Ramón y Cajal University Hospital (Madrid, Spain) and in 12 de Octubre University Hospital (Madrid, Spain) were selected, all had available paired samples (primary tumor and CM).
3.2. Clinicopathological Features
Clinicopathological features of all 33 primary samples are presented in supplementary Table S2 and summarized in Table 1. The median age of the patients at diagnosis was 63.5 years old (range 29 to 84), 57.6% of patients were older than 60 years.

Table 1.
Clinicopathological features of the 33 patients.
According to the immunohistochemical profile, 16 cases (48.5%) were Luminal HER2- surrogate molecular type, 3 cases (9%) were Luminal HER2+, 2 cases (6%) were HER2+ (non-Luminal), and 12 cases (36.4%) were TN.
Examining the distribution of histological types, 24 cases (72.7%) corresponded to invasive breast carcinomas of non-special type (BCNST), the rest were special histological types. The largest group of 7 cases (21.2%) corresponded to invasive lobular carcinoma (ILC), one case (3%) was a matrix-producing (chondroid) metaplastic carcinoma, and one case was an apocrine carcinoma (3%).
3.3. Immunohistochemistry and HER2 FISH
3.3.1. Surrogate Molecular Types
The surrogate molecular type of the tumors and metastases was confirmed by IHC and FISH. The most frequent type of primary tumor was Luminal HER2- followed by TN. The molecular type changed between the primary tumor and its CM in 5 patients (15%), the most common being from luminal to TN (Table 2).

Table 2.
Surrogate molecular type change between primary tumors and their cutaneous metastasis.
In case Pt14, HER2 amplification was lost in the metastasis; on the other hand, in case Pt32, HER2 was overexpressed in the metastasis due to polysomy of chromosome 17.
3.3.2. Androgen Receptor in Triple-Negative Cases
AR expression was studied in 12 TN primary tumors and their metastases by IHC, since AR expression in TNBC is related with the apocrine molecular type [23]. In this series, 3 out 12 TNBC expressed AR and expression was concordant in primary tumors and the associated metastases.
3.4. Molecular Characterization
3.4.1. Most Frequently Altered Genes in Primary Tumors and Cutaneous Metastases
The following molecular analysis is based on 66 samples (33 primary tumor and 33 CM) for mutation analysis and 40 samples (20 primary tumor and 20 CM) for CNV analysis. Figure 1 shows the mutations and CNVs found in 29 pairs. In 4 cases, no mutations or CNVs were found, and all were Luminal HER2- cases. Molecular alterations were detected in 12 TN, 12 Luminal HER2-, 3 Luminal HER2+ and 2 HER2+ (non-Luminal) primary tumors. The number of mutations ranged between 1–5. A summary table with the type of mutation and CNVs found in each of the samples is presented in supplementary Table S3.
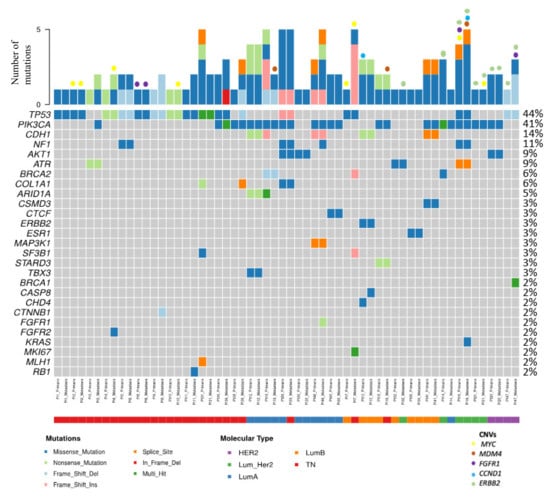
Figure 1.
Distribution of mutations and CNVs in this series. The figure shows paired samples corresponding to 29 paired patients, but the mutation frequencies were calculated considering the 33 paired cases with 66 samples.
Among the 33 matched cases, TP53 was mutated in 13 primary tumors (39.4%) and in 14 CMs (42.4%). PIK3CA was mutated in 13 primary tumors (39.4%) and 15 CMs (45.4%). NF1 was mutated in 3 of the primary tumors (9%) and 4 CMs (12.1%). AKT1 was mutated in 3 primary tumors (9%) and 3 CMs (9%). ERBB2 was amplified or there was polysomy in 5 primary tumors (15.15%) and 5 CM (15.15%). Among 20 matched cases, MYC was amplified in 3 primary tumors (15%) and 5 CM (25%). MDM4 was amplified in only 3 CM (15%). FGFR1 CNVs (amplification or polysomy) were observed in 2 primary tumors (10%) and 2 CMs (10%). Finally, CCND1 was amplified in 1 primary tumor (5%) and 1 CM (5%). Table 3 shows a summary of these alterations distributed by surrogate molecular types.

Table 3.
Pathogenic alterations in primary tumors and their cutaneous metastasis.
Analyzing TP53 and PIK3CA mutation frequencies in the CMs of BC cases diagnosed at early stages (I-II) versus those diagnosed at advanced stages (III-IV), we observed that they were very similar. The mutation frequency for both genes of cases diagnosed at early stages was 44.4%. In cases diagnosed at advanced stages the frequencies were 47.8% for TP53 and 52.2% for PIK3CA. Thus, there were no statistically significant differences between the two groups.
Enriched Molecular Alterations in Cutaneous Metastases
Additional molecular alterations were observed in the CM, either mutations or CNVs, in 16 patients (48.5%) (Figure 2). In 7 cases (21.2%) there was more than one additional alteration in the CM. Table 4 shows the distribution of these alterations by surrogate molecular types.
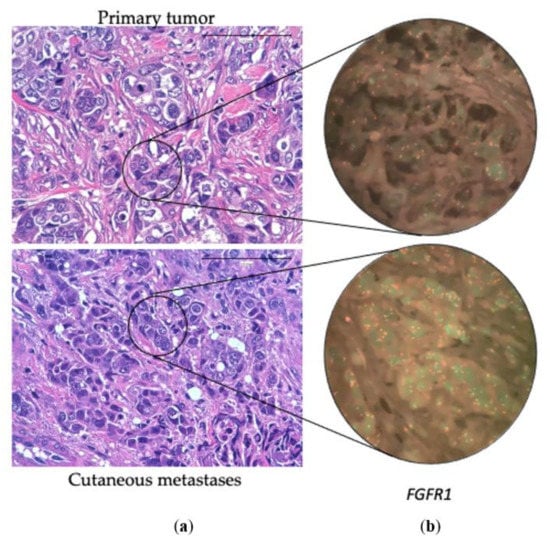
Figure 2.
(a) Hematoxylin-eosin of a primary breast tumor and its corresponding cutaneous metastasis. (b) Fluorescent In-Situ Hybridization of the FGFR1 gene in the primary breast tumor (without CNV) and in the cutaneous metastasis (with polysomy). 100×.

Table 4.
Additional molecular alteration in cutaneous metastases not found in primary tumors of paired cases.
There were some differences in the frequency of additional mutations depending on the local or distant nature of the CM. In the 17 paired cases that developed distant CM, additional alterations in the CM with respect to the primary tumors were found in 11 cases (64.7%) (5 TN, 4 Luminal HER2-, and 2 HER2+). In contrast, among the 15 paired cases that developed local CM, only 5 cases (33.3%) (3 TN, 1 Luminal HER-, and 1 HER2+) showed additional molecular alterations. (Table 4).
In addition, 11 mutations were confirmed in the TP53, PIK3CA, and ERBB2 genes. Figure 3 shows the different mutations found in the ERBB2 gene (L755S and S310F) between the primary tumor and the MC of Pt13, as well as the verification by Sanger.
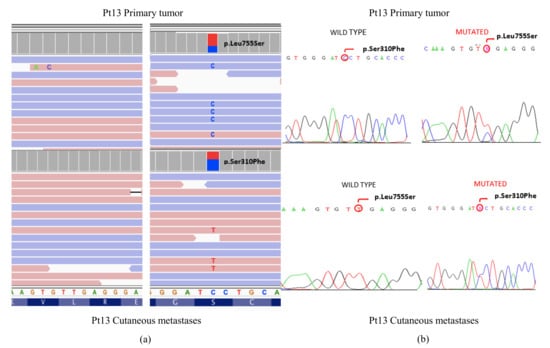
Figure 3.
(a) Visualization in the IGV software of different ERBB2 mutations found in the primary tumor and the cutaneous metastasis in patient Pt13. (b) Orthogonal validation by the Sanger sequencing.
3.5. Survival Analysis
In our series, 7 patients were alive and 26 had died when the data were censored. The median survival since the diagnosis of the disease was 53 months and the median survival since the diagnosis of the CM was 19.6 months. Of the patients who developed distant CM, the median survival was 14.5 months, with an overall survival in the first year of 46.7% of patients. In contrast, when MC was local, the median survival was 34.6 months, with an overall survival in the first year of 40% of patients.
The association between overall survival since the diagnosis of the primary tumor and age, location (local or distant), pT, pN, clinical stage, surrogate molecular type and histological type, histological grade, LIV, and neoadjuvant therapy were assessed. In addition, the association between overall survival and TP53 and PIK3CA mutations was assessed. Histological grade, surrogate molecular type, and TP53 mutations significantly affect overall survival (p = 0.015, p = 0.0011 and p = 0.019, respectively) (Figure 4a–c). The shortest overall survival was observed in the TN surrogate molecular type (Figure 4a). By multivariate analysis with the 3 significant variables, the only independent variable in the CoxPh analysis was surrogate molecular type, where HER2- Luminal was the best prognostic type. (Figure 4d).
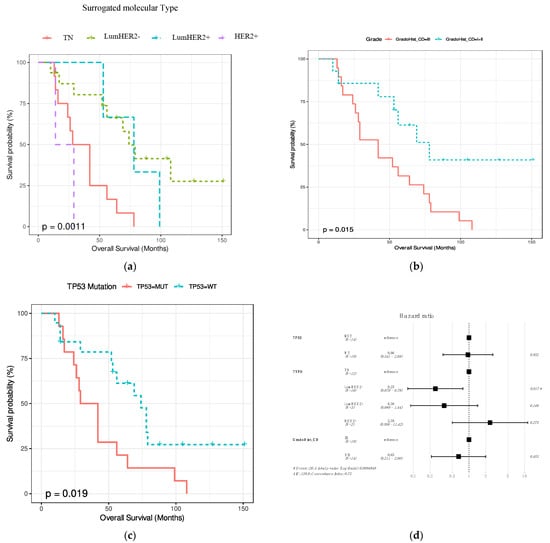
Figure 4.
Kaplan–Meyers graphs showing the association between overall survival and the surrogate molecular type (a), histological grade (b), and TP53 status (c). (d) Multivariate analysis showing the independent prognostic significance of the surrogate molecular type.
In contrast, the variables pT, pN, stage, permeation, age, histological type location (local or distant), neoadjuvant therapy, or PIK3CA mutations did not significantly affect overall survival.
4. Discussion
4.1. Clinicopathologic Features of Breast Carcinomas That Develop Cutaneous Relapses
In this study, the clinicopathological features were analyzed of 33 females with BC that developed cutaneous metastases, distantly in 53.1% and locally in 46.9% of the patients. Our observation that 46.9% of CM developed locally on the skin of the breast/thorax is in accordance with the review by Johnson et al. [24], who found in a study of 61 patients with CM, 57% showed metastases in the breast/thorax skin.
In the present series, CM developed at a median of 22.8 months after the initial diagnosis of the primary BC. CM usually appear at the end of the disease, late during cancer evolution. Thus, Lookingbill et al. [25] found that only 6.3% of patients with BC had cutaneous involvement at the time of diagnosis of the primary tumor. More recently, Johnson et al. [24] reported that approximately 13.7% of the patients in the 9 retrospective series reviewed [26,27,28,29,30,31,32,33,34] had a skin lesion before or simultaneously with the diagnosis of BC. Supporting these observations, we found that 18.2% of patient in the present series debuted with skin involvement.
The median age of women with BC who developed CM varies among series. Whereas the age in our series (63.5 years) was similar to the Johnson et al. [24] review of 41 patients, other studies have reported a mean of 74 years (n = 18) [9] and 48 years (n = 125) [15]. These differences may be related with the different number of patients included in each series or due to ethic/geographic differences, since these series originated from Italy, South Korea, and Spain.
In the present series, the histology of 24 cases (72.7%) corresponded to BCNST, and the rest to special histologic subtypes. Among them, the largest group corresponded to ILC (21.2%). However, considering only the 16 Luminal HER2- cases in our series, 43.7% were ILC. Since the frequency of ILC in Luminal HER2- BC is between 15% and 20% [35], our results suggested that ILC is a major risk factor for developing skin metastases. In accordance with our results, the study of Li et al. [36] that included only metastases from Luminal HER2- BC, reported that the proportion of ILC was 26% in the group of patients that developed CM.
We observed that the frequency of BC developing CM varied according to the subtype. Table 5 compares the data from our series with those from other published series [15,37,38,39], including our recently published CM review that includes samples (n = 58) of molecularly characterized CM [39,40,41,42,43].

Table 5.
Distribution of surrogated molecular types in breast cancer with cutaneous metastasis in different series.
Although there were differences between series, a finding common to all of them is the relative overrepresentation of TNBC, since its frequency in the general population of BC is around 15% [44], but it was between 23–39% in BC with CM, suggesting that this surrogate molecular type could be associated with a greater potential to metastasize to the skin.
In our series, 15.2% of BC that developed CM were HER2+, a frequency similar to the general population of BC, at least in Spain, suggesting that this surrogate molecular type does not have a special propensity to develop CM. However, Table 5 shows important differences in the frequency of HER2+ BC that develop CM among different series. These differences are probably due to differences in sample size, patient selection, methods of HER2 detection and the period of study.
4.2. Molecular Alterations Involved in the Development of Cutaneous Metastasis in Breast Cancer
In this series, we studied the mutational landscape of 33 matched primary tumors with their corresponding CM by NGS. In addition, changes in copy number of CCND1, FGFR1, MDM4 and MYC were analyzed by FISH in 20 paired samples. The molecular landscape of primary tumors in this series was concordant with many previous reports demonstrating different mutational patterns among different surrogate molecular types. Thus, whereas mutations in PIK3CA predominated among Luminal BC, TP53 mutation was the main molecular alteration in TNBC.
When primary tumors were compared with their respective metastasis, we found that 48.5% of CM exhibited additional pathogenic mutations and/or gene amplification in important oncogenes and/or tumor suppressor genes. Genes involved in the progression of more than one case in the present series included MYC, MDM4, PIK3CA and ERBB2. In spite of this high frequency of additional changes, we did not observe a specific mutational pattern related to tumor progression, indicating that CM is a very individual process in each tumor. These results add to the observations of three previous series analyzing paired primary BCs and their CMs including a total of only 15 patients [39] (Table 6).

Table 6.
Additional mutations in cutaneous metastases reported in different series.
There are several studies analyzing molecular alterations in metastatic BC including samples from CM, but without comparison with their primary tumors (see González- Martínez et al. [37]). In general, these studies also confirmed that there is no specific pattern of mutations that predispose to CM. Only Rinaldi et al. [39] observed that alterations in NOTCH1 were overrepresented in CM when compared to other metastatic locations.
We observed that the number of CM with additional molecular alterations was higher in distant than local metastases. Thus, additional mutations and CNVs were observed in 11 out of 17 (64.7%) distant CM but in only 5 out 15 (33.3%) local CM. These differences seemed to indicate a more advanced molecular stage of distant CM, although they did not have prognostic implications.
It is also worth noting that there was hardly any difference between the mutation frequency of TP53 and PIK3CA in CM samples of patients diagnosed at early and advanced stages, so we assume that treatment has no effect on the mutation pattern.
4.3. Therapeutic Implications
An important question in the study of cancer metastasis is whether or not the molecular characteristics of metastatic samples modify patient treatment. In 5 patients in this study (15.1%), there was a conversion of the surrogate molecular type between the primary tumor and its CM. In 3 cases (9%), there was a change from luminal to TN, from Luminal HER2+ to Luminal HER2- in one case (3%), and from Luminal HER2- to Luminal HER2+ in another case (3%). All these changes would have impacted on the treatment decision regarding the use of hormone therapy or anti-HER2 therapy.
Regarding specific mutations, three CM developed additional PIK3CA mutations, which is a target for treatment with alpelisib in RH+ metastatic BC. However, the three primary and metastases were TN. We detected an ESR1 mutation in a primary HR+ tumor and the corresponding CM. ESR1 mutations are the most frequent additional mutations that develop in metastatic HR+ BC after hormone therapy, especially after the use of aromatase inhibitors, being infrequent in primary tumors. In this case, ESR1 mutations would have influenced the response to hormone therapy during the complete evolution of the disease.
An interesting case in this series was the tumor and the CM developed in patient Pt13, which was an ILC that carried the pathogenic L755S mutation in the primary tumor. This mutation was absent in the CM, which carried the pathogenic mutation S310F. ERBB2 mutations are more frequent in ILC (6%) than in IDC (1.5%), especially in ILC with pleomorphic features, and are associated with a poor prognosis in ILC [45,46]. In addition, response to different anti-HER2 therapies differed among mutations. Thus, whereas L755S seems to be resistant to trastuzumab and lapatinib but sensitive to neratinib and afatinib, S310F seems to be sensitive to all drugs.
4.4. Prognosis
Patients with CM have a very poor prognosis. Lookingbill et al. [47] observed an average survival of 31 months after the diagnosis of the CM. Kong et al. [15] observed a median survival of 32 months. In our series, the median survival was 19.6 months, and this difference may be related to the clinical differences between the series. Additionally, patients who developed distant CM had a shorter overall survival and died more frequently during the first year after the diagnosis of CM. However, these differences between survival according to location were not statistically significant. In the univariate survival analysis, the only three variables that showed an impact on prognosis were the histological grade, the surrogate molecular type, and TP53 mutations. However, in the multivariate analysis, only the surrogate molecular type remained statistically significant, and the Luminal HER2- surrogate molecular type had a better prognosis.
4.5. Study Limitations
One limitation of this study is the relatively low number of cases studied. However, in spite of this and to the best of our knowledge, this is the largest series analyzing matched primary tumors and CMs reported so far. In addition, a comprehensive NGS panel of 61 genes was used in this study, including the most frequently mutated genes in BC. However, there may be additional genes with a role in progression in individual tumors that were not included in the panel. Moreover, the panel was not designed to detect CNVs or gene rearrangements, although this limitation was resolved in part by analyzing the genes most frequently amplified in BC by FISH.
Further research in this field would require studies with a larger number of patients and samples and perhaps focused on patients with BC of the TN surrogated molecular type, as this seems to be not only the most likely to metastasize to the skin but also has the poorest prognosis.
5. Conclusions
The development of new strategies for the management of CMs is a major clinical challenge because of the poor prognosis. To advance in this field, a better understanding of the molecular alterations involved in the metastatic process is needed. In the present study, the clinicopathological characteristics of BCs developing CM was analyzed and compared to the molecular differences between primary breast tumors and their corresponding CMs. We observed that the surrogate molecular type of BC with a greater risk to metastasize to skin was TN. A change of tumor surrogate molecular type in metastases with an impact on treatment was found in 15% of patients. In addition, half of the CM presented some additional molecular alterations with respect to the primary tumors, but a characteristic molecular pattern related to tumor progression and CM development was not observed. In this series, survival was related to the tumor surrogate molecular type. The immunohistochemical and molecular analysis of BC CM is essential for a proper treatment of the patients.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/2072-6694/14/5/1151/s1, Table S1: Mutations and primers used for Sanger verifications. Table S2: Clinicopathological features of the 33 patients; Table S3: List of somatic mutations found in the breast cancer and cutaneous metastases subjected to NGS analysis.
Author Contributions
Conceptualization, S.G.-M., J.C. and J.P.; methodology, S.G.-M., D.P., T.C.-C. and B.P-M.; validation, S.G.-M., B.P.-M. and J.P.; formal analysis, S.G.-M. and D.P.; investigation, S.G.-M., resources, J.C., J.L.R.-P. and J.P.; data curation, S.G.-M.; writing—original draft preparation, S.G.-M.; writing—review and editing, J.C., J.P., S.G.-M., D.P., B.P.-M., T.C.-C., J.L.R.-P., G.C., M.G. and A.C.; visualization, S.G.-M., J.C. and J.P.; supervision, J.C. and J.P.; funding acquisition, J.C. and J.P.; project administration, J.C. and J.P. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by grants from the Instituto de Salud Carlos III (ISCIII) (PI19/01331) and CIBERONC (CB16/12/00316 and CB16/12/00449), co-financed by the European Development Regional Fund ‘A way to achieve Europe’ (FEDER), and by the Spanish Association Against Cancer Scientific Foundation (Grupos Coordinados Traslacionales aecc 2018).
Institutional Review Board Statement
The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of Hospital Ramón y Cajal (Madrid, Spain) (protocol code 30-21 19/05/2021).
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Acknowledgments
“Contigo contra el Cáncer de la Mujer” Foundation.
Conflicts of Interest
J.C.: Consulting/Advisor: Roche, Celgene, Cellestia, AstraZeneca, Biothera Pharmaceutical, Merus, Seattle Genetics, Daiichi Sankyo, Erytech, Athenex, Polyphor, Lilly, Servier, Merck Sharp&Dohme, GSK, Leuko, Bioasis, Clovis Oncology, Boehringer Ingelheim, Kyowa Kirin. Honoraria: Roche, Novartis, Celgene, Eisai, Pfizer, Samsung Bioepis, Lilly, Merck Sharp&Dohme, Daiichi Sankyo. Research funding to the Institution: Roche, Ariad pharmaceuticals, AstraZeneca, Baxalta GMBH/Servier Affaires, Bayer healthcare, Eisai, F.Hoffman-La Roche, Guardanth health, Merck Sharp&Dohme, Pfizer, Piqur Therapeutics, Puma C, Queen Mary University of London. Stock, patents, and intellectual property: MedSIR. Travel, accommodation, expenses: Roche, Novartis, Eisai, Pfizer, Daiichi Sankyo. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Torre, L.A.; Islami, F.; Siegel, R.L.; Ward, E.M.; Jemal, A. Global Cancer in Women: Burden and Trends. Cancer Epidemiol. Biomark. Prev. 2017, 26, 444–457. [Google Scholar] [CrossRef]
- Tsang, J.Y.S.; Tse, G.M. Molecular Classification of Breast Cancer. Adv. Anat. Pathol. 2020, 27, 27–35. [Google Scholar] [CrossRef]
- Yam, C.; Mani, S.A.; Moulder, S.L. Targeting the Molecular Subtypes of Triple Negative Breast Cancer: Understanding the Diversity to Progress the Field. Oncologist 2017, 22, 1086–1093. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization Classification of Tumours Editorial Board; International Agency for Research on Cancer; World Health Organization. WHO Classification of Tumours. Breast Tumours; International Agency for Research on Cancer: Lyon, France, 2019; ISBN 978-92-832-4500-1. [Google Scholar]
- Scully, O.J.; Bay, B.-H.; Yip, G.; Yu, Y. Breast cancer metastasis. Cancer Genom. Proteom. 2012, 9, 311–320. [Google Scholar]
- Chiang, A.C.; Massagué, J. Molecular Basis of Metastasis. N. Engl. J. Med. 2008, 359, 2814–2823. [Google Scholar] [CrossRef]
- Liotta, L.A. Cancer invasion and metastases. JAMA J. Am. Med. Assoc. 1990, 263, 1123–1126. [Google Scholar] [CrossRef]
- Arozullah, A.M.; Calhoun, E.A.; Wolf, M.; Finley, D.K.; Fitzner, K.A.; Heckinger, E.A.; Gorby, N.S.; Schumock, G.T.; Bennett, C.L. The financial burden of cancer: Estimates from a study of insured women with breast cancer. J. Support. Oncol. 2004, 2, 271–278. [Google Scholar] [PubMed]
- Guanziroli, E.; Coggi, A.; Venegoni, L.; Fanoni, D.; Ercoli, G.; Boggio, F.; Veraldi, S.; Berti, E.; Gianotti, R.; Ferrero, S.; et al. Cutaneous metastases of internal malignancies: An experience from a single institution. Eur. J. Dermatol. 2017, 27, 609–614. [Google Scholar] [CrossRef]
- de Bittencourt, M.J.S.; Carvalho, A.H.; do Nascimento, B.A.M.; Freitas, L.K.M.; de Parijós, A.M. Cutaneous metastasis of a breast cancer diagnosed 13 years before. An. Bras. Dermatol. 2015, 90, 134–137. [Google Scholar] [CrossRef]
- De Giorgi, V.; Grazzini, M.; Alfaioli, B.; Savarese, I.; Corciova, S.A.; Guerriero, G.; Lotti, T. Cutaneous manifestations of breast carcinoma: A clinical guide. Dermatol. Ther. 2010, 23, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Alcaraz, I.; Cerroni, L.; Rütten, A.; Kutzner, H.; Requena, L. Cutaneous Metastases From Internal Malignancies: A Clinicopathologic and Immunohistochemical Review. Am. J. Dermatopathol. 2012, 34, 347–393. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.R. Cutaneous manifestations of breast cancer. Semin. Oncol. 2016, 43, 331–334. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.C.-S.; Chen, G.-S.; Wu, C.-S.; Chai, C.-Y.; Chen, W.-T.; Lan, C.-C.E. Rates of cutaneous metastases from different internal malignancies: Experience from a Taiwanese medical center. J. Am. Acad. Dermatol. 2009, 60, 379–387. [Google Scholar] [CrossRef]
- Kong, J.H.; Park, Y.H.; Kim, J.A.; Kim, J.H.; Yun, J.; Sun, J.M.; Won, Y.W.; Lee, S.; Kim, S.T.; Cho, E.Y.; et al. Patterns of Skin and Soft Tissue Metastases from Breast Cancer according to Subtypes: Relationship between EGFR Overexpression and Skin Manifestations. Oncology 2011, 81, 55–62. [Google Scholar] [CrossRef]
- Krathen, R.A.; Orengo, I.F.; Rosen, T. Cutaneous Metastasis: A Meta-Analysis of Data. South Med. J. 2003, 96, 164–167. [Google Scholar] [CrossRef]
- Brownstein, M.H.; Helwig, E.B. Metastatic tumors of the skin. Cancer 1972, 29, 1298–1307. [Google Scholar] [CrossRef]
- Moore, S. Cutaneous Metastatic Breast Cancer. Clin. J. Oncol. Nurs. 2002, 6, 255–260. [Google Scholar] [CrossRef]
- Marcoval, J.; Gallego, M.I.; Moreno, A. Inflammatory Cutaneous Metastasis as a First Sign of Recurrence of Squamous Cell Carcinoma of the Lung. Actas Dermo-Sifiliográficas Engl. Ed. 2008, 99, 157–159. [Google Scholar] [CrossRef]
- Lin, L.; Sirohi, D.; Coleman, J.F.; Gulbahce, H.E. American Society of Clinical Oncology/College of American Pathologists 2018 Focused Update of Breast Cancer HER2 FISH Testing GuidelinesResults From a National Reference Laboratory. Am. J. Clin. Pathol. 2019, 152, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Koboldt, D.C.; Zhang, Q.; Larson, D.E.; Shen, D.; McLellan, M.D.; Lin, L.; Miller, C.A.; Mardis, E.R.; Ding, L.; Wilson, R.K. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012, 22, 568–576. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.C.; Reis-Filho, J.S. Tackling the Diversity of Triple-Negative Breast Cancer. Clin. Cancer Res. 2013, 19, 6380–6388. [Google Scholar] [CrossRef]
- Johnson, C. Cutaneous Manifestation as Initial Presentation of Metastatic Breast Cancer: A Systematic Review. Cutis 2021, 107, E29–E36. [Google Scholar] [CrossRef] [PubMed]
- Lookingbill, D.P.; Spangler, N.; Sexton, F.M. Skin involvement as the presenting sign of internal carcinoma. A retrospective study of 7316 cancer patients. J. Am. Acad. Dermatol. 1990, 22, 19–26. [Google Scholar] [CrossRef]
- Atış, G.; Tükenmez Demirci, G.; Kıvanç Atunay, İ.; Sakız, D. Derinin metastatik tümörlerinin primer deri tümörleri arasındaki sıklığı ve klinik özellikleri. Turkderm 2013, 47, 166–169. [Google Scholar] [CrossRef][Green Version]
- Bansal, R.; Patel, T.; Sarin, J.; Parikh, B.; Ohri, A.; Trivedi, P. Cutaneous and subcutaneous metastases from internal malignancies: An analysis of cases diagnosed by fine needle aspiration. Diagn. Cytopathol. 2011, 39, 882–887. [Google Scholar] [CrossRef]
- Benmously, R.; Souissi, A.; Badri, T.; Ben Jannet, S.; Marrak, H.; Mokhtar, I.; Fenniche, S. Cutaneous metastases from internal cancers. Acta Dermatovenerol. Alp. Pannonica Adriat. 2008, 17, 167–170. [Google Scholar] [PubMed]
- Chopra, R.; Chhabra, S.; Samra, S.G.; Thami, G.P.; Punia, R.P.S.; Mohan, H. Cutaneous metastases of internal malignancies: A clinicopathologic study. Indian J. Dermatol. Venereol. Leprol. 2010, 76, 125–131. [Google Scholar] [CrossRef]
- El Khoury, J.; Khalifeh, I.; Kibbi, A.-G.; Abbas, O. Cutaneous metastasis: Clinicopathological study of 72 patients from a tertiary care center in Lebanon. Int. J. Dermatol. 2014, 53, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Flores, A. Cutaneous metastases: A study of 78 biopsies from 69 patients. Am. J. Dermatopathol. 2010, 32, 222–239. [Google Scholar] [CrossRef]
- Gómez Sánchez, M.E.; Martinez Martinez, M.L.; Martín De Hijas, M.C.; López Villaescusa, M.T.; Faura Berruga, C.; Rodríguez Vázquez, M.; Pérez García, L.J. Metástasis cutáneas de tumores sólidos. Estudio descriptivo retrospectivo. Piel 2014, 29, 207–212. [Google Scholar] [CrossRef]
- Handa, U.; Kundu, R.; Dimri, K. Cutaneous Metastasis: A Study of 138 Cases Diagnosed by Fine-Needle Aspiration Cytology. Acta Cytol. 2017, 61, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Itin, P.; Tomaschett, S. Hautmetastasen bei viszeralen Malignomen: Eine epidemiologische Studie. Internist 2009, 50, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Razavi, P.; Chang, M.T.; Xu, G.; Bandlamudi, C.; Ross, D.S.; Vasan, N.; Cai, Y.; Bielski, C.M.; Donoghue, M.T.A.; Jonsson, P.; et al. The Genomic Landscape of Endocrine-Resistant Advanced Breast Cancers. Cancer Cell 2018, 34, 427–438.e6. [Google Scholar] [CrossRef]
- Li, Q.; Jiang, B.; Guo, J.; Shao, H.; Del Priore, I.S.; Chang, Q.; Kudo, R.; Li, Z.; Razavi, P.; Liu, B.; et al. INK4 tumor suppressor proteins mediate resistance to CDK4/6 kinase inhibitors. Cancer Discov. 2021, 12, 356–371. [Google Scholar] [CrossRef] [PubMed]
- González-Martínez, S.; Pizarro, D.; Pérez-Mies, B.; Caniego-Casas, T.; Curigliano, G.; Cortés, J.; Palacios, J. Clinical, Pathological, and Molecular Features of Breast Carcinoma Cutaneous Metastasis. Cancers 2021, 13, 5416. [Google Scholar] [CrossRef]
- Luna, A.; Rabassa, M.E.; Isla Larrain, M.; Cabaleiro, P.; Zwenger, A.; Canzoneri, R.; Segal-Eiras, A.; Abba, M.C.; Croce, M.V. Breast cancer cutaneous metastases are associated to uMUC1 and sialyl Lewis x and to highly malignant primary tumors. Pathol.-Res. Pract. 2020, 216, 152859. [Google Scholar] [CrossRef]
- Yates, L.R.; Knappskog, S.; Wedge, D.; Farmery, J.H.R.; Gonzalez, S.; Martincorena, I.; Alexandrov, L.B.; Van Loo, P.; Haugland, H.K.; Lilleng, P.K.; et al. Genomic Evolution of Breast Cancer Metastasis and Relapse. Cancer Cell 2017, 32, 169–184.e7. [Google Scholar] [CrossRef]
- Lefebvre, C.; Bachelot, T.; Filleron, T.; Pedrero, M.; Campone, M.; Soria, J.-C.; Massard, C.; Lévy, C.; Arnedos, M.; Lacroix-Triki, M.; et al. Mutational Profile of Metastatic Breast Cancers: A Retrospective Analysis. PLoS Med. 2016, 13, e1002201. [Google Scholar] [CrossRef]
- Schrijver, W.A.; Selenica, P.; Lee, J.Y.; Ng, C.K.Y.; Burke, K.; Piscuoglio, S.; Berman, S.H.; Reis-Filho, J.S.; Weigelt, B.; Van Diest, P.J.; et al. Mutation profiling of key cancer genes in primary breast cancers and their distant metastases. Cancer Res. 2018, 78, 3112–3121. [Google Scholar] [CrossRef]
- Paul, M.R.; Pant, D.K.; Shih, N.N.C.; Chen, Y.; Harvey, K.L.; Solomon, A.; Lieberman, D.; Morrissette, J.J.D.; Soucier-Ernst, D.; Goodman, N.G.; et al. Genomic landscape of metastatic breast cancer identifies preferentially dysregulated pathways and targets. J. Clin. Investig. 2020, 130, 4252–4265. [Google Scholar] [CrossRef]
- Muller, K.E.; Marotti, J.D.; de Abreu, F.B.; Peterson, J.D.; Miller, T.W.; Chamberlin, M.D.; Tsongalis, G.J.; Tafe, L.J. Targeted next-generation sequencing detects a high frequency of potentially actionable mutations in metastatic breast cancers. Exp. Mol. Pathol. 2016, 100, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.J.; Naidu, S.; Topham, A.K.; Guiles, F.; Xu, Y.; McCue, P.; Schwartz, G.F.; Park, P.K.; Rosenberg, A.L.; Brill, K.; et al. Differences in breast carcinoma characteristics in newly diagnosed African-American and Caucasian patients: A single-institution compilation compared with the National Cancer Institute’s Surveillance, Epidemiology, and End Results database. Cancer 2007, 110, 876–884. [Google Scholar] [CrossRef]
- Kurozumi, S.; Alsaleem, M.; Monteiro, C.J.; Bhardwaj, K.; Joosten, S.E.P.; Fujii, T.; Shirabe, K.; Green, A.R.; Ellis, I.O.; Rakha, E.A.; et al. Targetable ERBB2 mutation status is an independent marker of adverse prognosis in estrogen receptor positive, ERBB2 non-amplified primary lobular breast carcinoma: A retrospective in silico analysis of public datasets. Breast Cancer Res. 2020, 22, 85. [Google Scholar] [CrossRef] [PubMed]
- Rosa-Rosa, J.; Caniego-Casas, T.; Leskela, S.; Cristobal, E.; González-Martínez, S.; Moreno-Moreno, E.; López-Miranda, E.; Holgado, E.; Pérez-Mies, B.; Garrido, P.; et al. High Frequency of ERBB2 Activating Mutations in Invasive Lobular Breast Carcinoma with Pleomorphic Features. Cancers 2019, 11, 74. [Google Scholar] [CrossRef]
- Lookingbill, D.P.; Spangler, N.; Helm, K.F. Cutaneous metastases in patients with metastatic carcinoma: A retrospective study of 4020 patients. J. Am. Acad. Dermatol. 1993, 29, 228–236. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).